Molecular Identification of Dendrobium Species (Orchidaceae) Based on the DNA Barcode ITS2 Region and Its Application for Phylogenetic Study
Abstract
:1. Introduction
2. Results
2.1. PCR Amplification, Success Rate and Sequence Characteristics
2.2. Genetic Divergence within and between Species
| Measurement | Kimura 2-Parameter (K2P) Value |
|---|---|
| All inter-specific distance | 0.182 ± 0.036 |
| Theta prime | 0.192 ± 0.036 |
| The minimum inter-specific distance | 0.185 ± 0.035 |
| All intra-specific distance | 0.007 ± 0.004 |
| Theta | 0.006 ± 0.003 |
| Coalescent depth | 0.014 ± 0.006 |
2.3. Assessment of Barcoding Gap
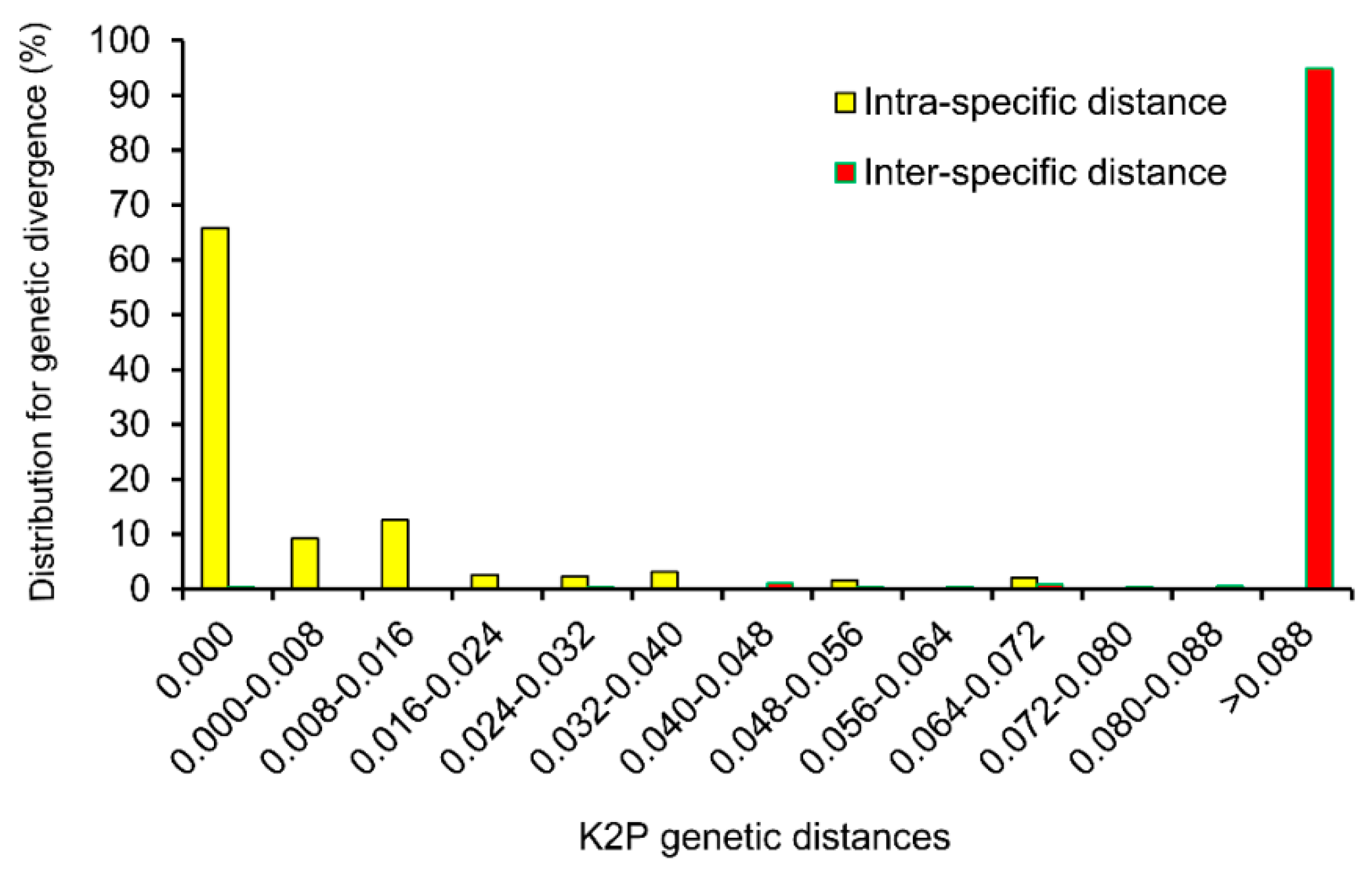
2.4. Efficacy of ITS2 for Authentication
| Methods of Identification | No. of Samples | No. of Species | Correct Identification (%) | Incorrect Identification (%) | Ambiguous Identification (%) |
|---|---|---|---|---|---|
| BLAST1 | 364 | 64 | 85.9 | 0 | 14.1 |
| Distance | 364 | 64 | 82.8 | 0 | 17.2 |
2.5. Evaluation of the Discriminatory Power of ITS2 Sequences
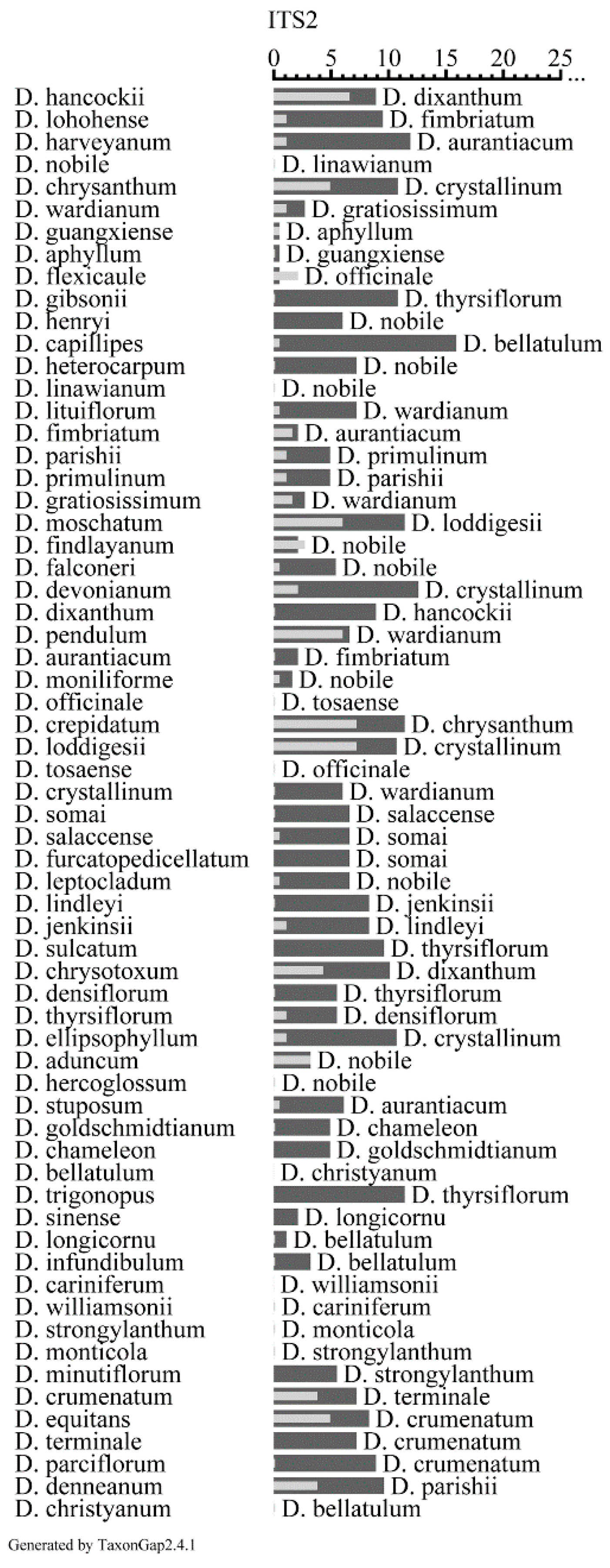

2.6. Phylogenetic Analysis
| Species Name | Voucher Number (No.) | Section | Collection Site | GenBank Accession No. |
|---|---|---|---|---|
| D. hancockii Rolfe | D-HZ001 | Dendrobium | Yunnan | KJ658296 |
| D. harveyanum Rchb. f. | D-HZ002 | Dendrobium | Yunnan | KJ658297 |
| D. nobile Lindl. | D-HZ003 | Dendrobium | Yunnan | KJ658298 |
| D. chrysanthum Wall. ex Lindl. | D-HZ004 | Dendrobium | Guangxi | KJ658299 |
| D. wardianum Warner | D-HZ005 | Dendrobium | Yunnan | KJ658300 |
| D. guangxiense S. J. Cheng et C. Z. Tang | D-HZ006 | Dendrobium | Yunnan | KJ658301 |
| D. guangxiense S. J. Cheng et C. Z. Tang | D-HZ007 | Dendrobium | Guangxi | KJ658302 |
| D. aphyllum (Rohb.) C. E. Fishcher | D-HZ008 | Dendrobium | Guangxi | KJ658303 |
| D. gibsonii Lindl. | D-HZ009 | Dendrobium | Yunnan | KJ658304 |
| D. gibsonii Lindl. | D-HZ010 | Dendrobium | Guangdong | KJ658305 |
| D. capillipes Rchb. | D-HZ011 | Dendrobium | Guangdong | KJ658306 |
| D. heterocarpum Lindl. | D-HZ012 | Dendrobium | Yunnan | KJ658307 |
| D. fimbriatum Hook. | D-HZ013 | Dendrobium | Guangxi | KJ658308 |
| D. primulinum Lindl. | D-HZ014 | Dendrobium | Yunnan | KJ658309 |
| D. falconeri Hook. | D-HZ015 | Dendrobium | Guangdong | KJ658310 |
