Biosynthesis of Essential Polyunsaturated Fatty Acids in Wheat Triggered by Expression of Artificial Gene
Abstract
:1. Introduction
2. Results and Discussion
2.1. Creating of Artificial Gene
2.2. Wheat Transformation by Artificial Gene
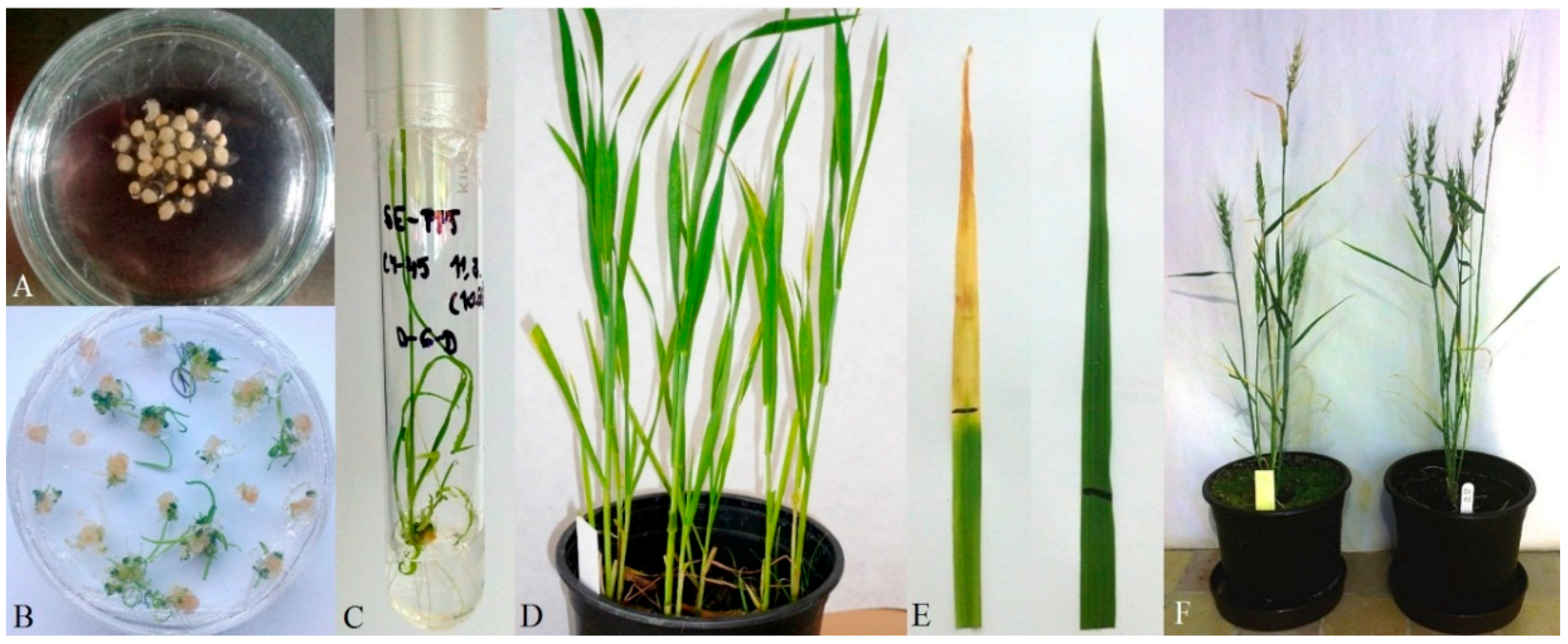

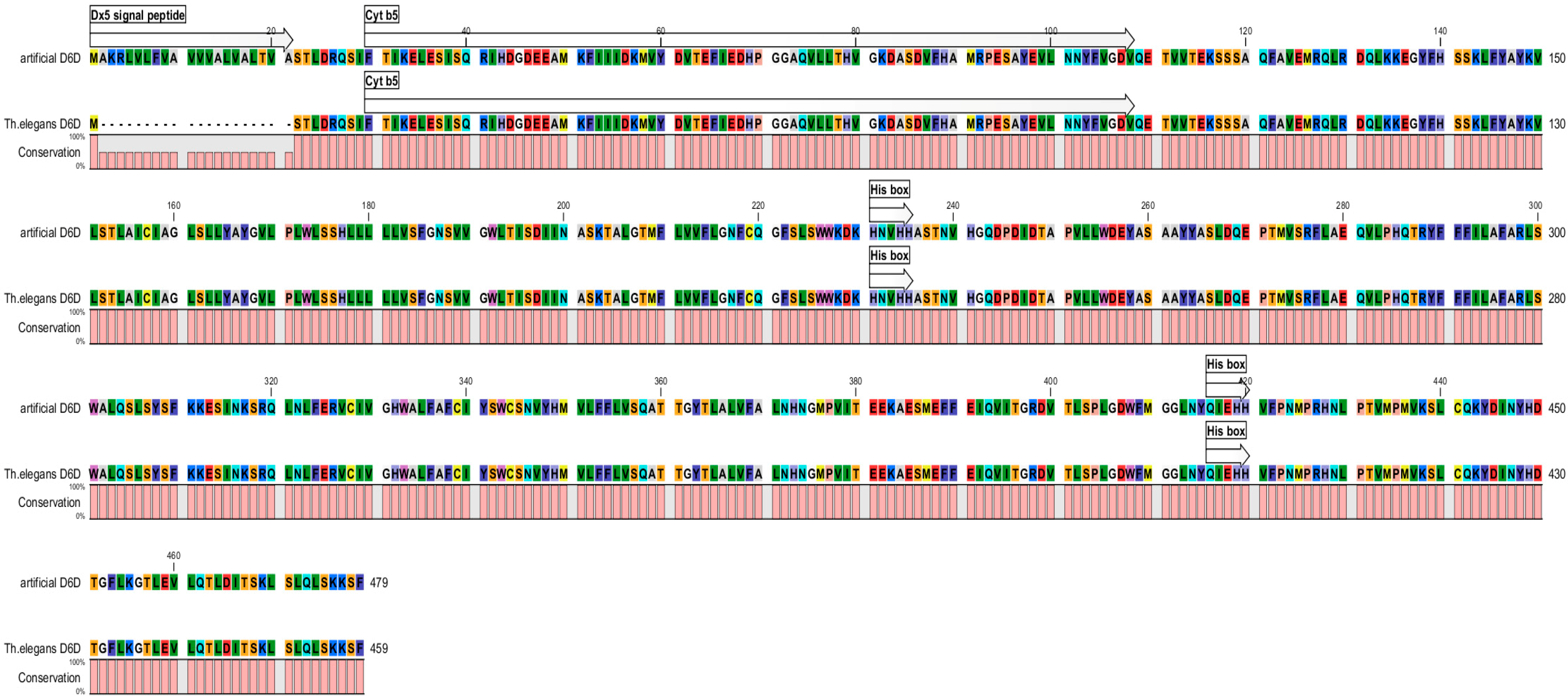

2.3. Analysis of Transgenic Wheats
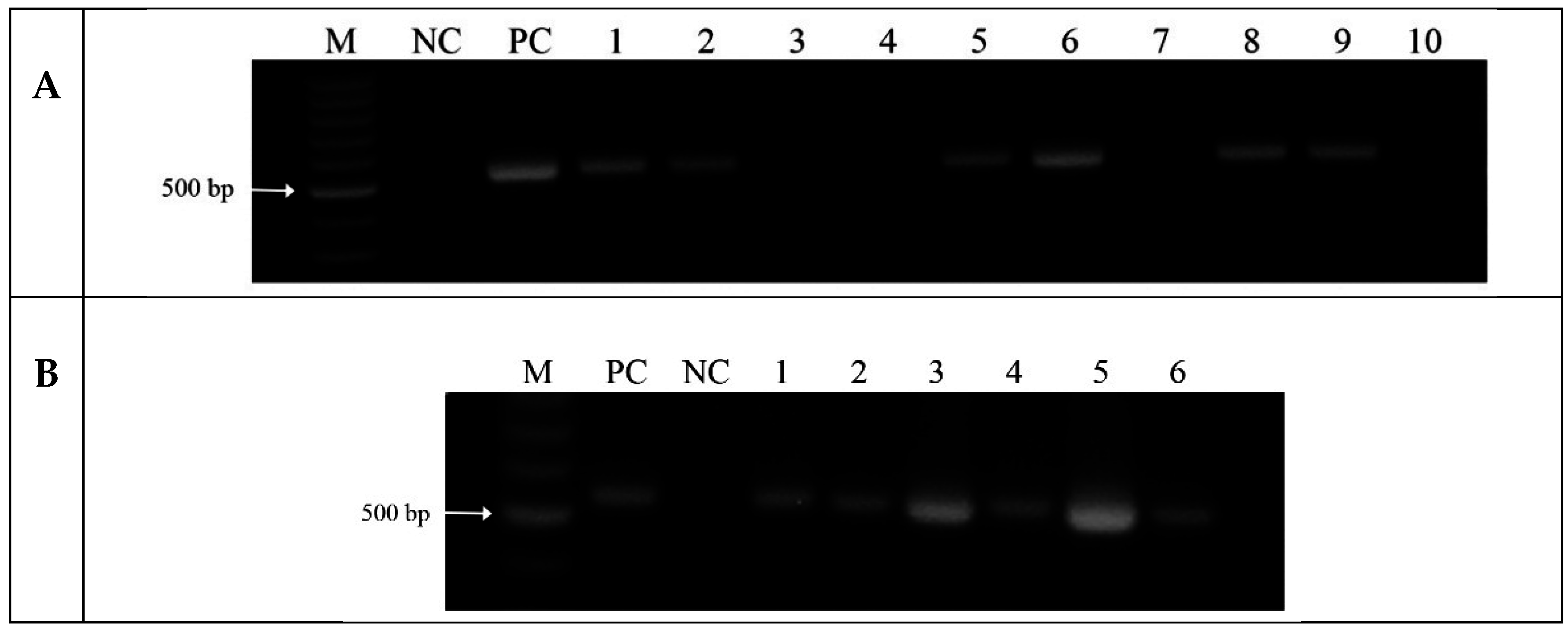

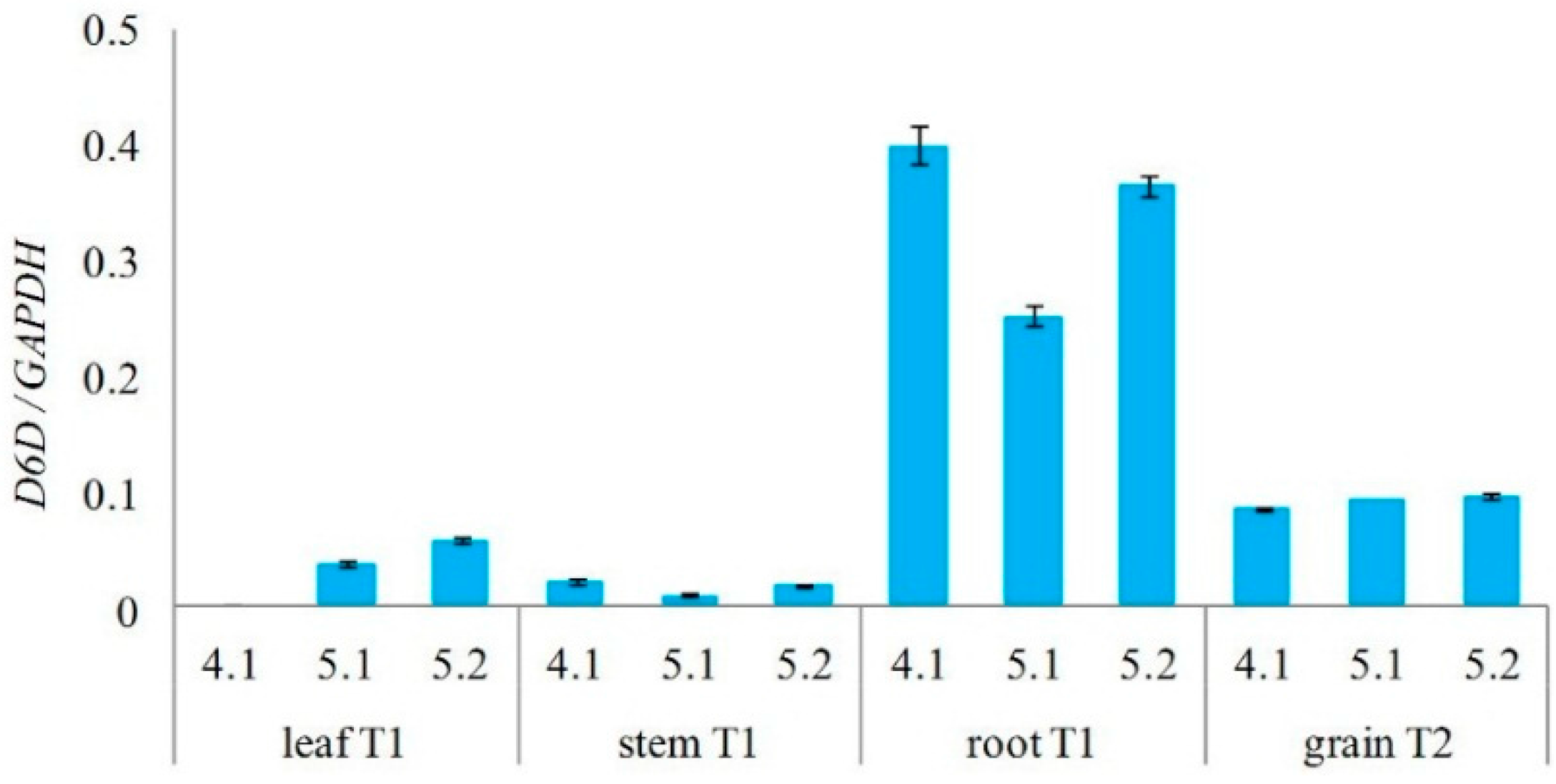
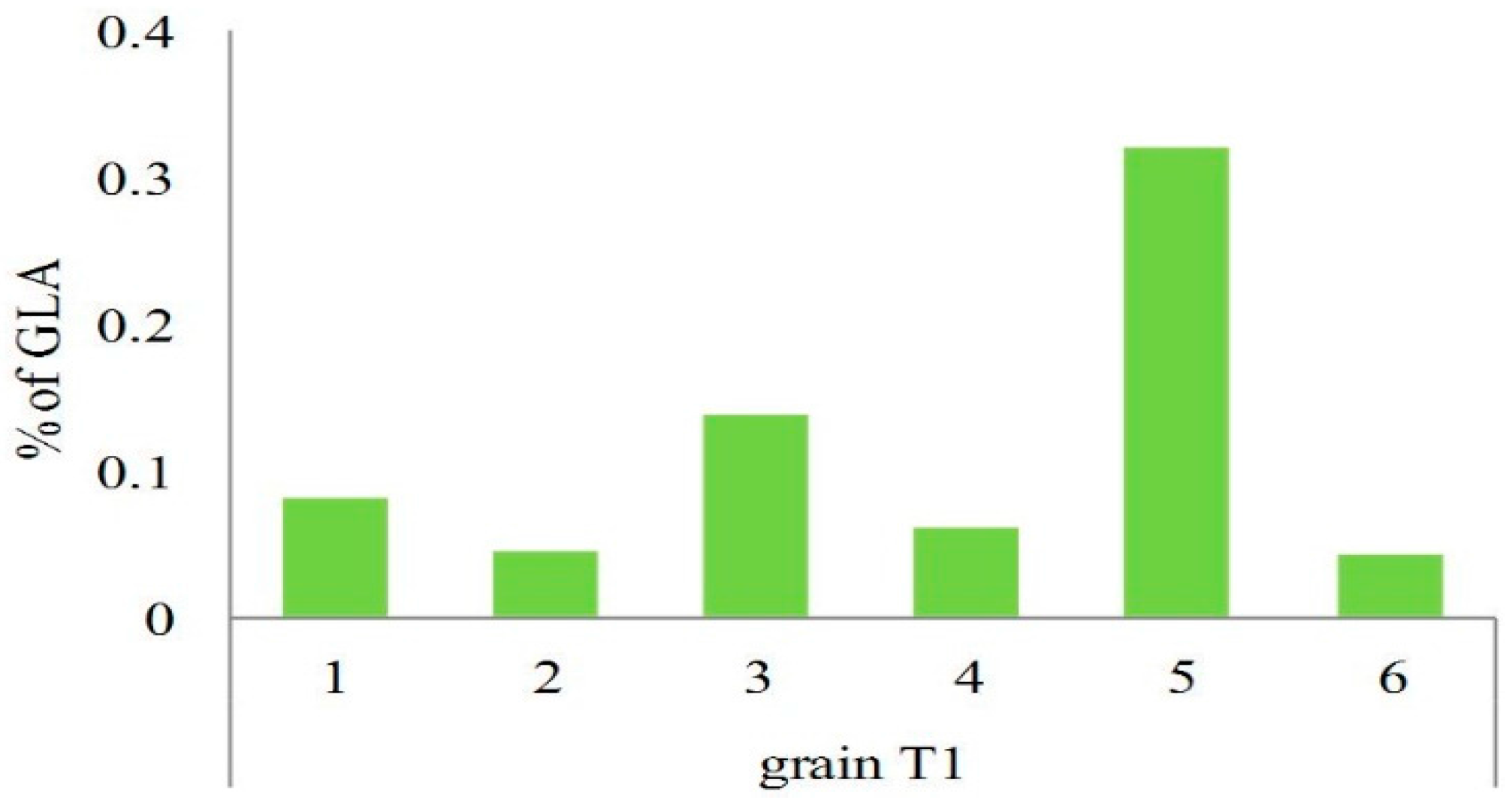
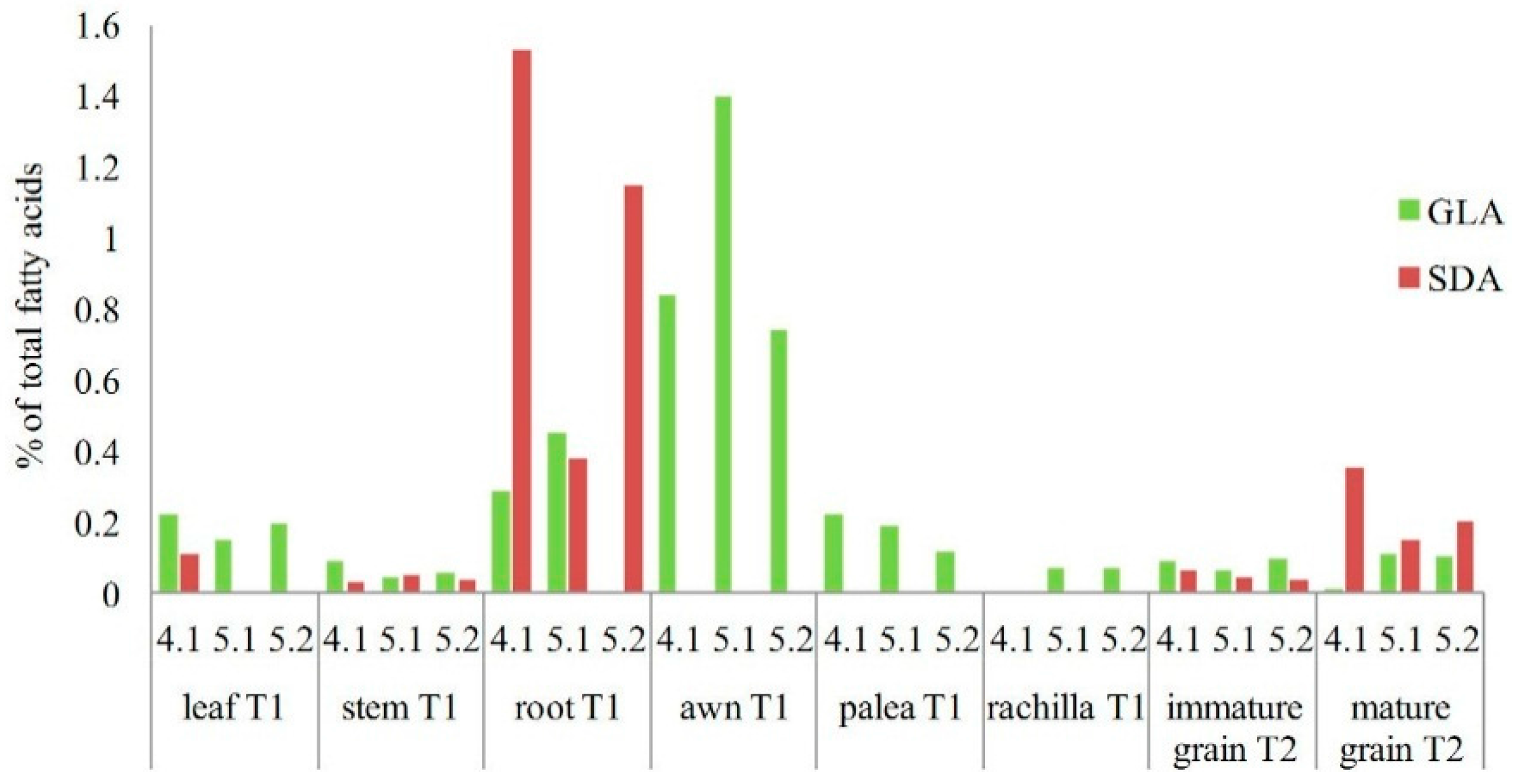
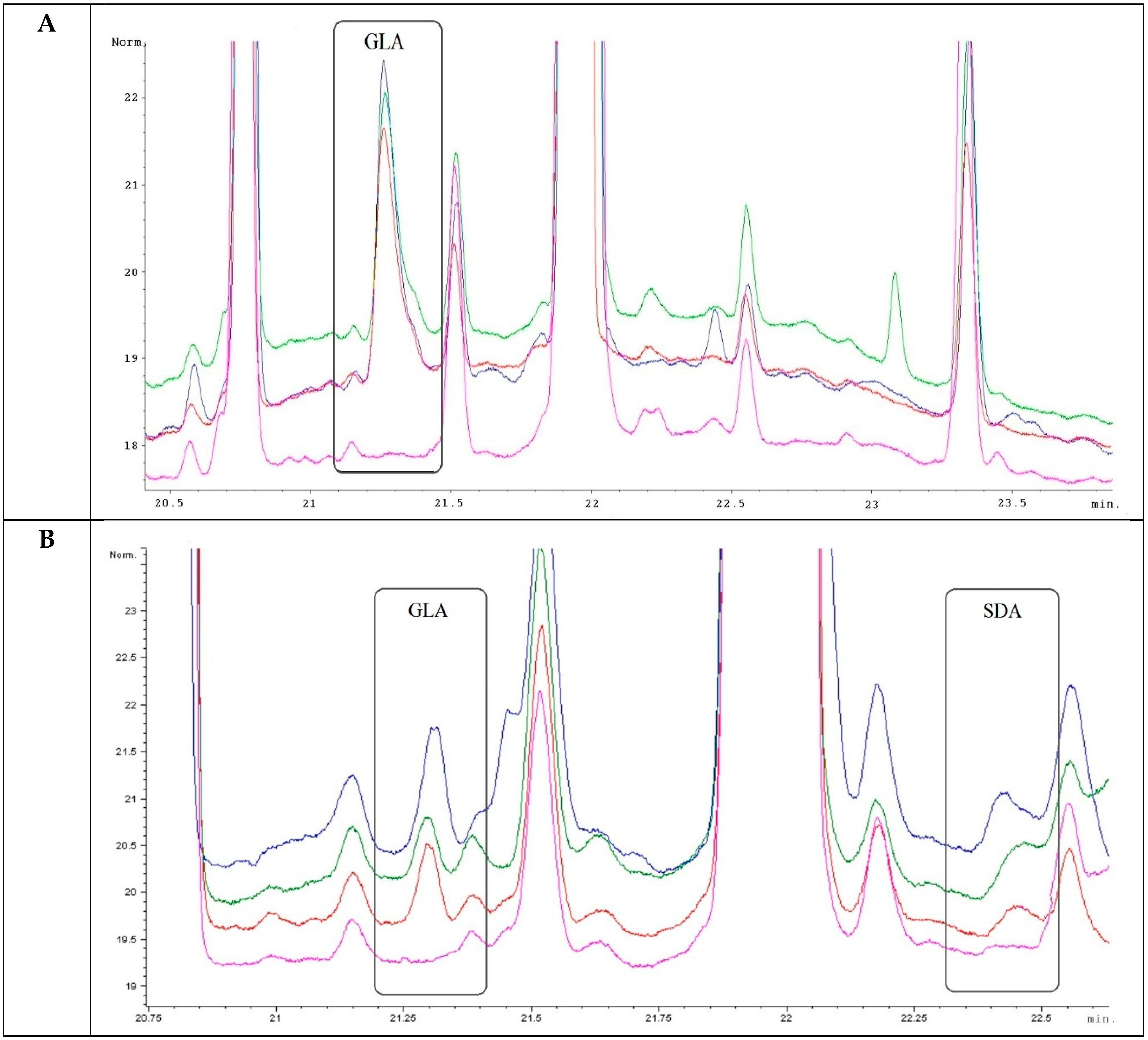
2.4. Analysis of PUFAs in Pathogens Invading Wheat
| Pathogen | Fatty Acid (%) | ||||||
|---|---|---|---|---|---|---|---|
| PA | SA | OA | VA | LA | ALA | Others | |
| Blumeria graminis | 37.1 | 24.3 | 2.8 | ND | 14.0 | 9.2 | 12.6 |
| Drechslera tritici-repentis | 24.1 | 3.5 | 34.8 | 6.7 | 23.0 | 4.4 | 3.5 |
| Stagonospora nodorum | 12.3 | 4.7 | 26.0 | 0.9 | 47.8 | 3.6 | 4.8 |
| Septoria tritici | 27.8 | 6.6 | 31.7 | 0.5 | 22.9 | 3.3 | 7.3 |
| Fusarium poae | 19.9 | 4.2 | 36.4 | 1.2 | 33.0 | 2.0 | 3.3 |
| Fusarium culmorum | 12.7 | 5.3 | 45.9 | 0.8 | 29.9 | 1.3 | 4.2 |
| Fusarium graminearum | 15.0 | 2.3 | 45.9 | 1.5 | 28.9 | 1.6 | 4.9 |
| Alternaria sp. | 17.2 | 4.8 | 33.1 | 2.7 | 37.0 | 1.2 | 4.0 |
3. Experimental Section
3.1. Biological Material
3.2. Artificial Gene and DNA Constructs
3.3. Wheat Transformation and Regeneration
3.4. DNA Analysis
3.5. RNA Analysis
3.6. Fatty Acids Analysis
4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Farré, G.; Twyman, R.M.; Changfu, Z.; Capell, T.; Christou, P. Nutritionally enhanced crops and food security: Scientific achievements versus political expediency. Curr. Opin. Biotechnol. 2011, 22, 245–251. [Google Scholar] [CrossRef] [PubMed]
- Dunford, N.T.; Zhang, M. Pressurized solvent extraction of wheat germ oil. Food Res. Int. 2003, 36, 905–909. [Google Scholar] [CrossRef]
