Rationale and Methodology of Reprogramming for Generation of Induced Pluripotent Stem Cells and Induced Neural Progenitor Cells
Abstract
:1. Introduction
2. Rationale of Reprogramming to Induced Pluripotent Stem Cells (iPSCs)
2.1. The Nature of Cellular Reprogramming
2.2. Intrinsic and Extrinsic Factors during iPSC Reprogramming
2.3. Stoichiometric Expression of Endogenous Pluripotency Genes in iPSC Reprogramming
3. Methodology of Reprogramming to iPSCs
3.1. Screening of Candidate Transcription Factors (TFs)
3.2. Transfection of Transcription Factors
3.2.1. Biological-Based Delivery System
3.2.2. Chemical-Based Delivery System
3.2.3. Physical-Based Delivery System
3.3. Activation of Endogenous Pluripotency Genes
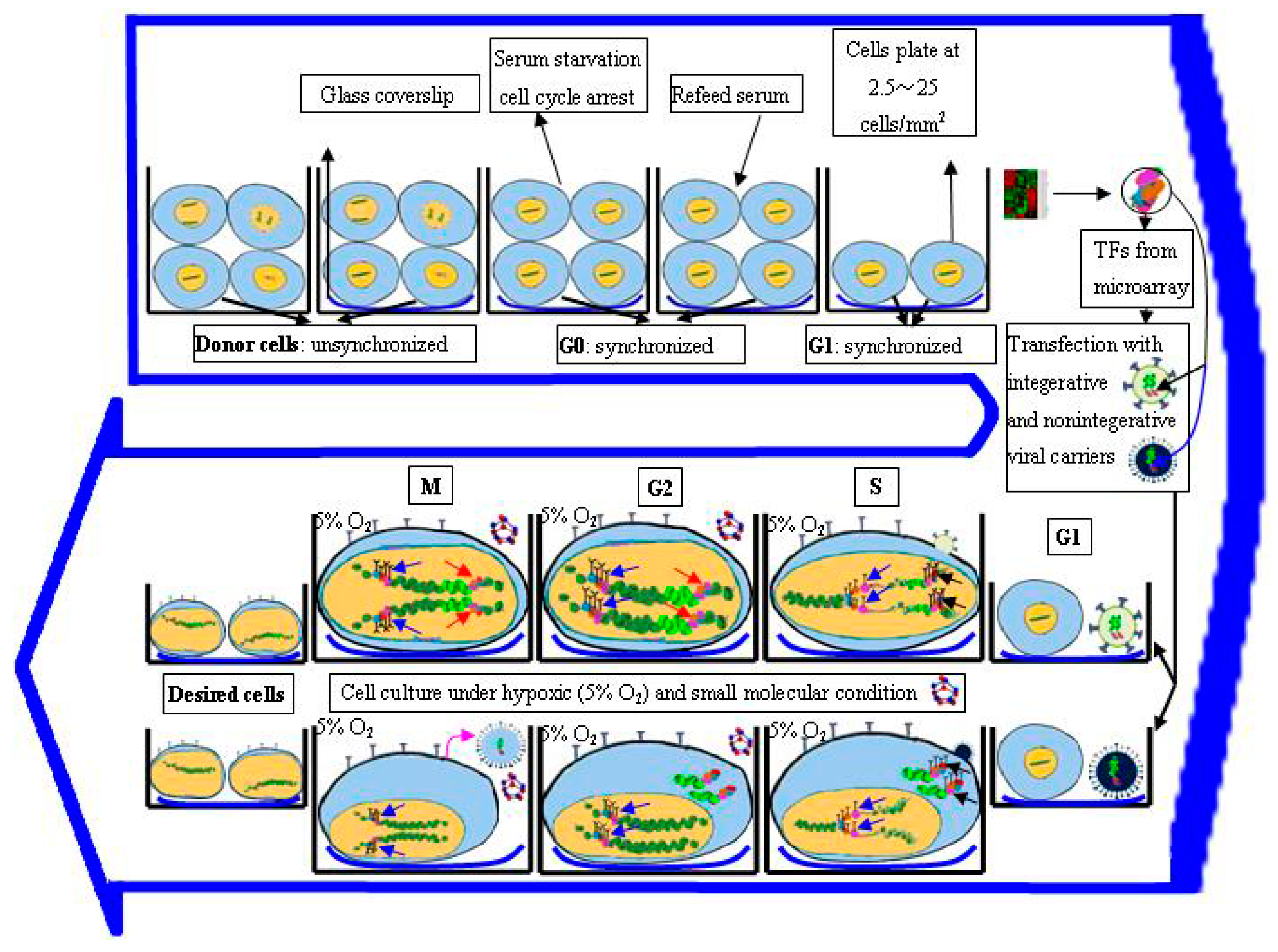
3.4. Regulation of Cell Cycle Status for Reprogramming
3.5. Optimization of Microenvironment for Reprogramming
4. Rationale of Reprogramming to iNPCs
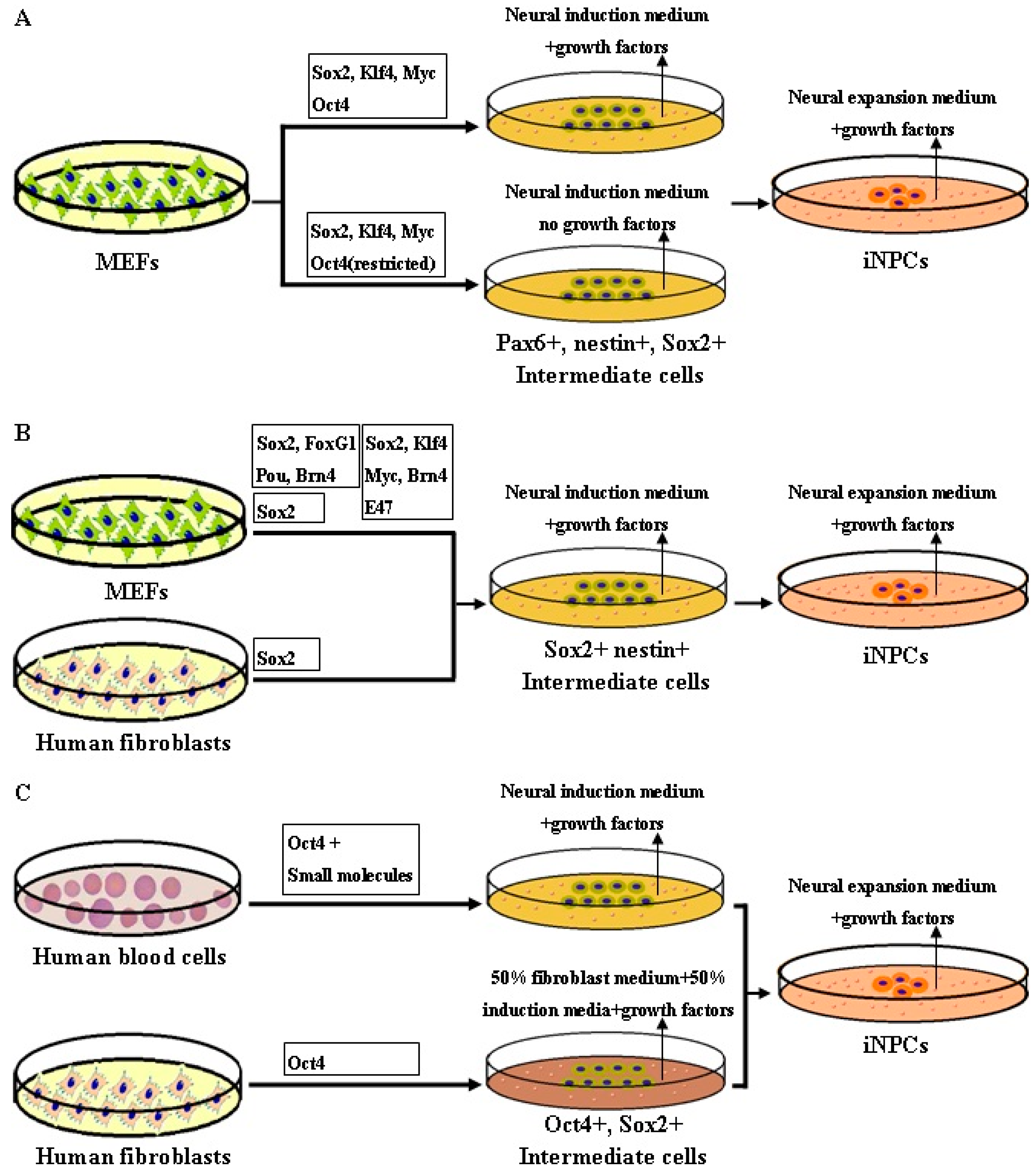
5. Methodology of Reprogramming to iNPCs
6. Chemical-Based Reprogramming to iPSCs and iNPCs
6.1. Chemical-Based Reprogramming of Somatic Cells to iPSCs
6.2. Chemical-Based Reprogramming to iNPCs
7. Concluding Remarks and Future Perspectives
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Takahashi, K.; Yamanaka, S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 2006, 126, 663–676. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, K.; Tanabe, K.; Ohnuki, M.; Narita, M.; Ichisaka, T.; Tomoda, K.; Yamanaka, S. Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell 2007, 131, 861–872. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, K.; Yamanaka, S. A decade of transcription factor-mediated reprogramming to pluripotency. Nat. Rev. Mol. Cell Biol. 2016, 17, 183–193. [Google Scholar] [CrossRef] [PubMed]
- Selvaraj, V.; Plane, J.M.; Williams, A.J.; Deng, W. Switching cell fate: The remarkable rise of induced pluripotent stem cells and lineage reprogramming technologies. Trends Biotechnol. 2010, 28, 214–223. [Google Scholar] [CrossRef] [PubMed]
- Song, B.; Sun, G.; Herszfeld, D.; Sylvain, A.; Campanale, N.V.; Hirst, C.E.; Caine, S.; Parkington, H.C.; Tonta, M.A.; Coleman, H.A.; et al. Neural differentiation of patient specific iPS cells as a novel approach to study the pathophysiology of multiple sclerosis. Stem Cell Res. 2012, 8, 259–273. [Google Scholar] [CrossRef] [PubMed]
- Israel, M.A.; Yuan, S.H.; Bardy, C.; Reyna, S.M.; Mu, Y.; Herrera, C.; Hefferan, M.P.; van Gorp, S.; Nazor, K.L.; Boscolo, F.S.; et al. Probing sporadic and familial Alzheimer’s disease using induced pluripotent stem cells. Nature 2012, 482, 216–220. [Google Scholar] [CrossRef] [PubMed]
- Dimos, J.T.; Rodolfa, K.T.; Niakan, K.K.; Weisenthal, L.M.; Mitsumoto, H.; Chung, W.; Croft, G.F.; Saphier, G.; Leibel, R.; Goland, R.; et al. Induced pluripotent stem cells generated from patients with ALS can be differentiated into motor neurons. Science 2008, 321, 1218–1221. [Google Scholar] [CrossRef] [PubMed]
- Ebert, A.D.; Yu, J.; Rose, F.F., Jr.; Mattis, V.B.; Lorson, C.L.; Thomson, J.A.; Svendsen, C.N. Induced pluripotent stem cells from a spinal muscular atrophy patient. Nature 2009, 457, 277–280. [Google Scholar] [CrossRef] [PubMed]
- Park, I.H.; Arora, N.; Huo, H.; Maherali, N.; Ahfeldt, T.; Shimamura, A.; Lensch, M.W.; Cowan, C.; Hochedlinger, K.; Daley, G.Q. Disease-specific induced pluripotent stem cells. Cell 2008, 134, 877–886. [Google Scholar] [CrossRef] [PubMed]
- Fong, C.Y.; Gauthaman, K.; Bongso, A. Teratomas from pluripotent stem cells: A clinical hurdle. J. Cell. Biochem. 2010, 111, 769–781. [Google Scholar] [CrossRef] [PubMed]
- Yamanaka, S. A fresh look at iPS cells. Cell 2009, 137, 13–17. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Efe, J.A.; Zhu, S.; Talantova, M.; Yuan, X.; Wang, S.; Lipton, S.A.; Zhang, K.; Ding, S. Direct reprogramming of mouse fibroblasts to neural progenitors. Proc. Natl. Acad. Sci. USA 2011, 108, 7838–7843. [Google Scholar] [CrossRef] [PubMed]
- Cheng, L.; Hu, W.; Qiu, B.; Zhao, J.; Yu, Y.; Guan, W.; Wang, M.; Yang, W.; Pei, G. Generation of neural progenitor cells by chemical cocktails and hypoxia. Cell Res. 2014, 24, 665–679. [Google Scholar] [CrossRef] [PubMed]
