miRNAs Regulation and Its Role as Biomarkers in Endometriosis
Abstract
:1. Introduction
| Symbol | Non-Coding RNAs | Functions |
|---|---|---|
| tRNA | Transfer RNA | mRNA translation (structural) |
| rRNA | Ribosomal RNA | mRNA translation (structural) |
| miRNA | micro RNAs | Post-transcriptional transposon repression |
| piRNA | Piwi-interacting RNA | DNA methylation, transposon repression |
| siRNA | Short interfering RNA | RNA interference |
| snoRNA | Small nucleolar RNAs | RNA modification, rRNA processing |
| PROMPT’s | Promoter upstream transcripts | Associated with chromatin changes |
| tiRNAs | Transcripton initation RNAs | Epigenetic regulation |
| lincRNAs | Long intergenic ncRNA | Epigenetic regulators of transcription |
| rasiRNA | Repeat associated small interfering RNA | Involved in the RNA interference (RNAi) pathway |
| eRNA | Enhancer-like ncRNA | Transcriptional gene activation |
| T-UCRs | Transcribed ultraconserved regions | Regulation of miRNA and mRNA levels |
| NATs | Natural antisense transcripts | mRNA stability |
| PALRs | Promoter-associated long RNAs | Chromatin changes |
| tasiRNA | Trans-acting siRNA | Represses gene expression |
| lncRNA | Long noncoding RNA | Regulation of gene transcription |
2. Non-Coding RNA Identification: The ENCODE Project
3. miRNAs
4. miRNAs Biogenesis
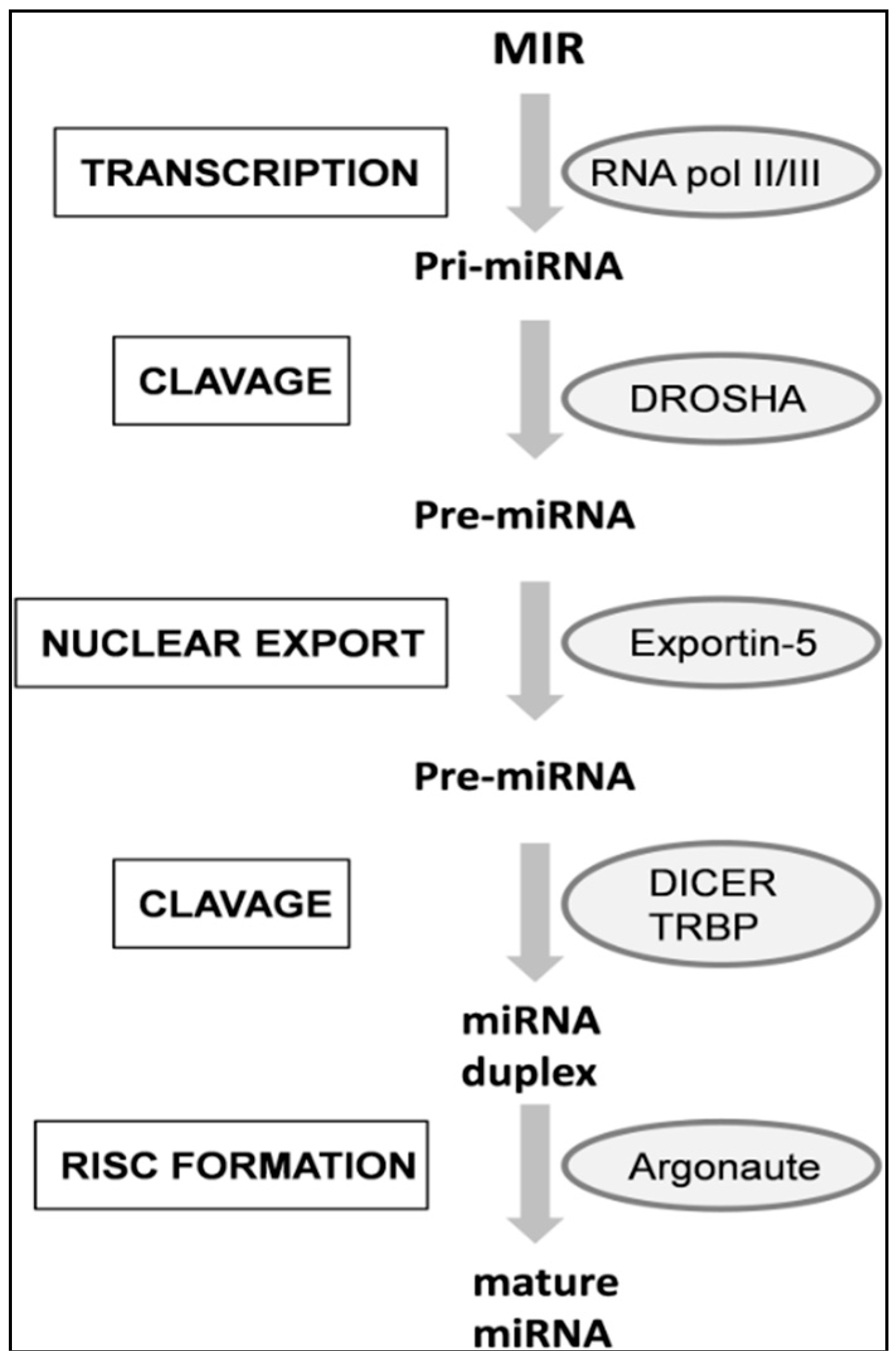
5. miRNAs Nomenclature
6. Studying miRNAs
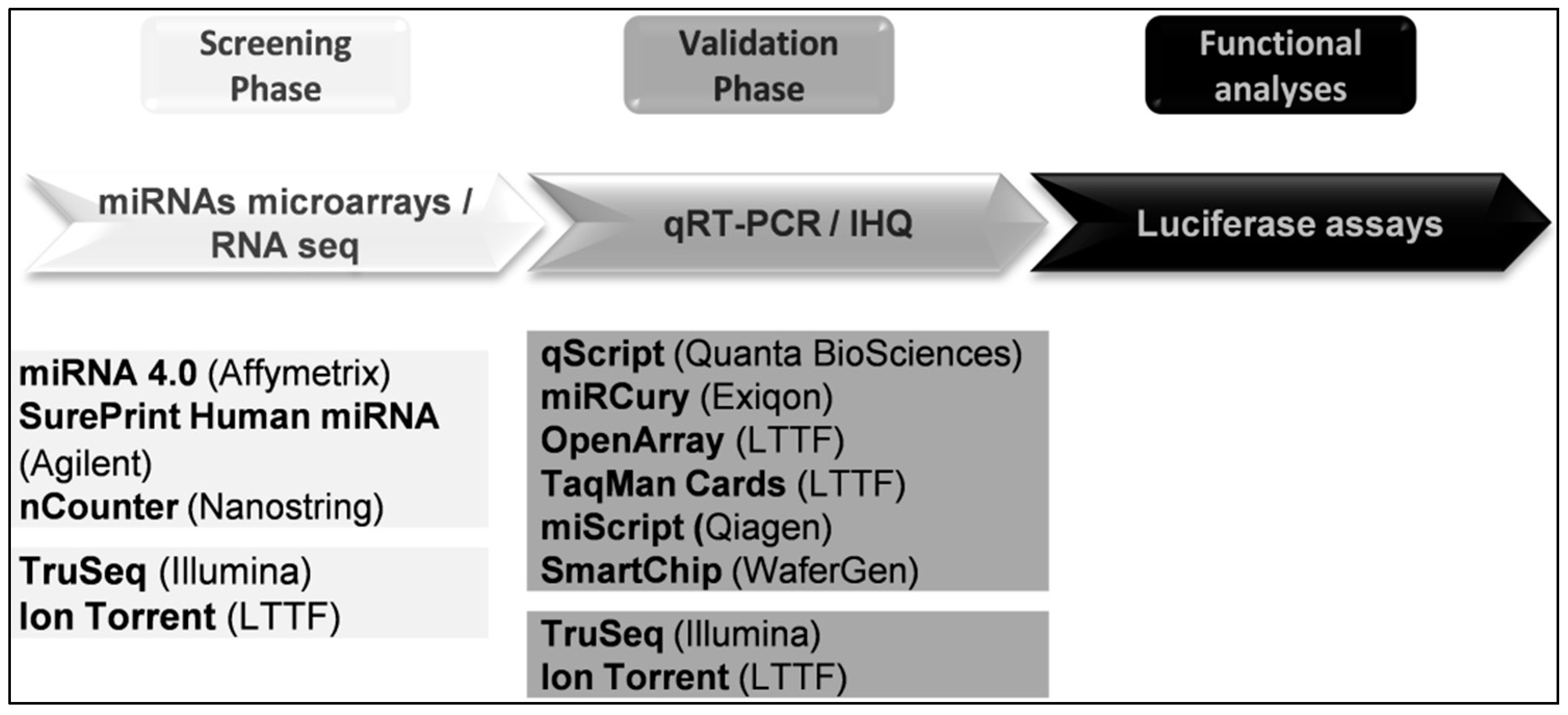
6.1. RNA Sequencing
6.2. miRNA Microarrays
6.3. Quantitative PCR or Real-Time PCR
6.4. In Situ Hybridization (ISH) and Live Cell miRNA Detection
7. miRNAs as Biomarkers
8. miRNAs Role in Endometriosis
9. Endometriosis
10. Studying New Biomarkers of Endometriosis
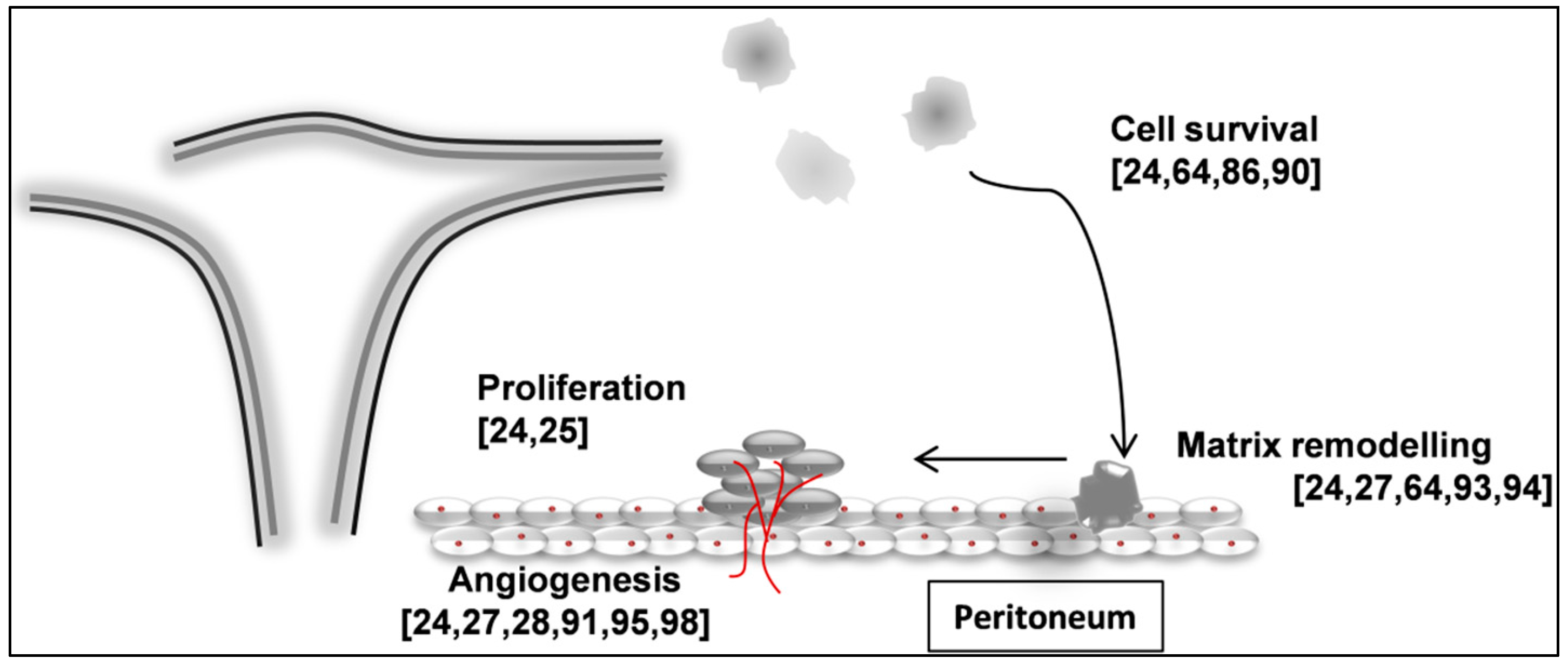
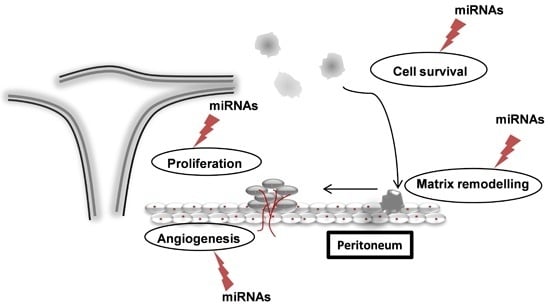
11. Circulating miRNAs as Biomarkers of Endometriosis
12. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Crick, F. Central dogma of molecular biology. Nature 1970, 227, 561–563. [Google Scholar] [CrossRef] [PubMed]
- Holley, R.W.; Apgar, J.; Everett, G.A.; Madison, J.T.; Marquisee, M.; Merrill, S.H.; Penswick, J.R.; Zamir, A. Structure of a ribonucleic acid. Science 1965, 147, 1462–1465. [Google Scholar] [CrossRef] [PubMed]
- Van Bakel, H.; Nislow, C.; Blencowe, B.J.; Hughes, T.R. Most “dark matter” transcripts are associated with known genes. PLoS Biol. 2010, 18, e1000371. [Google Scholar] [CrossRef] [PubMed]
- Phillips, T. Small non-coding RNA and gene expression. Nat. Educ. 2008, 1, 115. [Google Scholar]
- Desvignes, T.; Batze, P.; Berezikov, E.; Eilbeck, K.; Eppig, J.T.; McAndrews, M.S.; Singer, A.; Postlethwait, J.H. miRNA Nomenclature: A View Incorporating Genetic Origins, Biosynthetic Pathways, and Sequence Variants. Trends Genet. 2015, 31, 613–626. [Google Scholar] [CrossRef] [PubMed]
- The ENCODE Project Consortium. Identification and analysis of functional elements in 1% of the human genome by the ENCODE pilot project. Nature 2007, 447, 799–816. [Google Scholar]
- The ENCODE Project Consortium. An integrated encyclopedia of DNA elements in the human genome. Nature 2012, 489, 57–74. [Google Scholar]
- Nardone, J.; Lee, D.U.; Ansel, K.M.; Rao, A. Bioinformatics for the “bench biologist”: How to find regulatory regions in genomic DNA. Nat. Immunol. 2004, 5, 768–774. [Google Scholar] [CrossRef] [PubMed]
