The Potential of the Combination of CRISPR/Cas9 and Pluripotent Stem Cells to Provide Human Organs from Chimaeric Pigs
Abstract
:1. Introduction
2. Designer Endonucleases as a Tool for Precise and Efficient Gene Editing
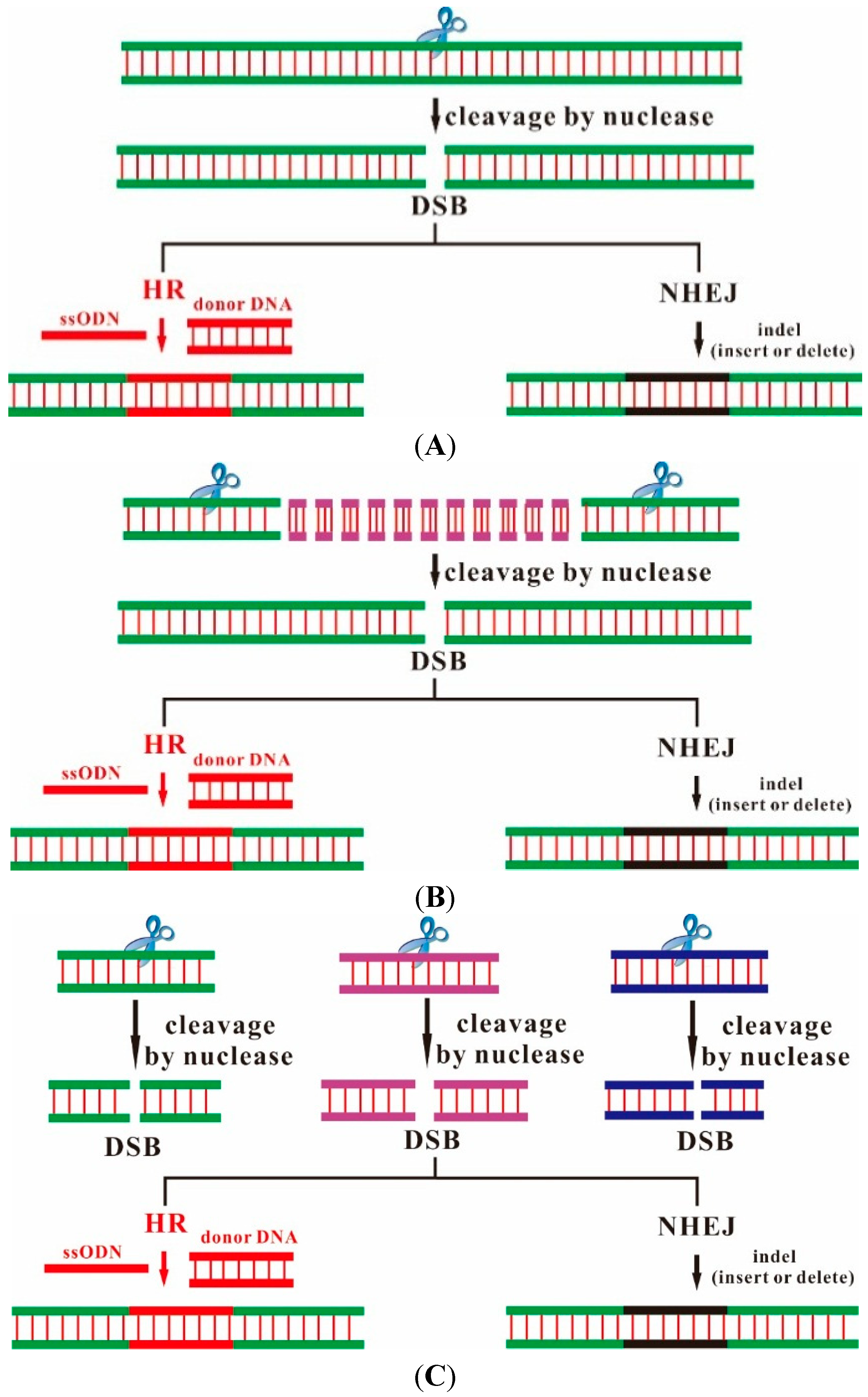
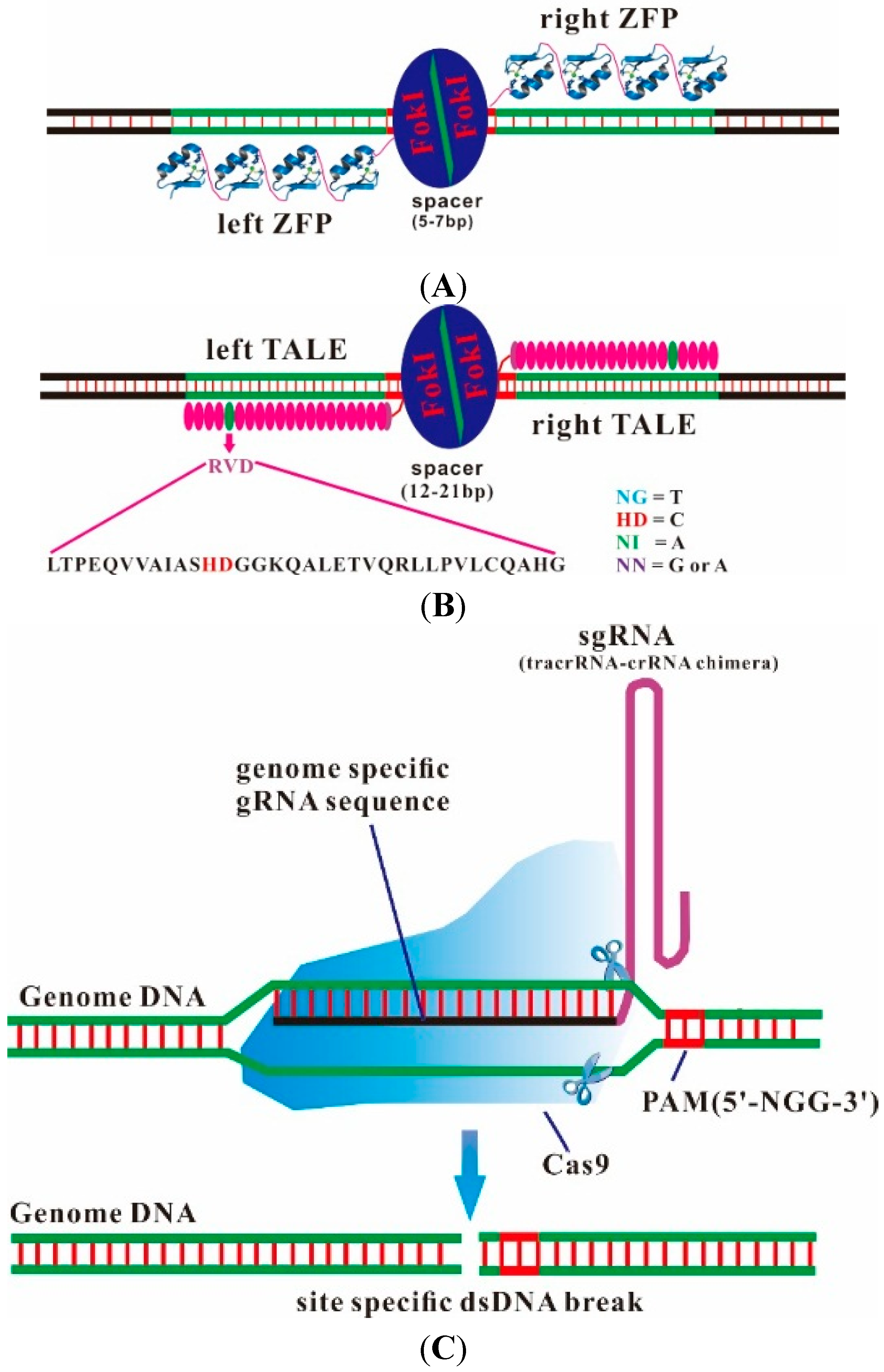
| Factors | ZFN | TALEN | CRISPR/Cas9 |
|---|---|---|---|
| Nuclease construction | significant | significant | simple |
| In vitro testing | significant | significant | simple |
| Target-efficient | limiting factor | average | good |
| Off-target-efficient | high | low | low |
| Target site choose | limited | limited | unlimited |
| Multiple gene mutations | limited | limited | unlimited |
| Designed component | protein | protein | RNA |
| Essential components | zinc finger proteins + FokI fusion protein | TALE and FokI fusion protein | guid RNA + Cas9 protein |
| Time consumption | long (7–15 days) | long (5–7 days) | short (1–3 days) |
| Cost | high | high | low |
3. From Bacterial CRISPR Immune Systems to Engineered RNA-Guided Endonucleases (CRISPR/Cas9)
4. Rapid and Efficient Generation of Genetically-Modified Animals
5. Making Human Organs from Human iPSCs and Genetically-Engineered Chimaeric Pigs
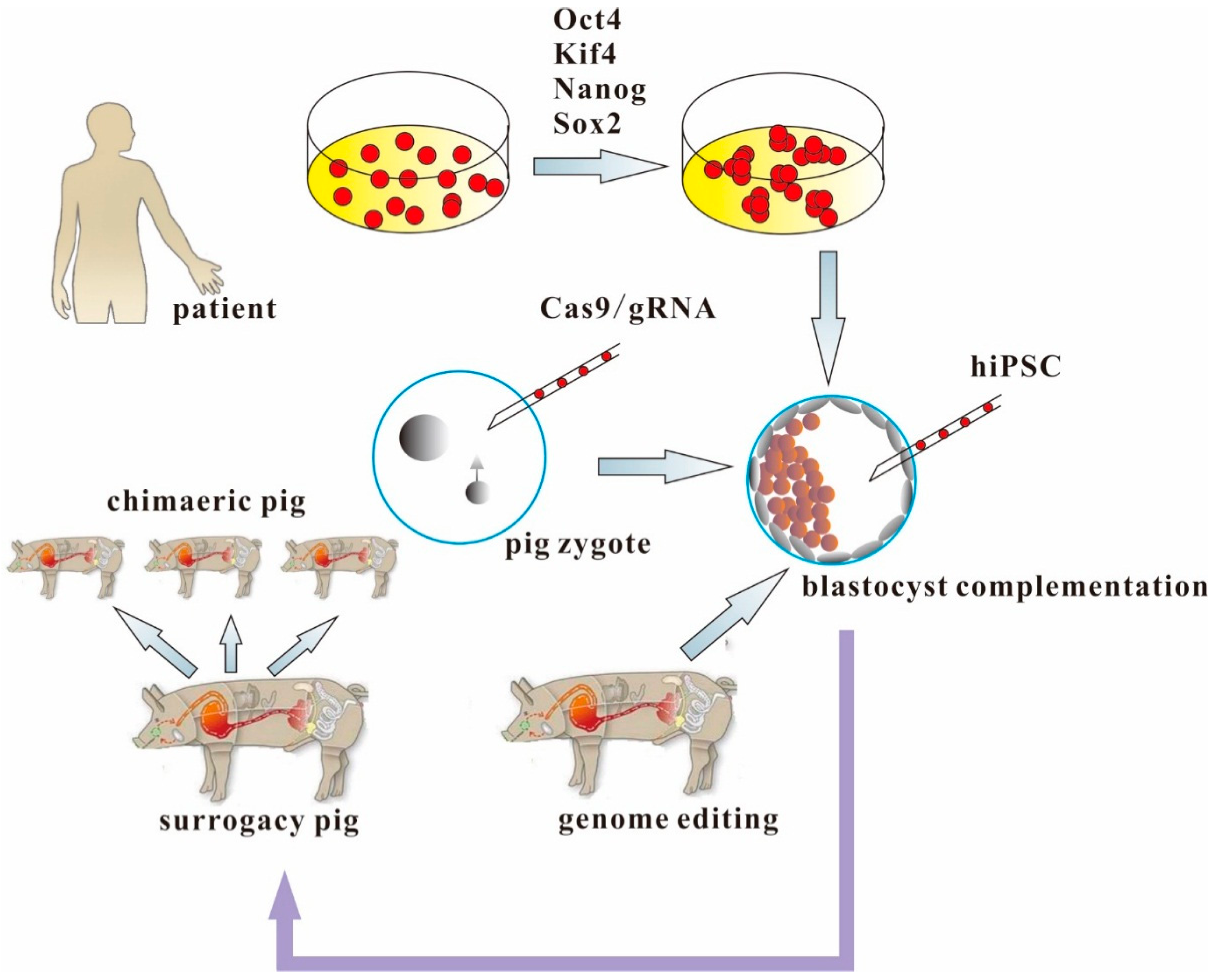
6. Conclusions
Acknowledgments
Abbreviations
| CRISPR | Clustered regularly interspaced short palindromic repeats |
| Cas | CRISPR-associated |
| PSC | pluripotent stem cells |
| TALENS | transcription activator-like effector nucleases |
| ZFNs | zinc-finger nucleases |
Conflicts of Interest
References
- Matsunari, H.; Nagashima, H.; Watanabe, M.; Umeyama, K.; Nakano, K.; Nagaya, M.; Kobayashi, T.; Yamaguchi, T.; Sumazaki, R.; Herzenberg, L.A. Blastocyst complementation generates exogenic pancreas in vivo in apancreatic cloned pigs. Proc. Natl. Acad. Sci. USA 2013, 110, 4557–4562. [Google Scholar] [CrossRef] [PubMed]
- Kang, L.; Wang, J.; Zhang, Y.; Kou, Z.; Gao, S. iPS cells can support full-term development of tetraploid blastocyst-complemented embryos. Cell Stem Cell 2009, 5, 135–138. [Google Scholar] [CrossRef] [PubMed]
