Interleukin-17A Differentially Induces Inflammatory and Metabolic Gene Expression in the Adipose Tissues of Lean and Obese Mice
Abstract
:1. Introduction
2. Results
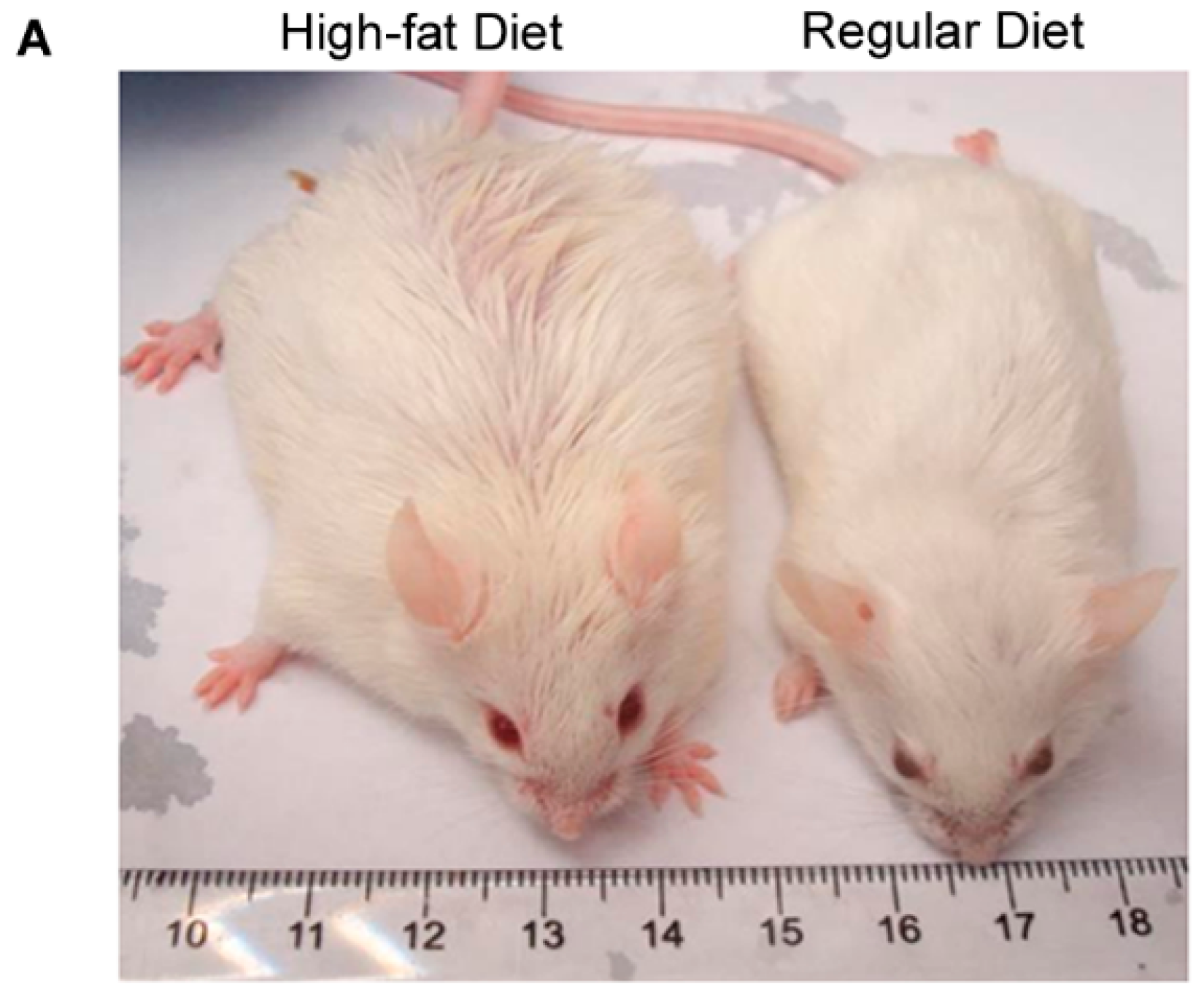
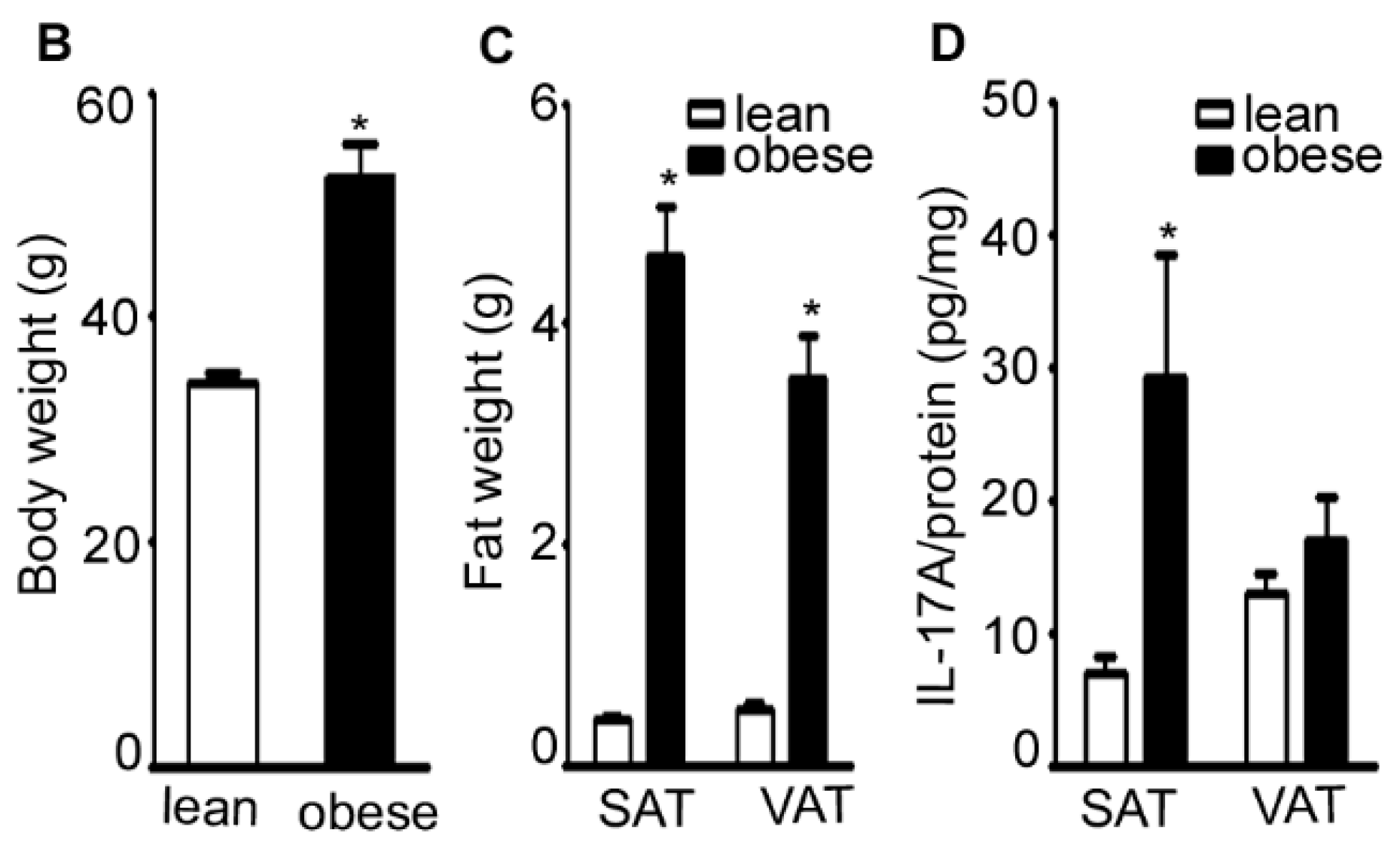
2.1. IL-17A Levels Are Increased in Subcutaneous Adipose Tissue (SAT) of Obese Mice
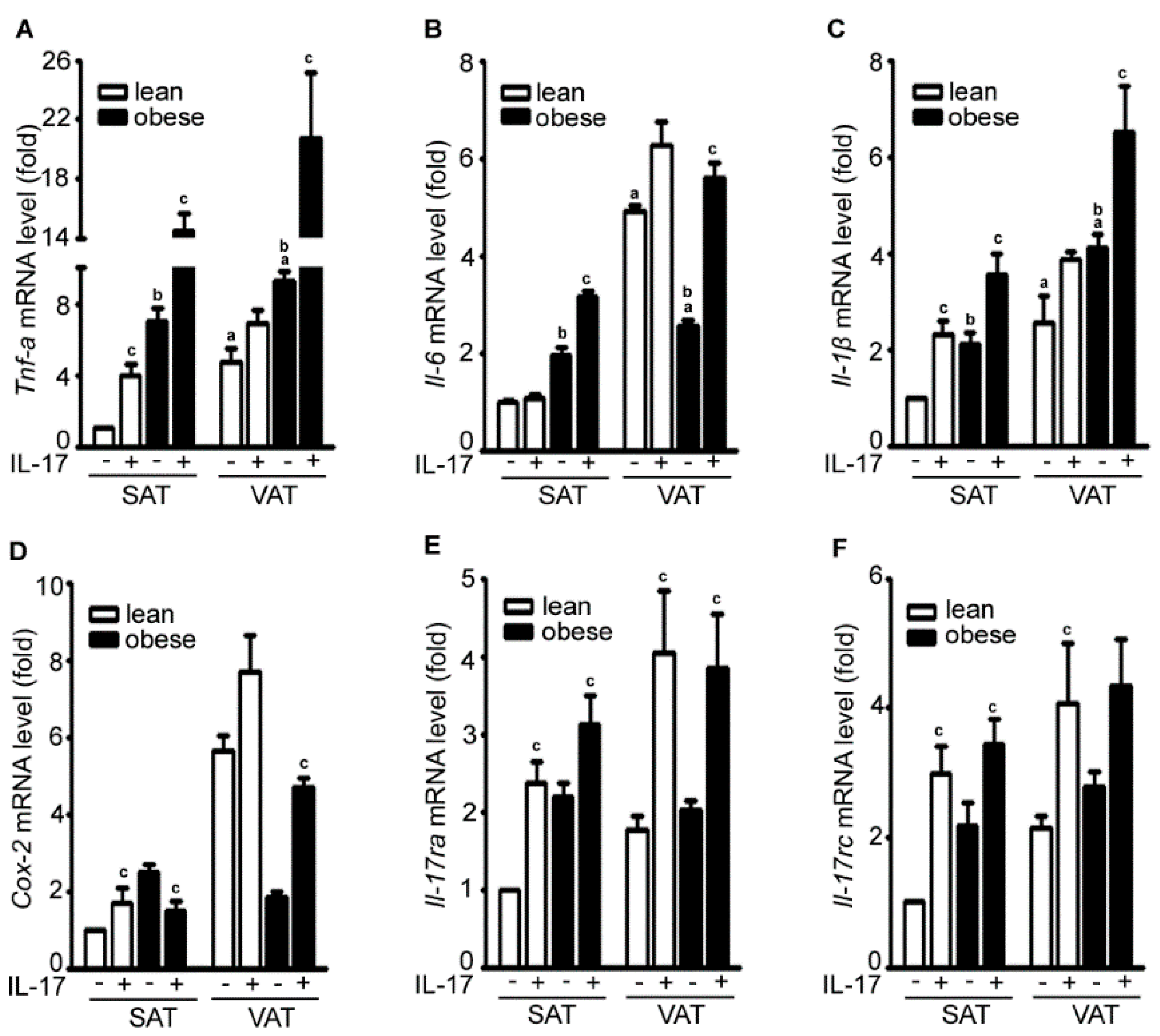