- Li, C.C.; Ramirez-Carrozzi, V.R.; Smale, S.T. Pursuing gene regulation “logic” via RNA interference and chromatin immunoprecipitation. Nat. Immunol. 2006, 7, 692–697. [Google Scholar] [CrossRef] [PubMed]
- Ozsolak, F.; Poling, L.L.; Wang, Z.; Liu, H.; Liu, X.S.; Roeder, R.G.; Zhang, X.; Song, J.S.; Fisher, D.E. Chromatin structure analyses identify miRNA promoters. Genes Dev. 2008, 15, 3172–3183. [Google Scholar] [CrossRef] [PubMed]
- Visel, A.; Rubin, E.M.; Pennacchio, L.A. Genomic views of distant-acting enhancers. Nature 2009, 10, 199–205. [Google Scholar] [CrossRef] [PubMed]
- Noonan, J.P.; McCallion, A.S. Genomics of long-range regulatory elements. Annu. Rev. Genom. Hum. Genet. 2010, 11, 1–23. [Google Scholar] [CrossRef] [PubMed]
- Christov, C.P.; Gardiner, T.J.; Szüts, D.; Krude, T. Functional requirement of noncoding Y RNAs for human chromosomal DNA replication. Mol. Cell. Biol. 2006, 26, 6993–7004. [Google Scholar] [CrossRef] [PubMed]
- Zhang, A.T.; Langley, A.R.; Christov, C.P.; Kheir, E.; Shafee, T.; Gardiner, T.J.; Krude, T. Dynamic interaction of Y RNAs with chromatin and initiation proteins during human DNA replication. J. Cell Sci. 2011, 15, 2058–2069. [Google Scholar] [CrossRef] [PubMed]
- Roberts, T.C. The MicroRNA biology of the mammalian nucleus. Mol. Ther. Nucleic Acids 2014, 19, e188. [Google Scholar] [CrossRef] [PubMed]
- Lee, R.C.; Feinbaum, R.L.; Ambros, V. The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 1993, 3, 843–854. [Google Scholar] [CrossRef]
- Bartel, D.P. MicroRNAs: Genomics, biogenesis, mechanism, and function. Cell 2004, 116, 281–297. [Google Scholar] [CrossRef]
- Eulalio, A.; Huntzinger, E.; Izaurralde, E. Getting to the root of miRNA-mediated gene silencing. Cell 2008, 132, 9–14. [Google Scholar] [CrossRef] [PubMed]
- Bartel, D.P. MicroRNAs: Target recognition and regulatory functions. Cell 2009, 23, 215–233. [Google Scholar] [CrossRef] [PubMed]
- Ambros, V.; Bartel, B.; Bartel, D.P.; Burge, C.B.; Carrington, J.C.; Chen, X.; Dreyfuss, G.; Eddy, S.R.; Griffiths-Jones, S.; Marshall, M.; et al. A uniform system for microRNA annotation. RNA 2003, 9, 277–279. [Google Scholar] [CrossRef] [PubMed]
- Burney, R.O.; Hamilton, A.E.; Aghajanova, L.; Vo, K.C.; Nezhat, C.N.; Lessey, B.A.; Giudice, L.C. MicroRNA expression profiling of eutopic secretory endometrium in women with versus without endometriosis. Mol. Hum. Reprod. 2009, 15, 625–631. [Google Scholar] [CrossRef] [PubMed]
- Ohlsson-Teague, E.M.; van der Hoek, K.H.; van der Hoek, M.B.; Perry, N.; Wagaarachchi, P.; Robertson, S.A.; Print, C.G.; Hull, L.M. MicroRNA-regulated pathways associated with endometriosis. Mol. Endocrinol. 2009, 23, 265–275. [Google Scholar] [CrossRef] [PubMed]
- Kuokkanen, S.; Chen, B.; Ojalvo, L.; Benard, L.; Santoro, N.; Pollard, J.W. Genomic profiling of microRNAs and messenger RNAs reveals hormonal regulation in microRNA expression in human endometrium. Biol. Reprod. 2010, 82, 791–801. [Google Scholar] [CrossRef] [PubMed]
- Ohlsson–Teague, E.M.; Print, C.G.; Hull, M.L. The role of microRNAs in endometriosis and associated reproductive conditions. Hum. Reprod. Update 2010, 16, 142–165. [Google Scholar] [CrossRef] [PubMed]