- Lee, K.; Kwon, D.-N.; Ezashi, T.; Choi, Y.-J.; Park, C.; Ericsson, A.C.; Brown, A.N.; Samuel, M.S.; Park, K.-W.; Walters, E.M. Engraftment of human iPS cells and allogeneic porcine cells into pigs with inactivated RAG2 and accompanying severe combined immunodeficiency. Proc. Natl. Acad. Sci. USA 2014, 111, 7260–7265. [Google Scholar] [CrossRef] [PubMed]
- Shaw, D.; Dondorp, W.; Geijsen, N.; de Wert, G. Creating human organs in chimaera pigs: An ethical source of immunocompatible organs? J. Med. Ethics 2014. [Google Scholar] [CrossRef]
- Rashid, T.; Kobayashi, T.; Nakauchi, H. Revisiting the flight of icarus: Making human organs from PSCs with large animal chimeras. Cell Stem Cell 2014, 15, 406–409. [Google Scholar] [CrossRef] [PubMed]
- Gaj, T.; Gersbach, C.A.; Barbas, C.F., III. ZFN, TALEN, and CRISPR/Cas-based methods for genome engineering. Trends Biotechnol. 2013, 31, 397–405. [Google Scholar] [CrossRef] [PubMed]
- Sanjana, N.E.; Cong, L.; Zhou, Y.; Cunniff, M.M.; Feng, G.; Zhang, F. A transcription activator-like effector toolbox for genome engineering. Nat. Protoc. 2012, 7, 171–192. [Google Scholar] [CrossRef] [PubMed]
- Joung, J.K.; Sander, J.D. Talens: A widely applicable technology for targeted genome editing. Nat. Rev. Mol. Cell Biol. 2012, 14, 49–55. [Google Scholar] [CrossRef] [PubMed]
- Zhang, F.; Cong, L.; Lodato, S.; Kosuri, S.; Church, G.M.; Arlotta, P. Efficient construction of sequence-specific TAL effectors for modulating mammalian transcription. Nat. Biotechnol. 2011, 29, 149–153. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Reyon, D.; Tsai, S.Q.; Khayter, C.; Foden, J.A.; Sander, J.D.; Joung, J.K. Flash assembly of TALENs for high-throughput genome editing. Nat. Biotechnol. 2012, 30, 460–465. [Google Scholar] [CrossRef] [PubMed]
- Cermak, T.; Doyle, E.L.; Christian, M.; Wang, L.; Zhang, Y.; Schmidt, C.; Baller, J.A.; Somia, N.V.; Bogdanove, A.J.; Voytas, D.F. Efficient design and assembly of custom TALEN and other TAL effector-based constructs for DNA targeting. Nucleic Acids Res. 2011, 39, e82. [Google Scholar] [CrossRef] [PubMed]
- Maeder, M.L.; Thibodeau-Beganny, S.; Sander, J.D.; Voytas, D.F.; Joung, J.K. Oligomerized pool engineering (open): An “open-source” protocol for making customized zinc-finger arrays. Nat. Protoc. 2009, 4, 1471–1501. [Google Scholar] [CrossRef] [PubMed]
