Droplet Digital PCR Based Androgen Receptor Variant 7 (AR-V7) Detection from Prostate Cancer Patient Blood Biopsies
Abstract
:1. Introduction
2. Results and Discussion
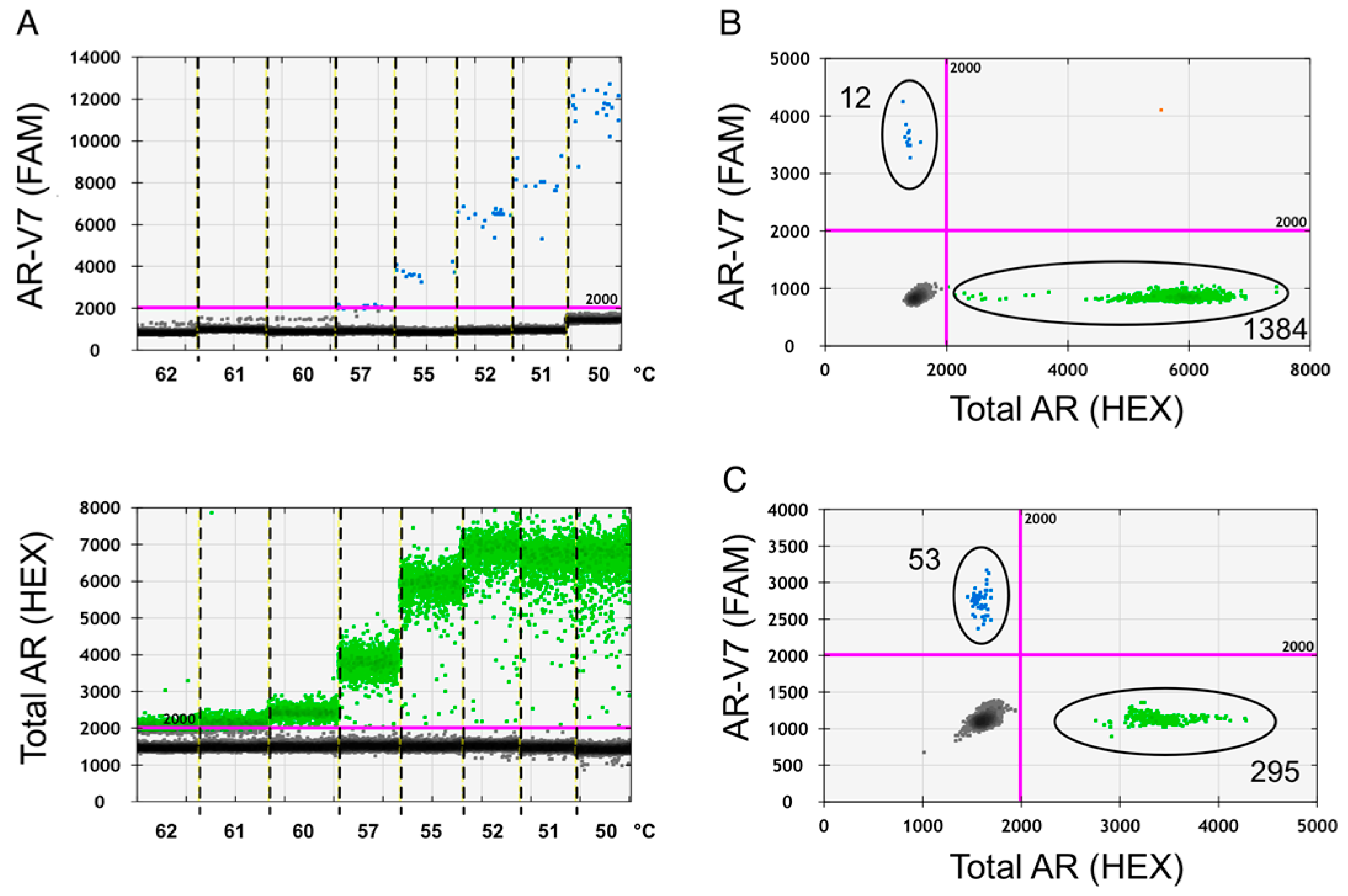
2.1. Assay Optimization
2.2. Assay Specificity
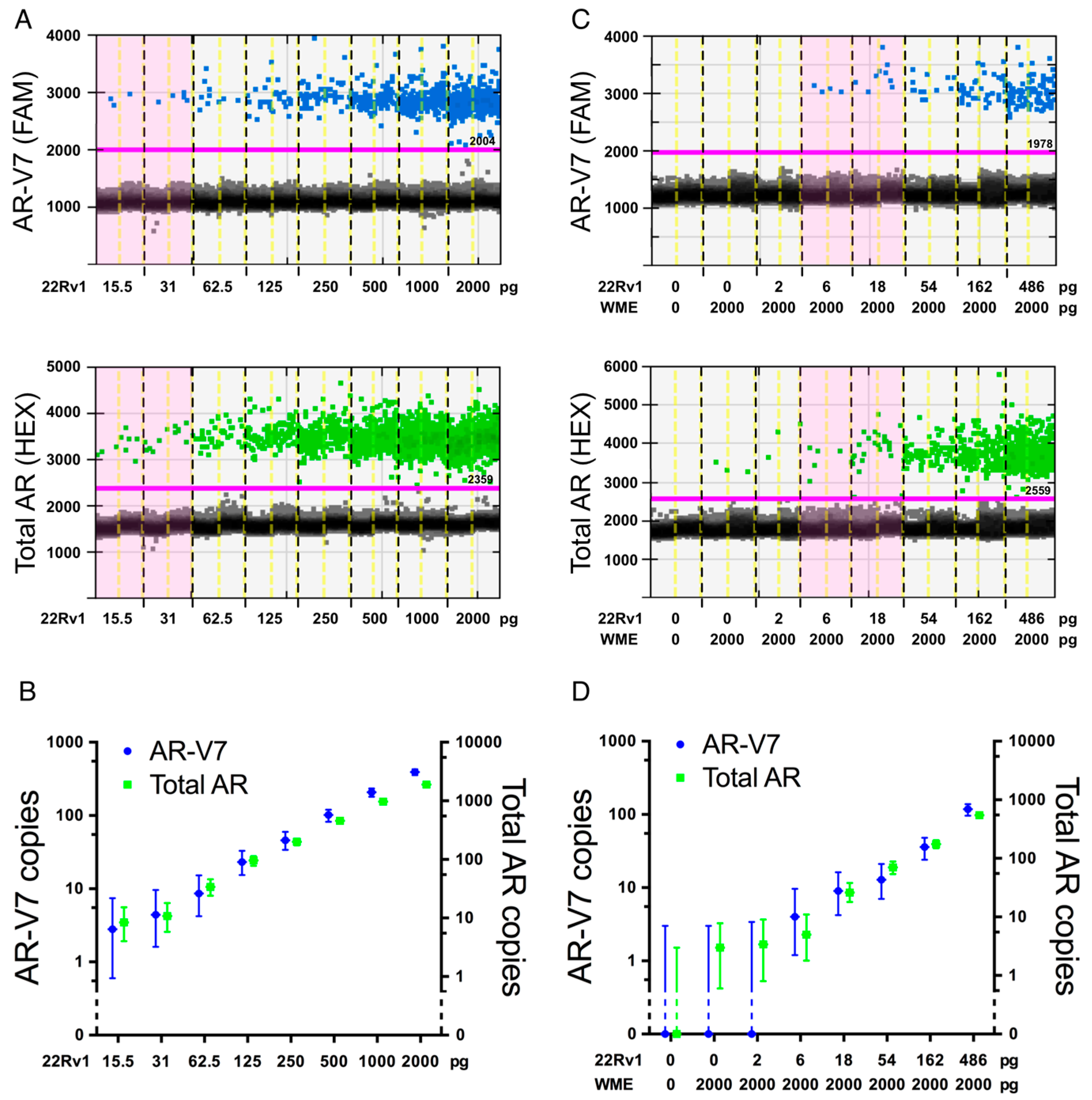
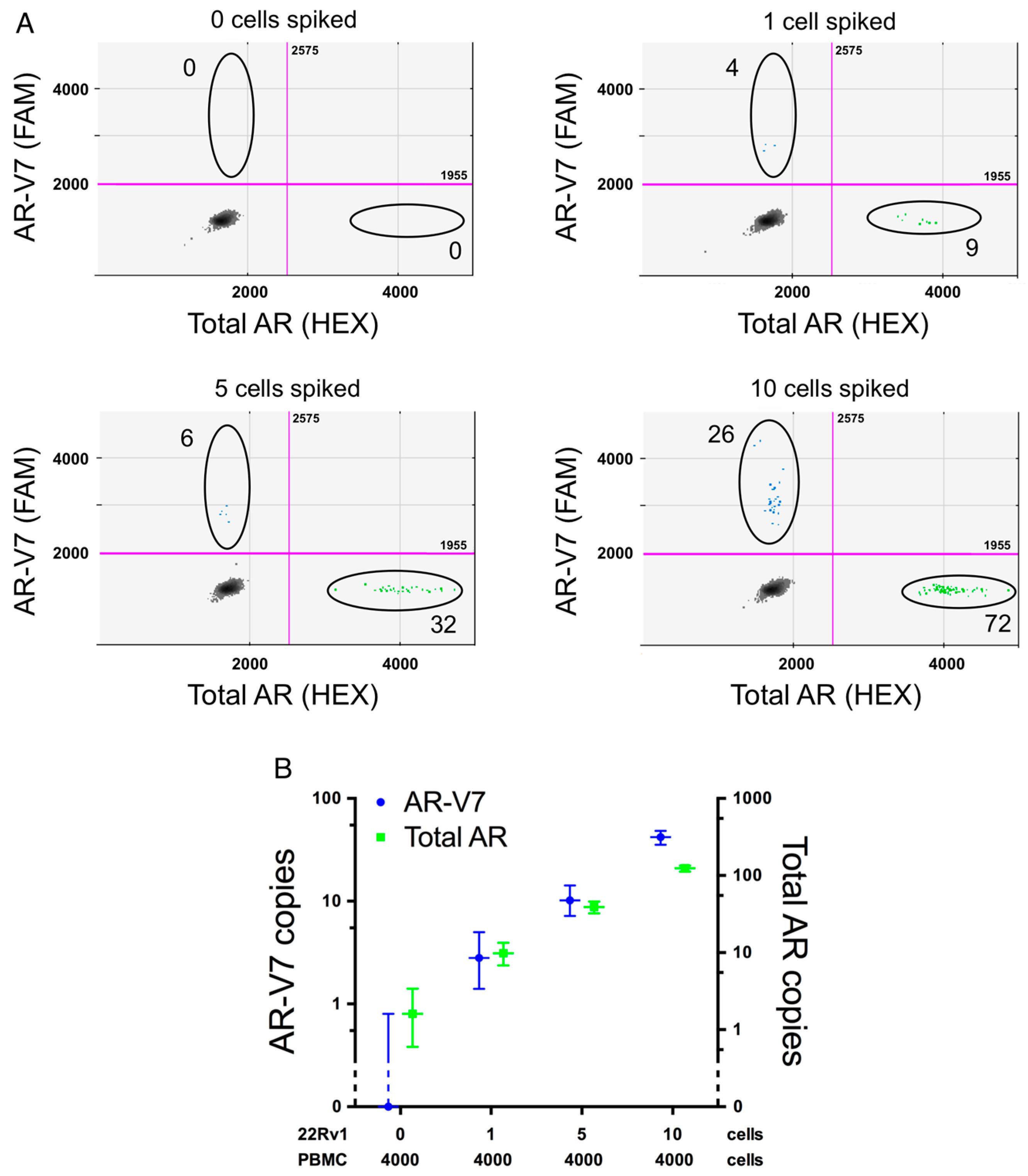
2.3. Assay Sensitivity
2.4. AR-V7 Expression in Patient Circulating Tumor Cell (CTC) Samples
3. Materials and Methods
3.1. Cell Lines
3.2. Patients
3.3. CTC Enrichment
3.4. Immunocytostaining and CTC Enumeration
3.5. RNA Extraction and cDNA Synthesis
3.6. Droplet Digital PCR (ddPCR)
3.7. Modeling CTC Samples and Single Cell Micromanipulation
3.8. Statistics
4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
| AR | androgen receptor |
| PCa | prostate cancer |
| CRPC | castrate resistant prostate cancer |
| HSPC | hormone sensitive prostate cancer |
| CTC | circulating tumor cell |
| ctNA | circulating tumor nucleic acid |
| PSA | prostate specific antigen |
| CEA | carcinoembryonic antigen |
| EGFR | epidermal growth factor receptor |
| ADT | androgen deprived therapy |
| EpCam | epithelial cell adhesion molecule |
| CK | cytokeratin |
| PBMC | peripheral blood mononuclear cell |
| ddPCR | droplet digital PCR |
| FP | forward primer |
| RP | reverse primer |
| FISH | fluorescent in situ hybridysation |
References
- Gerlinger, M.; Rowan, A.J.; Horswell, S.; Larkin, J.; Endesfelder, D.; Gronroos, E.; Martinez, P.; Matthews, N.; Stewart, A.; Tarpey, P.; et al. Intratumor heterogeneity and branched evolution revealed by multiregion sequencing. N. Engl. J. Med. 2012, 366, 883–892. [Google Scholar] [CrossRef] [PubMed]