- Huangfu, D.; Osafune, K.; Maehr, R.; Guo, W.; Eijkelenboom, A.; Chen, S.; Muhlestein, W.; Melton, D.A. Induction of pluripotent stem cells from primary human fibroblasts with only Oct4 and Sox2. Nat. Biotechnol. 2008, 26, 1269–1275. [Google Scholar] [CrossRef] [PubMed]
- Shi, Y.; Desponts, C.; Do, J.T.; Hahm, H.S.; Schöler, H.R.; Ding, S. Induction of pluripotent stem cells from mouse embryonic fibroblasts by Oct4 and Klf4 with small-molecule compounds. Cell Stem Cell 2008, 3, 568–574. [Google Scholar] [CrossRef] [PubMed]
- Lyssiotis, C.A.; Foreman, R.K.; Staerk, J.; Garcia, M.; Mathur, D.; Markoulaki, S.; Hanna, J.; Lairson, L.L.; Charette, B.D.; Bouchez, L.C.; et al. Reprogramming of murine fibroblasts to induced pluripotent stem cells with chemical complementation of Klf4. Proc. Natl. Acad. Sci. USA 2009, 106, 8912–8917. [Google Scholar] [CrossRef] [PubMed]
- Hou, P.; Li, Y.; Zhang, X.; Liu, C.; Guan, J.; Li, H.; Zhao, T.; Ye, J.; Yang, W.; Liu, K.; et al. Pluripotent stem cells induced from mouse somatic cells by small-molecule compounds. Science 2013, 341, 651–654. [Google Scholar] [CrossRef] [PubMed]
- Their, M.; Wörsdörfer, P.; Lakes, Y.B.; Gorris, R.; Herms, S.; Opitz, T.; Seiferling, D.; Quandel, T.; Hoffmann, P.; Nöthen, M.M.; et al. Direct conversion of fibroblasts into stably expandable neural stem cells. Cell Stem Cell 2012, 10, 473–479. [Google Scholar] [CrossRef] [PubMed]
- Lujan, E.; Chanda, S.; Ahlenius, H.; Südhof, T.C.; Wernig, M. Direct conversion of mouse fibroblasts to self-renewing, tripotent neural precursor cells. Proc. Natl. Acad. Sci. USA 2012, 109, 2527–2532. [Google Scholar] [CrossRef] [PubMed]
- Han, D.W.; Tapia, N.; Hermann, A.; Hemmer, K.; Höing, S.; Araúzo-Bravo, M.J.; Zaehres, H.; Wu, G.; Frank, S.; Moritz, S.; et al. Direct reprogramming of fibroblasts into neural stem cells by defined factors. Cell Stem Cell 2012, 10, 465–472. [Google Scholar] [CrossRef] [PubMed]
- Ring, K.L.; Tong, L.M.; Balestra, M.E.; Javier, R.; Andrews-Zwilling, Y.; Li, G.; Walker, D.; Zhang, W.R.; Kreitzer, A.C.; Huang, Y. Direct reprogramming of mouse and human fibroblasts into multipotent neural stem cells with a single factor. Cell Stem Cell 2012, 11, 100–109. [Google Scholar] [CrossRef] [PubMed]
- Mitchell, R.R.; Szabo, E.; Benoit, Y.D.; Case, D.T.; Mechael, R.; Alamilla, J.; Lee, J.H.; Fiebig-Comyn, A.; Gillespie, D.C.; Bhatia, M. Activation of neural cell fate programs toward direct conversion of adult human fibroblasts into tri-potent neural progenitors using Oct-4. Stem Cells Dev. 2014, 23, 1937–1946. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.H.; Mitchell, R.R.; McNicol, J.D.; Shapovalova, Z.; Laronde, S.; Tanasijevic, B.; Milsom, C.; Casado, F.; Fiebig-Comyn, A.; Collins, T.J.; et al. Single transcription factor conversion of human blood fate to NPCs with CNS and PNS developmental capacity. Cell Rep. 2015, 11, 1367–1376. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Wang, L.; Huang, W.; Su, H.; Xue, Y.; Su, Z.; Liao, B.; Wang, H.; Bao, X.; Qin, D.; et al. Generation of integration-free neural progenitor cells from cells in human urine. Nat. Methods 2013, 10, 84–89. [Google Scholar] [CrossRef] [PubMed]
- Campbell, K.H.; McWhir, J.; Ritchie, W.A.; Wilmut, I. Sheep cloned by nuclear transfer from a cultured cell line. Nature 1996, 380, 64–66. [Google Scholar] [CrossRef] [PubMed]
- Tsubouchi, T.; Soza-Ried, J.; Brown, K.; Piccolo, F.M.; Cantone, I.; Landeira, D.; Bagci, H.; Hochegger, H.; Merkenschlager, M.; Fisher, A.G. DNA synthesis is required for reprogramming mediated by stem cell fusion. Cell 2013, 152, 873–883. [Google Scholar] [CrossRef] [PubMed]
- Xie, F.; Tang, X.; Zhang, Q.; Deng, C. Reprogramming human adipose tissue stem cells using epidermal keratinocyte extracts. Mol. Med. Rep. 2015, 11, 182–188. [Google Scholar] [CrossRef] [PubMed]
- Kadengodlu, P.A.; Rai, V.R. Role of nanotechnology in epigenetic reprogramming. Stem Cells Dev. 2015, 24, 535–549. [Google Scholar] [CrossRef] [PubMed]
- Wilmut, I.; Schnieke, A.E.; McWhir, J.; Kind, A.J.; Campbell, K.H. Viable offspring derived from fetal and adult mammalian cells. Nature 1997, 385, 810–813. [Google Scholar] [CrossRef] [PubMed]