- Sander, J.D.; Dahlborg, E.J.; Goodwin, M.J.; Cade, L.; Zhang, F.; Cifuentes, D.; Curtin, S.J.; Blackburn, J.S.; Thibodeau-Beganny, S.; Qi, Y. Selection-free zinc-finger-nuclease engineering by context-dependent assembly (CoDA). Nat. Methods 2011, 8, 67–69. [Google Scholar] [CrossRef] [PubMed]
- Urnov, F.D.; Miller, J.C.; Lee, Y.-L.; Beausejour, C.M.; Rock, J.M.; Augustus, S.; Jamieson, A.C.; Porteus, M.H.; Gregory, P.D.; Holmes, M.C. Highly efficient endogenous human gene correction using designed zinc-finger nucleases. Nature 2005, 435, 646–651. [Google Scholar] [CrossRef] [PubMed]
- Mali, P.; Esvelt, K.M.; Church, G.M. Cas9 as a versatile tool for engineering biology. Nat. Methods 2013, 10, 957–963. [Google Scholar] [CrossRef] [PubMed]
- Ishino, Y.; Shinagawa, H.; Makino, K.; Amemura, M.; Nakata, A. Nucleotide sequence of the iap gene, responsible for alkaline phosphatase isozyme conversion in Escherichia coli, and identification of the gene product. J. Bacteriol. 1987, 169, 5429–5433. [Google Scholar] [PubMed]
- Westra, E.R.; Swarts, D.C.; Staals, R.H.; Jore, M.M.; Brouns, S.J.; van der Oost, J. The crisprs, they are a-changin': How prokaryotes generate adaptive immunity. Annu. Rev. Genet. 2012, 46, 311–339. [Google Scholar] [CrossRef] [PubMed]
- Wiedenheft, B.; Sternberg, S.H.; Doudna, J.A. RNA-guided genetic silencing systems in bacteria and archaea. Nature 2012, 482, 331–338. [Google Scholar] [CrossRef] [PubMed]
- Jinek, M.; Chylinski, K.; Fonfara, I.; Hauer, M.; Doudna, J.A.; Charpentier, E. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 2012, 337, 816–821. [Google Scholar] [CrossRef] [PubMed]
- Cong, L.; Ran, F.A.; Cox, D.; Lin, S.; Barretto, R.; Habib, N.; Hsu, P.D.; Wu, X.; Jiang, W.; Marraffini, L.A. Multiplex genome engineering using CRISPR/Cas systems. Science 2013, 339, 819–823. [Google Scholar] [CrossRef] [PubMed]
- Mali, P.; Yang, L.; Esvelt, K.M.; Aach, J.; Guell, M.; DiCarlo, J.E.; Norville, J.E.; Church, G.M. RNA-guided human genome engineering via Cas9. Science 2013, 339, 823–826. [Google Scholar] [CrossRef] [PubMed]
