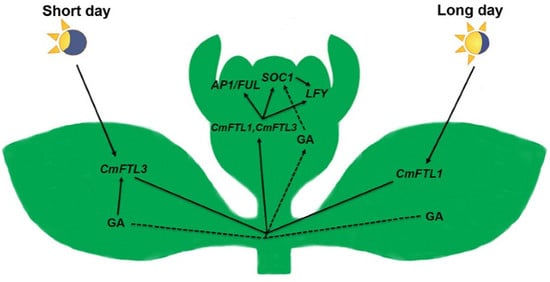
Gibberellic Acid Signaling Is Required to Induce Flowering of Chrysanthemums Grown under Both Short and Long Days
Abstract
:1. Introduction
2. Results
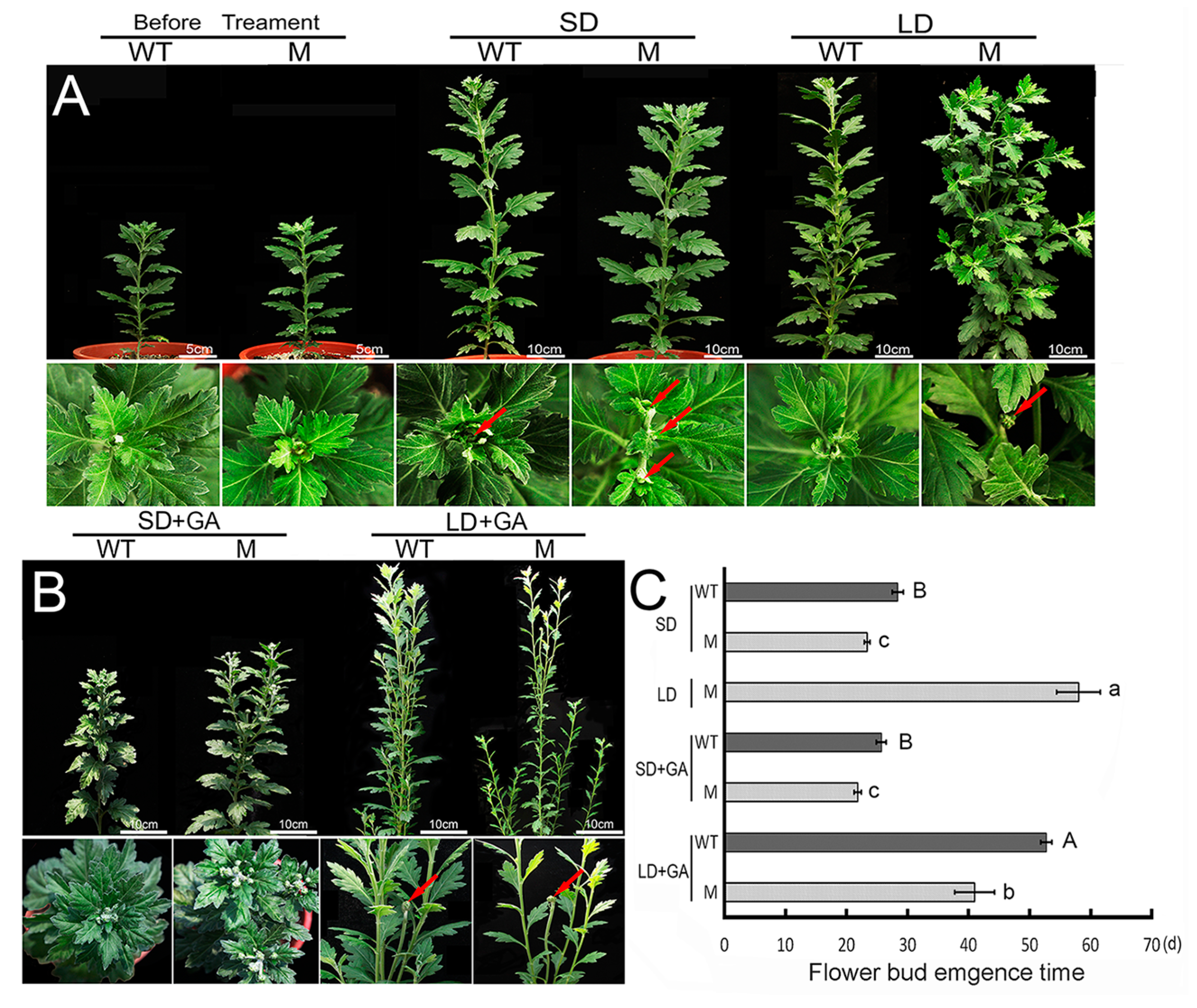
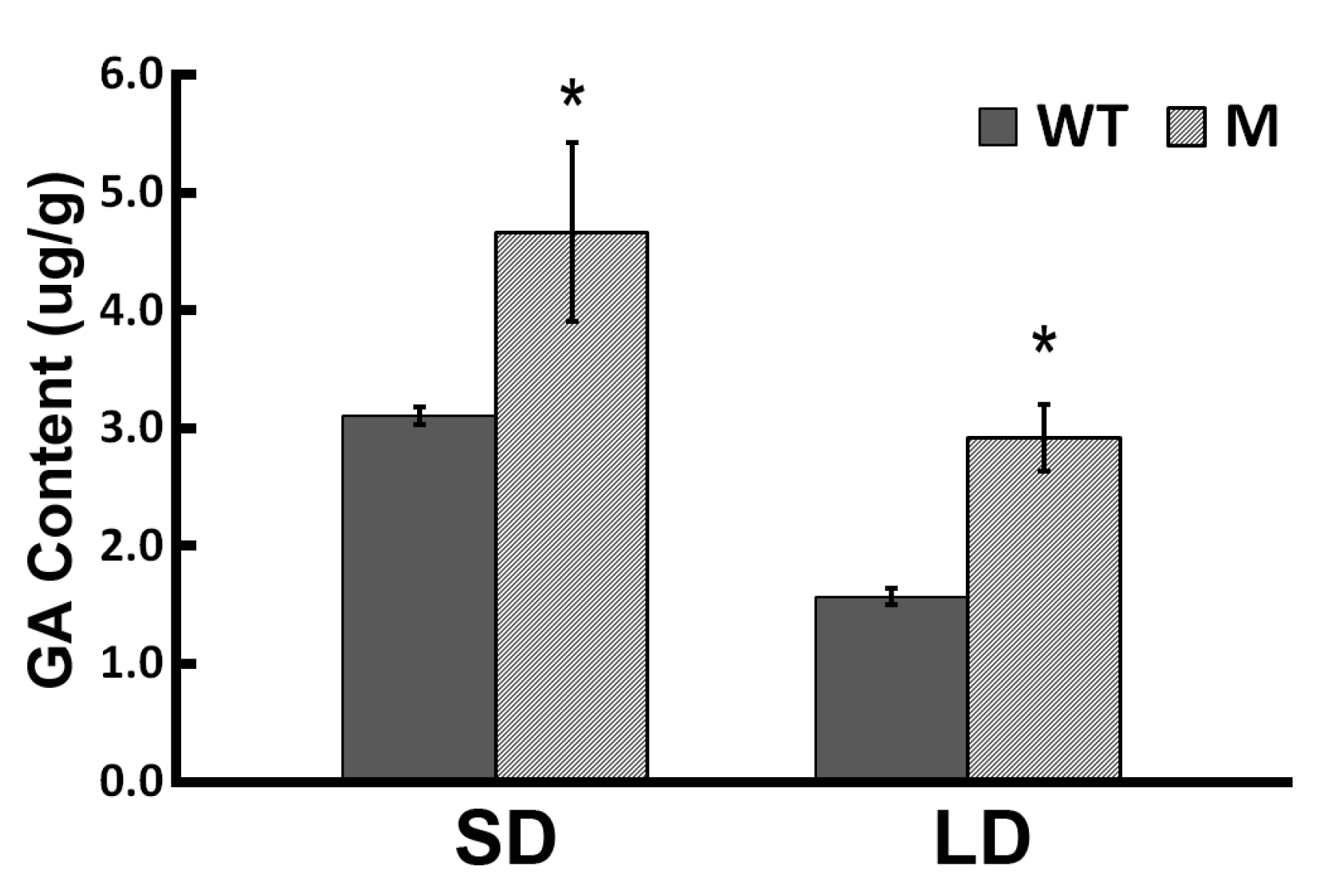
2.1. Flowering Behavior of the ‘Jinba’ WT and Mutant in Response to GA Treatment
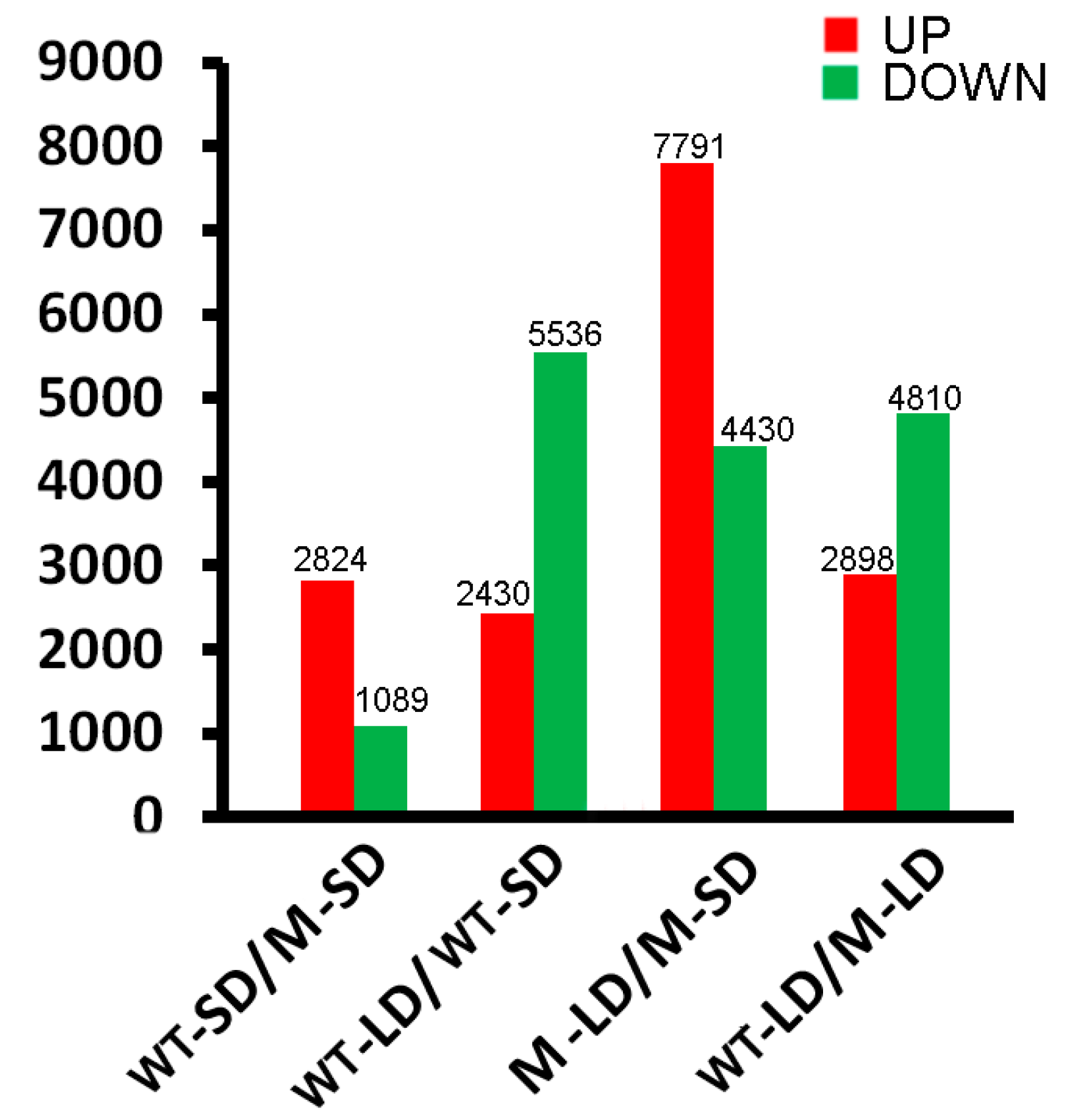
2.2. Transcriptome Sequencing and Bioinformatic Analysis
2.3. The Transcription Behavior of Genes Involved in the Determination of Flowering Time
2.4. Differentially Transcribed Transcription Factors
2.5. qRT-PCR-Based Validation of RNA-Seq Acquired Transcript Abundance
3. Discussion
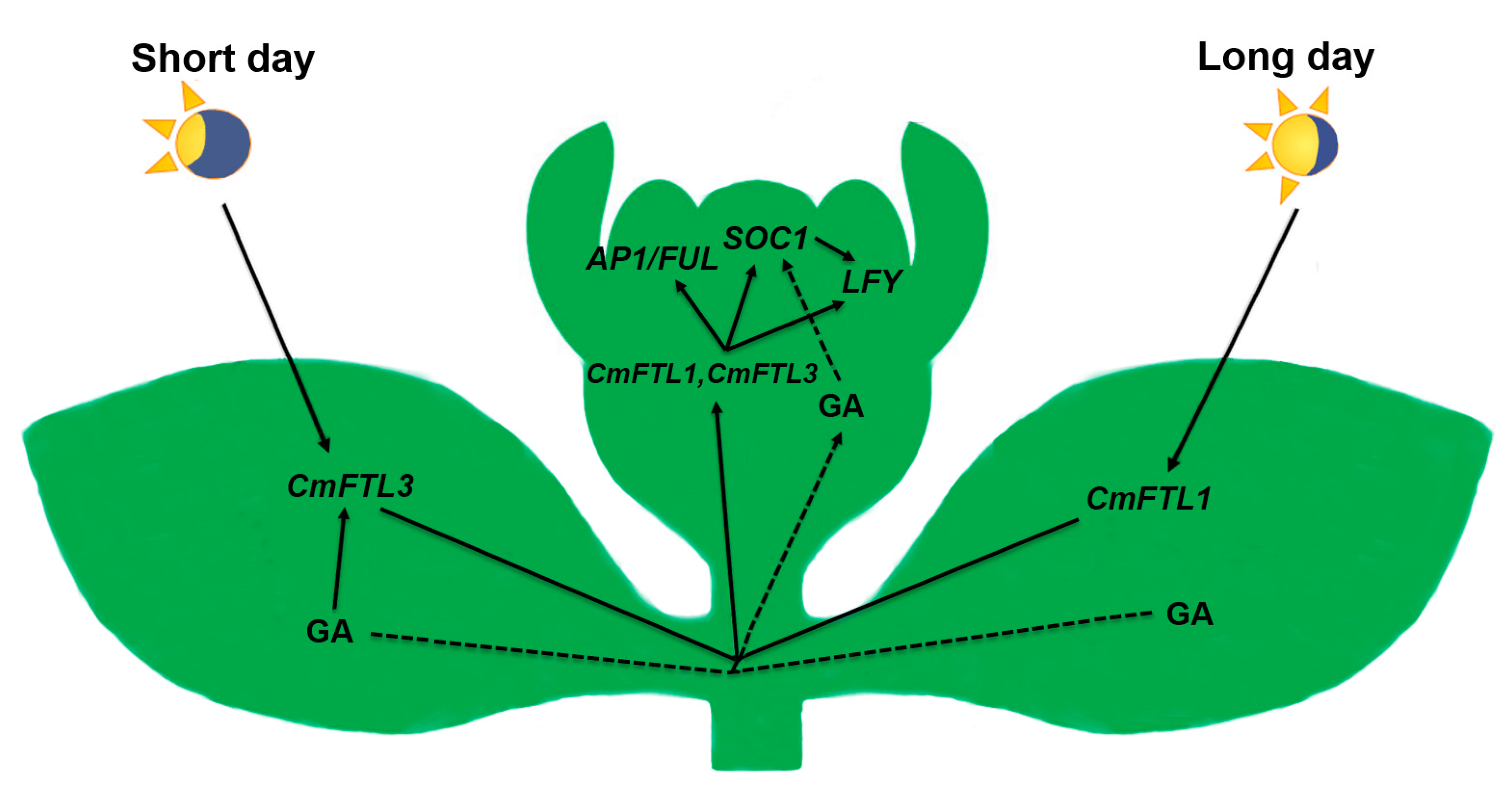
3.1. The Photoperiod and GA Pathways Act in Concert to Regulate Flowering Time under SD
3.2. GA Activates SOC1 to Promote Flowering under LD
3.3. Transcription Factors Involved in Floral Induction in Chrysanthemum
4. Materials and Methods
4.1. Plant Materials and Growth Conditions
4.2. RNA Extraction and cDNA Library Construction
4.3. Transcriptome Sequencing and Bioinformatic Analysis
4.4. qRT-PCR
4.5. Determination of Endogenous GA Content
4.6. GA Treatment
5. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Muller, C.J.A.E. Flowering time control and applications in plant breeding. Trends Plant Sci. 2009, 14, 563–573. [Google Scholar]
- Fornara, F.; de Montaigu, A.; Coupland, G. SnapShot: Control of flowering in Arabidopsis. Cell 2010, 141, 550–550.e2. [Google Scholar] [CrossRef] [PubMed]
- Amasino, R. Seasonal and developmental timing of flowering. Plant J 2010, 61, 1001–1013. [Google Scholar] [CrossRef] [PubMed]
- Andres, F.; Coupland, G. The genetic basis of flowering responses to seasonal cues. Nat. Rev. Genet. 2012, 13, 627–639. [Google Scholar] [CrossRef] [PubMed]