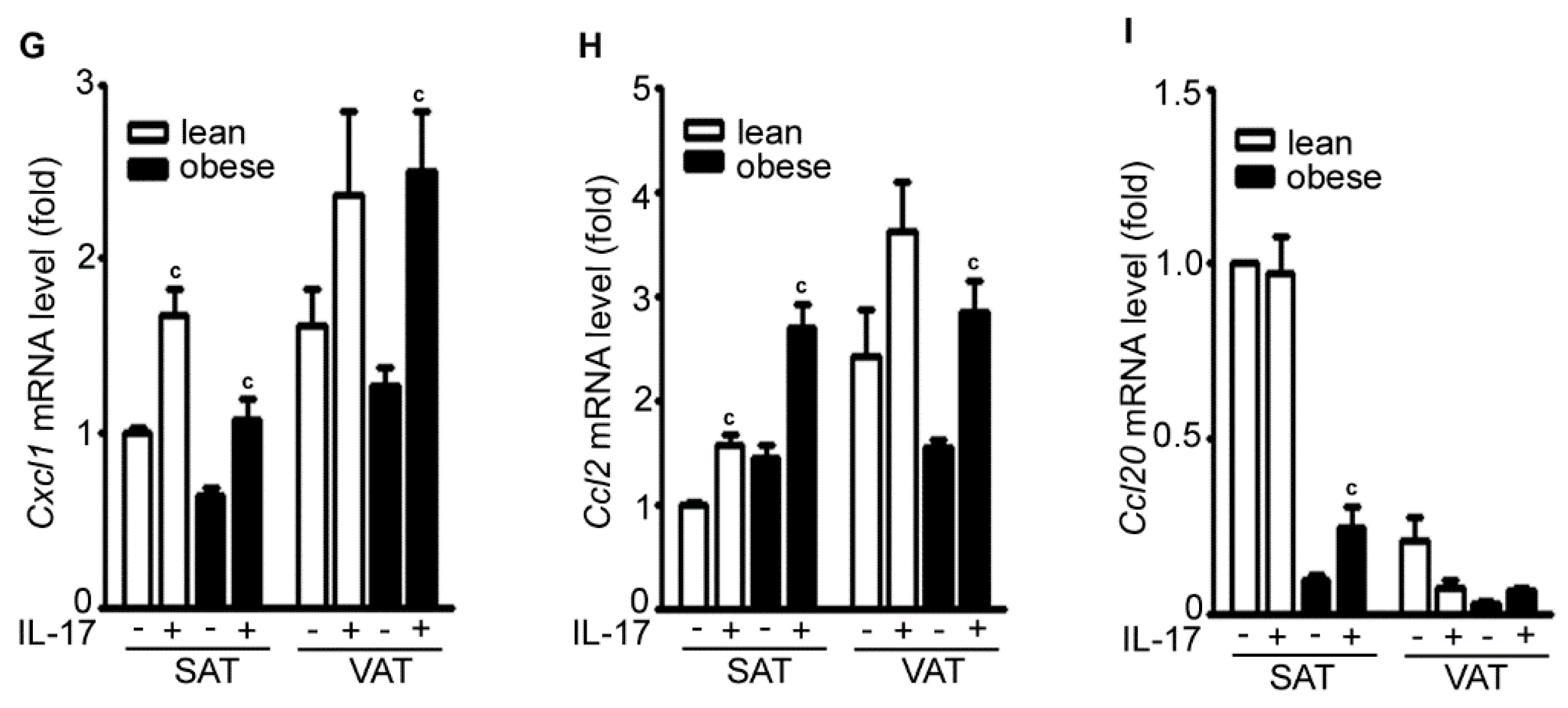
2.2. IL-17A Differentially Induces Inflammatory Gene Expression in the Adipose Tissues of Lean and Obese Mice
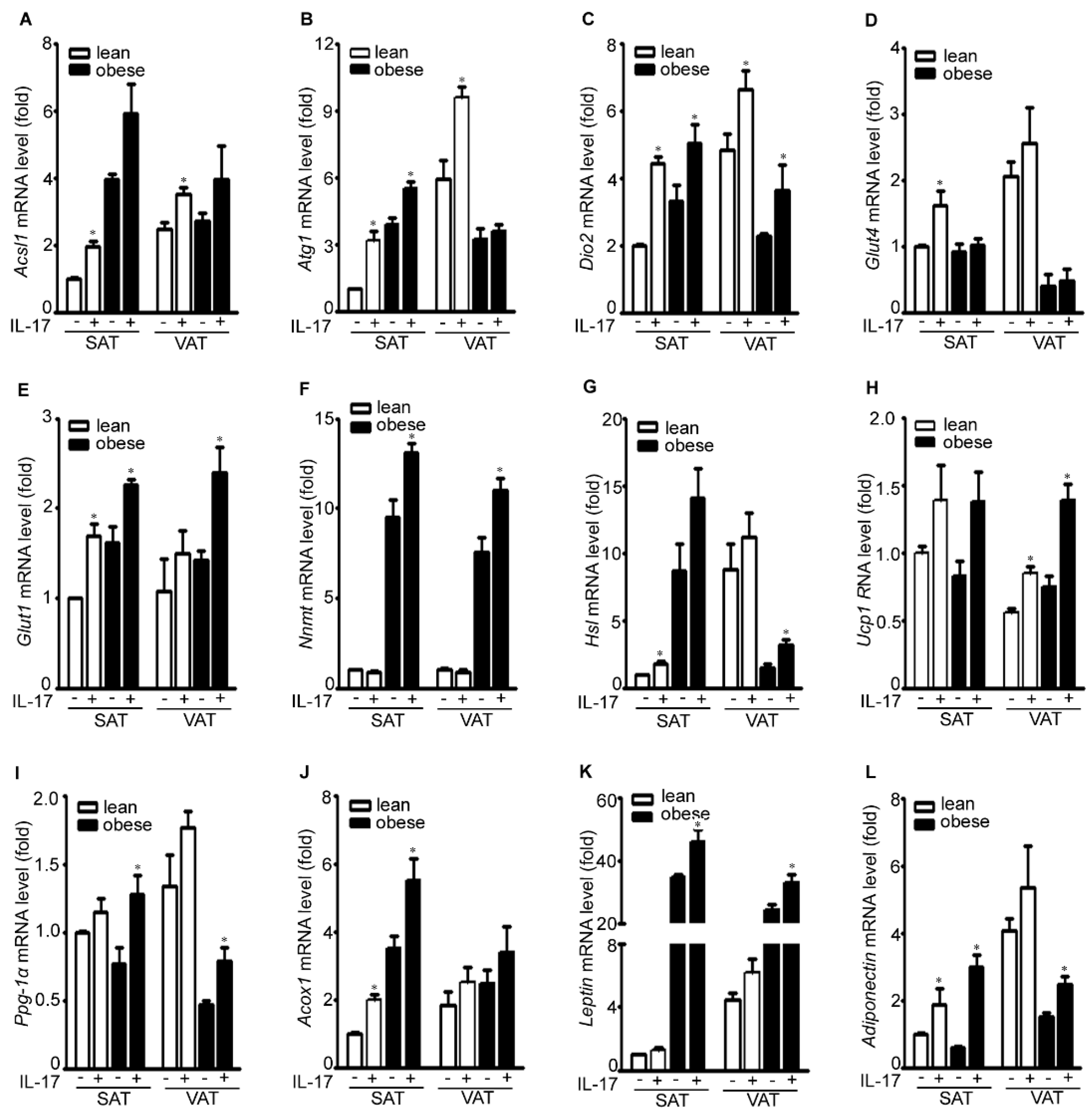
2.3. IL-17A Differentially Induces Metabolic Gene Expression in the Adipose Tissues of Lean and Obese Mice
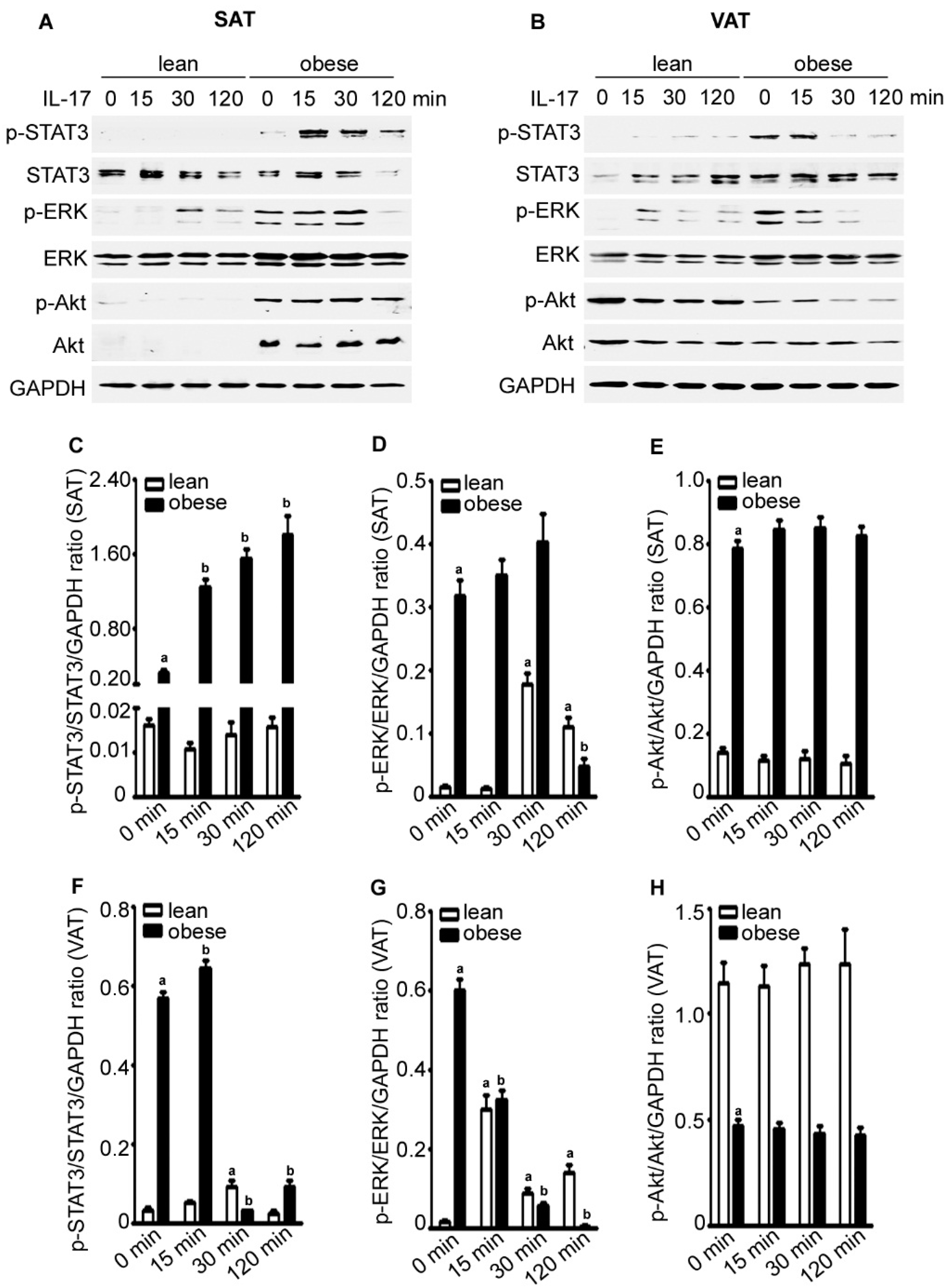
