Identification of Quantitative Trait Loci Conditioning the Main Biomass Yield Components and Resistance to Melampsora spp. in Salix viminalis × Salix schwerinii Hybrids
Abstract
:1. Introduction
2. Results
2.1. Yield-Associated Traits
2.2. Resistance/Susceptibility to Willow Rust
2.3. Linkage Analysis and Mapping
2.4. Identification of QTLs
3. Discussion
4. Materials and Methods
4.1. Plant Material
4.2. Evaluation of Yield-Associated Traits
4.3. Fungal Material
4.4. Evaluation of Plant Resistance
4.5. RAPD and ISSR Markers
4.6. Linkage Analysis, Genetic Mapping and QTL Mapping
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Karp, A. Willows as a source of renewable fuels and diverse products. In Challenges and Opportunities for the World's Forests in the 21st Century; Fenning, T., Ed.; Springer: Dordrecht, The Netherlands, 2014; Volume 81, pp. 617–641. [Google Scholar]
- Przyborowski, J.A.; Jędryczka, M.; Ciszewska-Marciniak, J.; Sulima, P.; Wojciechowicz, K.M.; Zenkteler, E. Evaluation of the yield potential and physicochemical properties of the biomass of Salix viminalis × Populus tremula hybrids. Ind. Crops Prod. 2012, 36, 549–554. [Google Scholar] [CrossRef]
- Stolarski, M.J.; Szczukowski, S.; Tworkowski, J.; Klasa, A. Yield, energy parameters and chemical composition of short-rotation willow biomass. Ind. Crops Prod. 2013, 46, 60–65. [Google Scholar] [CrossRef]
- Weih, M. 19 Willow. In Biofuel Crops: Production, Physiology and Genetics; Singh, B.P., Ed.; CABI: Wallingford, UK, 2013; p. 415. [Google Scholar]
- Karp, A.; Hanley, S.J.; Trybush, S.O.; Macalpine, W.; Pei, M.; Shield, I. Genetic improvement of willow for bioenergy and biofuels. J. Integr. Plant Biol. 2011, 53, 151–165. [Google Scholar] [CrossRef] [PubMed]
- Krzyżaniak, M.; Stolarski, M.J.; Waliszewska, B.; Szczukowski, S.; Tworkowski, J.; Załuski, D.; Śnieg, M. Willow biomass as feedstock for an integrated multi-product biorefinery. Ind. Crops Prod. 2014, 58, 230–237. [Google Scholar] [CrossRef]
