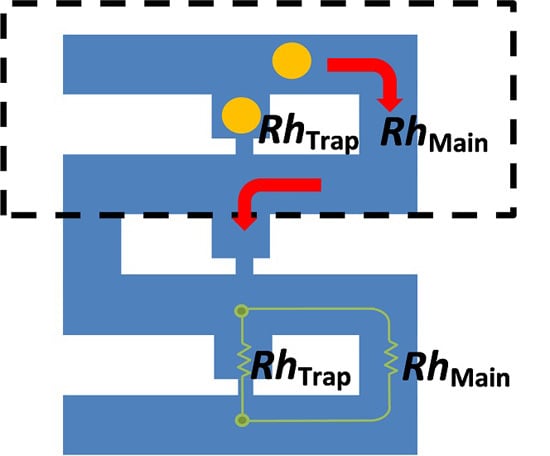
Numerical Analysis of Hydrodynamic Flow in Microfluidic Biochip for Single-Cell Trapping Application
Abstract
:1. Introduction
2. Results and Discussion
2.1. Verification of Hydrodynamic Trapping Concept
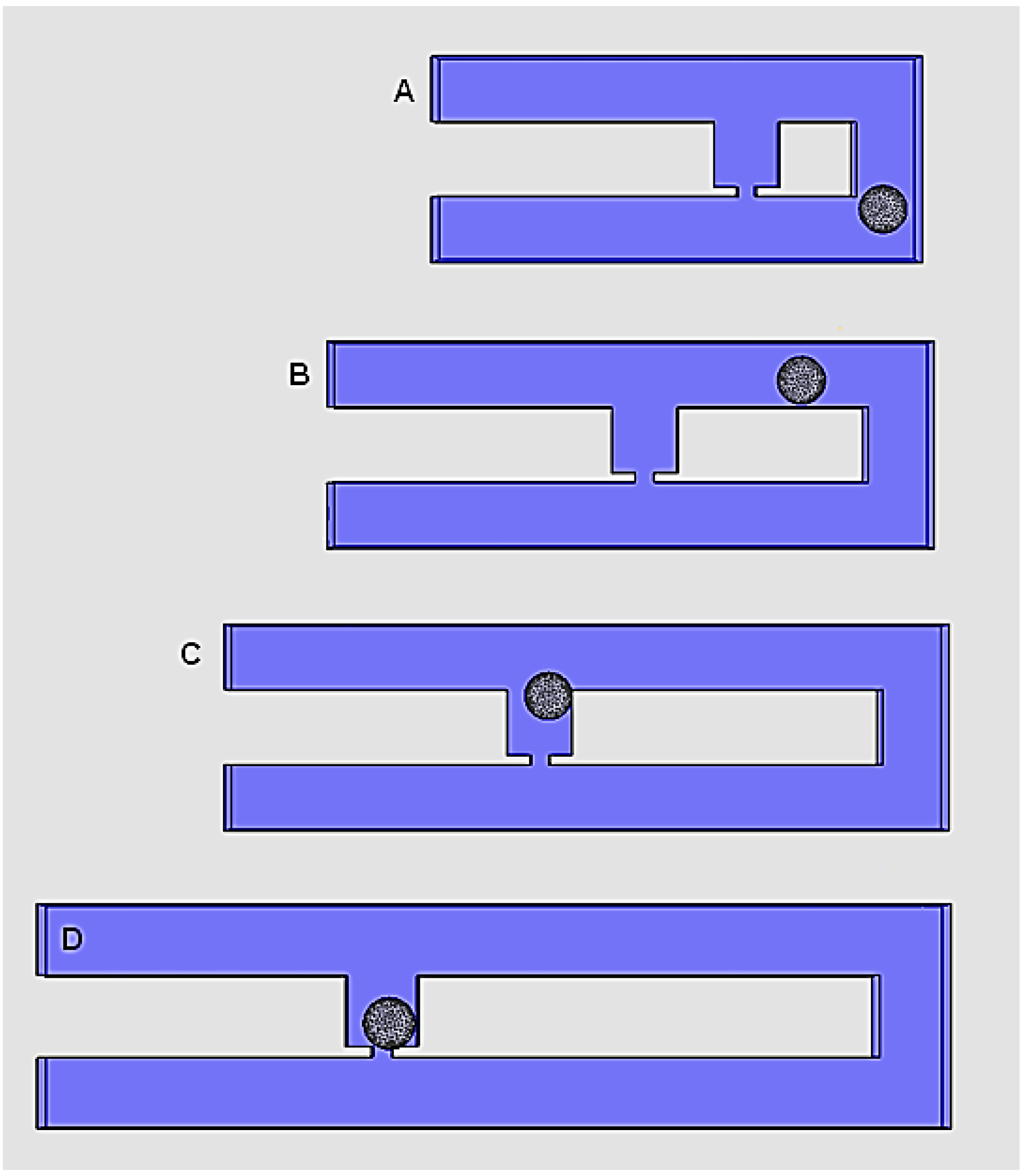
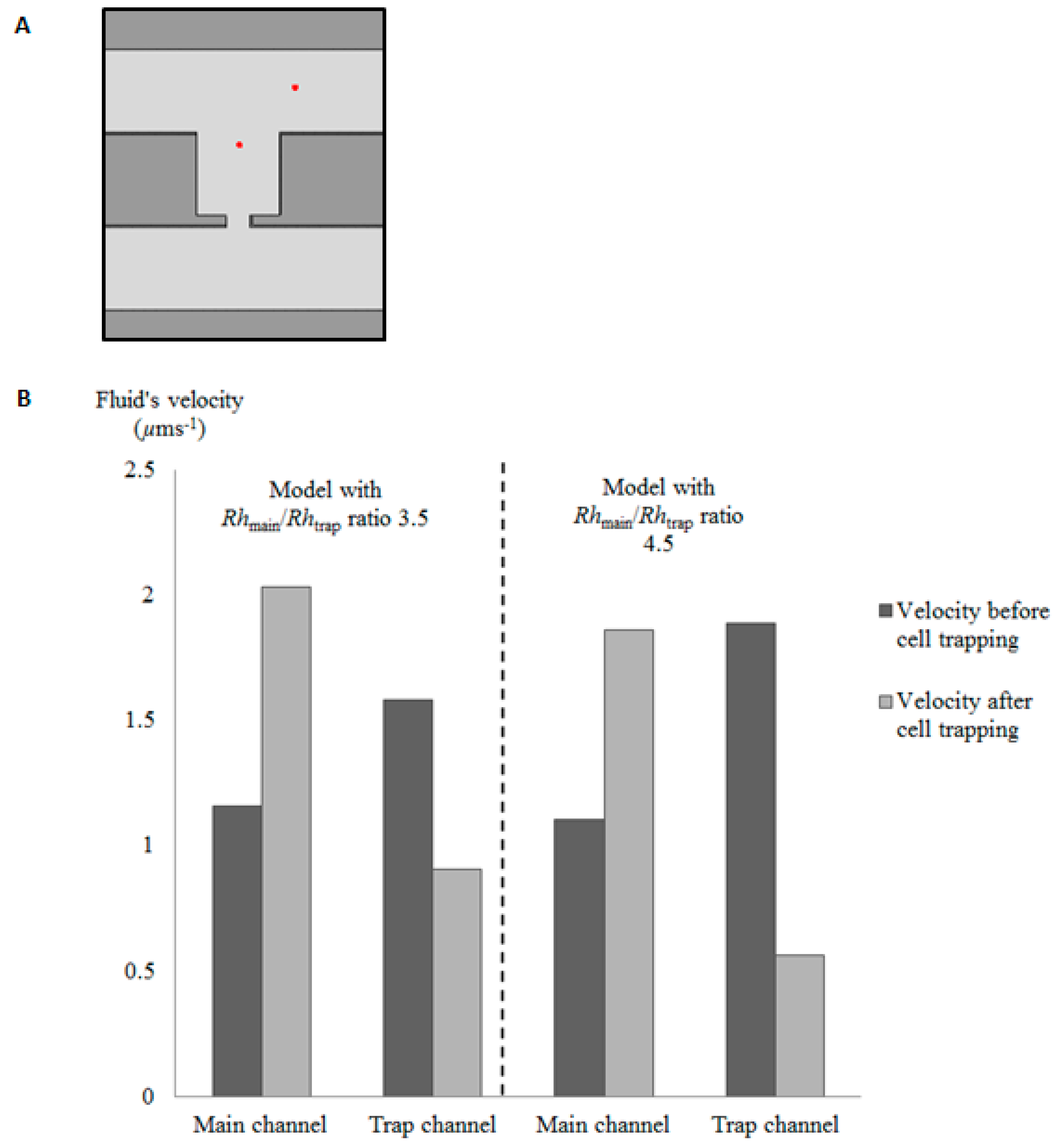
2.2. Effects of RhMain/RhTrap Ratio in Cell Trapping
| Ratio of RhMain/RhTrap | Ability to Trap Cells | |||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| WHole: 1.0 μm | WHole: 1.5 μm | WHole: 2.0 μm | WHole: 2.5 μm | WHole: 3.0 μm | WHole: 3.5 μm | HChan: 6.0 μm | HChan: 7.0 μm | HChan: 8.0 μm | HChan: 9.0 μm | LTrap: 3.0 μm | LTrap: 5.0 μm | LTrap: 7.0 μm | LTrap: 9.0 μm | |
| 1.0 | no | no | no | no | no | no | no | no | no | no | no | no | no | no |
| 1.5 | no | no | no | no | no | no | no | no | no | no | no | no | no | no |
| 2.0 | no | no | no | no | no | no | no | no | no | no | no | no | no | no |
| 2.5 | no | no | no | no | no | no | no | no | no | no | no | no | no | no |
| 3.0 | no | no | no | no | no | no | no | no | no | no | no | no | no | no |
| 3.5 | no | yes | yes | yes | yes | yes | yes | yes | yes | yes | yes | yes | yes | yes |
| 4.5 | no | yes | yes | yes | yes | yes | yes | yes | yes | yes | yes | yes | yes | yes |
| 5.0 | no | yes | yes | yes | yes | yes | yes | yes | yes | yes | yes | yes | yes | yes |
