A Multiple Interaction Analysis Reveals ADRB3 as a Potential Candidate for Gallbladder Cancer Predisposition via a Complex Interaction with Other Candidate Gene Variations
Abstract
:1. Introduction
2. Results
| Variables | Cases N (%) | Controls N (%) |
|---|---|---|
| Whole Subjects | 400 (100) | 246 (100) |
| Female | 278 (69.5) | 163 (66.3) |
| Male | 122 (30.5) | 83 (33.7) |
| Age ± SD | 52.65 ± 10.45 | 47.75 ± 10.65 |
| Stages | ||
| 0, I | None | NA |
| II | 21 (5.25) | |
| III | 199 (49.75) | |
| IV | 180 (45.0) | |
| Gallstone present | 200 (50.0) | None |
| Gallstone absent | 200 (50.0) | 246 (100) |
| Tobacco | ||
| No | 273 (68.9) | NA |
| Yes | 123 (31.1) |
2.1. Single Locus Analysis
| Pathway | Gene | SNP | MAFcontrols | MAFcases | ORhet a | ORhom a |
|---|---|---|---|---|---|---|
| Death receptor | Dr4 | rs20576 | 8 | 14 | 1.82 (1.18–2.83) | 3.27 (0.93–11.51) |
| rs6557634 | 27 | 33 | 1.61 (1.06–2.44) | 2.05 (0.90–4.70) | ||
| FAS | rs763110 | 39 | 41 | 0.94 (0.66–1.33) | 1.26 (0.78–2.02) | |
| FASL | rs2234767 | 20 | 22 | 0.99 (0.71–1.38) | 1.66 (0.70–4.12) | |
| Tumor suppressor | DCC | rs714 | 37 | 45 | 1.84 (1.29–2.63) | 1.72 (1.08–2.74) |
| rs2229080 | 32 | 24 | 0.64 (0.46–0.89) | 0.32 (0.15–0.68) | ||
| rs7504990 | 32 | 31 | 1.01 (0.72–1.40) | 0.92 (0.51–1.65) | ||
| rs4078288 | 34 | 39 | 0.98 (0.69–1.39) | 1.58 (1.01–2.49) | ||
| Prostate stem cell antigen | PSCA | rs2978974 | 32 | 30 | 0.91 (0.65–1.27) | 0.86 (0.50–1.48) |
| rs2294008 | 42 | 46 | 1.4 (0.97–2.02) | 1.25 (0.77–2.04) | ||
| Adrenergic pathway | ADRa2a | rs1800544 | 45 | 49 | 1.35 (0.92–1.97) | 1.41 (0.87–2.29) |
| ADRB3 | rs4994 | 10 | 21 | 2.58 (1.76–3.78) | 10.61 (1.38–81.92) | |
| ADRB1 | rs1801253 | 22 | 25 | 1.32 (0.95–1.84) | 1.12 (0.46–2.78) | |
| Estrogen metabolism pathway | CYP17 | rs2486758 | 26 | 27 | 1.04 (0.74–1.45) | 1.11 (0.59–2.09) |
2.2. Multifactor Dimensionality Reduction (MDR)
| No. of Risk Factors | Best Interaction Model | Testing Accuracy | # CVC | X² (p-Value) | OR (95% CI) |
|---|---|---|---|---|---|
| 1 | ADRB3rs4994 | 0.6003 | 10/10 | 28.5717 (p < 0.0001) | 2.7507 (1.8841–4.0158) |
| 2 | DCCrs2229080, ADRB3rs4994 | 0.5658 | 6/10 | 32.5889 (p < 0.0001) | 2.6238 (1.8762–3.6693) |
| 3 | DCCrs714, DCCrs2229080, ADRB3rs4994 | 0.5913 | 9/10 | 44.324 (p < 0.0001) | 3.0155 (2.1684–4.1935) |
| 4 | DCCrs714, DCCrs2229080, PSCArs2978974, ADRB3rs4994 | 0.5353 | 3/10 | 68.7203 (p < 0.0001) | 4.0443 (2.8834–5.6726) |
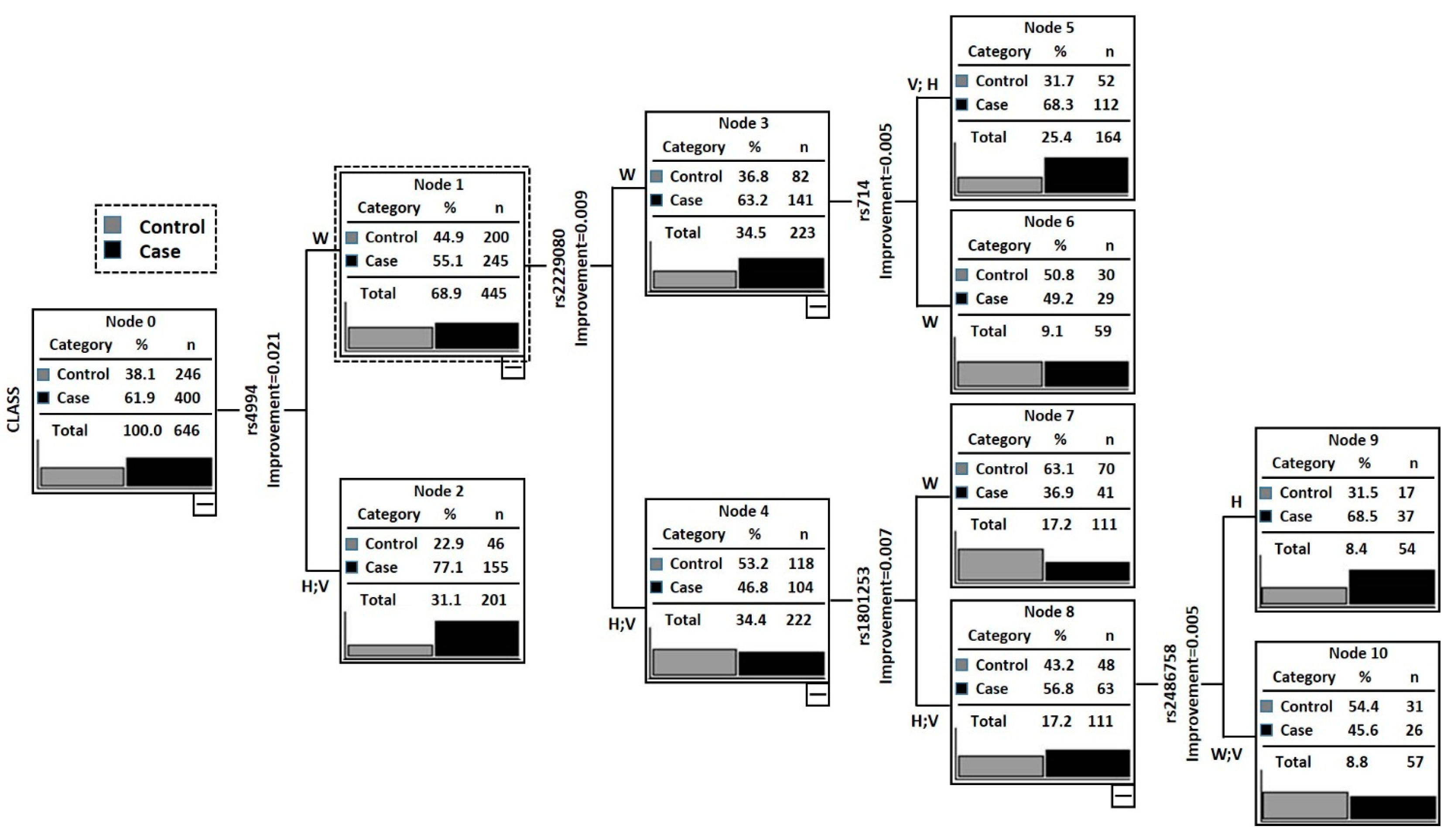
2.3. Classification and Regression Tree Analysis (CRT)
| Nodes | Genotype of Individuals in Each Node | Case | Control | Total | Case Rate (%) | p-Value | OR (95% CI) a |
|---|---|---|---|---|---|---|---|
| Node 1 | ADRB3rs4994 (W) + DCC rs2229080 (H + V) + ADRB1rs1801253 (W) | 41 | 70 | 111 | 36.94 | – | Reference |
