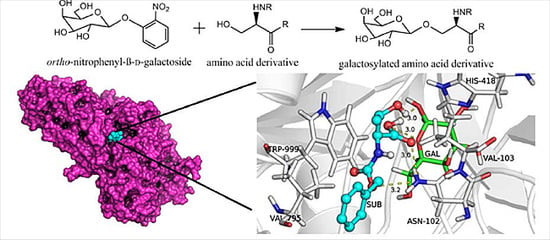
Enzymatic Synthesis of Galactosylated Serine/Threonine Derivatives by β-Galactosidase from Escherichia coli
Abstract
:1. Introduction

2. Results and Discussion
2.1. Effect of Selected Reaction Media on the Transgalactosylation of Serine Derivative
| Ratio of Sugar to Serine | Sugar | Reaction Time (h) | Bioconversion Yield (%) and Initial Rate of the Reaction (μmol/L.h) a | ||
|---|---|---|---|---|---|
| Buffer | Heptane:Buffer (70:30) | Heptanone:Buffer (70:30) | |||
| 1:1 | ONPG | 5 | <0.1 | 2.0 | 0.1 |
| 12 | 0.3 (0.01) b | 0.4 | <0.1 | ||
| 24 | <0.1 | <0.1 | <0.1 | ||
| Lactose | 5 | 3.9 (0.40) b | <0.1 | 0.2 (0.05) b | |
| 12 | 0.5 | <0.1 | 0.1 | ||
| 24 | <0.1 | <0.1 | <0.1 | ||
| 1:3 | ONPG | 5 | <0.1 | 3.9 (1.17) b | <0.1 |
| 12 | <0.1 | 23.2 | <0.1 | ||
| 24 | <0.1 | 19.2 | <0.1 | ||
| Lactose | 5 | <0.1 | 0.1 (0.01) b | <0.1 | |
| 12 | <0.1 | 0.6 | <0.1 | ||
| 24 | 0.1 | <0.1 | <0.1 | ||
| 3:1 | ONPG | 5 | <0.1 | 3.8 (0.40) b | 0.2 (0.02) b |
| 12 | <0.1 | 2.9 | <0.1 | ||
| 24 | <0.1 | <0.1 | <0.1 | ||
| Lactose | 5 | <0.1 | <0.1 | <0.1 | |
| 12 | <0.1 | <0.1 | <0.1 | ||
| 24 | <0.1 | <0.1 | <0.1 | ||
| 9:1 | ONPG | 5 | <0.1 | 0.9 (0.09) b | <0.1 |
| 12 | <0.1 | <0.1 | <0.1 | ||
| 24 | <0.1 | 0.8 | <0.1 | ||
| Lactose | 5 | 0.2 (0.02) b | 0.3 (0.03) b | <0.1 | |
| 12 | 0.1 | <0.1 | <0.1 | ||
| 24 | <0.1 | <0.1 | <0.1 | ||
2.2. Transgalactosylation of Serine/Threonine Derivatives
 ), 12 h (
), 12 h (  ), and 24 h (
), and 24 h (  ) of reaction and the initial rates of the reaction (right axis, ●).
) of reaction and the initial rates of the reaction (right axis, ●).
 ), 12 h (
), 12 h (  ), and 24 h (
), and 24 h (  ) of reaction and the initial rates of the reaction (right axis, ●).