- Gurdon, J.B.; Elsdale, T.R.; Fischberg, M. Sexually mature individuals of Xenopus laevis from the transplantation of single somatic nuclei. Nature 1958, 182, 64–65. [Google Scholar] [CrossRef] [PubMed]
- Gurdon, J.B. The developmental capacity of nuclei taken from intestinal epithelium cells of feeding tadpoles. J. Embryol. Exp. Morphol. 1962, 10, 622–640. [Google Scholar] [PubMed]
- Wernig, M.; Lengner, C.J.; Hanna, J.; Lodato, M.A.; Steine, E.; Foreman, R.; Staerk, J.; Markoulaki, S.; Jaenisch, R. A drug-inducible transgenic system for direct reprogramming of multiple somatic cell types. Nat. Biotechnol. 2008, 26, 916–924. [Google Scholar] [CrossRef] [PubMed]
- Hockemeyer, D.; Soldner, F.; Cook, E.G.; Gao, Q.; Mitalipova, M.; Jaenisch, R. A drug-inducible system for direct reprogramming of human somatic cells to pluripotency. Cell Stem Cell 2008, 3, 346–353. [Google Scholar] [CrossRef] [PubMed]
- Maherali, N.; Ahfeldt, T.; Rigamonti, A.; Utikal, J.; Cowan, C.; Hochedlinger, K. A high-efficiency system for the generation and study of human induced pluripotent stem cells. Cell Stem Cell 2008, 3, 340–345. [Google Scholar] [CrossRef] [PubMed]
- Yamaguchi, S.; Hirano, K.; Nagata, S.; Tada, T. Sox2 expression effects on direct reprogramming efficiency as determined by alternative somatic cell fate. Stem Cell Res. 2011, 6, 177–186. [Google Scholar] [CrossRef] [PubMed]
- Papapetrou, E.P.; Tomishima, M.J.; Chambers, S.M.; Mica, Y.; Reed, E.; Menon, J.; Tabar, V.; Mo, Q.; Studer, L.; Sadelain, M. Stoichiometric and temporal requirements of Oct4, Sox2, Klf4, and c-Myc expression for efficient human iPSC induction and differentiation. Proc. Natl. Acad. Sci. USA 2009, 106, 12759–12764. [Google Scholar] [CrossRef] [PubMed]
- Buganim, Y.; Faddah, D.A.; Cheng, A.W.; Itskovich, E.; Markoulaki, S.; Ganz, K.; Klemm, S.L.; van Oudenaarden, A.; Jaenisch, R. Single-cell expression analyses during cellular reprogramming reveal an early stochastic and a late hierarchic phase. Cell 2012, 150, 1209–1222. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, K.; Tanabe, K.; Ohnuki, M.; Narita, M.; Sasaki, A.; Yamamoto, M.; Nakamura, M.; Sutou, K.; Osafune, K.; Yamanaka, S. Induction of pluripotency in human somatic cells via a transient state resembling primitive streak-like mesendoderm. Nat. Commun. 2014, 5, 3678. [Google Scholar] [CrossRef] [PubMed]
- O’Malley, J.; Skylaki, S.; Iwabuchi, K.A.; Chantzoura, E.; Ruetz, T.; Johnsson, A.; Tomlinson, S.R.; Linnarsson, S.; Kaji, K. High-resolution analysis with novel cell-surface markers identifies routes to iPS cells. Nature 2013, 499, 88–91. [Google Scholar] [CrossRef] [PubMed]
- Ban, H.; Nishishita, N.; Fusaki, N.; Tabata, T.; Saeki, K.; Shikamura, M.; Takada, N.; Inoue, M.; Hasegawa, M.; Kawamata, S.; et al. Efficient generation of transgene-free human induced pluripotent stem cells (iPSCs) by temperature-sensitive Sendai virus vectors. Proc. Natl. Acad. Sci. USA 2011, 108, 14234–14239. [Google Scholar] [CrossRef] [PubMed]
- Najm, F.J.; Lager, A.M.; Zaremba, A.; Wyatt, K.; Caprariello, A.V.; Factor, D.C.; Karl, R.T.; Maeda, T.; Miller, R.H.; Tesar, P.J. Transcription factor-mediated reprogramming of fibroblasts to expandable, myelinogenic oligodendrocyte progenitor cells. Nat. Biotechnol. 2013, 31, 426–433. [Google Scholar] [CrossRef] [PubMed]
- Yang, N.; Zuchero, J.B.; Ahlenius, H.; Marro, S.; Ng, Y.H.; Vierbuchen, T.; Hawkins, J.S.; Geissler, R.; Barres, B.A.; Wernig, M. Generation of oligodendroglial cells by direct lineage conversion. Nat. Biotechnol. 2013, 31, 434–439. [Google Scholar] [CrossRef] [PubMed]
- Soufi, A.; Donahue, G.; Zaret, K.S. Facilitators and impediments of the pluripotency reprogramming factors’ initial engagement with the genome. Cell 2012, 151, 994–1004. [Google Scholar] [CrossRef] [PubMed]
- Koche, R.P.; Smith, Z.D.; Adli, M.; Gu, H.; Ku, M.; Gnirke, A.; Bernstein, B.E.; Meissner, A. Reprogramming factor expression initiates widespread targeted chromatin remodeling. Cell Stem Cell 2011, 8, 96–105. [Google Scholar] [CrossRef] [PubMed]
- Knoepfler, P.S.; Zhang, X.Y.; Cheng, P.F.; Gafken, P.R.; McMahon, S.B.; Eisenman, R.N. Myc influences global chromatin structure. EMBO J. 2006, 25, 2723–2734. [Google Scholar] [CrossRef] [PubMed]