- Song, Y.H.; Shim, J.S.; Kinmonth-Schultz, H.A.; Imaizumi, T. Photoperiodic flowering: Time measurement mechanisms in leaves. Annu. Rev. Plant Biol. 2015, 66, 441–464. [Google Scholar] [CrossRef] [PubMed]
- Miwa, K.; Serikawa, M.; Suzuki, S.; Kondo, T.; Oyama, T. Conserved expression profiles of circadian clock-related genes in two Lemna species showing long-day and short-day photoperiodic flowering responses. Plant Cell Physiol. 2006, 47, 601–612. [Google Scholar] [CrossRef] [PubMed]
- Shim, J.S.; Imaizumi, T. Circadian clock and photoperiodic response in Arabidopsis: From seasonal flowering to redox homeostasis. Biochemistry 2015, 54, 157–170. [Google Scholar] [CrossRef] [PubMed]
- Srikanth, A.; Schmid, M. Regulation of flowering time: All roads lead to Rome. Cell Mol. Life Sci. 2011, 68, 2013–2037. [Google Scholar] [CrossRef] [PubMed]
- Song, Y.H.; Ito, S.; Imaizumi, T. Flowering time regulation: Photoperiod- and temperature-sensing in leaves. Trends Plant Sci. 2013, 18, 575–583. [Google Scholar] [CrossRef] [PubMed]
- Olszewski, N.; Sun, T.; Gubler, F. Gibberellin signaling biosynthesis, catabolism, and response pathways. Plant Cell 2002, 14 (Suppl. S1), S61–S80. [Google Scholar]
- Hisamatsua, T.; Koshioka, M.; Kubotaa, S.; Fujimeb, Y.; King, R.W.; Mander, L.N. The role of gibberellin biosynthesis in the control of growth and flowering in Matthiola incana. Physiol. Plant 2000, 109, 97–105. [Google Scholar] [CrossRef]
- Mutasa-Gottgens, E.; Hedden, P. Gibberellin as a factor in floral regulatory networks. J. Exp. Bot. 2009, 60, 1979–1989. [Google Scholar] [CrossRef] [PubMed]
- Davière, J.-M.; Achard, P. Gibberellin signaling in plants. Development 2013, 140, 1147–1151. [Google Scholar]
- Moon, J.; Suh, S.S.; Lee, H.; Choi, K.R.; Hong, C.B.; Paek, N.C.; Kim, S.G.; Lee, I. The SOC1 MADS-box gene integrates vernalization and gibberellin signals for flowering in Arabidopsis. Plant J. 2003, 35, 613–623. [Google Scholar] [CrossRef] [PubMed]