| D. devonianum Paxt. | D-HZ016 | Dendrobium | Yunnan | KJ658311 |
| D. dixanthum Rchb. | D-HZ017 | Dendrobium | Zhejiang | KJ658312 |
| D. dixanthum Rchb. | D-HZ018 | Dendrobium | Yunnan | KJ658313 |
| D. pendulum Roxb. | D-HZ019 | Dendrobium | Yunnan | KJ658314 |
| D. moniliforme (Linn.) Sw. | D-HZ020 | Dendrobium | Yunnan | KJ658315 |
| D. officinale Kimura et Migo | D-HZ021 | Dendrobium | Yunnan | KJ658316 |
| D. officinale Kimura et Migo | D-HZ022 | Dendrobium | Guangxi | KJ658317 |
| D. officinale Kimura et Migo | D-HZ023 | Dendrobium | Zhejiang | KJ658318 |
| D. crepidatum Lindl. ex Paxt. | D-HZ024 | Dendrobium | Yunnan | KJ658319 |
| D. loddigesii Rolfe | D-HZ025 | Dendrobium | Yunnan | KJ658320 |
| D. crystallinum Tchb. F. | D-HZ026 | Dendrobium | Guangxi | KJ658321 |
| D. denneanum Kerr. | D-HZ041 | Dendrobium | Guangdong | KJ658336 |
| D. lindleyi Steudel | D-HZ027 | Chrysotoxae | Guangdong | KJ658322 |
| D. lindleyi Steudel | D-HZ028 | Chrysotoxae | Yunnan | KJ658323 |
| D. chrysotoxum Lindl. | D-HZ029 | Chrysotoxae | Yunnan | KJ658324 |
| D. densiflorum Lindl. | D-HZ030 | Chrysotoxae | Guangdong | KJ658325 |
| D. densiflorum Lindl. | D-HZ031 | Chrysotoxae | Guangxi | KJ658326 |
| D. thyrsiflorum Rchb. f. ex André | D-HZ032 | Chrysotoxae | Yunnan | KJ658327 |
| D. aduncum Wall. ex Lindl. | D-HZ033 | Breviflores | Gangdong | KJ658328 |
| D.hercoglossum Rehb. f. | D-HZ034 | Breviflores | Guizhou | KJ658329 |
| D. stuposum Lindl. | D-HZ035 | Stuposa | Yunnan | KJ658330 |
| D. stuposum Lindl. | D-HZ036 | Stuposa | Yunnan | KJ658331 |
| D. longicornu Lindl. | D-HZ037 | Formosae | Guangxi | KJ658332 |
| D. williamsonii Day et Rchb. F. | D-HZ038 | Formosae | Guangxi | KJ658333 |
| D. williamsonii Day et Rchb. F. | D-HZ039 | Formosae | Yunnan | KJ658334 |
| D. christyanum Rchb. F. | D-HZ042 | Formosae | Yunnan | KJ658337 |
| D. christyanum Rchb. F. | D-HZ043 | Formosae | Yunnan | KJ658338 |
| D. strongylanthum Rchb. F. | D-HZ040 | Stachyobium | Yunnan | KJ658335 |
3. Discussion
4. Materials and Methods
4.1. Plant Materials
4.2. DNA Extraction, Amplification, and Sequencing
4.3. Data Analysis
5. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Wang, H.Z.; Feng, S.G.; Lu, J.J.; Shi, N.N.; Liu, J.J. Phylogenetic study and molecular identification of 31 Dendrobium species using inter-simple sequence repeat (ISSR) markers. Sci. Hortic. 2009, 122, 440–447. [Google Scholar] [CrossRef]