| 5.5 | no | yes | yes | yes | yes | yes | yes | yes | yes | yes | yes | yes | yes | yes |
| 6.0 | no | yes | yes | yes | yes | yes | yes | yes | yes | yes | yes | yes | yes | yes |
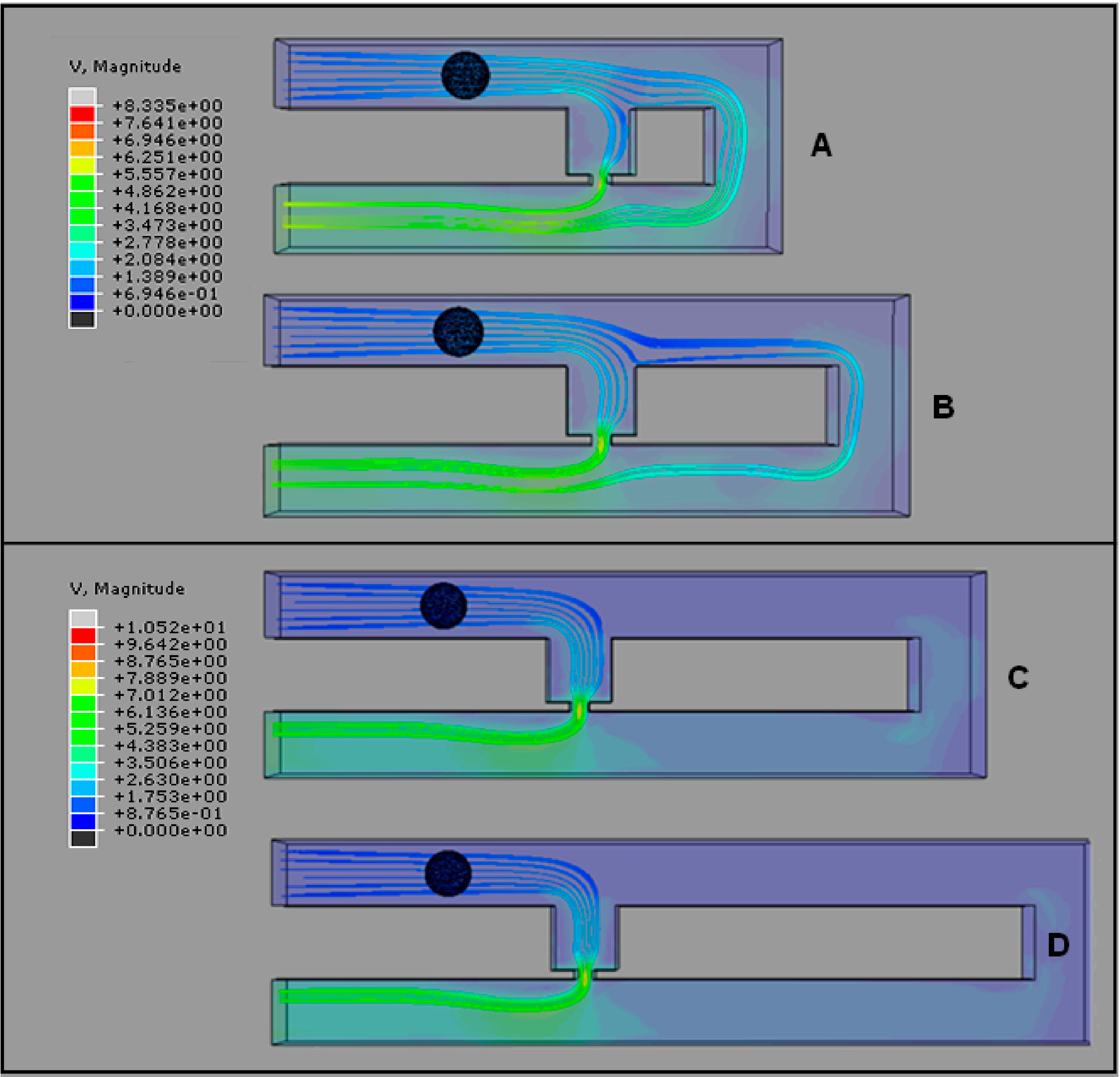
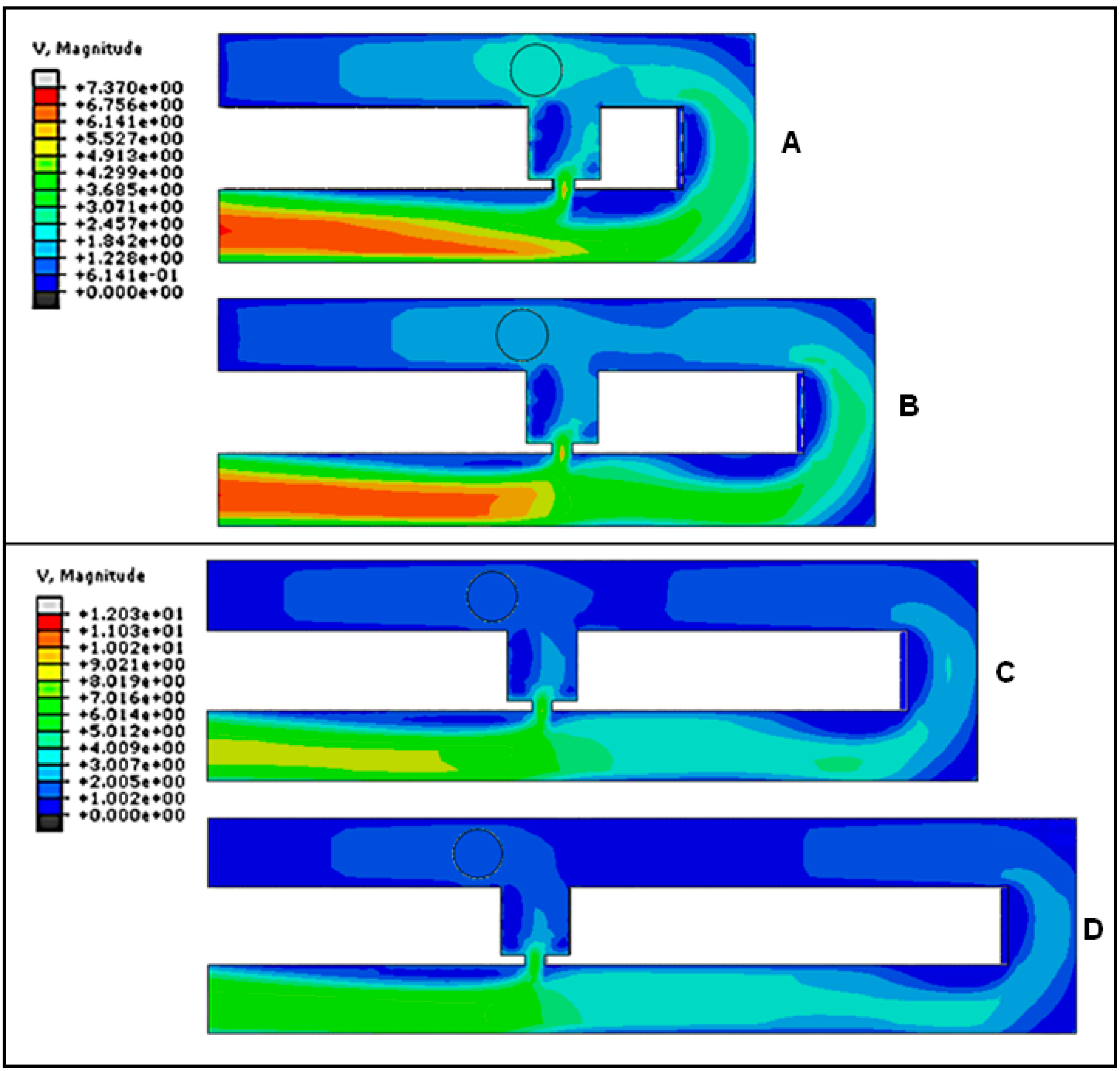
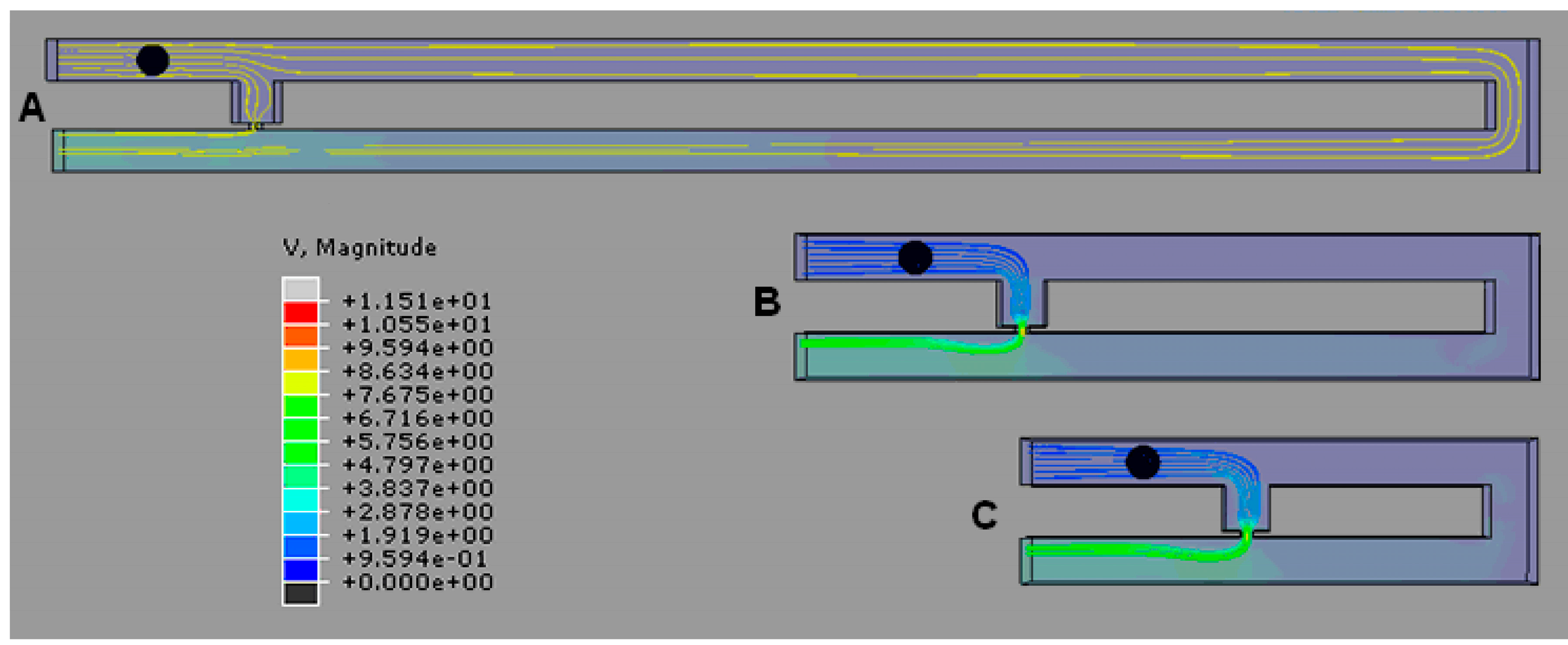
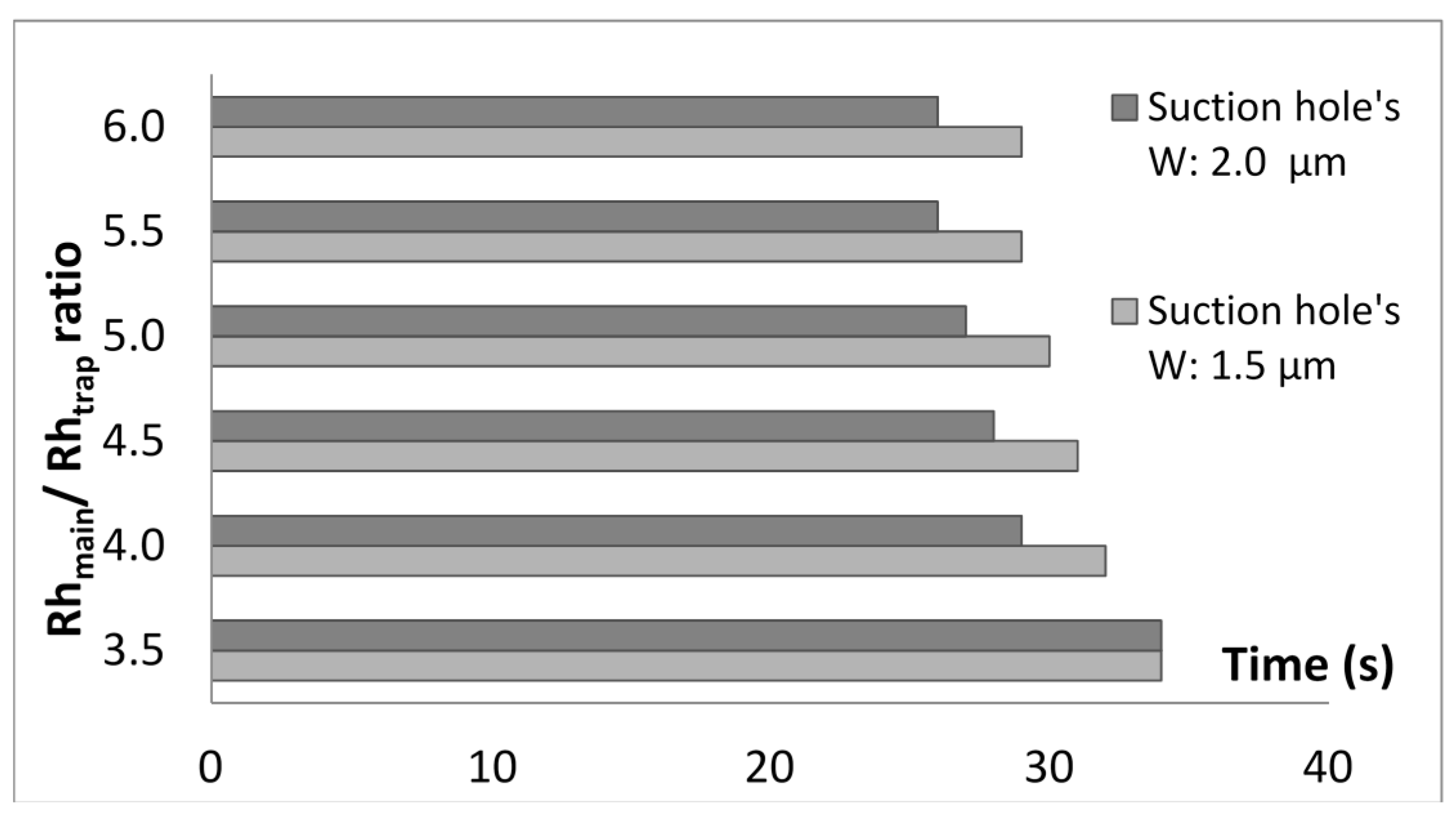
2.3. Optimization of Trap Channel’s Length
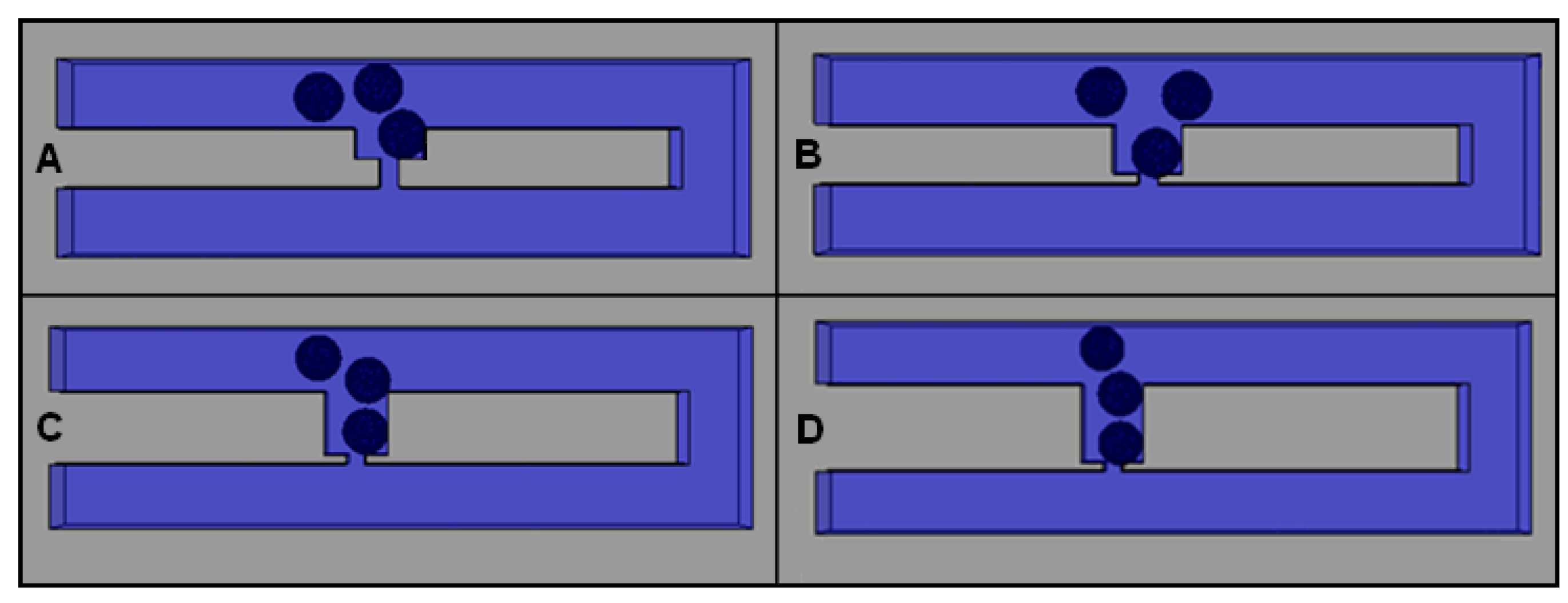
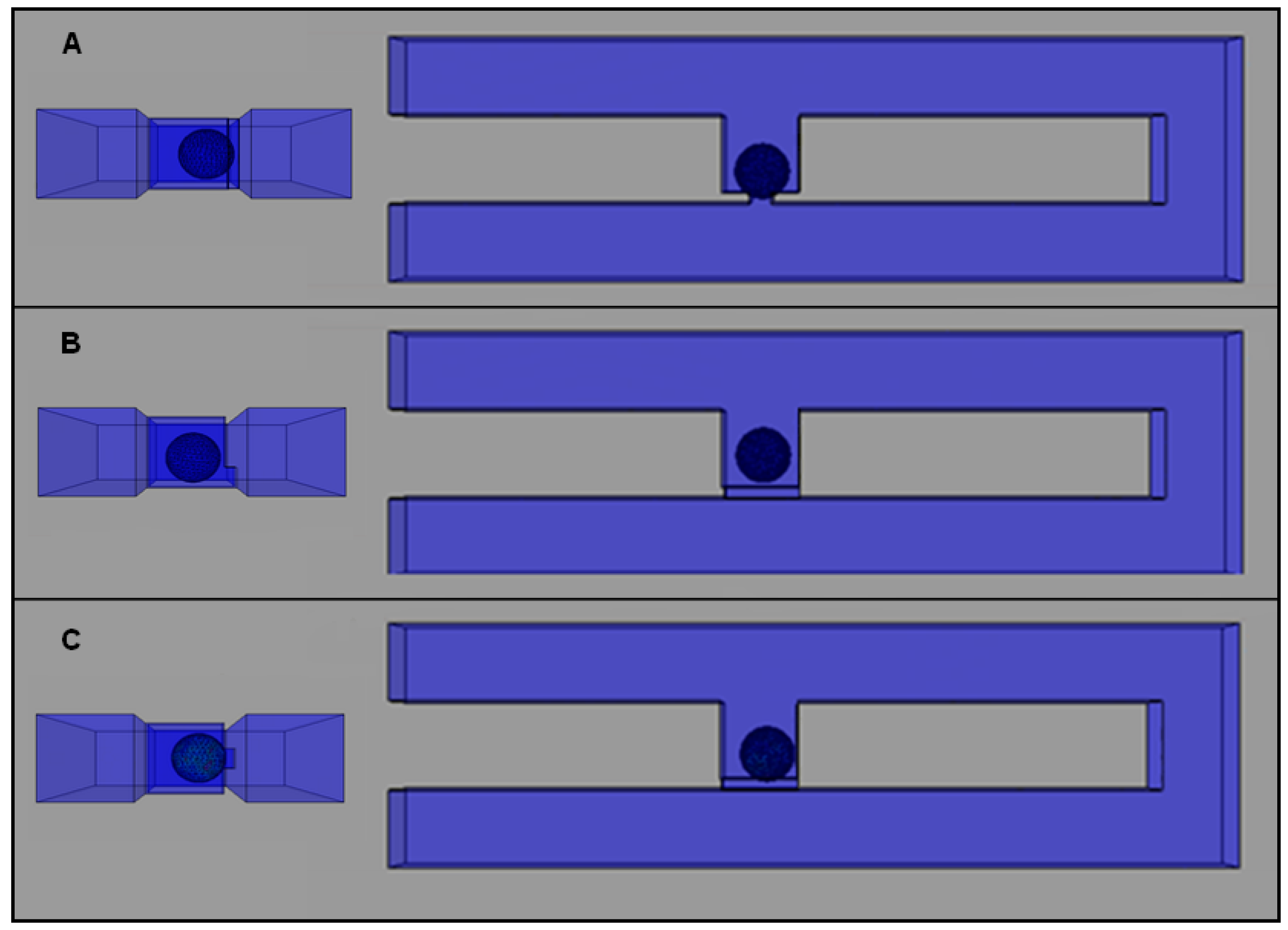
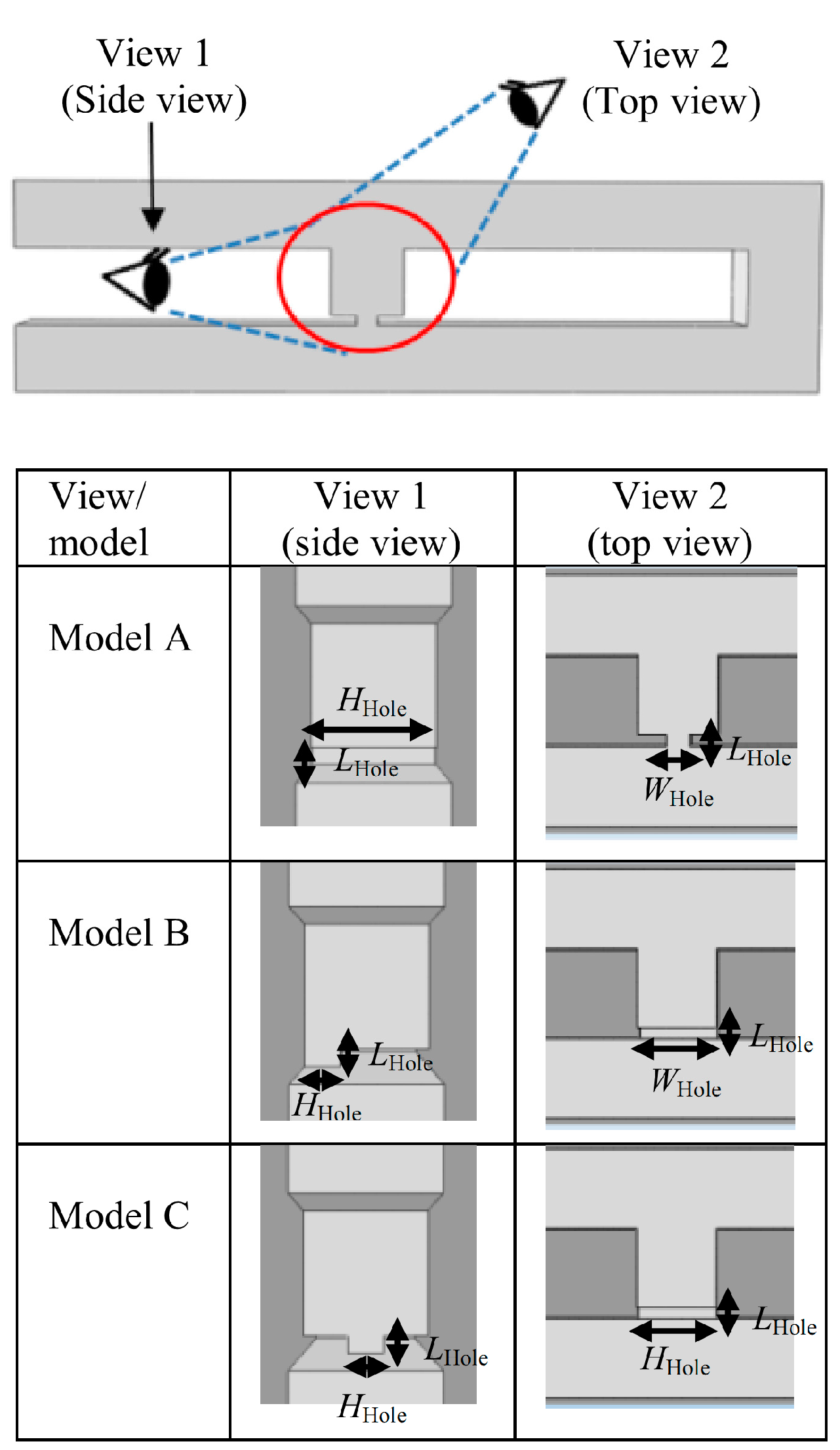
2.4. Effects of Different Trap Hole Positions
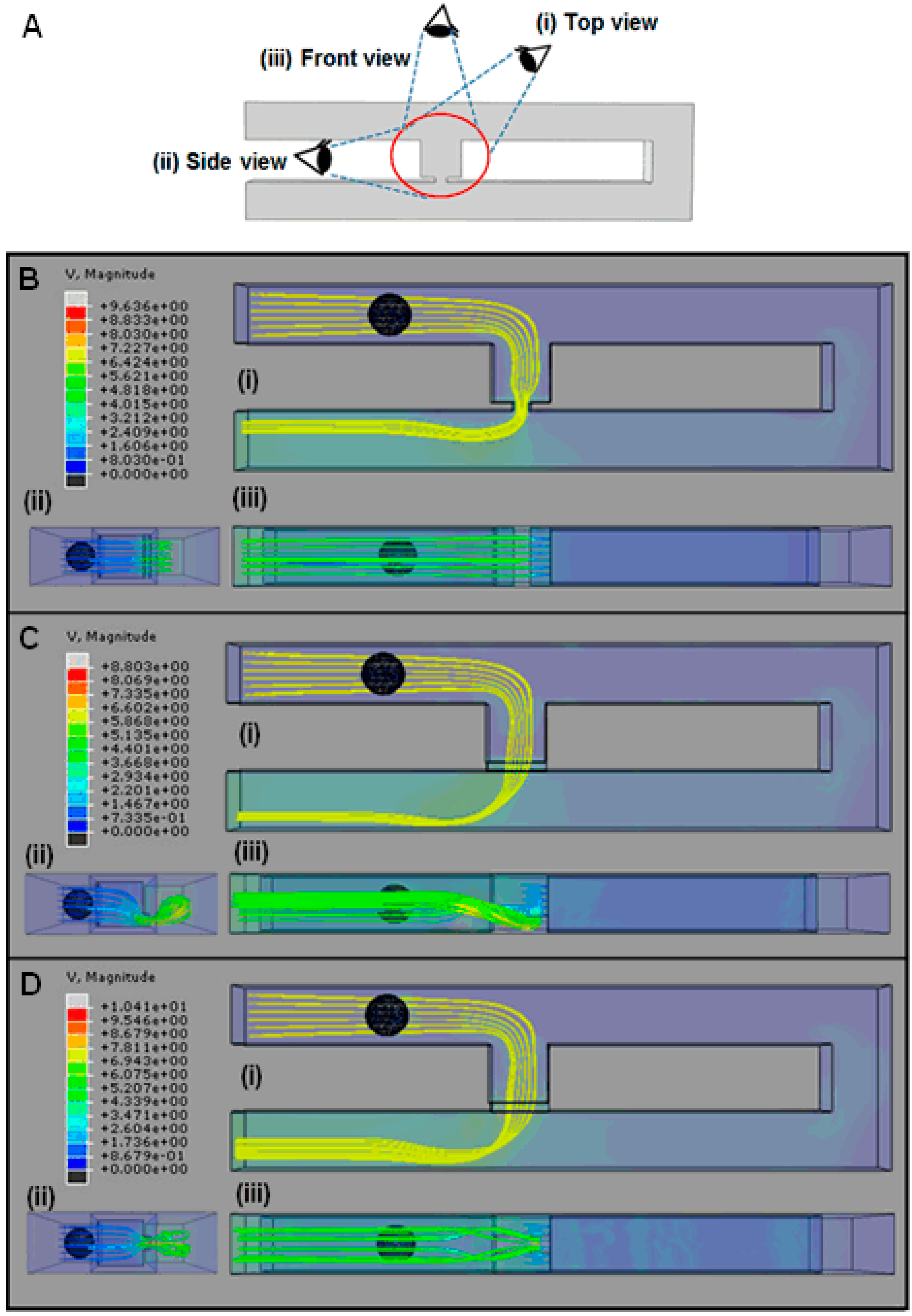
3. The Concept of the Model
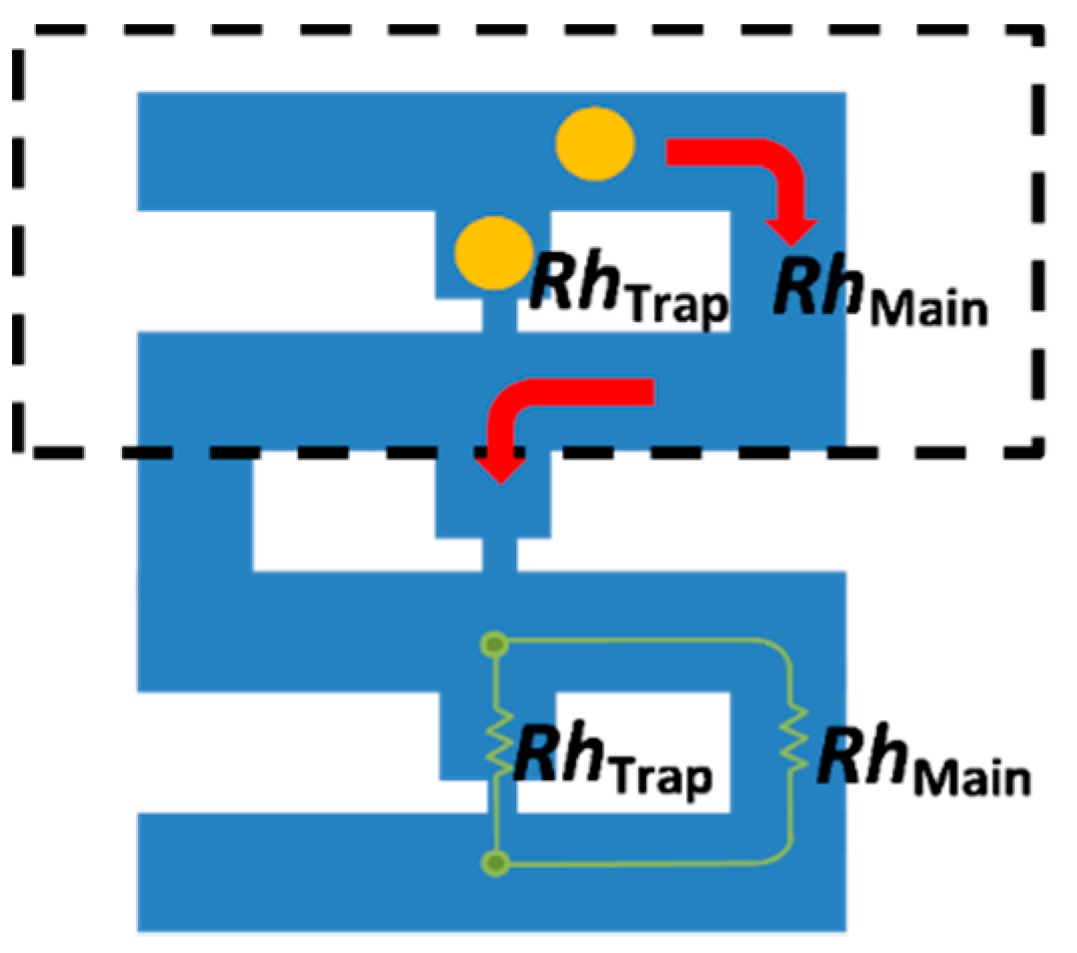
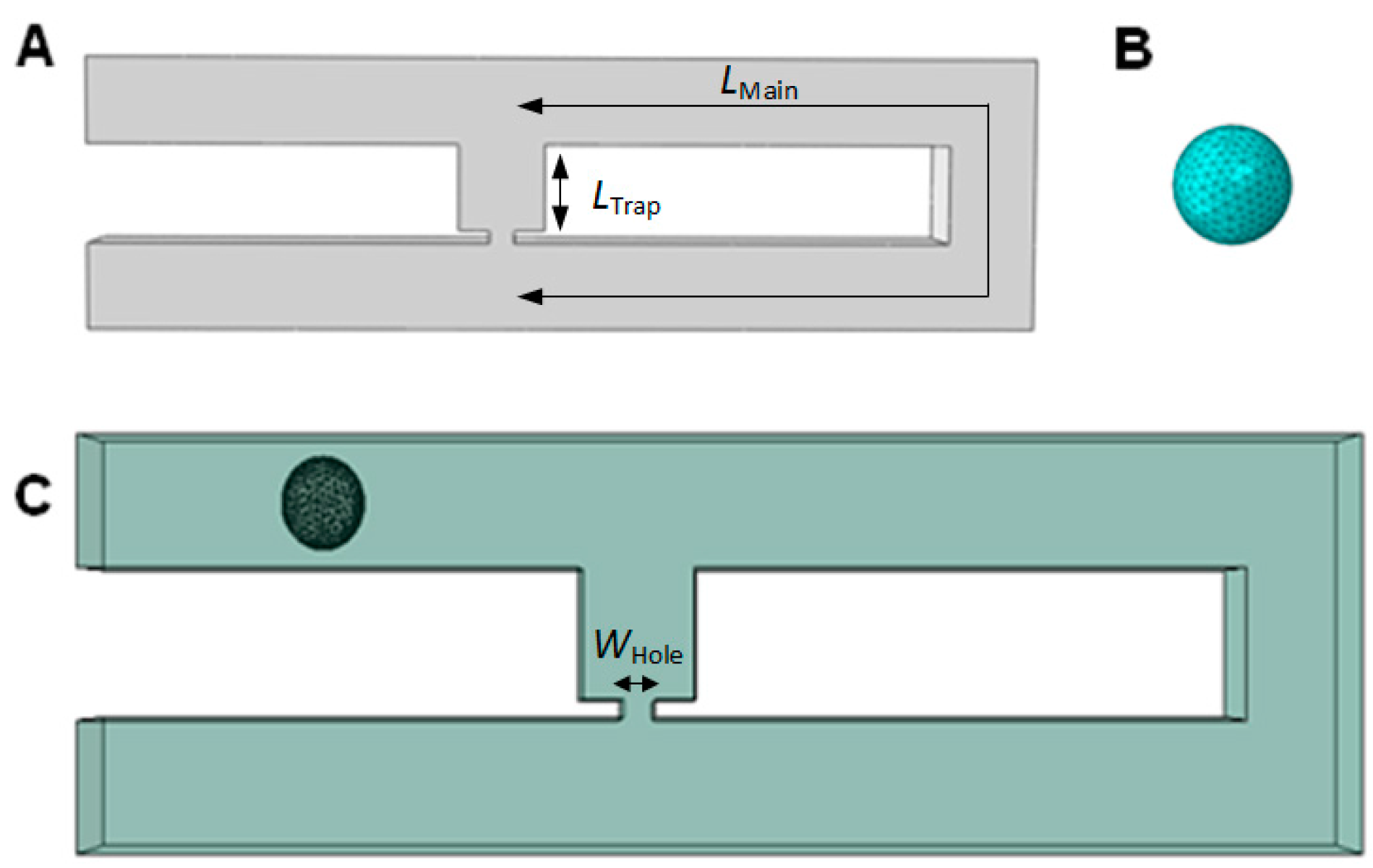
4. Simulation Setup
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Johann, R.M. Cell Trapping in Microfluidic Chips. Anal. Bioanal. Chem. 2006, 385, 408–412. [Google Scholar] [CrossRef] [PubMed]
- Lee, G.-H.; Kim, S.-H.; Kang, A.; Takayama, S.; Lee, S.-H.; Park, J.Y. Deformable L-shaped microwell array for trapping pairs of heterogeneous cells. J. Micromech. Microeng. 2015, 25. [Google Scholar] [CrossRef]
- Sun, T.; Kovac, J.; Voldman, J. Image-based single-cell sorting via dual-photopolymerized microwell arrays. Anal. Chem. 2014, 86, 977–981. [Google Scholar] [CrossRef] [PubMed]