- Gilabert-Estelles, J.; Braza-Boils, A.; Ramon, L.A.; Zorio, E.; Medina, P.; España, F.; Estellés, A. Role of microRNAs in gynecological pathology. Curr. Med. Chem. 2012, 19, 2406–2413. [Google Scholar] [CrossRef] [PubMed]
- Ramón, L.A.; Braza-Boïls, A.; Gilabert, J.; Chirivella, M.; España, F.; Estellés, A.; Gilabert-Estellés, J. microRNAs related to angiogenesis are dysregulated in endometrioid endometrial cancer. Hum. Reprod. 2012, 27, 3036–3045. [Google Scholar] [CrossRef] [PubMed]
- Braza-Boïls, A.; Marí-Alexandre, J.; Gilabert, J.; Sánchez-Izquierdo, D.; España, F.; Estellés, A.; Gilabert-Estellés, J. MicroRNA expression profile in endometriosis: Its relation to angiogenesis and fibrinolytic factors. Hum. Reprod. 2014, 29, 978–988. [Google Scholar] [CrossRef] [PubMed]
- Marí-Alexandre, J.; García-Oms, J.; Barceló-Molina, M.; Gilabert-Aguilar, J.; Estellés, A.; Braza-Boïls, A.; Gilabert-Estellés, J. MicroRNAs and angiogenesis in endometriosis. Thromb. Res. 2015, 135, S38–S40. [Google Scholar] [CrossRef]
- Lund, E.; Güttinger, S.; Calado, A.; Dahlberg, J.E.; Kutay, U. Nuclear export of MicroRNA precursors. Science 2004, 303, 95–98. [Google Scholar] [CrossRef] [PubMed]
- Meijer, H.A.; Smith, E.M.; Bushell, M. Regulation of miRNA strand selection: Follow the leader? Biochem. Soc. Trans. 2014, 42, 1135–1140. [Google Scholar] [CrossRef] [PubMed]
- Machlin, E.S.; Sarnow, P.; Sagan, S.M. Masking the 5’ terminal nucleotides of the hepatitis C virus genome by an unconventional microRNA-target RNA complex. Proc. Natl. Acad. Sci. USA 2011, 22, 3193–3198. [Google Scholar] [CrossRef] [PubMed]
- Neilsen, C.T.; Goodall, G.J.; Bracken, C.P. IsomiRs: The overlooked repertoire in the dynamic microRNAome. Trends Genet. 2012, 28, 544–549. [Google Scholar] [CrossRef] [PubMed]
- Westholm, J.O.; Lai, E.C. Mirtrons: MicroRNA biogenesis via splicing. Biochimie 2011, 93, 1897–1904. [Google Scholar] [CrossRef] [PubMed]
- Rodriguez, A.; Griffiths-Jones, S.; Ashurst, J.L.; Bradley, A. Identification of mammalian microRNA host genes and transcription units. Genome Res. 2004, 14, 1902–1910. [Google Scholar] [CrossRef] [PubMed]
- Baskerville, S.; Bartel, D.P. Microarray profiling of microRNAs reveals frequent coexpression with neighboring miRNAs and host genes. RNA 2005, 11, 241–247. [Google Scholar] [CrossRef] [PubMed]
- Gregory, R.I.; Shiekhattar, R. MicroRNA Biogenesis and Cancer. Cancer Res. 2005, 65, 3509–3512. [Google Scholar] [CrossRef] [PubMed]
- Kozomara, A.; Griffiths-Jones, S. miRBase: Annotating high confidence microRNAs using deep sequencing data. A uniform system for microRNA annotation. Nucleic Acids Res. 2014, 42, 68–73. [Google Scholar] [CrossRef] [PubMed]
- Hacia, J.G.; Fan, J.B.; Ryder, O.; Jin, L.; Edgemon, K.; Ghandour, G.; Mayer, R.A.; Sun, B.; Hsie, L.; Robbins, C.M.; et al. Determination of ancestral alleles for human single-nucleotide polymorphisms using high-density oligonucleotide arrays. Nat. Genet. 1999, 22, 164–167. [Google Scholar] [PubMed]
- Zampetaki, A.; Mayr, M. Analytical challenges and technical limitations in assessing circulating miRNAs. Thromb. Haemost. 2012, 108, 592–598. [Google Scholar] [CrossRef] [PubMed]
- Urbanek, M.O.; Nawrocka, A.U.; Krzyzosiak, W.J. Small RNA Detection by in Situ Hybridization Methods. Int. J. Mol. Sci. 2015, 16, 13259–13286. [Google Scholar] [CrossRef] [PubMed]