- Feng, S.G.; Lu, J.J.; Gao, L.; Liu, J.J.; Wang, H.-Z. Molecular phylogeny analysis and species identification of Dendrobium (Orchidaceae) in China. Biochem. Genet. 2014, 52, 127–136. [Google Scholar] [CrossRef] [PubMed]
- Tsi, Z.H.; Chen, S.C.; Luo, Y.B.; Zhu, G.H. Orchidaceae (3). In Flora Republicae Popularis Sinicae; Science Press: Beijing, China, 1999. [Google Scholar]
- Adams, P.B. Systematics of Dendrobiinae (Orchidaceae), with special reference to Australian taxa. Bot. J. Linn. Soc. 2011, 166, 105–126. [Google Scholar] [CrossRef]
- Xiang, X.G.; Schuiteman, A.; Li, D.Z.; Huang, W.C.; Chung, S.W.; Li, J.W.; Zhou, H.L.; Jin, W.T.; Lai, Y.J.; Li, Z.Y.; et al. Molecular systematics of Dendrobium (Orchidaceae, Dendrobieae) from mainland Asia based on plastid and nuclear sequences. Mol. Phylogenet. Evol. 2013, 69, 950–960. [Google Scholar] [CrossRef] [PubMed]
- Chinese Pharmmacopoeia Editorial Committee (CPEC). Pharmmacopoeia of the People’s Republic of China. In Theoretical and Applied Genetics; CPEC: Beijing, China, 2010. [Google Scholar]
- Bulpitt, C.J.; Li, Y.; Bulpitt, P.F. The use of orchids in Chinese medicine. Genome Natl. Res. Counc. Can. 2007, 100, 558–563. [Google Scholar] [CrossRef] [PubMed]
- Jin, W.; Yao, S. Cultivation and Appreciation of Noble Spring Orchid Cultivars; Guangdong Science and Technology Press: Guangzhou, China, 2006. [Google Scholar]
- Ding, G.; Li, X.; Ding, X.; Qian, L. Genetic diversity across natural populations of Dendrobium officinale, the endangered medicinal herb endemic to China, revealed by ISSR and RAPD markers. Genetika 2009, 45, 375–382. [Google Scholar] [CrossRef] [PubMed]
- Lau, D.T.; Shaw, P.C.; Wang, J.; But, P.P. Authentication of medicinal Dendrobium species by the internal transcribed spacer of ribosomal DNA. Planta Med. 2001, 67, 456–460. [Google Scholar] [PubMed]
- Feng, S.; He, R.; Yang, S.; Chen, Z.; Jiang, M.; Lu, J.; Wang, H. Start codon targeted (SCoT) and target region amplification polymorphism (TRAP) for evaluating the genetic relationship of Dendrobium species. Gene 2015, 567, 182–188. [Google Scholar] [CrossRef] [PubMed]
