Evaluation of Housekeeping Genes for Quantitative Real-Time PCR Analysis of Bradysia odoriphaga (Diptera: Sciaridae)
Abstract
:1. Introduction
2. Results
2.1. Amplification Efficiencies
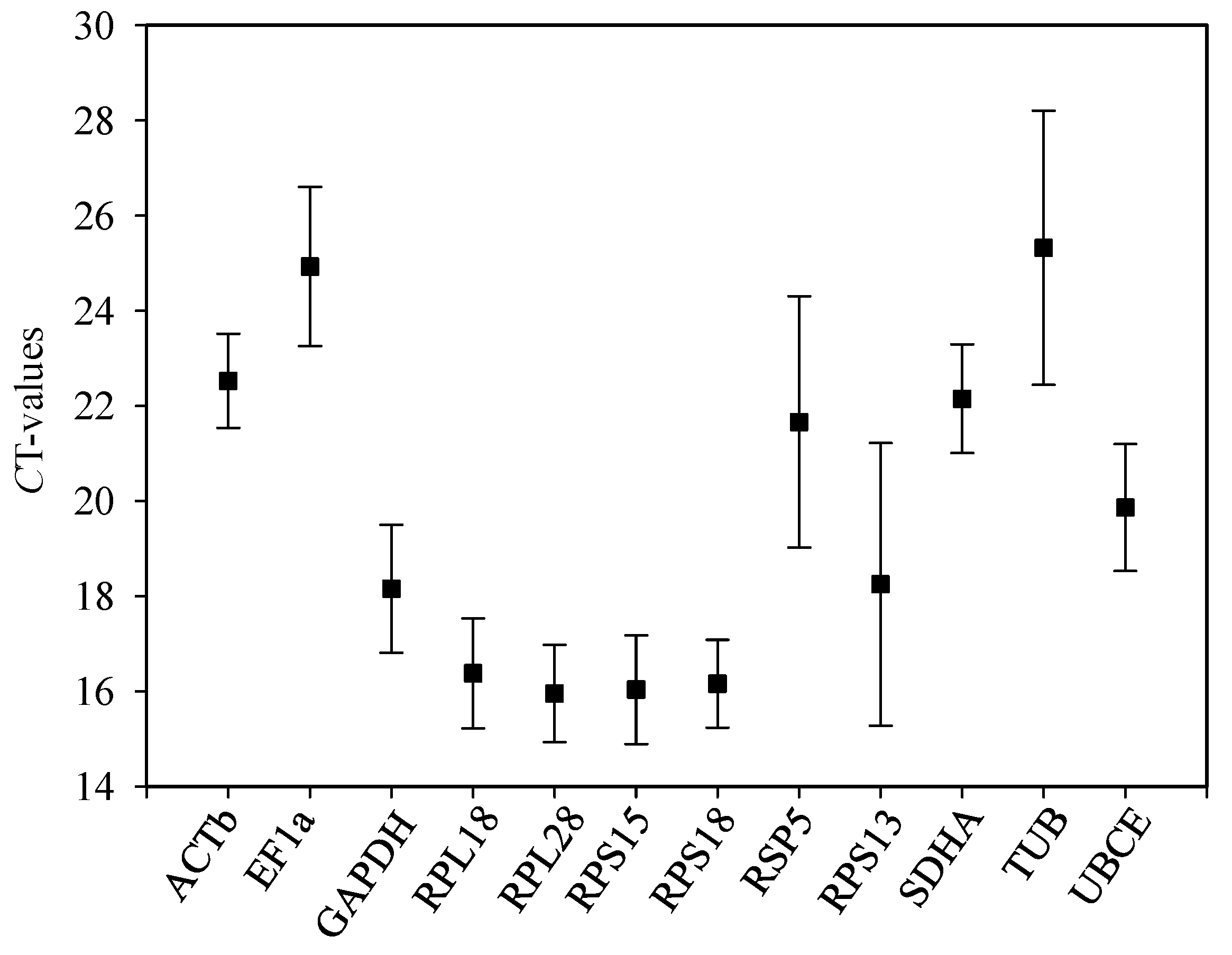
2.2. Expression Images of Candidate Reference Genes
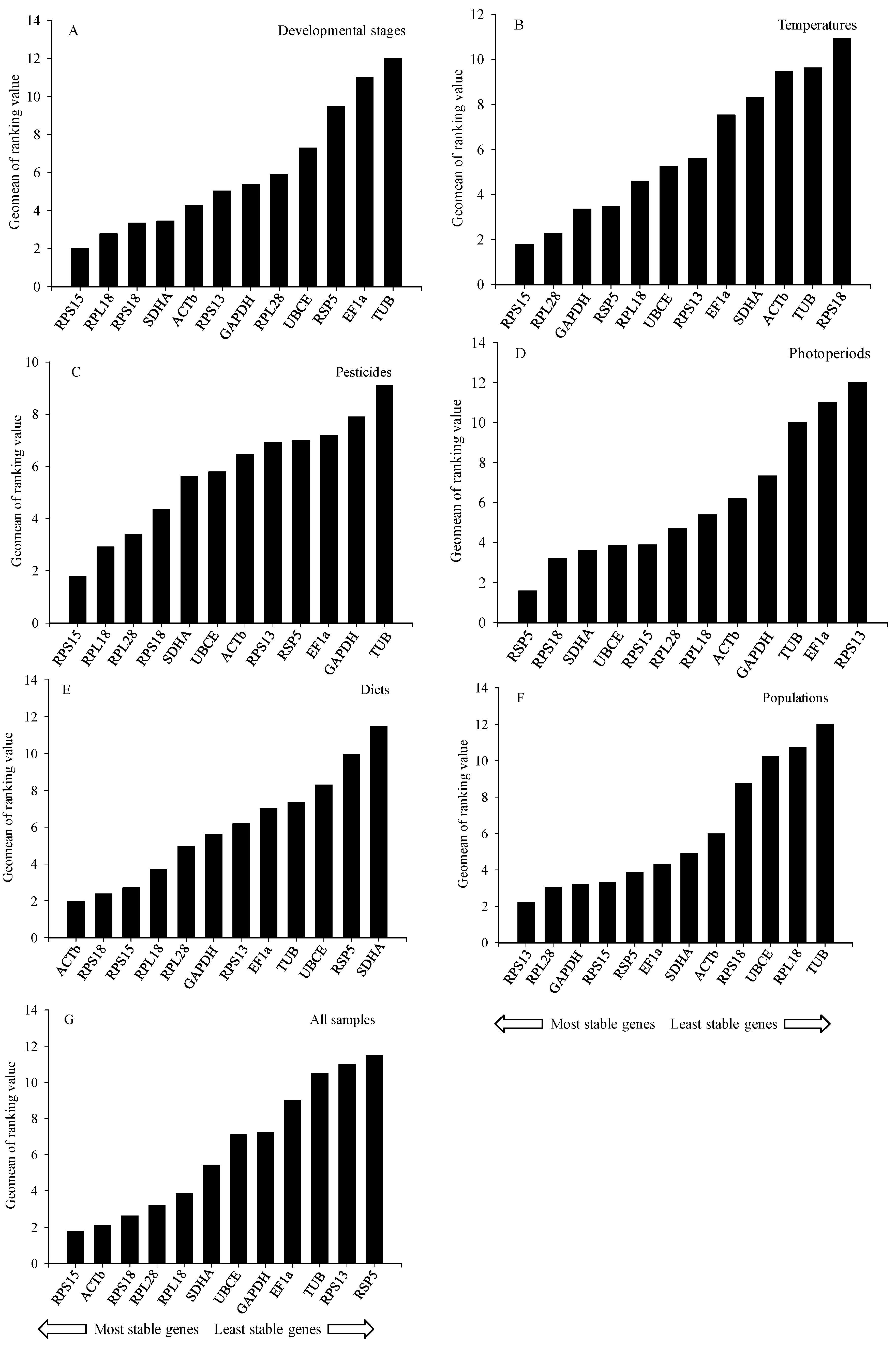
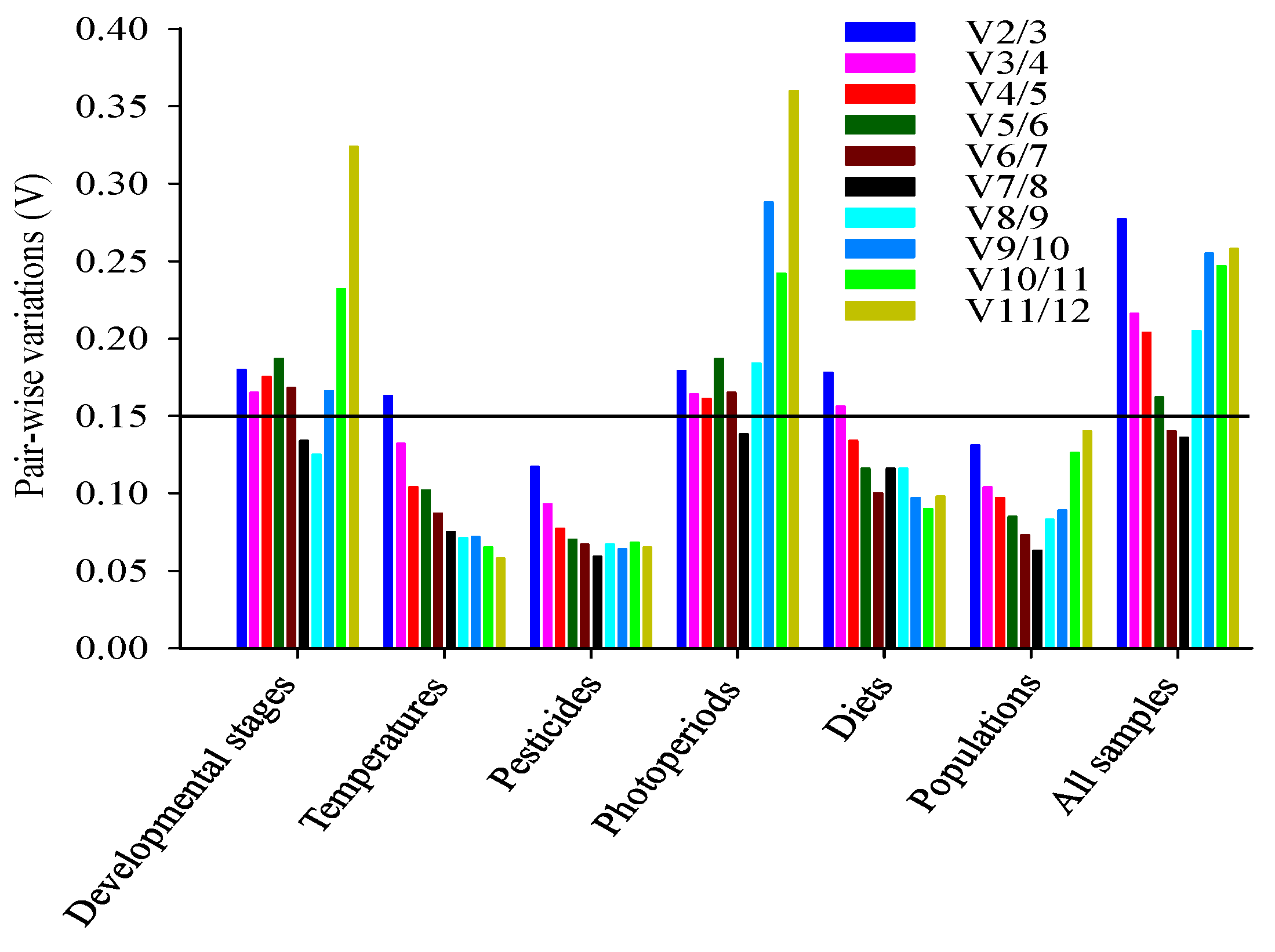
2.3. Stability of Reference Genes
2.3.1. Developmental Stages
2.3.2. Temperatures
2.3.3. Pesticides
2.3.4. Photoperiods
2.3.5. Diets
2.3.6. Populations
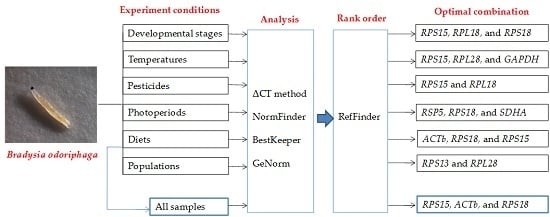
2.4. Ranking of Reference Genes for All Specimens
2.5. Target Gene Expression
3. Discussion
4. Materials and Methods
4.1. Insects
4.2. Factors that Could Affect the Expression of Housekeeping Genes
4.2.1. Developmental Stages
4.2.2. Temperatures
4.2.3. Pesticides
4.2.4. Photoperiods
4.2.5. Diets
4.2.6. Populations
4.3. Candidate Reference Genes
4.4. Total RNA Abstraction and cDNA Synthesis
4.5. qRT-PCR
4.6. Constancy of Gene Expression
4.7. Evaluation of a Target Gene Expression
4.8. Statistical Analysis
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Bustin, S.A. Absolute quantification of mRNA using real-time reverse transcription polymerase chain reaction assays. J. Mol. Endocrinol. 2000, 25, 169–193. [Google Scholar] [CrossRef] [PubMed]
- Bustin, S.A. Quantification of mRNA using real-time reverse transcription PCR (RT-PCR): Trends and problems. J. Mol. Endocrinol. 2002, 29, 23–39. [Google Scholar] [CrossRef] [PubMed]
- Kubista, M.; Andrade, J.M.; Bengtsson, M.; Forootan, A.; Jonak, J.; Lind, K.; Sindelka, R.; Sjobac, R.; Sjogreen, B.; Strombom, L.; et al. The real-time polymerase chain reaction. Mol. Asp. Med. 2006, 27, 95–125. [Google Scholar] [CrossRef] [PubMed]
- Hao, X.Y.; Horvath, D.P.; Chao, W.S.; Yang, Y.J.; Wang, X.C.; Xiao, B. Identification and evaluation of reliable reference genes for quantitative real-time PCR analysis in tea plant (Camellia sinensis (L.) O. Kuntze). Int. J. Mol. Sci. 2014, 15, 22155–22172. [Google Scholar] [CrossRef] [PubMed]
- Fleige, S.; Pfaffl, M.W. RNA integrity and the effect on the real-time qRT-PCR performance. Mol. Asp. Med. 2006, 27, 126–139. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, T.; Higgins, P.J.; Crawford, D.R. Control selection for RNA quantitation. Biotechniques 2000, 29, 332–337. [Google Scholar] [PubMed]
- Coulson, D.T.R.; Brockbank, S.; Quinn, J.G.Q.; Murphy, S.; Ravid, R.; Irvine, G.B.; Johnston, J.A. Identification of valid reference genes for the normalization of RT qPCR gene expression data in human brain tissue. BMC Mol. Biol. 2008, 9, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Su, Y.L.; He, W.B.; Wang, J.; Li, J.M.; Liu, S.S.; Wang, X.W. Selection of endogenous reference genes for gene expression analysis in the Mediterranean species of the Bemisia tabaci (Hemiptera: Aleyrodidae) complex. J. Econ. Entomol. 2013, 106, 1446–1455. [Google Scholar] [CrossRef] [PubMed]