- Franciscus, A. A Brief History of Hepatitis C. Hepat. C Support Proj. Fact Sheet 2015, 12, 1–8. [Google Scholar]
- Biomarkers Definitions Working Group. Biomarkers and surrogate endpoints: Preferred definitions and conceptual framework. Clin. Pharmacol. Ther. 2001, 69, 89–95. [Google Scholar]
- Mayr, M.; Zampetaki, A.; Willeit, P.; Willeit, J.; Kiechl, S. MicroRNAs within the continuum of postgenomics biomarker discovery. Arterioscler. Thromb. Vasc. Biol. 2013, 33, 206–214. [Google Scholar] [CrossRef] [PubMed]
- Zorio, E.; Medina, P.; Rueda, J.; Millán, J.M.; Arnau, M.A.; Beneyto, M.; Marín, F.; Gimeno, J.R.; Osca, J.; Salvador, A.; et al. Insights into the role of microRNAs in cardiac diseases: From biological signalling to therapeutic targets. Cardiovasc. Hematol. Agents Med. Chem. 2009, 7, 82–90. [Google Scholar] [CrossRef] [PubMed]
- Wittmann, J.; Jäck, H.M. Serum microRNAs as powerful cancer biomarkers. Biochim. Biophys. Acta 2010, 1806, 200–207. [Google Scholar] [CrossRef] [PubMed]
- Weber, J.A.; Baxter, D.H.; Zhang, S.; Huang, D.Y.; Huang, K.H.; Lee, M.J.; Galas, D.J.; Wang, K. The microRNA spectrum in 12 body fluids. Clin. Chem. 2010, 56, 1733–1741. [Google Scholar] [CrossRef] [PubMed]
- Silva, S.S.; Lopes, C.; Teixeira, A.L.; Carneiro de Sousa, M.J.; Medeiros, R. Forensic miRNA: Potential biomarker for body fluids? Forensic Sci. Int. Genet. 2015, 14, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Cortez, M.A.; Bueso-Ramos, C.; Ferdin, J.; Lopez-Berestein, G.; Sood, A.K.; Calin, G.A. MicroRNAs in body fluids: The mix of hormones and biomarkers. Nat. Rev. Clin. Oncol. 2011, 8, 467–477. [Google Scholar] [CrossRef] [PubMed]
- Shah, M.Y.; Calin, G.A. The Mix of Two Worlds: Non-Coding RNAs and Hormones. Nucleic Acid Ther. 2013, 23, 2–8. [Google Scholar] [CrossRef] [PubMed]
- Valadi, H.; Ekström, K.; Bossios, A.; Sjöstrand, M.; Lee, J.J.; Lötvall, J.O. Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nat. Cell Biol. 2007, 9, 654–659. [Google Scholar] [CrossRef] [PubMed]
- Hunter, M.P.; Ismail, N.; Zhang, X.; Aguda, B.D.; Lee, E.J.; Yu, L.; Xiao, T.; Schafer, J.; Ting Lee, M.L.; Schmittgen, T.D.; et al. Detection of microRNA Expression in Human Peripheral Blood Microvesicles. PLoS ONE 2008, 3, e3694. [Google Scholar] [CrossRef] [PubMed]
- Zernecke, A.; Bidzhekov, K.; Noels, H.; Shagdarsuren, E.; Gan, L.; Denecke, B.; Hristov, M.; Köppel, T.; Jahantigh, M.N.; Lutgens, E.; et al. Delivery of microRNA-126 by apoptotic bodies induces CXCL12-dependent vascular protection. Sci. Signal. 2009, 8, ra81. [Google Scholar] [CrossRef] [PubMed]
- Vickers, K.C.; Palmisano, B.T.; Shoucri, B.M.; Shamburek, R.D.; Remaley, A.T. MicroRNAs are transported in plasma and delivered to recipient cells by high-density lipoproteins. Nat. Cell Biol. 2011, 13, 423–433. [Google Scholar] [CrossRef] [PubMed]
- Arroyo, J.D.; Chevilleta, J.R.; Kroha, E.M.; Rufa, I.K.; Pritchardb, C.C.; Gibsonb, D.F.; Mitchella, P.S.; Bennetta, C.F.; Pogosova-Agadjanyand, E.L.; Stirewaltd, D.L.; et al. Argonaute2 complexes carry a population of circulating microRNAs independent of vesicles in human plasma. Prot. Nat. Acad. Sci. 2011, 108, 5003–5008. [Google Scholar] [CrossRef] [PubMed]
