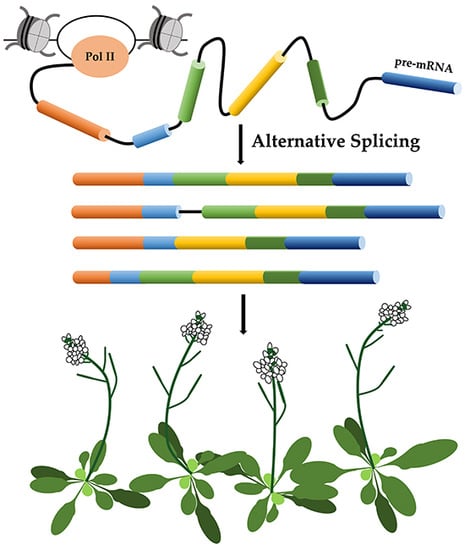
Alternative Splicing in Plant Genes: A Means of Regulating the Environmental Fitness of Plants
Abstract
:1. Introduction
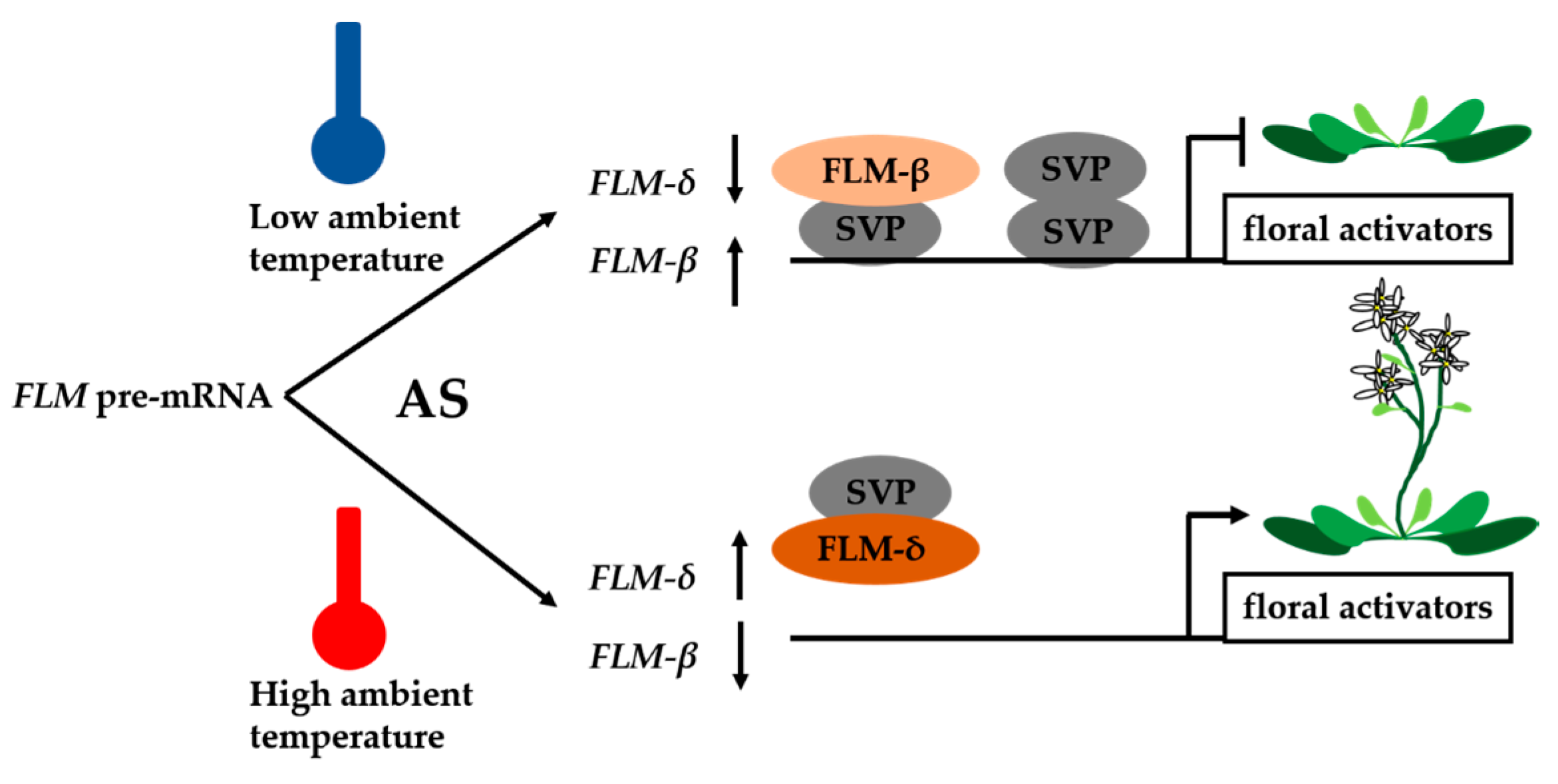
2. Ambient Temperature and Vernalization-Mediated Flowering
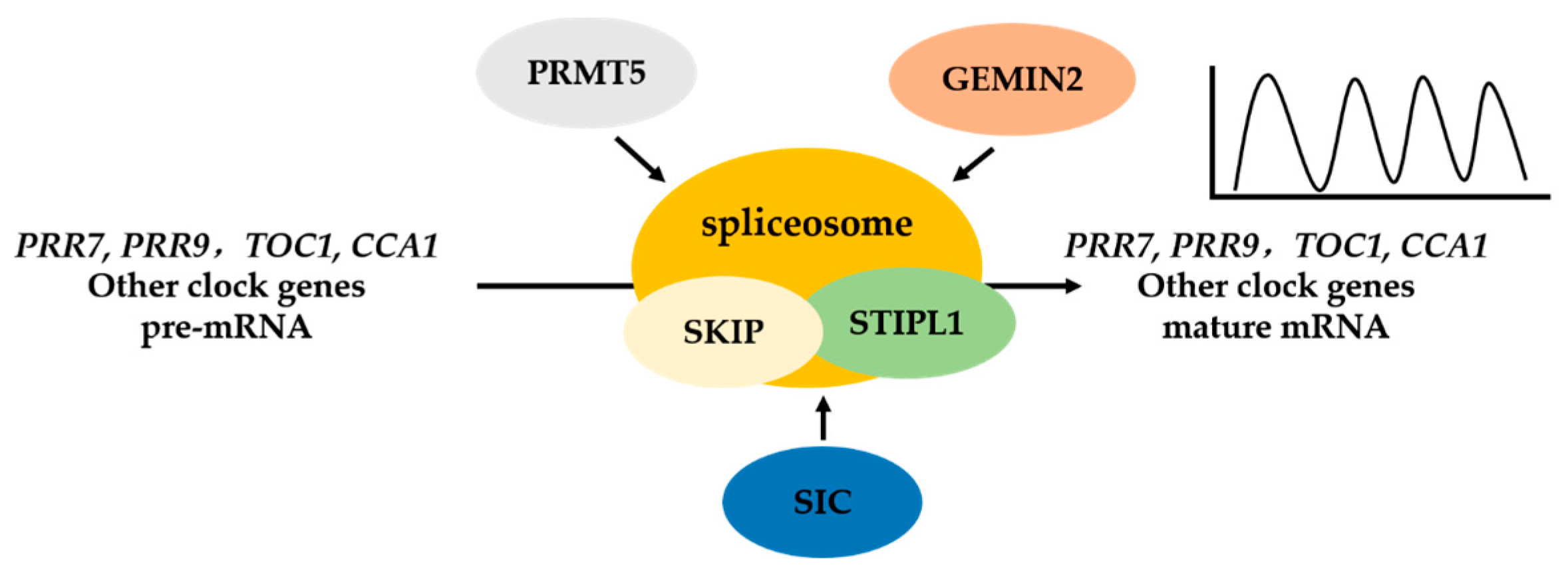
3. The Circadian Clock in Plants
4. Abiotic Stress Responses
5. Biotic Stress Responses
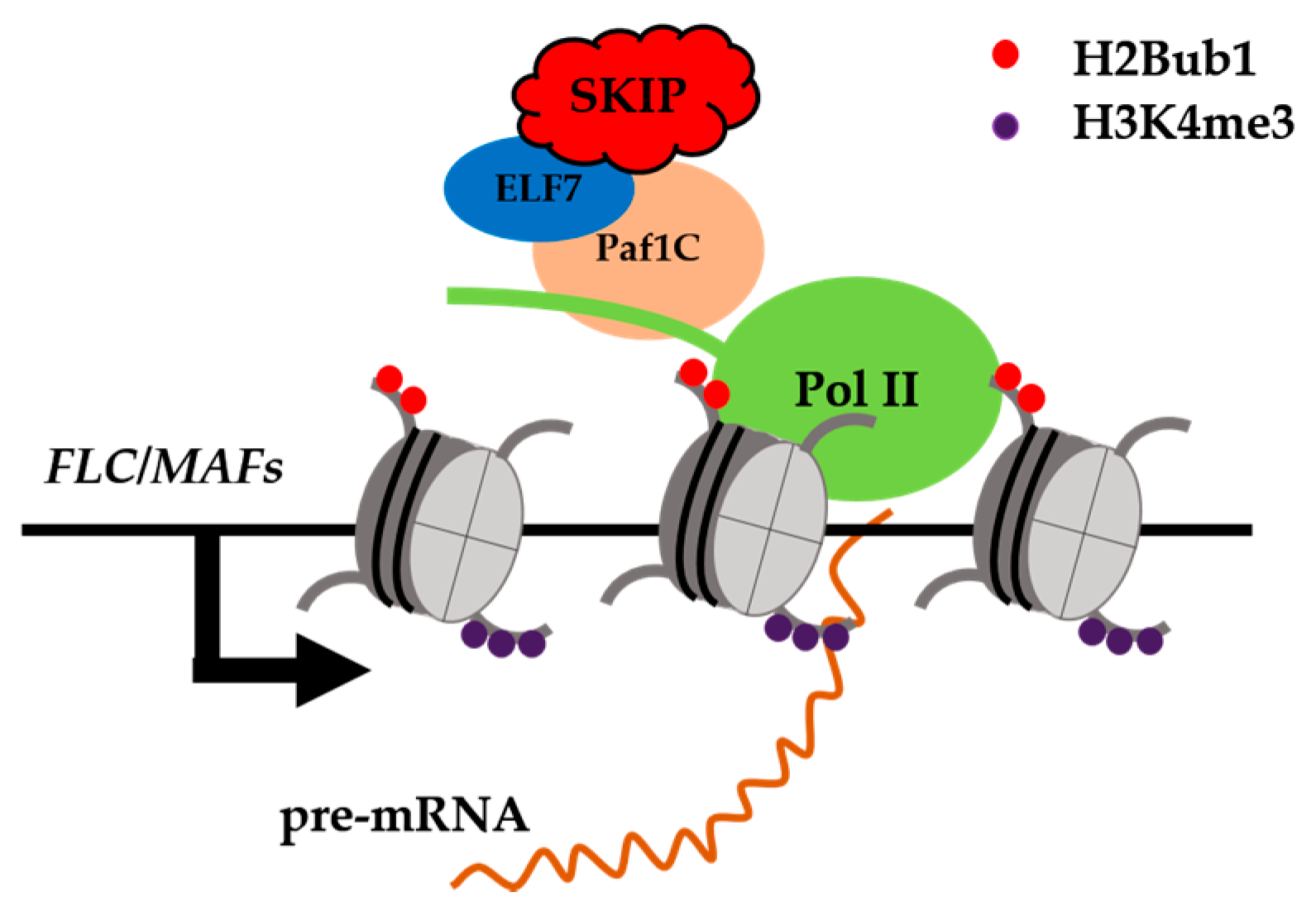
6. Splicing Factors or Transcriptional Co-Regulators in Plants
7. Perspectives and Conclusions
Acknowledgments
Conflicts of Interest
References
- Sharp, P.A. Split genes and RNA splicing. Cell 1994, 77, 805–815. [Google Scholar] [CrossRef]
- Sharp, P.A. The discovery of split genes and RNA splicing. Trends Biochem. Sci. 2005, 30, 279–281. [Google Scholar] [CrossRef] [PubMed]
- Moore, M.J.; Proudfoot, N.J. Pre-mRNA processing reaches back to transcription and ahead to translation. Cell 2009, 136, 688–700. [Google Scholar] [CrossRef] [PubMed]
- Sharp, P.A. The centrality of RNA. Cell 2009, 136, 577–580. [Google Scholar] [CrossRef] [PubMed]
- Fu, X.D.; Manuel Ares, J. Context-dependent control of alternative splicing by RNA-binding proteins. Nat. Rev. Genet. 2014, 15, 689–701. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Burge, C.B. Splicing regulation: From a parts list of regulatory elements to an integrated splicing code. RNA 2008, 14, 802–813. [Google Scholar] [CrossRef] [PubMed]
- Deckert, J.; Hartmuth, K.; Boehringer, D.; Behzadnia, N.; Will, C.L.; Kastner, B.; Stark, H.; Urlaub, H.; Lührmann, R. Protein composition and electron microscopy structure of affinity-purified human spliceosomal B complexes isolated under physiological conditions. Mol. Cell. Biol. 2006, 26, 5528–5543. [Google Scholar] [CrossRef] [PubMed]
- Jurica, M.S.; Moore, M.J. Pre-mRNA splicing: Awash in a sea of proteins. Mol. Cell 2003, 12, 5–14. [Google Scholar] [CrossRef]
- Wahl, M.C.; Will, C.L.; Lührmann, R. The spliceosome: Design principles of a dynamic RNP machine. Cell 2009, 136, 701–718. [Google Scholar] [CrossRef] [PubMed]
- Klepikova, A.V.; Kasianov, A.S.; Gerasimov, E.S.; Logacheva, M.D.; Penin, A.A. A high resolution map of the Arabidopsis thaliana developmental transcriptome based on RNA-seq profiling. Plant J. 2016, 88, 1058–1070. [Google Scholar] [CrossRef] [PubMed]
- Marquez, Y.; Brown, J.W.S.; Simpson, C.; Barta, A.; Kalyna, M. Transcriptome survey reveals increased complexity of the alternative splicing landscape in Arabidopsis. Genome Res. 2012, 22, 1184–1195. [Google Scholar] [CrossRef] [PubMed]
- Reddy, A.S.N.; Marquez, Y.; Kalyna, M.; Barta, A. Complexity of the alternative splicing landscape in plants. Plant Cell 2013, 25, 3657–3683. [Google Scholar] [CrossRef] [PubMed]
- McGlincy, N.J.; Smith, C.W. Alternative splicing resulting in nonsense-mediated mRNA decay: What is the meaning of nonsense? Trends Biochem. Sci. 2008, 33, 385–393. [Google Scholar] [CrossRef] [PubMed]
- Kalyna, M.; Simpson, C.G.; Syed, N.H.; Lewandowska, D.; Marquez, Y.; Kusenda, B.; Marshall, J.; Fuller, J.; Cardle, L.; McNicol, J.; et al. Alternative splicing and nonsense-mediated decay modulate expression of important regulatory genes in Arabidopsis. Nucleic Acids Res. 2012, 40, 2454–2469. [Google Scholar] [CrossRef] [PubMed]
- Syed, N.H.; Kalyna, M.; Marquez, Y.; Barta, A.; and Brown, J.W.S. Alternative splicing in plants—Coming of age. Trends Plant Sci. 2012, 17, 616–623. [Google Scholar] [CrossRef] [PubMed]
- Seo, P.J.; Hong, S.Y.; Kim, S.G.; Park, C.M. Competitive inhibition of transcription factors by small interfering peptides. Trends Plant Sci. 2011, 16, 541–549. [Google Scholar] [CrossRef] [PubMed]
- Michaels, S.D. Flowering time regulation produces much fruit. Curr. Opin. Plant Biol. 2009, 12, 75–80. [Google Scholar] [CrossRef] [PubMed]
- Song, J.; Angel, A.; Howard, M.; Dean, C. Vernalization—A cold induced epigenetic switch. J. Cell Sci. 2012, 125, 3723–3731. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.H.; Ryu, H.S.; Chung, K.S.; Posé, D.; Kim, S.; Schmid, M.; Ahn, J.H. Regulation of temperature-responsive flowering by MADS-box transcription factor repressors. Science 2013, 342, 628–632. [Google Scholar] [CrossRef] [PubMed]