- Hisamatsu, T.; King, R.W. The nature of floral signals in Arabidopsis. II. Roles for FLOWERING LOCUS T (FT) and gibberellin. J. Exp. Bot. 2008, 59, 3821–3829. [Google Scholar] [CrossRef] [PubMed]
- Itoh, H.; Matsuoka, M.; Steber, C.M. A role for the ubiquitin–26S-proteasome pathway in gibberellin signaling. Trends Plant Sci. 2003, 8, 492–497. [Google Scholar] [CrossRef] [PubMed]
- Fleet, C.M.; Sun, T.-p. A DELLAcate balance: The role of gibberellin in plant morphogenesis. Curr. Opin. Plant Biol. 2005, 8, 77–85. [Google Scholar] [CrossRef] [PubMed]
- Du, H.; Wang, Y.-B.; Xie, Y.; Liang, Z.; Jiang, S.-J.; Zhang, S.-S.; Huang, Y.-B.; Tang, Y.-X. Genome-Wide identification and evolutionary and expression analyses of MYB-related genes in land plants. DNA Res. 2013, 20, 437–448. [Google Scholar] [CrossRef] [PubMed]
- Ito, S.; Song, Y.H.; Josephson-Day, A.R.; Miller, R.J.; Breton, G.; Olmstead, R.G.; Imaizumi, T. FLOWERING BHLH transcriptional activators control expression of the photoperiodic flowering regulator CONSTANS in Arabidopsis. Proc. Natl. Acad. Sci. USA 2012, 109, 3582–3587. [Google Scholar] [CrossRef] [PubMed]
- Smaczniak, C.; Immink, R.G.; Muiño, J.M.; Blanvillain, R.; Busscher, M.; Busscher-Lange, J.; Dinh, Q.P.; Liu, S.; Westphal, A.H.; Boeren, S. Characterization of MADS-domain transcription factor complexes in Arabidopsis flower development. Proc. Natl. Acad. Sci. USA 2012, 109, 1560–1565. [Google Scholar] [CrossRef] [PubMed]
- Yu, Y.; Liu, Z.; Wang, L.; Kim, S.G.; Seo, P.J.; Qiao, M.; Wang, N.; Li, S.; Cao, X.; Park, C.M. WRKY71 accelerates flowering via the direct activation of FLOWERING LOCUS T and LEAFY in Arabidopsis thaliana. Plant J. 2016, 85, 96–106. [Google Scholar] [CrossRef] [PubMed]