- Rettig, J.R.; Folch, A. Large-scale single-cell trapping and imaging using microwell arrays. Anal. Chem. 2005, 77, 5628–5634. [Google Scholar] [CrossRef] [PubMed]
- Tang, J.; Peng, R.; Ding, J. The Regulation of Stem cell differentiation by cell-cell contact on micropatterned material surfaces. Biomaterials 2010, 31, 2470–2476. [Google Scholar] [CrossRef] [PubMed]
- Doh, J.; Kim, M.; Krummel, M.F. Cell-laden microwells for the study of multicellularity in lymphocyte fate decisions. Biomaterials 2010, 31, 3422–3428. [Google Scholar] [CrossRef] [PubMed]
- Chen, N.-C.; Chen, C.-H.; Chen, M.-K.; Jang, L.-S.; Wang, M.-H. Single-cell trapping and impedance measurement utilizing dielectrophoresis in a parallel-plate microfluidic device. Sens. Actuators B Chem. 2014, 190, 570–577. [Google Scholar] [CrossRef]
- Sen, M.; Ino, K.; Ramon-Azcon, J.; Shiku, H.; Matsue, T. Cell pairing using a dielectrophoresis-based device with interdigitated array electrodes. Lab Chip 2013, 13, 3650–3652. [Google Scholar] [CrossRef] [PubMed]
- Voldman, J.; Gray, M.L.; Toner, M.; Schmidt, M.A. A Microfabrication-based dynamic array cytometer. Anal. Chem. 2002, 74, 3984–3990. [Google Scholar] [CrossRef] [PubMed]
- Thomas, R.S.; Morgan, H.; Green, N.G. Negative DEP traps for single cell immobilisation. Lab Chip 2009, 9, 1534–1540. [Google Scholar] [CrossRef] [PubMed]