- Hanley, S.J.; Karp, A. Genetic strategies for dissecting complex traits in biomass willows (Salix spp.). Tree Physiol. 2013, 34, 1167–1180. [Google Scholar] [CrossRef] [PubMed]
- Berlin, S.; Trybush, S.O.; Fogelqvist, J.; Gyllenstrand, N.; Hallingbäck, H.R.; Åhman, I.; Hanley, S.J. Genetic diversity, population structure and phenotypic variation in European Salix viminalis L. (Salicaceae). Tree Genet. Genomes 2014, 10, 1595–1610. [Google Scholar] [CrossRef]
- Przyborowski, J.A.; Sulima, P. The analysis of genetic diversity of Salix viminalis genotypes as a potential source of biomass by RAPD markers. Ind. Crops Prod. 2010, 31, 395–400. [Google Scholar] [CrossRef]
- Trybush, S.O.; Jahodová, Š.; Čížková, L.; Karp, A.; Hanley, S.J. High levels of genetic diversity in Salix viminalis of the Czech Republic as revealed by microsatellite markers. Bioenergy Res. 2012, 5, 969–977. [Google Scholar] [CrossRef]
- Stolarski, M.J.; Szczukowski, S.; Tworkowski, J.; Wróblewska, H.; Krzyżaniak, M. Short rotation willow coppice biomass as an industrial and energy feedstock. Ind. Crops Prod. 2011, 33, 217–223. [Google Scholar] [CrossRef]
- Yang, H.; Li, C.; Lam, H.M.; Clements, J.; Yan, G.; Zhao, S. Sequencing consolidates molecular markers with plant breeding practice. Theor. Appl. Genet. 2015, 128, 779–795. [Google Scholar] [CrossRef] [PubMed]
- Berlin, S.; Lagercrantz, U.; von Arnold, S.; Öst, T.; Rönnberg-Wästljung, A.C. High-density linkage mapping and evolution of paralogs and orthologs in Salix and Populus. BMC Genom. 2010, 11, 129. [Google Scholar] [CrossRef] [PubMed]
- Ghelardini, L.; Berlin, S.; Weih, M.; Lagercrantz, U.; Gyllenstrand, N.; Rönnberg-Wästljung, A.C. Genetic architecture of spring and autumn phenology in Salix. BMC Plant Biol. 2014, 14, 31. [Google Scholar] [CrossRef] [PubMed]