- Navarro, C.; Cruz-Oró, E.; Prat, S. Conserved function of FLOWERING LOCUS T (FT) homologues as signals for storage organ differentiation. Curr. Opin. Plant Biol. 2015, 23, 45–53. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.H.; Yoo, S.J.; Park, S.H.; Hwang, I.; Lee, J.S.; Ahn, J.H. Role of SVP in the control of flowering time by ambient temperature in Arabidopsis. Genes Dev. 2007, 21, 397–402. [Google Scholar] [CrossRef] [PubMed]
- Moura, F.B.; Vieira, M.R.d.S.; Evangelista, W.S.; Silva, L.F.; Oliveira, R.H.; Rocha, A.T.; Costa, F.B. Cultivars, conduction, photoperiodic and quality chrysanthemum in Brazil. J. Hortic. For. 2014, 6, 53–57. [Google Scholar] [CrossRef]
- Oda, A.; Narumi, T.; Li, T.; Kando, T.; Higuchi, Y.; Sumitomo, K.; Fukai, S.; Hisamatsu, T. CsFTL3, a chrysanthemum FLOWERING LOCUS T-like gene, is a key regulator of photoperiodic flowering in chrysanthemums. J. Exp. Bot. 2012, 63, 1461–1477. [Google Scholar] [CrossRef] [PubMed]
- Achard, P.; Genschik, P. Releasing the brakes of plant growth: How GAs shutdown DELLA proteins. J. Exp. Bot. 2009, 60, 1085–1092. [Google Scholar] [CrossRef] [PubMed]
- Croker, S.J.; García-Lepe, R.; Lewis, M.J.; Hedden, P. Modification of gibberellin production and plant development in Arabidopsis by sense and antisense expression of gibberellin 20-oxidase genes. Plant J. 1999, 17, 547–556. [Google Scholar]
- Mouradov, A.; Cremer, F.; Coupland, G. Control of flowering time interacting pathways as a basis for diversity. Plant Cell 2002, 14 (Suppl. S1), S111–S130. [Google Scholar] [PubMed]
- Yang, Y.; Ma, C.; Xu, Y.; Wei, Q.; Imtiaz, M.; Lan, H.; Gao, S.; Cheng, L.; Wang, M.; Fei, Z.; et al. A zinc finger protein regulates flowering time and abiotic stress tolerance in chrysanthemum by modulating gibberellin biosynthesis. Plant Cell 2014, 26, 2038–2054. [Google Scholar] [CrossRef] [PubMed]
- Udvardi, M.K.; Kakar, K.; Wandrey, M.; Montanari, O.; Murray, J.; Andriankaja, A.; Zhang, J.-Y.; Benedito, V.; Hofer, J.M.; Chueng, F. Legume transcription factors: Global regulators of plant development and response to the environment. Plant Physiol. 2007, 144, 538–549. [Google Scholar] [CrossRef] [PubMed]