- Lin, C.Y.; Lovén, J.; Rahl, P.B.; Paranal, R.M.; Burge, C.B.; Bradner, J.E.; Lee, T.I.; Young, R.A. Transcriptional amplification in tumor cells with elevated c-Myc. Cell 2012, 151, 56–67. [Google Scholar] [CrossRef] [PubMed]
- Nie, Z.; Hu, G.; Wei, G.; Cui, K.; Yamane, A.; Resch, W.; Wang, R.; Green, D.R.; Tessarollo, L.; Casellas, R.; et al. c-Myc is a universal amplifier of expressed genes in lymphocytes and embryonic stem cells. Cell 2012, 151, 68–79. [Google Scholar] [CrossRef] [PubMed]
- Yu, J.; Vodyanik, M.A.; Smuga-Otto, K.; Antosiewicz-Bourget, J.; Frane, J.L.; Tian, S.; Nie, J.; Jonsdottir, G.A.; Ruotti, V.; Stewart, R.; et al. Induced pluripotent stem cell lines derived from human somatic cells. Science 2007, 318, 1917–1920. [Google Scholar] [CrossRef] [PubMed]
- Sablin, E.P.; Blind, R.D.; Uthayaruban, R.; Chiu, H.J.; Deacon, A.M.; Das, D.; Ingraham, H.A.; Fletterick, R.J. Structure of Liver Receptor Homolog-1 (NR5A2) with PIP3 hormone bound in the ligand binding pocket. J. Struct. Biol. 2015, 192, 342–348. [Google Scholar] [CrossRef] [PubMed]
- Heng, J.C.; Feng, B.; Han, J.; Jiang, J.; Kraus, P.; Ng, J.H.; Orlov, Y.L.; Huss, M.; Yang, L.; Lufkin, T.; et al. The nuclear receptor Nr5a2 can replace Oct4 in the reprogramming of murine somatic cells to pluripotent cells. Cell Stem Cell 2010, 6, 167–174. [Google Scholar] [CrossRef] [PubMed]
- Shu, J.; Wu, C.; Wu, Y.; Li, Z.; Shao, S.; Zhao, W.; Tang, X.; Yang, H.; Shen, L.; Zuo, X.; et al. Induction of pluripotency in mouse somatic cells with lineage specifiers. Cell 2013, 153, 963–975. [Google Scholar] [CrossRef] [PubMed]
- Nuria, M.; Emmanuel, N.; Ignacio, S.M.; Tomoaki, H.; Sachin, K.; Laia, M.; Carme, C.; Yuriko, H.; Yun, X. Tency with lineage specifiers. Cell Stem Cell 2013, 13, 341–350. [Google Scholar]
- Feng, B.; Jiang, J.; Kraus, P.; Ng, J.H.; Heng, J.C.; Chan, Y.S.; Yaw, L.P.; Zhang, W.; Loh, Y.H.; Han, J.; et al. Reprogramming of fibroblasts into induced pluripotent stem cells with orphan nuclear receptor Esrrb. Nat. Cell Biol. 2009, 11, 197–203. [Google Scholar] [CrossRef] [PubMed]
- Maekawa, M.; Yamaguchi, K.; Nakamura, T.; Shibukawa, R.; Kodanaka, I.; Ichisaka, T.; Kawamura, Y.; Mochizuki, H.; Goshima, N.; Yamanaka, S. Direct reprogramming of somatic cells is promoted by maternal transcription factor Glis1. Nature 2011, 474, 225–229. [Google Scholar] [CrossRef] [PubMed]
- Cheval, L.; Pierrat, F.; Rajerison, R.; Piquemal, D.; Doucet, A. Of mice and men: Divergence of gene expression patterns in kidney. PLoS ONE 2012, 7, e46876. [Google Scholar] [CrossRef] [PubMed]
- Balestrieri, C.; Vanoni, M.; Hautaniemi, S.; Alberghina, L.; Chiaradonna, F. Integrative transcriptional analysis between human and mouse cancer cells provides a common set of transformation associated genes. Biotechnol. Adv. 2012, 30, 16–29. [Google Scholar] [CrossRef] [PubMed]
- Gul-Uludag, H.; Xu, P.; Marquez-Curtis, L.A.; Xing, J.; Janowska-Wieczorek, A.; Chen, J. Cationic liposome-mediated CXCR4 gene delivery into hematopoietic stem/progenitor cells: Implications for clinical transplantation and gene therapy. Stem Cells Dev. 2012, 21, 1587–1596. [Google Scholar] [CrossRef] [PubMed]
- McLenachan, S.; Sarsero, J.P.; Ioannou, P.A. Flow-cytometric analysis of mouse embryonic stem cell lipofection using small and large DNA constructs. Genomics 2007, 89, 708–720. [Google Scholar] [CrossRef] [PubMed]
- Ellis, J. Silencing and variegation of gammaretrovirus and lentivirus vectors. Hum. Gene Ther. 2005, 16, 1241–1246. [Google Scholar] [CrossRef] [PubMed]
- Follenzi, A.; Naldini, L. Generation of HIV-1 derived lentiviral vectors. Methods Enzymol. 2002, 346, 454–465. [Google Scholar] [PubMed]
- Lotti, F.; Mavilio, F. The choice of a suitable lentivirus vector: Transcriptional targeting. Methods Mol. Biol. 2003, 229, 17–27. [Google Scholar] [PubMed]
- Stadtfeld, M.; Nagaya, M.; Utikal, J.; Weir, G.; Hochedlinger, K. Induced pluripotent stem cells generated without viral integration. Science 2008, 322, 945–949. [Google Scholar] [CrossRef] [PubMed]
- Zhou, W.; Freed, C.R. Adenoviral gene delivery can reprogram human fibroblasts to induced pluripotent stem cells. Stem Cells 2009, 27, 2667–2674. [Google Scholar] [CrossRef] [PubMed]