| Node 2 | ADRB3rs4994 (W) + DCC rs2229080 (H + V) + ADRB1rs1801253 (H + V) + Cyp17rs2486758 (W + V) | 26 | 31 | 57 | 45.61 | 0.2836 | 1.43 (0.74–2.75) |
| Node 3 | ADRB3rs4994 (W) + DCCrs2229080 (W) + DCCrs714 (W) | 29 | 30 | 59 | 49.15 | 0.1290 | 1.65 (0.87–3.14) |
| Node 4 | ADRB3rs4994 (W) + DCC rs2229080 (W) + DCCrs714 (H + V) | 112 | 52 | 164 | 68.29 | 0.0003 | 3.66 (2.21–6.12) |
| Node 5 | ADRB3rs4994 (W) + DCCrs2229080 (H + V) + ADRB1rs1801253 (H + V) + Cyp17rs2486758 (H) | 37 | 17 | 54 | 68.52 | 0.0001 | 3.69 (1.86–7.50) |
2.4. In-Silico Analysis
| SNPs | Result of F-SNP/FAST SNP | |||
|---|---|---|---|---|
| – | FS Score | Functional Category | Prediction Tool | Prediction Result |
| DR4rs20576 | 0 | Protein coding | Ensemble | Nonsynnymous |
| Polyphen | Possible damaging | |||
| DR4rs6557634 | 0.284 | Protein coding | Ensembl | Nonsynonymous |
| Polyphen | Probably damaging | |||
| Splicing regulation | ESE finder | Changed | ||
| ESR Search | Changed | |||
| FASLrs763110 | 0.434 | Protein coding | Ensembl | Frameshift-coding |
| Transcriptional regulation | TF-Search | Changed | ||
| Ensembl-TR | Regulatory region | |||
| FASrs2234767 | 0 | Protein coding | Ensembl | Nonsynnymous |
| DCCrs2229080 | 0.616 | Protein coding | Polyphen | Probably damaging |
| SNPeffect | Deleterious | |||
| LS-SNP | Deleterious | |||
| Missense (non-conservative) Medium–high (3,4) | ||||
| Splicing regulation | ESE finder | Changed | ||
| ESR Search | Changed | |||
| PESX | Changed | |||
| DCCrs4078288 | NA | Intronic enhancer Very low–low (1–2) | ||
| DCCrs7504990 | NA | Intronic with no known function | ||
| DCCrs714 | NA | Intronic with no known function | ||
| Cyp17rs2486758 | 0.176 | Transcriptional regulation | TFSearch | Changed |
| ADRA2Ars1800544 | 0.065 | Transcriptional regulation | Golden path | Exit |
| ADRB3rs4994 | 0.551 | Protein coding | Ensembl | Nonsynonymous |
| SIFT | Damaging | |||
| SNPeffect | Deleterious | |||
| Splicing regulation | ESE finder | Changed | ||
| ESR Search | Changed | |||
| PESX | Changed | |||
| Transcriptional regulation | Golden path | Exit | ||
| ADRB1rs1800544 | 0.774 | Protein coding | Ensembl | Nonsynonymous |
| CYP17rs2486758 | 0.176 | Transcriptional regulation | TFsearch | Changed |
3. Discussion
4. Materials and Methods
4.1. Ethics Statement and Study Population
4.2. Selected SNPs and Genotyping
5. Statistical Analysis
5.1. Single Locus Analysis
5.2. Multifactor Dimensionality Reduction (MDR)
5.3. Classification and Regression Tree Analysis (CRT)
5.4. In-Silico Analysis and Functional Prediction of SNPs
6. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Dwivedi, A.N.; Jain, S.; Dixit, R. Gall bladder carcinoma: Aggressive malignancy with protean loco-regional and distant spread. World J. Clin. Cases 2015, 3, 231–244. [Google Scholar] [CrossRef] [PubMed]
- Eslick, G.D. Epidemiology of gallbladder cancer. Gastroenterol. Clin. N. Am. 2010, 39, 307–330. [Google Scholar] [CrossRef] [PubMed]