- Shan, H.; Chen, S.; Jiang, J.; Chen, F.; Chen, Y.; Gu, C.; Li, P.; Song, A.; Zhu, X.; Gao, H. Heterologous expression of the chrysanthemum R2R3-MYB transcription factor CmMYB2 enhances drought and salinity tolerance, increases hypersensitivity to ABA and delays flowering in Arabidopsis thaliana. Mol. Biotechnol. 2012, 51, 160–173. [Google Scholar] [CrossRef] [PubMed]
- Litt, A.; Irish, V.F. Duplication and diversification in the APETALA1/FRUITFULL floral homeotic gene lineage: Implications for the evolution of floral development. Genetics 2003, 165, 821–833. [Google Scholar] [PubMed]
- Shchennikova, A.V.; Shulga, O.A.; Immink, R.; Skryabin, K.G.; Angenent, G.C. Identification and characterization of four chrysanthemum MADS-box genes, belonging to the APETALA1/FRUITFULL and SEPALLATA3 subfamilies. Plant Physiol. 2004, 134, 1632–1641. [Google Scholar] [CrossRef] [PubMed]
- Nakano, T.; Suzuki, K.; Fujimura, T.; Shinshi, H. Genome-Wide analysis of the ERF gene family in Arabidopsis and rice. Plant Physiol. 2006, 140, 411–432. [Google Scholar] [CrossRef] [PubMed]
- Mizoi, J.; Shinozaki, K.; Yamaguchi-Shinozaki, K. AP2/ERF family transcription factors in plant abiotic stress responses. Biochim. Biophys. Acta 2012, 1819, 86–96. [Google Scholar] [CrossRef] [PubMed]
- Singh, V.K.; Garg, R.; Jain, M. A global view of transcriptome dynamics during flower development in chickpea by deep sequencing. Plant Biotechnol. J. 2013, 11, 691–701. [Google Scholar] [CrossRef] [PubMed]
- Ren, L.; Sun, J.; Chen, S.; Gao, J.; Dong, B.; Liu, Y.; Xia, X.; Wang, Y.; Liao, Y.; Teng, N. A transcriptomic analysis of Chrysanthemum nankingense provides insights into the basis of low temperature tolerance. BMC Genom. 2014, 15, 1. [Google Scholar] [CrossRef] [PubMed]