- Jinek, M.; East, A.; Cheng, A.; Lin, S.; Ma, E.; Doudna, J. RNA-programmed genome editing in human cells. eLife 2013. [Google Scholar] [CrossRef]
- Hsu, P.D.; Lander, E.S.; Zhang, F. Development and applications of CRISPR-Cas9 for genome engineering. Cell 2014, 157, 1262–1278. [Google Scholar] [CrossRef] [PubMed]
- Doudna, J.A.; Charpentier, E. The new frontier of genome engineering with CRISPR-Cas9. Science 2014, 346. [Google Scholar] [CrossRef] [PubMed]
- Friedland, A.E.; Tzur, Y.B.; Esvelt, K.M.; Colaiácovo, M.P.; Church, G.M.; Calarco, J.A. Heritable genome editing in C. elegans via a CRISPR-Cas9 system. Nat. Methods 2013, 10, 741–743. [Google Scholar] [CrossRef] [PubMed]
- Bassett, A.R.; Tibbit, C.; Ponting, C.P.; Liu, J.-L. Highly efficient targeted mutagenesis of drosophila with the CRISPR/Cas9 system. Cell Rep. 2013, 4, 220–228. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.; Liang, D.; Wang, Y.; Bai, M.; Tang, W.; Bao, S.; Yan, Z.; Li, D.; Li, J. Correction of a genetic disease in mouse via use of CRISPR-Cas9. Cell Stem Cell 2013, 13, 659–662. [Google Scholar] [CrossRef] [PubMed]
- Hruscha, A.; Krawitz, P.; Rechenberg, A.; Heinrich, V.; Hecht, J.; Haass, C.; Schmid, B. Efficient CRISPR/Cas9 genome editing with low off-target effects in zebrafish. Development 2013, 140, 4982–4987. [Google Scholar] [CrossRef] [PubMed]
- Whitworth, K.M.; Lee, K.; Benne, J.A.; Beaton, B.P.; Spate, L.D.; Murphy, S.L.; Samuel, M.S.; Mao, J.; O’Gorman, C.; Walters, E.M. Use of the CRISPR/Cas9 system to produce genetically engineered pigs from in vitro-derived oocytes and embryos. Biol. Reprod. 2014. [Google Scholar] [CrossRef]
- Li, D.; Qiu, Z.; Shao, Y.; Chen, Y.; Guan, Y.; Liu, M.; Li, Y.; Gao, N.; Wang, L.; Lu, X. Heritable gene targeting in the mouse and rat using a CRISPR-Cas system. Nat. Biotechnol. 2013, 31, 681–683. [Google Scholar] [CrossRef] [PubMed]
- Yang, D.; Xu, J.; Zhu, T.; Fan, J.; Lai, L.; Zhang, J.; Chen, Y.E. Effective gene targeting in rabbits using RNA-guided Cas9 nucleases. J. Mol. Cell. Biol. 2014. [Google Scholar] [CrossRef]