- Gonzalez-Pons, M.; Cruz-Correa, M. Colorectal cancer biomarkers: Where are we now? BioMed Res. Int. 2015, 2015, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Pezaro, C.; Woo, H.H.; Davis, I.D. Prostate cancer: Measuring PSA. Intern. Med. J. 2014, 44, 433–440. [Google Scholar] [CrossRef] [PubMed]
- Chapman, P.B.; Hauschild, A.; Sondak, V.K. Frontline approach to metastatic braf-mutant melanoma diagnosis, molecular evaluation, and treatment choice. Am. Soc. Clin. Oncol. Educ. 2014, 34, e412–e421. [Google Scholar] [CrossRef] [PubMed]
- Hecht, J.R.; Mitchell, E.; Chidiac, T.; Scroggin, C.; Hagenstad, C.; Spigel, D.; Marshall, J.; Cohn, A.; McCollum, D.; Stella, P.; et al. A randomized phase iiib trial of chemotherapy, bevacizumab, and panitumumab compared with chemotherapy and bevacizumab alone for metastatic colorectal cancer. J. Clin. Oncol. 2009, 27, 672–680. [Google Scholar] [CrossRef] [PubMed]
- Crowley, E.; di Nicolantonio, F.; Loupakis, F.; Bardelli, A. liquid biopsy: Monitoring cancer-genetics in the blood. Nat. Rev. Clin. Oncol. 2013, 10, 472–484. [Google Scholar] [CrossRef] [PubMed]
- Becker, T.M.; Caixeiro, N.J.; Lim, S.H.; Tognela, A.; Kienzle, N.; Scott, K.F.; Spring, K.J.; de Souza, P. New frontiers in circulating tumor cell analysis: A reference guide for biomolecular profiling toward translational clinical use. Int. J. Cancer 2014, 134, 2523–2533. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.; Guan, Y.; Sun, Y.; Ai, D.; Guo, Q. Tumor heterogeneity and circulating tumor cells. Cancer Lett. 2016, 374, 216–223. [Google Scholar] [CrossRef] [PubMed]
- Yu, Z.; Chen, S.; Sowalsky, A.G.; Voznesensky, O.S.; Mostaghel, E.A.; Nelson, P.S.; Cai, C.; Balk, S.P. Rapid induction of androgen receptor splice variants by androgen deprivation in prostate cancer. Clin. Cancer Res. 2014, 20, 1590–1600. [Google Scholar] [CrossRef] [PubMed]
