Berberine Alleviates Olanzapine-Induced Adipogenesis via the AMPKα–SREBP Pathway in 3T3-L1 Cells
Abstract
:1. Introduction
2. Results
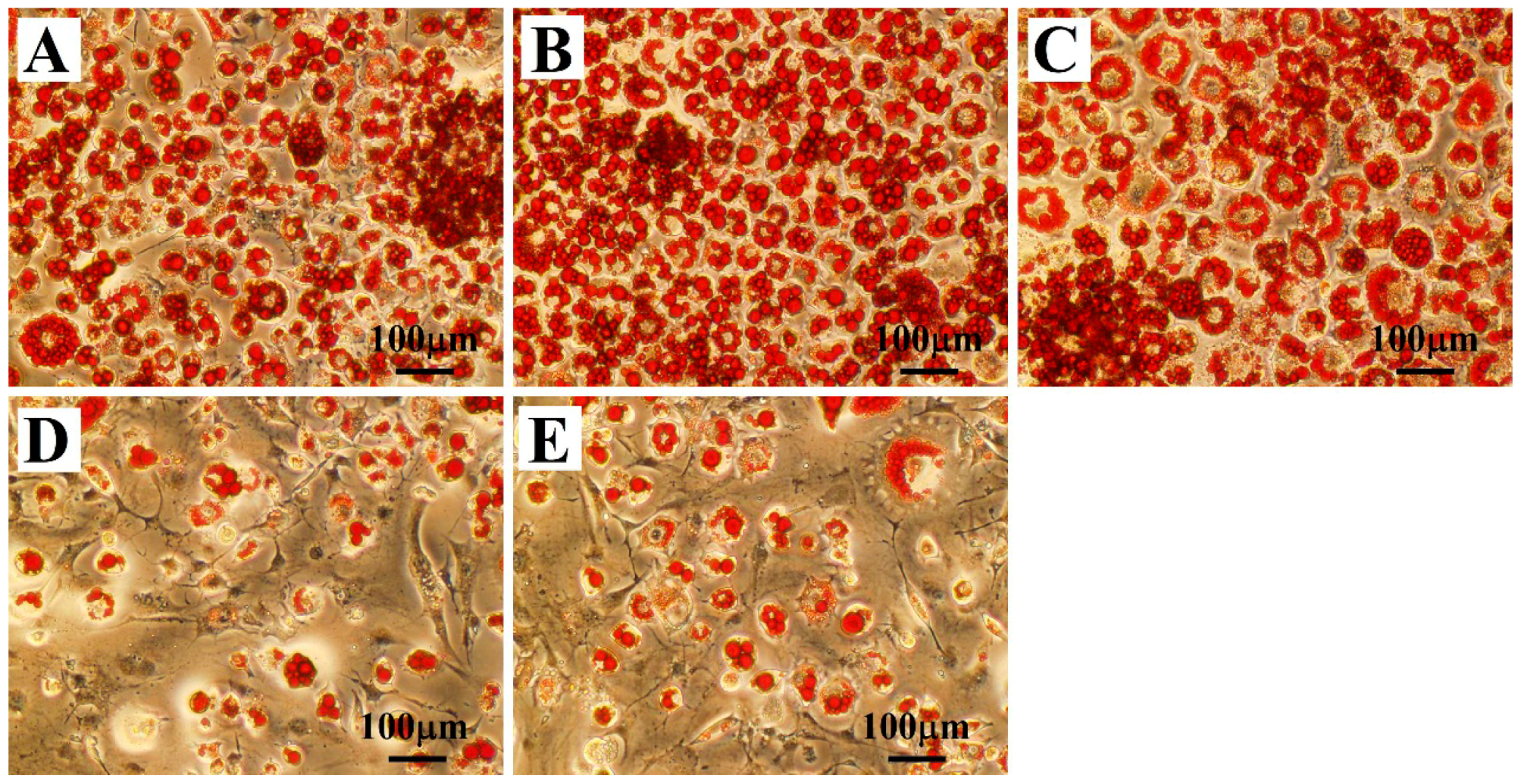
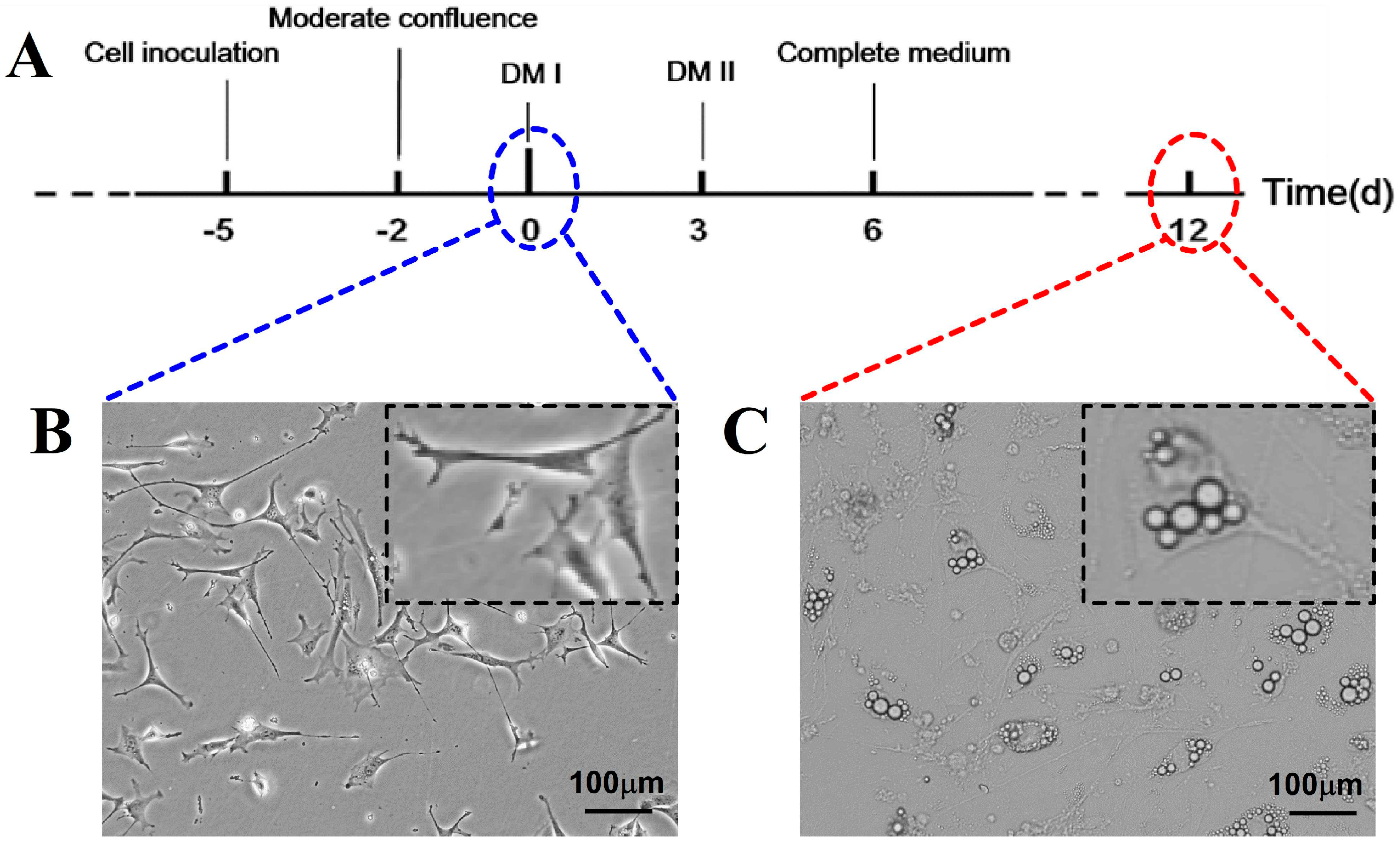
2.1. Effects of Treatment with OLZ, BBR, or Both on 3T3-L1 Differentiation
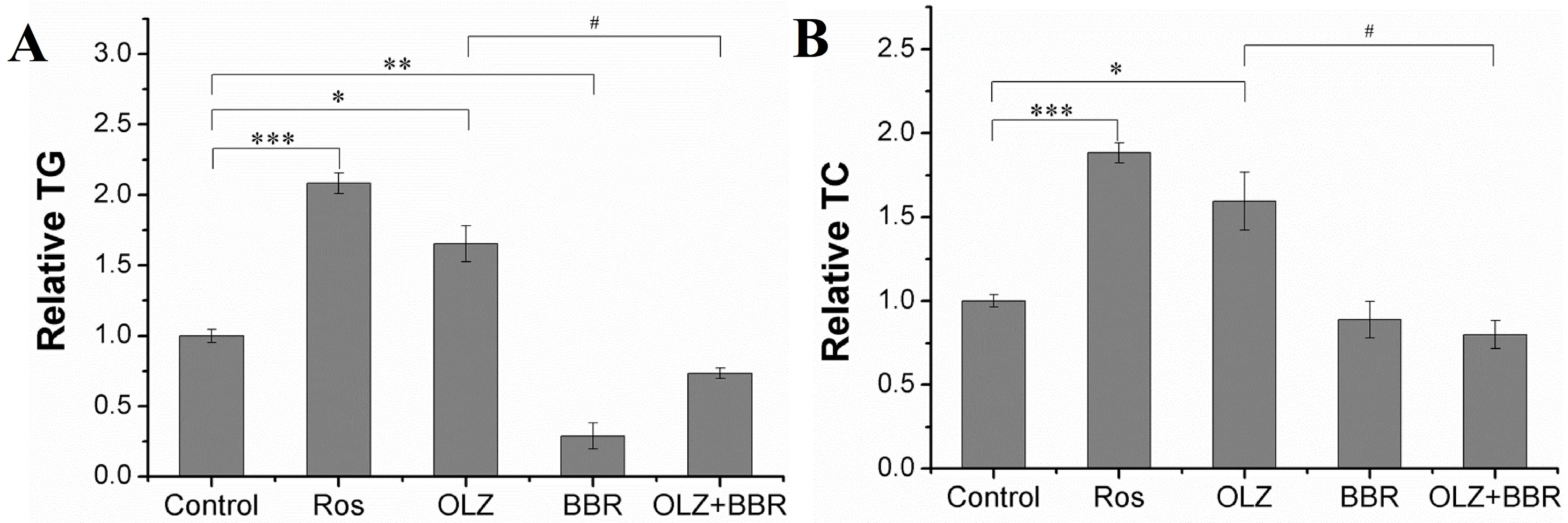
2.2. Effects of Treatment with OLZ, BBR, or Both on Biochemical Properties of 3T3-L1 Cells
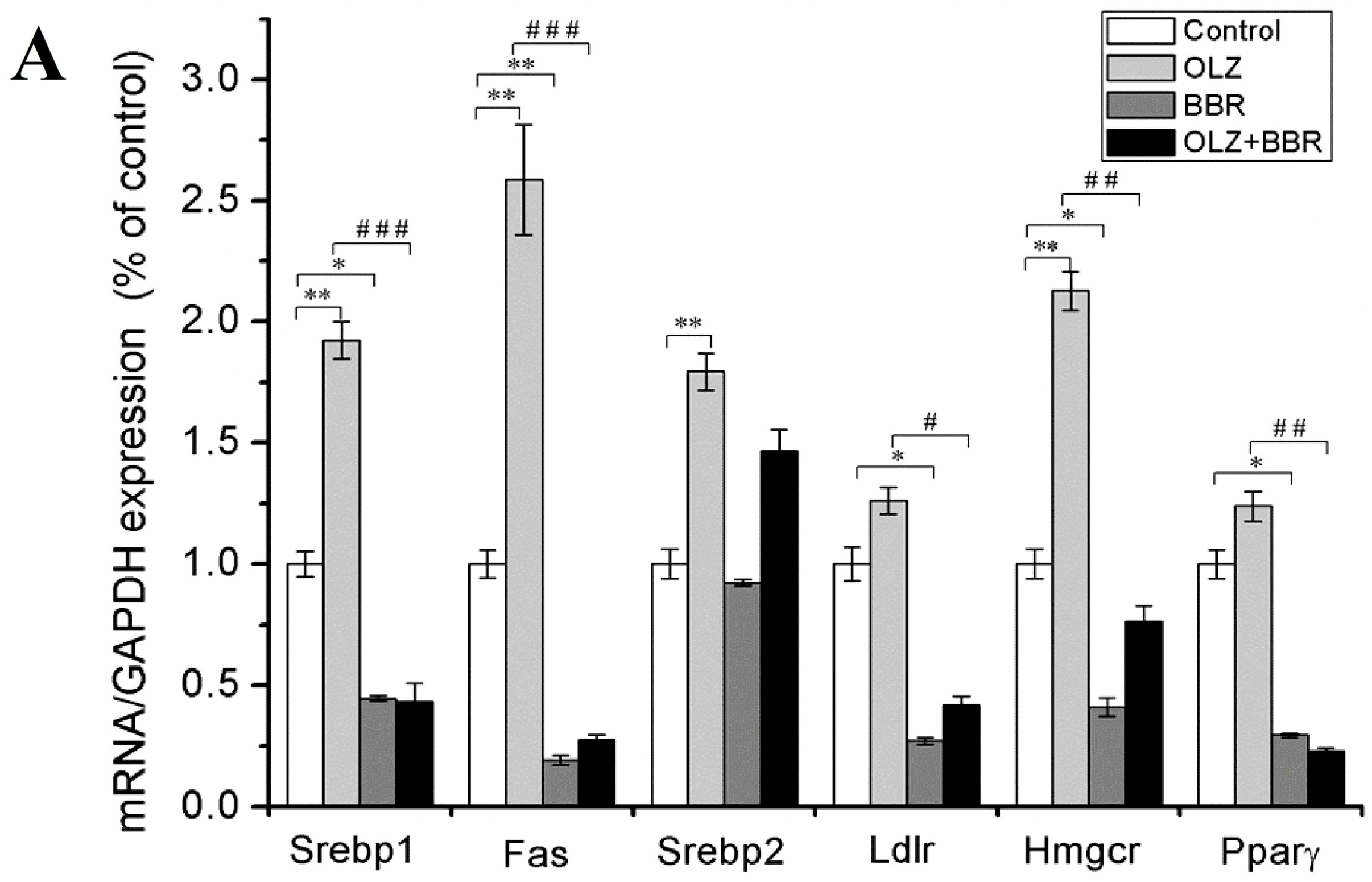
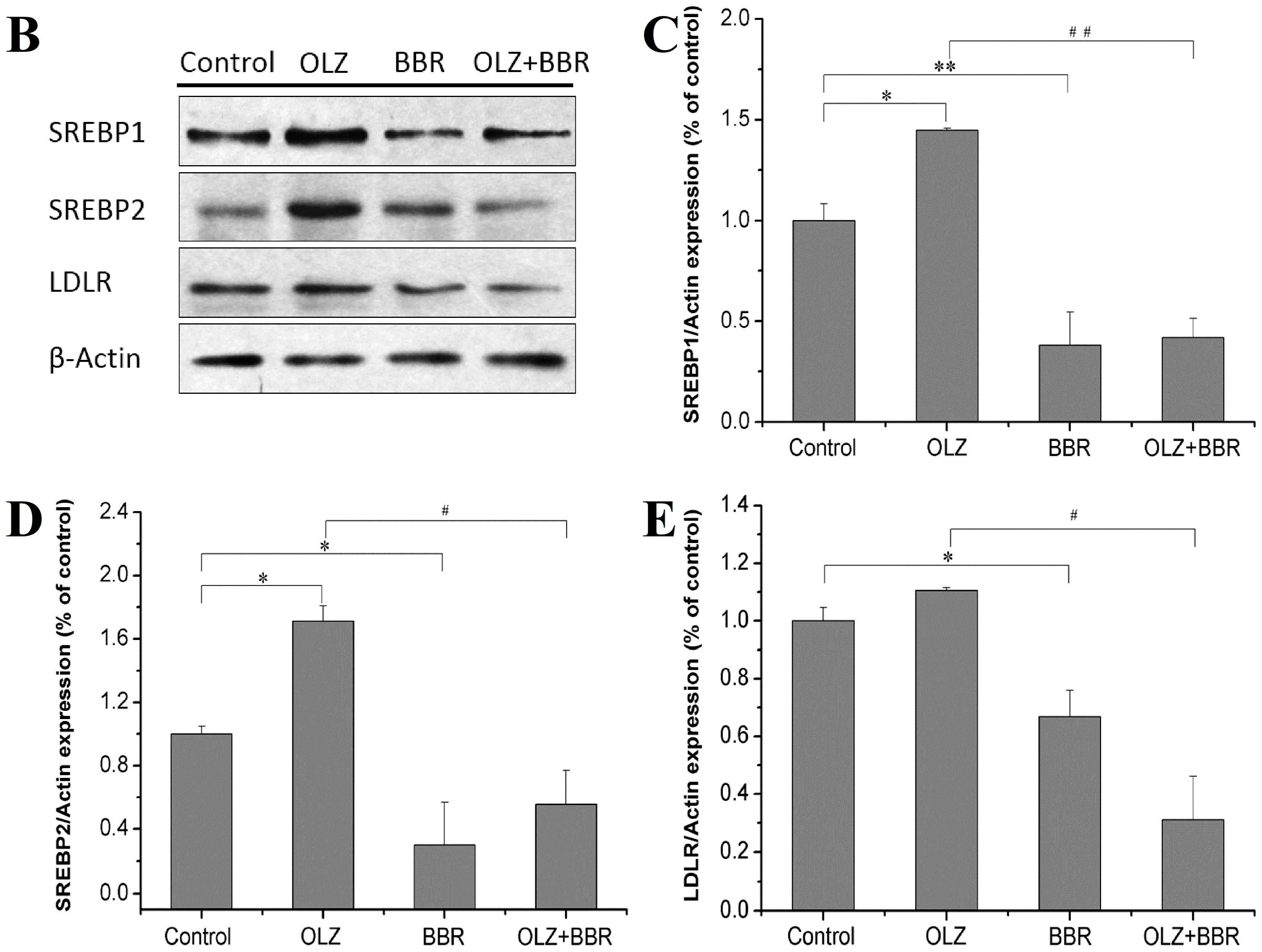
2.3. Effects of Treatment with OLZ, BBR, or Both on the Expression of SREBP-Related Genes for Adipogenesis in 3T3-L1 Cells
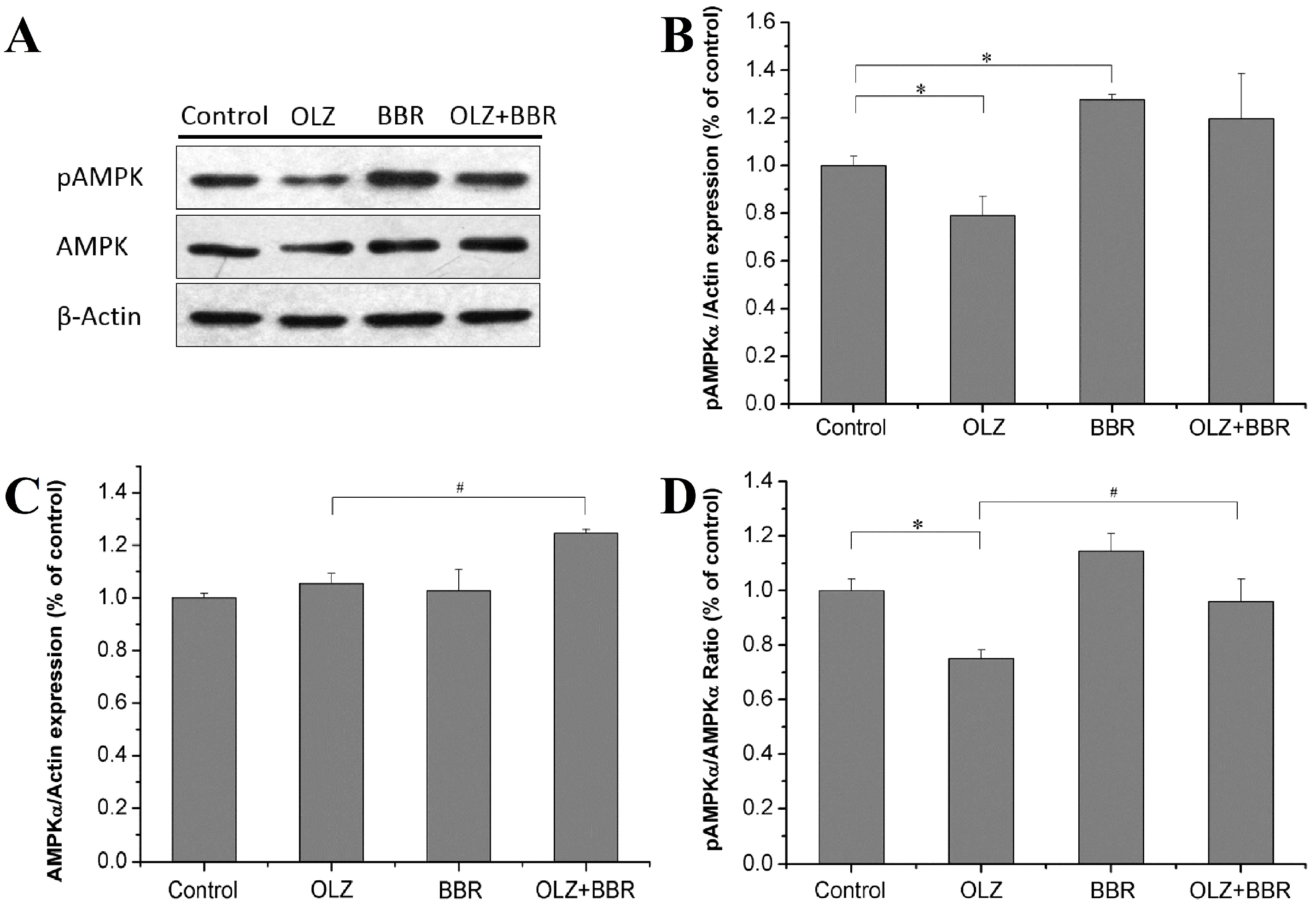
2.4. Effects of Treatment with OLZ, BBR, or Both on Expression of AMPKα-Related Adipogenesis in 3T3-L1 Cells
3. Discussion
4. Materials and Methods
4.1. Materials
4.2. Culturing 3T3-L1 Fibroblasts and Differentiation into Adipocytes
4.3. Oil-Red-O Staining
4.4. TG and TC Assay
4.5. Quantitative Real-Time PCR (qRT-PCR)
4.6. Western Blots
4.7. Statistics
5. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
| FGAs | first-generation antipsychotic drugs |
| SGAs | second-generation antipsychotic drugs |
| CNS | central nervous system |
| CREB | cAMP-response element-binding protein |
| C/EBPα | CCAAT/enhancer binding protein-α |
| SREBPs | sterol regulatory element binding proteins |
| LDLR | low-density lipoprotein receptor |
| FAS | fatty acid synthase |
| PPARγ | peroxisome proliferator activated receptor-γ |
| HMGR | hydroxymethylglutaryl-coenzyme A reductase |
| AMPKα | AMP-activated protein kinase-α |
| OLZ | olanzapine |
| BBR | berberine |
| Ros | rosiglitazone |
| TG | triglycerides |
| TC | total cholesterol |