- Ni, W.; Qiao, J.; Hu, S.; Zhao, X.; Regouski, M.; Yang, M.; Polejaeva, I.A.; Chen, C. Efficient gene knockout in goats using CRISPR/Cas9 system. PLoS One 2014, 9, e106718. [Google Scholar] [CrossRef] [PubMed]
- Cho, S.W.; Kim, S.; Kim, J.M.; Kim, J.-S. Targeted genome engineering in human cells with the cas9 RNA-guided endonuclease. Nat. Biotechnol. 2013, 31, 230–232. [Google Scholar] [CrossRef] [PubMed]
- Hai, T.; Teng, F.; Guo, R.; Li, W.; Zhou, Q. One-step generation of knockout pigs by zygote injection of CRISPR/Cas system. Cell Res. 2014, 24, 372–375. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Yang, H.; Shivalila, C.S.; Dawlaty, M.M.; Cheng, A.W.; Zhang, F.; Jaenisch, R. One-step generation of mice carrying mutations in multiple genes by CRISPR/Cas-mediated genome engineering. Cell 2013, 153, 910–918. [Google Scholar] [CrossRef] [PubMed]
- Yin, H.; Xue, W.; Chen, S.; Bogorad, R.L.; Benedetti, E.; Grompe, M.; Koteliansky, V.; Sharp, P.A.; Jacks, T.; Anderson, D.G. Genome editing with Cas9 in adult mice corrects a disease mutation and phenotype. Nat. Biotechnol. 2014, 32, 551–553. [Google Scholar] [CrossRef] [PubMed]
- Sánchez-Rivera, F.J.; Papagiannakopoulos, T.; Romero, R.; Tammela, T.; Bauer, M.R.; Bhutkar, A.; Joshi, N.S.; Subbaraj, L.; Bronson, R.T.; Xue, W. Rapid modeling of cooperating genetic events in cancer through somatic genome editing. Nature 2014, 516, 428–431. [Google Scholar] [CrossRef] [PubMed]
- Maddalo, D.; Manchado, E.; Concepcion, C.P.; Bonetti, C.; Vidigal, J.A.; Han, Y.-C.; Ogrodowski, P.; Crippa, A.; Rekhtman, N.; de Stanchina, E. In vivo engineering of oncogenic chromosomal rearrangements with the CRISPR/Cas9 system. Nature 2014, 516, 423–427. [Google Scholar] [CrossRef] [PubMed]
- Xue, W.; Chen, S.; Yin, H.; Tammela, T.; Papagiannakopoulos, T.; Joshi, N.S.; Cai, W.; Yang, G.; Bronson, R.; Crowley, D.G. CRISPR-mediated direct mutation of cancer genes in the mouse liver. Nature 2014, 514, 380–384. [Google Scholar] [CrossRef] [PubMed]
- Antonica, F.; Kasprzyk, D.F.; Opitz, R.; Iacovino, M.; Liao, X.-H.; Dumitrescu, A.M.; Refetoff, S.; Peremans, K.; Manto, M.; Kyba, M. Generation of functional thyroid from embryonic stem cells. Nature 2012, 491, 66–71. [Google Scholar] [CrossRef] [PubMed]
