Bacterial Responses to Glyoxal and Methylglyoxal: Reactive Electrophilic Species
Abstract
:1. Introduction
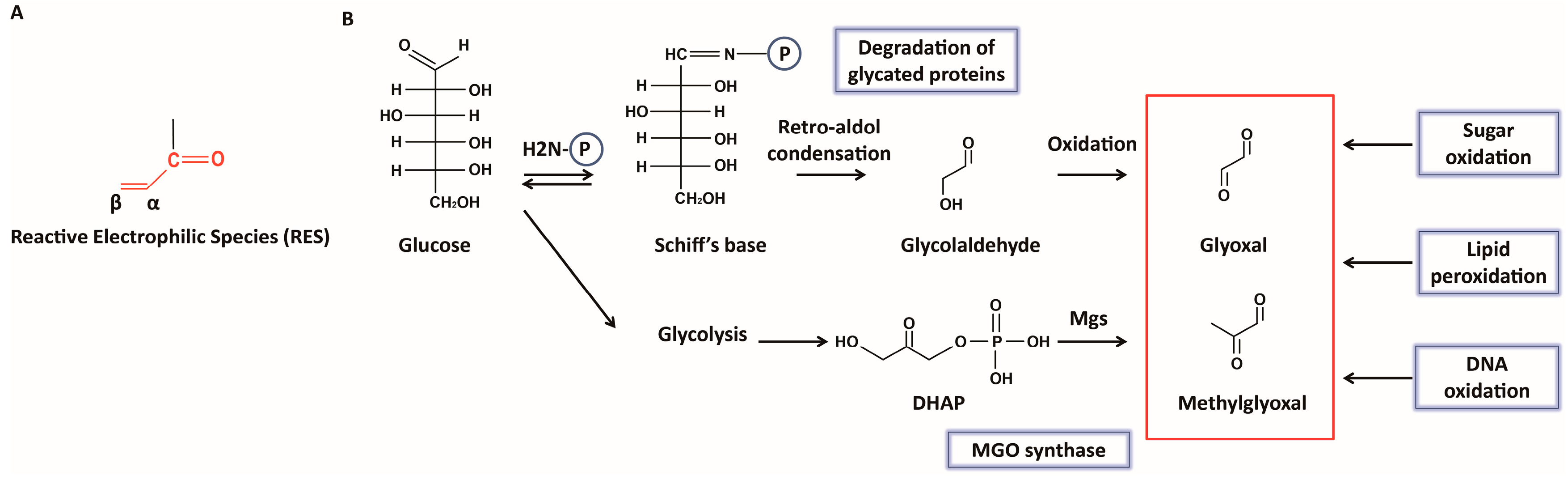
2. Formation of Glyoxals
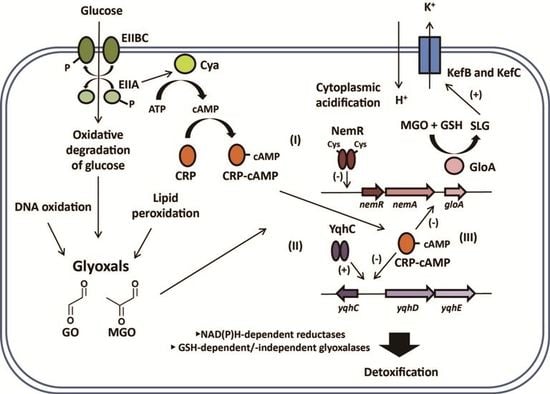
3. Bacterial Sensing of Glyoxals
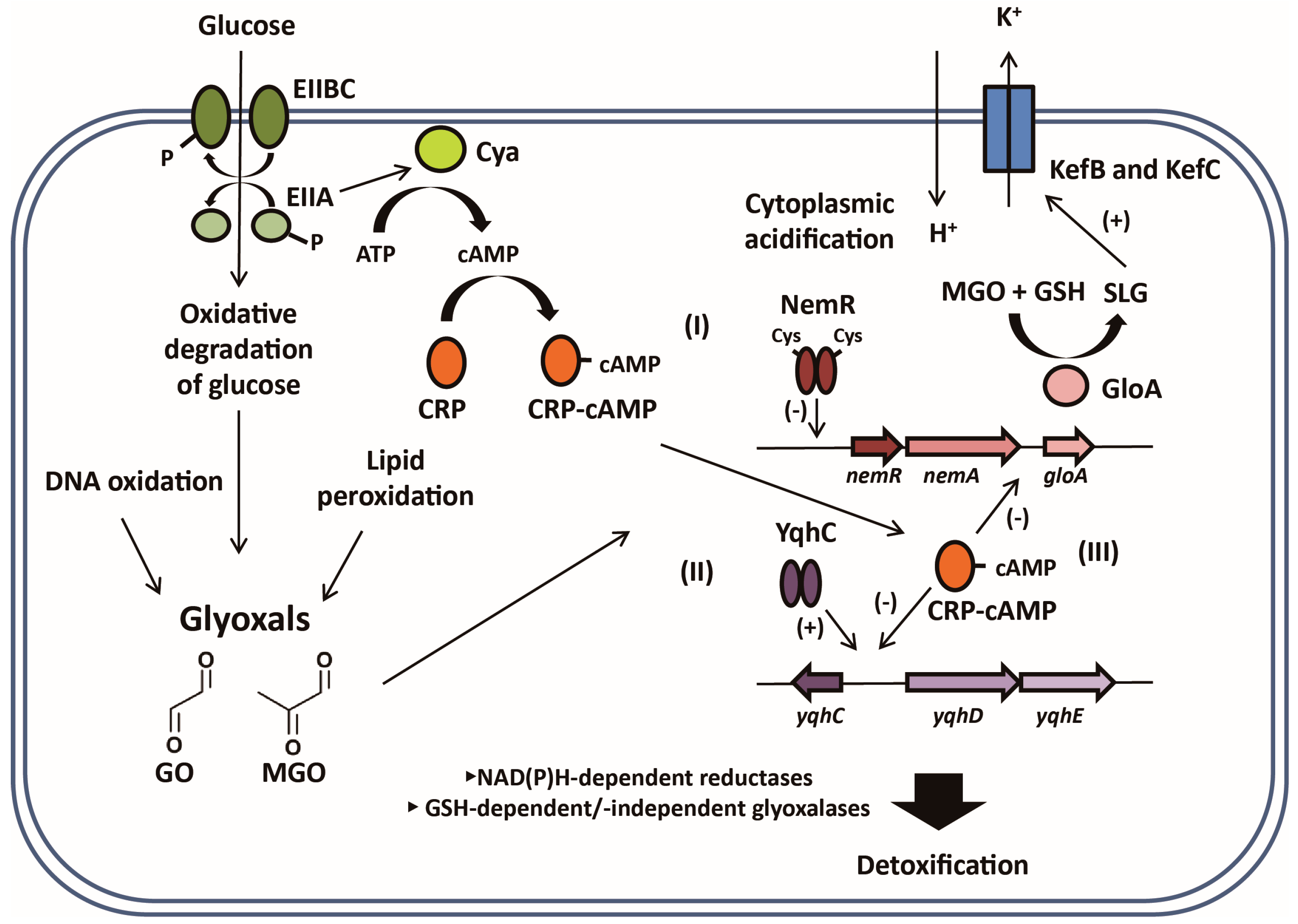
3.1. YqhC, a Major Regulator for Glyoxal (GO)-Detoxifying YqhDE
3.2. NemR, a Potential Reactive Electrophilic Species (RES) Sensor for Regulating the nemRA-gloA Operon
3.3. CRP/cAMP-Mediated Detoxification Pathway for Glyoxal
3.4. Fnr and NsrR—Regulators of YafB Aldo-Keto Reductase
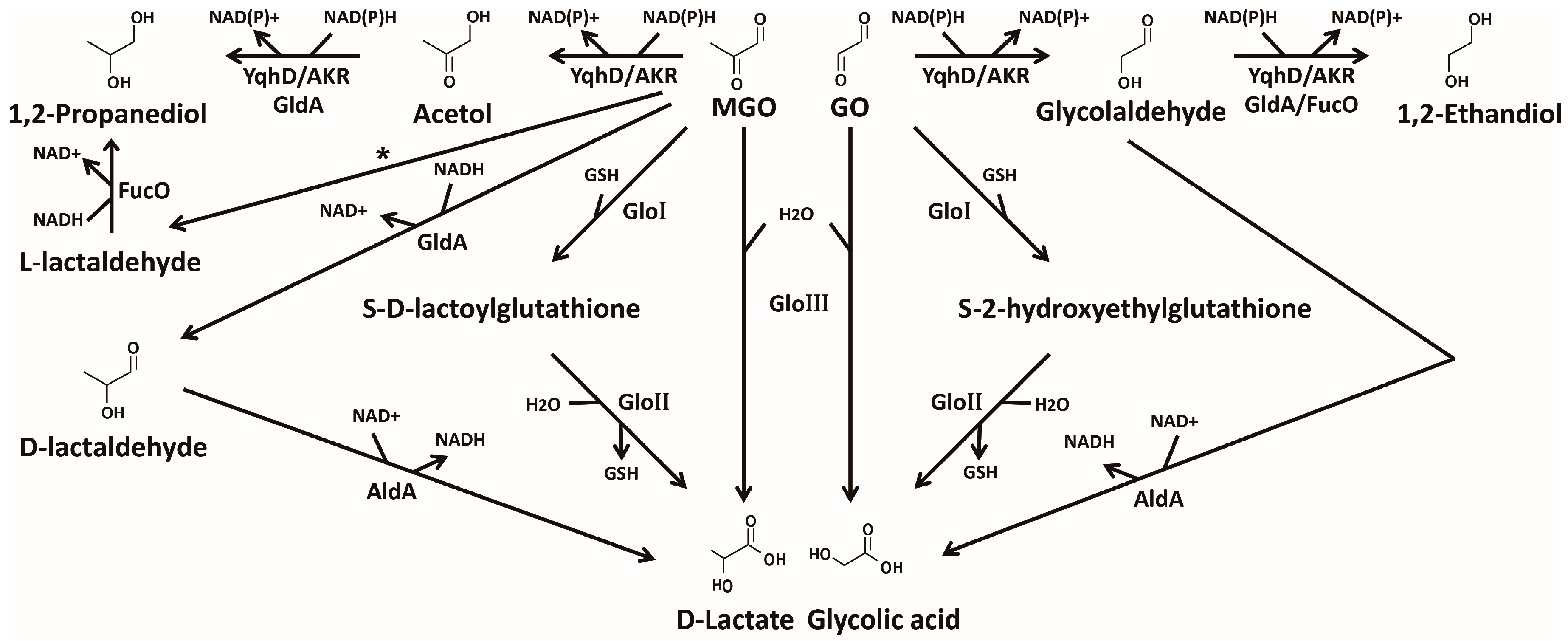
4. Detoxification of Glyoxals
4.1. GSH-Dependent Glyoxalase System
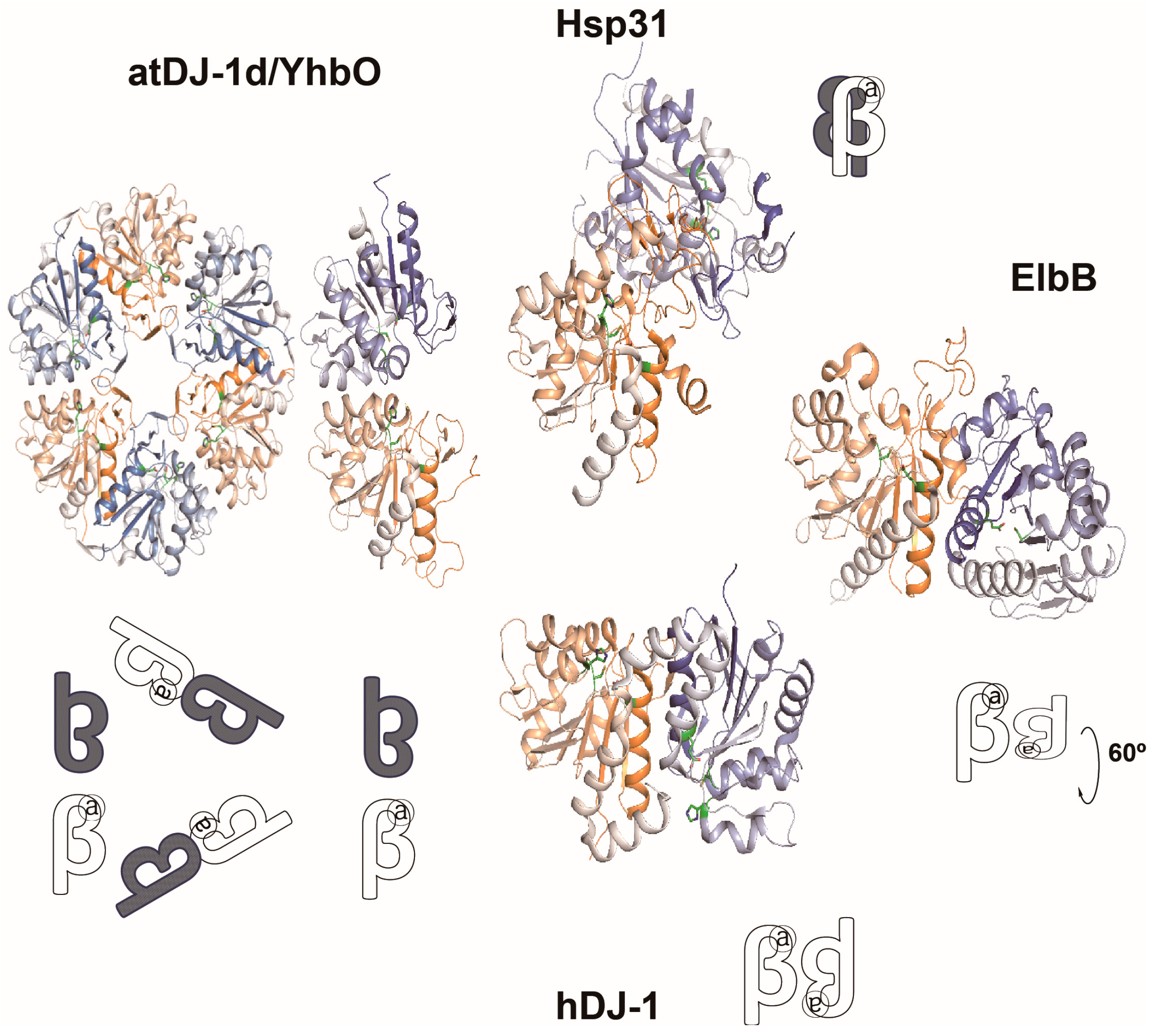
4.2. GSH-Independent Glyoxalase System
4.3. NAD(P)H-Dependent Detoxifying Enzymes
5. Toxicity of Glyoxals
5.1. Protein Damage
5.2. Nucleotide Damage
5.3. Relation to Oxidative Stress
6. Conclusions and Perspectives
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Farmer, E.E.; Davoine, C. Reactive electrophile species. Curr. Opin. Plant Biol. 2007, 10, 380–386. [Google Scholar] [CrossRef] [PubMed]
- Jacobs, A.T.; Marnett, L.J. Systems analysis of protein modification and cellular responses induced by electrophile stress. Acc. Chem. Res. 2010, 43, 673–683. [Google Scholar] [CrossRef] [PubMed]
- Marnett, L.J.; Riggins, J.N.; West, J.D. Endogenous generation of reactive oxidants and electrophiles and their reactions with DNA and protein. J. Clin. Investig. 2003, 111, 583–593. [Google Scholar] [CrossRef] [PubMed]
- Munch, G.; Luth, H.J.; Wong, A.; Arendt, T.; Hirsch, E.; Ravid, R.; Riederer, P. Crosslinking of α-synuclein by advanced glycation endproducts—An early pathophysiological step in Lewy body formation? J. Chem. Neuroanat. 2000, 20, 253–257. [Google Scholar] [CrossRef]
- Brownlee, M. Advanced protein glycosylation in diabetes and aging. Annu. Rev. Med. 1995, 46, 223–234. [Google Scholar] [CrossRef] [PubMed]
- Luth, H.J.; Ogunlade, V.; Kuhla, B.; Kientsch-Engel, R.; Stahl, P.; Webster, J.; Arendt, T.; Munch, G. Age- and stage-dependent accumulation of advanced glycation end products in intracellular deposits in normal and Alzheimer’s disease brains. Cereb. Cortex 2005, 15, 211–220. [Google Scholar] [CrossRef] [PubMed]