) of reaction and the initial rates of the reaction (right axis, ●).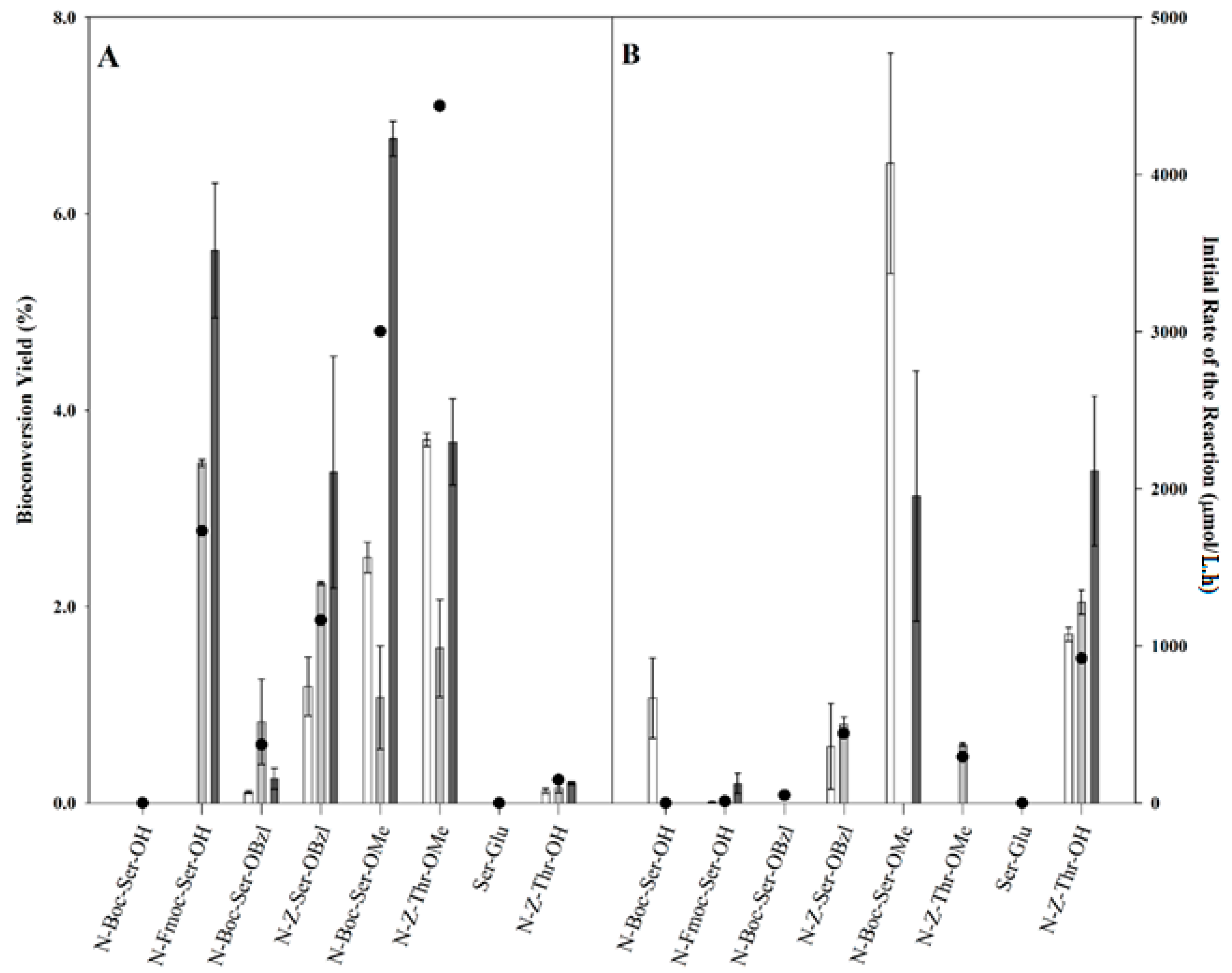
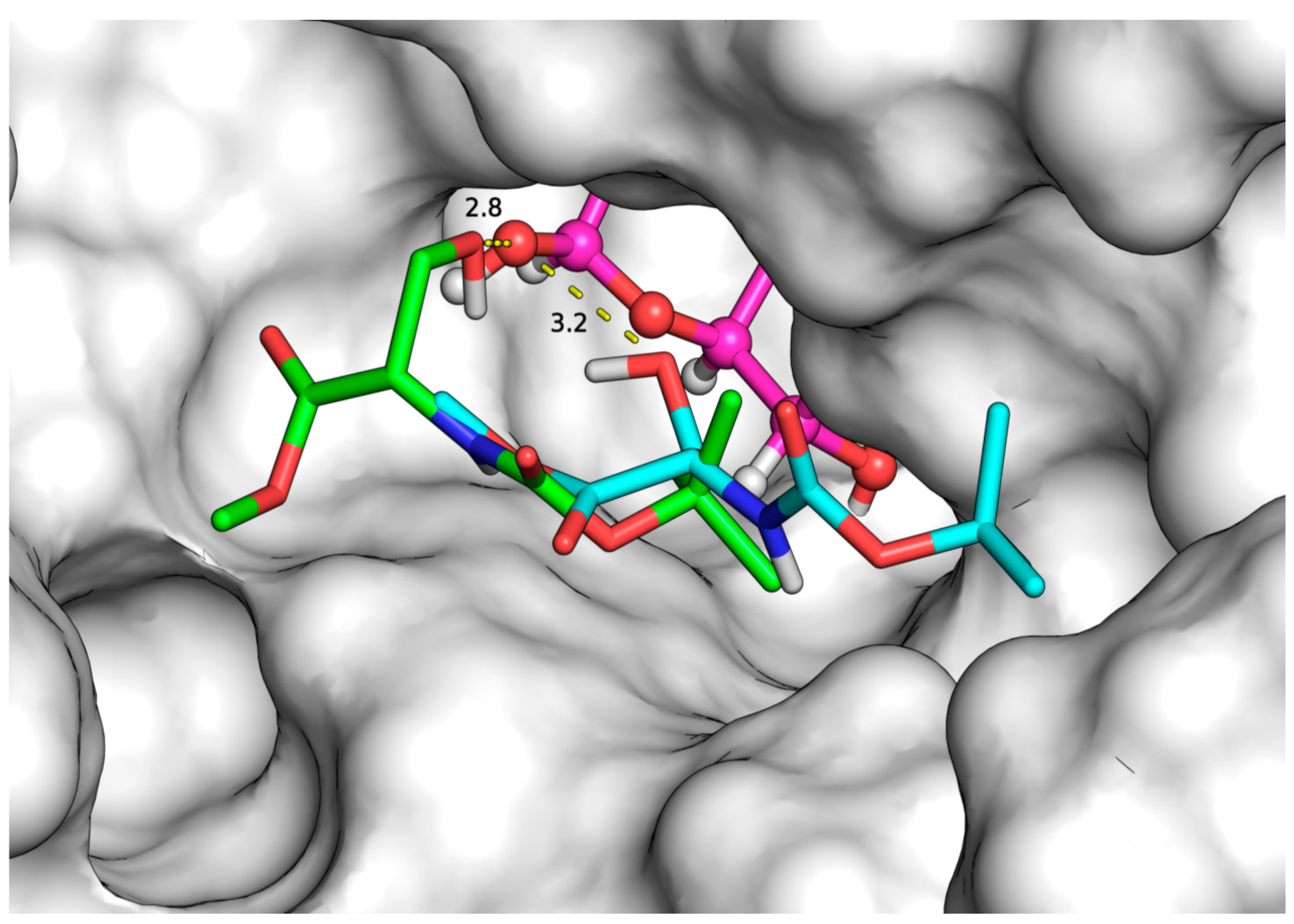
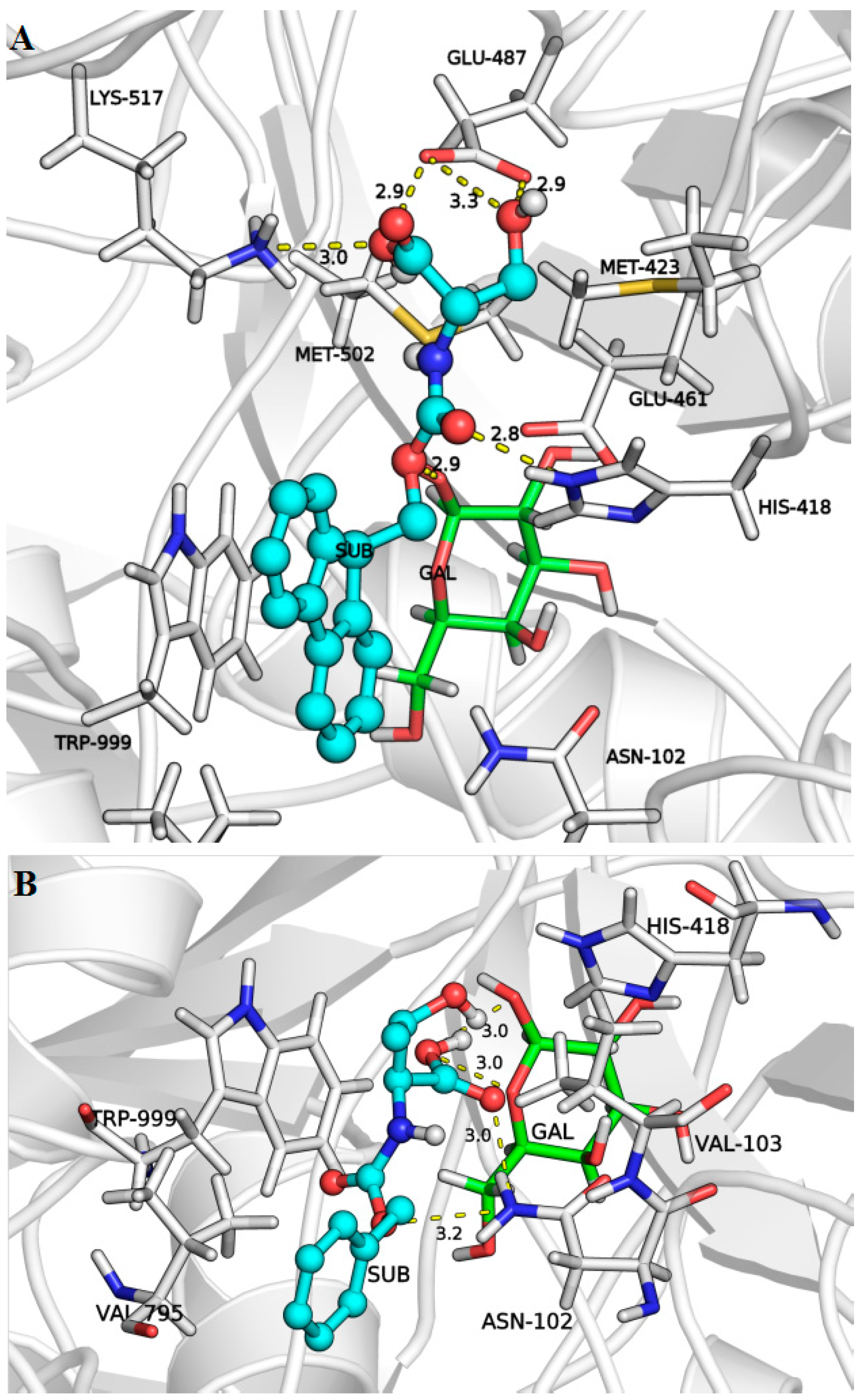
2.3. Docking Simulations
| Substrates (Acceptors) | Initial Rate (μmol/L.h) | Binding Modes | Binding Affinity (kcal/mol) | Distance (Å) |
|---|---|---|---|---|
| N-Fmoc-Ser-OH | 1731.2 a | 1 | −7.1 | 3.7 |
| 2 | −6.3 | 2.8 | ||
| N-Boc-Ser-OBzl | 370.8 a | 1 | −6.2 | 3.1 |
| 2 | −6.0 | 3.0 | ||
| 3 | −5.9 | 2.9 | ||
| 4 | −5.7 | 3.1 | ||
| 5 | −5.5 | 3.0 | ||
| N-Z-Ser-OBzl | 1163.6 a | 1 | −7.0 | 3.0 |
| 2 | −6.6 | 2.7 | ||
| 3 | −6.6 | 2.8 | ||
| N-Z-Ser-OMe | 4680.0 a | 1 | −4.9 | 3.1 |
| N-Boc-Ser-OMe | 7816.8 b | 1 | −4.7 | 2.8 |
| 2 | −4.6 | 3.2 | ||
| 3 | −4.5 | 3.0 | ||
| N-Z-Thr-OMe | 4437.2 a | 1 | −5.7 | 2.7 |
| 2 | −5.5 | 2.9 | ||
| N-Z-Thr-OH | 920.4 b | 1 | −5.7 | 3.6 |
3. Experimental Section
3.1. Materials
3.2. Enzymatic Transgalactosylation
3.3. Acceptor Specificity
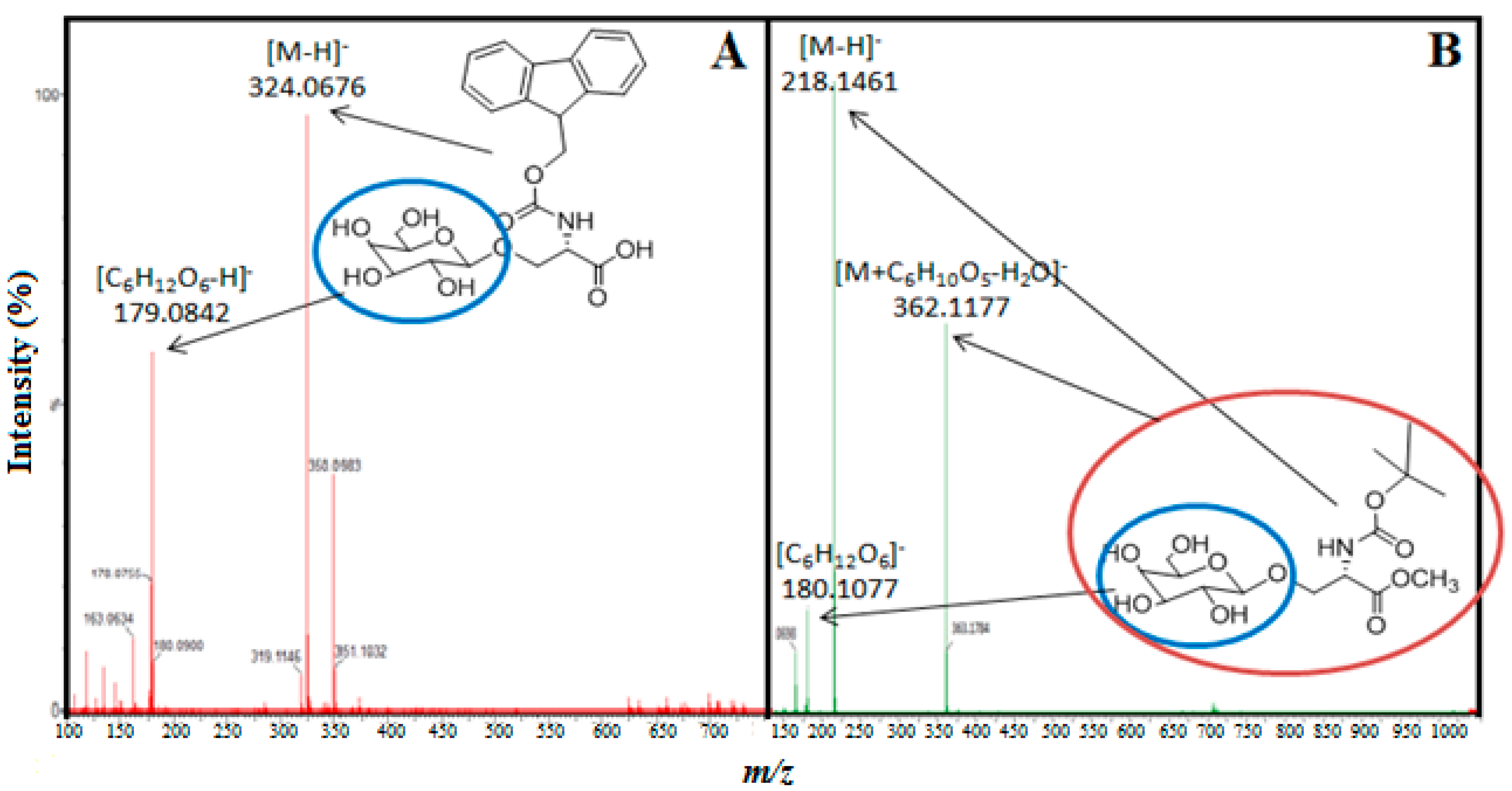
3.4. Analytical Methods
3.5. Computational Analysis
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Nishikawa, I.; Nakajima, Y.; Ito, M.; Fukuchi, S.; Homma, K.; Nishikawa, K. Computational prediction of O-linked glycosylation sites that preferentially map on intrinsically disordered regions of extracellular proteins. Int. J. Mol. Sci. 2010, 11, 4991–5008. [Google Scholar] [PubMed]