- Thomas, C.E.; Ehrhardt, A.; Kay, M.A. Progress and problems with the use of viral vectors for gene therapy. Nat. Rev. Genet. 2003, 4, 346–358. [Google Scholar] [CrossRef] [PubMed]
- Fusaki, N.; Ban, H.; Nishiyama, A.; Saeki, K.; Hasegawa, M. Efficient induction of transgene-free human pluripotent stem cells using a vector based on Sendai virus, an RNA virus that does not integrate into the host genome. Proc. Jpn. Acad. Ser. B Phys. Biol. Sci. 2009, 85, 348–362. [Google Scholar] [CrossRef] [PubMed]
- Ruggieri, M.; Riboldi, G.; Brajkovic, S.; Bucchia, M.; Bresolin, N.; Comi, G.P.; Corti, S. Induced neural stem cells: Methods of reprogramming and potential therapeutic applications. Prog. Neurobiol. 2014, 114, 15–24. [Google Scholar] [CrossRef] [PubMed]
- Yu, J.; Hu, K.; Smuga-Otto, K.; Tian, S.; Stewart, R.; Slukvin, I.I.; Thomson, J.A. Human induced pluripotent stem cells free of vector and transgene sequences. Science 2009, 324, 797–801. [Google Scholar] [CrossRef] [PubMed]
- McClintock, B. The origin and behavior of mutable loci in maize. Proc. Natl. Acad. Sci. USA 1950, 36, 344–355. [Google Scholar] [CrossRef] [PubMed]
- Jia, F.; Wilson, K.D.; Sun, N.; Gupta, D.M.; Huang, M.; Li, Z.; Panetta, N.J.; Chen, Z.Y.; Robbins, R.C.; Kay, M.A.; et al. A nonviral minicircle vector for deriving human iPS cells. Nat. Methods 2010, 7, 197–199. [Google Scholar] [CrossRef] [PubMed]
- Narsinh, K.H.; Jia, F.; Robbins, R.C.; Kay, M.A.; Longaker, M.T.; Wu, J.C. Generation of adult human induced pluripotent stem cells using nonviral minicircle DNA vectors. Nat. Protoc. 2011, 6, 78–88. [Google Scholar] [CrossRef] [PubMed]
- Schlaeger, T.M.; Daheron, L.; Brickler, T.R.; Entwisle, S.; Chan, K.; Cianci, A.; DeVine, A.; Ettenger, A.; Fitzgerald, K.; Godfrey, M.; et al. A comparison of non-integrating reprogramming methods. Nat. Biotechnol. 2015, 33, 58–63. [Google Scholar] [CrossRef] [PubMed]
- Ogawa, H.; Wu, X.; Kawamoto, K.; Nishida, N.; Konno, M.; Koseki, J.; Matsui, H.; Noguchi, K.; Gotoh, N.; Yamamoto, T.; et al. MicroRNAs Induce Epigenetic Reprogramming and Suppress Malignant Phenotypes of Human Colon Cancer Cells. PLoS ONE 2015, 10, e0127119. [Google Scholar] [CrossRef] [PubMed]
- Weide, B.; Garbe, C.; Rammensee, H.G.; Pascolo, S. Plasmid DNA- and messenger RNA-based anti-cancer vaccination. Immunol. Lett. 2008, 115, 33–42. [Google Scholar] [CrossRef] [PubMed]
- Zhou, H.; Wu, S.; Joo, J.Y.; Zhu, S.; Han, D.W.; Lin, T.; Trauger, S.; Bien, G.; Yao, S.; Zhu, Y.; et al. Generation of induced pluripotent stem cells using recombinant proteins. Cell Stem Cell 2009, 4, 381–384. [Google Scholar] [CrossRef] [PubMed]
- Li, G.H.; Li, W.; Mumper, R.J.; Nath, A. Molecular mechanisms in the dramatic enhancement of HIV-1 Tat transduction by cationic liposomes. FASEB J. 2012, 26, 2824–2834. [Google Scholar] [CrossRef] [PubMed]
- Neumann, E.; Schaefer-Ridder, M.; Wang, Y.; Hofschneider, P.H. Gene transfer into mouse lyoma cells by electroporation in high electric fields. EMBO J. 1982, 1, 841–845. [Google Scholar] [PubMed]
- Bhise, N.S.; Wahlin, K.J.; Zack, D.J.; Green, J.J. Evaluating the potential of poly (β-amino ester) nanoparticles for reprogramming human fibroblasts to become induced pluripotent stem cells. Int. J. Nanomed. 2013, 8, 4641–4658. [Google Scholar]
- Høffding, M.K.; Hyttel, P. Ultrastructural visualization of the Mesenchymal-to-Epithelial Transition during reprogramming of human fibroblasts to induced pluripotent stem cells. Stem Cell Res. 2015, 14, 39–53. [Google Scholar] [CrossRef] [PubMed]
- Hotta, A.; Ellis, J. Retroviral vector silencing during iPS cell induction: An epigenetic beacon that signals distinct pluripotent states. J. Cell. Biochem. 2008, 105, 940–948. [Google Scholar] [CrossRef] [PubMed]
- Tong, J.; Sun, D.; Yang, C.; Wang, Y.; Sun, S.; Li, Q.; Bao, J.; Liu, Y. Serum starvation and thymidine double blocking achieved efficient cell cycle synchronization and altered the expression of p27, p53, bcl-2 in canine breast cancer cells. Res. Vet. Sci. 2016, 105, 10–14. [Google Scholar] [CrossRef] [PubMed]
- Tsubouchi, T.; Fisher, A.G. Reprogramming and the pluripotent stem cell cycle. Curr. Top. Dev. Biol. 2013, 104, 223–241. [Google Scholar] [PubMed]