- Posé, D.; Verhage, L.; Ott, F.; Yant, L.; Mathieu, J.; Angenent, G.C.; Immink, R.G.H.; Schmid, M. Temperature-dependent regulation of flowering by antagonistic FLM variants. Nature 2013, 503, 414–417. [Google Scholar] [CrossRef] [PubMed]
- Amasino, R.M.; Michaels, S.D. The timing of flowering. Plant Physiol. 2010, 154, 516–520. [Google Scholar] [CrossRef] [PubMed]
- Ietswaart, R.; Wu, Z.; Dean, C. Flowering time control: Another window to the connection between antisense RNA and chromatin. Trends Genet. 2012, 28, 445–453. [Google Scholar] [CrossRef] [PubMed]
- Swiezewski, S.; Liu, F.; Magusin, A.; Dean, C. Cold-induced silencing mediated by long antisense RNA from a Polycomb target. Nature 2009, 462, 799–802. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Wu, Z.; Raitskin, O.; Sun, Q.; Dean, C. Antisense-mediated FLC transcriptional repression requires the p-TEFb transcription elongation factor. Proc. Natl. Acad. Sci. USA 2014, 111, 7468–7473. [Google Scholar] [CrossRef] [PubMed]
- Marquardt, S.; Raitskin, O.; Wu, Z.; Liu, F.; Sun, Q.; Dean, C. Functional consequences of splicing of the antisense transcript COOLAIR on FLC transcription. Mol. Cell 2014, 54, 156–165. [Google Scholar] [CrossRef] [PubMed]
- Li, P.; Tao, Z.; Dean, C. Phenotypic evolution through variation in splicing of the noncoding RNA, COOLAIR. Genes Dev. 2015, 29, 696–701. [Google Scholar] [CrossRef] [PubMed]
- Bardou, F.; Ariel, F.; Simpson, C.G.; Romero-Barrios, N.; Laporte, P.; Balzergue, S.; Brown, J.W.; Crespi, M. Long noncoding RNA modulates alternative splicing regulators in Arabidopsis. Dev. Cell 2014, 30, 166–176. [Google Scholar] [CrossRef] [PubMed]
- Jiao, Y.; Meyerowitz, E.M. Cell-type specific analysis of translating RNAs in developing flowers reveals new levels of control. Mol. Syst. Biol. 2010, 6, 419. [Google Scholar] [CrossRef] [PubMed]
- Balasubramanian, S.; Sureshkumar, S.; Lempe, J.; Weigel, D. Potent induction of Arabidopsis thaliana flowering by elevated growth temperature. PLoS Genet. 2006, 2, e106. [Google Scholar] [CrossRef] [PubMed]
- Balasubramanian, S.; Weigel, D. Temperature induced flowering in Arabidopsis thaliana. Plant Signal. Behav. 2006, 1, 227–228. [Google Scholar] [CrossRef]
- Lempe, J.; Balasubramanian, S.; Sureshkumar, S.; Singh, A.; Schmid, M.; Weigel, D. Diversity of flowering responses in wild Arabidopsis thaliana strains. PLoS Genet. 2005, 1, 109–118. [Google Scholar] [CrossRef]
- Rosloski, S.M.; Singh, A.; Jali, S.S.; Balasubramanian, S.; Weigel, D.; Grbic, V. Functional analysis of splice variant expression of MADS AFFECTING FLOWERING 2 of Arabidopsis thaliana. Plant Mol. Biol. 2012, 81, 57–69. [Google Scholar] [CrossRef] [PubMed]
- McClung, C.R. Plant circadian rhythms. Plant Cell 2006, 18, 792–803. [Google Scholar] [CrossRef]
- Harmer, S.L. The circadian system in higher plants. Annu. Rev. Plant Biol. 2009, 60, 357–377. [Google Scholar] [CrossRef] [PubMed]
- Schöning, J.C.; Staiger, D. At the pulse of time: Protein interactions determine the pace of circadian clocks. FEBS Lett. 2005, 579, 3246–3252. [Google Scholar] [CrossRef] [PubMed]
- Más, P. Circadian clock function in Arabidopsis thaliana: Time beyond transcription. Trends Cell Biol. 2008, 18, 273–281. [Google Scholar] [CrossRef] [PubMed]
- Herrero, E.; Davis, S.J. Time for a nuclear meeting: Protein trafficking and chromatin dynamics intersect in the plant circadian system. Mol. Plant 2012, 5, 554–565. [Google Scholar] [CrossRef]
- McClung, C.R. The genetics of plant clocks. Adv. Genet. 2011, 74, 105–139. [Google Scholar] [PubMed]
- Alabadi, D.; Oyama, T.; Yanovsky, M.J.; Harmon, F.G.; Mas, P.; Kay, S.A. Reciprocal regulation between TOC1 and LHY/CCA1 within the Arabidopsis circadian clock. Science 2001, 293, 880–883. [Google Scholar] [CrossRef] [PubMed]
- Green, R.M.; Tobin, E.M. The role of CCA1 and LHY in the plant circadian clock. Dev. Cell 2002, 2, 516–518. [Google Scholar] [CrossRef]