- Horrobin, D.F. Fatty acid metabolism in health and disease: The role of Δ6desaturase. Am. J. Clin. Nutr. 1993, 57, 723S–727S. [Google Scholar]
- Certik, M.; Shimizu, S. Biosynthesis and regulation of microbial polyunsaturated fatty acid production. J. Biosci. Bioeng. 1999, 87, 1–14. [Google Scholar] [CrossRef]
- Fakas, S.; Čertik, M.; Papanikolaou, S.; Aggelis, G.; Komaitis, M.; Galiotou-Panagotou, M. γ-Linolenic acid production by Cunninghamella echinulata growing on complex organic nitrogen sources. Biores. Technol. 2008, 99, 5986–5990. [Google Scholar] [CrossRef]
- Gill, I.; Valivety, R. Polyunsaturated fatty acids, Part 1: Occurrence, biological activities and applications. Trends Biotechnol. 1997, 15, 401–409. [Google Scholar] [CrossRef]
- Hiruta, O.; Futamura, T.; Takebe, H.; Satoh, A.; Kamisaka, Y.; Yokochi, T.; Nakahara, T.; Suzuki, O. Optimization and scale-up of γ-linolenic acid production by Mortierella ramanniana MM 15-1, a high γ-linolenic acid producing mutant. J. Ferm. Bioeng. 1996, 82, 366–370. [Google Scholar] [CrossRef]
- Ratledge, C. Fatty acid biosynthesis in microorganisms being used for single cell oil production. Biochimie 2004, 86, 807–815. [Google Scholar] [CrossRef] [PubMed]
- Zhao, X.; Kong, X.; Hua, Y.; Feng, B.; Zhao, Z.K. Medium optimization for lipid production through co-fermentation of glucose and xylose by oleaginous yeast Lipomyces starkeyi. Eur. J. Lipid Sci. Technol. 2008, 110, 405–412. [Google Scholar] [CrossRef]
- Traitler, H.; Wille, H.J.; Studer, A. Fractionation of black currant seed oil. J. Am. Oil Chem. Soc. 1988, 65, 755–760. [Google Scholar] [CrossRef]
- Stymne, S.; Stobart, A.K. Biosynthesis of γ-linolenic acid in cotyledons and microsomal preparation of the developing seeds of common borage (Borago officinalis). Biochem. J. 1986, 24, 385–393. [Google Scholar] [CrossRef]