- Alexander, S.; Lemmens, V.E.; Houterman, S.; Nollen, L.; Roumen, R.; Slooter, G.D. Gallbladder cancer, a vanishing disease? Cancer Causes Control 2012, 23, 1705–1709. [Google Scholar] [CrossRef] [PubMed]
- Rai, R.; Tewari, M.; Kumar, M.; Singh, T.B.; Shukla, H.S. Expression profile of cholecystokinin type-a receptor in gallbladder cancer and gallstone disease. Hepatobiliary Pancreat. Dis. Int. 2011, 10, 408–414. [Google Scholar] [CrossRef]
- Kanthan, R.; Senger, J.L.; Ahmed, S.; Kanthan, S.C. Gallbladder cancer in the 21st century. J. Oncol. 2015, 2015, 967472. [Google Scholar] [CrossRef] [PubMed]
- Hundal, R.; Shaffer, E.A. Gallbladder cancer: Epidemiology and outcome. Clin. Epidemiol. 2014, 6, 99–109. [Google Scholar] [PubMed]
- Rai, R.; Tewari, M.; Kumar, M.; Singh, A.K.; Shukla, H.S. P53: Its alteration and gallbladder cancer. Eur. J. Cancer Prev. 2011, 20, 77–85. [Google Scholar] [CrossRef] [PubMed]
- Rai, R.; Sharma, K.L.; Misra, S.; Kumar, A.; Mittal, B. CYP17 polymorphism (rs743572) is associated with increased risk of gallbladder cancer in tobacco users. Tumour Biol. 2014, 35, 6531–6537. [Google Scholar] [CrossRef] [PubMed]
- Rai, R.; Sharma, K.L.; Sharma, S.; Misra, S.; Kumar, A.; Mittal, B. Death receptor (DR4) haplotypes are associated with increased susceptibility of gallbladder carcinoma in North Indian population. PLoS ONE 2014, 9, e90264. [Google Scholar] [CrossRef] [PubMed]
- Rai, R.; Sharma, K.L.; Misra, S.; Kumar, A.; Mittal, B. Association of adrenergic receptor gene polymorphisms in gallbladder cancer susceptibility in a north Indian population. J. Cancer Res. Clin. Oncol. 2014, 140, 725–735. [Google Scholar] [CrossRef] [PubMed]
- Rai, R.; Sharma, K.L.; Misra, S.; Kumar, A.; Mittal, B. Psca gene variants (rs2294008 and rs2978974) confer increased susceptibility of gallbladder carcinoma in females. Gene 2013, 530, 172–177. [Google Scholar] [CrossRef] [PubMed]
- Rai, R.; Sharma, K.L.; Tiwari, S.; Misra, S.; Kumar, A.; Mittal, B. Dcc (deleted in colorectal carcinoma) gene variants confer increased susceptibility to gallbladder cancer (ref. No.: Gene-d-12-01446). Gene 2013, 518, 303–309. [Google Scholar] [CrossRef] [PubMed]
- Umar, M.; Upadhyay, R.; Mittal, B. PLCE1 rs2274223 A>G polymorphism and cancer risk: A meta-analysis. Tumour Biol. 2013, 34, 3537–3544. [Google Scholar] [CrossRef] [PubMed]
- Sharma, K.L.; Rai, R.; Srivastava, A.; Sharma, A.; Misra, S.; Kumar, A.; Mittal, B. A multigenic approach to evaluate genetic variants of PLCE1, LXRs, MMPs, TIMP, and CYP genes in gallbladder cancer predisposition. Tumour Biol. 2014, 35, 8597–8606. [Google Scholar] [CrossRef] [PubMed]
- Hahn, L.W.; Ritchie, M.D.; Moore, J.H. Multifactor dimensionality reduction software for detecting gene–gene and gene-environment interactions. Bioinformatics 2003, 19, 376–382. [Google Scholar] [CrossRef] [PubMed]