- Hanley, S.; Mallott, M.; Karp, A. Alignment of a Salix linkage map to the Populus genomic sequence reveals macrosynteny between willow and poplar genomes. Tree Genet. Genomes 2006, 3, 35–48. [Google Scholar] [CrossRef]
- Hanley, S.; Barker, J.; van Ooijen, J.; Aldam, C.; Harris, S.; Åhman, I.; Larsson, S.; Karp, A. A genetic linkage map of willow (Salix viminalis) based on AFLP and microsatellite markers. Theor. Appl. Genet. 2002, 105, 1087–1096. [Google Scholar] [PubMed]
- Tsarouhas, V.; Gullberg, U.; Lagercrantz, U. An AFLP and RFLP linkage map and quantitative trait locus (QTL) analysis of growth traits in Salix. Theor. Appl. Genet. 2002, 105, 277–288. [Google Scholar] [PubMed]
- Berlin, S.; Ghelardini, L.; Bonosi, L.; Weih, M.; Rönnberg-Wästljung, A.C. QTL mapping of biomass and nitrogen economy traits in willows (Salix spp.) grown under contrasting water and nutrient conditions. Mol. Breed. 2014, 34, 1987–2003. [Google Scholar] [CrossRef]
- Hallingbäck, H.R.; Fogelqvist, J.; Powers, S.J.; Turrion-Gomez, J.; Rossiter, R.; Amey, J.; Martin, T.; Weih, M.; Gyllenstrand, N.; Karp, A.; et al. Association mapping in Salix viminalis L. (Salicaceae)-identification of candidate genes associated with growth and phenology. GCB Bioenergy 2016, 8, 670–685. [Google Scholar] [CrossRef] [PubMed]
- Salmon, J.; Ward, S.P.; Hanley, S.J.; Leyser, O.; Karp, A. Functional screening of willow alleles in Arabidopsis combined with QTL mapping in willow (Salix) identifies SxMAX4 as a coppicing response gene. Plant Biotechnol. J. 2014, 12, 480–491. [Google Scholar] [CrossRef] [PubMed]
- Weih, M.; Rönnberg-Wästljung, A.C.; Glynn, C. Genetic basis of phenotypic correlations among growth traits in hybrid willow (Salix dasyclados × S. viminalis) grown under two water regimes. New Phytol. 2006, 170, 467–477. [Google Scholar] [CrossRef] [PubMed]
- Samils, B.; Rönnberg-Wästljung, A.C.; Stenlid, J. QTL mapping of resistance to leaf rust in Salix. Tree Genet. Genomes 2011, 7, 1219–1235. [Google Scholar] [CrossRef]