- Gray, D.S.; Tan, J.L.; Voldman, J.; Chen, C.S. Dielectrophoretic registration of living cells to a microelectrode array. Biosens. Bioelectron. 2004, 19, 771–780. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.-C.; Allen, S.G.; Ingram, P.N.; Buckanovich, R.; Merajver, S.D.; Yoon, E. Single-cell migration chip for chemotaxis-based microfluidic selection of heterogeneous cell populations. Sci. Rep. 2015, 5, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Jin, D.; Deng, B.; Li, J.X.; Cai, W.; Tu, L.; Chen, J.; Wu, Q.; Wang, W.H. A Microfluidic device enabling high-efficiency single cell trapping. Biomicrofluidics 2015, 9. [Google Scholar] [CrossRef] [PubMed]
- Benavente-Babace, A.; Gallego-Pérez, D.; Hansford, D.J.; Arana, S.; Pérez-Lorenzo, E.; Mujika, M. Single-cell trapping and selective treatment via co-flow within a microfluidic platform. Biosens. Bioelectron. 2014, 61, 298–305. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Erath, J.; Rodriguez, A.; Yang, C. A High-efficiency microfluidic device for size-selective trapping and sorting. Lab Chip 2014, 14, 2480–2490. [Google Scholar] [CrossRef] [PubMed]
- Lee, P.J.; Hung, P.J.; Shaw, R.; Jan, L.; Lee, L.P. Microfluidic application-specific integrated device for monitoring direct cell-cell communication via gap junctions between individual cell pairs. Appl. Phys. Lett. 2005, 86. [Google Scholar] [CrossRef]