- Yuan, M.; Lu, Y.H.; Zhu, X.; Wan, H.; Shakeel, M.; Zhan, S.; Jin, B.R.; Li, J.R. Selection and evaluation of potential reference genes for gene expression analysis in the brown planthopper, Nilaparvata lugens (Hemiptera: Delphacidae) using reverse-transcription quantitative PCR. PLoS ONE 2014, 9, e86503. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; He, L.L.; Fu, Q.T.; Xu, Z.F. Selection of reliable reference genes for gene expression studies in the biofuel plant Jatropha curcas using real-time quantitative CPR. Int. J. Mol. Sci. 2013, 14, 24338–24354. [Google Scholar] [CrossRef] [PubMed]
- Gutierrez, L.; Mauriat, M.; Guenin, S.; Pelloux, J.; Lefebre, J.E.; Louvet, R.; Rusterucci, C.; Moritz, T.; Guerineau, F.; Bellini, C.; et al. The lack of a systematic validation of reference genes: A serious pitfall undervalued in reverse transcription-polymerase chain reaction (RT-PCR) analysis in plants. Plant Biotechnol. J. 2008, 6, 609–618. [Google Scholar] [CrossRef] [PubMed]
- Thellin, O.; Zorzi, W.; Lakaye, B.; de Borman, B.; Coumans, B.; Hennen, G.; Grisar, T.; Igout, A.; Heinen, E. Housekeeping genes as internal standards: Use and limits. J. Biotechnol. 1999, 75, 291–295. [Google Scholar] [CrossRef]
- Stürzenbaum, S.R.; Kille, P. Control genes in quantitative molecular biological techniques: The variability of invariance. Comp. Biochem. Physiol. 2001, 130, 281–289. [Google Scholar] [CrossRef]
- Vandesompele, J.; de Preter, K.; Pattyn, F.; Poppe, B.; van Roy, N.; de Paepe, A.; Speleman, F. Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol. 2002, 3, 0034.1–0034.11. [Google Scholar] [CrossRef]
- Wang, E.L.; Wang, K.Y.; Chen, D.F.; Wang, J.; He, Y.; Long, B.; Yang, L.; Yang, Q.; Geng, Y.; Huang, X.L.; et al. Evaluation and selection of appropriate reference genes for real-time quantitative PCR analysis of gene expression in Nile Tilapia (Oreochromis niloticus) during vaccination and infection. Int. J. Mol. Sci. 2015, 16, 9998–10015. [Google Scholar] [CrossRef] [PubMed]
- Peters, I.R.; Peeters, D.; Helps, C.R.; Day, M.J. Development and application of multiple internal reference (housekeeper) gene assays for accurate normalisation of canine gene expression studies. Vet. Immunol. Immunopathol. 2007, 117, 55–66. [Google Scholar] [CrossRef] [PubMed]
- Yabuki, Y.; Mukaida, Y.; Saito, Y.; Oshima, K.; Takahashi, T.; Muroi, E.; Hashimoto, E.; Uda, Y. Characterisation of volatile sulphur-containing compounds generated in crushed leaves of Chinese chive (Allium tuberosum Rottler). Food Chem. 2010, 120, 343–348. [Google Scholar] [CrossRef]
- Misawa, T.; Kuninaga, S. First report of while leaf rot on Chinese chives caused by Rhizoctonia solani AG-2-1. J. Gen. Plant Pathol. 2013, 79, 280–283. [Google Scholar] [CrossRef]
- Yang, J.K.; Zhang, X.M. Notes on the fragrant onion gnats with descriptions of two new species of Bradysia (Diptera: Sciaridae). Acta Agric. Univ. Pekin. 1985, 11, 153–156. [Google Scholar]
- Yang, Y.T.; Li, W.X.; Xie, W.; Wu, Q.J.; Xu, B.Y.; Wang, S.L.; Li, C.R.; Zhang, Y.J. Development of Bradysia odoriphaga (Diptera: Sciaridae) as affected by humidity: An age-stage, two-sex, life-table study. Appl. Entomol. Zool. 2015, 50, 3–10. [Google Scholar] [CrossRef]