- Hebert, P.D.N.; Penton, E.H.; Burns, J.M.; Janzen, D.H.; Hallwachs, W. Ten species in one: DNA barcoding reveals cryptic species in the neotropical skipper butterfly Astraptes fulgerator. Proc. Natl. Acad. Sci. USA 2004, 101, 14812–14817. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.; Yao, H.; Han, J.; Liu, C.; Song, J.; Shi, L.; Zhu, Y.; Ma, X.; Gao, T.; Pang, X.; et al. Validation of the ITS2 region as a novel DNA barcode for identifying medicinal plant species. PLoS ONE 2010, 5, e8613. [Google Scholar] [CrossRef] [PubMed]
- Gao, T.; Yao, H.; Song, J.; Liu, C.; Zhu, Y.; Ma, X.; Pang, X.; Xu, H.; Chen, S. Identification of medicinal plants in the family Fabaceae using a potential DNA barcode ITS2. J. Ethnopharmacol. 2010, 130, 116–121. [Google Scholar] [CrossRef] [PubMed]
- Hajiahmadi, Z.; Talebi, M.; Sayed-Tabatabaei, B.E. Studying genetic variability of Pomegranate (Punica granatum L.) based on chloroplast DNA and barcode genes. Mol. Biotechnol. 2013, 55, 249–259. [Google Scholar] [CrossRef] [PubMed]
- Giudicelli, G.C.; Mader, G.; Brandao de Freitas, L. Efficiency of ITS sequences for DNA barcoding in Passiflora (Passifloraceae). Int. J. Mol. Sci. 2015, 16, 7289–7303. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Provan, J.; Gao, L.M.; Li, D.Z. Sampling strategy and potential utility of indels for DNA barcoding of closely related plant species: A case study in taxus. Int. J. Mol. Sci. 2012, 13, 8740–8751. [Google Scholar] [CrossRef] [PubMed]
- Sun, Z.Y.; Chen, S.L. Identification of cortex herbs using the DNA barcode nrITS2. J. Nat. Med. Tokyo 2013, 67, 296–302. [Google Scholar] [CrossRef] [PubMed]
- Gismondi, A.; Rolfo, M.F.; Leonardi, D.; Rickards, O.; Canini, A. Identification of ancient Olea europaea L. and Cornus mas L. seeds by DNA barcoding. Comptes Rendus Biol. 2012, 335, 472–479. [Google Scholar] [CrossRef] [PubMed]
- Gismondi, A.; Leonardi, D.; Enei, F.; Canini, A. Identification of plant remains in underwater archaeological areas by morphological analysis and DNA barcoding. Adv. Anthropol. 2013, 3, 240–248. [Google Scholar] [CrossRef]
- Gismondi, A.; Fanali, F.; Labarga, J.; Caiola, M.; Canini, A. Crocus sativus L. genomics and different DNA barcode applications. Plant Syst. Evol. 2013, 299, 1859–1863. [Google Scholar] [CrossRef]
- Liu, Z.; Zeng, X.; Yang, D.; Ren, G.; Chu, G.; Yuan, Z.; Luo, K.; Xiao, P.; Chen, S. Identification of medicinal vines by ITS2 using complementary discrimination methods. J. Ethnopharmacol. 2012, 141, 242–249. [Google Scholar] [CrossRef] [PubMed]