2.4. IL-17A Differentially Activates Signaling Pathways in the Adipose Tissues of Lean and Obese Mice
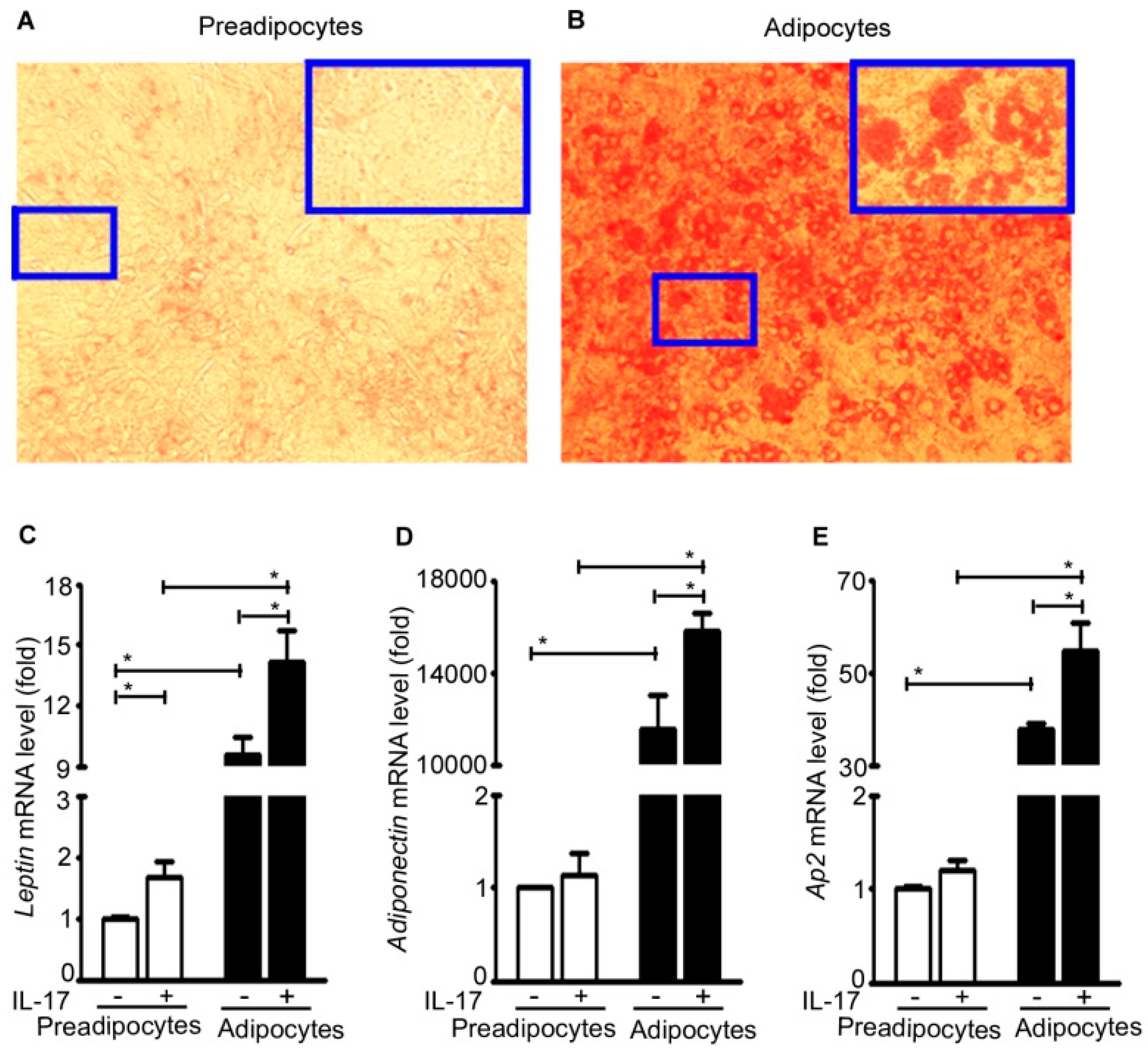
2.5. IL-17A Induces Adipokine Expression in Adipocytes
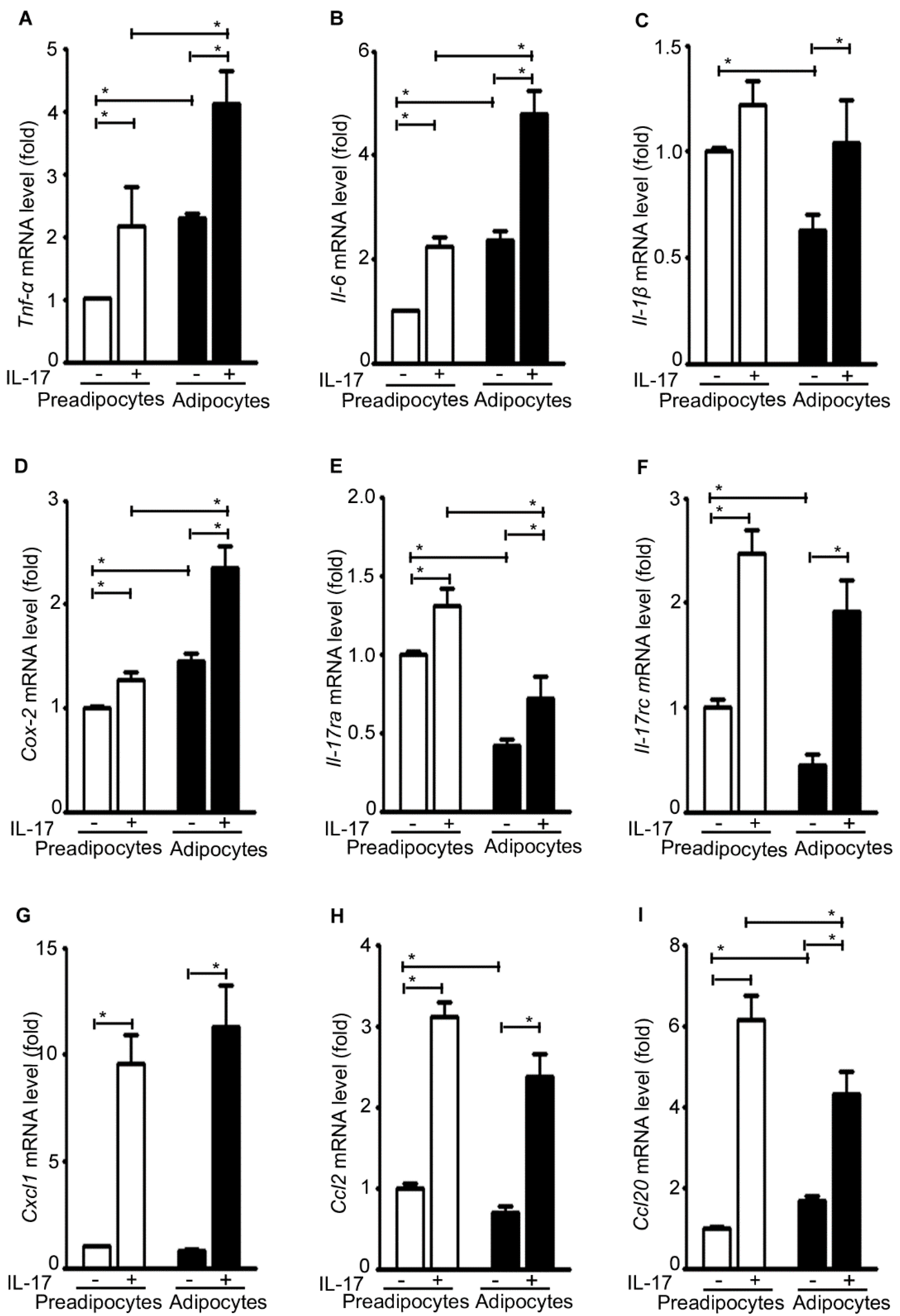
2.6. IL-17A Differentially Induces Inflammatory Gene Expression in Pre-Adipocytes and Adipocytes
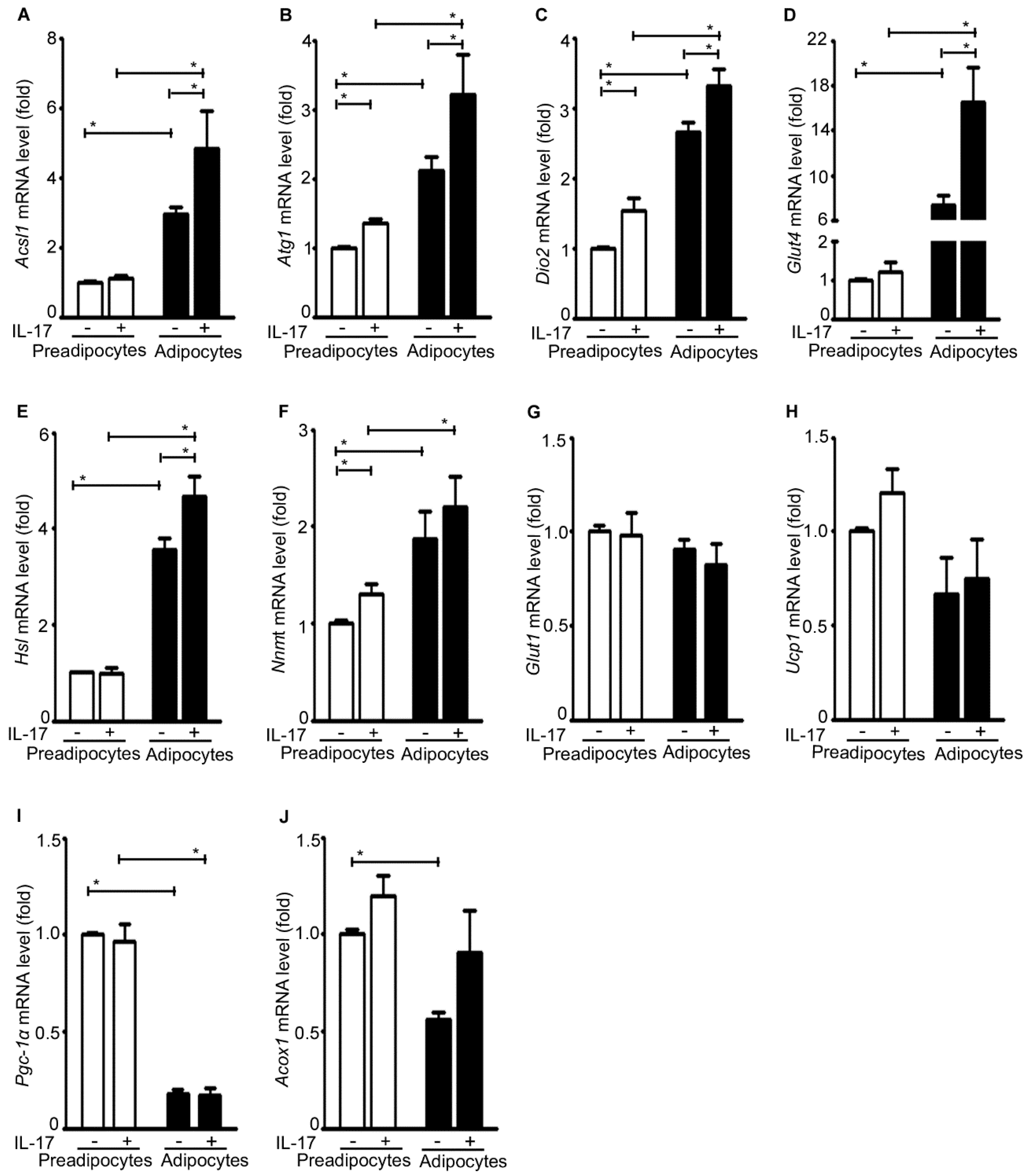