- Ritchie, M.D.; Hahn, L.W.; Roodi, N.; Bailey, L.R.; Dupont, W.D.; Parl, F.F.; Moore, J.H. Multifactor-dimensionality reduction reveals high-order interactions among estrogen-metabolism genes in sporadic breast cancer. Am. J. Hum. Genet. 2001, 69, 138–147. [Google Scholar] [CrossRef] [PubMed]
- Lunetta, K.L.; Hayward, L.B.; Segal, J.; van Eerdewegh, P. Screening large-scale association study data: Exploiting interactions using random forests. BMC Genet. 2004, 5, 32. [Google Scholar] [CrossRef] [PubMed]
- Piper, M.E.; Loh, W.Y.; Smith, S.S.; Japuntich, S.J.; Baker, T.B. Using decision tree analysis to identify risk factors for relapse to smoking. Subst. Use Misuse 2011, 46, 492–510. [Google Scholar] [CrossRef] [PubMed]
- Srivastava, A.; Sharma, K.L.; Srivastava, N.; Misra, S.; Mittal, B. Significant role of estrogen and progesterone receptor sequence variants in gallbladder cancer predisposition: A multi-analytical strategy. PLoS ONE 2012, 7, e40162. [Google Scholar] [CrossRef] [PubMed]
- Ferrer-Lorente, R.; Cabot, C.; Fernandez-Lopez, J.A.; Alemany, M. Combined effects of oleoyl-estrone and a β3-adrenergic agonist (cl316,243) on lipid stores of diet-induced overweight male wistar rats. Life Sci. 2005, 77, 2051–2058. [Google Scholar] [CrossRef] [PubMed]
- Anthony, A.; Schepelmann, S.; Guillaume, J.L.; Strosberg, A.D.; Dhillon, A.P.; Pounder, R.E.; Wakefield, A.J. Localization of the β(β)3-adrenoceptor in the human gastrointestinal tract: An immunohistochemical study. Aliment. Pharmacol. Ther. 1998, 12, 519–525. [Google Scholar] [CrossRef] [PubMed]
- Krief, S.; Lonnqvist, F.; Raimbault, S.; Baude, B.; van Spronsen, A.; Arner, P.; Strosberg, A.D.; Ricquier, D.; Emorine, L.J. Tissue distribution of β3-adrenergic receptor mRNA in man. J. Clin. Investig. 1993, 91, 344–349. [Google Scholar] [CrossRef] [PubMed]
- Ochoa, M.C.; Marti, A.; Azcona, C.; Chueca, M.; Oyarzabal, M.; Pelach, R.; Patino, A.; Moreno-Aliaga, M.J.; Martinez-Gonzalez, M.A.; Martinez, J.A.; et al. Gene–gene interaction between PPARγ2 and ADRβ3 increases obesity risk in children and adolescents. Int. J. Obes. 2004, 28, S37–S41. [Google Scholar] [CrossRef] [PubMed]
- Tan, W.; Gao, M.; Liu, N.; Zhang, G.; Xu, T.; Cui, W. Body mass index and risk of gallbladder cancer: Systematic review and meta-analysis of observational studies. Nutrients 2015, 7, 8321–8334. [Google Scholar] [CrossRef] [PubMed]
- Takezaki, T.; Hamajima, N.; Matsuo, K.; Tanaka, R.; Hirai, T.; Kato, T.; Ohashi, K.; Tajima, K. Association of polymorphisms in the β-2 and β-3 adrenoceptor genes with risk of colorectal cancer in Japanese. Int. J. Clin. Oncol. 2001, 6, 117–122. [Google Scholar] [CrossRef] [PubMed]
- Klass, D.M.; Lauer, N.; Hay, B.; Kratzer, W.; Fuchs, M.; Group, E.S. Arg64 variant of the β3-adrenergic receptor is associated with gallstone formation. Am. J. Gastroenterol. 2007, 102, 2482–2487. [Google Scholar] [CrossRef] [PubMed]