- Shaffer, D.R.; Leversha, M.A.; Danila, D.C.; Lin, O.; Gonzalez-Espinoza, R.; Gu, B.; Anand, A.; Smith, K.; Maslak, P.; Doyle, G.V.; et al. Circulating tumor cell analysis in patients with progressive castration-resistant prostate cancer. Clin. Cancer Res. 2007, 13, 2023–2029. [Google Scholar] [CrossRef] [PubMed]
- Leversha, M.A.; Han, J.; Asgari, Z.; Danila, D.C.; Lin, O.; Gonzalez-Espinoza, R.; Anand, A.; Lilja, H.; Heller, G.; Fleisher, M.; et al. Fluorescence in situ hybridization analysis of circulating tumor cells in metastatic prostate cancer. Clin. Cancer Res. 2009, 15, 2091–2097. [Google Scholar] [CrossRef] [PubMed]
- Jiang, Y.; Palma, J.F.; Agus, D.B.; Wang, Y.; Gross, M.E. Detection of androgen receptor mutations in circulating tumor cells in castration-resistant prostate cancer. Clin. Chem. 2010, 56, 1492–1495. [Google Scholar] [PubMed]
- Steinestel, J.; Luedeke, M.; Arndt, A.; Schnoeller, T.; Lennerz, J.K.; Maier, C.; Cronauer, M.; Steinestel, K.; Boegemann, M.; Schrader, A.J. Detecting predictive androgen receptor modifications in circulating prostate cancer cells. J. Clin. Oncol. 2015. [Google Scholar] [CrossRef]
- Thadani-Mulero, M.; Portella, L.; Sun, S.; Sung, M.; Matov, A.; Vessella, R.L.; Corey, E.; Nanus, D.M.; Plymate, S.R.; Giannakakou, P. Androgen receptor splice variants determine taxane sensitivity in prostate cancer. Cancer Res. 2014, 74, 2270–2282. [Google Scholar] [CrossRef] [PubMed]
- Qu, Y.; Dai, B.; Ye, D.; Kong, Y.; Chang, K.; Jia, Z.; Yang, X.; Zhang, H.; Zhu, Y.; Shi, G. Constitutively active AR-V7 plays an essential role in the development and progression of castration-resistant prostate cancer. Sci. Rep. 2015, 5, 7654. [Google Scholar] [CrossRef] [PubMed]
