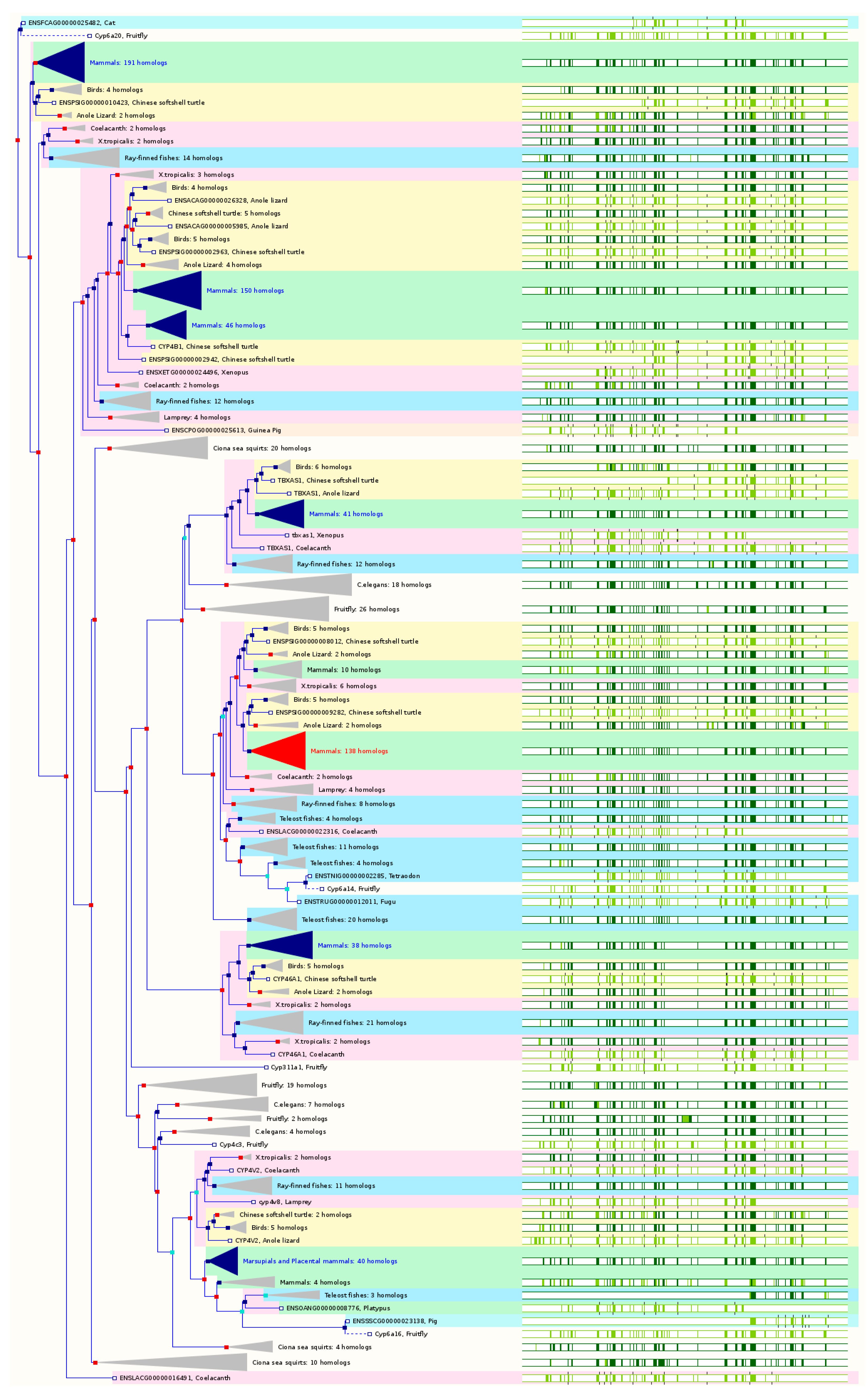
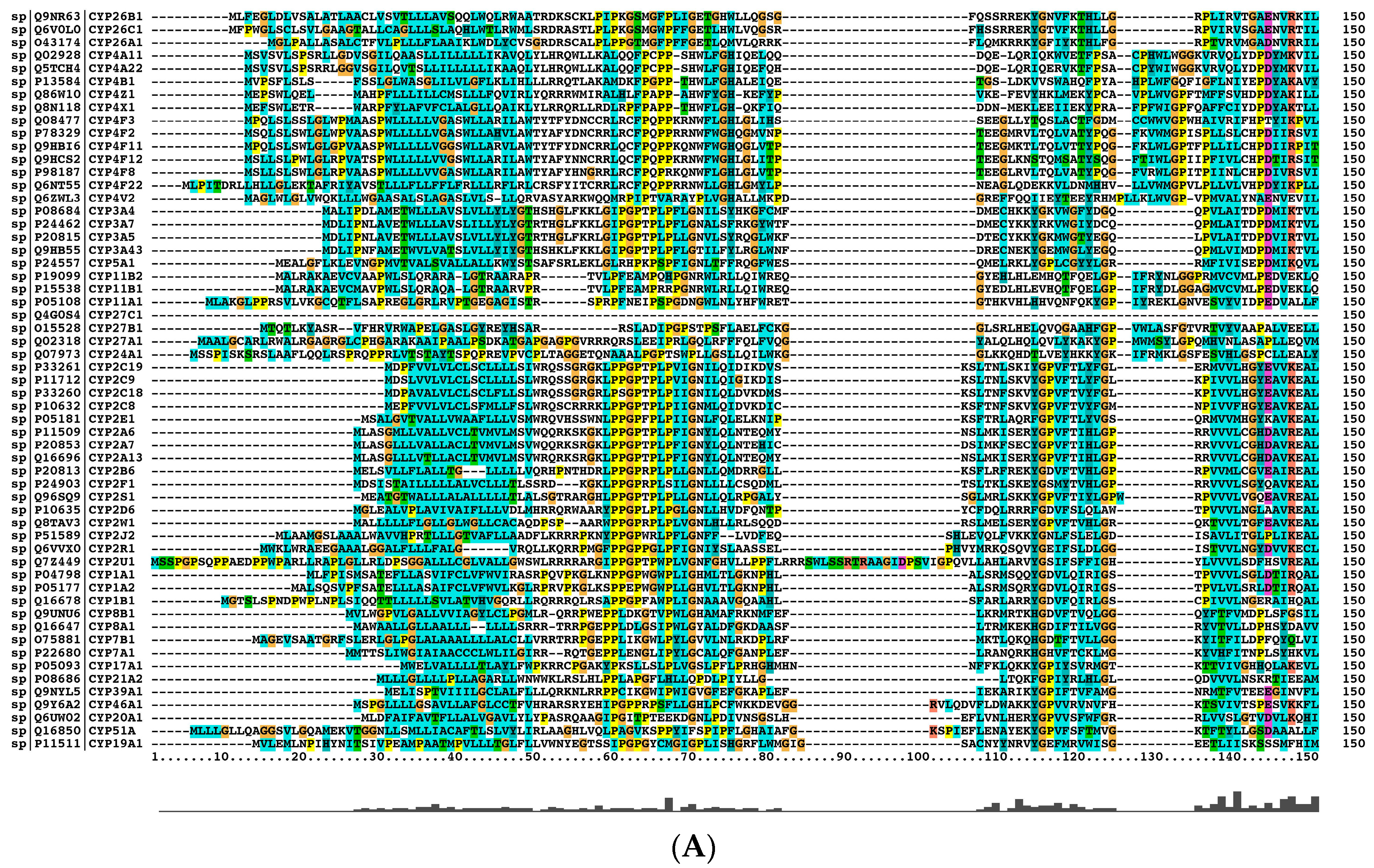
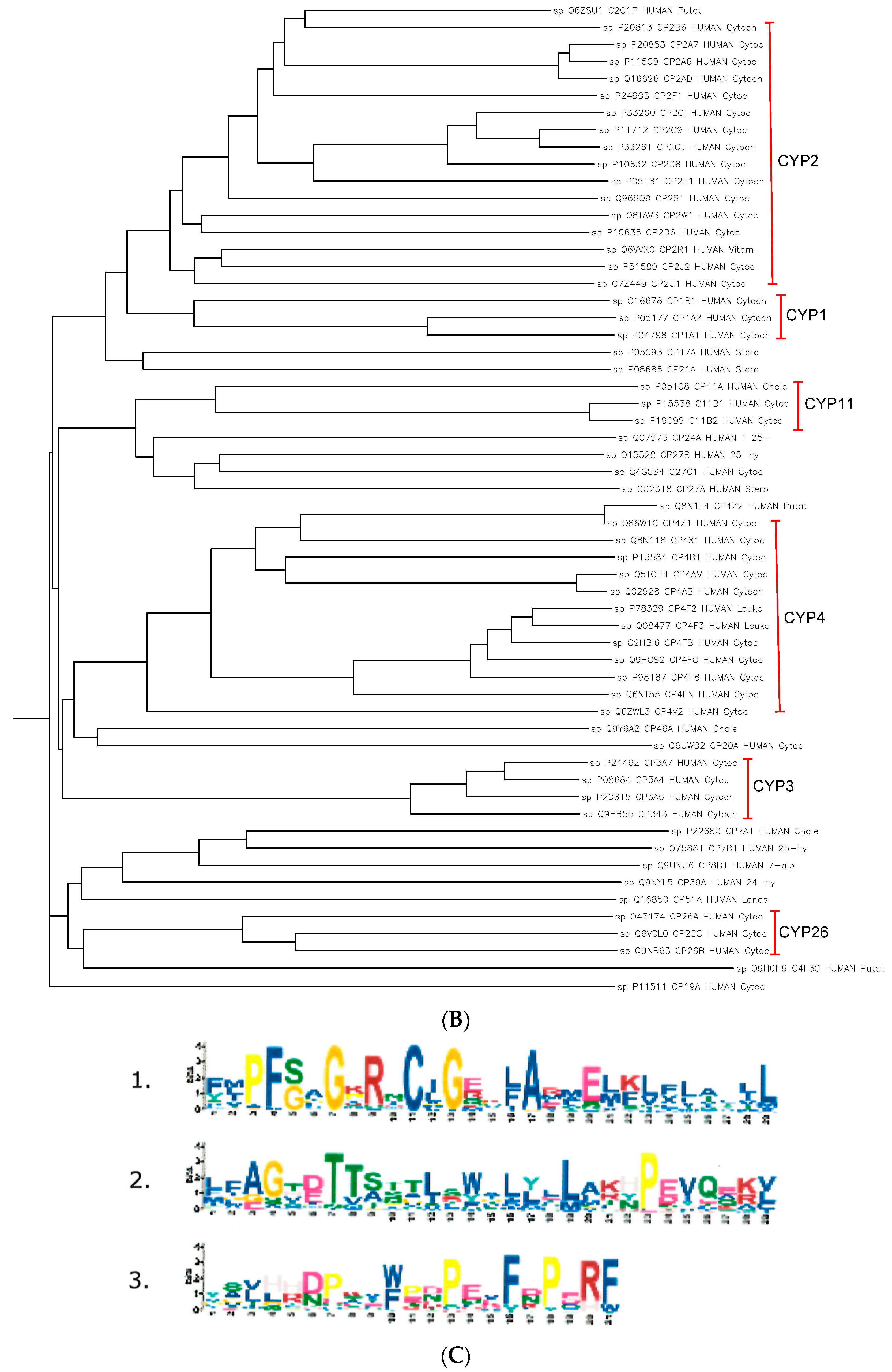
2.2. The Paralogs, Homologs and Orthologs of CYP1A1, 1B1, 1A2, 17A1, and 21A2
The human genome contains three functional
CYP1 genes including
CYP1A1,
1A2, and
1B1 and one pseudogene
CYP1D1P/1A8P.
CYP1D1 became a pseudogene in human and cattle due to five nonsense mutations in the putative coding region; however, several other mammals including chimpanzee, Rhesus monkey, and cynomolgus monkey possess a functional
CYP1D1.
CYP1D1 is also conserved in the zebrafish, frog,
Magnaporthe oryzae (
M. oryzae), and rice. In cynomolgus monkey,
CYP1D1 is 95% identical to human
CYP1D1P sequence and is mainly expressed in the liver, kidney, and jejunum. Cynomolgus monkey CYP1D1 heterologously expressed in
E. coli catalyzes ethoxyresorufin
O-deethylation and caffeine 8-hydroxylation, which human CYP1A1/1A2 also catalyze. Based on both GeneCards 4.1.1 and Ensembl 84,
CYP17A1 and
21A2 are the paralogs of
CYP1A1,
1A2 and
1B1 (
Figure 2 and
Table 1).