- Kress, W.J.; Erickson, D.L. A two-locus global DNA barcode for land plants: The coding rbcL gene complements the non-coding trnH-psbA spacer region. PLoS ONE 2007, 2, e508. [Google Scholar] [CrossRef] [PubMed]
- Lahaye, R.; van der Bank, M.; Bogarin, D.; Warner, J.; Pupulin, F.; Gigot, G.; Maurin, O.; Duthoit, S.; Barraclough, T.G.; Savolainen, V. DNA barcoding the floras of biodiversity hotspots. Proc. Natl. Acad. Sci. USA 2008, 105, 2923–2928. [Google Scholar] [CrossRef] [PubMed]
- Hadjiolova, K.V.; Nicoloso, M.; Mazan, S.; Hadjiolov, A.A.; Bachellerie, J.P. Alternative pre-rRNA processing pathways in human cells and their alteration by cycloheximide inhibition of protein synthesis. Eur. J. Biochem. 1993, 212, 211–215. [Google Scholar] [CrossRef] [PubMed]
- Van Nues, R.W.; Rientjes, J.M.J.; Morré, S.A.; Mollee, E.; Planta, R.J.; Venema, J.; Raué, H.A. Evolutionarily conserved structural elements are critical for processing of Internal Transcribed Spacer 2 from Saccharomyces cerevisie Precursor Ribosomal RNA. J. Mol. Biol. 1995, 250, 24–36. [Google Scholar] [CrossRef] [PubMed]
- Jorgensen, A.; Stothard, J.R.; Madsen, H.; Nalugwa, A.; Nyakaana, S.; Rollinson, D. The ITS2 of the genus Bulinus: Novel secondary structure among freshwater snails and potential new taxonomic markers. Acta Trop. 2012, 128, 218–225. [Google Scholar] [CrossRef] [PubMed]
- Pang, X.; Song, J.; Zhu, Y.; Xu, H.; Huang, L.; Chen, S. Applying plant DNA barcodes for Rosaceae species identification. Cladistics 2011, 27, 165–170. [Google Scholar] [CrossRef]
- Yao, H.; Song, J.; Liu, C.; Luo, K.; Han, J.; Li, Y.; Pang, X.; Xu, H.; Zhu, Y.; Xiao, P.; et al. Use of ITS2 region as the universal DNA barcode for plants and animals. PLoS ONE 2010, 5, e13102. [Google Scholar] [CrossRef] [PubMed]
- Gao, T.; Yao, H.; Song, J.; Zhu, Y.; Liu, C.; Chen, S. Evaluating the feasibility of using candidate DNA barcodes in discriminating species of the large Asteraceae family. BMC Evol. Biol. 2010, 10, 324. [Google Scholar] [PubMed]
- Pang, X.; Song, J.; Zhu, Y.; Xie, C.; Chen, S. Using DNA barcoding to identify species within Euphorbiaceae. Planta Med. 2010, 76, 1784–1786. [Google Scholar] [CrossRef] [PubMed]