- Thornalley, P.J. The glyoxalase system: New developments towards functional characterization of a metabolic pathway fundamental to biological life. Biochem. J. 1990, 269, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Thornalley, P.J.; Langborg, A.; Minhas, H.S. Formation of glyoxal, methylglyoxal and 3-deoxyglucosone in the glycation of proteins by glucose. Biochem. J. 1999, 344 Pt 1, 109–116. [Google Scholar] [CrossRef] [PubMed]
- Hopper, D.J.; Cooper, R.A. The regulation of Escherichia coli methylglyoxal synthase; a new control site in glycolysis? FEBS Lett. 1971, 13, 213–216. [Google Scholar] [CrossRef]
- Singh, R.; Barden, A.; Mori, T.; Beilin, L. Advanced glycation end-products: A review. Diabetologia 2001, 44, 129–146. [Google Scholar] [CrossRef] [PubMed]
- Thornalley, P.J. Pharmacology of methylglyoxal: Formation, modification of proteins and nucleic acids, and enzymatic detoxification—A role in pathogenesis and antiproliferative chemotherapy. Gen. Pharmacol. 1996, 27, 565–573. [Google Scholar] [CrossRef]
- Subedi, K.P.; Kim, I.; Kim, J.; Min, B.; Park, C. Role of GldA in dihydroxyacetone and methylglyoxal metabolism of Escherichia coli K12. FEMS Microbiol. Lett. 2008, 279, 180–187. [Google Scholar] [CrossRef] [PubMed]
- Kim, I.; Kim, E.; Yoo, S.; Shin, D.; Min, B.; Song, J.; Park, C. Ribose utilization with an excess of mutarotase causes cell death due to accumulation of methylglyoxal. J. Bacteriol. 2004, 186, 7229–7235. [Google Scholar] [CrossRef] [PubMed]
- Lee, C.; Kim, I.; Lee, J.; Lee, K.L.; Min, B.; Park, C. Transcriptional activation of the aldehyde reductase YqhD by YqhC and its implication in glyoxal metabolism of Escherichia coli K-12. J. Bacteriol. 2010, 192, 4205–4214. [Google Scholar] [CrossRef] [PubMed]
- Lee, C.; Shin, J.; Park, C. Novel regulatory system nemRA-gloA for electrophile reduction in Escherichia coli K-12. Mol. Microbiol. 2013, 88, 395–412. [Google Scholar] [CrossRef] [PubMed]
- Turner, P.C.; Miller, E.N.; Jarboe, L.R.; Baggett, C.L.; Shanmugam, K.T.; Ingram, L.O. YqhC regulates transcription of the adjacent Escherichia coli genes yqhD and dkgA that are involved in furfural tolerance. J. Ind. Microbiol. Biotechnol. 2011, 38, 431–439. [Google Scholar] [CrossRef] [PubMed]