- Gendron, J.M.; Pruneda-Paz, J.L.; Doherty, C.J.; Gross, A.M.; Kang, S.E.; Kay, S.A. Arabidopsis circadian clock protein, TOC1, is a DNA-binding transcription factor. Proc. Natl. Acad. Sci. USA 2012, 109, 3167–3172. [Google Scholar] [CrossRef] [PubMed]
- Huang, W.; Perez-Garcia, P.; Pokhilko, A.; Millar, A.J.; Antoshechkin, I.; Riechmann, J.L.; Mas, P. Mapping the core of the Arabidopsis circadian clock defines the network structure of the oscillator. Science 2012, 336, 75–79. [Google Scholar] [CrossRef] [PubMed]
- Pokhilko, A.; Fernandez, A.P.; Edwards, K.D.; Southern, M.M.; Halliday, K.J.; Millar, A.J. The clock gene circuit in Arabidopsis includes a repressilator with additional feedback loops. Mol. Syst. Biol. 2012, 8, 574. [Google Scholar] [CrossRef] [PubMed]
- Farre, E.M.; Harmer, S.L.; Harmon, F.G.; Yanovsky, M.J.; Kay, S.A. Overlapping and distinct roles of PRR7 and PRR9 in the Arabidopsis circadian clock. Curr. Biol. 2005, 15, 47–54. [Google Scholar] [CrossRef] [PubMed]
- Nakamichi, N.; Kiba, T.; Henriques, R.; Mizuno, T.; Chua, N.H.; Sakakibara, H. PSEUDO-RESPONSE REGULATORS 9, 7, and 5 are transcriptional repressors in the Arabidopsis circadian clock. Plant Cell 2010, 22, 594–605. [Google Scholar] [CrossRef]
- Pruneda-Paz, J.L.; Kay, S.A. An expanding universe of circadian networks in higher plants. Trends Plant Sci. 2010, 15, 259–265. [Google Scholar] [CrossRef] [PubMed]
- Helfer, A.; Nusinow, D.A.; Chow, B.Y.; Gehrke, A.R.; Bulyk, M.L.; Kay, S.A. LUX ARRHYTHMO encodes a night time repressor of circadian gene expression in the Arabidopsis core clock. Curr. Biol. 2011, 21, 126–133. [Google Scholar] [CrossRef] [PubMed]
- Herrero, E.; Kolmos, E.; Bujdoso, N.; Yuan, Y.; Wang, M.; Berns, M.C.; Uhlworm, H.; Coupland, G.; Saini, R.; Jaskolski, M.; et al. EARLY FLOWERING4 recruitment of EARLY FLOWERING3 in the nucleus sustains the Arabidopsis circadian clock. Plant Cell 2012, 24, 428–443. [Google Scholar] [CrossRef] [PubMed]
- Filichkin, S.A.; Priest, H.D.; Givan, S.A.; Shen, R.; Bryant, D.W.; Fox, S.E.; Wong, W.K.; Mockler, T.C. Genome-wide mapping of alternative splicing in Arabidopsis thaliana. Genome Res. 2010, 20, 45–58. [Google Scholar] [CrossRef] [PubMed]
- Seo, P.J.; Park, M.J.; Lim, M.H.; Kim, S.G.; Lee, M.; Baldwin, I.T.; Park, C.M. A self-regulatory circuit of CIRCADIANCLOCK-ASSOCIATED1 underlies the circadian clock regulation of temperature responses in Arabidopsis. Plant Cell 2011, 24, 2427–2442. [Google Scholar] [CrossRef] [PubMed]
- Daniel, X.; Sugano, S.; Tobin, E.M. CK2 phosphorylation of CCA1 is necessary for its circadian oscillator function in Arabidopsis. Proc. Natl. Acad. Sci. USA 2004, 101, 3292–3297. [Google Scholar] [CrossRef] [PubMed]
- Lu, S.X.; Knowles, S.M.; Andronis, C.; Ong, M.S.; Tobin, E.M. CIRCADIAN CLOCK ASSOCIATED1 and LATE ELONGATED HYPOCOTYL function synergistically in the circadian clock of Arabidopsis. Plant Physiol. 2009, 150, 834–843. [Google Scholar] [CrossRef] [PubMed]
- Filichkin, S.A.; Cumbie, J.S.; Dharmawardhana, J.P.; Jaiswal, P.; Chang, J.H.; Palusa, S.G.; Reddy, A.S.; Megraw, M.; Mockler, T.C. Environmental stresses modulate abundance and timing of alternatively spliced circadian transcripts in Arabidopsis. Mol. Plant 2015, 8, 207–227. [Google Scholar] [CrossRef] [PubMed]
- Sanchez, S.E.; Petrillo, E.; Beckwith, E.J.; Zhang, X.; Rugnone, M.L.; Hernando, C.E.; Cuevas, J.C.; GodoyHerz, M.A.; Depetris-Chauvin, A.; Simpson, C.G.; et al. A methyl transferase links the circadian clock to the regulation of alternative splicing. Nature 2010, 468, 112–116. [Google Scholar] [CrossRef] [PubMed]
- Deng, X.; Gu, L.; Liu, C.; Lu, T.; Lu, F.; Lu, Z.; Cui, P.; Pei, Y.; Wang, B.; Hu, S.; et al. Arginine methylation mediated by the Arabidopsis homolog of PRMT5 is essential for proper pre-mRNA splicing. Proc. Natl. Acad. Sci. USA 2010, 107, 19114–19119. [Google Scholar] [CrossRef] [PubMed]