- Xia, Y.; Nivet, E.; Sancho-Martinez, I.; Gallegos, T.; Suzuki, K.; Okamura, D.; Wu, M.-Z.; Dubova, I.; Esteban, C.R.; Montserrat, N. Directed differentiation of human pluripotent cells to ureteric bud kidney progenitor-like cells. Nat. Cell Biol. 2013, 15, 1507–1515. [Google Scholar] [CrossRef] [PubMed]
- Takebe, T.; Sekine, K.; Enomura, M.; Koike, H.; Kimura, M.; Ogaeri, T.; Zhang, R.-R.; Ueno, Y.; Zheng, Y.-W.; Koike, N. Vascularized and functional human liver from an iPSC-derived organ bud transplant. Nature 2013, 499, 481–484. [Google Scholar] [CrossRef] [PubMed]
- Eiraku, M.; Takata, N.; Ishibashi, H.; Kawada, M.; Sakakura, E.; Okuda, S.; Sekiguchi, K.; Adachi, T.; Sasai, Y. Self-organizing optic-cup morphogenesis in three-dimensional culture. Nature 2011, 472, 51–56. [Google Scholar] [CrossRef] [PubMed]
- Nakano, T.; Ando, S.; Takata, N.; Kawada, M.; Muguruma, K.; Sekiguchi, K.; Saito, K.; Yonemura, S.; Eiraku, M.; Sasai, Y. Self-formation of optic cups and storable stratified neural retina from human ESCs. Cell Stem Cell 2012, 10, 771–785. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mannoor, M.S.; Jiang, Z.; James, T.; Kong, Y.L.; Malatesta, K.A.; Soboyejo, W.O.; Verma, N.; Gracias, D.H.; McAlpine, M.C. 3D printed bionic ears. Nano Lett. 2013, 13, 2634–2639. [Google Scholar] [CrossRef] [PubMed]
- Murphy, S.V.; Atala, A. 3D bioprinting of tissues and organs. Nat. Biotechnol. 2014, 32, 773–785. [Google Scholar] [CrossRef] [PubMed]
- Skardal, A.; Atala, A. Biomaterials for integration with 3-d bioprinting. Ann. Biomed. Eng. 2014, in press. [Google Scholar]
- Song, J.J.; Ott, H.C. Organ engineering based on decellularized matrix scaffolds. Trends Mol. Med. 2011, 17, 424–432. [Google Scholar] [CrossRef] [PubMed]
- Zambon, J.P.; Magalhaes, R.S.; Ko, I.; Ross, C.L.; Orlando, G.; Peloso, A.; Atala, A.; Yoo, J.J. Kidney regeneration: Where we are and future perspectives. World J. Nephrol. 2014, 3, 24–30. [Google Scholar] [CrossRef] [PubMed]
- Moser, P.T.; Ott, H.C. Recellularization of organs: What is the future for solid organ transplantation? Curr. Opin. Organ Transplant. 2014, 19, 603–609. [Google Scholar] [CrossRef] [PubMed]