- Umezawa, Y.; Shimada, T.; Kori, A.; Yamada, K.; Ishihama, A. The uncharacterized transcription factor YdhM is the regulator of the nemA gene, encoding N-ethylmaleimide reductase. J. Bacteriol. 2008, 190, 5890–5897. [Google Scholar] [CrossRef] [PubMed]
- Ozyamak, E.; de Almeida, C.; de Moura, A.P.; Miller, S.; Booth, I.R. Integrated stress response of Escherichia coli to methylglyoxal: Transcriptional readthrough from the nemRA operon enhances protection through increased expression of glyoxalase I. Mol. Microbiol. 2013, 88, 936–950. [Google Scholar] [CrossRef] [PubMed]
- Miura, K.; Tomioka, Y.; Suzuki, H.; Yonezawa, M.; Hishinuma, T.; Mizugaki, M. Molecular cloning of the nemA gene encoding N-ethylmaleimide reductase from Escherichia coli. Biol. Pharm. Bull. 1997, 20, 110–112. [Google Scholar] [CrossRef] [PubMed]
- Williams, R.E.; Bruce, N.C. “New uses for an Old Enzyme”—The Old Yellow Enzyme family of flavoenzymes. Microbiology 2002, 148 Pt 6, 1607–1614. [Google Scholar] [CrossRef] [PubMed]
- Gray, M.J.; Wholey, W.Y.; Parker, B.W.; Kim, M.; Jakob, U. NemR is a bleach-sensing transcription factor. J. Biol. Chem. 2013, 288, 13789–13798. [Google Scholar] [CrossRef] [PubMed]
- Gray, M.J.; Wholey, W.Y.; Jakob, U. Bacterial responses to reactive chlorine species. Annu. Rev. Microbiol. 2013, 67, 141–160. [Google Scholar] [CrossRef] [PubMed]
- Gray, M.J.; Li, Y.; Leichert, L.I.; Xu, Z.; Jakob, U. Does the Transcription Factor NemR Use a Regulatory Sulfenamide Bond to Sense Bleach? Antioxid. Redox Signal. 2015, 23, 747–754. [Google Scholar] [CrossRef] [PubMed]
- Zheng, D.; Constantinidou, C.; Hobman, J.L.; Minchin, S.D. Identification of the CRP regulon using in vitro and in vivo transcriptional profiling. Nucleic Acids Res. 2004, 32, 5874–5893. [Google Scholar] [CrossRef] [PubMed]
- Lee, C.; Kim, J.; Kwon, M.; Lee, K.; Min, H.; Kim, S.H.; Kim, D.; Lee, N.; Kim, J.; Kim, D.; et al. Screening for Escherichia coli K-12 genes conferring glyoxal resistance or sensitivity by transposon insertions. FEMS Microbiol. Lett. 2016, 363. [Google Scholar] [CrossRef] [PubMed]
- Deutscher, J. The mechanisms of carbon catabolite repression in bacteria. Curr. Opin. Microbiol. 2008, 11, 87–93. [Google Scholar] [CrossRef] [PubMed]