- Srivastava, A.; Mishra, A.; Singh, R.; Rai, R.; Srivastava, N.; Mittal, B. Multi-analytic approach elucidates significant role of hormonal and hepatocanalicular transporter genetic variants in gallstone disease in north indian population. PLoS ONE 2013, 8, e59173. [Google Scholar] [CrossRef] [PubMed]
- Kuruma, S.; Egawa, N.; Kurata, M.; Honda, G.; Kamisawa, T.; Ueda, J.; Ishii, H.; Ueno, M.; Nakao, H.; Mori, M.; et al. Case-control study of diabetes-related genetic variants and pancreatic cancer risk in Japan. World J. Gastroenterol. 2014, 20, 17456–17462. [Google Scholar] [CrossRef] [PubMed]
- Fearon, E.R.; Cho, K.R.; Nigro, J.M.; Kern, S.E.; Simons, J.W.; Ruppert, J.M.; Hamilton, S.R.; Preisinger, A.C.; Thomas, G.; Kinzler, K.W.; et al. Identification of a chromosome 18q gene that is altered in colorectal cancers. Science 1990, 247, 49–56. [Google Scholar] [CrossRef] [PubMed]
- Forcet, C.; Ye, X.; Granger, L.; Corset, V.; Shin, H.; Bredesen, D.E.; Mehlen, P. The dependence receptor DCC (deleted in colorectal cancer) defines an alternative mechanism for caspase activation. Proc. Natl. Acad. Sci. USA 2001, 98, 3416–3421. [Google Scholar] [CrossRef] [PubMed]
- Arakawa, H. Netrin-1 and its receptors in tumorigenesis. Nat. Rev. Cancer 2004, 4, 978–987. [Google Scholar] [CrossRef] [PubMed]
- Inokuchi, K.; Yamaguchi, H.; Hanawa, H.; Tanosaki, S.; Nakamura, K.; Tarusawa, M.; Miyake, K.; Shimada, T.; Dan, K. Loss of DCC gene expression is of prognostic importance in acute myelogenous leukemia. Clin. Cancer Res. 2002, 8, 1882–1888. [Google Scholar] [PubMed]
- Sun, X.F.; Rutten, S.; Zhang, H.; Nordenskjold, B. Expression of the deleted in colorectal cancer gene is related to prognosis in DNA diploid and low proliferative colorectal adenocarcinoma. J. Clin. Oncol. 1999, 17, 1745–1750. [Google Scholar] [PubMed]
- Yoshida, Y.; Itoh, F.; Endo, T.; Hinoda, Y.; Imai, K. Decreased dcc mRNA expression in human gastric cancers is clinicopathologically significant. Int. J. Cancer 1998, 79, 634–639. [Google Scholar] [CrossRef]
- Bamias, A.T.; Bai, M.C.; Agnantis, N.J.; Michael, M.C.; Alamanos, Y.P.; Stefanaki, S.V.; Razi, E.D.; Skarlos, D.V.; Kappas, A.M.; Pavlidis, N.A. Prognostic significance of the deleted in colorectal cancer gene protein expression in high-risk resected gastric carcinoma. Cancer Investig. 2003, 21, 333–340. [Google Scholar] [CrossRef]
- Toma, M.; Stavarachi, M.; Cimponeriu, D.; Apostol, P.; Cojocaru, M.; Belusicaa, L.; Panduru, N.; Radu, I.; Gavrila, L. P53 and DCC polymorphisms and the risk for colorectal cancer in Romanian patients—A preliminary study. J. Analele Univ. Oradea Fasc. Biol. 2009, 16, 162–165. [Google Scholar]
- Djansugurova, L.; Zhunussova, G.; Khussainova, E.; Iksan, O.; Afonin, G.; Kaidarova, D.; Parker, M.I. Association of DCC, MLH1, GSTT1, GSTM1, and TP53 gene polymorphisms with colorectal cancer in Kazakhstan. Tumour Biol. 2015, 36, 279–289. [Google Scholar] [CrossRef] [PubMed]