- Hakomori, S.; Cummings, R.D. Glycosylation effects on cancer development. Glycoconj. J. 2012, 29, 565–566. [Google Scholar] [PubMed]
- Oliver, C.M.; Melton, L.D.; Stanley, R.A. Creating proteins with novel functionality via the maillard reaction: A review. Crit. Rev. Food Sci. Nutr. 2006, 46, 337–350. [Google Scholar] [CrossRef] [PubMed]
- Schmaltz, R.M.; Hanson, S.R.; Wong, C.-H. Enzymes in the synthesis of glycoonjugates. Chem. Rev. 2011, 111, 4259–4307. [Google Scholar] [CrossRef] [PubMed]
- Lombard, V.; Golaconda Ramulu, H.; Drula, E.; Coutinho, P.M.; Henrissat, B. The carbohydrate-active enzymes database (CAZy) in 2013. Nucleic Acids Res. 2014, 42, D490–D485. [Google Scholar] [CrossRef] [PubMed]
- Brás, N.F.; Fernandes, P.A.; Ramos, M.J. Qm/mm studies on the β-galactosidase catalytic mechanism: Hydrolysis and transglycosylation reactions. J. Chem. Theory Comput. 2010, 6, 421–433. [Google Scholar] [CrossRef]
- Juers, D.H.; Matthews, B.W.; Huber, R.E. LacZ β-galactosidase: Structure and function of an enzyme of historical and molecular biological importance. Protein Sci. 2012, 21, 1792–1807. [Google Scholar] [CrossRef] [PubMed]