- Ko, J.; Kim, I.; Yoo, S.; Min, B.; Kim, K.; Park, C. Conversion of methylglyoxal to acetol by Escherichia coli aldo-keto reductases. J. Bacteriol. 2005, 187, 5782–5789. [Google Scholar] [CrossRef] [PubMed]
- Lee, C.; Kim, I.; Park, C. Glyoxal detoxification in Escherichia coli K-12 by NADPH dependent aldo-keto reductases. J. Microbiol. 2013, 51, 527–530. [Google Scholar] [CrossRef] [PubMed]
- Kwon, M.; Lee, J.; Lee, C.; Park, C. Genomic rearrangements leading to overexpression of aldo-keto reductase YafB of Escherichia coli confer resistance to glyoxal. J. Bacteriol. 2012, 194, 1979–1988. [Google Scholar] [CrossRef] [PubMed]
- Beaumont, H.J.; Lens, S.I.; Reijnders, W.N.; Westerhoff, H.V.; van Spanning, R.J. Expression of nitrite reductase in Nitrosomonas europaea involves NsrR, a novel nitrite-sensitive transcription repressor. Mol. Microbiol. 2004, 54, 148–158. [Google Scholar] [CrossRef] [PubMed]
- Salmon, K.; Hung, S.P.; Mekjian, K.; Baldi, P.; Hatfield, G.W.; Gunsalus, R.P. Global gene expression profiling in Escherichia coli K12. The effects of oxygen availability and FNR. J. Biol. Chem. 2003, 278, 29837–29855. [Google Scholar] [CrossRef] [PubMed]
- Kang, Y.; Weber, K.D.; Qiu, Y.; Kiley, P.J.; Blattner, F.R. Genome-wide expression analysis indicates that FNR of Escherichia coli K-12 regulates a large number of genes of unknown function. J. Bacteriol. 2005, 187, 1135–1160. [Google Scholar] [CrossRef] [PubMed]
- Subedi, K.P.; Choi, D.; Kim, I.; Min, B.; Park, C. Hsp31 of Escherichia coli K-12 is glyoxalase III. Mol. Microbiol. 2011, 81, 926–936. [Google Scholar] [CrossRef] [PubMed]
- Lee, C.; Lee, J.; Lee, J.Y.; Park, C. Characterization of the Escherichia coli YajL, YhbO and ElbB glyoxalases. FEMS Microbiol. Lett. 2016, 363. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.M.; Zhu, Y.; Lin, E.C. NAD-linked aldehyde dehydrogenase for aerobic utilization of l-fucose and l-rhamnose by Escherichia coli. J. Bacteriol. 1987, 169, 3289–3294. [Google Scholar] [CrossRef] [PubMed]
- Baldoma, L.; Aguilar, J. Involvement of lactaldehyde dehydrogenase in several metabolic pathways of Escherichia coli K12. J. Biol. Chem. 1987, 262, 13991–13996. [Google Scholar] [PubMed]
- Altaras, N.E.; Cameron, D.C. Metabolic engineering of a 1,2-propanediol pathway in Escherichia coli. Appl. Environ. Microbiol. 1999, 65, 1180–1185. [Google Scholar] [PubMed]