- Starinsky, S.; Figer, A.; Ben-Asher, E.; Geva, R.; Flex, D.; Fidder, H.H.; Zidan, J.; Lancet, D.; Friedman, E. Genotype phenotype correlations in israeli colorectal cancer patients. Int. J. Cancer 2005, 114, 58–73. [Google Scholar] [CrossRef] [PubMed]
- Malik, M.A.; Gupta, A.; Zargar, S.A.; Mittal, B. Role of genetic variants of deleted in colorectal carcinoma (DCC) polymorphisms and esophageal and gastric cancers risk in kashmir valley and meta-analysis. Tumour Biol. 2013, 34, 3049–3057. [Google Scholar] [CrossRef] [PubMed]
- Wistuba, II; Albores-Saavedra, J. Genetic abnormalities involved in the pathogenesis of gallbladder carcinoma. J. Hepatobiliary Pancreat. Surg. 1999, 6, 237–244. [Google Scholar]
- Cha, P.C.; Zembutsu, H.; Takahashi, A.; Kubo, M.; Kamatani, N.; Nakamura, Y. A genome-wide association study identifies SNP in DCC is associated with gallbladder cancer in the Japanese population. J. Hum. Genet. 2012, 57, 235–237. [Google Scholar] [CrossRef] [PubMed]
- Mattar, R.; Nonogaki, S.; Silva, C.; Alves, V.; Gama-Rodrigues, J.J. P53 and RB tumor suppressor gene alterations in gastric cancer. Rev. Hosp. Clin. 2004, 59, 172–180. [Google Scholar] [CrossRef]
- Enomoto, T.; Fujita, M.; Cheng, C.; Nakashima, R.; Ozaki, M.; Inoue, M.; Nomura, T. Loss of expression and loss of heterozygosity in the DCC gene in neoplasms of the human female reproductive tract. Br. J. Cancer 1995, 71, 462–467. [Google Scholar] [CrossRef] [PubMed]
- Minami, R.; Aoyama, N.; Honsako, Y.; Kasuga, M.; Fujimori, T.; Maeda, S. Codon 201Arg/Gly polymorphism of DCC (deleted in colorectal carcinoma) gene in flat- and polypoid-type colorectal tumors. Dig. Dis. Sci. 1997, 42, 2446–2452. [Google Scholar] [CrossRef] [PubMed]
- Kong, X.T.; Choi, S.H.; Bessho, F.; Kobayashi, M.; Hanada, R.; Yamamoto, K.; Hayashi, Y. Codon 201 (Gly) polymorphic type of the DCC gene is related to disseminated neuroblastoma. Neoplasia 2001, 3, 267–272. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Arbman, G.; Sun, X.F. Codon 201 polymorphism of DCC gene is a prognostic factor in patients with colorectal cancer. Cancer Detect. Prev. 2003, 27, 216–221. [Google Scholar] [CrossRef]
- Kupcinskas, J.; Gyvyte, U.; Bruzaite, I.; Leja, M.; Kupcinskaite-Noreikiene, R.; Pauzas, H.; Tamelis, A.; Jonaitis, L.; Skieceviciene, J.; Kiudelis, G. Common genetic variants of PSCA, MUC1 and PLCE1 genes are not associated with colorectal cancer. Asian Pac. J. Cancer Prev. 2015, 16, 6027–6032. [Google Scholar] [CrossRef] [PubMed]
- Geng, P.; Li, J.; Wang, N.; Ou, J.; Xie, G.; Liu, C.; Zhao, X.; Xiang, L.; Liao, Y.; Liang, H. PSCA rs2294008 polymorphism with increased risk of cancer. PLoS ONE 2015, 10, e0136269. [Google Scholar] [CrossRef] [PubMed]