| ORO | Oil-Red-O |
References
- Newton, S.S.; Duman, R.S. Neurogenic actions of atypical antipsychotic drugs and therapeutic implications. CNS Drugs 2007, 21, 715–725. [Google Scholar] [CrossRef] [PubMed]
- Leucht, S.; Corves, C.; Arbter, D.; Engel, R.R.; Li, C.B.; Davis, J.M. Second-generation versus first-generation antipsychotic drugs for schizophrenia: A meta-analysis. Lancet 2009, 373, 31–41. [Google Scholar] [CrossRef]
- Potvin, S.; Zhornitsky, S.; Stip, E. Antipsychotic-induced changes in blood levels of leptin in schizophrenia: A meta-analysis. Can. J. Psychiatry 2015, 60, 26–34. [Google Scholar]
- Bartoli, F.; Crocamo, C.; Clerici, M.; Carrà, G. Second-generation antipsychotics and adiponectin levels in schizophrenia: A comparative meta-analysis. Eur. Neuropsychopharmacol. 2015, 25, 1767–1774. [Google Scholar] [CrossRef] [PubMed]
- De Leon, J.; Susce, M.T.; Johnson, M.; Hardin, M.; Pointer, L.; Ruano, G.; Windemuth, A.; Diaz, F.J. A clinical study of the association of antipsychotics with hyperlipidemia. Schizophr. Res. 2007, 92, 95–102. [Google Scholar] [CrossRef] [PubMed]
- Lieberman, J.A. Effectiveness of antipsychotic drugs in patients with chronic schizophrenia. N. Engl. J. Med. 2010, 363, 1092–1093. [Google Scholar]
- Stahl, S.M.; Mignon, L.; Meyer, J.M. Which comes first: Atypical antipsychotic treatment or cardiometabolic risk? Acta Psychiat. Scand. 2009, 119, 171–179. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.M.; Deng, C.; Cao, S.; Gong, J.W.; Wang, B.C.; Hu, C.H. Acute effects of oral olanzapine treatment on the expression of fatty acid and cholesterol metabolism-related gene in rats. Life Sci. 2015, 128, 72–78. [Google Scholar] [CrossRef] [PubMed]
