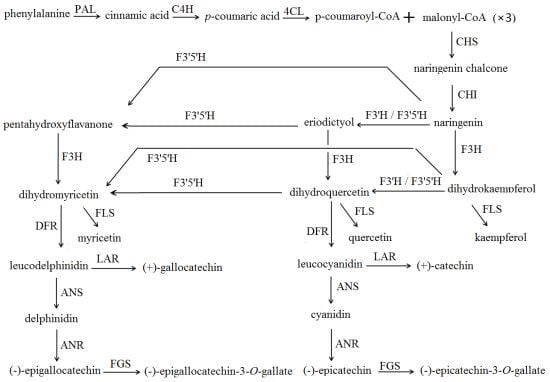
Cloning and Characterization of a Flavonoid 3′-Hydroxylase Gene from Tea Plant (Camellia sinensis)
Abstract
:1. Introduction
2. Results and Discussion
2.1. Cloning and Sequence Analysis of CsF3′H Gene
2.2. Substrate Specificity of CsF3′H
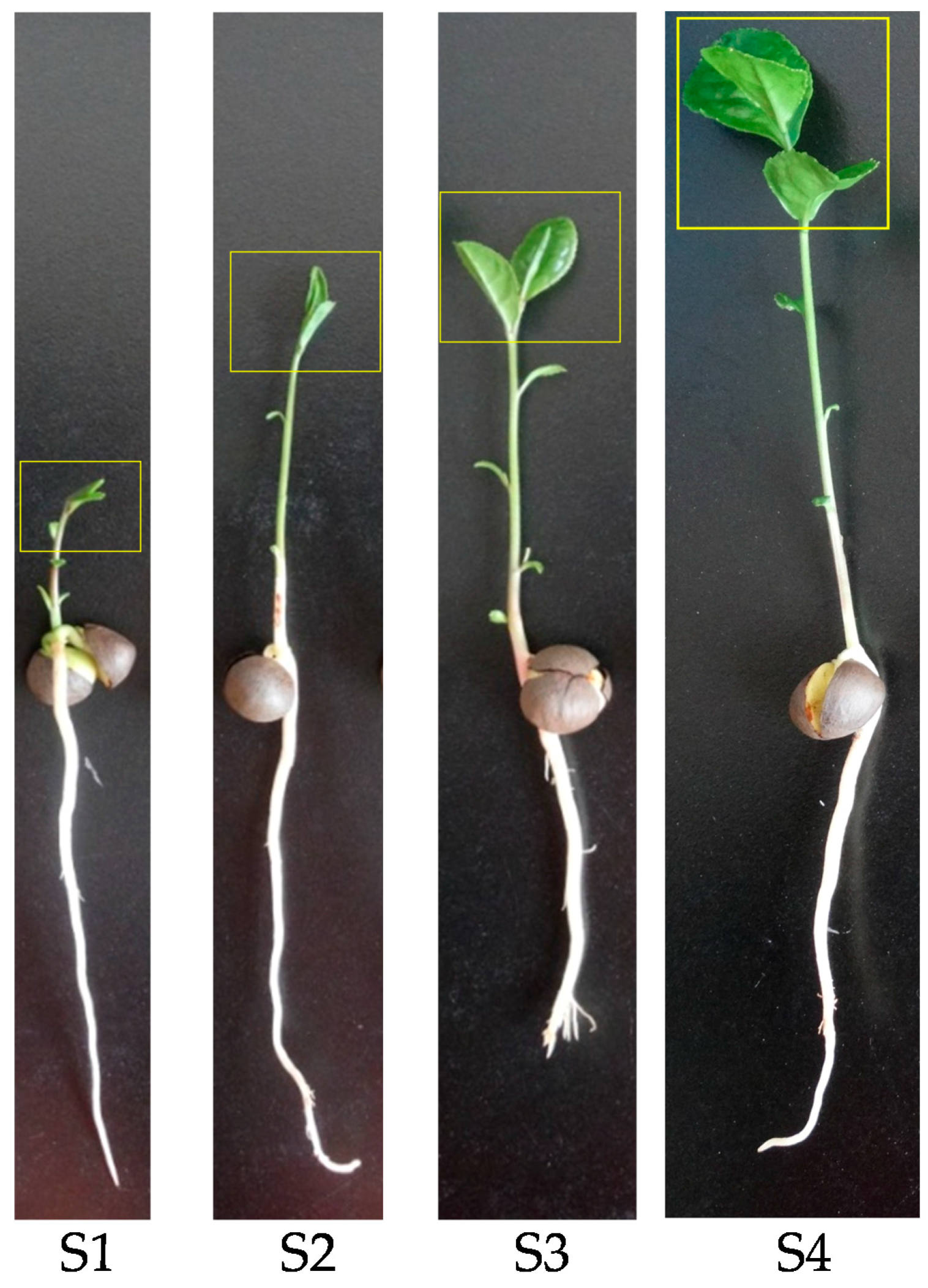
2.3. Gene Expression and Flavan 3-ol Accumulation in Tea Seed Germination
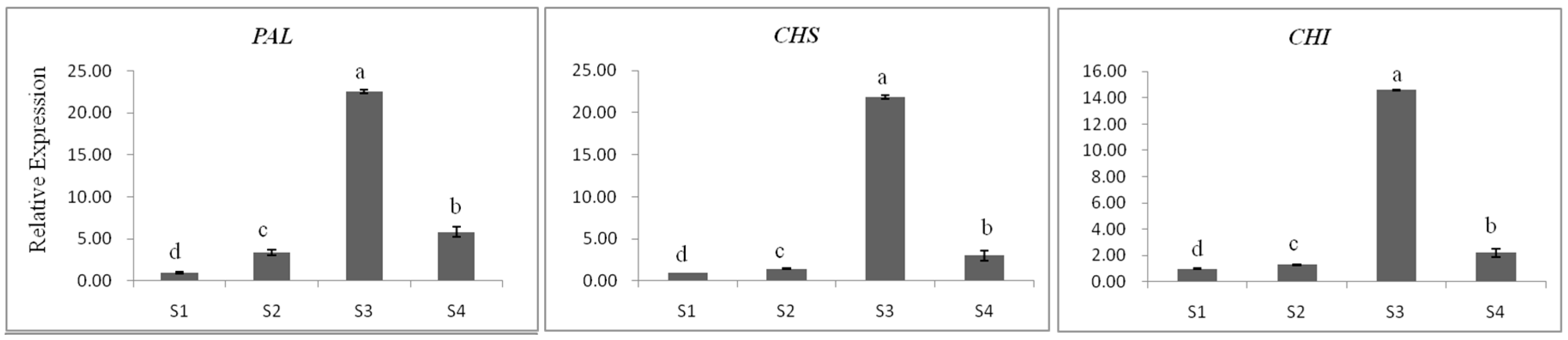
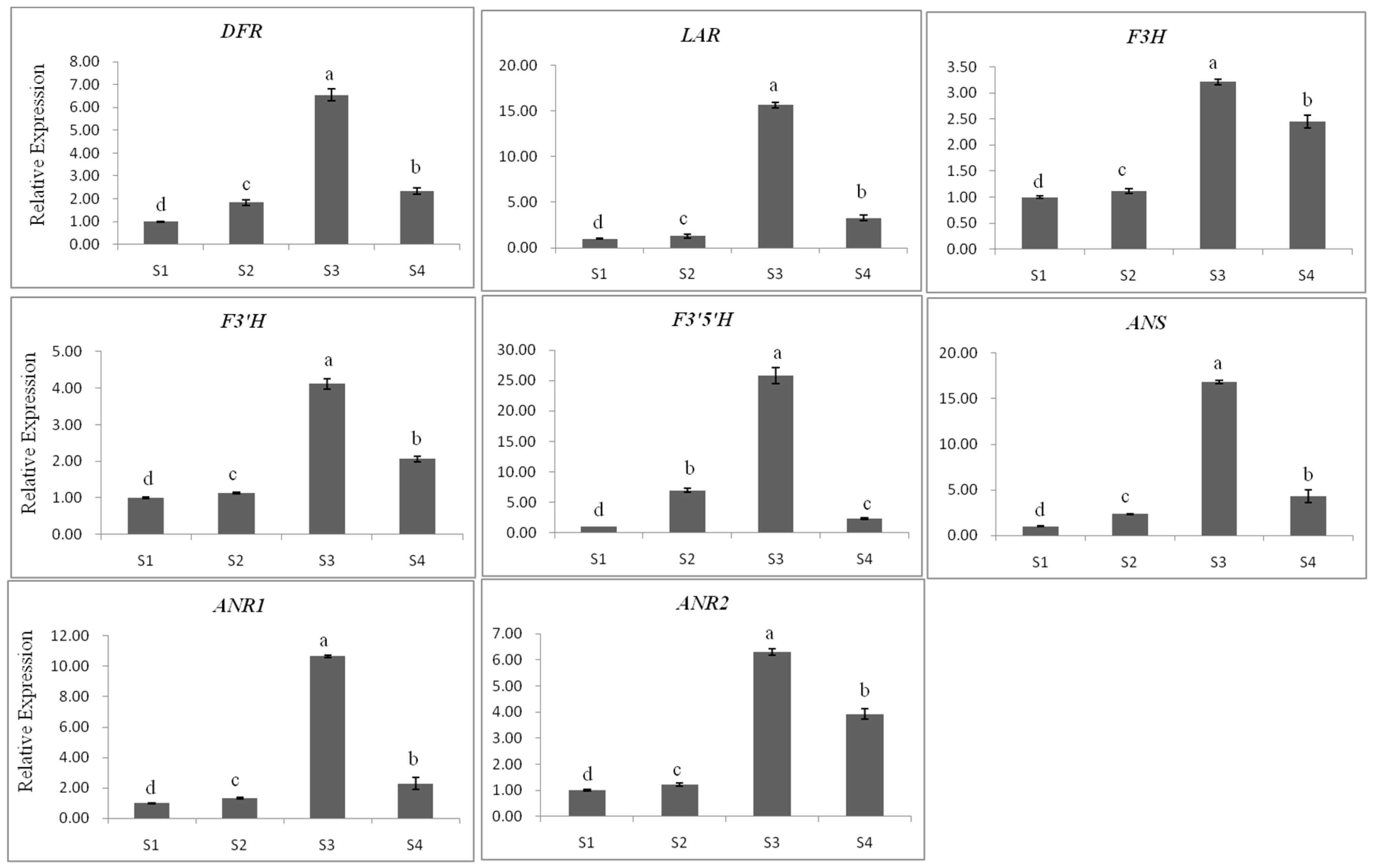
2.3.1. Gene Expression
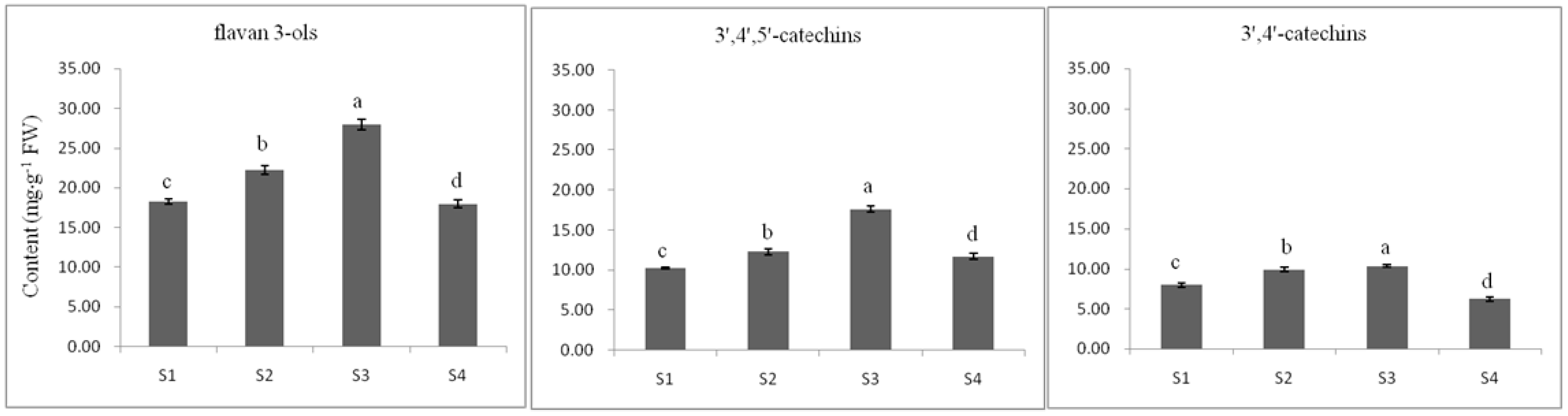
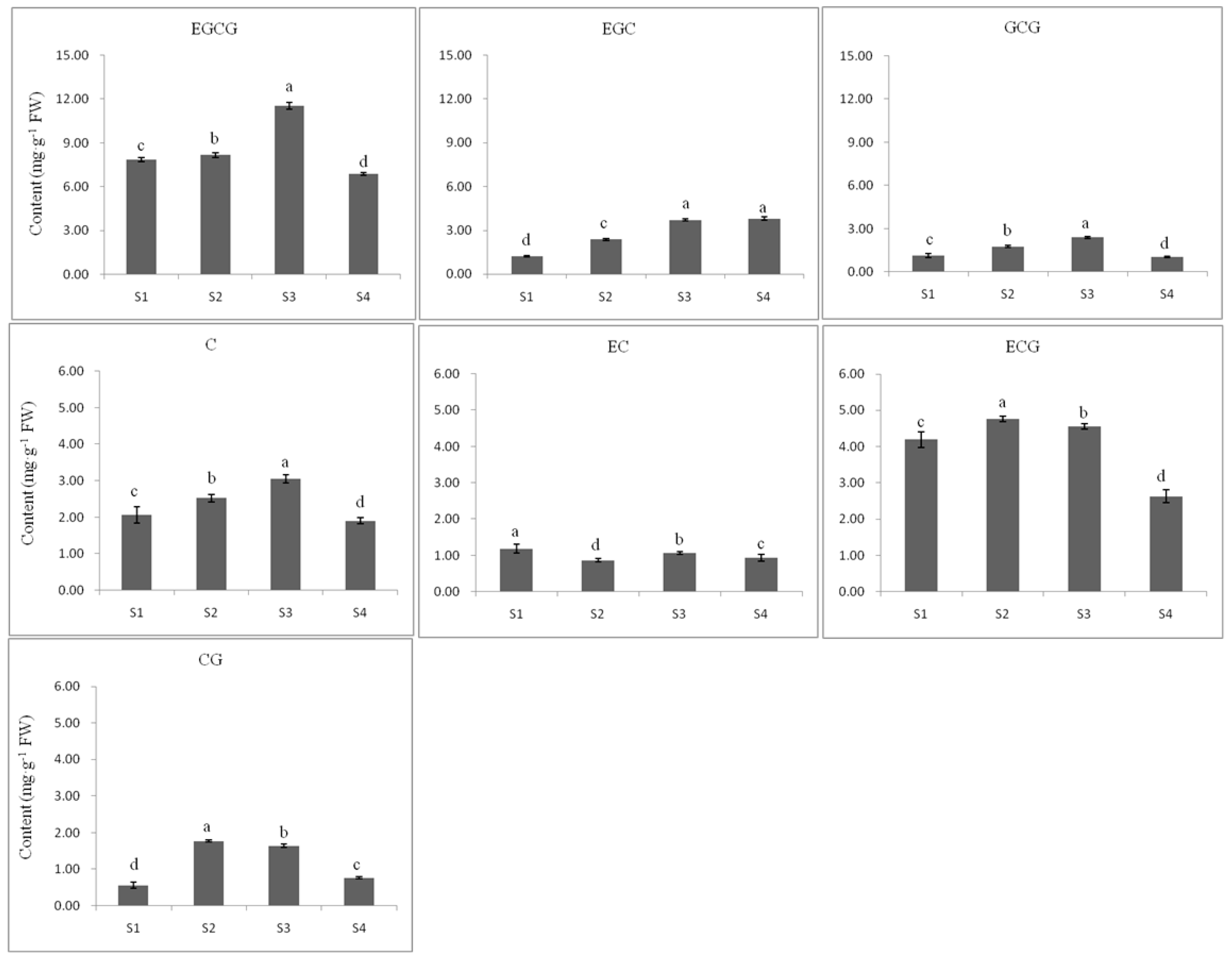
2.3.2. Flavan 3-ol Accumulation in Tea Seed Germination
2.3.3. The Relationship between Gene Expression and Accumulation of Flavan 3-ols
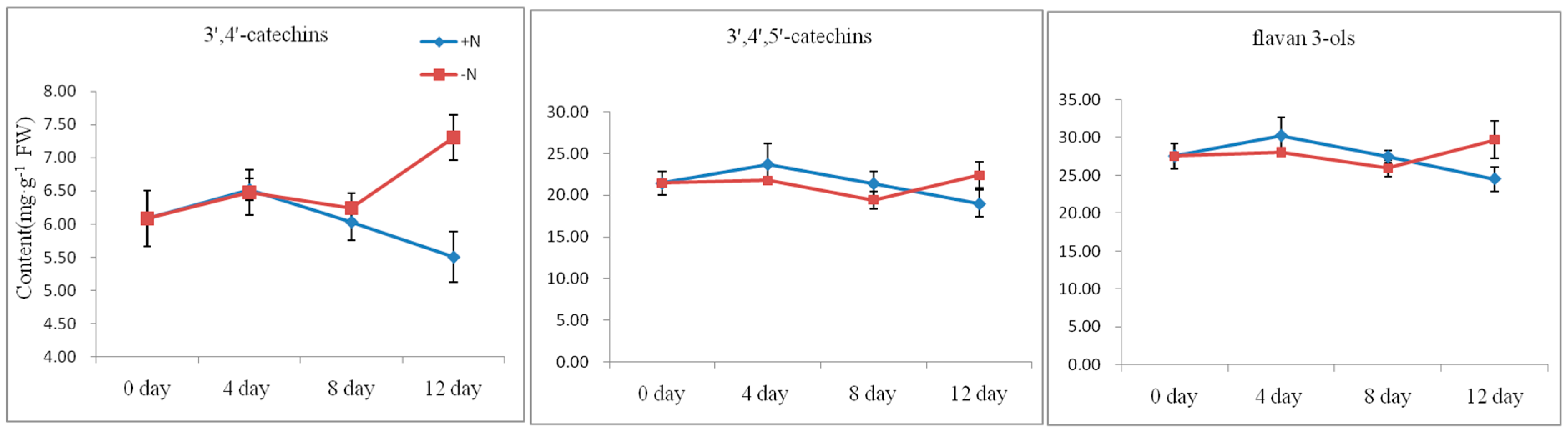
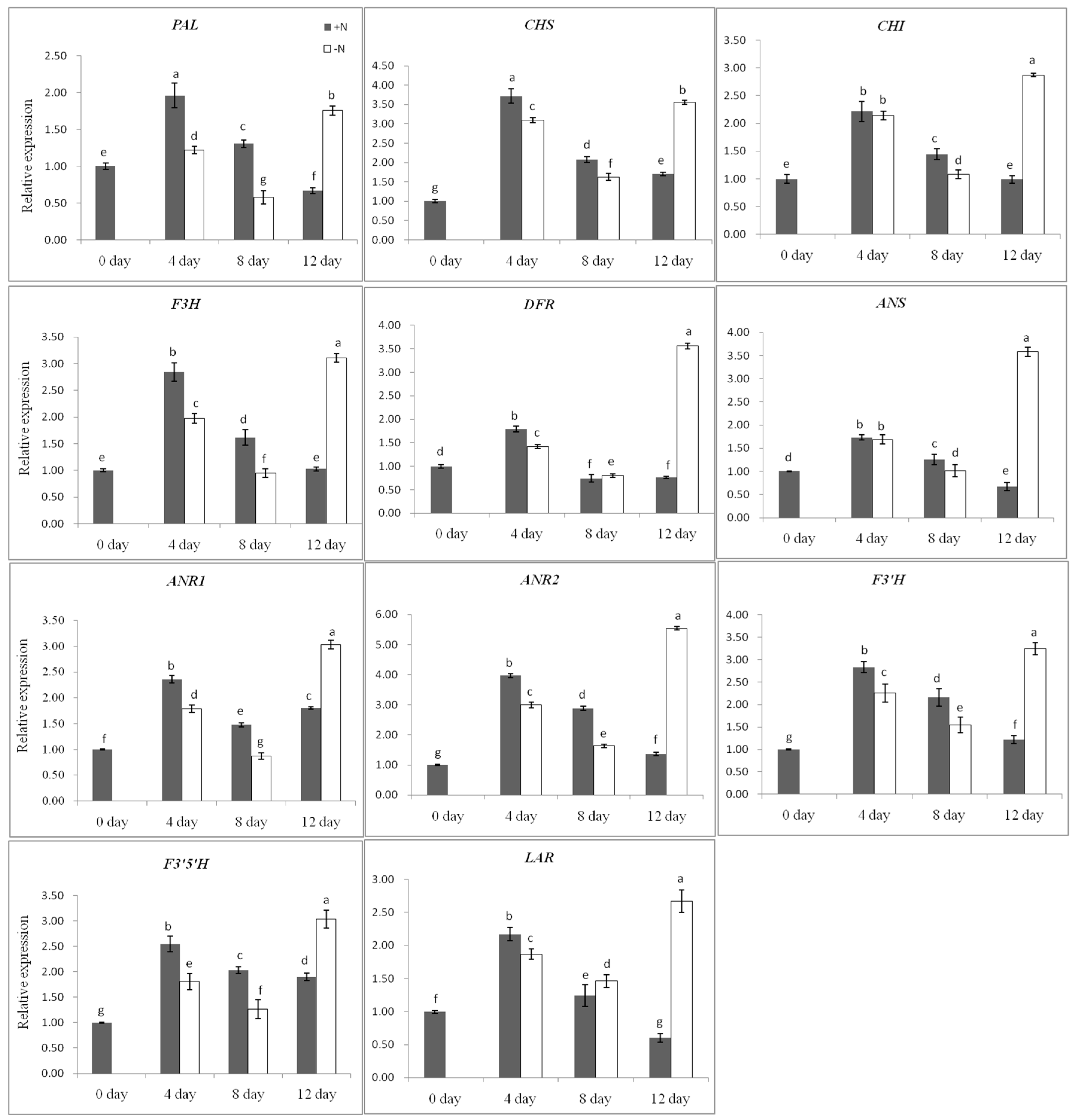
2.4. Flavan 3-ols Accumulation and Gene Expression Response to Nitrogen Dificiency
3. Experimental Section
3.1. Plant Materials
3.2. Isolation of the Flavonoid 3′-Hydroxylase Gene
3.3. Bioinformatic Analysis
3.4. Yeast Expression and Microsome Preparation
3.5. Enzyme Assays
3.6. HPLC Analysis
3.6.1. Flavonoid Standards
3.6.2. Analysis of Enzyme Substrates and Products
3.6.3. Analysis of Flavan 3-ols in Leaves
3.7. Expression of Genes Related to Flavan 3-ol Biosynthesis by Real-Time PCR
4. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Lin, Y.S.; Tsai, Y.J.; Tsay, J.S.; Lin, J.K. Factors affecting the levels of tea polyphenols and caffeine in tea leaves. J. Agric. Food Chem. 2003, 51, 1864–1873. [Google Scholar] [CrossRef] [PubMed]
- Bhattacharya, U.; Adak, S.; Majumder, N.S.; Bera, B.; Giri, A.K. Antimutagenic and anticancer activity of Darjeeling tea in multiple test systems. BMC Complement. Altern. Med. 2014, 14, 327. [Google Scholar] [CrossRef] [PubMed]
- Tijburg, L.B.M.; Mattern, T.; Folts, J.D.; Weisgerber, U.M.; Katan, M.B. Tea flavonoids and cardiovascular diseases: A review. Crit. Rev. Food Sci. 1997, 37, 771–785. [Google Scholar] [CrossRef] [PubMed]