- Hong, S.; Song, H.R.; Lutz, K.; Kerstetter, R.A.; Michael, T.P.; McClung, C.R. Type II protein arginine methyltransferase 5 (PRMT5) is required for circadian period determination in Arabidopsis thaliana. Proc. Natl. Acad. Sci. USA 2010, 107, 21211–21216. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Wu, F.; Xie, Q.; Wang, H.; Wang, Y.; Yue, Y.; Gahura, O.; Ma, S.; Liu, L.; Cao, Y.; et al. SKIP is a component of the spliceosome linking alternative splicing and the circadian clock in Arabidopsis. Plant Cell 2012, 24, 3278–3295. [Google Scholar] [CrossRef] [PubMed]
- Jones, M.A.; Williams, B.A.; McNicol, J.; Simpson, C.G.; Brown, J.W.; Harmer, S.L. Mutation of Arabidopsis spliceosomal timekeeper locus1 causes circadian clock defects. Plant Cell 2012, 24, 4066–4082. [Google Scholar] [CrossRef] [PubMed]
- Figueroa, J.D.; Hayman, M.J. The human Ski-interacting protein functionally substitutes for the yeast PRP45 gene. Biochem. Biophys. Res. Commun. 2004, 319, 1105–1109. [Google Scholar] [CrossRef] [PubMed]
- Golovkin, M.; Reddy, A.S. Structure and expression of a plant U1 snRNP 70K gene: Alternative splicing of U1 snRNP 70K pre-mRNAs produces two different transcripts. Plant Cell 1996, 8, 1421–1435. [Google Scholar] [CrossRef] [PubMed]
- Day, I.S.; Golovkin, M.; Palusa, S.G.; Link, A.; Ali, G.S.; Thomas, J.; Richardson, D.N.; Reddy, A.S. Interactions of SR45, an SR-like protein, with spliceosomal proteins and an intronic sequence: Insights into regulated splicing. Plant J. Cell Mol. Biol. 2012, 71, 936–947. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Xia, C.; Feng, J.; Yang, D.; Wu, F.; Cao, Y.; Li, L.; Ma, L. The SNW domain of SKIP is required for itsintegration into the spliceosome and its interaction with the Pall complex in Arabidopsis. Mol. Plant 2016, 7, 1040–1050. [Google Scholar] [CrossRef] [PubMed]
- Schlaen, R.G.; Mancini, E.; Sanchez, S.E.; Perezsantángelo, S.; Rugnone, M.L.; Simpson, C.G.; Brown, J.W.; Zhang, X.; Chernomoretz, A.; Yanovsky, M.J. The spliceosome assembly factor GEMIN2 attenuates the effects of temperature on alternative splicing and circadian rhythms. Proc. Natl. Acad. Sci. USA 2015, 112, 9382–9387. [Google Scholar] [CrossRef] [PubMed]
- Marshall, C.M.; Tartaglio, V.; Duarte, M.; Harmon, F.G. The Arabidopsis sickle mutant exhibits altered circadian clock responses to cool temperatures and temperature-dependent alternative splicing. Plant Cell 2016, 10, 2560–2575. [Google Scholar] [CrossRef] [PubMed]
- Hegele, A.; Kamburov, A.; Grossmann, A.; Sourlis, C.; Wowro, S.; Weimann, M.; Will, C.L.; Pena, V.; Lührmann, R.; Stelzl, U. Dynamic protein–protein interaction wiring of the human spliceosome. Mol. Cell 2012, 45, 567–580. [Google Scholar] [CrossRef] [PubMed]
- Xiong, L.; Schumaker, K.S.; Zhu, J.K. Cell signaling during cold, drought, and salt stress. Plant Cell 2002, 14, S165–S183. [Google Scholar] [PubMed]
- Zhu, J.K. Salt and drought stress signal transduction in plants. Annu. Rev. Plant Biol. 2002, 53, 247–273. [Google Scholar] [CrossRef] [PubMed]
- Munns, R.; Tester, M. Mechanisms of salinity tolerance. Annu. Rev. Plant Biol. 2008, 59, 651–681. [Google Scholar] [CrossRef] [PubMed]
- Von Koskull-Döring, P.; Scharf, K.D.; Nover, L. The diversity of plant heat stress transcription factors. Trends Plant Sci. 2007, 12, 452–457. [Google Scholar] [CrossRef] [PubMed]
- Sugio, A.; Dreos, R.; Aparicio, F.; Maule, A.J. The cytosolic protein response as a subcomponent of the wider heat shock response in Arabidopsis. Plant Cell 2009, 21, 642–654. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Sun, N.; Liu, M.; Liu, J.; Du, B.; Wang, X.; Qi, X. An autoregulatory loop controlling Arabidopsis HsfA2 expression: Role of heat shock-induced alternative splicing. Plant Physiol. 2013, 162, 512–521. [Google Scholar] [CrossRef] [PubMed]
- Torres, M.A.; Dangl, J.L.; Jones, J.D. Arabidopsis gp91phox homologues AtrbohD and AtrbohF are required for accumulation of reactive oxygen intermediates in the plant defense response. Proc. Natl. Acad. Sci. USA 2002, 99, 517–522. [Google Scholar] [CrossRef]