CYP1A1 is primarily distributed in extrahepatic tissues, but it is involved in the metabolism of a number of clinical drugs (e.g., acetaminophen, caffeine, propranolol, and theophylline), certain procarcinogens (e.g., polycyclic aromatic hydrocarbons and aristolochic acid), toxicants and environmental compounds, and some endogenous substrates such as arachidonic acid (AA) and 17β-estradiol. The gene is highly inducible and mainly expressed in fetal liver. It has a low level in adult liver, lung, skin, intestine, skin, and gallbladder. In NCBI HomoloGene 68,
CYP1A1 has 14 homologenes in 11 species, including chimpanzee, Rhesus monkey, dog, mouse, rat, Xenopus, zebrafish, etc. Based on NCBI Annotation Pipeline, 98 organisms have orthologs with
CYP1A1 (
Table S2). These include non-human primates, rodents, even-toed ungulates and whales, other mammals, birds, fishes, other vertebrates, etc. In Ensembl 84,
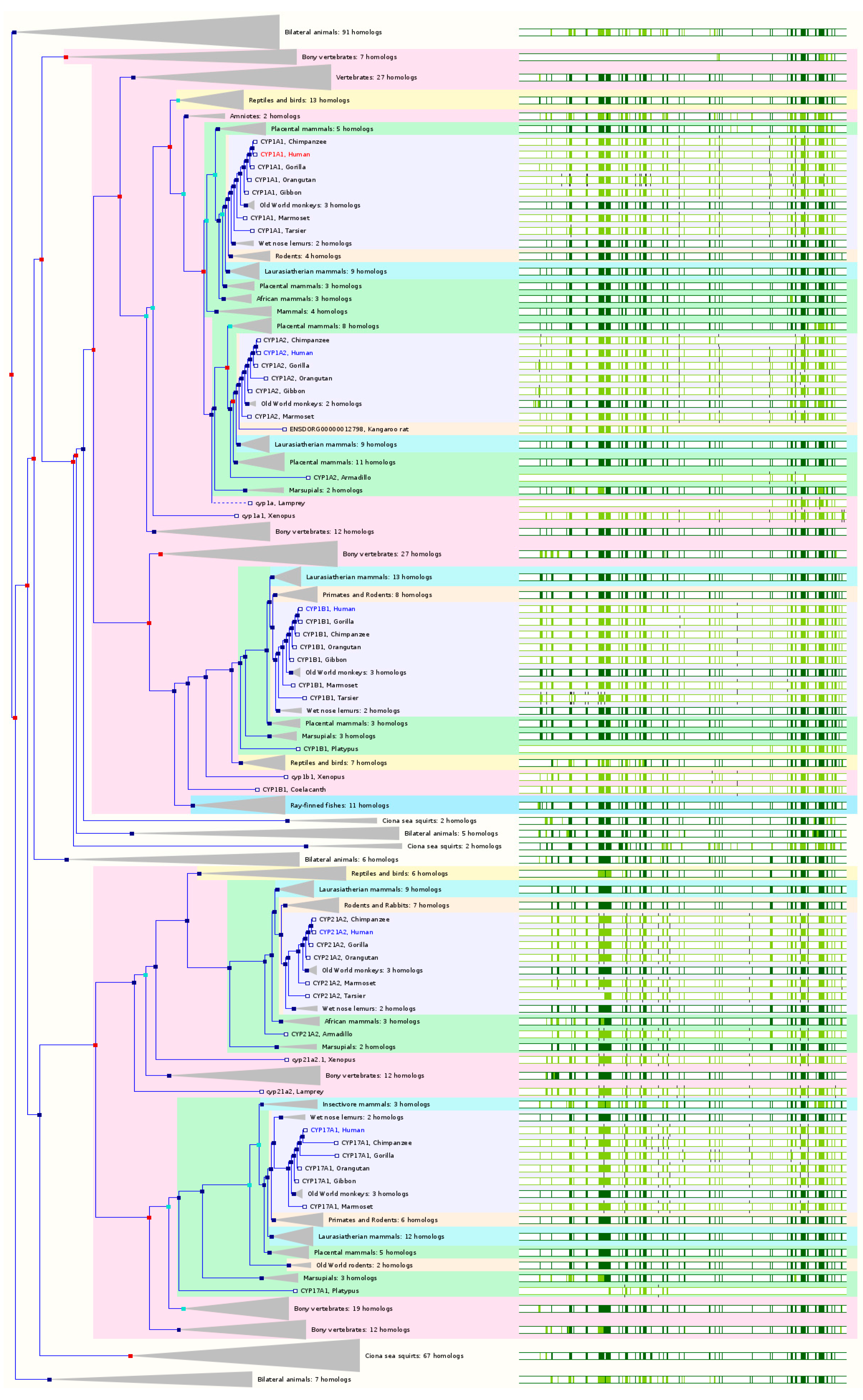
CYP1A1 has 71 orthologs from 63 species of chordates including 11 species of non-human primates, 7 species of rodents, 12 species of Laurasiatheria, 35 species of placental mammals, 7 species of Sauropsida, and 11 species of fishes (
Figure 2 and
Table S3). In GeneCards version 4.1.1,
CYP1A1 has orthologs in 15 species including chimpanzee, cattle, dog, mouse, rat, etc. (
Table S4). Similar results with remarkable differences are observed with OMA, PANTHER, and EggNOG (
Tables S5–S7).
CYP1A2 is a hepatic enzyme that oxidizes drugs (e.g., caffeine, clozapine, tacrine, theophylline, propranolol, and acetaminophen), procarcinogens and environmental compounds (e.g., benzopyrene, aflatoxin B
1, and nicotine), and several groups of endogenous substrates (e.g., steroids and AA). Hepatic CYP1A2 is highly inducible by proton pump inhibitors such as omeprazole, smoking, polyamine hydrocarbons from grilled meats, and dietary cruciferous vegetables. Furafylline is a selective and potent inhibitor for human CYP1A2, but it is a weak inhibitor for mouse, rat and dog CYP1A2/1a2 and no inhibitory effect on monkey CYP1A2. In NCBI HomoloGene 68,
CYP1A2 has 7 homologenes in 7 species, including chimpanzee, dog, cattle, mouse, rat, etc. Based on NCBI Annotation Pipeline, 69 organisms have orthologs with
CYP1A2 (
Table S2). These include non-human primates, rodents, even-toed ungulates and whales, other mammals, birds, fishes, other vertebrates, etc. In Ensembl release 84,
CYP1A2 has 69 orthologs from 60 species of chordates including 9 species of non-human primates, 8 species of rodents, 11 species of Laurasiatheria, 33 species of placental mammals, 7 species of Sauropsida, and 11 species of fishes (
Figure 2 and
Table S3). In GeneCards 4.1.1,
CYP1A2 has orthologs in 15 species including chimpanzee, dog, mouse, rat, etc. (
Table S4).
The
CYP1B1 gene located on chromosome 2p22.2 contains 3 exons and 2 introns, encoding a 543-amino acid protein. CYP1B1 can metabolize some drugs and several endogenous substances, but its role points to the bioactivation of certain procarcinogens such as polycyclic aromatic hydrocarbons.
CYP1B1 is highly expressed in the nasal epithelium and lung, with a low expression in the liver, intestine, and kidney. In NCBI HomoloGene 68,
CYP1B1 has 11 homologenes in 11 species, including chimpanzee, Rhesus monkey, dog, cattle, mouse, rat, Xenopus, zebrafish, etc. Based on NCBI Annotation Pipeline, 156 organisms have orthologs with
CYP1B1 (
Table S2). These include non-human primates, rodents, even-toed ungulates and whales, other mammals, birds, fishes, other vertebrates, etc. In Ensembl 84,
CYP1B1 has 64 orthologs from 61 species of chordates including 11 species of non-human primates, 8 species of rodents, 11 species of Laurasiatheria, 33 species of placental mammals, 7 species of Sauropsida, and 11 species of fishes (
Figure 2 and
Table S3). In GeneCards 4.1.1,
CYP1B1 has orthologs in 15 species including chimpanzee, cattle, dog, mouse, rat, etc. (
Table S4).
CYP17A1 converts pregnenolone and progesterone to their corresponding 17α-hydroxylated metabolites and then to dehydroepiandrosterone (DHEA) and androstenedione. CYP17A1 maps to chromosome 10q24.3. CYP17A1 is conserved in chimpanzee, Rhesus monkey, dog, cow, mouse, rat,
Arabidopsis thaliana and rice. In NCBI HomoloGene 68, CYP17A1 has 15 homologs in 8 species including chimpanzee, Rhesus monkey, dog, chicken, frog, zebrafish, etc. Based on NCBI Annotation Pipeline, 155 organisms have orthologs with CYP17A1 (
Table S2). These include non-human primates, rodents, even-toed ungulates and whales, other mammals, birds, fishes, other vertebrates, etc. In Ensembl 84,
CYP17A1 has 129 orthologs from 62 species including 10 species of non-human primates, 8 species of rodents, 13 species of Laurasiatheria, 36 species of placental mammals, 6 species of Sauropsida, and 11 species of fishes (
Figure 2 and
Table S3). In GeneCards 4.1.1,
CYP17A1 has orthologs in 15 species including chimpanzee, cattle, dog, mouse, rat, opossum, etc. (
Table S4).
CYP21A2 catalyzes steroid 21-hydroxylation, which is needed for the synthesis of mineralocorticoids and glucocorticoids in adrenal gland.
CYP21A2 contains 10 exons and 9 introns and displays relatively low sequence identity compared to other
CYP members.
CYP21A2 resides a multiallelic, complex and tandem copy number variation of the major histocompatibility complex region on chromosome 6p21.3.
CYP21A2 is expressed primarily in the adrenal cortex but has a low level in brain and lymphocytes. Mutations in
CYP21A2 cause congenital adrenal hyperplasia.
CYP21A1P is pseudogene also located on 6p21.3.
CYP21A2 is conserved in chimpanzee, Rhesus monkey, dog, cow, mouse, rat, chicken, zebrafish, and frog (
Figure 2). In NCBI HomoloGene 68,
CYP21A2 has 9 homologs in 9 species including chimpanzee, Rhesus monkey, dog, mouse, rat, etc. Based on NCBI Annotation Pipeline, 106 organisms have orthologs with
CYP21A2 (
Table S2). These include non-human primates, rodents, even-toed ungulates and whales, other mammals, birds, fishes, other vertebrates, etc. In Ensembl 84,
CYP21A2 has 120 orthologs from 53 species including 10 species of non-human primates, 7 species of rodents, 8 species of Laurasiatheria, 29 species of placental mammals, 6 species of Sauropsida, and 10 species of fish (
Figure 2 and
Table S3). In GeneCards 4.1.1,
CYP21A2 has orthologs in 11 species including chimpanzee, cattle, dog, mouse, rat, opossum, chicken, lizard, tropical clawed frog, zebrafish, and fruitfly (
Table S4).
2.3. The Paralogs, Homologs and Orthologs of CYP2 Family
The
CYP2ABFGST cluster located on chromosome 19q13.2 includes
CYP2A,
2B,
2F,
2G,
2S, and
2T subfamilies. It contains six functional genes including
CYP2A6,
2A7,
2A13,
2B6,
2F1, and
2S1 and seven pseudogenes including
CYP2A7P1,
2B7P,
2F2P,
2G1P,
2G2P,
2T2P, and
2T3P.
CYP2G1P carries a single nucleotide deletion in exon 2 and a 2.4-kb deletion between exons 3 and 7.
CYP2G2P harbors two nonsense mutations in exons 1 and 3. The
CYP2ABFGST cluster diverged through duplication events and inversions in the 80 Mya since the human and rodent lineages separated, resulting in 14 genes and 4 pseudogenes in rats, 12 active genes and 10 pseudogenes in mice, and 6 genes and 7 pseudogenes in humans. All the
CYP2 members are paralogs of each other (
Figure 3 and
Table 1).
CYP2A6 is a coumarin 7-hydroxylase that hydroxylates many drugs (e.g., tegafur, efavirenz, pilocarpine, and cyclophosphamide) and environmental and toxic compounds such as coumarin, nicotine, and nitrosamines.
CYP2A6 maps to chromosome 19q13.2 and consists of 9 exons. This gene is located within a 350-kb gene cluster on chromosome 19q13 together with
CYP2A7 and
2A13, two
CYP2A7P pseudogenes, and
CYP2B and
CYP2F subfamilies. CYP2A6 is mainly expressed in the liver. In NCBI HomoloGene 68,
CYP2A6 has 14 homologenes in 6 species including human
CYP2A7 and
2A13, Rhesus monkey
CYP2A24, mouse
Cyp2a4 and
2a5, rat
Cyp2a3, Xenopus
cyp2f2 and
2a6, etc. Based on NCBI Annotation Pipeline, 3 species have orthologs with
CYP2A6 (
Table S2). These include crab-eating macaque, Rhesus monkey, and western gorilla. In Ensembl 84,
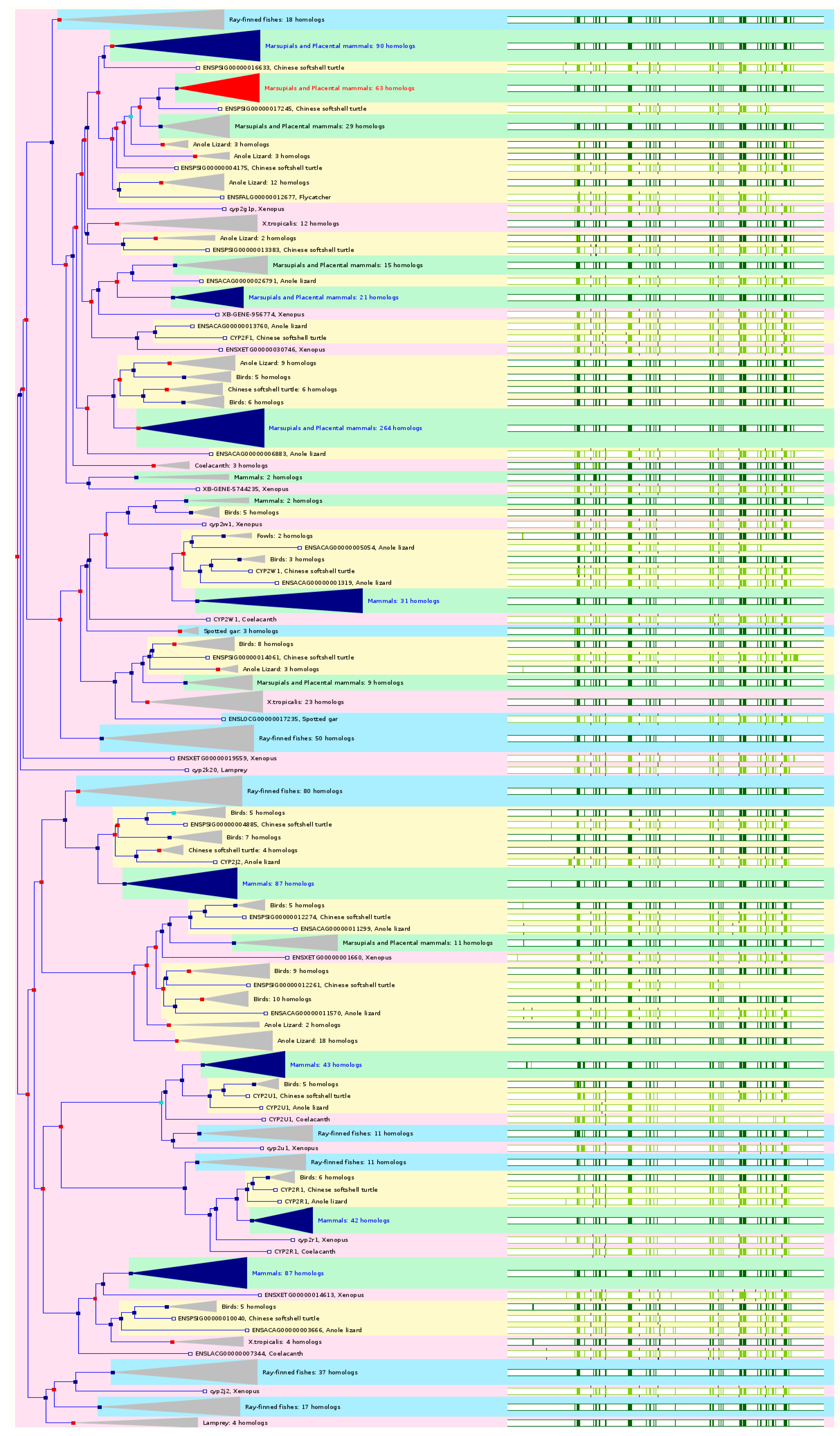
CYP2A6 has 71 orthologs from 50 species of chordates including 7 species of non-human primates, 7 species of rodents, 12 species of Laurasiatheria, 29 species of placental mammals, 3 species of Sauropsida, and 11 species of fishes (
Figure 3 and
Table S3). In GeneCards 4.1.1,
CYP2A6 has orthologs in 9 species including chimpanzee, cattle, dog, mouse, rat, etc. (
Table S4).