- Raeder, M.B.; Fernø, J.; Vik-Mo, A.O.; Steen, V.M. SREBP activation by antipsychotic-and antidepressant-drugs in cultured human liver cells: Relevance for metabolic side-effects? Mol. Cell. Biochem. 2006, 289, 167–173. [Google Scholar] [CrossRef] [PubMed]
- Oh, K.-J.; Park, J.; Lee, S.Y.; Hwang, I.; Kim, J.B.; Park, T.-S.; Lee, H.-J.; Koo, S.-H. Atypical antipsychotic drugs perturb AMPK-dependent regulation of hepatic lipid metabolism. Am. J. Physiol. Endocrinol. Metab. 2011, 300, 624–632. [Google Scholar] [CrossRef] [PubMed]
- Fernø, J.; Raeder, M.; Vik-Mo, A.; Skrede, S.; Glambek, M.; Tronstad, K.; Breilid, H.; Løvlie, R.; Berge, R.; Stansberg, C. Antipsychotic drugs activate SREBP-regulated expression of lipid biosynthetic genes in cultured human glioma cells: A novel mechanism of action? Pharmacogenom. J. 2005, 5, 298–304. [Google Scholar] [CrossRef] [PubMed]
- Yang, L.H.; Chen, T.M.; Yu, S.T.; Chen, Y.H. Olanzapine induces SREBP1-related adipogenesis in 3T3-L1 cells. Pharmacol. Res. 2007, 56, 202–208. [Google Scholar] [CrossRef] [PubMed]
- Lauressergues, E.; Staels, B.; Valeille, K.; Majd, Z.; Hum, D.W.; Duriez, P.; Cussac, D. Antipsychotic drug action on SREBPs-related lipogenesis and cholesterogenesis in primary rat hepatocytes. Naunyn-Schmiedebergs Arch. Pharmacol. 2010, 381, 427–439. [Google Scholar] [CrossRef] [PubMed]
- Brown, M.S.; Goldstein, J.L. The SREBP pathway: Regulation of cholesterol metabolism by proteolysis of a membrane-bound transcription factor. Cell 1997, 89, 331–340. [Google Scholar] [CrossRef]
- Brown, M.S.; Goldstein, J.L. Sterol regulatory element binding proteins (SREBPs): Controllers of lipid synthesis and cellular uptake. Nutr. Rev. 1998, 56, 1–3. [Google Scholar] [CrossRef]