- Juers, D.H.; Heightman, T.D.; Vasella, A.; McCarter, J.D.; Mackenzie, L.; Withers, S.G.; Matthews, B.W. A structural view of the action of Escherichia coli (lacz) β-galactosidase. Biochemistry 2001, 40, 14781–14794. [Google Scholar] [CrossRef] [PubMed]
- Matthews, B.W. The structure of E. coli β-galactosidase. CR Biol. 2005, 328, 549–556. [Google Scholar] [CrossRef] [PubMed]
- Attal, S.; Bay, S.; Cantacuzene, D. Enzymatic synthesis of β-galactosyldipeptides and of β-1,3-digalactosylserine derivatives using β-galactosidase. Tetrahedron 1992, 48, 9251–9260. [Google Scholar] [CrossRef]
- Bay, S.; Namane, A.; Cantacuzene, D. Enzymatic synthesis of some o-β-d-digalactosyl glycopeptides, using β-d-galactosidase. Carbohydr. Res. 1993, 248, 317–325. [Google Scholar] [CrossRef]
- Becker, K.-C.; Kuhl, P. Synthesis of O-β-galactopyranosyl-l-serine derivatives using β-galactosidase in aqueous-organic reaction systems. J. Carbohydr. Chem. 1999, 18, 121–129. [Google Scholar] [CrossRef]
- Cantacuzene, D.; Attal, S. Enzymic synthesis of galactopyranosyl-l-serine derivatives using galactosidases. Carbohydr. Res. 1991, 211, 327–331. [Google Scholar] [CrossRef]
- Holla, E.W.; Schudok, M.; Weber, A.; Zulauf, M. Enzyme-catalyzed synthesis of O-glycopeptide building blocks. J. Carbohydr. Chem. 1992, 11, 659–663. [Google Scholar] [CrossRef]