- Antonarakis, E.S.; Lu, C.; Luber, B.; Wang, H.; Chen, Y.; Nakazawa, M.; Nadal, R.; Paller, C.J.; Denmeade, S.R.; Carducci, M.A.; et al. Androgen receptor splice variant 7 and efficacy of taxane chemotherapy in patients with metastatic castration-resistant prostate cancer. JAMA Oncol. 2015, 1, 582–591. [Google Scholar] [CrossRef] [PubMed]
- Antonarakis, E.S.; Lu, C.; Wang, H.; Luber, B.; Nakazawa, M.; Roeser, J.C.; Chen, Y.; Mohammad, T.A.; Chen, Y.; Fedor, H.L.; et al. AR-V7 and resistance to enzalutamide and abiraterone in prostate cancer. N. Engl. J. Med. 2014, 371, 1028–1038. [Google Scholar] [CrossRef] [PubMed]
- Onstenk, W.; Sieuwerts, A.M.; Kraan, J.; Van, M.; Nieuweboer, A.J.; Mathijssen, R.H.; Hamberg, P.; Meulenbeld, H.J.; de Laere, B.; Dirix, L.Y.; et al. Efficacy of cabazitaxel in castration-resistant prostate cancer is independent of the presence of AR-V7 in circulating tumor cells. Eur. Urol. 2015, 68, 939–945. [Google Scholar] [CrossRef] [PubMed]
- Reid, A.L.; Freeman, J.B.; Millward, M.; Ziman, M.; Gray, E.S. Detection of BRAF-V600E and V600K in melanoma circulating tumour cells by droplet digital PCR. Clin. Biochem. 2015, 48, 999–1002. [Google Scholar] [CrossRef] [PubMed]
- Sun, H.; Mediwala, S.N.; Szafran, A.T.; Mancini, M.A.; Marcelli, M. Cudc-101, a novel inhibitor of full-length androgen receptor (FLAR) and androgen receptor variant 7 (AR-V7) activity: Mechanism of action and in vivo efficacy. Horm. Cancer 2016, 7, 196–210. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.; Lou, W.; Zhu, Y.; Nadiminty, N.; Schwartz, C.T.; Evans, C.P.; Gao, A.C. Niclosamide inhibits androgen receptor variants expression and overcomes enzalutamide resistance in castration-resistant prostate cancer. Clin. Cancer Res. 2014, 20, 3198–3210. [Google Scholar] [CrossRef] [PubMed]