- Zou, Q.; Yang, L.; Yang, Z.; Huang, J.; Fu, X. PSCA and OCT-4 expression in the benign and malignant lesions of gallbladder: Implication for carcinogenesis, progression, and prognosis of gallbladder adenocarcinoma. BioMed Res. Int. 2013, 2013, 648420. [Google Scholar] [CrossRef] [PubMed]
- Saeki, N.; Ono, H.; Sakamoto, H.; Yoshida, T. Down-regulation of immune-related genes by PSCA in gallbladder cancer cells implanted into mice. Anticancer Res. 2015, 35, 2619–2625. [Google Scholar] [PubMed]
- Ono, H.; Chihara, D.; Chiwaki, F.; Yanagihara, K.; Sasaki, H.; Sakamoto, H.; Tanaka, H.; Yoshida, T.; Saeki, N.; Matsuo, K. Missense allele of a single nucleotide polymorphism rs2294008 attenuated antitumor effects of prostate stem cell antigen in gallbladder cancer cells. J. Carcinog. 2013, 12, 4. [Google Scholar] [CrossRef] [PubMed]
- Ono, H.; Hiraoka, N.; Lee, Y.S.; Woo, S.M.; Lee, W.J.; Choi, I.J.; Saito, A.; Yanagihara, K.; Kanai, Y.; Ohnami, S.; et al. Prostate stem cell antigen, a presumable organ-dependent tumor suppressor gene, is down-regulated in gallbladder carcinogenesis. Genes Chromosomes Cancer 2012, 51, 30–41. [Google Scholar] [CrossRef] [PubMed]
- Fu, Y.P.; Kohaar, I.; Rothman, N.; Earl, J.; Figueroa, J.D.; Ye, Y.; Malats, N.; Tang, W.; Liu, L.; Garcia-Closas, M.; et al. Common genetic variants in the PSCA gene influence gene expression and bladder cancer risk. Proc. Natl. Acad. Sci. USA 2012, 109, 4974–4979. [Google Scholar] [CrossRef] [PubMed]
- Qiao, L.; Feng, Y. Genetic variations of prostate stem cell antigen (PSCA) contribute to the risk of gastric cancer for eastern Asians: A meta-analysis based on 16792 individuals. Gene 2012, 493, 83–91. [Google Scholar] [CrossRef] [PubMed]
- Wu, X.; Ye, Y.; Kiemeney, L.A.; Sulem, P.; Rafnar, T.; Matullo, G.; Seminara, D.; Yoshida, T.; Saeki, N.; Andrew, A.S.; et al. Genetic variation in the prostate stem cell antigen gene PSCA confers susceptibility to urinary bladder cancer. Nat. Genet. 2009, 41, 991–995. [Google Scholar] [CrossRef] [PubMed]
- Matsuo, K.; Tajima, K.; Suzuki, T.; Kawase, T.; Watanabe, M.; Shitara, K.; Misawa, K.; Ito, S.; Sawaki, A.; Muro, K.; et al. Association of prostate stem cell antigen gene polymorphisms with the risk of stomach cancer in Japanese. Int. J. Cancer 2009, 125, 1961–1964. [Google Scholar] [CrossRef] [PubMed]
- Kohaar, I.; Porter-Gill, P.; Lenz, P.; Fu, Y.P.; Mumy, A.; Tang, W.; Apolo, A.B.; Rothman, N.; Baris, D.; Schned, A.R.; et al. Genetic variant as a selection marker for anti-prostate stem cell antigen immunotherapy of bladder cancer. J. Natl. Cancer Inst. 2013, 105, 69–73. [Google Scholar] [CrossRef] [PubMed]
- Saeki, N.; Ono, H.; Yanagihara, K.; Aoyagi, K.; Sasaki, H.; Sakamoto, H.; Yoshida, T. Rs2294008t, a risk allele for gastric and gallbladder cancers, suppresses the PSCA promoter by recruiting the transcription factor YY1. Genes Cells Devoted Mol. Cell. Mech. 2015, 20, 382–391. [Google Scholar] [CrossRef] [PubMed]