- Fukai, K.; Ishigami, T.; Hara, Y. Antibacterial activity of tea polyphenols against phytopathogenic bacteria. Agric. Biol. Chem. 1991, 55, 1895–1897. [Google Scholar] [CrossRef]
- Cavet, M.E.; Harrington, K.L.; Vollmer, T.R.; Ward, K.W.; Zhang, J.Z. Anti-inflammatory and anti-oxidative effects of the green tea polyphenol epigallocatechin gallate in human corneal epithelial cells. Mol. Vis. 2011, 17, 533–542. [Google Scholar] [PubMed]
- Matsuo, N.; Yamada, K.; Shoji, K.; Mori, M.; Sugano, M. Effect of tea polyphenols on histamine release from rat basophilic leukemia (RBL-2H3) cells: The structure-inhibitory activity relationship. Allergy Eur. J. Allergy Clin. Immunol. 1997, 52, 58–64. [Google Scholar] [CrossRef]
- Dixon, R.A.; Pasinetti, G.M. Flavonoids and isoflavonoids: From plant biology to agriculture and neuroscience. Plant Physiol. 2010, 154, 453–457. [Google Scholar] [CrossRef] [PubMed]
- Soobrattee, M.A.; Neergheen, V.S.; Luximon, R.A.; Aruoma, O.I.; Bahorun, T. Phenolics as potential antioxidant therapeutic agents: Mechanism and actions. Mutat. Res. Fund Mol. M 2005, 579, 200–213. [Google Scholar] [CrossRef] [PubMed]
- Winkel-Shirley, B. Flavonoid biosynthesis: A colorful model for genetics, biochemistry, cell biology, and biotechnology. Plant Physiol. 2001, 126, 485–493. [Google Scholar] [CrossRef] [PubMed]
- Seitz, C.; Eder, C.; Deiml, B.; Kellner, S.; Martens, S.; Forkmann, G. Cloning, functional identification and sequence analysis of flavonoid 3′-hydroxylase and flavonoid 3′,5′-hydroxylase cDNAs reveals independent evolution of flavonoid 3′,5′-hydroxylase in the Asteraceae family. Plant Mol. Biol. 2006, 61, 365–381. [Google Scholar] [CrossRef] [PubMed]