- Chen, M.; Huang, J.; Yang, X.; Liu, B.; Zhang, W.; Huang, L.; Deng, F.; Ma, J.; Bai, Y.; Lu, R.; et al. Serum starvation induced cell cycle synchronization facilitates human somatic cells reprogramming. PLoS ONE 2012, 7, e28203. [Google Scholar] [CrossRef] [PubMed]
- Ruiz, S.; Panopoulos, A.D.; Herrerías, A.; Bissig, K.D.; Lutz, M.; Berggren, W.T.; Verma, I.M.; Izpisua Belmonte, J.C. A high proliferation rate is required for cell reprogramming and maintenance of human embryonic stem cell identity. Curr. Biol. 2011, 21, 45–52. [Google Scholar] [CrossRef] [PubMed]
- Feng, J. Kinetic barriers in transdifferentiation. Cell Cycle 2016, 22, 1–2. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.; Go, Y.; Kang, I.; Han, Y.M.; Kim, J. Oct-4 controls cell-cycle progression of embryonic stem cells. Biochem. J. 2010, 426, 171–181. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Neganova, I.; Przyborski, S.; Yang, C.; Cooke, M.; Atkinson, S.P.; Anyfantis, G.; Fenyk, S.; Keith, W.N.; Hoare, S.F.; et al. A role for NANOG in G1 to S transition in human embryonic stem cells through direct binding of CDK6 and CDC25A. J. Cell Biol. 2009, 184, 67–82. [Google Scholar] [CrossRef] [PubMed]
- Card, D.A.; Hebbar, P.B.; Li, L.; Trotter, K.W.; Komatsu, Y.; Mishina, Y.; Archer, T.K. Oct4/Sox2-regulated miR-302 targets cyclin D1 in human embryonic stem cells. Mol. Cell. Biol. 2008, 28, 6426–6438. [Google Scholar] [CrossRef] [PubMed]
- Lange, C.; Huttner, W.B.; Calegari, F. Cdk4/cyclinD1 overexpression in neural stem cells shortens G1, delays neurogenesis, and promotes the generation and expansion of basal progenitors. Cell Stem Cell 2009, 5, 320–331. [Google Scholar] [CrossRef] [PubMed]
- Weinberg, R.A. The retinoblastoma protein and cell cycle control. Cell 1995, 81, 323–330. [Google Scholar] [CrossRef]
- Malumbres, M.; Barbacid, M. Cell cycle, CDKs and cancer: A changing paradigm. Nat. Rev. Cancer 2009, 9, 153–166. [Google Scholar] [CrossRef] [PubMed]
- Ma, Y.; Kanakousaki, K.; Buttitta, L. How the cell cycle impacts chromatin architecture and influences cell fate. Front. Genet. 2015, 6, 19. [Google Scholar] [CrossRef] [PubMed]
- Chung, H.C.; Lin, R.C.; Logan, G.J.; Alexander, I.E.; Sachdev, P.S.; Sidhu, K.S. Human induced pluripotent stem cells derived under feeder-free conditions display unique cell cycle and DNA replication gene profiles. Stem Cells Dev. 2012, 21, 206–216. [Google Scholar] [CrossRef] [PubMed]
- Petruk, S.; Sedkov, Y.; Johnston, D.M.; Hodgson, J.W.; Black, K.L.; Kovermann, S.K.; Beck, S.; Canaani, E.; Brock, H.W.; Mazo, A. TrxG and PcG proteins but not methylated histones remain associated with DNA through replication. Cell 2012, 150, 922–933. [Google Scholar] [CrossRef] [PubMed]
- Shibahara, K.; Stillman, B. Replication-dependent marking of DNA by PCNA facilitates CAF-1-coupled inheritance of chromatin. Cell 1999, 96, 575–585. [Google Scholar] [CrossRef]
- Rountree, M.R.; Bachman, K.E.; Baylin, S.B. DNMT1 binds HDAC2 and a new co-repressor, DMAP1, to form a complex at replication foci. Nat. Genet. 2000, 25, 269–277. [Google Scholar] [CrossRef] [PubMed]
- Yoshida, Y.; Takahashi, K.; Okita, K.; Ichisaka, T.; Yamanaka, S. Hypoxia enhances the generation of induced pluripotent stem cells. Cell Stem Cell 2009, 5, 237–241. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.P.; Fu, R.H.; Wu, D.C.; Hsu, C.Y.; Chang, C.H.; Lee, W.; Lee, Y.D.; Liu, C.H.; Chien, Y.J.; Lin, S.Z.; et al. Mouse-induced pluripotent stem cells generated under hypoxic conditions in the absence of viral infection and oncogenic factors and used for ischemic stroke therapy. Stem Cells Dev. 2014, 23, 421–433. [Google Scholar] [CrossRef] [PubMed]
- Mikkelsen, T.S.; Hanna, J.; Zhang, X.; Ku, M.; Wernig, M.; Schorderet, P.; Bernstein, B.E.; Jaenisch, R.; Lander, E.S.; Meissner, A. Dissecting direct reprogramming through integrative genomic analysis. Nature 2008, 454, 49–55. [Google Scholar] [CrossRef] [PubMed]
- Xu, X.L.; Yang, J.P.; Fu, L.N.; Ren, R.T.; Yi, F.; Suzuki, K.; Liu, K.; Ding, Z.C.; Qu, J.; Zhang, W.Q.; et al. Direct reprogramming of porcine fibroblasts to neural progenitor cells. Protein Cell 2014, 5, 4–7. [Google Scholar] [CrossRef] [PubMed]