- Mei, Z.X.; Wu, Q.J.; Zhang, Y.J.; Hua, L. The biology, ecology and management of Bradysia odoriphaga. Entomol. Knowl. 2003, 40, 396–398. [Google Scholar]
- Dang, Z.H.; Dong, J.H.; Gao, Z.L.; Jia, H.M.; Zhang, K.J.; Pan, W.L. Biology and injury of Bradysia odoriphaga on leek in different types of cultivation. J. Agric. Univ. Hebei 2002, 24, 65–68. [Google Scholar]
- Li, H.J.; He, X.K.; Zeng, A.J.; Liu, Y.J.; Jiang, S.R. Bradysia odoriphaga copulatory behavior and evidence of a female sex pheromone. J. Agric. Urban Entomol. 2007, 24, 27–34. [Google Scholar] [CrossRef]
- Li, W.X.; Yang, Y.T.; Xie, W.; Wu, Q.J.; Xu, B.Y.; Wang, S.L.; Zhu, X.; Wang, S.J.; Zhang, Y.J. Effects of Temperature on the Age-Stage, Two-Sex Life Table of Bradysia odoriphaga (Diptera: Sciaridae). J. Econ. Entomol. 2015, 108, 126–134. [Google Scholar] [CrossRef] [PubMed]
- Zhang, P.; Liu, F.; Mu, W.; Wang, Q.H.; Li, H.; Chen, C.Y. Life table study of the effects of sublethal concentrations of thiamethoxam on Bradysia odoriphage Yang and Zhang. Pestic. Biochem. Phys. 2014, 111, 31–37. [Google Scholar] [CrossRef] [PubMed]
- Bustin, S.A.; Benes, V.; Garson, J.A.; Hellemans, J.; Huggett, J.; Kubista, M.; Mueller, R.; Nolan, T.; Pfaffl, M.W.; Shipley, G.L.; et al. The MIQE guidelines: Minimum information for publication of quantitative real-time PCR experiments. Clin. Chem. 2009, 55, 611–622. [Google Scholar] [CrossRef] [PubMed]
- Li, R.M.; Xie, W.; Wang, S.L.; Wu, Q.J.; Yang, N.N.; Yang, X.; Pan, H.P.; Zhou, X.M.; Bai, L.Y.; Xu, B.Y.; et al. Reference gene selection for qRT-PCR analysis in the sweet potato whitefly, Bemisia tabaci. (Hemiptera: Aleyrodidae). PLoS ONE 2013, 8, e53006. [Google Scholar] [CrossRef] [PubMed]
- Zhu, X.; Yuan, M.; Shakeel, M.; Zhang, Y.J.; Wang, S.L.; Wang, X.; Zhan, S.; Kang, T.H.; Li, J.H. Selection and evaluation of reference genes for expression analysis using qRT-PCR in the beet armyworm Spodoptera exigua (Hübner) (Lepidoptera: Noctuidae). PLoS ONE 2014, 9, e84730. [Google Scholar] [CrossRef] [PubMed]
- Yang, C.X.; Pan, H.P.; Liu, Y.; Zhou, X.G. Selection of reference genes for expression analysis using quantitative real-time PCR in the pea aphid, Acyrthosiphon pisum (Harris) (Hemiptera: Aphidiae). PLoS ONE 2014, 9, e110454. [Google Scholar] [CrossRef] [PubMed]
- Radonic, A.; Thulke, S.; Mackay, I.M.; Landt, O.; Siegert, W.; Nitsche, A. Guideline to reference gene selection for quantitative real-time PCR. Biochem. Biophys. Res. Commun. 2004, 313, 856–862. [Google Scholar] [CrossRef] [PubMed]
- Ladror, D.T.; Frey, B.L.; Scalf, M.; Levenstein, M.E.; Artymiuk, J.M.; Smith, L.M. Methylation of yeast ribosomal protein S2 is elevated during stationary phase growth conditions. Biochem. Biophys. Res. Commun. 2014, 445, 535–541. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.D.; An, S.H.; Li, Z.; Wu, F.M.; Yang, Q.P.; Liu, Y.C.; Cao, J.J.; Zhang, H.J.; Zhang, Q.W.; Liu, X.X. Identification and validation of reference genes for normalization of gene expression analysis using qRT-PCR in Helicoverpa armigera (Lepidoptera: Noctuidae). Gene 2015, 555, 393–402. [Google Scholar] [CrossRef] [PubMed]