- Ashihara, H.; Deng, W.W.; Mullen, W.; Crozier, A. Distribution and biosynthesis of flavan-3-ols in Camellia sinensis seedlings and expression of genes encoding biosynthetic enzymes. Phytochemistry 2010, 71, 559–566. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.S.; Xu, Y.J.; Gao, L.P.; Yu, O.; Wang, X.Z.; He, X.J.; Jiang, X.L.; Liu, Y.J.; Xia, T. Functional analysis of flavonoid 3′,5′-hydroxylase from Tea plant (Camellia sinensis): Critical role in the accumulation of catechins. BMC Plant Biol. 2014, 14, 374. [Google Scholar] [CrossRef] [PubMed]
- Wei, K.; Wang, L.Y.; Zhou, J.; He, W.; Zeng, J.M.; Jiang, Y.W.; Cheng, H. Catechin contents in tea (Camellia sinensis) as affected by cultivar and environment and their relation to chlorophyll contents. Food Chem. 2011, 125, 44–48. [Google Scholar] [CrossRef]
- Jin, J.Q.; Ma, J.Q.; Ma, C.L.; Yao, M.Z.; Chen, L. Determination of catechin content in representative Chinese tea germplasms. J. Agric. Food Chem. 2014, 62, 9436–9441. [Google Scholar] [CrossRef] [PubMed]
- Owuor, P.O.; Obanda, M. The use of green tea (Camellia sinensis) leaf flavan-3-ol composition in predicting plain black tea quality potential. Food Chem. 2007, 100, 873–884. [Google Scholar] [CrossRef]
- Olsen, K.M.; Hehn, A.; Jugde, H.; Slimestad, R.; Larbat, R.; Bourgaud, F.; Lillo, C. Identification and characterisation of CYP75A31, a new flavonoid 3′,5′-hydroxylase, isolated from Solanum. lycopersicum. BMC Plant Biol. 2010, 10, 21. [Google Scholar] [CrossRef] [PubMed]
- Brugliera, F.; Barri-Rewell, G.; Holton, T.A.; Mason, J.G. Isolation and characterization of a flavonoid 3′-hydroxylase cDNA clone corresponding to the Ht1 locus of Petunia hybrida. Plant J. 1999, 19, 441–451. [Google Scholar] [CrossRef] [PubMed]
- Schoenbohm, C.; Martens, S.; Eder, C.; Forkmann, G.; Weisshaar, B. Identification of the Arabidopsis thaliana flavonoid 3′-hydroxylase gene and functional expression of the encoded P450 enzyme. Biol. Chem. 2000, 381, 749–753. [Google Scholar] [CrossRef] [PubMed]
- Werck-Reichhart, D.; Bak, S.; Paquette, S. Cytochromes P450. In The Arabidopsis Book; Somerville, C.R., Meyerowitz, E.M., Eds.; NCBI: Rockville, MD, USA, 2002. [Google Scholar]