CYP2A7 is a hepatic enzyme, but its substrate specificity is unclear. The gene is located about 25 kb upstream of
CYP2A6. It maps to chromosome 19q13.2 consisting of 9 exons. In NCBI HomoloGene 68,
CYP2A7 has 14 homologs in 6 species including human
CYP2A6 and
2A13, Rhesus monkey
CYP2A24, mouse
Cyp2a4 and
2a5, rat
Cyp2a3, etc. Based on NCBI Annotation Pipeline, 2 organisms have orthologs with
CYP2A7, namely pig-tailed macaque and pygmy chimpanzee. In Ensembl 84,
CYP2A7 has 71 orthologs from 50 species of chordates including 7 species of non-human primates, 7 species of rodents, 12 species of Laurasiatheria, 29 species of placental mammals, 3 species of Sauropsida, and 11 species of fishes (
Figure 3 and
Table S3). In GeneCards 4.1.1,
CYP2A7 has orthologs in 9 species including chimpanzee, cattle, dog, mouse, rat, etc. (
Table S4).
CYP2A13 is responsible for metabolic activation of hexamethylphosphoramide,
N,
N-dimethylaniline, and 4-(methylnitrosamino)-1-(3-pyridyl)-1-butanone, a tobacco-specific procarcinogen in the liver and other tissues.
CYP2A13 located in the
CYP2 cluster maps to chromosome 19q13.2. The nucleotide and protein sequences of CYP2A13 are highly similar to CYP2A6 with 93.5%–95.3% identity.
CYP2A13 is mainly expressed in the liver and also in nasal mucosa, lung, brain, and trachea. In NCBI HomoloGene 68,
CYP2A13 has 14 homologs in 6 species, including human
CYP2A6 and
2A7, Rhesus monkey
CYP2A24, mouse
Cyp2a4 and
2a5, rat
Cyp2a3, Xenopus
cyp2f2 and
2a6, etc. Based on NCBI Annotation Pipeline, 3 species have orthologs with
CYP2A13, namely Weddell seal, pygmy chimpanzee, and western gorilla (
Table S2). In Ensembl 84,
CYP2A13 has 75 orthologs from 53 species of chordates including 10 species of non-human primates, 7 species of rodents, 12 species of Laurasiatheria, 32 species of placental mammals, 3 species of Sauropsida, and 11 species of fishes (
Figure 3 and
Table S3). In GeneCards 4.1.1,
CYP2A13 has orthologs in 12 species including chimpanzee, cattle, dog, mouse, rat, etc. (
Table S4).
CYP2B6 metabolizes many drugs such as efavirenz, nevirapine, bupropion, and cyclophosphamide and activate certain procarcinogens and environmental compounds such as benzo(a)pyrene and some herbicides and pesticides. The gene maps to chromosome 19q13.2 together with
CYP2B7P. It contains 9 exons encoding a 491-amino acid protein.
CYP2B6 is conserved in chimpanzee, Rhesus monkey, dog, cow, mouse, and rat. In NCBI HomoloGene 68,
CYP2B6 has 7 homologs in 7 species, including chimpanzee, Rhesus monkey, dog, cattle, mouse, and rat. Based on NCBI Annotation Pipeline, 25 organisms have orthologs with
CYP2B6 (
Table S2). These include the
CYP2B6-like or
CYP2B4 gene in non-human primates, pika, hedgehog, Arabian camel, degu, etc. In Ensembl 84,
CYP2B6 has 79 orthologs from 48 species of chordates including 8 species of non-human primates, 6 species of rodents, 11 species of Laurasiatheria, 29 species of placental mammals, 1 species of Sauropsida (Chinese softshell turtle), and 11 species of fishes (
Figure 3 and
Table S3). In GeneCards 4.1.1,
CYP2B6 has orthologs in 9 species including chimpanzee, cattle, dog, mouse, rat, etc. (
Table S4).
CYP2F1 catalyzes the dehydrogenation of 3-methylindole, an endogenous toxin derived from the fermentation of tryptophan and bioactivates lung toxicants 4-ipomeanol, naphthalene, and styrene. This gene is mainly expressed in the lung, with very low or no expression in the liver, kidney, and intestine.
CYP2F1 was present in the common ancestor of chordates and is conserved in chimpanzee, Rhesus monkey, dog, mouse, and rat. In NCBI HomoloGene 68,
CYP2F1 has 5 homologs in 5 species, including chimpanzee, Rhesus monkey, dog, mouse, and rat. Based on NCBI Annotation Pipeline, 46 organisms have orthologs with
CYP2F1 (
Table S2). These include non-human primates, rodents, other mammals, fishes, other vertebrates, etc. In Ensembl 84,
CYP2F1 has 45 orthologs from 36 species of chordates including 3 species of non-human primates, 5 species of rodents, 9 species of Laurasiatheria, 17 species of placental mammals, 1 species of Sauropsida, and 11 species of fishes (
Figure 3 and
Table S3). In GeneCards 4.1.1,
CYP2F1 has orthologs in 9 species including chimpanzee, cattle, dog, mouse, rat, etc. (
Table S4).
Human CYP2S1 oxidizes a number of carcinogens via the peroxide shunt. CYP2S1 also metabolizes PGG
2, PGH
2, hydroperoxyeicosatetraenoic acids, and 13
S-hydroperoxyoctadecadienoic acid, resulting in thromboxane A
2 and other epoxyalcohols. The gene is mainly expressed in nasal epithelium, trachea, lung, stomach, small intestine, and spleen.
CYP2S1 is conserved in chimpanzee, dog, cow, mouse, and rat. In NCBI HomoloGene release 68,
CYP2S1 has 5 homologs in 5 species, including chimpanzee, dog, cattle, mouse, and rat. Based on NCBI Annotation Pipeline, 83 organisms have orthologs with
CYP2S1 (
Table S2). These include non-human primates, rodents, even-toed ungulates and whales, other mammals, birds, fishes, other vertebrates, etc. In Ensembl 84,
CYP2S1 has 56 orthologs from 47 species of chordates including 9 species of non-human primates, 8 species of rodents, 4 species of Laurasiatheria, 28 species of placental mammals, 1 species of Sauropsida, and 11 species of fishes (
Figure 3 and
Table S3). In GeneCards 4.1.1,
CYP2S1 has orthologs in 10 species including chimpanzee, cattle, dog, mouse, rat, opossum, etc. (
Table S4).
The human CYP2C genes located on chromosome 10q24 align with an order of Centriole–2C18–2C19–2C9–2C8–Telemere. The human CYP2C genes have a significant potential to recombine, since they contains many L1 LINE repetitive DNA sequences located primarily in intron 5. Both CYP2C9 and 2C19 contain L1PA7, L1M4, L1MB5 and L1PA16 repeats in this intron. CYP2C18 and 2C19 share L1PA5 repeats. Both CYP2C8 and 2C19 carry an L1P repeat, but the two genes are on opposite strands. In mice, 14 of the 15 Cyp2c genes are located within a single cluster except for Cyp2c44 which is 3.8 Mb away from the locus and has unique catalytic property, expression profile and regulation. In the current rat genome assembly Rnor_6.0, there are 13 Cyp2c genes including 2c6/2c37, 2c7/2c39, 2c11, 2c12/2c40, 2c13/2c38, 2c22-2c24, 2c26, 2c29, 2c66, 2c79/2c65, and 2c80/2c55, which are located in a single cluster on rat chromosome 1q.
CYP2C8 metabolizes many drugs such as paclitaxel, amodiaquine, and methadone and several endogenous compounds such as AA and retinoid acid (RA).
CYP2C8 is conserved in chimpanzee, Rhesus monkey, mouse, and rat. In NCBI HomoloGene release 68,
CYP2C8 has 6 homologs in 4 species, including chimpanzee, Rhesus monkey, mouse, and rat. Based on NCBI Annotation Pipeline, 10 organisms have orthologs with
CYP2C8 (
Table S2). These include chimpanzee, pygmy chimpanzee, Bolivian squirrel monkey, horse, etc. In Ensembl 84,
CYP2C8 has 163 orthologs from 59 species of chordates including 10 species of non-human primates, 7 species of rodents, 13 species of Laurasiatheria, 35 species of placental mammals, 7 species of Sauropsida, and 11 species of fishes (
Figure 3 and
Table S3). In GeneCards 4.1.1,
CYP2C8 has orthologs in 10 species including chimpanzee, cattle, dog, mouse, rat, opossum, etc. (
Table S4).
CYP2C9 is abundant in the liver and metabolizes 15% of drugs that are metabolized by CYPs such as
S-flurbiprofen,
S-warfarin, tolbutamide, phenytoin, and diclofenac.
CYP2C9 is conserved in chimpanzee, Rhesus monkey, cow, mouse, and rat. In NCBI HomoloGene 68,
CYP2C9 has 8 homologs in 5 species, including chimpanzee, Rhesus monkey, cattle, mouse, rat, etc. Based on NCBI Annotation Pipeline, 7 organisms have orthologs with
CYP2C9. These include chimpanzee, pygmy chimpanzee, Rhesus monkey, macaque, etc. In Ensembl 84,
CYP2C9 has 160 orthologs from 58 species of chordates including 8 species of non-human primates, 8 species of rodents, 13 species of Laurasiatheria, 34 species of placental mammals, 7 species of Sauropsida, and 11 species of fishes (
Figure 3 and
Table S3). In GeneCards 4.1.1,
CYP2C9 has orthologs in 7 species including chimpanzee, cattle, mouse, rat, etc. (
Table S4).
CYP2C18 can metabolize several drugs including tolbutamide, phenytoin, and verapamil. The gene is mainly expressed in the liver, esophagus, stomach, and small intestine. The
CYP2C18 gene is conserved in Rhesus monkey, cow, mouse, rat, chicken, and mosquito. In NCBI HomoloGene 68,
CYP2C18 has 7 homologs in 6 species, including Rhesus monkey, cattle, mouse, rat, etc. Based on NCBI Annotation Pipeline, 27 organisms have orthologs with
CYP2C18 (
Table S2). These include non-human primates, even-toed ungulates and whales, other mammals, etc. In Ensembl 84,
CYP2C18 has 123 orthologs from 53 species of chordates including 8 species of non-human primates, 5 species of rodents, 11 species of Laurasiatheria, 29 species of placental mammals, 7 species of Sauropsida, and 11 species of fishes (
Figure 3 and
Table S3). In GeneCards 4.1.1,
CYP2C18 has orthologs in 10 species including chimpanzee, dog, mouse, rat, etc. (
Table S4).
CYP2C19 metabolizes about 10% of drugs that are metabolized by CYPs, such as phenytoin, omeprazole, and voriconazole. The gene is mainly expressed in the liver, small intestine, and gallbladder.
CYP2C19 is conserved in chimpanzee, Rhesus monkey, dog, cow, and rat. In NCBI HomoloGene 68,
CYP2C19 has 7 homologs in 6 species. These include chimpanzee, Rhesus monkey, dog, cow, rat, etc. Based on NCBI Annotation Pipeline, 7 organisms have orthologs with
CYP2C19 (
Table S2). These include gray short-tailed opossum, pig, goat, etc. In Ensembl 84,
CYP2C19 has 157 orthologs from 55 species of chordates including 5 species of non-human primates, 8 species of rodents, 13 species of Laurasiatheria, 31 species of placental mammals, 7 species of Sauropsida, and 11 species of fishes (
Figure 3 and
Table S3). In GeneCards 4.1.1,
CYP2C19 has orthologs in 8 species including chimpanzee, cattle, dog, mouse, rat, etc. (
Table S4).
CYP2D6 belongs to a gene cluster consisting of highly homologous 2 functional genes and 1 pseudogene on chromosome 22q13. CYP2D8P encompasses multiple deletions and insertions in its exons. In Ensembl 84, this pseudogene produces one single transcript only that does not encode any functional proteins. The evolution of the human CYP2D locus results in inactivation of CYP2D7 and 2D8P and partial inactivation of CYP2D6 (in ~10% Caucasian). Based on the identification and characterization of a non-functional CYP2D7 gene and a 2D8P pseudogene, gene duplication events may give rise to CYP2D6 and 2D7, and that gene conversion events occur later to generate CYP2D8P.