- Zhang, T.; Chen, Y.N.; Wang, Z.; Chen, J.Q.; Huang, S. Effect of PSCA gene polymorphisms on gastric cancer risk and survival prediction: A meta-analysis. Exp. Ther. Med. 2012, 4, 158–164. [Google Scholar] [PubMed]
- Miller, S.A.; Dykes, D.D.; Polesky, H.F. A simple salting out procedure for extracting DNA from human nucleated cells. Nucleic Acids Res. 1988, 16, 1215. [Google Scholar] [CrossRef] [PubMed]
- MDR software. Available online: www.multifactordimensionalityreduction.org (accessed on16 October 2015).
- Yang, C.H.; Lin, Y.D.; Yang, C.S.; Chuang, L.Y. An efficiency analysis of high-order combinations of gene-gene interactions using multifactor-dimensionality reduction. BMC Genom. 2015, 16, 489. [Google Scholar] [CrossRef] [PubMed]
- Lee, P.H.; Shatkay, H. F-SNP: Computationally predicted functional SNPs for disease association studies. Nucleic Acids Res. 2008, 36, 820–824. [Google Scholar] [CrossRef] [PubMed]
- Yuan, H.Y.; Chiou, J.J.; Tseng, W.H.; Liu, C.H.; Liu, C.K.; Lin, Y.J.; Wang, H.H.; Yao, A.; Chen, Y.T.; Hsu, C.N. FASTSNP: An always up-to-date and extendable service for SNP function analysis and prioritization. Nucleic Acids Res. 2006, 34, 635–641. [Google Scholar] [CrossRef] [PubMed]
- FASTSNP. Available online: http://fastsnp.ibms.sinica.edu.tw (accessed on16 October 2015).
- F-SNP. Available online: http://compbio.cs.queensu.ca/F-SNP/ (accessed on 16 October 2015).
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons by Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rai, R.; Kim, J.J.; Misra, S.; Kumar, A.; Mittal, B. A Multiple Interaction Analysis Reveals ADRB3 as a Potential Candidate for Gallbladder Cancer Predisposition via a Complex Interaction with Other Candidate Gene Variations. Int. J. Mol. Sci. 2015, 16, 28038-28049. https://doi.org/10.3390/ijms161226077
Rai R, Kim JJ, Misra S, Kumar A, Mittal B. A Multiple Interaction Analysis Reveals ADRB3 as a Potential Candidate for Gallbladder Cancer Predisposition via a Complex Interaction with Other Candidate Gene Variations. International Journal of Molecular Sciences. 2015; 16(12):28038-28049. https://doi.org/10.3390/ijms161226077
Chicago/Turabian StyleRai, Rajani, Jong Joo Kim, Sanjeev Misra, Ashok Kumar, and Balraj Mittal. 2015. "A Multiple Interaction Analysis Reveals ADRB3 as a Potential Candidate for Gallbladder Cancer Predisposition via a Complex Interaction with Other Candidate Gene Variations" International Journal of Molecular Sciences 16, no. 12: 28038-28049. https://doi.org/10.3390/ijms161226077
APA StyleRai, R., Kim, J. J., Misra, S., Kumar, A., & Mittal, B. (2015). A Multiple Interaction Analysis Reveals ADRB3 as a Potential Candidate for Gallbladder Cancer Predisposition via a Complex Interaction with Other Candidate Gene Variations. International Journal of Molecular Sciences, 16(12), 28038-28049. https://doi.org/10.3390/ijms161226077