- Frimat, J.-P.; Becker, M.; Chiang, Y.-Y.; Marggraf, U.; Janasek, D.; Hengstler, J.G.; Franzke, J.; West, J. A microfluidic array with cellular valving for single cell co-culture. Lab Chip 2011, 11, 231–237. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.; Lee, S.; Kim, J. Hydrodynamic trap-and-release of single particles using dual-function elastomeric valves: design, fabrication, and characterization. Microfluid. Nanofluid. 2012, 13, 835–844. [Google Scholar] [CrossRef]
- Arakawa, T.; Noguchi, M.; Sumitomo, K.; Yamaguchi, Y.; Shoji, S. High-throughput single-cell manipulation system for a large number of target cells. Biomicrofluidics 2011, 5. [Google Scholar] [CrossRef] [PubMed]
- Kobel, S.; Valero, A.; Latt, J.; Renaud, P.; Lutolf, M. Optimization of microfluidic single cell trapping for long-term on-chip culture. Lab Chip 2010, 10, 857–863. [Google Scholar] [CrossRef]
- Hong, S.; Pan, Q.; Lee, L.P. Single-cell level co-culture platform for intercellular communication. Integr. Biol. 2012, 4, 374–380. [Google Scholar] [CrossRef] [PubMed]
- Shi, W.; Qin, J.; Ye, N.; Lin, B. Droplet-based microfluidic system for individual caenorhabditis elegans assay. Lab Chip 2008, 8, 1432–1435. [Google Scholar] [CrossRef] [PubMed]