- Gu, W.; Song, J.; Cao, Y.; Sun, Q.; Yao, H.; Wu, Q.; Chao, J.; Zhou, J.; Xue, W.; Duan, J. Application of the ITS2 region for barcoding medicinal plants of Selaginellaceae in Pteridophyta. PLoS ONE 2013, 8, e67818. [Google Scholar] [CrossRef] [PubMed]
- Coleman, A.W. ITS2 is a double-edged tool for eukaryote evolutionary comparisons. Trends Genet. 2003, 19, 370–375. [Google Scholar] [CrossRef]
- Schultz, J.; Wolf, M. ITS2 sequence-structure analysis in phylogenetics: A how-to manual for molecular systematics. Mol. Phylogenet. Evol. 2009, 52, 520–523. [Google Scholar] [CrossRef] [PubMed]
- Chiou, S.J.; Yen, J.H.; Fang, C.L.; Chen, H.L.; Lin, T.Y. Authentication of medicinal herbs using PCR-amplified ITS2 with specific primers. Planta Med. 2007, 73, 1421–1426. [Google Scholar] [CrossRef] [PubMed]
- Schlechter, R. The Orchidaceae of German New Guinea; Blaxell, D.F., Translator; The Australian Orchid Foundation: Melbourne, Australia, 1982. [Google Scholar]
- Cowan, R.S.; Chase, M.W.; Kress, W.J.; Savolainen, V. 300,000 species to identify: Problems, progress, and prospects in DNA barcoding of land plants. Taxon 2006, 55, 611–616. [Google Scholar] [CrossRef]
- Li, H.Q.; Chen, J.Y.; Wang, S.; Xiong, S.Z. Evaluation of six candidate DNA barcoding loci in Ficus (Moraceae) of China. Mol. Ecol. Resour. 2012, 12, 783–790. [Google Scholar] [CrossRef] [PubMed]
- Hebert, P.D.; Cywinska, A.; Ball, S.L.; deWaard, J.R. Biological identifications through DNA barcodes. Proc. Biol. Sci. 2003, 270, 313–321. [Google Scholar] [CrossRef] [PubMed]
- Yukawa, T.; Uehara, K. Vegetative diversification and radiation in subtribe Dendrobiinae (Orchidaceae): Evidence from chloroplast DNA phylogeny and anatomical characters. Plant Syst. Evol. 1996, 201, 1–14. [Google Scholar] [CrossRef]
- Morris, M.W.; Steen, W.L.; Judd, W.S. Vegetative anatomy and systematics of subtribe Dendrobiinae (Orchidaceae). Bot. J. Linn. Soc. 1996, 120, 89–144. [Google Scholar] [CrossRef]
- Burke, J.M.; Bayly, M.J.; Adams, P.B.; Ladiges, P.Y. Molecular phylogenetic analysis of Dendrobium (Orchidaceae), with emphasis on the Australian section Dendrocoryne, and implications for generic classification. Aust. Syst. Bot. 2008, 21, 1–14. [Google Scholar] [CrossRef]
- Schuiteman, A. Dendrobium (Orchidaceae): To split or not split. Gard. Bull. Singap. 2011, 63, 245–257. [Google Scholar]