- Makkonen, H.; Kauhanen, M.; Jaaskelainen, T.; Palvimo, J.J. Androgen receptor amplification is reflected in the transcriptional responses of vertebral-cancer of the prostate cells. Mol. Cell. Endocrinol. 2011, 331, 57–65. [Google Scholar] [CrossRef] [PubMed]
- Azad, A.A.; Volik, S.V.; Wyatt, A.W.; Haegert, A.; Le Bihan, S.; Bell, R.H.; Anderson, S.A.; McConeghy, B.; Shukin, R.; Bazov, J.; et al. Androgen receptor gene aberrations in circulating cell-free DNA: Biomarkers of therapeutic resistance in castration-resistant prostate cancer. Clin. Cancer Res. 2015, 21, 2315–2324. [Google Scholar] [CrossRef] [PubMed]
- Yu, Y.; Zhang, Y.; Guan, W.; Huang, T.; Kang, J.; Sheng, X.; Qi, J. Androgen receptor promotes the oncogenic function of overexpressed Jagged1 in prostate cancer by enhancing cyclin B1 expression via Akt phosphorylation. Mol. Cancer Res. 2014, 12, 830–842. [Google Scholar] [CrossRef] [PubMed]
- Mantalaris, A.; Panoskaltsis, N.; Sakai, Y.; Bourne, P.; Chang, C.; Messing, E.M.; David Wu, J.H. Localization of androgen receptor expression in human bone marrow. J. Pathol. 2001, 193, 361–366. [Google Scholar] [CrossRef]
- Sambrook, J.; Russell, D.W. Molecular Cloning—A Laboratory Manual, 3rd ed.; Cold Spring, Harbor Laboratory Press: Cold Spring Harbor, NY, USA, 2001. [Google Scholar]
- Scher, H.I.; Lu, D.; Schreiber, N.A.; Louw, J.; Graf, R.P.; Vargas, H.A.; Johnson, A.; Jendrisak, A.; Bambury, R.; Danila, D.; et al. Association of AR-V7 on circulating tumor cells as a treatment-specific biomarker with outcomes and survival in castration-resistant prostate cancer. JAMA Oncol. 2016. [Google Scholar] [CrossRef] [PubMed]