- Zampetaki, A.; Willeit, P.; Drozdov, I.; Kiechl, S.; Mayr, M. Profiling of circulating microRNAs: From single biomarkers to re-wired networks. Cardiovasc. Res. 2012, 3, 555–562. [Google Scholar] [CrossRef] [PubMed]
- Kosaka, N.; Yoshioka, Y.; Hagiwara, K.; Tominaga, N.; Katsuda, T.; Ochiya, T. Trash or Treasure: extracellular microRNAs and cell-to-cell communication. Front. Genet. 2013, 4, 173. [Google Scholar] [CrossRef] [PubMed]
- Turchinovich, A.; Samatov, T.R.; Tonevitsky, A.G.; Burwinkel, B. Circulating miRNAs: Cell-cell communication function? Front. Genet. 2013, 4, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Bang, C.; Batkai, S.; Dangwal, S.; Gupta, S.K.; Foinquinos, A.; Holzmann, A.; Just, A.; Remke, J.; Zimmer, K.; Zeug, A.; et al. Cardiac fibroblast-derived microRNA passenger strand-enriched exosomes mediate cardiomyocyte hypertrophy. J. Clin. Investig. 2014, 124, 2136–2146. [Google Scholar] [CrossRef] [PubMed]
- Fabbri, M. TLRs as miRNA Receptors. Cancer Res. 2012, 72, 6333–6337. [Google Scholar] [CrossRef] [PubMed]
- Fabbri, M.; Paone, A.; Calore, F.; Galli, R.; Gaudio, E.; Santhanam, R.; Lovat, F.; Fadda, P.; Mao, C.; Nuovo, G.J.; et al. MicroRNAs bind to Toll-like receptors to induce prometastatic inflammatory response. Proc. Natl. Acad. Sci. USA 2012, 109, E2110–E2116. [Google Scholar] [CrossRef] [PubMed]
- Mitchell, P.S.; Parkin, R.K.; Kroh, E.M.; Fritz, B.R.; Wyman, S.K.; Pogosova-Agadjanyan, E.L.; Peterson, A.; Noteboom, J.; O’Briant, K.C.; Allen, A.; et al. Circulating microRNAs as stable blood-based markers for cancer detection. Prot. Nat. Acad. Sci. 2008, 105, 10513–10518. [Google Scholar] [CrossRef] [PubMed]
- Hawkins, S.M.; Creighton, C.J.; Han, D.Y.; Zariff, A.; Anderson, M.L.; Gunaratne, P.H.; Matzuk, M.M. Functional microRNA involved in endometriosis. Mol. Endocrinol. 2011, 25, 821–832. [Google Scholar] [CrossRef] [PubMed]
- Nisolle, M.; Donnez, J. Peritoneal endometriosis, ovarian endometriosis, and adenomyotic nodules of the rectovaginal septum are three different entities. Fertil. Steril. 1997, 68, 585–596. [Google Scholar] [CrossRef]
- Giudice, L.C. Endometriosis. N. Engl. J. Med. 2010, 362, 2389–2398. [Google Scholar] [CrossRef] [PubMed]
- Berkley, K.J.; Rapkin, A.J.; Papka, R.E. The pains of endometriosis. Science 2005, 308, 1587–1589. [Google Scholar] [CrossRef] [PubMed]
- Stilley, J.A.; Birt, J.A.; Sharpe-Timms, K.L. Cellular and molecular basis for endometriosis-associated infertility. Cell Tissue Res. 2012, 349, 849–862. [Google Scholar] [CrossRef] [PubMed]
- Giudice, L.C.; Kao, L.C. Endometriosis. Lancet 2004, 364, 1789–1799. [Google Scholar] [CrossRef]
- Scully, R.E. Classification of human ovarian tumors. Environ. Health Perspect. 1987, 73, 15–25. [Google Scholar] [CrossRef] [PubMed]
- Guo, S.W. Endometriosis and ovarian cancer: Potential benefits and harms of screening and risk-reducing surgery. Fertil. Steril. 2015, 104, 813–830. [Google Scholar] [CrossRef] [PubMed]
- Sampson, J. Endometrial carcinoma of the ovary, arising in endometrial tissue in that organ. Arch. Surg. 1925, 10, 1–72. [Google Scholar] [CrossRef]
- Wei, J.J.; William, J.; Bulun, S. Endometriosis and ovarian cancer: A review of clinical, pathologic, and molecular aspects. Int. J. Gynecol. Pathol. 2011, 30, 553–568. [Google Scholar] [CrossRef] [PubMed]