- Hudson, B.J. Evening primrose (Oenothera spp.) oil and seed. J. Am. Oil Chem. Soc. 1984, 61, 540–542. [Google Scholar] [CrossRef]
- Sayanova, O.; Napier, J.A. Transgenic oilseed crops as an alternative do fish oil. Prost. Leukotr. Essent. Fatty Acids 2011, 85, 253–260. [Google Scholar] [CrossRef] [PubMed]
- Ucciani, E. Potential sources of γ-linolenic acid. Ocl-Ol. Corps Gras Lipides 1995, 2, 319–322. [Google Scholar]
- Reddy, A.S.; Nuccio, M.L.; Gross, L.M. Isolation of a Δ6desaturase gene from the cyanobacterium Synechocystis sp. strain PCC6803 by gain-of-function expression in Anabaena sp. strain PCC7120. Plant Mol. Biol. 1993, 22, 293–300. [Google Scholar] [CrossRef] [PubMed]
- Reddy, A.S.; Thomas, T.L. Expression of a cyanobacterial Δ6desaturase gene results in γ-linolenic acid production in transgenic plants. Nat. Biotechnol. 1996, 14, 639–642. [Google Scholar] [CrossRef] [PubMed]
- Sayanova, O.; Smith, M.A.; Lapinskas, P.; Stobart, A.K.; Dobson, G.; Christie, W.W.; Shewry, P.R.; Napier, J.A. Expression of borage desaturase cDNA containing an N-terminal cytochrom b5 domain results in the accumulation of high levels of Δ6desaturated fatty acids in transgenic tobacco. Proc. Natl. Acad. Sci. USA 1997, 94, 4211–4216. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.W.; DeMichele, S.; Bergana, M.; Bobik, E.; Hastilow, C.; Chuang, L.T.; Mukerji, P.; Huang, Y.S. Characterization of oil exhibiting high γ-linolenic acid from a genetically transformed canola strain. J. Am. Oil Chem. Soc. 2001, 78, 489–493. [Google Scholar] [CrossRef]
- García-Maroto, F.; Garrido-Cárdenas, J. A.; Rodríguez-Ruiz, J.; Vilches-Ferrón, M.; Adam, A.C.; Polaina, J.; López Alonso, D. Cloning and molecular characterization of the Δ6desaturase from two Echium plant species: Production of GLA by heterologous expression in yeast and tobacco. Lipids 2002, 37, 417–426. [Google Scholar] [CrossRef] [PubMed]