- Meyer, S.; Wörsdörfer, P.; Günther, K.; Thier, M.; Edenhofer, F. Derivation of Adult Human Fibroblasts and their Direct Conversion into Expandable Neural Progenitor Cells. J. Vis. Exp. 2015, e52831. [Google Scholar] [CrossRef] [PubMed]
- Su, G.; Zhao, Y.; Wei, J.; Xiao, Z.; Chen, B.; Han, J.; Chen, L.; Guan, J.; Wang, R.; Dong, Q.; et al. Direct conversion of fibroblasts into neural progenitor-like cells by forced growth into 3D spheres on low attachment surfaces. Biomaterials 2013, 34, 5897–5906. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Li, K.; Wei, W.; Ding, S. Chemical approaches to stem cell biology and therapeutics. Cell Stem Cell 2013, 13, 270–283. [Google Scholar] [CrossRef] [PubMed]
- Yu, C.; Liu, K.; Tang, S.; Ding, S. Chemical approaches to cell reprogramming. Curr. Opin. Genet. Dev. 2014, 28, 50–56. [Google Scholar] [CrossRef] [PubMed]
- Biswas, D.; Jiang, P. Chemically induced reprogramming of somatic cells to pluripotent stem cells and neural cells. Int. J. Mol. Sci. 2016, 17. [Google Scholar] [CrossRef] [PubMed]
- Soufi, A. Mechanisms for enhancing cellular reprogramming. Curr. Opin. Genet. Dev. 2014, 25, 101–109. [Google Scholar] [CrossRef] [PubMed]
- Huangfu, D.; Maehr, R.; Guo, W.; Eijkelenboom, A.; Snitow, M.; Chen, A.E.; Melton, D.A. Induction of pluripotent stem cells by defined factors is greatly improved by small-molecule compounds. Nat. Biotechnol. 2008, 26, 795–797. [Google Scholar] [CrossRef] [PubMed]
- Mali, P.; Chou, B.K.; Yen, J.; Ye, Z.; Zou, J.; Dowey, S.; Brodsky, R.A.; Ohm, J.E.; Yu, W.; Baylin, S.B.; et al. Butyrate greatly enhances derivation of human induced pluripotent stem cells by promoting epigenetic remodeling and the expression of pluripotency-associated genes. Stem Cells 2010, 28, 713–720. [Google Scholar] [CrossRef] [PubMed]
- He, Y.F.; Li, B.Z.; Li, Z.; Liu, P.; Wang, Y.; Tang, Q.; Ding, J.; Jia, Y.; Chen, Z.; Li, L.; et al. Tet-mediated formation of 5-carboxylcytosine and its excision by TDG in mammalian DNA. Science 2011, 333, 1303–1307. [Google Scholar] [CrossRef] [PubMed]
- Tahiliani, M.; Koh, K.P.; Shen, Y.; Pastor, W.A.; Bandukwala, H.; Brudno, Y.; Agarwal, S.; Iyer, L.M.; Liu, D.R.; Aravind, L.; et al. Conversion of 5-methylcytosine to 5-hydroxymethylcytosine in mammalian DNA by MLL partner TET1. Science 2009, 324, 930–935. [Google Scholar] [CrossRef] [PubMed]
- Gao, Y.; Chen, J.; Li, K.; Wu, T.; Huang, B.; Liu, W.; Kou, X.; Zhang, Y.; Huang, H.; Jiang, Y.; et al. Replacement of Oct4 by Tet1 during iPSC induction reveals an important role of DNA methylation and hydroxymethylation in reprogramming. Cell Stem Cell 2013, 12, 453–469. [Google Scholar] [CrossRef] [PubMed]
- Blaschke, K.; Ebata, K.T.; Karimi, M.M.; Zepeda-Martínez, J.A.; Goyal, P.; Mahapatra, S.; Tam, A.; Laird, D.J.; Hirst, M.; Rao, A.; et al. Vitamin C induces Tet-dependent DNA demethylation and a blastocyst-like state in ES cells. Nature 2013, 500, 222–226. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Zhang, Q.; Yin, X.; Yang, W.; Du, Y.; Hou, P.; Ge, J.; Liu, C.; Zhang, W.; Zhang, X.; et al. Generation of iPSCs from mouse fibroblasts with a single gene, Oct4, and small molecules. Cell Res. 2011, 21, 196–204. [Google Scholar] [CrossRef] [PubMed]
- Miranda, T.B.; Cortez, C.C.; Yoo, C.B.; Liang, G.; Abe, M.; Kelly, T.K.; Marquez, V.E.; Jones, P.A. DZNep is a global histone methylation inhibitor that reactivates developmental genes not silenced by DNA methylation. Mol. Cancer Ther. 2009, 8, 1579–1588. [Google Scholar] [CrossRef] [PubMed]
- Tam, E.K.; Li, Z.; Goh, Y.L.; Cheng, X.; Wong, S.Y.; Santhanakrishnan, S.; Chai, C.L.; Yao, S.Q. Cell-based proteome profiling using an affinity-based probe (AfBP) derived from 3-deazaneplanocin A (DzNep). Chem. Asian J. 2013, 8, 1818–1828. [Google Scholar] [CrossRef] [PubMed]
- Folmes, C.D.; Nelson, T.J.; Martinez-Fernandez, A.; Arrell, D.K.; Lindor, J.Z.; Dzeja, P.P.; Ikeda, Y.; Perez-Terzic, C.; Terzic, A. Somatic oxidative bioenergetics transitions into pluripotency- dependent glycolysis to facilitate nuclear reprogramming. Cell Metab. 2011, 14, 264–271. [Google Scholar] [CrossRef] [PubMed]
- Zhu, S.; Ambasudhan, R.; Sun, W.; Kim, H.J.; Talantova, M.; Wang, X.; Zhang, M.; Zhang, Y.; Laurent, T.; Parker, J.; et al. Small molecules enable Oct4-mediated direct reprogramming into expandable human neural stem cells. Cell Res. 2014, 24, 126–129. [Google Scholar] [CrossRef] [PubMed]