- Torres, M.A.; Jones, J.D.; Dangl, J.L. Pathogen-induced, NADPH oxidase-derived reactive oxygen intermediates suppress spread of cell death in Arabidopsis thaliana. Nat. Genet. 2005, 37, 1130–1134. [Google Scholar] [CrossRef] [PubMed]
- Lin, F.; Zhang, Y.; Jiang, M.Y. Alternative splicing and differential expression of two transcripts of nicotine adenine dinucleotide phosphate oxidase B gene from Zea mays. J. Integr. Plant Biol. 2009, 51, 287–298. [Google Scholar] [CrossRef] [PubMed]
- Matsukura, S.; Mizoi, J.; Yoshida, T.; Todaka, D.; Ito, Y.; Maruyama, K.; Shinozaki, K.; Yamaguchi-Shinozaki, K. Comprehensive analysis of rice DREB2-type genes that encode transcription factors involved in the expression of abiotic stress-responsive genes. Mol. Genet. Genom. 2010, 283, 185–196. [Google Scholar] [CrossRef] [PubMed]
- Xue, G.P.; Loveridge, C.W. HvDRF1 is involved in abscisic acid-mediated gene regulation in barley and produces two forms of AP2 transcriptional activators, interacting preferably with a CT-rich element. Plant J. 2004, 37, 326–339. [Google Scholar] [CrossRef] [PubMed]
- Egawa, C.; Kobayashi, F.; Ishibashi, M.; Nakamura, T.; Nakamura, C.; Takumi, S. Differential regulation of transcript accumulation and alternative splicing of a DREB2 homolog under abiotic stress conditions in common wheat. Genes Genet. Syst. 2006, 81, 77–91. [Google Scholar] [CrossRef] [PubMed]
- Qin, F.; Kakimoto, M.; Sakuma, Y.; Maruyama, K.; Osakabe, Y.; Tran, L.S.; Shinozaki, K.; Yamaguchi-Shinozaki, K. Regulation and functional analysis of ZmDREB2A in response to drought and heat stresses in Zea mays L. Plant J. 2007, 50, 54–69. [Google Scholar] [CrossRef] [PubMed]
- Yan, K.; Liu, P.; Wu, C.A.; Yang, G.D.; Xu, R.; Guo, Q.H.; Huang, J.G.; Zheng, C.C. Stress-induced alternative splicing provides a mechanism for the regulation of microRNA processing in Arabidopsis thaliana. Mol. Cell 2012, 48, 521–531. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Ji, H.; Yuan, B.; Wang, S.; Su, C.; Yao, B.; Zhao, H.; Li, X. ABA signalling is fine-tuned by antagonistic HAB1 variants. Nat. Commun. 2015, 6, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Zhan, X.; Qian, B.; Cao, F.; Wu, W.; Yang, L.; Guan, Q.; Gu, X.; Wang, P.; Okusolubo, T.A.; Dunn, S.L.; et al. An Arabidopsis PWI and RRM motif-containing protein is critical for pre-mRNA splicing and ABA responses. Nat. Commun. 2015, 6, 8139. [Google Scholar] [CrossRef] [PubMed]
- Ali, G.S.; Palusa, S.G.; Golovkin, M.; Prasad, J.; Manley, J.L.; Reddy, A.S.N. Regulation of plant developmental processes by a novel splicing factor. PLoS ONE. 2007, 2, e471. [Google Scholar] [CrossRef] [PubMed]
- Carvalho, R.F.; Carvalho, S.D.; Duque, P. The plant specific SR45 protein negatively regulates glucose and ABA signaling during early seedling development in Arabidopsis. Plant Physiol. 2010, 154, 772–783. [Google Scholar] [CrossRef] [PubMed]
- Carvalho, R.F.; Szakonyi, D.; Simpson, C.G.; Barbosa, I.C.; Brown, J.W.; Baena-González, E.; Duque, P. The Arabidopsis SR45 splicing factor, a negative regulator of sugar signaling, modulates SNF1-related protein kinase 1 stability. Plant Cell 2016, 28, 1910–1925. [Google Scholar] [CrossRef]
- Palusa, S.G.; Ali, G.S.; Reddy, A.S. Alternative splicing of pre-mRNAs of Arabidopsis serine/arginine-rich proteins: Regulation by hormones and stresses. Plant J. 2007, 49, 1091–1107. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.N.; Mount, S.M. Two alternatively spliced isoforms of the Arabidopsis SR45 protein have distinct roles during normal plant development. Plant Physiol. 2009, 150, 1450–1458. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.H. Quorum quenching and proactive host defense. Trends Plant Sci. 2003, 8, 238–244. [Google Scholar] [CrossRef]
- Deyoung, B.J.; Innes, R.W. Plant NBS-LRR proteins in pathogen sensing and host defense. Nat. Immunol. 2006, 7, 1243–1249. [Google Scholar] [CrossRef] [PubMed]
- Staiger, D.; Brown, J.W.S. Alternative splicing at the intersection of biological timing, development, and stress responses. Plant Cell 2013, 25, 3640–3656. [Google Scholar] [CrossRef] [PubMed]