- Horton, J.D.; Goldstein, J.L.; Brown, M.S. SREBPs: Activators of the complete program of cholesterol and fatty acid synthesis in the liver. J. Clin. Investig. 2002, 109, 1125–1132. [Google Scholar] [CrossRef] [PubMed]
- Goldstein, J.L.; DeBose-Boyd, R.A.; Brown, M.S. Protein sensors for membrane sterols. Cell 2006, 124, 35–46. [Google Scholar] [CrossRef] [PubMed]
- Ferre, P.; Azzout-Marniche, D.; Foufelle, F. AMP-activated protein kinase and hepatic genes involved in glucose metabolism. Biochem. Soc. Trans. 2003, 31, 220–223. [Google Scholar] [CrossRef] [PubMed]
- Park, K.G.; Min, A.K.; Koh, E.H.; Kim, H.S.; Kim, M.O.; Park, H.S.; Kim, Y.D.; Yoon, T.S.; Jang, B.K.; Hwang, J.S. α-lipoic acid decreases hepatic lipogenesis through adenosine monophosphate-activated protein kinase (AMPK)-dependent and AMPK-independent pathways. Hepatology 2008, 48, 1477–1486. [Google Scholar] [CrossRef] [PubMed]
- Viollet, B.; Foretz, M.; Guigas, B.; Horman, S.; Dentin, R.; Bertrand, L.; Hue, L.; Andreelli, F. Activation of AMP-activated protein kinase in the liver: A new strategy for the management of metabolic hepatic disorders. J. Physiol. 2006, 574, 41–53. [Google Scholar] [CrossRef] [PubMed]
- Kroeze, W.K.; Hufeisen, S.J.; Popadak, B.A.; Renock, S.M.; Steinberg, S.; Ernsberger, P.; Jayathilake, K.; Meltzer, H.Y.; Roth, B.L. H1-histamine receptor affinity predicts short-term weight gain for typical and atypical antipsychotic drugs. Neuropsychopharmacology 2003, 28, 519–526. [Google Scholar] [CrossRef] [PubMed]
- Jiang, W.; Wei, W.; Gaertig, M.A.; Li, S.; Li, X.-J. Therapeutic Effect of Berberine on Huntington’s Disease Transgenic Mouse Model. PLoS ONE 2015, 10, e0134142. [Google Scholar] [CrossRef] [PubMed]