- Munksgaard, P.S.; Blaakaer, J. The association between endometriosis and ovarian cancer: A review of histological, genetic and molecular alterations. Gynecol. Oncol. 2012, 124, 164–169. [Google Scholar] [CrossRef] [PubMed]
- Worley, M.J.; Welch, W.R.; Berkowitz, R.S.; Ng, S.W. Endometriosis-associated ovarian cancer: A review of pathogenesis. Int. J. Mol. Sci. 2013, 14, 5367–5379. [Google Scholar] [CrossRef] [PubMed]
- Siufi-Neto, J.; Kho, R.M.; Siufi, D.F.; Baracat, E.C.; Anderson, K.S.; Abrão, M.S. Cellular, histologic, and molecular changes associated with endometriosis and ovarian cancer. J. Minim. Invasive Gynecol. 2014, 21, 55–63. [Google Scholar] [CrossRef] [PubMed]
- Burney, R.O.; Giudice, L.C. Pathogenesis and pathophysiology of endometriosis. Fertil. Steril. 2012, 98, 511–519. [Google Scholar] [CrossRef] [PubMed]
- Dunselman, G.A.; Vermeulen, N.; Becker, C.; Calhaz-Jorge, C.; D'Hooghe, T.; De Bie, B.; Heikinheimo, O.; Horne, A.W.; Kiesel, L.; Nap, A.; et al. European Society of Human Reproduction and Embryology. ESHRE guideline: Management of women with endometriosis. Hum. Reprod. 2014, 29, 400–412. [Google Scholar] [CrossRef] [PubMed]
- Ballard, K.; Lowton, K.; Wright, J. What’s the delay? A qualitative study of women’s experiences of reaching a diagnosis of endometriosis. Fertil. Steril. 2006, 86, 1296–1301. [Google Scholar] [CrossRef] [PubMed]
- Fassbender, A.; Vodolazkaia, A.; Saunders, P.; Lebovic, D.; Waelkens, E.; De Moor, B.; D’Hooghe, T. Biomarkers of endometriosis. Fertil. Steril. 2013, 99, 1135–1145. [Google Scholar] [CrossRef] [PubMed]
- D’Hooghe, T.M.; Debrock, S. Endometriosis, retrograde menstruation and peritoneal inflammation in women and in baboons. Hum. Reprod. Update 2002, 8, 84–88. [Google Scholar] [CrossRef] [PubMed]
- Nnoaham, K.E.; Hummelshoj, L.; Kennedy, S.H.; Jenkinson, C.; Zondervan, K.T. Developing symptom-based predictive models of endometriosis as a clinical screening tool: Results from a multicenter study. Fertil. Steril. 2012, 98, 692–701. [Google Scholar] [CrossRef] [PubMed]
- Jia, S.Z.; Yang, Y.; Lang, J.; Sun, P.; Leng, J. Plasma miR-17–5p, miR-20a and miR-22 are down-regulated in women with endometriosis. Hum. Reprod. 2013, 28, 322–330. [Google Scholar] [CrossRef] [PubMed]
- Cho, S.; Mutlu, L.; Grechukhina, O.; Taylor, H.S. Circulating microRNAs as potential biomarkers for endometriosis. Fertil. Steril. 2015, 103, 1252–1260. [Google Scholar] [CrossRef] [PubMed]
- Rekker, K.; Saare, M.; Roost, A.M.; Kaart, T.; Sõritsa, D.; Karro, H.; Sõritsa, A.; Simón, C.; Salumets, A.; Peters, M. Circulating miR-200-family micro-RNAs have altered plasma levels in patients with endometriosis and vary with blood collection time. Fertil. Steril. 2015, 104, 938–946. [Google Scholar] [CrossRef] [PubMed]
- Rogers, P.A.; D'Hooghe, T.M.; Fazleabas, A.; Gargett, C.E.; Giudice, L.C.; Montgomery, G.W.; Rombauts, L.; Salamonsen, L.A.; Zondervan, K.T. Priorities for endometriosis research: Recommendations from an international consensus workshop. Reprod. Sci. 2009, 16, 335–346. [Google Scholar] [CrossRef] [PubMed]
- Halme, J.; Hammond, M.G.; Hulka, J.F.; Raj, S.G.; Talbert, L.M. Retrograde menstruation in healthy women and in patients with endometriosis. Obstet. Gynecol. 1984, 64, 151–154. [Google Scholar] [PubMed]