- Lu, Y.H.; Yuan, M.; Gao, X.W.; Kang, T.H.; Zhan, S.; Wan, H.; Li, J.H. Identification and validation of reference genes for gene expression analysis using quantitative PCR in Spodoptera litura (Lepidoptera: Noctuidae). PLoS ONE 2013, 8, e68059. [Google Scholar] [CrossRef] [PubMed]
- Sun, W.; Jin, Y.; He, L.; Lu, W.C.; Li, M. Suitable reference gene selection for different strains and developmental stages of the carmine spider mite, Tetranychus cinnabarinus, using quantitative real-time PCR. J. Insect Sci. 2010, 10, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Chen, F.; Lu, Y.Y. Selection of reference genes in Phenacoccus solenopsis (Hemiptera: Pseudococcidae) under heat stress. Acta Entomol. Sin. 2014, 57, 1146–1154. [Google Scholar]
- Van Hiel, M.B.; Wielendaele, P.V.; Temmerman, L.; Soest, S.V.; Vuerinckx, K.; Huybrechts, R.; Broeck, J.V.; Simonet, G. Identification and validation of housekeeping genes in brains of the desert locust Schistocerca gregaria under different developmental conditions. BMC Mol. Biol. 2009, 10, 56. [Google Scholar] [CrossRef] [PubMed]
- Mamidala, P.; Rajarapu, S.P.; Jones, S.C.; Mittapalli, O. Identification and Validation of Reference Genes for Quantitative Real-Time Polymerase Chain Reaction in Cimex lectularius. J. Med. Entomol. 2011, 48, 947–951. [Google Scholar] [CrossRef] [PubMed]
- Fu, W.; Xie, W.; Zhang, Z.; Wang, S.; Wu, Q.; Liu, Y.; Zhou, X.M.; Zhou, X.G.; Zhang, Y. Exploring valid reference genes for quantitative real-time PCR analysis in Plutella xylostella (Lepidoptera: Plutellidae). Int. J. Biol. Sci. 2013, 9, 792–802. [Google Scholar] [CrossRef] [PubMed]
- Lord, J.C.; Hartzer, K.; Toutges, M.; Oppert, B. Evaluation of quantitative PCR reference genes for gene expression studies in Tribolium castaneum after fungal challenge. J. Microbiol. Methods 2010, 80, 219–221. [Google Scholar] [CrossRef] [PubMed]
- Shakeel, M.; Zhu, X.; Kang, T.H.; Wan, H.; Li, J.H. Selection and evaluation of reference genes for quantitative gene expression studies in cotton bollworm, Helicoverpa armigera (Lepidoptera: Noctuidae). J. Asia-Pac. Entomol. 2015, 18, 123–130. [Google Scholar] [CrossRef]
- Paim, R.M.; Pereira, M.H.; Ponzio, R.D.; Rodrigues, J.O.; Guarneri, A.A.; Gontijo, N.F.; Araújo, R.N. Validation of reference genes for expression analysis in the salivary gland and the intestine of Rhodnius prolixus (Hemiptera: Reduviidae) under different experimental conditions by quantitative real-time PCR. BMC Res. Notes 2012, 5, 128. [Google Scholar] [CrossRef] [PubMed]
- Hunter, T.; Garrels, J.I. Characterization of the mRNAs for α-, β- and γ-actin. Cell 1977, 12, 767–781. [Google Scholar] [CrossRef]
- Ponton, F.; Chapuis, M.P.; Pernice, M.; Sword, G.A.; Simpson, S.J. Evaluation of potential reference genes for reverse transcription-qPCR studies of physiological responses in Drosophila melanogaster. J. Insect Physiol. 2011, 57, 840–850. [Google Scholar] [CrossRef] [PubMed]
- Teng, X.; Zhang, Z.; He, G.; Yang, L.; Li, F. Validation of reference genes for quantitative expression analysis by real-time RT-PCR in four lepidopteran insects. J. Insect Sci. 2012, 12, 1–17. [Google Scholar] [CrossRef] [PubMed]