- MacLean, M.J.; Ness, L.S.; Ferguson, G.P.; Booth, I.R. The role of glyoxalase I in the detoxification of methylglyoxal and in the activation of the KefB K+ efflux system in Escherichia coli. Mol. Microbiol. 1998, 27, 563–571. [Google Scholar] [CrossRef] [PubMed]
- Ferguson, G.P.; Battista, J.R.; Lee, A.T.; Booth, I.R. Protection of the DNA during the exposure of Escherichia coli cells to a toxic metabolite: The role of the KefB and KefC potassium channels. Mol. Microbiol. 2000, 35, 113–122. [Google Scholar] [CrossRef] [PubMed]
- Healy, J.; Ekkerman, S.; Pliotas, C.; Richard, M.; Bartlett, W.; Grayer, S.C.; Morris, G.M.; Miller, S.; Booth, I.R.; Conway, S.J.; et al. Understanding the structural requirements for activators of the Kef bacterial potassium efflux system. Biochemistry 2014, 53, 1982–1992. [Google Scholar] [CrossRef] [PubMed]
- Chandrangsu, P.; Dusi, R.; Hamilton, C.J.; Helmann, J.D. Methylglyoxal resistance in Bacillus subtilis: Contributions of bacillithiol-dependent and independent pathways. Mol. Microbiol. 2014, 91, 706–715. [Google Scholar] [CrossRef] [PubMed]
- Misra, K.; Banerjee, A.B.; Ray, S.; Ray, M. Glyoxalase III from Escherichia coli: A single novel enzyme for the conversion of methylglyoxal into d-lactate without reduced glutathione. Biochem. J. 1995, 305 Pt 3, 999–1003. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.Y.; Song, J.; Kwon, K.; Jang, S.; Kim, C.; Baek, K.; Kim, J.; Park, C. Human DJ-1 and its homologs are novel glyoxalases. Hum. Mol. Genet. 2012, 21, 3215–3225. [Google Scholar] [CrossRef] [PubMed]
- Choi, D.; Kim, J.; Ha, S.; Kwon, K.; Kim, E.H.; Lee, H.Y.; Ryu, K.S.; Park, C. Stereospecific mechanism of DJ-1 glyoxalases inferred from their hemithioacetal-containing crystal structures. FEBS J. 2014, 281, 5447–5462. [Google Scholar] [CrossRef] [PubMed]
- Kwon, K.; Choi, D.; Hyun, J.K.; Jung, H.S.; Baek, K.; Park, C. Novel glyoxalases from Arabidopsis thaliana. FEBS J. 2013, 280, 3328–3339. [Google Scholar] [CrossRef] [PubMed]
- Abdallah, J.; Kern, R.; Malki, A.; Eckey, V.; Richarme, G. Cloning, expression, and purification of the general stress protein YhbO from Escherichia coli. Protein Expr. Purif. 2006, 47, 455–460. [Google Scholar] [CrossRef] [PubMed]