- Han, Y.C.; Lim, Y.; Duffieldl, M.D.; Li, H.; Liu, J.; Abdul Manaph, N.P.; Yang, M.; Keating, D.J.; Zhou, X.F. Direct reprogramming of mouse fibroblasts to neural stem cells by small molecules. Stem Cells Int. 2016, 2016, 4304916. [Google Scholar] [CrossRef] [PubMed]


| Items | Method | Donor Cells | Duration | Characteristics of iPSCs/iNPCs |
|---|---|---|---|---|
| iPSC Studies | ||||
| Takahashi, et al. [1] | Retrovirus | Mouse embryonic (MEF) and adult fibroblast | 16 days | Could differentiate into all three germ layers in vitro |
| Takahashi, et al. [2] | Retrovirus | Adult human fibroblasts | 30 days | Could differentiate into cell types of the three germ layers in vitro |
| Hockemeyer, et al. [14] | Lentivirus + doxycycline | Primary and secondary human fibroblasts | 20–25 days | Primary and secondary human iPSCs |
| Huangfu, et al. [15] | Retrovirus +Valproic acid VPA | Human fibroblasts | 30 days | Resemble human ESCs in pluripotency and global gene expression profiles |
| Shi, et al. [16] | Retrovirus+BIX-01294, BayK8644 | MEF | 14–21 days | Phenotypically and functionally similar to the classic mESCs |
| Lyssiotis, et al. [17] | Retrovirus+ kenpaullone | MEF | 20 days | Generate germline-competent chimeras |
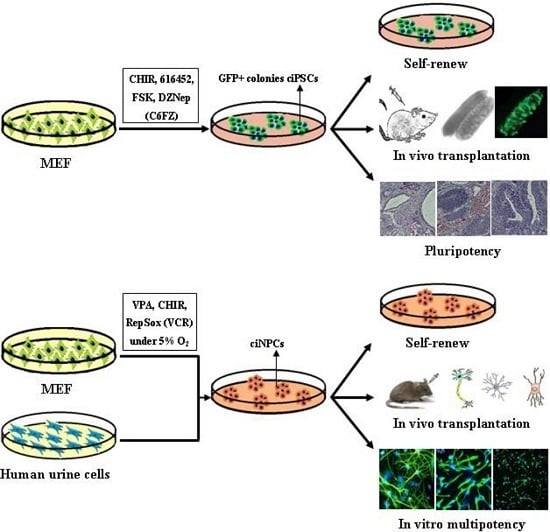
| Hou, et al. [12] | CHIR, 616452, FSK and DZNep (C6FZ) | MEF and adult fibroblasts | 40 days | Differentiate into tissues of all three germ layers |
| iNPC Studies | ||||
| Kim, et al. [12] | doxycycline | Doxycycline-inducible secondary MEF | 7 days | Lose capacity to self-renew after 3–5 passages in vitro and can not differentiate into oligodendrocytes |
| Their, et al. [18] | Retrovirus and lentivirus | MEF | 18 days | Differentiate into neurons, astrocytes, and oligodendrocytes. |
| Lujan, et al. [19] | doxycycline-inducible lentiviral + tetO promoter | MEF | 24 days | Tripotent in vitro, but without evidence of deriving neurons and astrocytes in vivo |
| Han, et al. [20] | Retrovirus | MEF | 4–5 weeks | Exhibit functionality similar to those of wild-type NPCs in vitro and in vivo |
| Ring, et al. [21] | Retrovirus | MEF and human fetal fibroblasts | 41 days | Differentiate into neurons, astrocytes, and oligodendrocytes |
| Mitchell, et al. [22] | Lentivirus | adult human fibroblasts | 14 days | Gives rise to all three major subtypes of neural cells with functional capacity |
| Lee, et al. [23] | Lentivirus + SB431542, Noggin, DN-193189, CHIR99021 | Human cord blood or adult peripheral blood cells | 10–14 days | Produce astrocytes and oligodendrocytes and multiple neuronal subtypes |
| Wang, et al. [24] | episomal vectors + microRNA + CHIR99021, PD0325901, A83-01, thiazovivin and DMH1 | human urine cells | 15 days | differentiated into neurons and glial cells in vitro |
| Cheng, et al. [13] | VPA, CHIR99021 and Repsox | MEFs and human urinary cells | Mouse 10 days; Human, 20 days | Mouse tripotent iNPCs; Human iNPC could differentiate into neurons and astrocytes |
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tian, Z.; Guo, F.; Biswas, S.; Deng, W. Rationale and Methodology of Reprogramming for Generation of Induced Pluripotent Stem Cells and Induced Neural Progenitor Cells. Int. J. Mol. Sci. 2016, 17, 594. https://doi.org/10.3390/ijms17040594
Tian Z, Guo F, Biswas S, Deng W. Rationale and Methodology of Reprogramming for Generation of Induced Pluripotent Stem Cells and Induced Neural Progenitor Cells. International Journal of Molecular Sciences. 2016; 17(4):594. https://doi.org/10.3390/ijms17040594
Chicago/Turabian StyleTian, Zuojun, Fuzheng Guo, Sangita Biswas, and Wenbin Deng. 2016. "Rationale and Methodology of Reprogramming for Generation of Induced Pluripotent Stem Cells and Induced Neural Progenitor Cells" International Journal of Molecular Sciences 17, no. 4: 594. https://doi.org/10.3390/ijms17040594
APA StyleTian, Z., Guo, F., Biswas, S., & Deng, W. (2016). Rationale and Methodology of Reprogramming for Generation of Induced Pluripotent Stem Cells and Induced Neural Progenitor Cells. International Journal of Molecular Sciences, 17(4), 594. https://doi.org/10.3390/ijms17040594