- Clements, M.A. Molecular phylogenetic systematics in Dendrobieae (Orchidaceae). Aliso 2006, 22, 465–480. [Google Scholar]
- Queiroz Cde, S.; Batista, F.R.; de Oliveira, L.O. Evolution of the 5.8S nrDNA gene and internal transcribed spacers in Carapichea ipecacuanha (Rubiaceae) within a phylogeographic context. Mol. Phylogenet. Evol. 2011, 59, 293–302. [Google Scholar] [CrossRef] [PubMed]
- Nieto Feliner, G.; Rossello, J.A. Better the devil you know? Guidelines for insightful utilization of nrDNA ITS in species-level evolutionary studies in plants. Mol. Phylogenet. Evol. 2007, 44, 911–919. [Google Scholar] [CrossRef] [PubMed]
- Feng, S.; Zhao, H.; Lu, J.; Liu, J.; Shen, B.; Wang, H. Preliminary genetic linkage maps of Chinese herb Dendrobium nobile and D. moniliforme. J. Genet. 2013, 92, 205–212. [Google Scholar] [CrossRef] [PubMed]
- Keller, A.; Schleicher, T.; Schultz, J.; Muller, T.; Dandekar, T.; Wolf, M. 5.8S–28S rRNA interaction and HMM-based ITS2 annotation. Gene 2009, 430, 50–57. [Google Scholar] [CrossRef] [PubMed]
- Koetschan, C.; Hackl, T.; Muller, T.; Wolf, M.; Forster, F.; Schultz, J. ITS2 Database IV: Interactive taxon sampling for internal transcribed spacer 2 based phylogenies. Mol. Phylogenet. Evol. 2012, 63, 585–588. [Google Scholar] [CrossRef] [PubMed]
- Thompson, J.D.; Gibson, T.; Higgins, D.G. Multiple sequence alignment using ClustalW and ClustalX. Curr. Protoc. Bioinform. 2002. [Google Scholar] [CrossRef]
- Tamura, K.; Peterson, D.; Peterson, N.; Stecher, G.; Nei, M.; Kumar, S. MEGA5: Molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol. 2011, 28, 2731–2739. [Google Scholar] [CrossRef] [PubMed]
- Meier, R.; Zhang, G.; Ali, F. The use of mean instead of smallest interspecific distances exaggerates the size of the “barcoding gap” and leads to misidentification. Syst. Biol. 2008, 57, 809–813. [Google Scholar] [CrossRef] [PubMed]
- Meyer, C.P.; Paulay, G. DNA barcoding: Error rates based on comprehensive sampling. PLoS Biol. 2005, 3, e422. [Google Scholar] [CrossRef] [PubMed]
- Ross, H.A.; Murugan, S.; Li, W.L. Testing the reliability of genetic methods of species identification via simulation. Syst. Biol. 2008, 57, 216–230. [Google Scholar] [CrossRef] [PubMed]