2.7. IL-17A Differentially Induces Metabolic Gene Expression in Pre-Adipocytes and Adipocytes
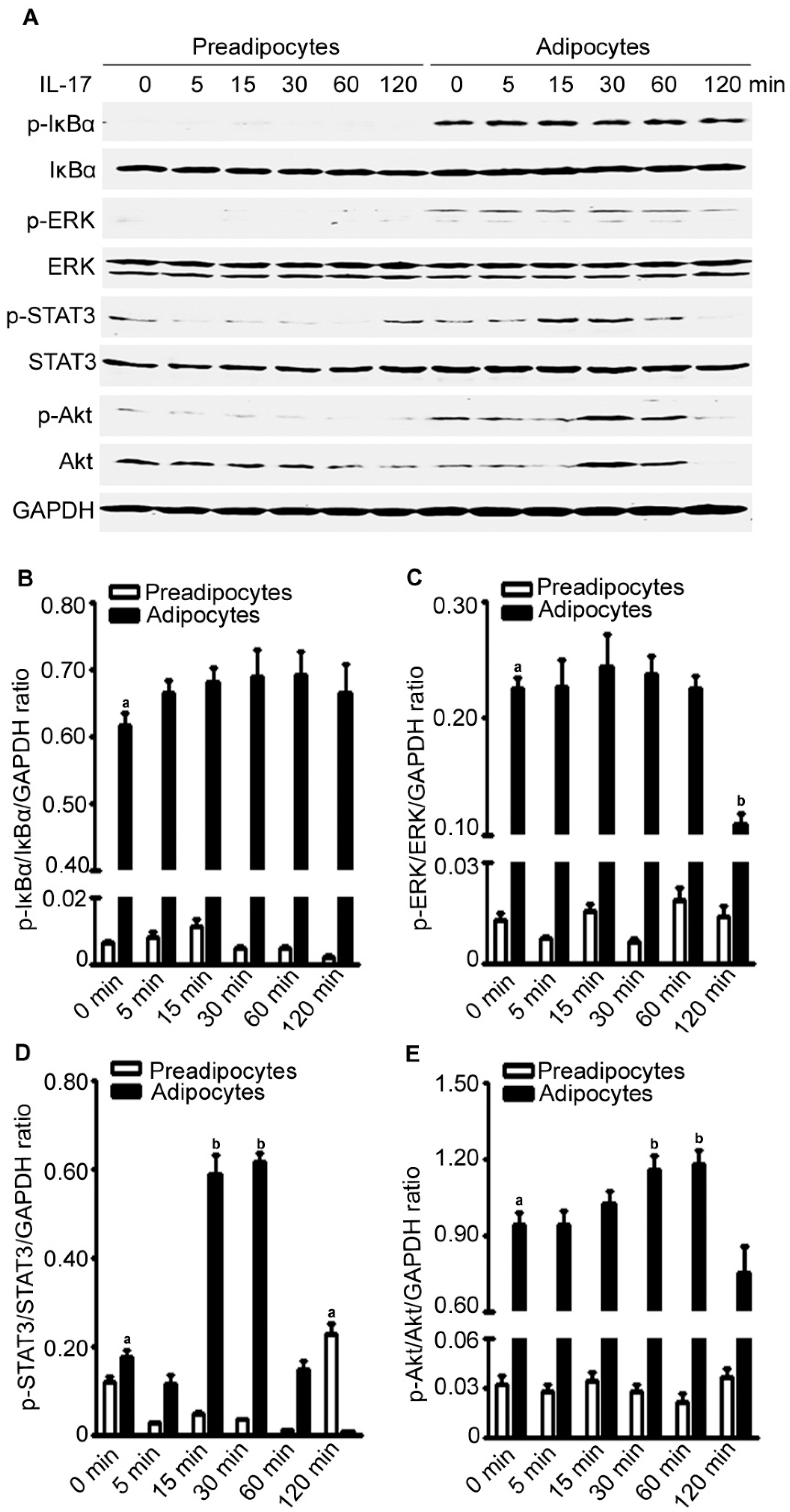
2.8. IL-17A Differentially Activates Signaling Pathways in Preadipocytes and Adipocytes
3. Discussion
4. Materials and Methods
4.1. Animals and Adipose Tissue Culture
4.2. Cell Culture and Adipocyte Differentiation
4.3. Quantitative Real-Time Reverse Transcriptase Polymerase Chain Reaction (qRT-PCR)
4.4. Western Blot Analysis
4.5. ELISA Assay
4.6. Statistical Analysis
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- National Center for Health Statistics. Health, United States, 2012: With Special Feature on Emergency Care; National Center for Health Statistics: Hyattsville, MD, USA, 2013; pp. 12–13.
- Schmidt, A.M. The growing problem of obesity: Mechanisms, consequences, and therapeutic approaches. Arterioscler. Thromb. Vasc. Biol. 2015, 35, e19–e23. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Stewart, B.W.; Wild, C.P. World Cancer Report 2014. Available online: http://www.thehealthwell.info/ (accessed on 11 February 2016).