- Johansson, E.; Hedbys, L.; Larsson, P.O. Enzymatic synthesis of monosaccharide-amino acid conjugates. Enzym. Microb. Technol. 1991, 13, 781–787. [Google Scholar] [CrossRef]
- Layer, A.; Fischer, F. Enzymatic O-galactosylation of protected serine and threonine by β-galactosidase employing high lactose concentrations. Ind. Eng. Chem. Res. 2006, 45, 6619–6621. [Google Scholar] [CrossRef]
- Honarparvar, B.; Govender, T.; Maguire, G.E.M.; Soliman, M.E.S.; Kruger, H.G. Integrated approach to structure-based enzymatic drug design: Molecular modeling, spectroscopy, and experimental bioactivity. Chem. Rev. 2014, 114, 493–537. [Google Scholar] [CrossRef] [PubMed]
- Klibanov, A.M. Enzymatic catalysis in anhydrous organic solvents. Trends Biochem. Sci. 1989, 14, 141–144. [Google Scholar] [CrossRef]
- Yang, L.; Dordick, J.S.; Garde, S. Hydration of enzyme in nonaqueous media is consistent with solvent dependence of its activity. Biophys. J. 2004, 87, 812–821. [Google Scholar] [CrossRef] [PubMed]
- Cantacuzene, D.; Attal, S.; Bay, S. Stereospecific chemoenzymatic synthesis of galactopyranosyl-l-serine. Bioorg. Med. Chem. Lett. 1991, 1, 197–200. [Google Scholar] [CrossRef]
- Mann, M.; Jensen, O.N. Proteomic analysis of post-translational modifications. Nat. Biotechnol. 2003, 21, 255–261. [Google Scholar] [CrossRef] [PubMed]