CYP2D6 accounts for only a small percentage of total hepatic CYPs (1.3%–4.3%), however, it metabolizes ~25% of the drugs that are metabolized by CYPs such as tamoxifen, imipramine, codeine, and dextromethorphan.
CYP2D6 is conserved in chimpanzee, Rhesus monkey, rat, chicken, and frog. In NCBI HomoloGene 68,
CYP2D6 has 8 homologs in 5 species. These include chimpanzee, Rhesus monkey, rat, chicken, and frog. Based on NCBI Annotation Pipeline, 77 organisms have orthologs with
CYP2D6 (
Table S2). These include non-human primates, rodents, even-toed ungulates and whales, other mammals, birds, fishes, other vertebrates, etc. In Ensembl 84,
CYP2D6 has 97 orthologs from 49 species of chordates including 11 species of non-human primates, 8 species of rodents, 12 species of Laurasiatheria, 35 species of placental mammals, 7 species of Sauropsida, and 0 species of fishes (
Figure 3 and
Table S3). In GeneCards 4.1.1,
CYP2D6 has orthologs in 12 species including chimpanzee, mouse, rat, etc. (
Table S4).
CYP2D7 is primarily expressed in brain cortex. In NCBI HomoloGene 68 and Orthologs from Annotation Pipeline,
CYP2D7 has no homolog and ortholog. In Ensembl 84,
CYP2D7 has 97 orthologs from 49 species of chordates including 11 species of non-human primates, 8 species of rodents, 12 species of Laurasiatheria, 35 species of placental mammals, 7 species of Sauropsida, and 0 species of fishes (
Figure 3 and
Table S3). In GeneCards 4.1.1,
CYP2D7 has only one ortholog in mouse (
Cyp2d37-ps). The origin of the
CYP2D subfamily could be traced back to before the divergence between amniotes and amphibians about 312 Mya.
CYP2D7 was derived from
CYP2D6 duplication in a stem lineage of humans and great apes. In fact, the origin of
CYP2D6 and
2D8P in humans can be tracked back to a stem lineage of the New World monkeys and Catarrhini at the latest. Two functional CYP2Ds have been found in marmosets and macaques.
CYP2E1 metabolizes many low molecular weight xenobiotics such as acetaminophen, chlorzoxazone, halothane, and benzene.
CYP2E1 is located on chromosome 10q26.3 with 9 exons. CYP2E1 is mainly expressed in the liver.
CYP2E1 is conserved in chimpanzee, Rhesus monkey, dog, cow, mouse, and rat. In NCBI HomoloGene release 68,
CYP2E1 has 6 homologs in 6 species including chimpanzee, Rhesus monkey, dog, cattle, mouse, and rat. Based on NCBI Annotation Pipeline, 71 organisms have orthologs with
CYP2E1 (
Table S2). These include non-human primates, rodents, even-toed ungulates and whales, other mammals, birds, fishes, other vertebrates, etc. In Ensembl 84,
CYP2E1 has 84 orthologs from 55 species of chordates including 9 species of non-human primates, 8 species of rodents, 12 species of Laurasiatheria, 32 species of placental mammals, 7 species of Sauropsida, and 11 species of fishes (
Figure 3 and
Table S3). In GeneCards 4.1.1,
CYP2E1 has orthologs in 6 species including chimpanzee, cattle, dog, mouse, rat, and opossum (
Table S4).
CYP2J2 metabolizes AA, linoleic acid, and various drugs including certain antihistamine drugs (e.g., ebastine and terfenadine), mesoridazine, danazol, certain tyrosine kinase inhibitors (e.g., imatinib and sunitinib), fenbendazole, etc. In the human genome and GRCh38.p2, there is only one single
CYP2J2 gene, which maps to chromosome 1p31.3-p31.2.
CYP2J2 is highly expressed in heart, present at a lower level in the liver, intestine, brain, bladder, pancreas, placenta, and kidney.
CYP2J2 is conserved in chimpanzee, Rhesus monkey, dog, cattle, mouse, rat, chicken, zebrafish, frog, fruitfly, mosquito, and
C. elegans. In NCBI HomoloGene 68,
CYP2J2 has as many as 51 homologs in 12 species, including chimpanzee, Rhesus monkey, dog, cattle, mouse, rat, etc. Based on NCBI Annotation Pipeline, 81 organisms have orthologs with
CYP2J2 (
Table S2). These include non-human primates, rodents, even-toed ungulates and whales, other mammals, birds, fishes, other vertebrates, etc. In Ensembl 84,
CYP2J2 has 188 orthologs from 60 species of chordates including 11 species of non-human primates, 7 species of rodents, 14 species of Laurasiatheria, 37 species of placental mammals, 7 species of Sauropsida, and 11 species of fishes (
Figure 3 and
Table S3). In GeneCards 4.1.1,
CYP2J2 has orthologs in 15 species including chimpanzee, dog, mouse, rat, etc. (
Table S4). Unlike humans, the mouse
Cyp2j cluster has 7 functional genes including
Cyp2j5,
2j6,
2j8,
2j9, and
2j11-2j13 and 3 pseudogenes including
Cyp2j7-ps,
2j14-ps, and
2j15-ps. This cluster has the unusual property that all the genes and pseudogene fragments are oriented in the same direction, which is not the case for other six
Cyp clusters.
CYP2R1 is also known as vitamin D
3 25-hydroxylase that converts vitamin D into 25-hydroxyvitamin D
3 (calcidiol). Calcidiol is subsequently converted by CYP27B1 (i.e., 25-hydroxyvitamin D
3 1-α-hydroxylase) to calcitriol, the active form of vitamin D
3 that binds to vitamin D
3 receptor which mediates most of the physiological actions of vitamin D
3.
CYP2R1 maps to chromosome 11p15.2. CYP2R1 is mainly expressed in the liver and pancreas with the highest expression in the testes.
CYP2R1 is conserved from zebrafish and frog to human. In NCBI HomoloGene 68,
CYP2R1 has 9 homologs in 9 species including chimpanzee, Rhesus monkey, dog, cattle, mouse, rat, frog, etc. Based on NCBI Annotation Pipeline, 161 organisms have orthologs with
CYP2R1 (
Table S2). These include non-human primates, rodents, even-toed ungulates and whales, other mammals, birds, fishes, other vertebrates, etc. In Ensembl 84,
CYP2R1 has 66 orthologs from 62 species of chordates including 10 species of non-human primates, 7 species of rodents, 14 species of Laurasiatheria, 37 species of placental mammals, 7 species of Sauropsida, and 11 species of fishes (
Figure 3 and
Table S3). In GeneCards 4.1.1,
CYP2J2 has orthologs in 11 species including chimpanzee, cattle, dog, mouse, etc. (
Table S4).
CYP2U1 metabolizes AA, docosahexaenoic acid, other long-chain fatty acids, and endogenous
N-arachidonoylserotonin.
CYP2U1 maps to chromosome 4q25. There is a high mRNA expression of
CYP2U1 in human thymus, with lesser expression in the heart, brain (mainly amygdala and prefrontal cortex), and platelets. The
CYP2U1 gene is conserved in chimpanzee, dog, cow, mouse, rat, chicken, zebrafish, frog, fruit fly, and
A. thaliana. In NCBI HomoloGene 68,
CYP2U1 has 11 homologs in 10 species including chimpanzee, dog, cattle, mouse, rat, frog, etc. Based on NCBI Annotation Pipeline, 160 organisms have orthologs with
CYP2U1 (
Table S2). These include non-human primates, rodents, even-toed ungulates and whales, other mammals, birds, fishes, other vertebrates, etc. In Ensembl 84,
CYP2U1 has 66 orthologs from 62 species of chordates including 11 species of non-human primates, 8 species of rodents, 13 species of Laurasiatheria, 37 species of placental mammals, 7 species of Sauropsida, and 11 species of fishes (
Figure 3 and
Table S3). In GeneCards 4.1.1,
CYP2U1 has orthologs in 13 species including chimpanzee, dog, mouse, rat, etc. (
Table S4).
CYP2W1 catalyzes the oxidation of indole and certain lipids including lysolecithin and their stereoisomers and shows monooxygenase activity towards 3-methylindole and chlorzoxazone, but not AA.
CYP2W1 maps to chromosome 7p22.3. The gene contains 10 exons and encode a 490-amino acid protein. The 5-prime flanking region, first exon, and first intron of
CYP2W1 carry abundant CpG dinucleotides including 2 CpG islands.
CYP2W1 is mainly expressed in colorectal, hepatic and adrenal gland tumors, but it is rarely detected in normal tissues.
CYP2W1 is an ancient member of the
CYP superfamily and it is conserved in chimpanzee, Rhesus monkey, dog, cow, mouse, rat, and chicken. In NCBI HomoloGene 68,
CYP2W1 has 7 homologs in 7 species including chimpanzee, Rhesus monkey, dog, cattle, mouse, rat, and chicken. Based on NCBI Annotation Pipeline, 108 organisms have orthologs with
CYP2W1 (
Table S2). These include non-human primates, rodents, even-toed ungulates and whales, other mammals, birds, fishes, other vertebrates, etc. In Ensembl 84,
CYP2W1 has 93 orthologs from 52 species of chordates including 10 species of non-human primates, 7 species of rodents, 9 species of Laurasiatheria, 28 species of placental mammals, 7 species of Sauropsida, and 11 species of fishes (
Figure 3 and
Table S3). In GeneCards 4.1.1,
CYP2W1 has orthologs in 10 species including chimpanzee, cattle, dog, mouse, etc. (
Table S4).
2.4. The Paralogs, Homologs and Orthologs of CYP3, 4, 5, and 46 Families
The
CYP3A gene cluster is located on chromosome 7q21.1 (Ensembl cytogenetic band: 7q22.1) and spans ~231 kb, containing 4
CYP3A genes:
CYP3A4,
3A5,
3A7 and
3A43, as well as 2 pseudogenes including (
CYP3A51P/3A5P1 and
3A52P/
3A5P2).
CYP3A54P and
3A137P are two additional pseudogenes in
CYP3 family, which map to chromosome 7q22.1. The human CYP3A subfamily is involved in the oxidative metabolism of a wide range of substrates, including more than 50% of all currently marketed drugs, endogenous steroids and xenobiotics.
CYP3A4 and
3A5 are mainly expressed in the liver and intestine, while
CYP3A5 appears to be primarily expressed in extrahepatic tissues.
CYP3A4 is most abundantly expressed in the liver while
CYP3A5 expression at the protein level is only about 10.6% of that of CYP3A4. Both CYP3A4 and 3A5 share substrate specificity and so it is often difficult to identify their relative contribution to the overall metabolism of a substrate.
CYP3A4,
3A5,
3A7, and
3A43 share paralogs from human
CYP superfamily and orthologs from various species with slight differences only. CYP3A7 is a fetal-specific CYP. CYP3A43 has very low expression in the liver. In GeneCards 4.1.1,
CYP3,
4, and
5 members are paralogs to each other (
Table 1). Ensembl 84 also includes
CYP46A1 as the paralog of
CYP3,
4, and
5 families (
Figure 4 and
Table 1).
CYP3A4 is conserved in chimpanzee, Rhesus monkey, dog, cow, rat, chicken, frog, fruitfly, mosquito,
C. elegans, and
M. oryzae. In NCBI HomoloGene 68,
CYP3A4 has 27 homologs in 11 species, including chimpanzee, Rhesus monkey, dog, cattle, rat, etc. Based on NCBI Annotation Pipeline, 6 organisms have orthologs with
CYP3A4 (
Table S2). These include chimpanzee, white-tufted-ear marmoset, drill, Bolivian squirrel monkey, etc. In Ensembl 84,
CYP3A4 has 148 orthologs from 60 species of chordates including 8 species of non-human primates, 8 species of rodents, 14 species of Laurasiatheria, 35 species of placental mammals, 7 species of Sauropsida, and 11 species of fishes (
Figure 4 and
Table S3). In GeneCards 4.1.1,
CYP3A4 has orthologs in 16 species including chimpanzee, cattle, dog, mouse, rat, etc. (
Table S4).
CYP3A5 is conserved in chimpanzee, Rhesus monkey, mouse, rat, and fruitfly. In NCBI HomoloGene 68,
CYP3A5 has 13 homologs in 5 species including chimpanzee, Rhesus monkey, mouse, rat, etc. Based on NCBI Annotation Pipeline, 10 organisms have orthologs with
CYP3A5 (
Table S2). These include chimpanzee, Rhesus monkey, etc. In Ensembl 84,
CYP3A5 has 153 orthologs from 62 species of chordates including 10 species of non-human primates, 8 species of rodents, 14 species of Laurasiatheria, 35 species of placental mammals, 7 species of Sauropsida, and 11 species of fishes (
Figure 4 and
Table S3). In GeneCards 4.1.1,
CYP3A5 has orthologs in 13 species including chimpanzee, cattle, dog, mouse, rat, opossum, etc. (
Table S4).
CYP3A7 is conserved in chimpanzee, mouse, rat, chicken, zebrafish, frog, and rice. In NCBI HomoloGene 68,
CYP3A7 has 9 homologs in 7 species including chimpanzee, mouse, rat, chicken, Xenopus, etc. Based on NCBI Annotation Pipeline, 7 organisms have orthologs with
CYP3A7, including chimpanzee, Rhesus monkey, macaque, and gorilla, etc. In Ensembl 84,
CYP3A7 has 150 orthologs from 61 species of chordates including 9 species of non-human primates, 8 species of rodents, 14 species of Laurasiatheria, 36 species of placental mammals, 7 species of Sauropsida, and 11 species of fishes (
Figure 4 and
Table S3). In GeneCards 4.1.1,
CYP3A7 has orthologs in 14 species including chimpanzee, cattle, dog, mouse, rat, opossum,
C. elegans, etc. (
Table S4).