- Filigheddu, N.; Gregnanin, I.; Porporato, P.; Surico, D.; Perego, B.; Galli, L.; Patrignani, C.; Graziani, A.; Surico, N. Differential expression of micrornas between eutopic and ectopic endometrium in ovarian endometriosis. J. Biomed. Biotechnol. 2010, 2010, 369549. [Google Scholar] [CrossRef] [PubMed]
- Laudanski, P.; Charkiewicz, R.; Kuzmicki, M.; Szamatowicz, J.; Charkiewicz, A.; Niklinski, J. MicroRNAs expression profiling of eutopic proliferative endometrium in women with ovarian endometriosis. Reprod. Biol. Endocrinol. 2013, 11, 78. [Google Scholar] [CrossRef] [PubMed]
- Houshdaran, S.; Zelenko, Z.; Irwin, J.C.; Giudice, L.C. Human endometrial DNA methylome is cycle-dependent and is associated with gene expression regulation. Mol. Endocrinol. 2014, 28, 1118–1135. [Google Scholar] [CrossRef] [PubMed]
- Munro, S.K.; Farquhar, C.M.; Mitchell, M.D.; Ponnampalam, A.P. Epigenetic regulation of endometrium during the menstrual cycle. Mol. Hum. Reprod. 2010, 16, 297–310. [Google Scholar] [CrossRef] [PubMed]
- Harada, T.; Kaponis, A.; Iwabe, T.; Taniguchi, F.; Makrydimas, G.; Sofikitis, N.; Paschopoulos, M.; Paraskevaidis, E.; Terakawa, N. Apoptosis in human endometrium and endometriosis. Hum. Reprod. Update 2004, 10, 29–38. [Google Scholar] [CrossRef] [PubMed]
- Rocha, A.L.; Reis, F.M.; Taylor, R.N. Angiogenesis and endometriosis. Obstet. Gynecol. Int. 2013, 2013, 859619. [Google Scholar] [CrossRef] [PubMed]
- Laschke, M.W.; Menger, M.D. In vitro and in vivo approaches to study angiogenesis in the pathophysiology and therapy of endometriosis. Hum. Reprod. Update 2007, 13, 331–342. [Google Scholar] [CrossRef] [PubMed]
- Gilabert-Estelles, J.; Estelles, A.; Gilabert, J.; Castelló, R.; España, F.; Falcó, C.; Romeu, A.; Chirivella, M.; Zorio, E.; Aznar, J. Expression of several components of the plasminogen activator and matrix metalloproteinase systems in endometriosis. Hum. Reprod. 2003, 18, 1516–1522. [Google Scholar] [CrossRef] [PubMed]
- Gilabert-Estelles, J.; Ramon, L.A.; Espana, F.; Gilabert, J.; Vila, V.; Réganon, E.; Castelló, R.; Chirivella, M.; Estellés, A. Expression of angiogenic factors in endometriosis: Its relation to fibrinolytic and metalloproteinase (MMP) systems. Hum. Reprod. 2007, 22, 2120–2127. [Google Scholar] [CrossRef] [PubMed]
- Caporali, A.; Emanueli, C. MicroRNA regulation in angiogenesis. Vascul. Pharmacol. 2011, 55, 79–86. [Google Scholar] [CrossRef] [PubMed]
- Braza-Boils, A.; Salloum-Asfar, S.; Marí-Alexandre, J.; Arroyo, A.B.; González-Conejero, R.; Barceló-Molina, M.; García-Oms, J.; Vicente, V.; Estellés, A.; Gilabert-Estellés, J.; et al. Peritoneal fluid modifies the microRNA expression profile in endometrial and endometriotic cells from women with endometriosis. Hum. Reprod. 2015, 30, 2292–2302. [Google Scholar] [CrossRef] [PubMed]
- Olive, V.; Jiang, I.; He, L. mir-17–92, a cluster of miRNAs in the midst of the cancer network. Int. J. Biochem. Cell Biol. 2010, 42, 1348–1354. [Google Scholar] [CrossRef] [PubMed]
- Braza-Boïls, A.; Gilabert-Estellés, J.; Ramón, L.A.; Gilabert, J.; Marí-Alexandre, J.; Chirivella, M.; España, F.; Estellés, A. Peritoneal fluid reduces angiogenesis-related microRNA expression in cell cultures of endometrial and endometriotic tissues from women with endometriosis. PLoS ONE 2013, 8, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.T.; Zhao, Y.N.; Han, B.W.; Hong, S.J.; Chen, Y.Q. Circulating microRNAs identified in a genome-wide serum microRNA expression analysis as noninvasive biomarkers for endometriosis. J. Clin. Endocrinol. Metab. 2013, 98, 281–289. [Google Scholar] [CrossRef] [PubMed]