- Wei, K.; Wang, L.Y.; Zhang, C.C.; Wu, L.Y.; Li, H.L.; Zhang, F.; Cheng, H. Transcriptome analysis reveals key flavonoid 3′-hydroxylase and flavonoid 3′,5′-hydroxylase genes in affecting the ratio of dihydroxylated to trihydroxylated catechins in Camellia sinensis. PLoS ONE 2015. [Google Scholar] [CrossRef] [PubMed]
- Lillo, C.; Lea, U.S.; Ruoff, P. Nutrient depletion as a key factor for manipulating gene expression and product formation in different branches of the flavonoid pathway. Plant Cell Environ. 2008, 31, 587–601. [Google Scholar] [CrossRef] [PubMed]
- Olsen, K.M.; Slimestad, R.; Lea, U.S.; Brede, C.; Løvdal, T.; Ruoff, P.; Verheul, M.; Lillo, C. Temperature and nitrogen effects on regulators and products of the flavonoid pathway: Experimental and kinetic model studies. Plant Cell Environ. 2009, 32, 286–299. [Google Scholar] [CrossRef] [PubMed]
- Yamazaki, S.; Sato, K.; Suhara, K.; Sakaguchi, M.; Mihara, K.; Omura, T. Importance of the proline-rich region following signal-anchor sequence in the formation of correct conformation of microsomal cytochrome P450s. J. Biochem. 1993, 114, 652–657. [Google Scholar] [PubMed]
- Murakami, K.; Mihara, K.; Omura, T. The transmembrane region of microsomal cytochrome P450 identified as the endoplasmic reticulum retention signal. J. Biochem. 1994, 116, 164–175. [Google Scholar] [PubMed]
- Werck-Reichhart, D.; Feyereisen, R. Cytochromes P450: A success story. Genome Biol. 2000, 1, 3003.1–3003.9. [Google Scholar] [CrossRef] [PubMed]
- Chapple, C. Molecular-genetic analysis of plant cytochrome P450-dependent monooxygenases. Annu. Rev. Plant Physiol. Plant Mol. Biol. 1998, 49, 311–343. [Google Scholar] [CrossRef] [PubMed]
- Hasemann, C.A.; Kurumbail, R.G.; Boddupalli, S.S.; Peterson, J.A.; Deisenhofer, J. Structure and function of cytochrome P450: A comparative analysis of three crystal structures. Structure 1995, 3, 41–62. [Google Scholar] [CrossRef]