CYP3A43 is conserved in primates only. In NCBI HomoloGene 68,
CYP3A43 has only 2 homologs in two primates including chimpanzee and Rhesus monkey. Based on NCBI Annotation Pipeline, 13 organisms have orthologs with
CYP3A43 (
Table S2). These mainly include non-human primates and Ord’s kangaroo rat. In Ensembl 84,
CYP3A43 has 148 orthologs from 60 species of chordates including 8 species of non-human primates, 8 species of rodents, 14 species of Laurasiatheria, 35 species of placental mammals, 7 species of Sauropsida, and 11 species of fishes (
Figure 4 and
Table S3). In GeneCards 4.1.1,
CYP3A43 has orthologs in 12 species including chimpanzee, dog, mouse, zebrafish, fruitfly,
C. elegans, etc. (
Table S4).
There is a cluster of human
CYP4 genes on chromosome 1p33, including
CYP4A11,
4A22,
4B1,
4A26P,
4A27P,
4A43P,
4A44P,
4X1,
4Z1, and
4Z2P. The CYP4 proteins are the major fatty acid ω-hydorxylation which can hydroxylate the terminal ω-carbon and the ω-1 position of saturated and unsaturated fatty acids. They can also catalyze the ω-hydroxylation of various prostaglandins (PGs).
CYP4A11 is conserved in chimpanzee, mouse, and rat. In NCBI HomoloGene 68,
CYP4A11 has 5 homologs in 3 species including chimpanzee, mouse, and rat. Based on NCBI Annotation Pipeline, 26 organisms have orthologs with
CYP4A11 (
Table S2). These mainly include non-human primates, rodents, even-toed ungulates and whales, other mammals, birds, fishes, other vertebrates, etc. In Ensembl 84,
CYP4A11 has 134 orthologs from 57 species of chordates including 10 species of non-human primates, 8 species of rodents, 11 species of Laurasiatheria, 32 species of placental mammals, 7 species of Sauropsida, and 11 species of fishes (
Figure 4 and
Table S3). In GeneCards 4.1.1,
CYP4A11 has orthologs in 13 species including chimpanzee, dog, mouse, rat, fruitfly,
C. elegans, etc. (
Table S4).
CYP4A22 is conserved in chimpanzee, Rhesus monkey, dog, cow, mouse, rat, chicken,
A. thaliana, and rice. In NCBI HomoloGene 68,
CYP4A22 has 13 homologs in 9 species, including chimpanzee, Rhesus monkey, dog, cow, mouse, rat, etc. Based on NCBI Annotation Pipeline, 3 organisms have orthologs with
CYP4A22, including chimpanzee, pygmy chimpanzee, and Rhesus monkey (
Table S2). In Ensembl 84,
CYP4A22 has 136 orthologs from 58 species of chordates including 11 species of non-human primates, 8 species of rodents, 11 species of Laurasiatheria, 33 species of placental mammals, 7 species of Sauropsida, and 11 species of fishes (
Figure 4 and
Table S3). In GeneCards 4.1.1,
CYP4A22 has orthologs in 11 species including chimpanzee, cattle, dog, mouse, rat,
A. thaliana, etc. (
Table S4).
CYP4B1 is conserved in chimpanzee, Rhesus monkey, dog, cow, mouse, rat, chicken, zebrafish, and frog. In NCBI HomoloGene 68,
CYP4B1 has 12 homologs in 9 species, including chimpanzee, Rhesus monkey, dog, cattle, mouse, rat, chicken, zebrafish, etc. Based on NCBI Annotation Pipeline, 79 organisms have orthologs with
CYP4B1 (
Table S2). These include non-human primates, rodents, even-toed ungulates and whales, other mammals, birds, fishes, other vertebrates, etc. In Ensembl 84,
CYP4B1 has 67 orthologs from 54 species of chordates including 10 species of non-human primates, 8 species of rodents, 12 species of Laurasiatheria, 35 species of placental mammals, 1 species of Sauropsida, and 11 species of fishes (
Figure 4 and
Table S3). In GeneCards 4.1.1,
CYP4B1 has orthologs in 13 species including chimpanzee, cattle, dog, mouse, rat, zebrafish,
C. elegans, etc. (
Table S4).
CYP4X1 is a so-called “orphan” enzyme, but it converts the natural endocannabinoid anandamide to a single monooxygenated product and also metabolizes AA.
CYP4X1 is mainly detected in the trachea, aorta, heart, liver, breast, brain, and prostate.
CYP4X1 is conserved in chimpanzee, Rhesus monkey, dog, cow, mouse, rat,
A. thaliana, and rice. In NCBI HomoloGene 68,
CYP4X1 has 13 homologs in 8 species, including chimpanzee, dog, mouse, rat, etc. Based on NCBI Annotation Pipeline, 53 organisms have orthologs with
CYP4X1 (
Table S2). These include non-human primates, rodents, even-toed ungulates and whales, other mammals, birds, fishes, other vertebrates, etc. In Ensembl 84,
CYP4X1 has 86 orthologs from 62 species of chordates including 10 species of non-human primates, 8 species of rodents, 14 species of Laurasiatheria, 37 species of placental mammals, 7 species of Sauropsida, and 11 species of fishes (
Figure 4 and
Table S3). In GeneCards 4.1.1,
CYP4X1 has orthologs in 10 species including chimpanzee, cattle, dog, mouse, rat,
A. thaliana, etc. (
Table S4).
CYP4Z1 is responsible for the in-chain hydroxylation of myristic acid and lauric acid.
CYP4Z1 is primarily expressed in mammary tissue.
CYP4Z1 is conserved in fish, mammal, and primate. In NCBI HomoloGene 68,
CYP4Z1 has only one homolog—chimpanzee
CYP4Z1. Based on NCBI Annotation Pipeline, 14 organisms have orthologs with
CYP4Z1 (
Table S2). These mainly include non-human primates, bottlenosed dolphin, etc. In Ensembl 84,
CYP4Z1 has 54 orthologs from 38 species of chordates including 5 species of non-human primates, 0 species of rodents, 0 species of Laurasiatheria, 5 species of placental mammals, 7 species of Sauropsida, and 11 species of fishes (
Figure 4 and
Table S3). In GeneCards 4.1.1,
CYP4Z1 has orthologs in 5 species including chimpanzee, mouse, opossum, platypus, and lizard (
Table S4).
The human genome has 6 functional CYP4F genes including CYP4F2, 4F3, 4F8, 4F11, 4F12, and 4F22 and 5 pseudogenes including 4F9P, 4F10P, 4F23P, 4F24P, and 4F36P, which are located at chromosome 19p13.12. The Entrez Gene cytogenetic band is chromosome 19p13.1. The CYP4F subfamily is able to metabolize several important endogenous eicosanoids such as AA, PGs, and leukotriene B4 (LTB4). These compounds can regulate many physiological functions such as inflammation and vasoconstriction. Both CYP4F3A and 4F3B catalyze LTB4 and AA ω-hydroxylation. CYP4Fs can convert AA to 20-HETE that regulates renal tubular and vascular functions. The ω-hydroxylated LTB4 is further metabolized to form 20-carboxy-LTB4, which can undergo B-oxidation from its ω-side and along with traditional β-oxidation from the C1 carbon, thus inactivating this pro-inflammatory agent. CYP4Fs appear to have a minor role in the biotransformation of therapeutic drugs. CYP4F2 can ω-hydroxylate vitamin E and vitamin K1 phytyl side chains, suggesting that this enzyme may play a role in the regulation of vitamin E status and synthesis of vitamin K-dependent clotting factors. CYP4F11 metabolizes erythromycin and ethylmorphine and CYP4F12 metabolizes ebastine with contribution from CYP2J2.
CYP4F2 is a major hepatic ω-hydroxylase that hydroxylates LTB
4 and AA, inactivating LTB
4 and forming the potent vasoconstrictor 20-hydroxyeicosatetraenoic acid (HETE), respectively.
CYP4F2 is part of a cluster of the
CYP4F genes located on the band of chromosome 19p13.12.
CYP4F2 is conserved in chimpanzee, Rhesus monkey, cow, mouse, rat, and zebrafish. In NCBI HomoloGene 68,
CYP4F2 has 8 homologs, including chimpanzee, Rhesus monkey, cattle, mouse, rat, zebrafish, etc. Based on NCBI Annotation Pipeline, 5 organisms have orthologs with
CYP4F2 (
Table S2). These include Rhesus monkey, yak, western gorilla, pygmy chimpanzee, and horse. In Ensembl 84,
CYP4F2 has 56 orthologs from 27 species of chordates including 6 species of non-human primates, 4 species of rodents, 8 species of Laurasiatheria, 18 species of placental mammals, 6 species of Sauropsida, and 0 species of fishes (
Figure 4 and
Table S3). In GeneCards 4.1.1,
CYP4F2 has orthologs in 12 species including chimpanzee, cattle, mouse, rat, zebrafish, fruitfly,
C. elegans, etc. (
Table S4).
CYP4F3 catalyzes the ω-hydroxylation of LTB
4 and AA. The gene produces two tissue-specific splice variants, CYP4F3A (myeloid form) and CYP4F3B (liver form), but both transcripts are present in other tissues.
CYP4F3 is conserved in chimpanzee, Rhesus monkey, dog, cow, mouse, and rat. In NCBI HomoloGene 68,
CYP4F3 has 6 homologs in 6 species including chimpanzee, Rhesus monkey, cow, dog, mouse, and rat. Based on NCBI Annotation Pipeline, 15 organisms have orthologs with
CYP4F3 (
Table S2). These mainly include non-human primates, rodents, even-toed ungulates and whales, other mammals, birds, fishes, other vertebrates, etc. In Ensembl 84,
CYP4F3 has 30 orthologs from 19 species of chordates including 2 species of non-human primates, 4 species of rodents, 3 species of Laurasiatheria, 10 species of placental mammals, 6 species of Sauropsida, and 0 species of fishes (
Figure 4 and
Table S3). In GeneCards 4.1.1,
CYP4F3 has orthologs in 12 species including chimpanzee, dog, mouse, rat, zebrafish, fruitfly,
C. elegans, etc. (
Table S4).
CYP4F8 catalyzes the ω-2 hydroxylation of AA and three stable PGH
2 analogs but not PGD
2, E
1, E
2, and F
2α and LTB
4.
CYP4F8 is mainly expressed in epidermis, hair follicles, sweat glands, corneal epithelium, proximal renal tubules, and epithelial lining of the gut and urinary tract.
CYP4F8 is conserved in chimpanzee, Rhesus monkey, mouse, rat, and
A. thaliana. In NCBI HomoloGene 68,
CYP4F8 has 12 homologs in 5 species including chimpanzee, Rhesus monkey, mouse, rat, etc. Based on NCBI Annotation Pipeline, 7 organisms have orthologs with
CYP4F8 (
Table S2). These include Rhesus monkey, chimpanzee, green monkey, etc. In Ensembl 84,
CYP4F8 has 56 orthologs from 28 species of chordates including 7 species of non-human primates, 4 species of rodents, 8 species of Laurasiatheria, 19 species of placental mammals, 6 species of Sauropsida, and 0 species of fishes (
Figure 4 and
Table S3). In GeneCards 4.1.1,
CYP4F8 has orthologs in 6 species including chimpanzee, mouse, rat, fruitfly,
C. elegans, and
A. thaliana (
Table S4).
CYP4F11 metabolizes LTB
4 and AA but at a much lower activity than CYP4F2 and 4F3A/B. The
CYP4F11 gene is conserved in chimpanzee, mouse, rat, and chicken. In NCBI HomoloGene 68,
CYP4F11 has 4 homologs in 4 species including chimpanzee
CYP4F11, mouse and rat
Cyp4f40, and chicken
CYP4F22. Based on NCBI Annotation Pipeline, 12 organisms have orthologs with
CYP4F11 (
Table S2). These include chimpanzee
CYP4F11, Sumatran orangutan, sooty mangabey, pygmy chimpanzee, etc. In Ensembl 84,
CYP4F11 has 58 orthologs from 28 species of chordates including 7 species of non-human primates, 4 species of rodents, 8 species of Laurasiatheria, 19 species of placental mammals, 6 species of Sauropsida, and 0 species of fishes (
Figure 4 and
Table S3). In GeneCards 4.1.1,
CYP4F11 has orthologs in 11 species including chimpanzee, cattle, mouse, rat, zebrafish, fruitfly,
C. elegans, etc. (
Table S4).
CYP4F12 catalyzes the hydroxylation of AA at C18 and C20, ω1 through ω3-hydroxylations of 9,11-diazo-PGH
2 and 9,11-methanoepoxy-PGH
2, and ω-oxidation of LTB
4. In addition, CYP4F12 contributes to astemizole
O-demethylation and hydroxylation of ebastine and terfenadine, with contributions from CYP3A4 and 2J2.
CYP4F12 is mainly expressed in the liver, kidney, colon, small intestine, urogenital tract, and heart.
CYP4F12 is conserved in chimpanzee, Rhesus monkey, mouse, rat,
A. thaliana, and rice. In NCBI HomoloGene 68,
CYP4F12 has 7 homologs in 6 species including chimpanzee, Rhesus monkey, mouse, rat, etc. Based on NCBI Annotation Pipeline, 7 organisms have orthologs with
CYP4F12 (
Table S2). These include chimpanzee, golden snub-nosed monkey, western gorilla, olive baboon, pygmy chimpanzee, etc. In Ensembl 84,
CYP4F12 has 58 orthologs from 28 species of chordates including 7 species of non-human primates, 4 species of rodents, 8 species of Laurasiatheria, 19 species of placental mammals, 6 species of Sauropsida, and 0 species of fishes (
Figure 4 and
Table S3). In GeneCards 4.1.1,
CYP4F12 has orthologs in 14 species including chimpanzee, dog, mouse, rat, zebrafish, fruitfly,
C. elegans, etc. (
Table S4).