- Di Carlo, D.; Aghdam, N.; Lee, L.P. Single-cell enzyme concentrations, kinetics, and inhibition analysis using high-density hydrodynamic cell isolation arrays. Anal. Chem. 2006, 78, 4925–4930. [Google Scholar] [CrossRef] [PubMed]
- Skelley, A.M.; Kirak, O.; Suh, H.; Jaenisch, R.; Voldman, J. Microfluidic control of cell pairing and fusion. Nat. Methods 2009, 6, 147–152. [Google Scholar] [CrossRef] [PubMed]
- Di Carlo, D.; Wu, L.Y.; Lee, L.P. Dynamic single cell culture array. Lab Chip 2006, 6, 1445–1449. [Google Scholar] [CrossRef] [PubMed]
- Tan, W.-H.; Takeuchi, S. A Trap-and-release Integrated microfluidic system for dynamic microarray applications. Proc. Natl. Acad. Sci. USA 2007, 104, 1146–1151. [Google Scholar] [CrossRef] [PubMed]
- Chung, K.; Rivet, C.A.; Kemp, M.L.; Lu, H.; States, U. Imaging single-cell signaling dynamics with a deterministic high-density single-cell trap array. Anal. Chem. 2011, 83, 7044–7052. [Google Scholar] [CrossRef] [PubMed]
- Khalili, A.A.; Basri, M.A.M.; Ahmad, M.R. Simulation of single cell trapping via hydrodynamic manipulation. J. Teknol. 2014, 69, 121–126. [Google Scholar]
- Teshima, T.; Ishihara, H.; Iwai, K.; Adachi, A.; Takeuchi, S. A dynamic microarray device for paired bead-based analysis. Lab Chip 2010, 10, 2443–2448. [Google Scholar] [CrossRef] [PubMed]