- Boddu, J.; Svabek, C.; Sekhon, R.; Gevens, A.; Nicholson, R.L.; Jones, A.D.; Pedersen, J.F.; Gustine, D.L.; Chopra, S. Expression of a putative flavonoid 3′-hydroxylase in sorghum mesocotyls synthesizing 3-deoxyanthocyanidin phytoalexins. Physiol. Mol. Plant Pathol. 2004, 65, 101–113. [Google Scholar] [CrossRef]
- Han, Y.P.; Vimolmangkang, S.; Soria-Guerra, R.E.; Rosales-Mendoza, S.; Zheng, D.M.; Lygin, A.V.; Korban, S.S. Ectopic expression of apple F3′H genes contribute to anthocyanin accumulation in the Arabidopsis tt7 mutant grown under nitrogen stress. Plant Physiol. 2010, 153, 806–820. [Google Scholar] [CrossRef] [PubMed]
- Takatori, Y.; Shimizu, K.; Ogata, J.; Endo, H.; Ishimaru, K.; Okamoto, S.; Hashimoto, F. Cloning of the flavonoid 3′-hydroxylase gene of Eustoma grandiflorum (Raf.) Shinn. (EgF3′H) and complementation of an F3′H-deficient Mutant of Ipomoea nil (L.) Roth. by heterologous expression of EgF3′H. Horic. J. 2015, 84, 131–139. [Google Scholar] [CrossRef]
- Zhou, W.; Gong, Y.F.; Lu, X.; Huang, C.T.; Gao, F. Molecular cloning and characterization of a flavonoid 3′-hydroxylase gene from purple-fleshed sweet potato (Ipomoea batatas). Mol. Biol. Rep. 2012, 39, 295–302. [Google Scholar] [CrossRef] [PubMed]
- Pompon, D.; Louerat, B.; Bronine, A.; Urban, P. Yeast expression of animal and plant P450s in optimized redox environments. Methods Enzymol. 1996, 272, 51–64. [Google Scholar] [PubMed]
- Teoh, K.H.; Polichuk, D.R.; Reed, D.W.; Nowak, G.; Covello, P.S. Artemisia annua L. (Asteraceae) trichome-specific cDNAs reveal CYP71AV1, a cytochrome P450 with a key role in the biosynthesis of the antimalarial sesquiterpene lactone artemisinin. FEBS Lett. 2006, 580, 1411–1416. [Google Scholar] [CrossRef] [PubMed]
- Ueyama, Y.; Suzuki, K.; Fukuchi-Mizutani, F.; Fukui, Y.; Miyazaki, K.; Ohkawa, H.; Kusumi, T.; Tanaka, Y. Molecular and biochemical characterization of torenia flavonoid 3′-hydroxylase and flavone synthase and modification of flower color by modulating the expression of these genes. Plant Sci. 2002, 163, 253–263. [Google Scholar] [CrossRef]