CYP4F22 catalyzes the ω-3 hydroxylation of 20:4
n-6 and synthesizes the ω-hydroxy fatty acids in the ceramide.
CYP4F22 is conserved in chimpanzee, Rhesus monkey, dog, cow, mouse, rat, frog,
A. thaliana, and rice. In NCBI HomoloGene 68,
CYP4F22 has 9 homologs in 9 species including chimpanzee, Rhesus monkey, dog, cow, mouse, rat, frog, etc. Based on NCBI Annotation Pipeline, 92 organisms have orthologs with
CYP4F22 (
Table S2). These include non-human primates, rodents, even-toed ungulates and whales, other mammals, birds, fishes, other vertebrates, etc. In Ensembl 84,
CYP4F22 has 44 orthologs from 43 species of chordates including 9 species of non-human primates, 6 species of rodents, 13 species of Laurasiatheria, 33 species of placental mammals, 6 species of Sauropsida, and 0 species of fishes (
Figure 4 and
Table S3). In GeneCards 4.1.1,
CYP4F22 has orthologs in 15 species including chimpanzee, dog, mouse, rat, zebrafish,
A. thaliana, etc. (
Table S4).
CYP4V2 is a selective ω-hydroxylase of saturated, medium-chain fatty acids with relatively high catalytic efficiency toward myristic acid and also hydroxylates the ω-3 polyunsaturated fatty acids such as docosahexaenoic acid and eicosapentaenoic acid.
CYP4V2 is an unusual
CYP4 member in that it resides on chromosome 4q35.2, separate from the
CYP4ABXZ and
CYP4F clusters on chromosomes 1 and 19. The mRNA of
CYP4V2 is found in the heart, brain, placenta, lung, liver, skeletal muscle, kidney, pancreas, retina, retinal pigment epithelium, and lymphocytes. The protein has very low sequence identity (31%–37%) to other CYP4 members.
CYP4V2 is conserved in chimpanzee, Rhesus monkey, dog, cow, mouse, rat, chicken, fruitfly, mosquito, frog, and
C. elegans. In NCBI HomoloGene 68,
CYP4V2 has 12 homologs in 11 species including chimpanzee, Rhesus monkey, dog, cow, mouse, rat, frog, etc. Based on NCBI Annotation Pipeline, 153 organisms have orthologs with
CYP4V2 (
Table S2). These include non-human primates, rodents, even-toed ungulates and whales, other mammals, birds, fishes, other vertebrates, etc. In Ensembl 84,
CYP4V2 has 76 orthologs from 61 species of chordates including 10 non-human primates, 8 species of rodents, 13 species of Laurasiatheria, 34 species of placental mammals, 7 species of Sauropsida, and 10 species of fishes (
Figure 4 and
Table S3). In GeneCards 4.1.1,
CYP4V2 has orthologs in 16 species including chimpanzee, dog, mouse, rat, zebrafish, fruitfly,
C. elegans, etc. (
Table S4).
TBXAS1 catalyzes the conversion of PGH
2 to thromboxane, which causes vasoconstriction and platelet aggregation.
TBXAS1 maps to chromosome 7q34-q35. TBXAS1 is expressed in platelets, erythroleukemia cells, human monocytes, leukocytes, and kidney interstitial dendritic reticulum cells, with a low expression in the liver and lung.
TBXAS1 is conserved in dog, cow, mouse, rat, chicken, zebrafish, fruitfly,
A. thaliana, rice, and frog. In NCBI HomoloGene 68,
TBXAS1 has 18 homologs in 10 species including dog, cattle, mouse, rat, chicken, Xenopus, zebrafish etc. Based on NCBI Annotation Pipeline, 154 organisms have orthologs with
TBXAS1 (
Table S2). These include non-human primates, rodents, even-toed ungulates and whales, other mammals, birds, fishes, other vertebrates, etc. In Ensembl 84,
TBXAS1 has 106 orthologous genes from 61 species of chordates including 11 species of non-human primates, 8 species of rodents, 12 species of Laurasiatheria, 35 species of placental mammals, 7 species of Sauropsida, and 11 species of fishes (
Figure 4 and
Table S3). In GeneCards 4.1.1,
TBXAS1 has orthologs in 16 species including chimpanzee, cattle, dog, mouse, zebrafish, fruitfly,
C. elegans, etc. (
Table S4).
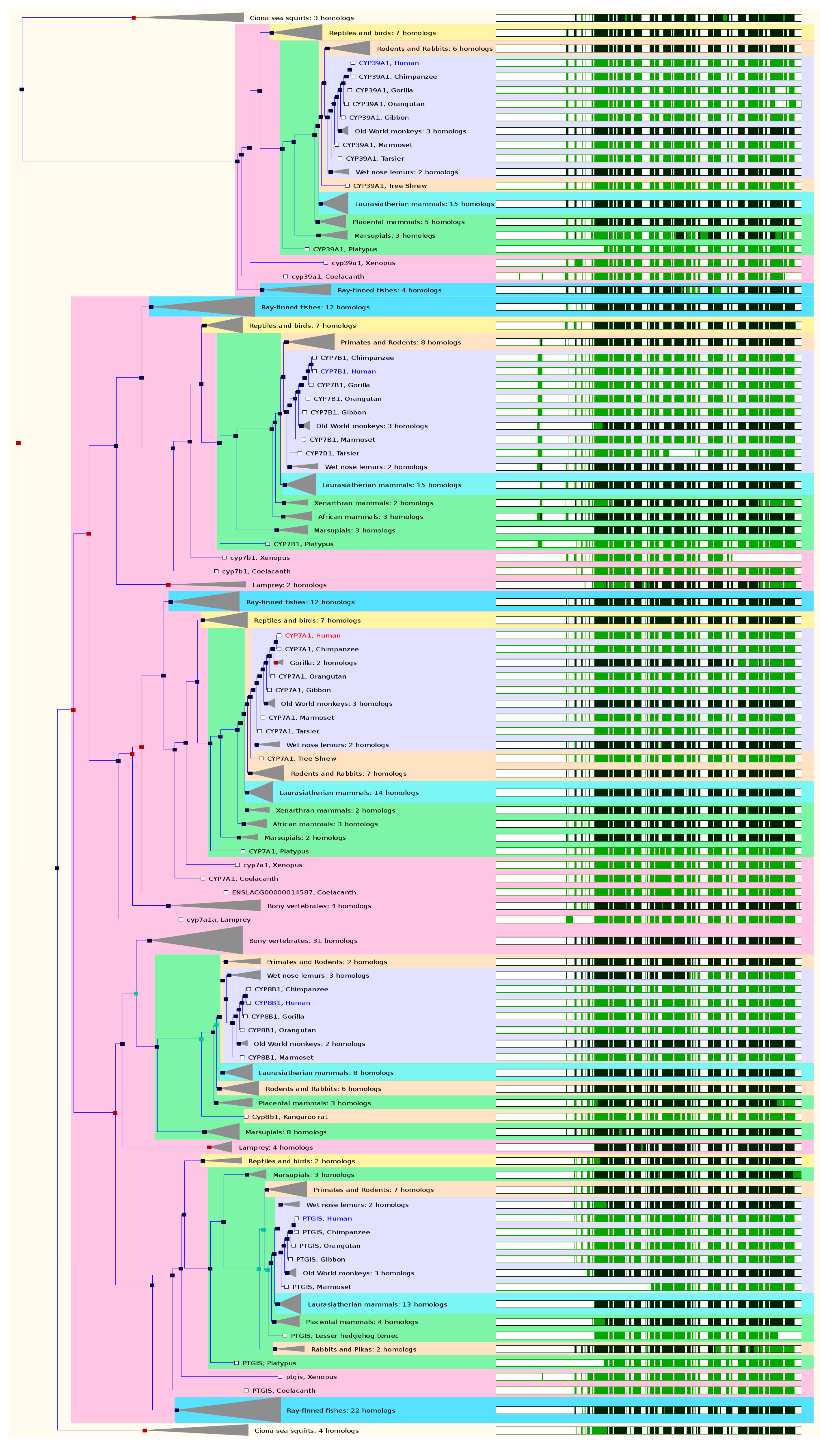
2.5. The Paralogs, Homologs and Orthologs of CYP7, 8, and 39 Families
CYP7B1 shares 40% amino acid sequence identity with CYP7A1. Human
CYP7A1 and
7B1 share identical paralogs and orthologs. CYP7A1 (called cholesterol 7α-hydroxylase) catalyzes the first and major rate-limiting step in the classical, neutral pathway for bile acid biosynthesis. The
CYP7A1 gene maps to chromosome 8q11-q12 and its promoter region contains recognition sequences for a number of liver-specific transcription factors. The gene spans 10,059 bases with 7 exons, encoding a 504-amino acid enzyme. In both GeneCards 4.1.1 and Ensembl 84,
CYP7A1 has 4 paralogs:
CYP7B1,
PTGIS,
8B1, and
39A1 (
Figure 5 and
Table 1).
CYP7A1 is conserved in chimpanzee, Rhesus monkey, dog, cow, mouse, rat, chicken, zebrafish, and frog. In NCBI HomoloGene 68,
CYP7A1 has 9 homologs in 9 species including chimpanzee, Rhesus monkey, dog, cow, mouse, rat, chicken, Xenopus, and zebrafish. Based on NCBI Annotation Pipeline, 163 organisms have orthologs with
CYP7A1 (
Table S2). These include non-human primates, rodents, even-toed ungulates and whales, other mammals, birds, fishes, other vertebrates, etc. In Ensembl 84,
CYP7A1 has 68 orthologs from 64 species including 11 species of non-human primates, 8 species of rodents, 14 species of Laurasiatheria, 38 species of placental mammals, 7 species of Sauropsida, and 11 species of fishes (
Figure 5 and
Table S3). In GeneCards 4.1.1,
CYP7A1 has orthologs in 12 species including chimpanzee, cattle, dog, mouse, rat, zebrafish, etc. (
Table S4).
CYP7B1 is involved in metabolism of oxysterols, sex hormones and neurosteroids. The gene spans 211,029 bases with 6 exons, encoding a 506-amino acid enzyme. CYP7B1 shares 40% amino acid sequence identity with CYP7A1.
CYP7B1 is conserved in chimpanzee, Rhesus monkey, dog, cow, mouse, rat, chicken, zebrafish, and frog. In NCBI HomoloGene 68,
CYP7B1 has 9 homologs in 9 species including chimpanzee, Rhesus monkey, dog, mouse, rat, etc. Based on NCBI Annotation Pipeline, 150 organisms have orthologs with
CYP7B1 (
Table S2). These include non-human primates, rodents, even-toed ungulates and whales, other mammals, birds, fishes, other vertebrates, etc. In Ensembl 84,
CYP7A1 has 70 orthologs from 65 species including 11 species of non-human primates, 8 species of rodents, 14 species of Laurasiatheria, 38 species of placental mammals, 7 species of Sauropsida, and 11 species of fishes (
Figure 5 and
Table S3). In GeneCards 4.1.1,
CYP7B1 has orthologs in 12 species including chimpanzee, cattle, dog, mouse, rat, zebrafish, etc. (
Table S4).PTGIS (prostaglandin I
2 synthase) catalyzes the conversion of PGH
2 to prostacyclin, a potent vasodilator and inhibitor of platelet aggregation.
PTGIS maps to chromosome 20q13.13.
PTGIS is conserved in chimpanzee, Rhesus monkey, dog, cow, mouse, rat, zebrafish, and frog. In NCBI HomoloGene 68,
PTGIS has 8 homologs in 8 species including chimpanzee, Rhesus monkey, dog, cattle, mouse, rat, frog, and zebrafish. Based on NCBI Annotation Pipeline, 100 organisms have orthologs with
PTGIS/
CYP8A1 (
Table S2). These include non-human primates, rodents, even-toed ungulates and whales, other mammals, birds, fishes, other vertebrates, etc. In Ensembl 84,
PTGIS has 70 orthologs from 56 species including 9 species of non-human primates, 8 species of rodents, 13 species of Laurasiatheria, 35 species of placental mammals, 2 species of Sauropsida (Anole lizard and Chinese softshell turtle), and 11 species of fishes (
Figure 5 and
Table S3). In GeneCards 4.1.1, human
PTGIS has orthologs in 11 species including chimpanzee, cattle, dog, mouse, rat, tropical clawed frog, zebrafish, etc. (
Table S4).
CYP8B1 is involved in bile acid synthesis and is responsible for the conversion of 7α-hydroxy-4-cholesten-3-one into 7α,12α-dihydroxy-4-cholesten-3-one.
CYP8B1 maps to chromosome 3p22.1 and it contains only one single exon and lacks introns.
CYP8B1 is conserved in chimpanzee, Rhesus monkey, dog, mouse, rat, chicken, zebrafish, and frog. In NCBI HomoloGene 68,
CYP8B1 has 11 homologs in 8 species including chimpanzee, Rhesus monkey, dog, mouse, rat, etc. Based on NCBI Annotation Pipeline, 120 organisms have orthologs with
CYP8B1 (
Table S2). These include non-human primates, rodents, even-toed ungulates and whales, other mammals, birds, fishes, other vertebrates, etc. In Ensembl 84,
CYP8B1 has 71 orthologs from 50 species including 8 species of non-human primates, 8 species of rodents, 8 species of Laurasiatheria, 27 species of placental mammals, 6 species of Sauropsida, 10 species of fishes (
Figure 5 and
Table S3). In GeneCards 4.1.1,
CYP8B1 has orthologs in 13 species including chimpanzee, cattle, dog, mouse, rat, opossum, zebrafish, etc. (
Table S4).
CYP39A1 catalyzes 7α-hydroxylation of oxysterols including 25-hydroxycholesterol, 27-hydroxycholesterol and 24-hydroxycholesterol, while CYP7B1 is called oxysterol 7α-hydroxylase 1.
CYP39A1 maps to chromosome 6p12.3, spans 103,079 bases, and contains 14 exons.