- Hong, H.; Datla, N.; Reed, D.W.; Covello, P.S.; MacKenzie, S.L.; Qiu, X. High-level production of γ-linolenic acid in Brassica juncea using a Δ6desaturase from Pythium irregulare. Plant Physiol. 2002, 129, 354–362. [Google Scholar] [CrossRef] [PubMed]
- Sato, S.; Xing, A.; Ye, X.; Schweiger, B.; Kinney, A.; Graef, G.; Clemente, T. Production of γ-linolenic acid and stearidoinic acid in seeds of marker-free transgenic soybean. Crop Sci. 2004, 44, 646–652. [Google Scholar] [CrossRef]
- Nykiforuk, C.L.; Shewmaker, C.; Harry, I.; Yurchenko, O.P.; Zhang, M.; Reed, C.; Oinam, G.S.; Zaplachinski, S.; Fidantsef, A.; Boothe, J.G.; et al. High level accumulation of γ-linolenic acid (C18:3Δ6.9,12 cis) in transgenic safflower (Carthamus tinctorius) seeds. Transgenic Res. 2012, 21, 367–381. [Google Scholar] [CrossRef] [PubMed]
- Brenner, R.R. Regulatory function of Δ6desaturase—A key enzyme of polyunsaturated fatty acid synthesis. Adv. Exp. Med. Biol. 1976, 83, 85–101. [Google Scholar]
- Gunstone, F.D. γ-linolenic acid–occurrence and physical and chemical properties. Prog. Lipid Res. 1992, 31, 145–161. [Google Scholar] [CrossRef]
- Certik, M.; Klempova, T.; Guothova, L.; Mihalik, D.; Kraic, J. Biotechnology for the functional improvement of cereal-based materials enriched with PUFAs and pigments. Eur. J. Lipid Sci. Technol. 2013, 115, 1247–1256. [Google Scholar] [CrossRef]
- Pérez-Massot, E.; Banakar, R.; Gómez-Galera, S.; Zorrilla-López, U.; Sanahuja, G.; Arjó, G.; Miralpeix, B.; Vamvaka, E.; Farré, G.; Rivera, S.M.; et al. The contribution of transgenic plants to better health through improved nutrition: Opportunities and constraints. Genes Nutr. 2013, 8, 29–41. [Google Scholar] [CrossRef] [PubMed]