- Kumano, I.; Hosoda, K.; Suzuki, H.; Hirata, K.; Yomo, T. Hydrodynamic trapping of tetrahymena thermophila for the long-term monitoring of cell behaviors. Lab Chip 2012, 12, 3451–3457. [Google Scholar] [CrossRef] [PubMed]
- Gervais, T.; El-Ali, J.; Günther, A.; Jensen, K.F. Flow-induced deformation of shallow microfluidic channels. Lab Chip 2006, 6, 500–507. [Google Scholar] [CrossRef] [PubMed]
- Bryan, A.K.; Goranov, A.; Amon, A.; Manalis, S.R. Measurement of mass, density, and volume during the cell cycle of yeast. Proc. Natl. Acad. Sci. USA 2010, 107, 999–1004. [Google Scholar] [CrossRef] [PubMed]
- Ahmad, M.R.; Nakajima, M.; Kojima, S.; Homma, M.; Fukuda, T. The Effects of cell sizes, environmental conditions, and growth phases on the strength of individual W303 yeast cells inside ESEM. IEEE Trans. Nanobiosci. 2008, 7, 185–193. [Google Scholar] [CrossRef] [PubMed]
- Smith, A.E.; Zhang, Z.; Thomas, C.R.; Moxham, K.E.; Middelberg, A.P. The Mechanical properties of saccharomyces cerevisiae. Proc. Natl. Acad. Sci. USA 2000, 97, 9871–9874. [Google Scholar] [CrossRef] [PubMed]
- Stenson, J.D.; Thomas, C.R.; Hartley, P. Modelling the mechanical properties of yeast cells. Chem. Eng. Sci. 2009, 64, 1892–1903. [Google Scholar] [CrossRef]
- Stenson, J.D.; Hartley, P.; Wang, C.; Thomas, C.R. Determining the mechanical properties of yeast cell walls. Biotechnol. Prog. 2011, 27, 505–512. [Google Scholar] [PubMed]