CYP39A1 is conserved in chimpanzee, Rhesus monkey, dog, cow, mouse, rat, chicken, zebrafish,
M. oryzae, and frog. In NCBI HomoloGene 68,
CYP39A1 has 10 homologs in 10 species including chimpanzee, Rhesus monkey, dog, mouse, rat, frog, etc. Based on NCBI Annotation Pipeline, 145 organisms have orthologs with
CYP39A1 (
Table S2). These include non-human primates, rodents, even-toed ungulates and whales, other mammals, birds, fishes, other vertebrates, etc. In Ensembl 84,
CYP39A1 has 58 orthologs from 55 species including 10 species of non-human primates, 7 species of rodents, 14 species of Laurasiatheria, 37 species of placental mammals, 7 species of Sauropsida (birds and reptiles), and 3 species of fishes (
Figure 5 and
Table S3). In GeneCards 4.1.1,
CYP39A1 has orthologs in 13 species including chimpanzee, dog, mouse, rat, zebrafish, etc. (
Table S4).
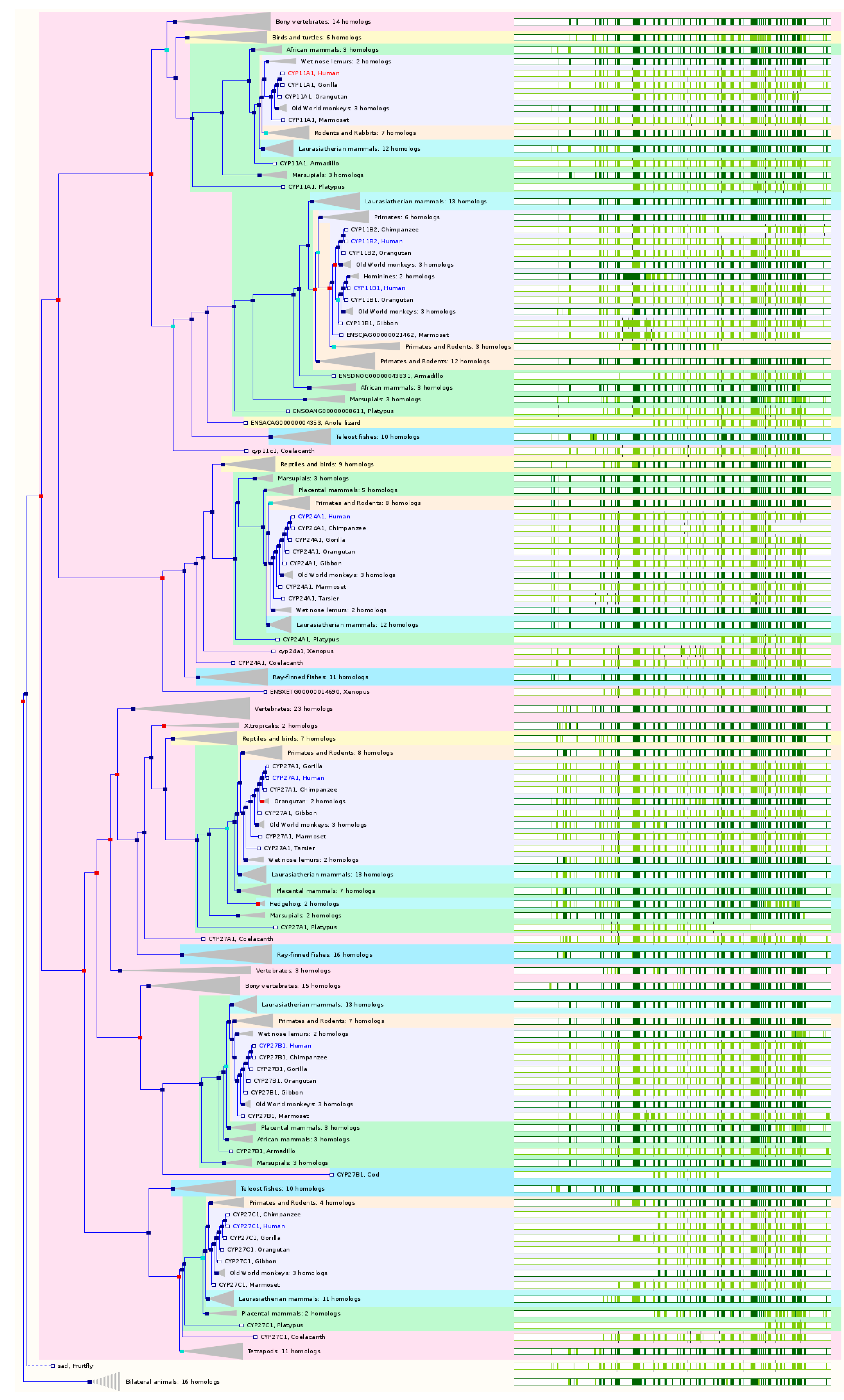
2.6. The Paralogs, Homologs and Orthologs of CYP11, 19, 24, 27, and 46 Families
The CYP11 members are important enzymes that participate in steroid biosynthesis and metabolism. The production of glucocorticoids and mineralocorticoids occurs in the adrenal gland and the final steps are catalyzed by three mitochondrial CYPs, namely CYP11A1, 11B1, and 11B2. CYP11B1 shows close homology to the CYP11B2 gene, which encodes aldosterone synthase and is normally expressed only in the zona glomerulosa. Both CYP11B genes map to chromosome 8q21, while CYP11A1 is located on chromosome 15q24.1. All CYP11 members are mitochondrial enzymes. CYP19A1, 24A1, CYP27 family, and 46A1 are the paralogs of CYP11 family members.
CYP11A1 catalyzes the conversion of cholesterol to pregnenolone, which is the first and rate-limiting step in the synthesis of the steroid hormones.
CYP11A1 maps to chromosome 15q23-q24.
CYP11A1 is conserved in chimpanzee, dog, cow, mouse, rat, chicken, zebrafish, and frog. In NCBI HomoloGene 68,
CYP11A1 has 9 homologs in 8 species including chimpanzee, dog, cow, mouse, rat, etc. Based on NCBI Annotation Pipeline, 148 organisms have orthologs with
CYP11A1. These include non-human primates, rodents, even-toed ungulates and whales, other mammals, birds, fishes, other vertebrates, etc. In Ensembl 84,
CYP11A1 has 61 orthologs from 58 species including 8 species of non-human primates, 7 species of rodents, 12 species of Laurasiatheria, 31 species of placental mammals, 6 species of Sauropsida, 11 species of fishes (
Figure 6 and
Table S3). In GeneCards 4.1.1,
CYP11A1 has orthologs in 14 species including chimpanzee, dog, mouse, rat, zebrafish, fruitfly,
C. elegans, etc. (
Table S4).
CYP11B1 catalyzes the 11β-hydroxylation of deoxycortisol to form cortisol and also catalyzes 18 or 19-hydroxylation of steroids and the aromatization of androstendione to estrone.
CYP11B1 maps to chromosome 8q21. The gene spans 7,491 bases with 11 exons and 10 introns.
CYP11B1 is conserved in chimpanzee, Rhesus monkey, mouse, rat, zebrafish, and frog. In NCBI HomoloGene release 68,
CYP11B1 has 6 homologs: chimpanzee, Rhesus monkey, mouse, rat, frog, and zebrafish. Based on NCBI Annotation Pipeline, 23 organisms have orthologs with
CYP11B1. These include chimpanzee, macaque, Rhesus monkey, green monkey, etc. In Ensembl 84,
CYP11B1 has 64 orthologs from 52 species including 11 species of non-human primates, 8 species of rodents, 9 species of Laurasiatheria, 32 species of placental mammals, 1 species of Sauropsida (Anole lizard), and 10 species of fishes (
Figure 6 and
Table S3). In GeneCards 4.1.1,
CYP11B1 has orthologs in 13 species including chimpanzee, dog, mouse, rat, zebrafish, fruitfly,
C. elegans, etc. (
Table S4).
CYP11B2 catalyzes the conversion of 11-deoxycorticosterone to aldosterone via corticosterone and 18-hydroxycorticosterone. The gene spans 7285 bases with 10 exons and 9 introns, encoding a 503-amino acid enzyme. CYP11B2 is expressed in the adrenal cortex.
CYP11B2 is conserved in Rhesus monkey, dog, cow, mouse, and rat. In NCBI HomoloGene 68,
CYP11B2 has 6 homologs in 5 species including Rhesus monkey, dog, cattle, mouse, and rat. Based on NCBI Annotation Pipeline, 13 organisms have orthologs with
CYP11B2 (
Table S2). These include Rhesus monkey, crab-eating macaque, pygmy chimpanzee, domestic cat, cattle, etc. In Ensembl 84,
CYP11B2 has 62 orthologs from 50 species including 9 species of non-human primates, 8 species of rodents, 9 species of Laurasiatheria, 30 species of placental mammals, 1 species of Sauropsida (Anole lizard), and 10 species of fishes (
Figure 6 and
Table S3). In GeneCards 4.1.1,
CYP11B2 has orthologs in 12 species including chimpanzee, dog, mouse, rat, zebrafish, fruitfly,
C. elegans, etc. (
Table S4).
CYP19A1 catalyzes the formation of aromatic C18 estrogens from C19 androgens.
CYP19A1 maps to chromosome 15q21.1 and contains 13 exons. CYP19A1 is expressed in the ovaries, testes, placenta, fetal liver, adipose tissue, bone, vasculature smooth muscle, and brain.
CYP19A1 is conserved in chimpanzee, Rhesus monkey, dog, cow, mouse, rat, chicken, zebrafish, and frog. In NCBI HomoloGene 68,
CYP19A1 has 10 homologs in 9 species including chimpanzee, Rhesus monkey, dog, cattle, mouse, rat, etc. Based on NCBI Annotation Pipeline, 155 organisms have orthologs with
CYP19A1 (
Table S2). These include non-human primates, rodents, even-toed ungulates and whales, other mammals, birds, fishes, other vertebrates, etc. In Ensembl 84,
CYP19A1 has 75 orthologs from 62 species including 11 species of non-human primates, 8 species of rodents, 14 species of Laurasiatheria, 38 species of placental mammals, 7 species of Sauropsida, and 11 species of fishes (
Table S3). In GeneCards 4.1.1,
CYP19A1 has orthologs in 14 species including chimpanzee, dog, mouse, rat, zebrafish, fruitfly, etc. (
Table S4).
CYP24A1 is called vitamin D
3 24-hydroxylase that catalyzes the 24-hydroxylation of calcidiol (25-hydroxyvitamin D
3) and calcitriol (1α,25-dihydroxyvitamin D
3). This gene maps to chromosome 20q13.2 and contains 12 exons. The enzyme is present in both proximal and distal kidney tubules.
CYP24A1 is conserved in chimpanzee, Rhesus monkey, dog, cow, mouse, rat, chicken, zebrafish, frog, fruitfly, mosquito, and
C. elegans. In NCBI HomoloGene 68,
CYP24A1 has 15 homologs in 12 species, including chimpanzee, Rhesus monkey, dog, cattle, mouse, rat, frog, zebrafish, etc. Based on NCBI Annotation Pipeline, 163 organisms have orthologs with
CYP24A1 (
Table S2). These include non-human primates, rodents, even-toed ungulates and whales, other mammals, birds, fishes, other vertebrates, etc. In Ensembl 84,
CYP24A1 has 68 orthologs from 64 species including 11 species of non-human primates, 8 species of rodents, 12 species of Laurasiatheria, 36 species of placental mammals, 7 species of Sauropsida, and 11 species of fishes (
Figure 6 and
Table S3). In GeneCards 4.1.1,
CYP24A1 has orthologs in 14 species including chimpanzee, cattle, dog, mouse, rat, fruitfly,
C. elegans, etc. (
Table S4).
The human
CYP27 family contains three functional members:
CYP27A1,
27B1, and
27C1. Both sterol 27-hydroxylase and 25-hydroxy-D
3 1α-hydroxylase are assigned to the CYP27 family since they share >40% sequence identity, while sterol 27-hydroxylase is assigned to the A subfamily and 25-hydroxy-D
3 1α-hydroxylase to the B subfamily of CYP27 since their protein sequences are <55% identical. The paralogs of
CYP27 family include
CYP11 family,
19A1,
24A1, and
46A1 (
Figure 6). CYP27A1 is called vitamin D
3 25-hydroxylase and sterol 27-hydroxylase catalyzing the first step in the oxidation of the side chain of sterol intermediates.
CYP27A1 maps to chromosome 2q35 and contains 9 exons. Mutations in
CYP27A1 cause cerebrotendinous xanthomatosis, a rare autosomal recessive lipid storage disease.
CYP27A1 is conserved in chimpanzee, Rhesus monkey, dog, cow, mouse, rat, chicken, zebrafish, frog, fruit fly, and mosquito. In NCBI HomoloGene 68,
CYP27A1 has 16 homologs in 11 species including chimpanzee, Rhesus monkey, dog, cattle, mouse, rat, zebrafish etc. Based on NCBI Annotation Pipeline, 139 organisms have orthologs with
CYP27A1 (
Table S2). These include non-human primates, rodents, even-toed ungulates and whales, other mammals, birds, fishes, other vertebrates, etc. In Ensembl 84,
CYP27A1 has 78 orthologs from 65 species including 11 species of non-human primates, 8 species of rodents, 13 species of Laurasiatheria, 37 species of placental mammals, 7 species of Sauropsida, and 11 species of fishes (
Figure 6 and
Table S3). In GeneCards 4.1.1,
CYP27A1 has orthologs in 14 species including chimpanzee, cattle, dog, mouse, rat, zebrafish, fruitfly, etc. (
Table S4).
CYP27B1 is called 25-hydroxyvitamin D
3 1α-hydroxylase that hydroxylates 25-hydroxyvitamin D
3 at the 1α position, resulting in 1α,25-dihydroxyvitamin D
3, the active form of vitamin D
3.