- Wakita, Y.; Otani, M.; Iba, K.; Shimada, T. Co-integration, co-expression and co-segregation of an unlinked selectable marker gene and NtFAD3 gene in transgenic rice plants produced by particle bombardment. Genet. Syst. 1998, 73, 219–226. [Google Scholar] [CrossRef] [Green Version]
- Shimada, T.; Wakita, Y.; Otani, M.; Iba, K. Modification of fatty acid composition in rice plants by transformation with a tobacco microsomal ω-3 fatty acid desaturase gene (NtFAD3). Plant Biotechnol. 2000, 17, 43–48. [Google Scholar] [CrossRef]
- Anai, T.; Koga, M.; Tanaka, H.; Kinoshita, T.; Rahman, S.M.; Takagi, Y. Improvement of rice (Oryza sativa L.) seed oil quality through introduction of a soybean microsomal ω-3 fatty acid desaturase gene. Plant Cell Rep. 2003, 21, 988–992. [Google Scholar] [PubMed]
- Liu, H.L.; Yin, Z.J.; Xiao, L.; Xu, Y.N.; Qu, L.Q. Identification and evaluation of ω-3 fatty acid desaturase genes for hyperfortifying α-linolenic acid in transgenic rice seed. J. Exp. Bot. 2012, 63, 3279–3287. [Google Scholar] [CrossRef] [PubMed]
- Zaplin, E.S.; Liu, Q.; Li, Z.; Butardo, V.M., Jr.; Blanchard, C.L.; Rahman, S. Production of high oleic rice grains by suppressing the expression of the OsFAD2-1 gene. Funct. Plant Biol. 2013, 40, 996–1004. [Google Scholar] [CrossRef]
- Mihálik, D.; Gubišová, M.; Klempová, T.; Čertík, M.; Ondreičková, K.; Hudcovicová, M.; Klčová, L.; Gubiš, J.; Dokupilová, I.; Ohnoutková, L.; et al. Transgenic barley producing essential polyunsaturated fatty acids. Biol. Plant. 2014, 58, 348–354. [Google Scholar] [CrossRef]
- Liu, W.; Steward, C.N., Jr. Plant synthetic biology. Trends Plant Sci. 2015, 20, 309–317. [Google Scholar] [CrossRef] [PubMed]
- Baltes, N.J.; Voytas, D.F. Enabling plant synthetic biology through genome engineering. Trends Biotechnol. 2015, 33, 120–131. [Google Scholar] [CrossRef] [PubMed]