- Traut, T.W. The functions and consensus motifs of nine types of peptide segments that form different types of nucleotide-binding sites. Eur. J. Biochem. 1994, 222, 9–19. [Google Scholar] [CrossRef] [PubMed]
- Kobe, B.; Deisenhofer, J. The leucine-rich repeat: A versatile binding motif. Trends Biochem. Sci. 1994, 19, 415–421. [Google Scholar] [CrossRef]
- Ellis, J.G.; Lawrence, G.J.; Luck, J.E.; Dodds, P.N. Identification of regions in alleles of the flax rust resistance gene L that determine differences in gene-for-gene specificity. Plant Cell 1999, 11, 495–506. [Google Scholar] [CrossRef] [PubMed]
- Leister, R.T.; Katagiri, F. A resistance gene product of the nucleotide binding site—Leucine rich repeats class can form a complex with bacterial avirulence proteins in vivo. Plant J. 2000, 22, 345–354. [Google Scholar] [CrossRef] [PubMed]
- Jia, Y.; McAdams, S.A.; Bryan, G.T.; Hershey, H.P.; Valent, B. Direct interaction of resistance gene and avirulence gene productsconfers rice blast resistance. EMBO J. 2000, 19, 4004–4014. [Google Scholar] [CrossRef] [PubMed]
- Whitham, S.; Dinesh-Kumar, S.P.; Choi, D.; Hehl, R.; Corr, C.; Baker, B. The product of the tobacco mosaic virus resistance gene N: Similarity to toll and the interleukin-1 receptor. Cell 1994, 78, 1101–1115. [Google Scholar] [CrossRef]
- Kim, S.H.; Kwon, S.I.; Saha, D.; Anyanwu, N.C.; Gassmann, W. Resistance to the Pseudomonas syringae effector HopA1 is governed by the TIR-NBS-LRR protein RPS6 and is enhanced by mutations in SRFR1. Plant Physiol. 2009, 150, 1723–1732. [Google Scholar] [CrossRef] [PubMed]
- Borhan, M.H.; Holub, E.B.; Beynon, J.L.; Rozwadowski, K.; Rimmer, S.R. The Arabidopsis TIR-NB-LRR gene RAC1 confers resistance to Albugo candida (white rust) and is dependent on EDS1 but not PAD4. Mol. Plant Microbe Interact. 2004, 17, 711–719. [Google Scholar] [CrossRef] [PubMed]
- Parker, J.E.; Coleman, M.J.; Szabo, V.; Frost, L.N.; Schmidt, R.; van der Biezen, E.A.; Moores, T.; Dean, C.; Daniels, M.J.; Jones, J.D. The Arabidopsis downy mildew resistance gene RPP5 shares similarity to the toll and interleukin-1 receptors with N and L6. Plant Cell 1997, 9, 879–894. [Google Scholar] [CrossRef] [PubMed]
- Gassmann, W.; Hinsch, M.E.; Staskawicz, B.J. The Arabidopsis RPS4 bacterial-resistance gene is a member of the TIR-NBS-LRR family of disease-resistance genes. Plant J. 1999, 20, 265–277. [Google Scholar] [CrossRef] [PubMed]
- Yi, H.; Richards, E.J. A cluster of disease resistance genes in Arabidopsis is coordinately regulated by transcriptional activation and RNA silencing. Plant Cell 2007, 19, 2929–2939. [Google Scholar] [CrossRef] [PubMed]
- Schornack, S.; Ballvora, A.; Gurlebeck, D.; Peart, J.; Baulcombe, D.; Ganal, M.; Baker, B.; Bonas, U.; Lahaye, T. The tomato resistance protein Bs4 is a predicted non-nuclear TIR-NB-LRR protein that mediates defense responses to severely truncated derivatives of AvrBs4 and overexpressed AvrBs3. Plant J. 2004, 37, 46–60. [Google Scholar] [CrossRef] [PubMed]
- Dinesh-Kumar, S.P.; Baker, B.J. Alternatively spliced N resistance gene transcripts: Their possible role in tobacco mosaic virus resistance. Proc. Natl. Acad. Sci. USA 2000, 97, 1908–1913. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.C.; Gassmann, W. RPS4-mediated disease resistance requires the combined presence of RPS4 transcripts with full-length and truncated open reading frames. Plant Cell 2003, 15, 2333–2342. [Google Scholar] [CrossRef] [PubMed]
- Mastrangelo, A.M.; Marone, D.; Laido, G.; De Leonardis, A.M.; De Vita, P. Alternative splicing: Enhancing ability to cope with stress via transcriptome plasticity. Plant Sci. 2012, 185, 40–49. [Google Scholar] [CrossRef] [PubMed]
- Palma, K.; Zhao, Q.; Cheng, Y.T.; Bi, D.; Monaghan, J.; Cheng, W.; Zhang, Y.; Li, X. Regulation of plant innate immunity by three proteins in a complex conserved across the plant and animal kingdoms. Genes Dev. 2007, 21, 1484–1493. [Google Scholar] [CrossRef] [PubMed]