- Lourenco, A.P.; Mackert, A.; Cristino, A.S.; Simões, Z.L.P. Validation of reference genes for gene expression studies in the honey bee, Apis mellifera, by quantitative real-time RT-PCR. Apidologie 2008, 39, 372–385. [Google Scholar] [CrossRef]
- Silver, N.; Best, S.; Jiang, J.; Thein, S.L. Selection of housekeeping genes for gene expression studies in human reticulocytes using real-time PCR. BMC Mol. Biol. 2006, 7, 33. [Google Scholar] [CrossRef] [PubMed]
- Dheda, K.; Huggett, J.F.; Chang, J.S.; Kim, L.U.; Bustin, S.A. The implications of using an inappropriate reference gene for real-time reverse transcription PCR data normalization. Anal. Biochem. 2005, 344, 141–143. [Google Scholar] [CrossRef] [PubMed]
- Ling, D.J.; Salvaterra, P.M. Robust RT-qPCR data normalization: Validation and selection of internal reference genes during post-experimental data analysis. PLoS ONE 2011, 6, e17762. [Google Scholar] [CrossRef] [PubMed]
- Ferguson, B.S.; Nam, H.; Hopkins, R.G.; Morrison, R.F. Impact of reference gene selection for target gene normalization on experimental outcome using real-time qRT-PCR in adipocytes. PLoS ONE 2010, 5, e15208. [Google Scholar] [CrossRef] [PubMed]
- Veazey, K.J.; Golding, M.C. Selection of stable reference genes for quantitative RT-PCR comparisons of mouse embryonic and extra-embryonic stem cells. PLoS ONE 2011, 6, e27592. [Google Scholar] [CrossRef] [PubMed]
- Jiang, H.C.; Qian, Z.J.; Lu, W.; Ding, H.Y.; Yu, H.W.; Wang, H.; Li, J. Identification and characterization of reference genes for normalizing expression data from red swamp crawfish Procambarus clarkii. Int. J. Mol. Sci. 2015, 16, 21591–21605. [Google Scholar] [CrossRef] [PubMed]
- Colinet, H.; Hoffmann, A.A. Comparing phenotypic effects and molecular correlates of developmental, gradual and rapid cold acclimation responses in Drosophila melanogaster. Funct. Ecol. 2012, 26, 84–93. [Google Scholar] [CrossRef]
- Zhou, X.H.; Zhang, S.C.; Zhuang, Q.Y.; Zhang, A.S.; Li, L.L.; Men, X.Y.; Zhai, Y.F.; Yu, Y. Screening and evaluation of the artificial diets of Bradysia odoriphaga Yang et Zhang (Diptera: Sciaridae). Acta Entomol. Sin. 2015, 58, 1245–1252. [Google Scholar]
- Markham, N.R.; Zuker, M. DNAMelt web server for nucleic acid melting prediction. Nucleic Acids Res. 2005, 33, W577–W581. [Google Scholar] [CrossRef] [PubMed]
- Pfaffl, M.W. A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res. 2001, 29, 2002–2007. [Google Scholar] [CrossRef]
- Pfaffl, M.W.; Tichopad, A.; Prgomet, C.; Neuvians, T.P. Determination of stable housekeeping genes, differentially regulated target genes and sample integrity: BestKeeper-Excel-Based tool using pair-wise correlations. Biotechnol. Lett. 2004, 26, 509–515. [Google Scholar] [CrossRef] [PubMed]
- Xie, F.; Sun, G.; Stiller, J.W.; Zhang, B. Genome-wide functional analysis of the cotton transcriptome by creating an integrated EST database. PLoS ONE 2011, 6, e26980. [Google Scholar] [CrossRef] [PubMed]