- Richarme, G.; Mihoub, M.; Dairou, J.; Bui, L.C.; Leger, T.; Lamouri, A. Parkinsonism-associated protein DJ-1/Park7 is a major protein deglycase that repairs methylglyoxal- and glyoxal-glycated cysteine, arginine, and lysine residues. J. Biol. Chem. 2015, 290, 1885–1897. [Google Scholar] [CrossRef] [PubMed]
- Richarme, G.; Marguet, E.; Forterre, P.; Ishino, S.; Ishino, Y. DJ-1 family Maillard deglycases prevent acrylamide formation. Biochem. Biophys. Res. Commun. 2016, 478, 1111–1116. [Google Scholar] [CrossRef] [PubMed]
- Jarboe, L.R. YqhD: A broad-substrate range aldehyde reductase with various applications in production of biorenewable fuels and chemicals. Appl. Microbiol. Biotechnol. 2011, 89, 249–257. [Google Scholar] [CrossRef] [PubMed]
- Boronat, A.; Caballero, E.; Aguilar, J. Experimental evolution of a metabolic pathway for ethylene glycol utilization by Escherichia coli. J. Bacteriol. 1983, 153, 134–139. [Google Scholar] [PubMed]
- Basta, G.; Schmidt, A.M.; de Caterina, R. Advanced glycation end products and vascular inflammation: Implications for accelerated atherosclerosis in diabetes. Cardiovasc. Res. 2004, 63, 582–592. [Google Scholar] [CrossRef] [PubMed]
- Sousa Silva, M.; Gomes, R.A.; Ferreira, A.E.; Ponces Freire, A.; Cordeiro, C. The glyoxalase pathway: The first hundred years... and beyond. Biochem. J. 2013, 453, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Kasai, H.; Iwamoto-Tanaka, N.; Fukada, S. DNA modifications by the mutagen glyoxal: Adduction to G and C, deamination of C and GC and GA cross-linking. Carcinogenesis 1998, 19, 1459–1465. [Google Scholar] [CrossRef] [PubMed]
- Murata-Kamiya, N.; Kamiya, H.; Kaji, H.; Kasai, H. Mutational specificity of glyoxal, a product of DNA oxidation, in the lacI gene of wild-type Escherichia coli W3110. Mutat. Res. 1997, 377, 255–262. [Google Scholar] [CrossRef]
- Courcelle, J.; Hanawalt, P.C. RecA-dependent recovery of arrested DNA replication forks. Annu. Rev. Genet. 2003, 37, 611–646. [Google Scholar] [CrossRef] [PubMed]
- Urbonavicius, J.; Qian, Q.; Durand, J.M.; Hagervall, T.G.; Bjork, G.R. Improvement of reading frame maintenance is a common function for several tRNA modifications. EMBO J. 2001, 20, 4863–4873. [Google Scholar] [CrossRef] [PubMed]