- Pucholt, P.; Rönnberg-Wästljung, A.C.; Berlin, S. Single locus sex determination and female heterogamety in the basket willow (Salix viminalis L.). Heredity 2015, 114, 575–583. [Google Scholar] [CrossRef] [PubMed]
- Brereton, N.J.B.; Pitre, F.E.; Hanley, S.J.; Ray, M.J.; Karp, A.; Murphy, R.J. QTL mapping of enzymatic saccharification in short rotation coppice willow and its independence from biomass yield. Bioenergy Res. 2010, 3, 251–261. [Google Scholar] [CrossRef]
- Rönnberg-Wästljung, A.C.; Glynn, C.; Weih, M. QTL analyses of drought tolerance and growth for a Salix dasyclados × Salix viminalis hybrid in contrasting water regimes. Theor. Appl. Genet. 2005, 110, 537–549. [Google Scholar] [CrossRef] [PubMed]
- Hanley, S.J.; Pei, M.H.; Powers, S.J.; Ruiz, C.; Mallott, M.D.; Barker, J.H.; Karp, A. Genetic mapping of rust resistance loci in biomass willow. Tree Genet. Genomes 2011, 7, 597–608. [Google Scholar] [CrossRef]
- Höglund, S.; Rönnberg-Wästljung, A.C.; Lagercrantz, U.; Larsson, S. A rare major plant QTL determines non-responsiveness to a gall-forming insect in willow. Tree Genet. Genomes 2012, 8, 1051–1060. [Google Scholar] [CrossRef]
- Parker, S.R.; Royle, D.J.; Hunter, T. Impact of Melampsora rust on yield of biomass willows. In Proceedings of the 6th International Congress of Plant Pathology, Montreal, QC, Canada, 28 July–6 August 1993; National Research Council Canada: Ottawa, ON, Canada, 1993; p. 117. [Google Scholar]
- Jędryczka, M.; Ciszewska-Marciniak, J.; Przyborowski, J. The search for genetic sources of willow resistance to rust (Melampsora epitea). Phytopathol. Pol. 2008, 49, 5–19. [Google Scholar]
- Van Ooijen, J. MapQTL 6. Software for the Mapping of Quantitative Trait Loci in Experimental Populations of Diploid Species; Kyazma B.V.: Wageningen, The Netherlands, 2009. [Google Scholar]
- Rönnberg-Wästljung, A.C.; Samils, B.; Tsarouhas, V.; Gulberg, U. Resistance to Melampsora larici-epitea leaf rust in Salix; analysis quantitative trait loci. J. Appl. Genet. 2008, 49, 321–331. [Google Scholar] [CrossRef] [PubMed]