- Zhu, X.; Li, Y.L.; Chen, D.X.; Wu, P.; Yi, T.; Chen, T.; Zhang, J.S.; Chu, W.Y. Selection of reference genes for microRNA quantitative expression analysis in Chinese perch, Siniperca chuatsi. Int. J. Mol. Sci. 2015, 16, 8310–8323. [Google Scholar] [CrossRef] [PubMed]


| Gene Symbol | Gene Name | Forward Primer (5′→3′) | Reverse Primer (5′→3′) | Product Length (bp) | Efficiency (%) | R2 * |
|---|---|---|---|---|---|---|
| ACTb | β-actin | CGCCCCCGAAGAAATTGTTG | GTCACGACCGGCAATGTCTA | 128 | 107.01 | 1.000 |
| EF1a | Elongation factor 1 alpha | TGCAACTGCACTGCGAAAAG | ACACTTTGCCCTACCGTCTG | 153 | 102.23 | 0.991 |
| GAPDH | Glyceraldehyde-3-phosphate | GCTAGTGCCGGTGCTGAATA | GACGCCACAGACGAACATTG | 144 | 100.20 | 1.000 |
| RPL18 | Ribosomal protein L18 | CCAACTGGCAAGGGAACTCT | AGCTACGTCTGCGACCTCTA | 160 | 101.26 | 0.998 |
| RPL28 | Ribosomal protein L28 | CGTGCCCGACATTTTCATCA | GACCAAGCCACTGTAACGGA | 180 | 105.18 | 1.000 |
| RPS15 | Ribosomal protein S15 | ATCGTGGCGTCGATTTGGAT | CTCATTTGGTGGGGCTTCCT | 164 | 101.03 | 0.997 |
| RPS18 | Ribosomal protein S18 | AACGAGCTGGTGAATGTACCG | TGGACGACGTCAATTGTGTG | 144 | 101.84 | 0.999 |
| RSP5 | Similar to ubiquity family member | TCTACCAAAGGCGCACACAT | CAACCGCAAATCCACACGTT | 116 | 103.85 | 1.000 |
| RPS13 | Ribosomal protein S13 | AAGTACGTTTCGTCAGCGGT | GTTTGCGAATAGCGACAGCC | 117 | 97.35 | 0.999 |
| SDHA | Succinate dehydrogenase | TTGCCTGCTGAACAATTGGC | GTCGGTACGCCACCCATATT | 134 | 95.10 | 0.998 |
| TUB | Alpha tubulin | ACAGTGCAAGGGCTTACAGG | GCTGTTGATACTCTGGGCGA | 159 | 101.80 | 1.000 |
| UBCE | Ubiquitin-conjugating enzyme | ACTACGGGCCGATTTAGCTG | CATTTGGTCGCTTCTCGCTG | 101 | 102.58 | 0.998 |
| HSP23 | Small heat shock protein | GAGAGCTATGCATCGCGACA | GCATTCTGCGGGTCGATTTC | 140 | 106.86 | 0.997 |
| Experimental Condition | Rank | ΔCt | BestKeeper | NormFinder | GeNorm | ||||
|---|---|---|---|---|---|---|---|---|---|
| Gene Name | Standard Value | Gene Name | Standard Value | Gene Name | Standard Value | Gene Name | Standard Value | ||
| Developmental stages | 1 | RPS15 | 1.460 | RPS18 | 0.559 | SDHA | 0.455 | RPL18/RPS15 | 0.429 |
| 2 | RPL18 | 1.510 | RPS13 | 0.628 | ACTb | 0.481 | |||
| 3 | ACTb | 1.520 | RPL28 | 0.742 | GAPDH | 0.729 | RPS18 | 0.530 | |
| 4 | SDHA | 1.530 | RPS15 | 0.745 | RPS15 | 0.757 | RPS13 | 0.626 | |
| 5 | GAPDH | 1.620 | RPL18 | 0.757 | UBCE | 0.810 | RPL28 | 0.756 | |
| 6 | RPS18 | 1.620 | SDHA | 0.824 | RPL18 | 0.927 | SDHA | 0.908 | |
| 7 | UBCE | 1.640 | GAPDH | 0.856 | RPS18 | 1.140 | ACTb | 1.020 | |
| 8 | RPS13 | 1.710 | ACTb | 0.970 | RSP5 | 1.221 | GAPDH | 1.080 | |
| 9 | RPL28 | 1.770 | UBCE | 1.238 | RPL28 | 1.264 | UBCE | 1.130 | |
| 10 | RSP5 | 1.950 | RSP5 | 1.652 | RPS13 | 1.273 | RSP5 | 1.259 | |
| 11 | EF1a | 2.860 | EF1a | 1.754 | EF1a | 2.514 | EF1a | 1.520 | |
| 12 | TUB | 3.990 | TUB | 3.942 | TUB | 3.870 | TUB | 1.931 | |
| Temperatures | 1 | RPS15 | 0.640 | RPL28 | 0.298 | RPS15 | 0.307 | RPL18/RSP5 | 0.476 |
| 2 | GAPDH | 0.680 | RPS15 | 0.432 | GAPDH | 0.397 | |||
| 3 | RPL28 | 0.690 | UBCE | 0.457 | RPL28 | 0.415 | RPL28 | 0.521 | |
| 4 | RSP5 | 0.720 | SDHA | 0.457 | RSP5 | 0.478 | GAPDH | 0.564 | |
| 5 | RPS13 | 0.750 | RPS13 | 0.468 | RPS13 | 0.515 | RPS15 | 0.581 | |