- Zurbriggen, M.D.; Moor, A.; Weber, W. Plant and bacterial systems biology as platform for plant synthetic bio(techno)logy. J. Biotechnol. 2012, 160, 80–90. [Google Scholar] [CrossRef] [PubMed]
- Napier, J.A.; Haslam, R.P.; Beaudoin, F.; Cahoon, E.B. Understanding and manipulating plant lipid composition: Metabolic engineering leads the way. Curr. Opin. Plant Biol. 2014, 19, 68–75. [Google Scholar] [CrossRef] [PubMed]
- Wang, D.; Li, M.; Wei, D.; Cai, Y.; Zhang, Y.; Xing, L. Identification and functional characterization of the Δ6-fatty acid desaturase gene from Thamnidium elegans. J. Eukaryot. Microbiol. 2007, 54, 110–117. [Google Scholar] [CrossRef] [PubMed]
- He, G.Y.; Jones, H.D.; D’Ovidio, R.; Masci, S.; Chen, M.; West, J.; Butow, B.; Anderson, O.D.; Lazzeri, P.; Fido, R.; et al. Expression of an extended HMW subunit in transgenic wheat and the effect on dough mixing properties. J. Cereal Sci. 2005, 42, 225–231. [Google Scholar] [CrossRef]
- Benson, D.A.; Cavanaugh, M.; Clark, K.; Karsch-Mizrachi, I.; Lipman, D.J.; Ostell, J.; Sayers, E.W. GenBank. Nucleic Acid Res. 2013, 41, D36–D42. [Google Scholar] [CrossRef] [PubMed]
- Furtado, A.; Henry, R.J.; Pellegrineschi, A. Analysis of promoters in transgenic barley and wheat. Plant Biotechnol. J. 2009, 7, 240–253. [Google Scholar] [CrossRef] [PubMed]
- Furtado, A.; Henry, R.; Takaiwa, F. Comparison of promoters in transgenic rice. Plant Biotechnol. J. 2008, 6, 679–693. [Google Scholar] [CrossRef] [PubMed]
- Mrízová, K.; Holasková, E.; Tufan Öz, M.; Jiskrová, E.; Frébort, I.; Galuszka, P. Transgenic barley: A prospective tool for biotechnology and agriculture. Biotechnol. Adv. 2014, 32, 137–157. [Google Scholar] [CrossRef] [PubMed]
- Sung, D.-Y.; Kaplan, F.; Lee, K.-J.; Guy, C.L. Acquired tolerance to temperature extrems. Trend Plant Sci. 2003, 8, 179–187. [Google Scholar] [CrossRef]