- Sharma, A.; Yeow, W.S.; Ertel, A.; Coleman, I.; Clegg, N.; Thangavel, C.; Morrissey, C.; Zhang, X.; Comstock, C.E.; Witkiewicz, A.K.; et al. The retinoblastoma tumor suppressor controls androgen signaling and human prostate cancer progression. J. Clin. Investig. 2010, 120, 4478–4492. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Morrissey, C.; Sun, S.; Ketchandji, M.; Nelson, P.S.; True, L.D.; Vakar-Lopez, F.; Vessella, R.L.; Plymate, S.R. Androgen receptor variants occur frequently in castration resistant prostate cancer metastases. PLoS ONE 2011, 6, e27970. [Google Scholar] [CrossRef] [PubMed]
- Scher, H.I.; Morris, M.J.; Stadler, W.M.; Higano, C.; Basch, E.; Fizazi, K.; Antonarakis, E.S.; Beer, T.M.; Carducci, M.A.; Chi, K.N.; et al. Trial design and objectives for castration-resistant prostate cancer: Updated recommendations from the prostate cancer clinical trials working group 3. J. Clin. Oncol. 2016, 34, 1402–1418. [Google Scholar] [CrossRef] [PubMed]
- Neumann, M.H.; Schneck, H.; Decker, Y.; Schomer, S.; Franken, A.; Endris, V.; Pfarr, N.; Weichert, W.; Niederacher, D.; Fehm, T.; et al. Isolation and characterization of circulating tumor cells using a novel workflow combining the cellsearch(r) system and the cellcelector. Biotechnol. Prog. 2016. [Google Scholar] [CrossRef] [PubMed]
| 22Rv1 | VCaP | C4-2 | LNCaP | C4-2B | LAPC4 | PC3 | WME099 | |
|---|---|---|---|---|---|---|---|---|
| AR-V7 (copies/cell) | 4.6 | 1.2 | 0.5 | 0.4 | 0.4 | 0.0 | 0.0 | 0.0 |
| AR-V7 95% CI | 2.6–6.6 | 0.8–1.8 | 0.2–0.9 | 0.1–0.8 | 0.1–0.7 | 0.0–0.2 | 0.0–0.3 | 0.0–0.2 |