- Slabbinck, B.; Dawyndt, P.; Martens, M.; de Vos, P.; de Baets, B. TaxonGap: A visualization tool for intra-and inter-species variation among individual biomarkers. Bioinformatics 2008, 24, 866–867. [Google Scholar] [CrossRef]
- Ding, X.Y.; Wang, Z.T.; Xu, H.; Xu, L.S.; Zhou, K.Y. Database establishment of the whole rDNA ITS region of Dendrobium species of “fengdou” and authentication by analysis of their sequences. Yao Xue Xue Bao 2002, 37, 567–573. (In Chinese) [Google Scholar] [PubMed]
- Yang, L.; Wang, Z.; Xu, L. Simultaneous determination of phenols (bibenzyl, phenanthrene, and fluorenone) in Dendrobium species by high-performance liquid chromatography with diode array detection. J. Chromatogr. A 2006, 1104, 230–237. [Google Scholar] [CrossRef] [PubMed]
- Yao, H.; Song, J.Y.; Ma, X.Y.; Liu, C.; Li, Y.; Xu, H.X.; Han, J.P.; Duan, L.S.; Chen, S.L. Identification of Dendrobium species by a candidate DNA barcode sequence: The chloroplast psbA-trnH intergenic region. Planta Med. 2009, 75, 667–669. [Google Scholar] [CrossRef] [PubMed]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Feng, S.; Jiang, Y.; Wang, S.; Jiang, M.; Chen, Z.; Ying, Q.; Wang, H. Molecular Identification of Dendrobium Species (Orchidaceae) Based on the DNA Barcode ITS2 Region and Its Application for Phylogenetic Study. Int. J. Mol. Sci. 2015, 16, 21975-21988. https://doi.org/10.3390/ijms160921975
Feng S, Jiang Y, Wang S, Jiang M, Chen Z, Ying Q, Wang H. Molecular Identification of Dendrobium Species (Orchidaceae) Based on the DNA Barcode ITS2 Region and Its Application for Phylogenetic Study. International Journal of Molecular Sciences. 2015; 16(9):21975-21988. https://doi.org/10.3390/ijms160921975
Chicago/Turabian StyleFeng, Shangguo, Yan Jiang, Shang Wang, Mengying Jiang, Zhe Chen, Qicai Ying, and Huizhong Wang. 2015. "Molecular Identification of Dendrobium Species (Orchidaceae) Based on the DNA Barcode ITS2 Region and Its Application for Phylogenetic Study" International Journal of Molecular Sciences 16, no. 9: 21975-21988. https://doi.org/10.3390/ijms160921975
APA StyleFeng, S., Jiang, Y., Wang, S., Jiang, M., Chen, Z., Ying, Q., & Wang, H. (2015). Molecular Identification of Dendrobium Species (Orchidaceae) Based on the DNA Barcode ITS2 Region and Its Application for Phylogenetic Study. International Journal of Molecular Sciences, 16(9), 21975-21988. https://doi.org/10.3390/ijms160921975