- Hogg, R.; McGrail, J.C.; Keefe, R.T. The function of the NineTeen Complex (NTC) in regulating spliceosome conformations and fidelity during pre-mRNA splicing. Biochem. Soc. Trans. 2010, 38, 1110–1115. [Google Scholar] [CrossRef] [PubMed]
- Xu, F.; Xu, S.; Wiermer, M.; Zhang, Y.; Li, X. The cyclin L homolog MOS12 and the MOS4-associated complex are required for the proper splicing of plant resistance genes. Plant J. 2012, 70, 916–928. [Google Scholar] [CrossRef] [PubMed]
- Monaghan, J.; Xu, F.; Xu, S.; Zhang, Y.; Li, X. Two putative RNA-binding proteins function with unequal genetic redundancy in the MOS4-associated complex. Plant Physiol. 2010, 154, 1783–1793. [Google Scholar] [CrossRef] [PubMed]
- Koncz, C.; Dejong, F.; Villacorta, N.; Szakonyi, D.; Koncz, Z. The spliceosome-activating complex: Molecular mechanisms underlying the function of a pleiotropic regulator. Front. Plant Sci. 2012, 3, 9. [Google Scholar] [CrossRef] [PubMed]
- He, Y.; Doyle, M.R.; Amasino, R.M. PAF1-complex mediated histone methylation of FLOWERING LOCUS C chromatin is required or the vernalization- responsive, winter-annual habit in Arabidopsis. Genes Dev. 2004, 18, 2774–2784. [Google Scholar] [CrossRef] [PubMed]
- Cao, Y.; Wen, L.; Wang, Z.; Ma, L. SKIP interacts with the Paf1 complex to regulate flowering via the activation of FLC, transcription in Arabidopsis. Mol. Plant 2015, 8, 1816–1819. [Google Scholar] [CrossRef] [PubMed]
- Kowalik, K.M.; Shimada, Y.; Flury, V.; Stadler, M.B.; Batki, J.; Buhler, M. The Paf1 complex represses small-RNA-mediated epigenetic gene silencing. Nature 2015, 520, 248–252. [Google Scholar] [CrossRef] [PubMed]
- Tomson, B.N.; Arndt, K.M. The many roles of the conserved eukaryotic Paf1 complex in regulating transcription, histone modifications, and disease states. Biochim. Biophys. Acta 2013, 1829, 116–126. [Google Scholar] [CrossRef] [PubMed]
- Slotkin, R.K.; Martienssen, R. Transposable elements and the epigenetic regulation of the genome. Nat. Rev. Genet. 2007, 8, 272–285. [Google Scholar] [CrossRef] [PubMed]
- Matzke, M.; Kanno, T.; Daxinger, L.; Huettel, B.; Matzke, A.J. RNA-mediated chromatin-based silencing in plants. Curr. Opin. Cell Biol. 2009, 21, 367–376. [Google Scholar] [CrossRef] [PubMed]
- Law, J.A.; Jacobsen, S.E. Establishing, maintaining and modifying DNA methylation patterns in plants and animals. Nat. Rev. Genet. 2010, 11, 204–220. [Google Scholar] [CrossRef] [PubMed]
- Gong, Z.; Morales-Ruiz, T.; Ariza, R.R.; Roldán-Arjona, T.; David, L.; Zhu, J.K. ROS1, a repressor of transcriptional gene silencing in Arabidopsis, encodes a DNA glycosylase/lyase. Cell 2002, 111, 803–814. [Google Scholar] [CrossRef]
- Liu, J.; Bai, G.; Zhang, C.; Chen, W.; Zhou, J.; Zhang, S.; Chen, Q.; Deng, X.; He, X.J.; Zhu, J.K. An atypical component of RNA-directed DNA methylation machinery has both DNA methylation-dependent and -independent roles in locus-specific transcriptional gene silencing. Cell Res. 2011, 21, 1691–1700. [Google Scholar] [CrossRef] [PubMed]
- Ausin, I.; Greenberg, M.V.; Li, C.F.; Jacobsen, S.E. The splicing factor SR45 affects the RNA-directed DNA methylation pathway in Arabidopsis. Epigenetics 2012, 7, 29–33. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.J.; Zhou, J.X.; Liu, J.; Ma, Z.Y.; Zhang, S.W.; Dou, K.; Huang, H.W.; Cai, T.; Liu, R.; Zhu, J.K.; et al. The splicing machinery promotes RNA-directed DNA methylation and transcriptional silencing in Arabidopsis. EMBO J. 2013, 32, 1128–1140. [Google Scholar] [CrossRef] [PubMed]
- Dou, K.; Huang, C.F.; Ma, Z.Y.; Zhang, C.J.; Zhou, J.X.; Huang, H.W.; Cai, T.; Tang, K.; Zhu, J.K.; He, X.J. The PRP6-like splicing factor STA1 is involved in RNA-directed DNA methylation by facilitating the production of Pol V-dependent scaffold RNAs. Nucleic Acids Res. 2013, 41, 8489–8502. [Google Scholar] [CrossRef] [PubMed]
- Du, J.L.; Zhang, S.W.; Huang, H.W.; Cai, T.; Li, L.; Chen, S.; He, X.J. The splicing factor PRP31 is involved in transcriptional gene silencing and stress response in Arabidopsis. Mol. Plant 2015, 8, 1053–1068. [Google Scholar] [CrossRef] [PubMed]