- Huber, R.E.; Hakda, S.; Cheng, C.; Cupples, C.G.; Edwards, R.A. Trp-999 of β-galactosidase (Escherichia coli) is a key residue for binding, catalysis, and synthesis of allolactose, the natural lac operon inducer. Biochemistry 2003, 42, 1796–1803. [Google Scholar] [CrossRef] [PubMed]
- Martinez-Bilbao, M.; Huber, R.E. Substitutions for Gly-794 show that binding interactions are important determinants of the catalytic action of β-galactosidase (Escherichia coli). Biochem. Cell Biol. 1994, 72, 313–319. [Google Scholar] [CrossRef] [PubMed]
- Brás, N.F.; Fernandes, P.A.; Ramos, M.J. Docking and molecular dynamics studies on the stereoselectivity in the enzymatic synthesis of carbohydrates. Theor. Chem. Acc. 2009, 122, 283–296. [Google Scholar] [CrossRef]
- Pérez-Sánchez, M.; Cabrera, Á.C.; García-Martín, H.; Sinisterra, J.V.; García, J.I.; Hernáiz, M.J. Improved synthesis of disaccharides with Escherichia coli β-galactosidase using bio-solvents derived from glycerol. Tetrahedron 2011, 67, 7708–7712. [Google Scholar] [CrossRef]
- Trott, O.; Olson, A.J. Autodock vina: Improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Comput. Chem. 2010, 31, 455–461. [Google Scholar] [CrossRef] [PubMed]