- Calle, E.E.; Rodriguez, C.; Walker-Thurmond, K.; Thun, M.J. Overweight, obesity, and mortality from cancer in a prospectively studied cohort of U.S. adults. N. Engl. J. Med. 2003, 348, 1625–1638. [Google Scholar] [CrossRef] [PubMed]
- Cohen, D.H.; LeRoith, D. Obesity, type 2 diabetes, and cancer: The insulin and IGF connection. Endocr. Relat. Cancer 2012, 19, F27–F45. [Google Scholar] [CrossRef] [PubMed]
- Vandanmagsar, B.; Youm, Y.H.; Ravussin, A.; Galgani, J.E.; Stadler, K.; Mynatt, R.L.; Ravussin, E.; Stephens, J.M.; Dixit, V.D. The NLRP3 inflammasome instigates obesity-induced inflammation and insulin resistance. Nat. Med. 2011, 17, 179–188. [Google Scholar] [CrossRef] [PubMed]
- Xu, H. Obesity and metabolic inflammation. Drug Discov. Today Dis. Mech. 2013, 10, e21–e25. [Google Scholar] [CrossRef] [PubMed]
- Catalan, V.; Gomez-Ambrosi, J.; Rodriguez, A.; Fruhbeck, G. Adipose tissue immunity and cancer. Front. Physiol. 2013, 4. [Google Scholar] [CrossRef] [PubMed]
- Winer, S.; Paltser, G.; Chan, Y.; Tsui, H.; Engleman, E.; Winer, D.; Dosch, H.M. Obesity predisposes to Th17 bias. Eur. J. Immunol. 2009, 39, 2629–2635. [Google Scholar] [CrossRef] [PubMed]
- Pini, M.; Fantuzzi, G. Enhanced production of IL-17A during zymosan-induced peritonitis in obese mice. J. Leukoc. Biol. 2010, 87, 51–58. [Google Scholar] [CrossRef] [PubMed]
- Sumarac-Dumanovic, M.; Stevanovic, D.; Ljubic, A.; Jorga, J.; Simic, M.; Stamenkovic-Pejkovic, D.; Starcevic, V.; Trajkovic, V.; Micic, D. Increased activity of interleukin-23/interleukin-17 proinflammatory axis in obese women. Int. J. Obes. 2009, 33, 151–156. [Google Scholar] [CrossRef] [PubMed]
- Song, X.; Qian, Y. IL-17 family cytokines mediated signaling in the pathogenesis of inflammatory diseases. Cell Signal. 2013, 25, 2335–2347. [Google Scholar] [CrossRef] [PubMed]
- Onishi, R.M.; Gaffen, S.L. Interleukin-17 and its target genes: Mechanisms of interleukin-17 function in disease. Immunology 2010, 129, 311–321. [Google Scholar] [CrossRef] [PubMed]
- Moseley, T.A.; Haudenschild, D.R.; Rose, L.; Reddi, A.H. Interleukin-17 family and IL-17 receptors. Cytokine Growth Factor Rev. 2003, 14, 155–174. [Google Scholar] [CrossRef]
- Ahmed, M.; Gaffen, S.L. IL-17 inhibits adipogenesis in part via C/EBPα, PPARγ and Kruppel-like factors. Cytokine 2013, 61, 898–905. [Google Scholar] [CrossRef] [PubMed]
- Ge, D.; You, Z. Expression of interleukin-17RC protein in normal human tissues. Int. Arch. Med. 2008, 1. [Google Scholar] [CrossRef] [PubMed]
- Vernia, S.; Cavanagh-Kyros, J.; Barrett, T.; Jung, D.Y.; Kim, J.K.; Davis, R.J. Diet-induced obesity mediated by the JNK/DIO2 signal transduction pathway. Genes Dev. 2013, 27, 2345–2355. [Google Scholar] [CrossRef] [PubMed]
- Kraus, D.; Yang, Q.; Kong, D.; Banks, A.S.; Zhang, L.; Rodgers, J.T.; Pirinen, E.; Pulinilkunnil, T.C.; Gong, F.; Wang, Y.C.; et al. Nicotinamide N-methyltransferase knockdown protects against diet-induced obesity. Nature 2014, 508, 258–262. [Google Scholar] [CrossRef] [PubMed]
- Alvehus, M.; Buren, J.; Sjostrom, M.; Goedecke, J.; Olsson, T. The human visceral fat depot has a unique inflammatory profile. Obesity 2010, 18, 879–883. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Tian, J.; Tian, X.; Tang, X.; Rui, K.; Tong, J.; Lu, L.; Xu, H.; Wang, S. Adipose tissue dendritic cells enhances inflammation by prompting the generation of Th17 cells. PLoS ONE 2014, 9, e92450. [Google Scholar] [CrossRef] [PubMed]
- Shen, F.; Ruddy, M.J.; Plamondon, P.; Gaffen, S.L. Cytokines link osteoblasts and inflammation: Microarray analysis of interleukin-17- and TNF-α-induced genes in bone cells. J. Leukoc. Biol. 2005, 77, 388–399. [Google Scholar] [CrossRef] [PubMed]
- Green, H.; Meuth, M. An established pre-adipose cell line and its differentiation in culture. Cell 1974, 3, 127–133. [Google Scholar] [CrossRef]
- Reed, B.C.; Lane, M.D. Insulin receptor synthesis and turnover in differentiating 3T3-L1 preadipocytes. Proc. Natl. Acad. Sci. USA 1980, 77, 285–289. [Google Scholar] [CrossRef] [PubMed]
- Zhang, M.; Zhang, Y.; Ma, J.; Guo, F.; Cao, Q.; Zhou, B.; Chai, J.; Zhao, W.; Zhao, R. The demethylase activity of FTO (Fat mass and obesity associated protein) is required for preadipocyte differentiation. PLoS ONE 2015, 10, e0133788. [Google Scholar] [CrossRef] [PubMed]
- Kramer, A.H.; Joos-Vandewalle, J.; Edkins, A.L.; Frost, C.L.; Prinsloo, E. Real-time monitoring of 3T3-L1 preadipocyte differentiation using a commercially available electric cell-substrate impedance sensor system. Biochem. Biophys. Res. Commun. 2014, 443, 1245–1250. [Google Scholar] [CrossRef] [PubMed]
| Nutrients | Minerals | Vitamins | |||||
|---|---|---|---|---|---|---|---|
| Protein, % | 21.0 | Fat (ether extract), % | 5.0 | Ash, % | 5.9 | Carotene, ppm | 1.5 |
| Arginine, % | 1.28 | Fat (acid hydrolysis), % | 6.3 | Calcium, % | 0.81 | Vitamin K, ppm | 3.3 |
| Cystine, % | 0.36 | Cholesterol, ppm | 142 | Phosphorus, % | 0.65 | Thiamin hydrochloride, ppm | 17 |