| 6 | UBCE | 0.760 | TUB | 0.486 | UBCE | 0.522 | EF1a | 0.625 | |
| 7 | EF1a | 0.770 | RPL18 | 0.498 | EF1a | 0.544 | UBCE | 0.654 | |
| 8 | RPL18 | 0.800 | GAPDH | 0.564 | RPL18 | 0.612 | RPS13 | 0.674 | |
| 9 | ACTb | 0.830 | RSP5 | 0.585 | ACTb | 0.645 | RPS18 | 0.696 | |
| 10 | SDHA | 0.850 | ACTb | 0.608 | TUB | 0.682 | ACTb | 0.726 | |
| 11 | RPS18 | 0.860 | EF1a | 0.712 | SDHA | 0.683 | SDHA | 0.748 | |
| 12 | TUB | 0.860 | RPS18 | 0.721 | RPS18 | 0.695 | TUB | 0.767 | |
| Pesticides | 1 | RPS15 | 0.550 | SDHA | 0.277 | RPS15 | 0.297 | RPL28/RPS15 | 0.300 |
| 2 | RPL18 | 0.580 | EF1a | 0.305 | RPL18 | 0.323 | |||
| 3 | RPL28 | 0.580 | ACTb | 0.402 | RPS18 | 0.356 | RPL18 | 0.351 | |
| 4 | RPS18 | 0.600 | TUB | 0.496 | RPL28 | 0.373 | GAPDH | 0.387 | |
| 5 | UBCE | 0.610 | RPS18 | 0.506 | UBCE | 0.385 | UBCE | 0.413 | |
| 6 | RPS13 | 0.620 | RPL18 | 0.511 | RPS13 | 0.387 | RPS18 | 0.438 | |
| 7 | RSP5 | 0.630 | RSP5 | 0.518 | RSP5 | 0.424 | RSP5 | 0.470 | |
| 8 | ACTb | 0.670 | RPS13 | 0.585 | ACTb | 0.471 | RPS13 | 0.492 | |
| 9 | GAPDH | 0.670 | UBCE | 0.632 | GAPDH | 0.536 | ACTb | 0.535 | |
| 10 | SDHA | 0.750 | RPS15 | 0.656 | SDHA | 0.591 | SDHA | 0.575 | |
| 11 | EF1a | 0.830 | RPL28 | 0.684 | EF1a | 0.704 | EF1a | 0.622 | |
| 12 | TUB | 0.880 | GAPDH | 0.774 | TUB | 0.765 | TUB | 0.664 | |
| Photoperiods | 1 | RSP5 | 1.620 | RSP5 | 0.526 | RSP5 | 0.324 | RPS18/UBCE | 0.542 |
| 2 | RPS15 | 1.680 | ACTb | 0.700 | SDHA | 0.363 | |||
| 3 | SDHA | 1.720 | RPS18 | 0.967 | RPL28 | 0.442 | RPL18 | 0.580 | |
| 4 | RPL28 | 1.740 | SDHA | 0.998 | RPS15 | 0.523 | RPS15 | 0.655 | |
| 5 | RPL18 | 1.760 | RPL28 | 1.035 | RPS18 | 0.849 | GAPDH | 0.746 | |
| 6 | UBCE | 1.770 | UBCE | 1.047 | UBCE | 0.850 | RSP5 | 0.903 | |
| 7 | RPS18 | 1.780 | RPS15 | 1.212 | RPL18 | 0.899 | SDHA | 1.009 | |
| 8 | GAPDH | 1.900 | RPL18 | 1.335 | GAPDH | 1.071 | RPL28 | 1.074 | |
| 9 | ACTb | 2.040 | GAPDH | 1.592 | ACTb | 1.337 | ACTb | 1.225 | |
| 10 | TUB | 3.040 | TUB | 1.874 | TUB | 2.899 | TUB | 1.564 | |
| 11 | EF1a | 3.090 | EF1a | 2.075 | EF1a | 2.956 | EF1a | 1.778 | |
| 12 | RPS13 | 4.370 | RPS13 | 4.172 | RPS13 | 4.300 | RPS13 | 2.210 | |
| Diets | 1 | ACTb | 0.850 | RPS15 | 0.596 | ACTb | 0.333 | RPL18/RPS18 | 0.470 |
| 2 | RPS18 | 0.860 | EF1a | 0.604 | RPS18 | 0.435 | |||
| 3 | RPS15 | 0.920 | GAPDH | 0.638 | RPS15 | 0.550 | ACTb | 0.546 | |
| 4 | RPL18 | 0.960 | TUB | 0.665 | RPL18 | 0.621 | RPL28 | 0.613 | |
| 5 | RPL28 | 0.980 | ACTb | 0.777 | RPL28 | 0.683 | RPS13 | 0.673 | |
| 6 | RPS13 | 1.020 | RPL28 | 0.803 | GAPDH | 0.728 | RPS15 | 0.719 | |
| 7 | GAPDH | 1.050 | RPS13 | 0.805 | RPS13 | 0.735 | UBCE | 0.752 | |
| 8 | UBCE | 1.060 | RPS18 | 0.864 | UBCE | 0.801 | GAPDH | 0.825 | |
| 9 | TUB | 1.130 | RSP5 | 0.928 | TUB | 0.870 | TUB | 0.900 | |
| 10 | RSP5 | 1.160 | SDHA | 0.956 | RSP5 | 0.920 | EF1a | 0.945 | |
| 11 | EF1a | 1.190 | UBCE | 0.980 | EF1a | 0.977 | RSP5 | 0.984 | |
| 12 | SDHA | 1.340 | RPL18 | 1.056 | SDHA | 1.154 | SDHA | 1.042 | |
| Populations | 1 | RPS13 | 0.760 | RPL28 | 0.200 | RPS13 | 0.189 | EF1a/RSP5 | 0.405 |
| 2 | RPS15 | 0.770 | SDHA | 0.214 | RPS15 | 0.247 | |||