- Rekker, K.; Saare, M.; Roost, A.M.; Salumets, A.; Peters, M. Circulating microRNA profile throughout the menstrual cycle. PLoS ONE 2013, 8, 1–6. [Google Scholar] [CrossRef] [PubMed]
- Becker, C.M.; Laufer, M.R.; Stratton, P.; Hummelshoj, L.; Missmer, S.A.; Zondervan, K.T.; Adamson, G.D.; WERF EPHect Working Group. World Endometriosis Research Foundation Endometriosis Phenome and Biobanking Harmonisation Project: I. Surgical phenotype data collection in endometriosis research. Fertil. Steril. 2014, 102, 1213–1222. [Google Scholar] [CrossRef] [PubMed]
- Vitonis, A.F.; Vincent, K.; Rahmioglu, N.; Fassbender, A.; Buck Louis, G.M.; Hummelshoj, L.; Giudice, L.C.; Stratton, P.; Adamson, G.D.; Becker, C.M.; et al. World Endometriosis Research Foundation Endometriosis Phenome and Biobanking Harmonization Project: II. Clinical and covariate phenotype data collection in endometriosis research. Fertil. Steril. 2014, 102, 1223–1232. [Google Scholar] [CrossRef] [PubMed]
- Rahmioglu, N.; Fassbender, A.; Vitonis, A.F.; Tworoger, S.S.; Hummelshoj, L.; D'Hooghe, T.M.; Adamson, G.D.; Giudice, L.C.; Becker, C.M.; Zondervan, K.T.; et al. World Endometriosis Research Foundation Endometriosis Phenome and Biobanking Harmonization Project: III. Fluid biospecimen collection, processing, and storage in endometriosis research. Fertil. Steril. 2014, 102, 1233–1243. [Google Scholar] [CrossRef] [PubMed]
- Fassbender, A.; Rahmioglu, N.; Vitonis, A.F.; Viganò, P.; Giudice, L.C.; D’Hooghe, T.M.; Hummelshoj, L.; Adamson, G.D.; Becker, C.M.; Missmer, S.A.; et al. World Endometriosis Research Foundation Endometriosis Phenome and Biobanking Harmonisation Project: IV. Tissue collection, processing, and storage in endometriosis research. Fertil. Steril. 2014, 102, 1244–1253. [Google Scholar] [CrossRef] [PubMed]
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons by Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Marí-Alexandre, J.; Sánchez-Izquierdo, D.; Gilabert-Estellés, J.; Barceló-Molina, M.; Braza-Boïls, A.; Sandoval, J. miRNAs Regulation and Its Role as Biomarkers in Endometriosis. Int. J. Mol. Sci. 2016, 17, 93. https://doi.org/10.3390/ijms17010093
Marí-Alexandre J, Sánchez-Izquierdo D, Gilabert-Estellés J, Barceló-Molina M, Braza-Boïls A, Sandoval J. miRNAs Regulation and Its Role as Biomarkers in Endometriosis. International Journal of Molecular Sciences. 2016; 17(1):93. https://doi.org/10.3390/ijms17010093
Chicago/Turabian StyleMarí-Alexandre, Josep, Dolors Sánchez-Izquierdo, Juan Gilabert-Estellés, Moisés Barceló-Molina, Aitana Braza-Boïls, and Juan Sandoval. 2016. "miRNAs Regulation and Its Role as Biomarkers in Endometriosis" International Journal of Molecular Sciences 17, no. 1: 93. https://doi.org/10.3390/ijms17010093
APA StyleMarí-Alexandre, J., Sánchez-Izquierdo, D., Gilabert-Estellés, J., Barceló-Molina, M., Braza-Boïls, A., & Sandoval, J. (2016). miRNAs Regulation and Its Role as Biomarkers in Endometriosis. International Journal of Molecular Sciences, 17(1), 93. https://doi.org/10.3390/ijms17010093