- Doostdar, H.; Shapiro, J.P.; Niedz, R.; Burke, M.D.; McCollum, T.G.; McDonald, R.E.; Mayer, R.T. A cytochrome P450 mediated naringenin 3′-hydroxylase from sweet orange cell cultures. Plant Cell Physiol. 1995, 36, 69–77. [Google Scholar]
- Schwinn, K.; Miosic, S.; Davies, K.; Thill, J.; Gotame, T.P.; Stich, K.; Halbwirth, H. The B-ring hydroxylation pattern of anthocyanins can be determined through activity of the flavonoid 3′-hydroxylase on leucoanthocyanidins. Planta 2014, 240, 1003–1010. [Google Scholar] [CrossRef] [PubMed]
- Fritsch, H.; Grisebach, H. Biosynthesis of cyanidin in cell cultures of Haplopappus gracilis. Phytochemistry 1975, 14, 2437–2442. [Google Scholar] [CrossRef]
- Hagmann, M.L.; Heller, W.; Grisebach, H. Induction and characterization of a microsomal Flavonoid 3′-hydroxylase from parsley cell cultures. FEBS J. 1983, 134, 547–554. [Google Scholar] [CrossRef]
- Xiong, L.G.; Li, J.; Li, Y.H.; Yuan, L.; Liu, S.Q.; Huang, J.A.; Liu, Z.H. Dynamic changes in catechin levels and catechin biosynthesis-related gene expression in albino tea plants (Camellia sinensis L). Plant Physiol. Biochem. 2013, 71, 132–143. [Google Scholar] [CrossRef] [PubMed]
- Løvdal, T.; Olsen, K.M.; Slimestad, R.; Verheul, M.; Lillo, C. Synergetic effects of nitrogen depletion, temperature, and light on the content of phenolic compounds and gene expression in leaves of tomato. Phytochemistry 2010, 71, 605–613. [Google Scholar] [CrossRef] [PubMed]
- Larbat, R.; Olsen, K.M.; Slimestad, R.; Løvdal, T.; Bénard, C.; Verheul, M.; Bourgaud, F.; Robin, C.; Lillo, C. Influence of repeated short-term nitrogen limitations on leaf phenolics metabolism in tomato. Phytochemistry 2012, 77, 119–128. [Google Scholar] [CrossRef] [PubMed]
- Nemie-Feyissa, D.; Olafsdottir, S.M.; Heidari, B.; Lillo, C. Nitrogen depletion and small R3-MYB transcription factors affecting anthocyanin accumulation in Arabidopsis leaves. Phytochemistry 2014, 98, 34–40. [Google Scholar] [CrossRef] [PubMed]