- Soto-Gutierrez, A.; Wertheim, J.A.; Ott, H.C.; Gilbert, T.W. Perspectives on whole-organ assembly: Moving toward transplantation on demand. J. Clin. Investig. 2012, 122, 3817–3823. [Google Scholar] [CrossRef] [PubMed]
- Madariaga, M.L.; Ott, H.C. Bioengineering kidneys for transplantation. Semin. Nephrol. 2014, 34, 384–393. [Google Scholar] [CrossRef] [PubMed]
- Ekser, B.; Ezzelarab, M.; Hara, H.; van der Windt, D.J.; Wijkstrom, M.; Bottino, R.; Trucco, M.; Cooper, D.K. Clinical xenotransplantation: The next medical revolution? Lancet 2012, 379, 672–683. [Google Scholar] [CrossRef] [PubMed]
- Cooper, D.K.; Ekser, B.; Burlak, C.; Ezzelarab, M.; Hara, H.; Paris, L.; Tector, A.J.; Phelps, C.; Azimzadeh, A.M.; Ayares, D.; et al. Clinical lung xenotransplantation—What donor genetic modifications may be necessary? Xenotransplantation 2012, 19, 144–158. [Google Scholar] [CrossRef] [PubMed]
- Cooper, D.K.; Hara, H.; Ezzelarab, M.; Bottino, R.; Trucco, M.; Phelps, C.; Ayares, D.; Dai, Y. The potential of genetically-engineered pigs in providing an alternative source of organs and cells for transplantation. J. Biomed. Res. 2013, 27, 249–253. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Lansford, R.; Stewart, V.; Young, F.; Alt, F.W. Rag-2-deficient blastocyst complementation: An assay of gene function in lymphocyte development. Proc. Natl. Acad. Sci. USA 1993, 90, 4528–4532. [Google Scholar] [CrossRef] [PubMed]
- Kobayashi, T.; Yamaguchi, T.; Hamanaka, S.; Kato-Itoh, M.; Yamazaki, Y.; Ibata, M.; Sato, H.; Lee, Y.-S.; Usui, J.-I.; Knisely, A. Generation of rat pancreas in mouse by interspecific blastocyst injection of pluripotent stem cells. Cell 2010, 142, 787–799. [Google Scholar] [CrossRef] [PubMed]
- Isotani, A.; Hatayama, H.; Kaseda, K.; Ikawa, M.; Okabe, M. Formation of a thymus from rat ES cells in xenogeneic nude mouse↔rat ES chimeras. Genes Cells 2011, 16, 397–405. [Google Scholar] [CrossRef] [PubMed]
- Takashima, Y.; Guo, G.; Loos, R.; Nichols, J.; Ficz, G.; Krueger, F.; Oxley, D.; Santos, F.; Clarke, J.; Mansfield, W.; et al. Resetting transcription factor control circuitry toward ground-state pluripotency in human. Cell 2014, 158, 1254–1269. [Google Scholar] [CrossRef] [PubMed]