| Glycine, % | 0.96 | Linoleic acid, % | 2.14 | Phosphorus (non-phytate), % | 0.34 | Riboflavin, ppm | 8.0 |
| Histidine, % | 0.52 | Linolenic acid, % | 0.27 | Potassium, % | 1.09 | Niacin, ppm | 85 |
| Isoleucine, % | 0.86 | Arachidonic acid, % | <0.01 | Magnessium, % | 0.23 | Pantothenic acid, ppm | 17 |
| Leucine, % | 1.56 | Omega-3 fatty acids, % | 0.44 | Sulfur, % | 0.33 | Choline chloride, ppm | 2000 |
| Lysine, % | 1.17 | Total saturated fatty acids, % | 0.78 | Sodium, % | 0.30 | Folic acid, ppm | 3.0 |
| Methionine, % | 0.62 | Total monounsaturated fatty acids, % | 0.97 | Chloride, % | 0.52 | Pyridoxine, ppm | 9.6 |
| Phenylalanine, % | 0.91 | Fiber (crude), % | 4.5 | Fluorine, ppm | 9.3 | Biotin, ppm | 0.30 |
| Tyrosine, % | 0.59 | Neutral detergent fiber, % | 15.8 | Iron, ppm | 200 | B12, mcg/kg | 51 |
| Threonine, % | 0.78 | Acid detergent fiber, % | 5.8 | Zinc, ppm | 89 | Vitamin A, IU/gm | 15 |
| Tryptophan, % | 0.24 | Nitrogen-free extract, % | 53.5 | Manganese, ppm | 84 | Vitamin D3, IU/gm | 2.3 |
| Valine, % | 0.96 | Starch, % | 28.6 | Copper, ppm | 14 | Vitamin E, IU/kg | 99 |
| Serine, % | 0.98 | Glucose, % | 0.19 | Cobalt, ppm | 0.71 | - | - |
| Aspartic acid, % | 2.18 | Fructose, % | 0.24 | Iodine, ppm | 0.97 | Calories provided by | |
| Glutamic acid, % | 4.16 | Sucrose, % | 3.24 | Chromium, ppm | 0.01 | Protein, % | 24.496 |
| Alanine, % | 1.19 | Lactose, % | 1.34 | Selenium, ppm | 0.37 | Fat, % | 13.123 |
| Proline, % | 1.31 | Total digestible nutrients, % | 75.1 | - | - | Carbohydrates, % | 62.380 |
| Taurine, % | 0.03 | - | - | - | - | - | - |
| Composition | Gram (gm)% | Kcal % |
|---|---|---|
| Protein | 26.2 | 20 |
| Carbohydrate | 26.3 | 20 |
| Fat | 34.9 | 60 |
| Total | - | 100 |
| kcal/gm | 5.24 | - |
| Ingredient | gm | kcal |
| Casein, 30 mesh | 200 | 800 |
| l-Cysteine | 3 | 12 |
| Corn starch | 0 | 0 |
| Maltodextrin 10 | 125 | 500 |
| Sucrose | 68.8 | 275.2 |
| Cellulose, BW200 | 50 | 0 |
| Soybean oil | 25 | 225 |
| Lard | 245 | 2205 |
| Mineral mix S10026 | 10 | 0 |
| Dicalcium phosphate | 13 | 0 |
| Potassium citrate, 1 H2O | 16.5 | 0 |
| Vitamin mix V10001 | 10 | 40 |
| Choline bitartrate | 2 | 0 |
| Brillant Blue For Coloring Food (FD & C Blue Dye # 1) | 0.05 | 0 |
| Total | 773.85 | 4057 |
| Gene | Forward (5′ to 3′) | Reverse (5′ to 3′) |
|---|---|---|
| Tnf-a | CTACTCCCAGGTTCTCTTCAA | GCAGAGAGGAGGTTGACTTTC |
| Il-6 | CCACTTCACAAGTCGGAGGCTTA | GCAAGTGCATCATCGTTGTTCATAC |
| Il-1β | TCTTCTTTGGGTATTGCTTGG | TGTAATGAAGACGGCACACC |
| Cox-2 | TGGAAAAGGTTCTTCTACGGAG | TGAACCCAGGTCCTCGCT |
| Il-17ra | CGGAGAATTAGTCCCTGTGTTG | GAACAGTCACTTCATACTCCTGG |
| Il-17rc | TTCTGCGGTATTTGACTGTTTCG | GTCCCGGACTTCAAGACCC |
| Cxcl1 | CACCCAAACCGAAGTCATAG | AAGCCAGCGTTCACCAGA |
| Ccl2 | GCCTGCTGTTCACAGTTGC | TGTATGTCTGGACCCATTCCT |
| Ccl20 | GTCCAATTCCATCCCAAAAA | AACTGGGTGAAAAGGGCTGT |
| Acsl1 | GATCTGGTGGAACGAGGCAA | CTTCGGGTTCTGGAGGCTTG |
| Atgl | CAGCACATTTATCCCGGTGTAC | AAATGCCGCCATCCACATAG |
| Dio2 | GTCCGCAAATGACCCCTTT | CCCACCCACTCTCTGACTTTC |
| Glut1 | TCAACACGGCCTTCACTG | CACGATGCTCAGATAGGACATC |
| Glut4 | TTGGCTCCCTTCAGTTTGG | CTACCCAGCCACGTTGCAT |
| Nnmt | AGGAACCAGGAGCCTTTGACT | CCTGAGGGCAGTGCGATAGG |
| Ucp1 | GGCTGGTGGTGGTCGGAGAT | CCGAAGGCAGAAGTGAAGTG |
| Pgc1a | AGACAAATGTGCTTCGAAAAAGAA | GAAGAGATAAAGTTGTTGGTTTGGC |
| Acox1 | GCCCAACTGTGACTTCCATTAA | GTAGCACTCCCCTCGAGTGAT |
| Leptin | CTGCCCCCCAGTTTGATG | GCCAGGCTGCCAGAATTG |
| Adiponectin | AACTTGTGCAGGTTGGATGG | GCCCTTCAGCTCCTGTCATT |
| Ap2 | ACACCGAGATTTCCTTCA AACTG | CCATCTAGGGTTATG TGCTCTTCA |
| Gapdh | TGCACCACCAACTGCTTAG | GGATGCAGGGATGATGTTC |
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons by Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Qu, Y.; Zhang, Q.; Ma, S.; Liu, S.; Chen, Z.; Mo, Z.; You, Z. Interleukin-17A Differentially Induces Inflammatory and Metabolic Gene Expression in the Adipose Tissues of Lean and Obese Mice. Int. J. Mol. Sci. 2016, 17, 522. https://doi.org/10.3390/ijms17040522
Qu Y, Zhang Q, Ma S, Liu S, Chen Z, Mo Z, You Z. Interleukin-17A Differentially Induces Inflammatory and Metabolic Gene Expression in the Adipose Tissues of Lean and Obese Mice. International Journal of Molecular Sciences. 2016; 17(4):522. https://doi.org/10.3390/ijms17040522
Chicago/Turabian StyleQu, Yine, Qiuyang Zhang, Siqi Ma, Sen Liu, Zhiquan Chen, Zhongfu Mo, and Zongbing You. 2016. "Interleukin-17A Differentially Induces Inflammatory and Metabolic Gene Expression in the Adipose Tissues of Lean and Obese Mice" International Journal of Molecular Sciences 17, no. 4: 522. https://doi.org/10.3390/ijms17040522
APA StyleQu, Y., Zhang, Q., Ma, S., Liu, S., Chen, Z., Mo, Z., & You, Z. (2016). Interleukin-17A Differentially Induces Inflammatory and Metabolic Gene Expression in the Adipose Tissues of Lean and Obese Mice. International Journal of Molecular Sciences, 17(4), 522. https://doi.org/10.3390/ijms17040522