- Quinn, P.J.; Joo, F.; Vigh, L. The role of unsaturated lipids in membrane structure and stability. Prog. Biophys. Mol. Biol. 1989, 53, 71–103. [Google Scholar] [CrossRef]
- Orsavova, J.; Misurcova, L.; Vavra Ambrozova, J.; Vicha, R.; Mlcek, J. Fatty acids composition of vegetable oils and its contribution to dietary energy intake and dependence of cardiovascular mortality on dietary intake of fatty acids. Int. J. Mol. Sci. 2015, 16, 12871–12890. [Google Scholar] [CrossRef] [PubMed]
- Anderson, O.D.; Greene, F.C.; Yip, R.E.; Halford, N.G.; Shewry, P.R.; Malpica-Romero, J.M. Nucleotide sequences of the two high-molecular weight glutenin genes from the D-genome of a hexaploid bread wheat, Triticum aestivum L. cv Cheyenne. Nucleic Acids Res. 1989, 17, 461–462. [Google Scholar] [CrossRef] [PubMed]
- Puigbó, P.; Guzmán, E.; Romeu, A.; Garcia-Vallvé, S. OPTIMIZER: A web server for optimizing the codon usage of DNA sequences. Nucleic Acids Res. 2007, 35, W126–W131. [Google Scholar] [CrossRef] [PubMed]
- Christensen, A.H.; Quail, P.H. Ubiquitin promoter-based vectors for high-level expression of selectable and/or screenable marker genes in monocotyledonous plants. Transgenic Res. 1996, 5, 213–218. [Google Scholar] [CrossRef] [PubMed]
- Thompson, C.J.; Rao Movva, N.; Tizard, R.; Crameri, R.; Davies, J.E.; Lauwereys, M.; Botterman, J. Characterization of the herbicide-resistance gene bar from Streptomyces hygroscopicus. EMBO J. 1987, 6, 2519–2523. [Google Scholar] [PubMed]
- Gubišová, M.; Mihálik, D.; Konôpková, Ľ. Regeneration efficiency of Slovak spring barley cultivars and winter wheat cultivars. Agriculture 2011, 57, 76–83. [Google Scholar]
- Sparks, C.A.; Jones, H.D. Biolistic transformation of wheat. In Methods in Molecular Biology, Transgenic Wheat, Barley and Oats; Jones, H.D., Shewry, P.R., Eds.; Humana Press: New York, NY, USA, 2009; Volume 478, pp. 71–92. [Google Scholar]