- Theunissen, T.W.; Powell, B.E.; Wang, H.; Mitalipova, M.; Faddah, D.A.; Reddy, J.; Fan, Z.P.; Maetzel, D.; Ganz, K.; Shi, L.; et al. Systematic identification of culture conditions for induction and maintenance of naive human pluripotency. Cell Stem Cell 2014, 15, 471–487. [Google Scholar] [CrossRef] [PubMed]
- Gafni, O.; Weinberger, L.; Mansour, A.A.; Manor, Y.S.; Chomsky, E.; Ben-Yosef, D.; Kalma, Y.; Viukov, S.; Maza, I.; Zviran, A.; et al. Derivation of novel human ground state naive pluripotent stem cells. Nature 2013, 504, 282–286. [Google Scholar] [CrossRef] [PubMed]
- Kobayashi, T.; Kato-Itoh, M.; Nakauchi, H. Targeted organ generation using Mixl1-inducible mouse pluripotent stem cells in blastocyst complementation. Stem Cells Dev. 2015, 24, 182–189. [Google Scholar] [CrossRef] [PubMed]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Feng, W.; Dai, Y.; Mou, L.; Cooper, D.K.C.; Shi, D.; Cai, Z. The Potential of the Combination of CRISPR/Cas9 and Pluripotent Stem Cells to Provide Human Organs from Chimaeric Pigs. Int. J. Mol. Sci. 2015, 16, 6545-6556. https://doi.org/10.3390/ijms16036545
Feng W, Dai Y, Mou L, Cooper DKC, Shi D, Cai Z. The Potential of the Combination of CRISPR/Cas9 and Pluripotent Stem Cells to Provide Human Organs from Chimaeric Pigs. International Journal of Molecular Sciences. 2015; 16(3):6545-6556. https://doi.org/10.3390/ijms16036545
Chicago/Turabian StyleFeng, Wanyou, Yifan Dai, Lisha Mou, David K. C. Cooper, Deshun Shi, and Zhiming Cai. 2015. "The Potential of the Combination of CRISPR/Cas9 and Pluripotent Stem Cells to Provide Human Organs from Chimaeric Pigs" International Journal of Molecular Sciences 16, no. 3: 6545-6556. https://doi.org/10.3390/ijms16036545
APA StyleFeng, W., Dai, Y., Mou, L., Cooper, D. K. C., Shi, D., & Cai, Z. (2015). The Potential of the Combination of CRISPR/Cas9 and Pluripotent Stem Cells to Provide Human Organs from Chimaeric Pigs. International Journal of Molecular Sciences, 16(3), 6545-6556. https://doi.org/10.3390/ijms16036545