- Čerňáková, M.; Košťálová, D. Antimicrobial activity of berberine—A constituent of Mahonia aquifolium. Folia Microbiol. 2002, 47, 375–378. [Google Scholar] [CrossRef]
- Piyanuch, R.; Sukhthankar, M.; Wandee, G.; Baek, S.J. Berberine, a natural isoquinoline alkaloid, induces NAG-1 and ATF3 expression in human colorectal cancer cells. Cancer Lett. 2007, 258, 230–240. [Google Scholar] [CrossRef] [PubMed]
- Ho, Y.T.; Yang, J.S.; Lu, C.C.; Chiang, J.H.; Li, T.C.; Lin, J.J.; Lai, K.C.; Liao, C.L.; Lin, J.G.; Chung, J.G. Berberine inhibits human tongue squamous carcinoma cancer tumor growth in a murine xenograft model. Phytomedicine 2009, 16, 887–890. [Google Scholar] [CrossRef] [PubMed]
- Kong, W.; Wei, J.; Abidi, P.; Lin, M.; Inaba, S.; Li, C.; Wang, Y.; Wang, Z.; Si, S.; Pan, H. Berberine is a novel cholesterol-lowering drug working through a unique mechanism distinct from statins. Nat. Med. 2004, 10, 1344–1351. [Google Scholar] [CrossRef] [PubMed]
- Huang, C.; Zhang, Y.; Gong, Z.; Sheng, X.; Li, Z.; Zhang, W.; Qin, Y. Berberine inhibits 3T3-L1 adipocyte differentiation through the PPARγ pathway. Biochem. Biophys. Res. Commun. 2006, 348, 571–578. [Google Scholar] [CrossRef] [PubMed]
- Hu, Y.; Kutscher, E.; Davies, G.E. Berberine inhibits SREBP1-related clozapine and risperidone induced adipogenesis in 3T3-L1 cells. Phytother. Res. 2010, 24, 1831–1838. [Google Scholar] [CrossRef] [PubMed]
- Kim, W.S.; Lee, Y.S.; Cha, S.H.; Jeong, H.W.; Choe, S.S.; Lee, M.R.; Oh, G.T.; Park, H.S.; Lee, K.U.; Lane, M.D. Berberine improves lipid dysregulation in obesity by controlling central and peripheral AMPK activity. Am. J. Physiol. Endocrinol. Metab. 2009, 296, 812–819. [Google Scholar] [CrossRef] [PubMed]