- Audic, S.; Claverie, J.M. The significance of digital gene expression profiles. Genome Res. 1997, 7, 986–995. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Chen, S.; Jiang, J.; Zhang, F.; Chen, F. Reference gene selection for cross-species and cross-ploidy level comparisons in Chrysanthemum spp. Sci. Rep. 2015, 5, 8094. [Google Scholar] [CrossRef] [PubMed]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−∆∆Ct method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]

| Samples | Total Raw Reads | Total Clean Reads | Total Clean Nucleotides (nt) | Q20 Percentage | Q30 Percentage | N Percentage | GC Percentage |
|---|---|---|---|---|---|---|---|
| WT-SD | 117,328,048 | 114,265,744 | 5,141,958,480 | 97.30% | 91.39% | 0.00 | 43.19% |
| WT-LD | 117,809,964 | 114,693,344 | 5,161,200,480 | 97.19% | 91.16% | 0.00 | 43.59% |
| M-SD | 121,981,564 | 118,657,316 | 5,339,579,220 | 97.16% | 91.08% | 0.00 | 43.18% |
| M-LD | 120,340,980 | 116,781,172 | 5,255,152,740 | 97.19% | 91.18% | 0.00 | 43.53% |
| Gene | Annotation | WT-SD RPKM | WT-LD RPKM | M-SD RPKM | M-LD RPKM | Fold Change | |||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| M-SD/WT-SD | p Value | WT-SD/WT-LD | p Value | M-SD/M-LD | p Value | M-LD/WT-LD | p Value | ||||||
| Photoperiod pathway | |||||||||||||
| CL80.Contig1l | CmTOC1 | 2.64 | 1.30 | 3.51 | 1.50 | 1.33 | 4.0 × 10−2 | 2.04 | 1.0 × 10−4 | 2.36 | 2.0 × 10−7 | 1.15 | 5.2 × 10−1 |
| CL6838.Contig1 | PRR5 | 36.32 | 33.30 | 52.36 | 26.07 | 1.44 | 1.8 × 10−45 | 1.09 | 3.0 × 10−3 | 2.01 | 4.0 × 10−133 | 0.78 | 3.0 × 10−14 |
| CL64.Contig1 | LHY | 3.25 | 2.35 | 4.20 | 1.65 | 1.29 | 2.0 × 10−2 | 1.38 | 1.0 × 10−2 | 2.54 | 2.0 × 10−12 | 0.70 | 2.0 × 10−2 |
| CL64.Contig2 | LHY | 4.57 | 3.29 | 6.34 | 2.30 | 1.39 | 4.0 × 10−4 | 1.39 | 3.0 × 10−3 | 2.76 | 7.0 × 10−20 | 0.70 | 7.0 × 10−2 |
| Unigene30909 | LHY | 3.74 | 1.97 | 5.06 | 2.43 | 1.35 | 3.0 × 10−3 | 1.90 | 7.0 × 10−7 | 2.09 | 8.0 × 10−11 | 1.23 | 1.5 × 10−1 |