- Matsumoto, T.; Tanaka, T.; Sakai, H.; Amano, N.; Kanamori, H.; Kurita, K.; Kikuta, A.; Kamiya, K.; Yamamoto, M.; Ikawa, H.; Fujii, N.; Hori, K.; Itoh, T.; Sato, K. Comprehensive sequence analysis of 24,783 barley full-length cDNAs derived from 12 clone libraries. Plant Physiol. 2011, 156, 20–28. [Google Scholar] [CrossRef] [PubMed]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and condi–tions of the Creative Commons by Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mihálik, D.; Klčová, L.; Ondreičková, K.; Hudcovicová, M.; Gubišová, M.; Klempová, T.; Čertík, M.; Pauk, J.; Kraic, J. Biosynthesis of Essential Polyunsaturated Fatty Acids in Wheat Triggered by Expression of Artificial Gene. Int. J. Mol. Sci. 2015, 16, 30046-30060. https://doi.org/10.3390/ijms161226137
Mihálik D, Klčová L, Ondreičková K, Hudcovicová M, Gubišová M, Klempová T, Čertík M, Pauk J, Kraic J. Biosynthesis of Essential Polyunsaturated Fatty Acids in Wheat Triggered by Expression of Artificial Gene. International Journal of Molecular Sciences. 2015; 16(12):30046-30060. https://doi.org/10.3390/ijms161226137
Chicago/Turabian StyleMihálik, Daniel, Lenka Klčová, Katarína Ondreičková, Martina Hudcovicová, Marcela Gubišová, Tatiana Klempová, Milan Čertík, János Pauk, and Ján Kraic. 2015. "Biosynthesis of Essential Polyunsaturated Fatty Acids in Wheat Triggered by Expression of Artificial Gene" International Journal of Molecular Sciences 16, no. 12: 30046-30060. https://doi.org/10.3390/ijms161226137
APA StyleMihálik, D., Klčová, L., Ondreičková, K., Hudcovicová, M., Gubišová, M., Klempová, T., Čertík, M., Pauk, J., & Kraic, J. (2015). Biosynthesis of Essential Polyunsaturated Fatty Acids in Wheat Triggered by Expression of Artificial Gene. International Journal of Molecular Sciences, 16(12), 30046-30060. https://doi.org/10.3390/ijms161226137