- Burg, T.P.; Godin, M.; Knudsen, S.M.; Shen, W.; Carlson, G.; Foster, J.S.; Babcock, K.; Manalis, S.R. Weighing of Biomolecules, Single Cells and Single Nanoparticles in Fluid. Nature 2007, 446, 1066–1069. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.; Chunara, R.; Shen, W.; Payer, K.; Babcock, K.; Burg, T.P.; Manalis, S.R. Suspended microchannel resonators with piezoresistive sensors. Lab Chip 2011, 11, 645–651. [Google Scholar] [CrossRef]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons by Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Khalili, A.A.; Ahmad, M.R. Numerical Analysis of Hydrodynamic Flow in Microfluidic Biochip for Single-Cell Trapping Application. Int. J. Mol. Sci. 2015, 16, 26770-26785. https://doi.org/10.3390/ijms161125987
Khalili AA, Ahmad MR. Numerical Analysis of Hydrodynamic Flow in Microfluidic Biochip for Single-Cell Trapping Application. International Journal of Molecular Sciences. 2015; 16(11):26770-26785. https://doi.org/10.3390/ijms161125987
Chicago/Turabian StyleKhalili, Amelia Ahmad, and Mohd Ridzuan Ahmad. 2015. "Numerical Analysis of Hydrodynamic Flow in Microfluidic Biochip for Single-Cell Trapping Application" International Journal of Molecular Sciences 16, no. 11: 26770-26785. https://doi.org/10.3390/ijms161125987