- Olsen, K.M.; Lea, U.S.; Slimestad, R.; Verheul, M.; Lillo, C. Differential expression of four Arabidopsis PAL genes; PAL1 and PAL2 have functional specialization in abiotic environmental-triggered flavonoid synthesis. J. Plant Physiol. 2008, 165, 1491–1499. [Google Scholar] [CrossRef] [PubMed]
- Konishi, S.; Miyamoto, S.; Taki, T. Stimulatory effects of aluminum on tea plants grown under low and high phosphorus supply. Soil Sci. Plant Nutr. 1985, 31, 361–368. [Google Scholar] [CrossRef]
- Tamura, K.; Peterson, D.; Peterson, N.; Stecher, G.; Nei, M.; Kumar, S. Mega 5: Molecular evolutinary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol. 2011, 28, 2731–2739. [Google Scholar] [CrossRef] [PubMed]
- Pfaffl, M.W. A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res. 2001, 29, 2002–2007. [Google Scholar] [CrossRef]


| Substrate | Naringenin | Dihydrokaempferol | Kaempferol |
|---|---|---|---|
| Km (μM) | 17.08 ± 0.46 | 143.64 ± 0.69 | 68.06 ± 0.43 |
| Vmax (pM·min−1) | 0.98 ± 0.01 | 0.19 ± 0.01 | 0.44 ± 0.01 |
| kcat (pM·min−1·mg−1·microsome) | 49.09 ± 0.21 | 9.86 ± 0.13 | 21.88 ± 0.35 |
| kcat / Km (×10−3 min−1·mg−1·microsome) | 2.88 ± 0.07 | 0.07 ± 0.01 | 0.32 ± 0.01 |
| Correlation Coefficient | 3′,4′-Catechins | 3′,4′,5′-Catechins | Flavan 3-ols |
|---|---|---|---|
| PAL | 0.513 | 0.981 * | 0.885 |
| CHS | 0.552 | 0.970 * | 0.893 |
| CHI | 0.557 | 0.970 * | 0.895 |
| F3H | 0.077 | 0.812 | 0.591 |
| DFR | 0.526 | 0.986 * | 0.893 |
| ANS | 0.508 | 0.977 * | 0.879 |
| ANR1 | 0.531 | 0.971 * | 0.885 |
| ANR2 | 0.166 | 0.861 | 0.661 |
| LAR | 0.506 | 0.967 * | 0.872 |
| F3′H | 0.364 | 0.941 | 0.796 |
| F3′5′H | 0.723 | 0.990 * | 0.975 |
| Gene | Accession No. | Forward Primer (5′–3′) | Reverse Primer (5′–3′) | Product Length (bp) |
|---|---|---|---|---|
| PAL | D26596 | TCCAATTCCTTGCCAATCC | AACTGCCTCGGCTGTCTTTC | 106 |
| CHS | AY169403 | ACAAAGGCAATCAAAGAATGG | ATGGGCGAAGACCGAGTAG | 124 |