| CL7048.Contig2 | CONSTANS-like | 10.17 | 9.32 | 14.99 | 6.42 | 1.47 | 2.3 × 10−9 | 1.09 | 1.4 × 10−1 | 2.33 | 2.0 × 10−36 | 0.69 | 2.0 × 10−7 |
| CL10258.Contig3 | CONSTANS-like | 41.44 | 23.72 | 54.60 | 13.61 | 1.38 | 6.8 × 10−19 | 1.68 | 1.0 × 10−29 | 3.74 | 2.0 × 10−170 | 0.62 | 9.0 × 10−16 |
| Unigene16657 | CONSTANS-like | 93.59 | 77.41 | 100.21 | 43.79 | 1.07 | 5.0 × 10−4 | 1.21 | 4.0 × 10−19 | 2.29 | 1.0 × 10−260 | 0.57 | 3.5 × 10−107 |
| GA biosynthesis and signaling | |||||||||||||
| CL10783.Contig2 | Ent-copalyl diphosphate synthase (CPS) | 0.09 | 0.79 | 2.06 | 0.28 | 22.89 | 2.0 × 10−6 | 0.11 | 1.5 × 10−2 | 7.36 | 9.5 × 10−5 | 0.35 | 1.3 × 10−1 |
| CL2973.Contig1 | Gibberellin 20-oxidase (GA20ox) | 38.62 | 7.74 | 30.22 | 18.62 | 0.78 | 6.0 × 10−14 | 4.99 | 6.0 × 10−261 | 1.62 | 2.0 × 10−34 | 2.41 | 1.0 × 10−54 |
| CL8331.Contig2 | Gibberellin 2-oxidase (GA2ox) | 0.14 | 3.94 | 0.20 | 0.92 | 1.38 | 5.6 × 10−1 | 0.04 | 2.0 × 10−40 | 0.21 | 8.0 × 10−6 | 0.23 | 7.0 × 10−19 |
| Unigene27395 | Gibberellin receptor (GID1) | 3.36 | 3.19 | 2.36 | 8.18 | 0.70 | 1.2 × 10−2 | 1.05 | 7.1 × 10−1 | 0.29 | 5.0 × 10−28 | 2.56 | 1.0 × 10−18 |
| Unigene27748 | GAI | 14.43 | 45.74 | 12.22 | 23.59 | 0.85 | 1.0 × 10−3 | 0.32 | 2.0 × 10−214 | 0.52 | 2.0 × 10−47 | 0.52 | 6.0 × 10−89 |
| Flowering integrators | |||||||||||||
| CL1484.Contig1 | CmFTL1 | 0.39 | 12.17 | 0.41 | 12.12 | 0.93 | 8.9 × 10−1 | 0.03 | 6.0 × 10−72 | 0.03 | 3.0 × 10−72 | 1.0 | 9.7 × 10−1 |
| CL1484.Contig2 | CmFTL3 | 16.88 | 1.68 | 31.81 | 2.08 | 1.88 | 6.0 × 10−51 | 3.94 | 5.0 × 10−148 | 15.32 | 0 | 1.23 | 1.69 × 10−1 |
| Unigene40592 | SOC1 | 8.68 | 3.60 | 17.65 | 16.82 | 2.03 | 2.0 × 10−21 | 2.41 | 9.0 × 10−15 | 1.05 | 4.5 × 10−1 | 4.67 | 5.0 × 10−57 |
| Unigene25380 | FLO/LFY-like | 0.75 | 1.28 | 0.57 | 8.50 | 0.76 | 3.3 × 10−1 | 0.58 | 2.0 × 10−2 | 0.07 | 1.0 × 10−71 | 6.66 | 2.0 × 10−50 |
| Unigene23898 | AP1/FUL like | 47.87 | 1.54 | 76.84 | 2.03 | 1.61 | 1.0 × 10−56 | 31.12 | 0 | 37.89 | 0 | 1.32 | 1.3 × 10−1 |