- Cochran, W.G.; Cox, G.M. Experimental Designs; Wiley Publications in Statistics: New York, NY, USA, 1955; pp. 1–455. [Google Scholar]
- Shapiro, S.S.; Wilk, M.B. An analysis of variance test for normality (complete samples). Biometrika 1965, 52, 591–611. [Google Scholar] [CrossRef]
- StatSoft, Inc. 2014 STATISTICA (Data Analysis Software System), Version 12.5. Available online: http://www.statsoft.com (accessed on 23 December 2016).
- Feau, N.; Vialle, A.; Allaire, M.; Tanguay, P.; Hamelin, R.C. Fungal pathogen (mis-) identifications: A case study with DNA barcodes on Melampsora rusts of aspen and white poplar. Mycol. Res. 2009, 113, 713–724. [Google Scholar] [CrossRef] [PubMed]
- Sokal, R.R.; Rohlf, F.J. Biometry: The Principles and Practice of Statistics in Biological Research, 3rd ed.; W.H. Freeman and Company: New York, NY, USA, 1995. [Google Scholar]
- Sulima, P.; Przyborowski, J.A.; Załuski, D. RAPD markers reveal genetic diversity in Salix purpurea. Crop Sci. 2009, 49, 857–863. [Google Scholar] [CrossRef]
- Williams, J.G.K.; Kubelik, A.R.; Livak, K.J.; Rafalski, J.A.; Tingey, S.V. DNA polymorphism amplified by arbitrary primers are useful as genetic markers. Nucleic Acids Res. 1990, 18, 6531–6535. [Google Scholar] [CrossRef] [PubMed]
- Zietkiewicz, E.; Rafalski, A.; Labuda, D. Genome fingerprinting by simple sequence repeat (SSR)-anchored polymerase chain reaction amplification. Genomics 1994, 20, 176–183. [Google Scholar] [CrossRef] [PubMed]
- McGregor, C.E.; Lambert, C.A.; Greyling, M.M.; Louw, J.H.; Warnich, L. A comparative assessment of DNA fingerprinting techniques (RAPD, ISSR, AFLP and SSR) in tetraploid potato (Solanum tuberosum L.) germplasm. Euphytica 2000, 113, 135–144. [Google Scholar] [CrossRef]
- Broman, K.W.; Wu, H.; Sen, S.; Churchill, G.A. R/QTL: QTL mapping in experimental crosses. Bioinformatics 2003, 19, 889–890. [Google Scholar] [CrossRef] [PubMed]