- Zhou, G.; Myers, R.; Li, Y.; Chen, Y.; Shen, X.; Fenyk-Melody, J.; Wu, M.; Ventre, J.; Doebber, T.; Fujii, N.; et al. Role of AMP-activated protein kinase in mechanism of metformin action. J. Clin. Investig. 2001, 108, 1167–1174. [Google Scholar] [CrossRef] [PubMed]
- Skrede, S.; Fernø, J.; Vázquez, M.J.; Fjær, S.; Pavlin, T.; Lunder, N.; Vidal-Puig, A.; Diéguez, C.; Berge, R.K.; López, M. Olanzapine, but not aripiprazole, weight-independently elevates serum triglycerides and activates lipogenic gene expression in female rats. Int. J. Neuropsychopharmacol. 2012, 15, 163–179. [Google Scholar] [CrossRef] [PubMed]
- Stanley, J.C. Feedback inhibition of cholesterol synthesis and uptake. Eur. J. Lipid Sci. Technol. 2008, 20, 17–18. [Google Scholar] [CrossRef]
- Canfrán-Duque, A.; Casado, M.E.; Pastor, Ó.; Sánchez-Wandelmer, J.; de la Peña, G.; Lerma, M.; Mariscal, P.; Bracher, F.; Lasunción, M.A.; Busto, R. Atypical antipsychotics alter cholesterol and fatty acid metabolism in vitro. J. Lipid Res. 2013, 54, 310–324. [Google Scholar] [CrossRef] [PubMed]
- Kristiana, I.; Sharpe, L.; Catts, V.; Lutze-Mann, L.; Brown, A. Antipsychotic drugs upregulate lipogenic gene expression by disrupting intracellular trafficking of lipoprotein-derived cholesterol. Pharmacogenom. J. 2010, 10, 396–407. [Google Scholar] [CrossRef] [PubMed]
- Lian, J.; Huang, X.-F.; Pai, N.; Deng, C. Betahistine ameliorates olanzapine-induced weight gain through modulation of histaminergic, NPY and AMPK pathways. Psychoneuroendocrino 2014, 48, 77–86. [Google Scholar] [CrossRef] [PubMed]