- Huang, C.F.; Miki, D.; Tang, K.; Zhou, H.R.; Zheng, Z.; Chen, W.; Ma, Z.Y.; Yang, L.; Zhang, H.; Liu, R.; et al. A Pre-mRNA splicing factor is required for RNA-directed DNA methylation in Arabidopsis. PLoS Genet. 2013, 9, e1003779. [Google Scholar] [CrossRef] [PubMed]
- Golovkin, M.; Reddy, A.S. An SC35-like protein and a novel serine/arginine-rich protein interact with Arabidopsis U1–70K protein. J. Biol. Chem. 1999, 274, 36428–36438. [Google Scholar] [CrossRef] [PubMed]
- Zhuang, Y.; Weiner, A.M. A compensatory base change in U1 snRNA suppresses a 5′ splice site mutation. Cell 1986, 46, 827–835. [Google Scholar] [CrossRef]
- Kohtz, J.D.; Jamison, S.F.; Will, C.L.; Zuo, P.; Lührmann, R.; Garcia-Blanco, M.A.; Manley, J.L. Protein-protein interactions and 5′-splice-site recognition in mammalian mRNA precursors. Nature 1994, 368, 119–124. [Google Scholar] [CrossRef] [PubMed]
- Wu, S.; Romfo, C.M.; Nilsen, T.W.; Green, M.R. Functional recognition of the 3′ splice site AG by the splicing factor U2A.F.35. Nature 1999, 402, 832–835. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.N.; Mo, C.; Garrett, W.M.; Cooper, B. Phosphothreonine 218 is required for the function of SR45.1 in regulating flower petal development in Arabidopsis. Plant Signal. Behav. 2014, 9, e29134. [Google Scholar] [CrossRef] [PubMed]
- Stankovic, N.; Schloesser, M.; Joris, M.; Sauvage, E.; Hanikenne, M.; Motte, P. Dynamic distribution and interaction of the Arabidopsis SRSF1 subfamily splicing factors. Plant Physiol. 2016, 170, 1000–1013. [Google Scholar] [CrossRef] [PubMed]
- Feng, J.; Li, J.; Gao, Z.; Lu, Y.; Yu, J.; Zheng, Q.; Yan, S.; Zhang, W.; He, H.; Ma, L.; et al. SKIP confers osmotic tolerance during salt stress by controlling alternative gene splicing in Arabidopsis. Mol. Plant 2015, 8, 1038–1052. [Google Scholar] [CrossRef] [PubMed]
- Baldwin, K.L.; Dinh, E.M.; Hart, B.M.; Masson, P.H. CACTIN is an essential nuclear protein in Arabidopsis and may be associated with the eukaryotic spliceosome. FEBS Lett. 2013, 587, 873–879. [Google Scholar] [CrossRef] [PubMed]
- Schwerk, C.; Prasad, J.; Degenhardt, K.; Erdjument-Bromage, H.; White, E.; Tempst, P.; Kidd, V.J.; Manley, J.L.; Lahti, J.M.; Reinberg, D. Asap, a novel protein complex involved in RNA processing and apoptosis. Mol. Cell. Biol. 2003, 23, 2981–2990. [Google Scholar] [CrossRef] [PubMed]
- Qüesta, J.I.; Song, J.; Geraldo, N.; An, H.; Dean, C. Arabidopsis transcriptional repressor val1 triggers polycomb silencing at FLC during vernalization. Science 2016, 353, 485–488. [Google Scholar] [CrossRef] [PubMed]
- Xing, D.; Wang, Y.; Hamilton, M.; Benhur, A.; Reddy, A.S. Transcriptome-wide identification of RNA targets of Arabidopsis serine/arginine-rich45 uncovers the unexpected roles of this RNA binding protein in RNA processing. Plant Cell 2015, 27, 3294. [Google Scholar] [CrossRef] [PubMed]
- Reddy, A.S. Alternative splicing of pre-messenger RNAs in plants in the genomic era. Annu. Rev. Plant Biol. 2007, 58, 267–294. [Google Scholar] [CrossRef] [PubMed]


© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license ( http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Shang, X.; Cao, Y.; Ma, L. Alternative Splicing in Plant Genes: A Means of Regulating the Environmental Fitness of Plants. Int. J. Mol. Sci. 2017, 18, 432. https://doi.org/10.3390/ijms18020432
Shang X, Cao Y, Ma L. Alternative Splicing in Plant Genes: A Means of Regulating the Environmental Fitness of Plants. International Journal of Molecular Sciences. 2017; 18(2):432. https://doi.org/10.3390/ijms18020432
Chicago/Turabian StyleShang, Xudong, Ying Cao, and Ligeng Ma. 2017. "Alternative Splicing in Plant Genes: A Means of Regulating the Environmental Fitness of Plants" International Journal of Molecular Sciences 18, no. 2: 432. https://doi.org/10.3390/ijms18020432
APA StyleShang, X., Cao, Y., & Ma, L. (2017). Alternative Splicing in Plant Genes: A Means of Regulating the Environmental Fitness of Plants. International Journal of Molecular Sciences, 18(2), 432. https://doi.org/10.3390/ijms18020432