- Broman, K.W.; Sen, S. A Guide to QTL Mapping with R/QTL; Springer: New York, NY, USA, 2009; pp. 1–400. [Google Scholar]
- Xu, S. Mapping quantitative trait loci using four-way crosses. Genet. Res. 1996, 68, 175–181. [Google Scholar] [CrossRef]
- Lander, E.S.; Botstein, D. Mapping Mendelian factors underlying quantitative traits using RFLP linkage maps. Genetics 1989, 121, 185–199. [Google Scholar] [PubMed]
- Churchill, G.A.; Doerge, R.W. Empirical threshold values for quantitative trait mapping. Genetics 1994, 138, 963–971. [Google Scholar] [PubMed]
- Hoeschele, I.; van Raden, P.M. Bayesian analysis of linkage between genetic markers and quantitative trait loci. I. Prior knowledge. Theor. Appl. Genet. 1993, 85, 953–960. [Google Scholar] [CrossRef] [PubMed]
- Voorrips, R.E. MapChart: Software for the graphical presentation of linkage maps and QTLs. J. Hered. 2002, 93, 77–78. [Google Scholar] [CrossRef] [PubMed]
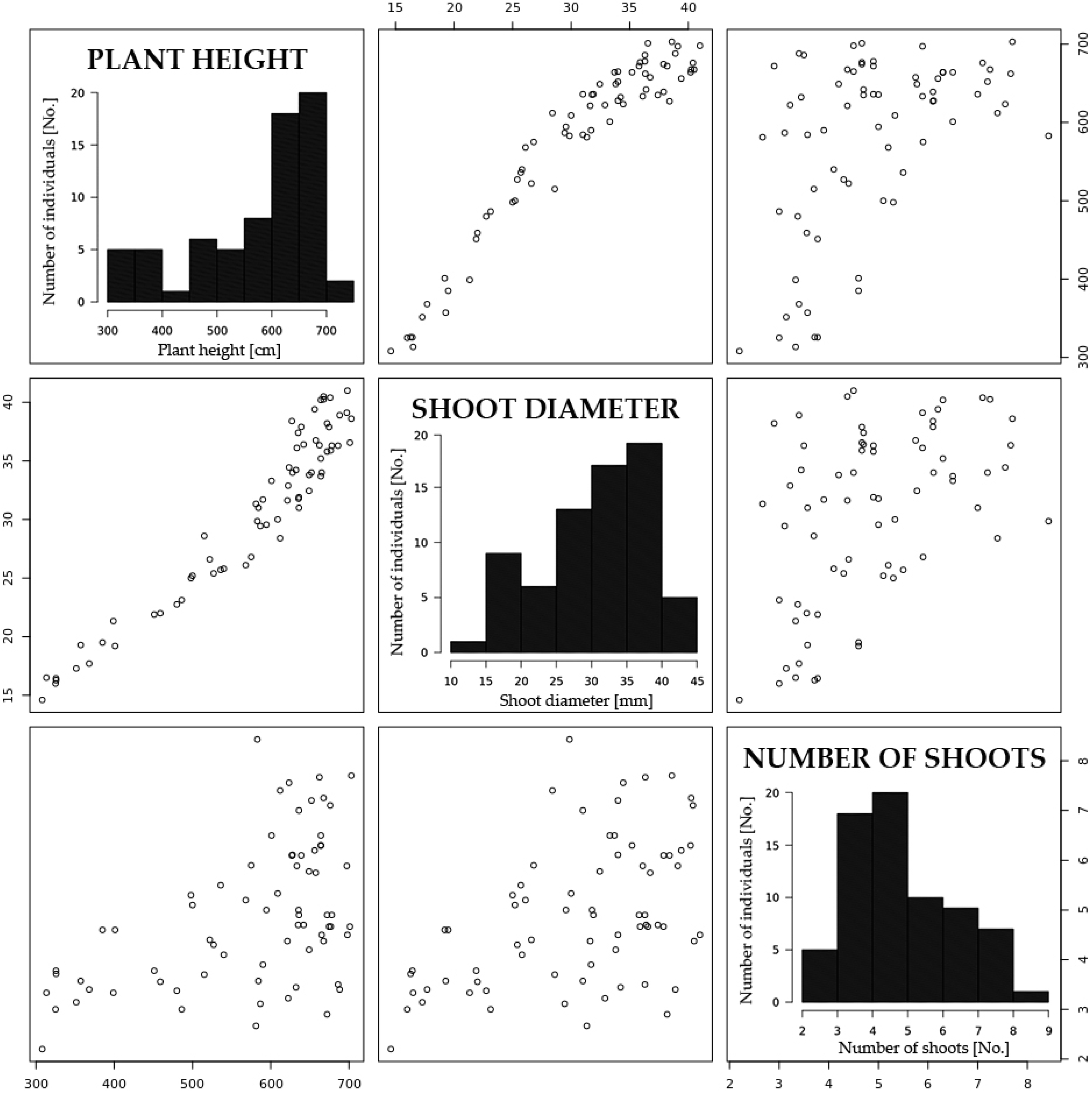
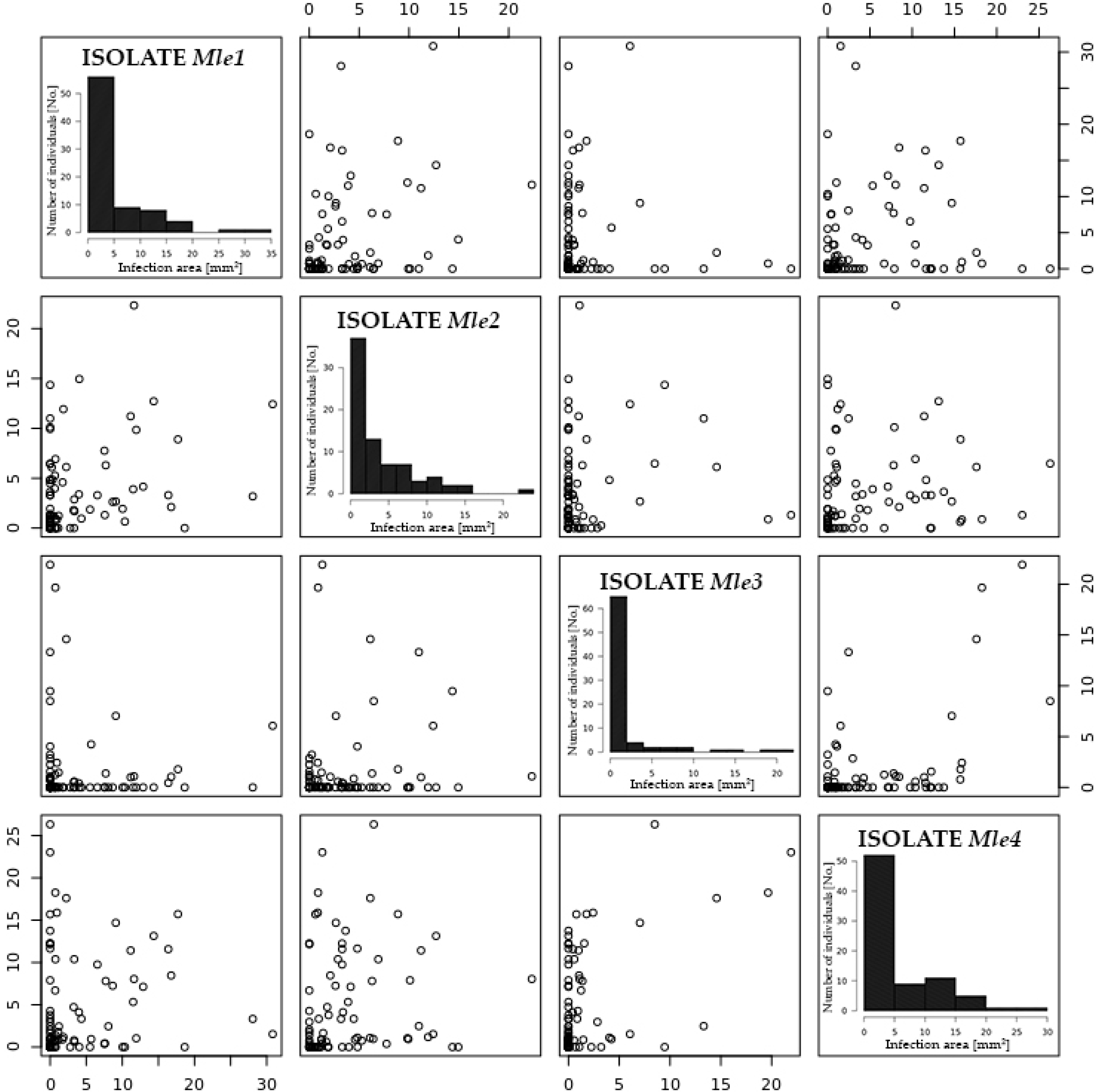
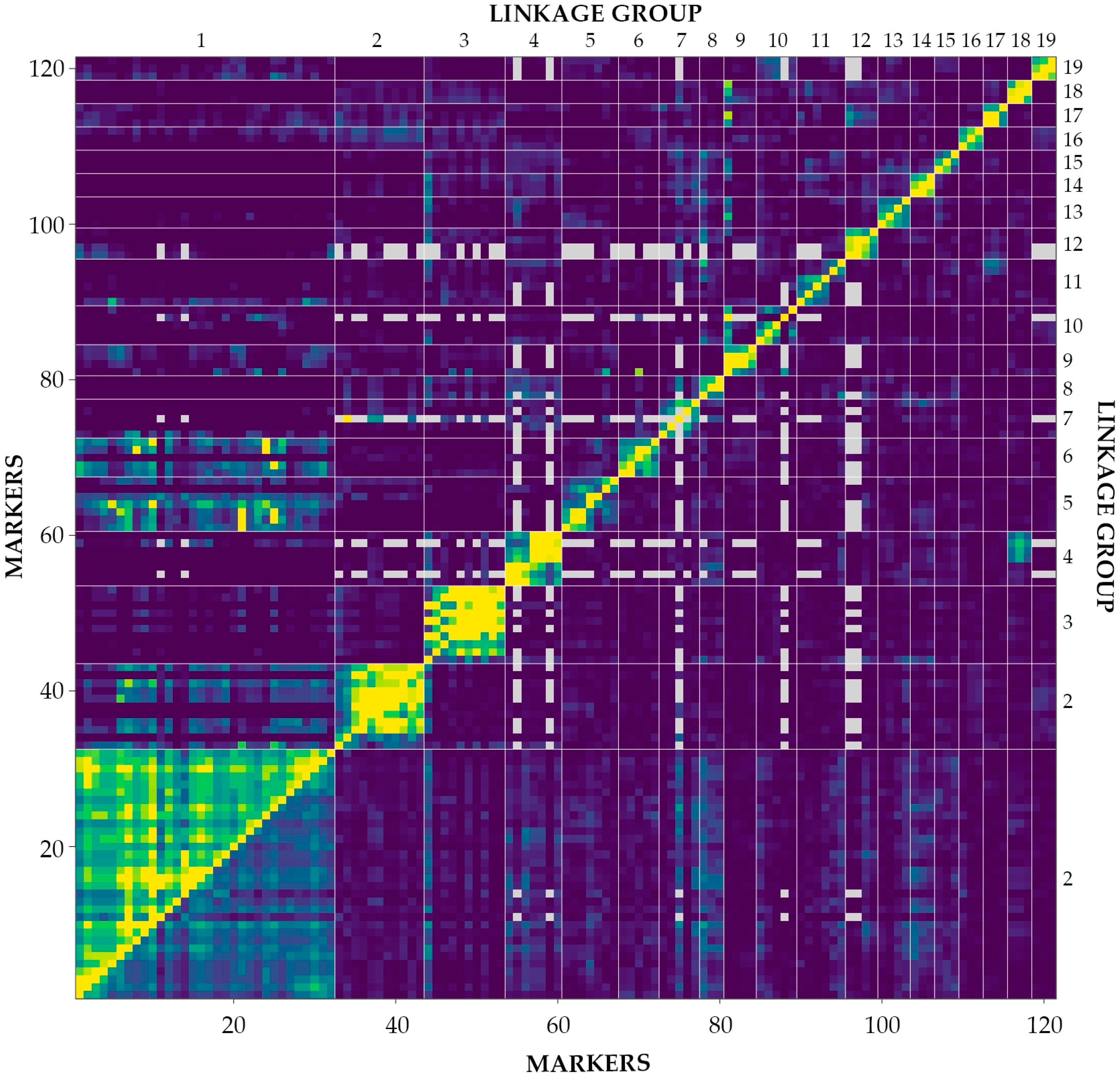
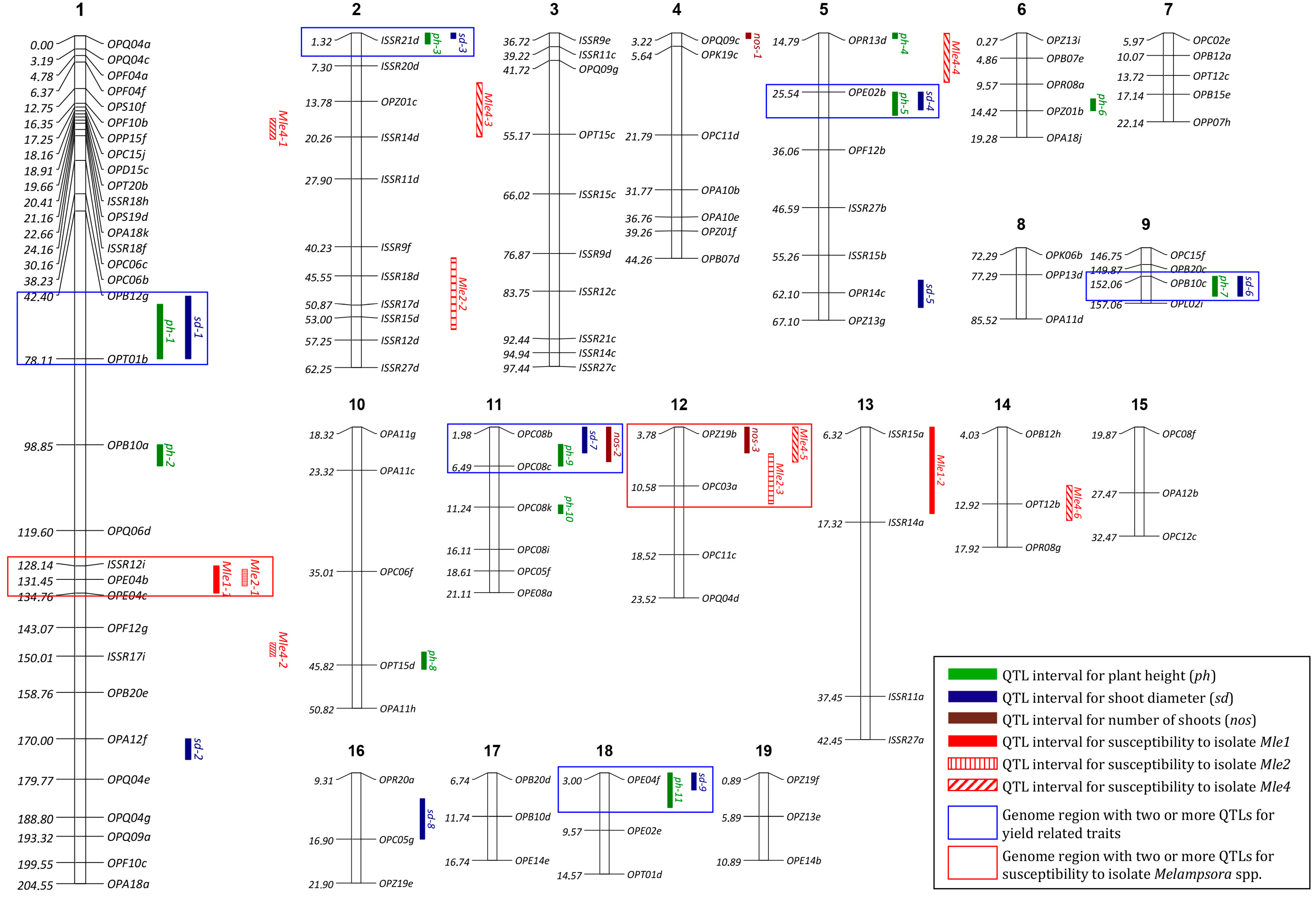
| Trait | Plants One-Year | Plants Two-Year | Plants Three-Year |
|---|---|---|---|
| Plant height (cm) | 346.3 ± 34.2 (9.9%) 1 | 436.6 ± 57.1 (13.1%) | 576.9 ± 114.6 (19.9%) |
| Shoot diameter (mm) | 19.1 ± 2.8 (14.7%) | 21.6 ± 3.5 (16.2%) | 30.7 ± 7.5 (24.4%) |
| Number of shoots (No.) | 8.6 ± 2.4 (27.9%) | 6.8 ± 1.7 (25.0%) | 4.9 ± 1.5 (30.6%) |
| Isolate Name | Leaf Area Covered with Uredinia (%) | Mean Area of a Pustule (mm2) |
|---|---|---|
| Mle1 | 25.41 c 1 | 1.773 c |
| Mle2 | 32.14 b | 1.405 b |
| Mle3 | 10.71 d | 0.617 d |
| Mle4 | 40.49 a | 2.355 a |
| Parameter | Mapping Population |
|---|---|
| P5 | |
| Total markers | 724 |
| Average markers per primer | 5.7 |
| RAPD 1 markers used for linkage analysis | 93 |
| ISSR 2 markers used for linkage analysis | 28 |