| Total-AR (copies/cell) | 17.6 | 97.0 | 24.7 | 44.4 | 47.8 | 13.4 | 0.0 | 0.1 |
| Total-AR 95% CI | 14.4–21.0 | 82.7–111.4 | 22.3–27.0 | 38.9–49.9 | 44.4–51.1 | 11.5–15.2 | 0.0–0.3 | 0.0–0.4 |
| V7/AR (%) | 26.0 | 1.3 | 1.9 | 0.9 | 0.7 | 0.0 | 0.0 | 0.0 |
| PBMC-1 | PBMC-2 | PBMC-3 | PBMC-4 | PBMC-5 | PBMC-6 | |
|---|---|---|---|---|---|---|
| AR-V7 (copies/cell) | 0 | 0 | 0 | 0 | 0 | 0 |
| AR-V7 95% CI | 0–0.004 | 0–0.003 | 0–0.003 | 0–0.003 | 0–0.003 | 0–0.002 |
| Total-AR (copies/cell) | 0 | 0 | 0.002 | 0 | 0.001 | 0 |
| Total-AR 95% CI | 0–0.004 | 0–0.003 | 0–0.00 | 0–0.003 | 0–0.006 | 0–0.003 |
| Hormone Sensitivity | Patient | AR-V7 Copies | Total AR Copies | %AR-V7 of Total AR | CTC Count | Total Cell Number |
|---|---|---|---|---|---|---|
| HSPC | 1 | 0 | 96 | 0 | 31 | 2700 |
| 2 | 0 | 0 | n/a | 7 | 2897 | |
| 3 * | 0 | 0 | n/a | 3 | 6919 | |
| 4 | 0 | 0 | n/a | 7 | 6800 | |
| 5 | 0 | 40 | 0 | 9 | 3848 | |
| 6 | 0 | 0 | n/a | 56 | 3400 | |
| 7 | 0 | 80 | 0 | 8 | 3380 | |
| 8 | 0 | 0 | n/a | 7 | 2732 | |
| 9 | 0 | 8 | 0 | 6 | 5464 | |
| 10 | 0 | 24 | 0 | 65 | 7366 | |
| CRPC | 11 | 0 | 296 | 0 | 25 | 3182 |
| 12 | 0 | 360 | 0 | 28 | 6229 | |
| 13 | 0 | 0 | n/a | 102 | 3997 | |
| 14 | 0 | 88 | 0 | 35 | 1566 | |
| 15 | 0 | 16 | 0 | 184 | 3224 | |
| 16a | 0 | 24 | 0 | 82 | 3715 | |
| 16b | 0 | 960 | 0 | 81 | 2058 | |
| 17a | 0 | 0 | n/a | 122 | 1163 | |
| 17b | 32 | 1152 | 2.5 | 12 | 3686 | |
| 18 | 8 | 1000 | 0.8 | 47 | 1505 | |
| 19 | 16 | 768 | 2.3 | 70 | 1820 | |
| 20 | 104 | 5336 | 1.9 | 10 | 8900 | |
| 21 | 264 | 37,008 | 0.7 | 39 | 1418 | |
| 22 | 360 | 20,880 | 1.7 | 44 | 4600 | |
| 23 | 880 | 153,120 | 0.6 | 12 | 2077 | |
| 24 | 1632 | 74,824 | 2.2 | 56 | 4434 |
| AR-V7 | HSPC | CRPC | Total |