- Minokoshi, Y.; Kim, Y.-B.; Peroni, O.D.; Fryer, L.G.; Müller, C.; Carling, D.; Kahn, B.B. Leptin stimulates fatty-acid oxidation by activating AMP-activated protein kinase. Nature 2002, 415, 339–343. [Google Scholar] [CrossRef] [PubMed]
- Steinberg, G.R.; Rush, J.W.; Dyck, D.J. AMPK expression and phosphorylation are increased in rodent muscle after chronic leptin treatment. Am. J. Physiol. 2003, 284, 648–654. [Google Scholar] [CrossRef] [PubMed]
- You, M.; Matsumoto, M.; Pacold, C.M.; Cho, W.K.; Crabb, D.W. The role of AMP-activated protein kinase in the action of ethanol in the liver. Gastroenterology 2004, 127, 1798–1808. [Google Scholar] [CrossRef] [PubMed]
- Hu, Y.S.; Young, A.J.; Ehli, E.A.; Nowotny, D.; Davies, P.S.; Droke, E.A.; Soundy, T.J.; Davies, G.E. Metformin and berberine prevent olanzapine-induced weight gain in rats. PLoS ONE 2014, 9, e93310. [Google Scholar] [CrossRef] [PubMed]
- Davies, G.; Hu, Y. Pharmaceutical Compositions Containing Berberine for Treatment or Prevention of Weight Gain and Obesity Associated with Anti-Psychotic Drugs. U.S. Patent WO2011119649 A3, 23 February 2012. [Google Scholar]
- Li, Y.; Wang, P.; Zhuang, Y.; Lin, H.; Li, Y.; Liu, L.; Meng, Q.; Cui, T.; Liu, J.; Li, Z. Activation of AMPK by berberine promotes adiponectin multimerization in 3T3-L1 adipocytes. FEBS Lett. 2011, 585, 1735–1740. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Lian, J.; Hu, C.-H.; Deng, C. Betahistine co-treatment ameliorates dyslipidemia induced by chronic olanzapine treatment in rats through modulation of hepatic AMPKα–SREBP1 and PPARα-dependent pathways. Pharmacol. Res. 2015, 100, 36–46. [Google Scholar] [CrossRef] [PubMed]
| Oligonucleotide | Sequences (5’—3’) |
|---|---|