| Total markers used for linkage analysis | 121 |
| Unlinked or doublets markers | 340 |
| Number of groups in the framework | 19 |
| Smallest group (cM) of the framework | 10.0 |
| Largest group (cM) of the framework | 204.5 |
| Average length (cM) of group | 34.5 |
| Total length (cM) of the framework | 656.4 |
| Average distance between two framework markers (cM) ± SD | 6.3 ± 1.9 |
| Trait | LOD Threshold | QTL Name | Nearest Marker | LG 1 | Position (cM) | LOD | %PVE 2 |
|---|---|---|---|---|---|---|---|
| Plant height | 4.13 | ph-1 | OPT01b | 1 | 67.00 | 4.48 | 2.97 |
| ph-2 | OPB10a | 1 | 103.00 | 5.73 | 3.85 | ||
| ph-3 | ISSR21d | 2 | 2.32 | 6.77 | 4.61 | ||
| ph-4 | OPR13d | 5 | 14.79 | 8.27 | 5.73 | ||
| ph-5 | OPE02b | 5 | 28.79 | 6.99 | 4.78 | ||
| ph-6 | OPZ01b | 6 | 13.27 | 10.51 | 7.49 | ||
| ph-7 | OPB10c | 9 | 154.75 | 13.20 | 9.72 | ||
| ph-8 | OPT15d | 10 | 45.82 | 11.27 | 8.10 | ||
| ph-9 | OPC08c | 11 | 4.98 | 7.19 | 4.92 | ||
| ph-10 | OPC08k | 11 | 11.24 | 5.59 | 3.75 | ||
| ph-11 | OPE04f | 18 | 6.01 | 6.62 | 4.50 | ||
| Shoot diameter | 3.26 | sd-1 | OPT01b | 1 | 64.00 | 3.86 | 4.10 |
| sd-2 | OPA12f | 1 | 174.00 | 3.50 | 4.88 | ||
| sd-3 | ISSR21d | 2 | 1.32 | 5.03 | 4.40 | ||
| sd-4 | OPE02b | 5 | 27.79 | 6.07 | 6.46 | ||
| sd-5 | OPR14c | 5 | 60.79 | 3.71 | 7.90 | ||
| sd-6 | OPB10c | 9 | 154.75 | 3.95 | 4.52 | ||
| sd-7 | OPC08b | 11 | 3.98 | 3.27 | 4.68 | ||
| sd-8 | OPC05g | 16 | 13.31 | 3.59 | 4.65 | ||
| sd-9 | OPE04f | 18 | 4.00 | 3.69 | 4.99 | ||
| Number of shoots | 3.19 | nos-1 | OPQ09c | 4 | 4.22 | 3.45 | 10.60 |
| nos-2 | OPC08b | 11 | 1.98 | 5.08 | 16.16 | ||
| nos-3 | OPZ19b | 12 | 3.78 | 3.38 | 10.37 | ||
| IALD 3 by isolate Mle1 | 8.36 | Mle1-1 | OPE04b | 1 | 131.45 | 12.2 | 15.47 |
| Mle1-2 | ISSR15a | 13 | 16.32 | 8.76 | 10.82 | ||
| IALD 3 by isolate Mle2 | 5.88 | Mle2-1 | OPE04b | 1 | 131.00 | 6.36 | 5.89 |
| Mle2-2 | ISSR15d | 2 | 55.32 | 5.37 | 4.95 | ||