| Aging, ambient temperature and autonomous pathway | |||||||||||||
| Unigene29044 | SPL5 | 1.30 | 9.51 | 0.40 | 7.11 | 0.33 | 2.0 × 10−3 | 0.14 | 2.0 × 10−33 | 0.06 | 1.7 × 10−33 | 0.75 | 8.0 × 10−3 |
| CL14613.Contig14 | FLC-like | 1.01 | 1.59 | 2.96 | 1.60 | 2.93 | 1.0 × 10−7 | 0.63 | 6.0 × 10−2 | 1.85 | 8.0 × 10−4 | 1.00 | 9.9 × 10−1 |
| CL8773.Contig1 | FRIGIDA | 1.03 | 0.89 | 1.70 | 0.63 | 1.92 | 4.0 × 10−5 | 0.86 | 4.0 × 10−1 | 2.70 | 1.3 × 10−8 | 0.61 | 1.0 × 10−2 |
| Unigene20776 | SVP | 42.08 | 308.77 | 23.50 | 167.50 | 0.56 | 2.0 × 10−45 | 0.14 | 0 | 0.14 | 0 | 0.54 | 0 |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Dong, B.; Deng, Y.; Wang, H.; Gao, R.; Stephen, G.K.; Chen, S.; Jiang, J.; Chen, F. Gibberellic Acid Signaling Is Required to Induce Flowering of Chrysanthemums Grown under Both Short and Long Days. Int. J. Mol. Sci. 2017, 18, 1259. https://doi.org/10.3390/ijms18061259
Dong B, Deng Y, Wang H, Gao R, Stephen GK, Chen S, Jiang J, Chen F. Gibberellic Acid Signaling Is Required to Induce Flowering of Chrysanthemums Grown under Both Short and Long Days. International Journal of Molecular Sciences. 2017; 18(6):1259. https://doi.org/10.3390/ijms18061259
Chicago/Turabian StyleDong, Bin, Ye Deng, Haibin Wang, Ri Gao, Githeng’u K. Stephen, Sumei Chen, Jiafu Jiang, and Fadi Chen. 2017. "Gibberellic Acid Signaling Is Required to Induce Flowering of Chrysanthemums Grown under Both Short and Long Days" International Journal of Molecular Sciences 18, no. 6: 1259. https://doi.org/10.3390/ijms18061259
APA StyleDong, B., Deng, Y., Wang, H., Gao, R., Stephen, G. K., Chen, S., Jiang, J., & Chen, F. (2017). Gibberellic Acid Signaling Is Required to Induce Flowering of Chrysanthemums Grown under Both Short and Long Days. International Journal of Molecular Sciences, 18(6), 1259. https://doi.org/10.3390/ijms18061259