CYP27B1 maps to chromosome 12q12-q14 and has 9 exons. Mutations in this gene causes vitamin D-dependent rickets, type I and hypocalcemic vitamin D-dependent rickets.
CYP27B1 is conserved in in chimpanzee, Rhesus monkey, dog, cow, mouse, rat, zebrafish, and frog. In NCBI HomoloGene 68,
CYP27B1 has 9 homologs in 9 species including chimpanzee, Rhesus monkey, dog, cattle, mouse, rat, frog, etc. Based on NCBI Annotation Pipeline, 116 organisms have orthologs with
CYP27B1 (
Table S2). These include non-human primates, rodents, even-toed ungulates and whales, other mammals, birds, fishes, other vertebrates, etc. In Ensembl 84,
CYP27B1 has 61 orthologs from 58 species including 10 species of non-human primates, 8 species of rodents, 14 species of Laurasiatheria, 36 species of placental mammals, 2 species of Sauropsida, and 11 species of fishes (
Figure 6 and
Table S3). In GeneCards 4.1.1,
CYP27B1 has orthologs in 12 species including chimpanzee, dog, mouse, rat, zebrafish, fruitfly,
C. elegans, etc. (
Table S4).
CYP27C1 maps to chromosome 2q14.3 and contains 14 exons. The gene spans 36,243 bases and encodes a 372-amino acid protein.
CYP27C1 is conserved in chimpanzee, Rhesus monkey, dog, cow, chicken, zebrafish, frog, and
M. oryzae. In NCBI HomoloGene 68,
CYP27C1 has 11 homologs in 10 species including chimpanzee, Rhesus monkey, dog, mouse, rat, zebrafish, etc. Based on NCBI Annotation Pipeline, 132 organisms have orthologs with
CYP27C1 (
Table S2). These include non-human primates, rodents, even-toed ungulates and whales, other mammals, birds, fishes, other vertebrates, etc. In Ensembl 84,
CYP27C1 has 53 orthologs from 51 species including 9 species of non-human primates, 4 species of rodents, 11 species of Laurasiatheria, 25 species of placental mammals, 7 species of Sauropsida, and 10 species of fishes (
Figure 6 and
Table S3). In GeneCards 4.1.1,
CYP27C1 has orthologs in 11 species including chimpanzee, cattle, dog, zebrafish, fruitfly, etc. (
Table S4).
CYP46A1 (called cholesterol 24-hydroxylase) converts cholesterol to 24
S-hydroxycholesterol in the brain. Cholesterol cannot pass the blood-brain barrier, but 24
S-hydroxycholesterol can be secreted in the brain into the circulation to be returned to the liver for catabolism.
CYP46A1 maps to chromosome 14q32.1.
CYP46A1 is conserved in chimpanzee, Rhesus monkey, dog, cow, mouse, rat, chicken, frog,
M. oryzae,
A. thaliana, and rice. In NCBI HomoloGene 68,
CYP46A1 has 15 homologs in 12 species including chimpanzee, Rhesus monkey, dog, mouse, rat, frog, zebrafish, etc. Based on NCBI Annotation Pipeline, 148 organisms have orthologs with
CYP46A1 (
Table S2). These include non-human primates, rodents, even-toed ungulates and whales, other mammals, birds, fishes, other vertebrates, etc. In Ensembl 84,
CYP46A1 has 69 orthologs from 57 species including 11 species of non-human primates, 7 species of rodents, 12 species of Laurasiatheria, 34 species of placental mammals, 7 species of Sauropsida, and 11 species of fishes (
Figure 4 and
Table S3). In GeneCards 4.1.1,
CYP46A1 has orthologs in 15 species including chimpanzee, cattle, dog, mouse, rat, zebrafish, fruitfly,
A. thaliana, etc. (
Table S4).
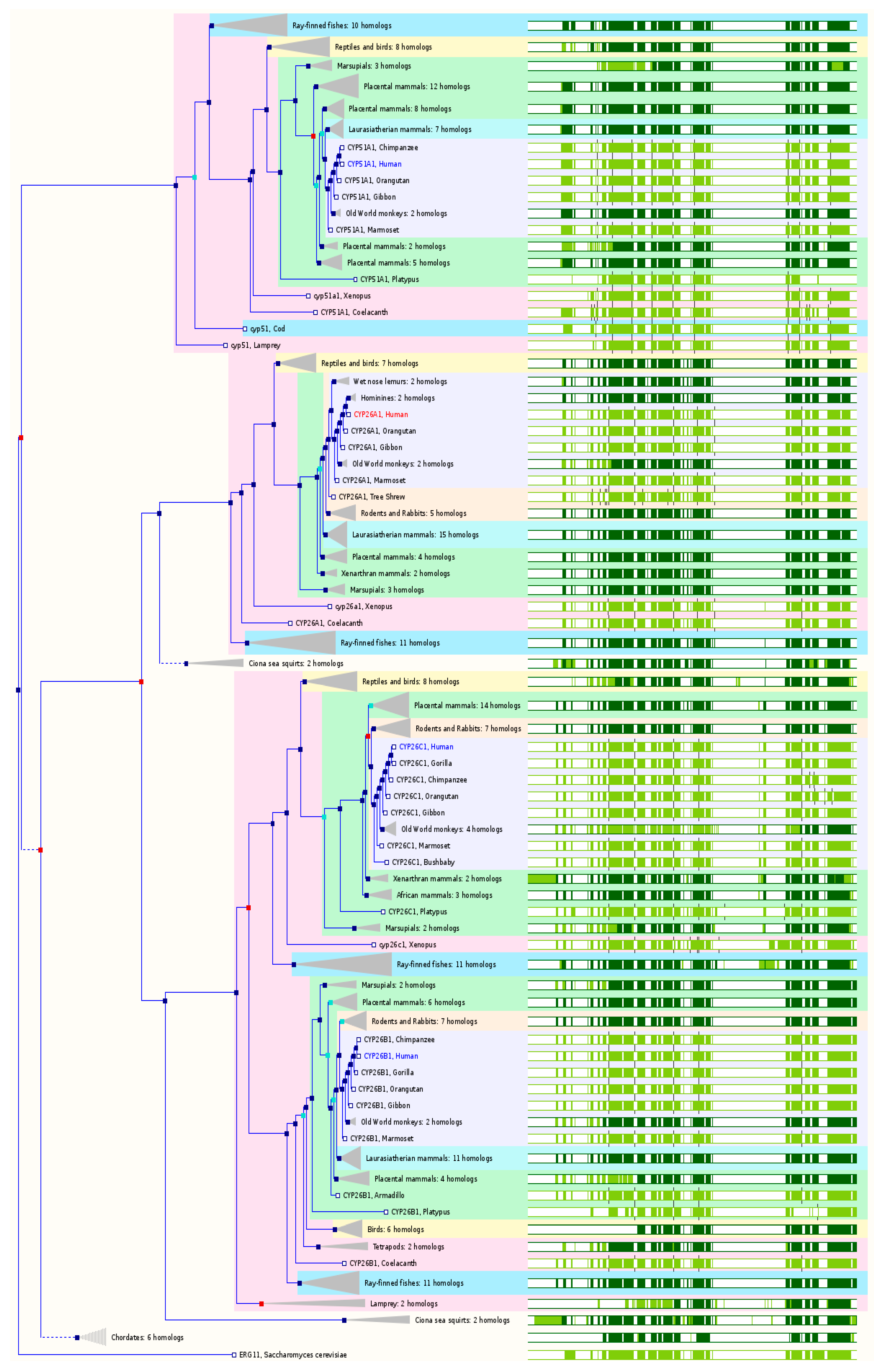
2.7. The Paralogs, Homologs and Orthologs of CYP26 and 51 Families
In the human genome, there are three members of in the
CYP26 family:
26A1,
26B1, and
26C1. These three members are all RA hydroxylases with similar substrate specificity but different tissue-specific expression patterns. CYP26A1 is called retinoic acid 4-hydroxylase with both 4-hydroxylation and 18-hydroxylation activities, acting on all-
trans-RA and its stereoisomer 9-
cis-RA.
CYP26A1 maps to chromosome 10q23-q24 and has 8 exons. In both GeneCards 4.1.1 and Ensembl 84,
CYP26 members are the paralogs of
CYP51A1 (
Figure 7 and
Table 1).
CYP26A1 has been detected in different cell lines with different tissue origins including kidney, liver, breast, intestine, and lung. Mutations in
CYP26A1 causes keratomalacia and caudal regression syndrome.
CYP26A1 is conserved in the chimpanzee, Rhesus monkey, dog, cow, mouse, rat, chicken, zebrafish, frog,
A. thaliana, and rice. In NCBI HomoloGene 68,
CYP26A1 has 19 homologs in 11 species including chimpanzee, Rhesus monkey, dog, mouse, rat, frog, zebrafish etc. Based on NCBI Annotation Pipeline, 156 organisms have orthologs with
CYP26A1 (
Table S2). These include non-human primates, rodents, even-toed ungulates and whales, other mammals, birds, fishes, other vertebrates, etc. In Ensembl 84,
CYP26A1 has 62 orthologs from 61 species including 9 species of non-human primates, 8 species of rodents, 14 species of Laurasiatheria, 35 species of placental mammals, 7 species of Sauropsida, and 11 species of fishes (
Figure 7 and
Table S3). In GeneCards 4.1.1,
CYP26A1 has orthologs in 16 species including chimpanzee, dog, mouse, rat, African clawed frog, zebrafish, fruitfly, baker’s yeast,
A. thaliana, etc. (
Table S4).
CYP26B1 is a critical regulator of all-
trans-RA levels by the specific inactivation of all-
trans-RA to hydroxylated forms. This gene maps to chromosome 2p13.2 and contains 8 exons. The gene spans 18,801 bases with 6 exons and encodes a 512-amino acid protein. Mutations in this gene are associated with radiohumeral fusions and other skeletal and craniofacial anomalies and lethal occipital encephalocele-skeletal dysplasia syndrome.
CYP26B1 is conserved in chimpanzee, Rhesus monkey, dog, cow, mouse, rat, zebrafish, frog, and
A. thaliana. In NCBI HomoloGene 68,
CYP26B1 has 19 homologs in 10 species including chimpanzee, Rhesus monkey, dog, mouse, rat, frog, zebrafish, etc. Based on NCBI Annotation Pipeline, 153 organisms have orthologs with
CYP26B1 (
Table S2). These include non-human primates, rodents, even-toed ungulates and whales, other mammals, birds, fishes, other vertebrates, etc. In Ensembl 84,
CYP26B1 has 64 orthologs from 62 species including 11 species of non-human primates, 8 species of rodents, 13 species of Laurasiatheria, 36 species of placental mammals, 6 species of Sauropsida, and 11 species of fishes (
Figure 7 and
Table S3). In GeneCards 4.1.1,
CYP26B1 has orthologs in 13 species including chimpanzee, cattle, dog, mouse, rat, tropical clawed frog, zebrafish, fruitfly, baker’s yeast,
A. thaliana, etc. (
Table S4).CYP26C1 is involved in the catabolism of all-
trans- and 9-
cis-RA, and thus contributes to the regulation of RA levels in cells and tissues. Like
CYP26A1, this gene maps to chromosome 10q23.33. The gene spans 7,434 bases with 6 exons and encodes a 522-amino acid protein Mutations of
CYP26C1 causes focal facial dermal dysplasia 4 and focal facial dermal dysplasia.
CYP26C1 is the conserved in chimpanzee, cow, mouse, rat, chicken, zebrafish, frog, and
A. thaliana. In NCBI HomoloGene 68,
CYP26C1 has 8 homologs in 8 species including chimpanzee, cattle, mouse, rat, chicken, frog, zebrafish, etc. Based on NCBI Annotation Pipeline, 139 organisms have orthologs with
CYP26C1 (
Table S2). These include non-human primates, rodents, even-toed ungulates and whales, other mammals, birds, fishes, other vertebrates, etc. In Ensembl 84,
CYP26C1 has 63 orthologs from 60 species including 10 species of non-human primates, 8 species of rodents, 11 species of Laurasiatheria, 34 species of placental mammals, 7 species of Sauropsida, and 11 species of fishes (
Figure 7 and
Table S3). In GeneCards 4.1.1,
CYP26C1 has orthologs in 14 species including chimpanzee, dog, mouse, rat, zebrafish, fruitfly,
A. thaliana, etc. (
Table S4).
CYP51A1 is called lanosterol 14α-demethylase/sterol 14α-demethylase which are found in yeast, plants, fungi, animals and even prokaryotes, suggesting this is among the oldest of the
CYP genes. CYP51A1 is a common target of antifungal drugs (e.g., miconazole and ketoconazole), which inhibit CYP51A1 activity and formation of ergosterol. This gene has 11 exons and maps to chromosome 7q21.2.
CYP51A1 is conserved in chimpanzee, Rhesus monkey, dog, cow, mouse, rat, chicken, zebrafish, frog,
Saccharomyces cerevisiae,
Kluyveromyces lactis,
Eremothecium gossypii,
Schizosaccharomyces pombe,
M. oryzae,
A. thaliana, and rice. In NCBI HomoloGene 68,
CYP51A1 has 16 homologs in 16 species including chimpanzee, Rhesus monkey, dog, cattle, mouse, rat, frog, zebrafish, etc. Based on NCBI Annotation Pipeline, 162 organisms have orthologs with
CYP51A1 (
Table S2). These include non-human primates, rodents, even-toed ungulates and whales, other mammals, birds, fishes, other vertebrates, etc. In Ensembl 84,
CYP51A1 has 64 orthologs from 63 species including 10 species of non-human primates, 8 species of rodents, 14 species of Laurasiatheria, 37 species of placental mammals, 7 species of Sauropsida, and 11 species of fishes (
Figure 7 and
Table S3). In GeneCards 4.1.1,
CYP51A1 has orthologs in 19 species including chimpanzee, cattle, dog, mouse, rat, zebrafish, baker’s yeast, etc. (
Table S4).