- Stierand, K.; Maab, P.; Rarey, M. Molecular complexes at a glance: Automated generation of two-dimentional complex diagrams. Bioinformatics 2006, 22, 1710–1716. [Google Scholar] [CrossRef] [PubMed]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Seo, S.; Rebehmed, J.; De Brevern, A.G.; Karboune, S. Enzymatic Synthesis of Galactosylated Serine/Threonine Derivatives by β-Galactosidase from Escherichia coli. Int. J. Mol. Sci. 2015, 16, 13714-13728. https://doi.org/10.3390/ijms160613714
Seo S, Rebehmed J, De Brevern AG, Karboune S. Enzymatic Synthesis of Galactosylated Serine/Threonine Derivatives by β-Galactosidase from Escherichia coli. International Journal of Molecular Sciences. 2015; 16(6):13714-13728. https://doi.org/10.3390/ijms160613714
Chicago/Turabian StyleSeo, Sooyoun, Joseph Rebehmed, Alexandre G. De Brevern, and Salwa Karboune. 2015. "Enzymatic Synthesis of Galactosylated Serine/Threonine Derivatives by β-Galactosidase from Escherichia coli" International Journal of Molecular Sciences 16, no. 6: 13714-13728. https://doi.org/10.3390/ijms160613714
APA StyleSeo, S., Rebehmed, J., De Brevern, A. G., & Karboune, S. (2015). Enzymatic Synthesis of Galactosylated Serine/Threonine Derivatives by β-Galactosidase from Escherichia coli. International Journal of Molecular Sciences, 16(6), 13714-13728. https://doi.org/10.3390/ijms160613714