| 3 | GAPDH | 0.770 | GAPDH | 0.366 | RPL28 | 0.324 | GAPDH | 0.430 | |
| 4 | RPL28 | 0.790 | RPS13 | 0.404 | GAPDH | 0.364 | ACTb | 0.457 | |
| 5 | RSP5 | 0.810 | ACTb | 0.406 | RSP5 | 0.445 | RPS15 | 0.498 | |
| 6 | SDHA | 0.830 | RPS15 | 0.473 | SDHA | 0.448 | RPS13 | 0.527 | |
| 7 | EF1a | 0.860 | EF1a | 0.474 | EF1a | 0.525 | RPL28 | 0.551 | |
| 8 | ACTb | 0.860 | RPS18 | 0.503 | ACTb | 0.546 | SDHA | 0.567 | |
| 9 | RPS18 | 0.960 | RSP5 | 0.517 | RPS18 | 0.604 | RPS18 | 0.625 | |
| 10 | UBCE | 1.080 | RPL18 | 0.834 | UBCE | 0.830 | UBCE | 0.694 | |
| 11 | RPL18 | 1.550 | UBCE | 0.937 | RPL18 | 1.472 | RPL18 | 0.829 | |
| 12 | TUB | 1.740 | TUB | 1.576 | TUB | 1.674 | TUB | 0.981 | |
| All samples | 1 | RPS15 | 1.630 | RPS18 | 0.744 | ACTb | 0.565 | RPL28/RPS15 | 0.893 |
| 2 | ACTb | 1.650 | ACTb | 0.811 | RPS15 | 0.668 | |||
| 3 | RPL18 | 1.660 | RPL28 | 0.828 | RPS18 | 0.763 | RPL18 | 0.926 | |
| 4 | RPS18 | 1.670 | SDHA | 0.917 | RPL18 | 0.768 | RPS18 | 0.968 | |
| 5 | RPL28 | 1.710 | RPS15 | 0.925 | UBCE | 0.810 | ACTb | 1.054 | |
| 6 | SDHA | 1.730 | RPL18 | 1.039 | SDHA | 0.826 | SDHA | 1.095 | |
| 7 | GAPDH | 1.740 | GAPDH | 1.057 | RPL28 | 0.848 | GAPDH | 1.127 | |
| 8 | UBCE | 1.760 | UBCE | 1.069 | GAPDH | 0.868 | UBCE | 1.171 | |
| 9 | EF1a | 2.330 | EF1a | 1.192 | EF1a | 1.116 | EF1a | 1.354 | |
| 10 | TUB | 2.990 | RSP5 | 2.125 | TUB | 2.623 | RPS13 | 1.620 | |
| 11 | RPS13 | 3.030 | TUB | 2.210 | RPS13 | 2.774 | TUB | 1.857 | |
| 12 | RSP5 | 3.320 | RPS13 | 2.274 | RSP5 | 3.062 | RSP5 | 2.101 | |
| Experimental Condition | Reference Genes | ||
|---|---|---|---|
| Developmental stages | RPS15 | RPL18 | RPS18 |
| Temperatures | RPS15 | RPL28 | GAPDH |
| Pesticides | RPS15 | RPL18 | |
| Photoperiods | RSP5 | RPS18 | SDHA |
| Diets | ACTb | RPS18 | RPS15 |
| Populations | RPS13 | RPL28 | |
| All samples | RPS15 | ACTb | RPS18 |
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Shi, C.; Yang, F.; Zhu, X.; Du, E.; Yang, Y.; Wang, S.; Wu, Q.; Zhang, Y. Evaluation of Housekeeping Genes for Quantitative Real-Time PCR Analysis of Bradysia odoriphaga (Diptera: Sciaridae). Int. J. Mol. Sci. 2016, 17, 1034. https://doi.org/10.3390/ijms17071034
Shi C, Yang F, Zhu X, Du E, Yang Y, Wang S, Wu Q, Zhang Y. Evaluation of Housekeeping Genes for Quantitative Real-Time PCR Analysis of Bradysia odoriphaga (Diptera: Sciaridae). International Journal of Molecular Sciences. 2016; 17(7):1034. https://doi.org/10.3390/ijms17071034
Chicago/Turabian StyleShi, Caihua, Fengshan Yang, Xun Zhu, Erxia Du, Yuting Yang, Shaoli Wang, Qingjun Wu, and Youjun Zhang. 2016. "Evaluation of Housekeeping Genes for Quantitative Real-Time PCR Analysis of Bradysia odoriphaga (Diptera: Sciaridae)" International Journal of Molecular Sciences 17, no. 7: 1034. https://doi.org/10.3390/ijms17071034
APA StyleShi, C., Yang, F., Zhu, X., Du, E., Yang, Y., Wang, S., Wu, Q., & Zhang, Y. (2016). Evaluation of Housekeeping Genes for Quantitative Real-Time PCR Analysis of Bradysia odoriphaga (Diptera: Sciaridae). International Journal of Molecular Sciences, 17(7), 1034. https://doi.org/10.3390/ijms17071034