- Kurata, S.; Weixlbaumer, A.; Ohtsuki, T.; Shimazaki, T.; Wada, T.; Kirino, Y.; Takai, K.; Watanabe, K.; Ramakrishnan, V.; Suzuki, T. Modified uridines with C5-methylene substituents at the first position of the tRNA anticodon stabilize U.G wobble pairing during decoding. J. Biol. Chem. 2008, 283, 18801–18811. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Liu, J.; Wu, L. Methylglyoxal-induced mitochondrial dysfunction in vascular smooth muscle cells. Biochem. Pharmacol. 2009, 77, 1709–1716. [Google Scholar] [CrossRef] [PubMed]
- Benov, L.; Fridovich, I. Induction of the soxRS regulon of Escherichia coli by glycolaldehyde. Arch. Biochem. Biophys. 2002, 407, 45–48. [Google Scholar] [CrossRef]
- Okado-Matsumoto, A.; Fridovich, I. The role of α,β-dicarbonyl compounds in the toxicity of short chain sugars. J. Biol. Chem. 2000, 275, 34853–34857. [Google Scholar] [CrossRef] [PubMed]
- Bankapalli, K.; Saladi, S.; Awadia, S.S.; Goswami, A.V.; Samaddar, M.; D’Silva, P. Robust glyoxalase activity of Hsp31, a ThiJ/DJ-1/PfpI family member protein, is critical for oxidative stress resistance in Saccharomyces cerevisiae. J. Biol. Chem. 2015, 290, 26491–26507. [Google Scholar] [CrossRef] [PubMed]
- Brouwers, O.; Niessen, P.M.; Ferreira, I.; Miyata, T.; Scheffer, P.G.; Teerlink, T.; Schrauwen, P.; Brownlee, M.; Stehouwer, C.D.; Schalkwijk, C.G. Overexpression of glyoxalase-I reduces hyperglycemia-induced levels of advanced glycation end products and oxidative stress in diabetic rats. J. Biol. Chem. 2011, 286, 1374–1380. [Google Scholar] [CrossRef] [PubMed]
| Proteins | Substrate | Product | Km (mM) | kcat (min−1) | kcat/Km (min−1·M−1) | Reference |
|---|---|---|---|---|---|---|
| YqhD | GO | Glycolaldehyde | 11.53 | 618 | 5.36 × 104 | [14] |
| Glycolaldehyde | 1,2-Ethandiol | 28.28 | 3258 | 1.09 × 105 | [14] | |
| MGO | Acetol | 2.6 | 284 | 1.09 × 105 | [14] | |
| Acetol | 1,2-Propanediol | 76.9 | 144 | 1.87 × 103 | [14] | |
| YqhE | GO | Glycolaldehyde | 22 | 296 | 1.34 × 104 | [28] |
| Glycolaldehyde | 1,2-Ethandiol | 12 | 15 | 1.25 × 103 | [28] | |
| MGO | Acetol | 2.05 | 1657 | 8.09 × 109 | [27] | |
| YafB | GO | Glycolaldehyde | 60 | 368 | 6.13 × 103 | [28] |
| Glycolaldehyde | 1,2-Ethandiol | 17 | 22 | 1.29 × 103 | [28] | |