| Mle2-3 | OPC03a | 12 | 8.78 | 9.40 | 8.86 | ||
| IALD 3 by isolate Mle4 | 5.23 | Mle4-1 | OPA18k | 1 | 22.00 | 14.09 | 10.15 |
| Mle4-2 | ISSR17i | 1 | 149.00 | 10.68 | 7.52 | ||
| Mle4-3 | OPZ01c | 2 | 15.32 | 19.50 | 14.57 | ||
| Mle4-4 | OPE02b | 5 | 24.79 | 17.05 | 12.53 | ||
| Mle4-5 | OPZ19b | 12 | 4.78 | 6.50 | 4.46 | ||
| Mle4-6 | OPT12b | 14 | 12.92 | 10.94 | 7.72 |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license ( http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sulima, P.; Przyborowski, J.A.; Kuszewska, A.; Załuski, D.; Jędryczka, M.; Irzykowski, W. Identification of Quantitative Trait Loci Conditioning the Main Biomass Yield Components and Resistance to Melampsora spp. in Salix viminalis × Salix schwerinii Hybrids. Int. J. Mol. Sci. 2017, 18, 677. https://doi.org/10.3390/ijms18030677
Sulima P, Przyborowski JA, Kuszewska A, Załuski D, Jędryczka M, Irzykowski W. Identification of Quantitative Trait Loci Conditioning the Main Biomass Yield Components and Resistance to Melampsora spp. in Salix viminalis × Salix schwerinii Hybrids. International Journal of Molecular Sciences. 2017; 18(3):677. https://doi.org/10.3390/ijms18030677
Chicago/Turabian StyleSulima, Paweł, Jerzy A. Przyborowski, Anna Kuszewska, Dariusz Załuski, Małgorzata Jędryczka, and Witold Irzykowski. 2017. "Identification of Quantitative Trait Loci Conditioning the Main Biomass Yield Components and Resistance to Melampsora spp. in Salix viminalis × Salix schwerinii Hybrids" International Journal of Molecular Sciences 18, no. 3: 677. https://doi.org/10.3390/ijms18030677
APA StyleSulima, P., Przyborowski, J. A., Kuszewska, A., Załuski, D., Jędryczka, M., & Irzykowski, W. (2017). Identification of Quantitative Trait Loci Conditioning the Main Biomass Yield Components and Resistance to Melampsora spp. in Salix viminalis × Salix schwerinii Hybrids. International Journal of Molecular Sciences, 18(3), 677. https://doi.org/10.3390/ijms18030677