| CHI | DQ904329 | TGAGACTGAACCCAAGACCG | TAGATTTTGATGCCGATGCC | 114 |
| F3H | AY641730 | TACCATCACCCTGCTCCTCC | CATTCTTGAACCTCCCATTGC | 153 |
| F3′H | KT180309 | TCGACCAGAACGATTCCTACC | ACTGGACCATACGCAACCCTA | 134 |
| F3′5′H | DQ194358 | TCTCAATCTTCCCAGAGTCGC | CAGTCTTCGCATTCTTTCCAC | 173 |
| DFR | AB018685 | ATTCCCACCAAGCCTAATCAC | CCTGAGGACGCTCATACAAGA | 137 |
| ANS | AY830416 | TTCAAGGGTATGGGAGCAAA | TGCAGGAATGTAGTCGGTTG | 139 |
| LAR | GU992401 | AACTCACCCTAGTCCATGCCA | CACCCTCCTCTTTTCGTTGTA | 134 |
| ANR1 | GU992402 | CATAGCCGGTTGTGACCTTG | TGACACGTTTAACCGTTCCTG | 147 |
| ANR2 | GU992400 | CGAGACCCAGGCAATCAGA | ACCAGGTCACAACCCGCTA | 131 |
| β-actin | HQ420251.1 | GCCATCTTTG ATTGGAATGG | GGTGCCACAACCTTGATCTT | 175 |
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons by Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhou, T.-S.; Zhou, R.; Yu, Y.-B.; Xiao, Y.; Li, D.-H.; Xiao, B.; Yu, O.; Yang, Y.-J. Cloning and Characterization of a Flavonoid 3′-Hydroxylase Gene from Tea Plant (Camellia sinensis). Int. J. Mol. Sci. 2016, 17, 261. https://doi.org/10.3390/ijms17020261
Zhou T-S, Zhou R, Yu Y-B, Xiao Y, Li D-H, Xiao B, Yu O, Yang Y-J. Cloning and Characterization of a Flavonoid 3′-Hydroxylase Gene from Tea Plant (Camellia sinensis). International Journal of Molecular Sciences. 2016; 17(2):261. https://doi.org/10.3390/ijms17020261
Chicago/Turabian StyleZhou, Tian-Shan, Rui Zhou, You-Ben Yu, Yao Xiao, Dong-Hua Li, Bin Xiao, Oliver Yu, and Ya-Jun Yang. 2016. "Cloning and Characterization of a Flavonoid 3′-Hydroxylase Gene from Tea Plant (Camellia sinensis)" International Journal of Molecular Sciences 17, no. 2: 261. https://doi.org/10.3390/ijms17020261
APA StyleZhou, T.-S., Zhou, R., Yu, Y.-B., Xiao, Y., Li, D.-H., Xiao, B., Yu, O., & Yang, Y.-J. (2016). Cloning and Characterization of a Flavonoid 3′-Hydroxylase Gene from Tea Plant (Camellia sinensis). International Journal of Molecular Sciences, 17(2), 261. https://doi.org/10.3390/ijms17020261