|---|---|---|---|
| +ve | 0 | 8 | 8 |
| −ve | 10 | 8 | 18 |
| total | 10 | 16 | 26 |
| AR-Species | Primers | Probes |
|---|---|---|
| Total AR | FP: 5′-GGAATTCCTGTGCATGAAAGC-3′ | 5′-[HEX]CTTCAGCATTATTCCAGTG[BHQ1]-3′ |
| RP: 5′-CGATCGAGTTCCTTGATGTAGTTC-3′ | ||
| AR-V7 | FP: 5′-CGGAAATGTTATGAAGCAGGGATGA-3′ | 5′-[6FAM]CGGAATTTTTCTCCCAGA[BHQ1]-3′ |
| RP: 5′-CTGGTCATTTTGAGATGCTTGCAAT-3′ |
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ma, Y.; Luk, A.; Young, F.P.; Lynch, D.; Chua, W.; Balakrishnar, B.; De Souza, P.; Becker, T.M. Droplet Digital PCR Based Androgen Receptor Variant 7 (AR-V7) Detection from Prostate Cancer Patient Blood Biopsies. Int. J. Mol. Sci. 2016, 17, 1264. https://doi.org/10.3390/ijms17081264
Ma Y, Luk A, Young FP, Lynch D, Chua W, Balakrishnar B, De Souza P, Becker TM. Droplet Digital PCR Based Androgen Receptor Variant 7 (AR-V7) Detection from Prostate Cancer Patient Blood Biopsies. International Journal of Molecular Sciences. 2016; 17(8):1264. https://doi.org/10.3390/ijms17081264
Chicago/Turabian StyleMa, Yafeng, Alison Luk, Francis P. Young, David Lynch, Wei Chua, Bavanthi Balakrishnar, Paul De Souza, and Therese M. Becker. 2016. "Droplet Digital PCR Based Androgen Receptor Variant 7 (AR-V7) Detection from Prostate Cancer Patient Blood Biopsies" International Journal of Molecular Sciences 17, no. 8: 1264. https://doi.org/10.3390/ijms17081264
APA StyleMa, Y., Luk, A., Young, F. P., Lynch, D., Chua, W., Balakrishnar, B., De Souza, P., & Becker, T. M. (2016). Droplet Digital PCR Based Androgen Receptor Variant 7 (AR-V7) Detection from Prostate Cancer Patient Blood Biopsies. International Journal of Molecular Sciences, 17(8), 1264. https://doi.org/10.3390/ijms17081264