| GAPDH-Forward | GGTGAAGGTCGGTGTGAACG |
| GAPDH-Reverse | CTCGCTCCTGGAAGATGGTG |
| SREBP1-Forward | ACAAAAGCAAATCACTGAAGGACC |
| SREBP1-Reverse | CGGGCTCAGAGTCACTACCACC |
| FAS-Forward | GCACTGACTGTCTGTTTTCCAA |
| FAS-Reverse | AGCATCAAGAGCAGCATTTTTA |
| SREBP2-Forward | ACAACACTGACCAGCACCCATAC |
| SREBP2-Reverse | AAGACGCTCAAGACAATCACACC |
| LDLR-Forward | GGGTTGATTCCAAACTCCACTCTA |
| LDLR-Reverse | ACTGAAAATGGCTTCGTTTATGAC |
| HmgcR-Forward | CCAAACCCCGTAACCCAAAG |
| HmgcR-Reverse | GATAAAACTGCCAGAGAGAAACACT |
| PPARγ-Forward | ACAGGAAAGACAACGGACAAATCA |
| PPARγ-Reverse | CTTCTACGGATCGAAACTGGCAC |
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Li, Y.; Zhao, X.; Feng, X.; Liu, X.; Deng, C.; Hu, C.-H. Berberine Alleviates Olanzapine-Induced Adipogenesis via the AMPKα–SREBP Pathway in 3T3-L1 Cells. Int. J. Mol. Sci. 2016, 17, 1865. https://doi.org/10.3390/ijms17111865
Li Y, Zhao X, Feng X, Liu X, Deng C, Hu C-H. Berberine Alleviates Olanzapine-Induced Adipogenesis via the AMPKα–SREBP Pathway in 3T3-L1 Cells. International Journal of Molecular Sciences. 2016; 17(11):1865. https://doi.org/10.3390/ijms17111865
Chicago/Turabian StyleLi, Yanjie, Xiaomin Zhao, Xiyu Feng, Xuemei Liu, Chao Deng, and Chang-Hua Hu. 2016. "Berberine Alleviates Olanzapine-Induced Adipogenesis via the AMPKα–SREBP Pathway in 3T3-L1 Cells" International Journal of Molecular Sciences 17, no. 11: 1865. https://doi.org/10.3390/ijms17111865
APA StyleLi, Y., Zhao, X., Feng, X., Liu, X., Deng, C., & Hu, C.-H. (2016). Berberine Alleviates Olanzapine-Induced Adipogenesis via the AMPKα–SREBP Pathway in 3T3-L1 Cells. International Journal of Molecular Sciences, 17(11), 1865. https://doi.org/10.3390/ijms17111865