| MGO | Acetol | 2.46 | 1749 | 7.13 × 105 | [27] | |
| YghZ | GO | Glycolaldehyde | 104 | 458 | 4.40 × 103 | [28] |
| Glycolaldehyde | 1,2-Ethandiol | 104 | 140 | 1.34 × 103 | [28] | |
| MGO | Acetol | 6.91 | 661 | 0.96 × 105 | [27] | |
| YeaE | GO | Glycolaldehyde | 10 | 15 | 1.50 × 103 | [28] |
| Glycolaldehyde | 1,2-Ethandiol | 7 | 15 | 2.14 × 103 | [28] | |
| MGO | Acetol | 2.09 | 171 | 0.82 × 105 | [27] | |
| YajO | GO | Glycolaldehyde | N.D. | N.D. | N.D. | [28] |
| Glycolaldehyde | 1,2-Ethandiol | N.D. | N.D. | N.D. | [28] | |
| MGO | Acetol | N.D. | N.D. | N.D. | [27] | |
| NemA | GO | Glycolaldehyde | 318.88 | 1008 | 3.15 × 103 | [15] |
| MGO | Acetol | 25.63 | 1276 | 4.98 × 104 | [15] | |
| Hsp31 | GO | Glycolaldehyde | 5.94 | 16 | 2.78 × 103 | [33] |
| MGO | Acetol | 1.43 | 156 | 1.09 × 105 | [33] | |
| YhbO | GO | Glycolaldehyde | 2.97 | 70 | 3.11 × 105 | [34] |
| MGO | Acetol | 0.06 | 20.8 | 3.47 × 105 | [34] | |
| YajL | GO | Glycolaldehyde | 2.97 | 70 | 0.24 × 105 | [34] |
| MGO | Acetol | N.D. | N.D. | N.D. | [34] | |
| ElbB | GO | Glycolaldehyde | 1.21 | 0.48 | 0.40 × 103 | [34] |
| MGO | Acetol | N.D. | N.D. | N.D. | [34] | |
| GldA | MGO | d-lactaldehyde | 0.5 | 0.1 | 2.03 × 102 | [12] |
| Glycolaldehyde | 1,2-Ethandiol | 0.85 | 0.53 | 6.23 × 102 | [12] | |
| Acetol a | 1,2-Propanediol | N.D. | N.D. | N.D. | [12] |
© 2017 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lee, C.; Park, C. Bacterial Responses to Glyoxal and Methylglyoxal: Reactive Electrophilic Species. Int. J. Mol. Sci. 2017, 18, 169. https://doi.org/10.3390/ijms18010169
Lee C, Park C. Bacterial Responses to Glyoxal and Methylglyoxal: Reactive Electrophilic Species. International Journal of Molecular Sciences. 2017; 18(1):169. https://doi.org/10.3390/ijms18010169
Chicago/Turabian StyleLee, Changhan, and Chankyu Park. 2017. "Bacterial Responses to Glyoxal and Methylglyoxal: Reactive Electrophilic Species" International Journal of Molecular Sciences 18, no. 1: 169. https://doi.org/10.3390/ijms18010169
APA StyleLee, C., & Park, C. (2017). Bacterial Responses to Glyoxal and Methylglyoxal: Reactive Electrophilic Species. International Journal of Molecular Sciences, 18(1), 169. https://doi.org/10.3390/ijms18010169