Molecular Cloning and Characterization of DXS and DXR Genes in the Terpenoid Biosynthetic Pathway of Tripterygium wilfordii
Abstract
1. Introduction
2. Results
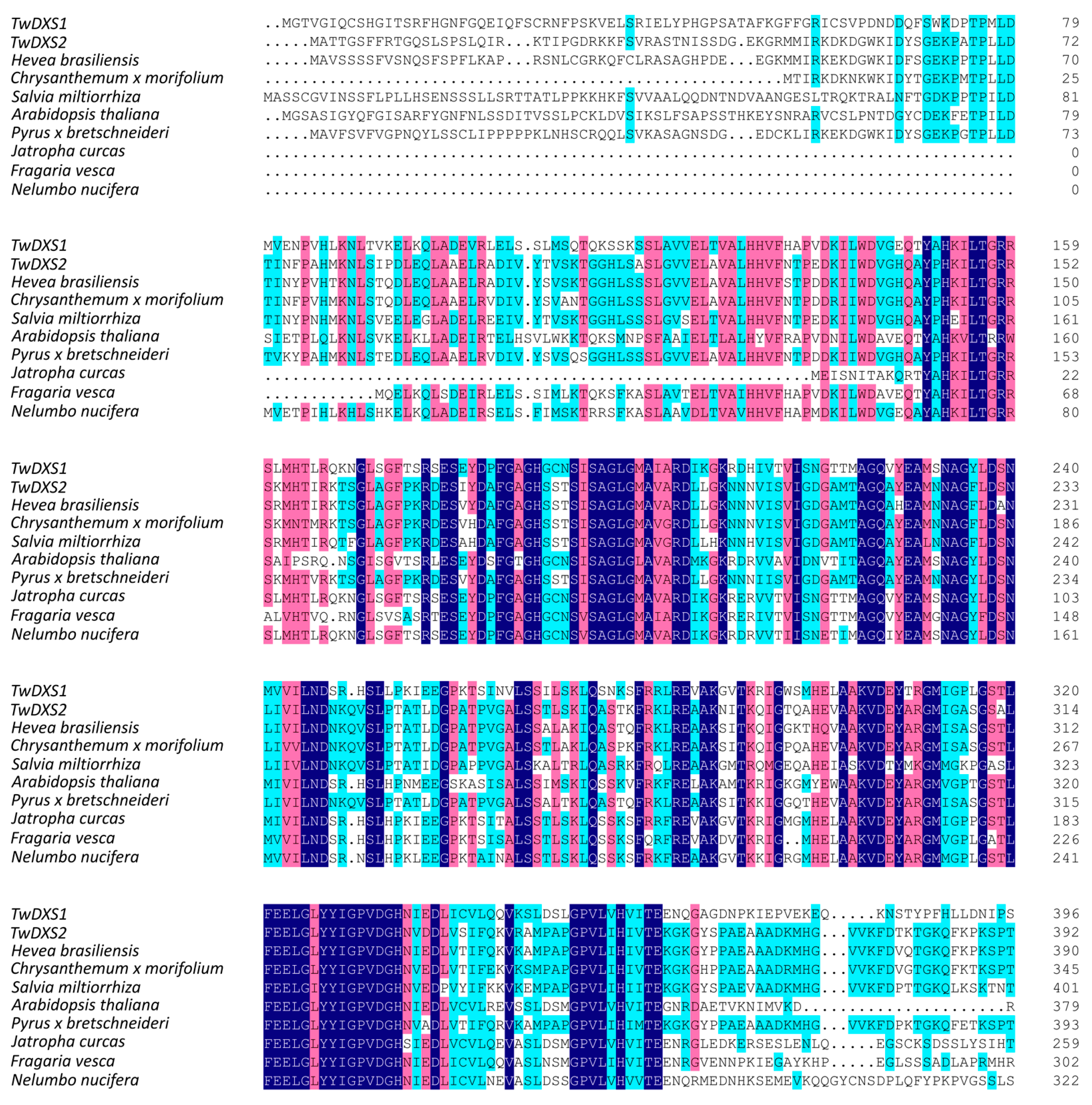
2.1. Cloning and Sequence Analysis of TwDXS1, TwDXS2, and TwDXR

| Gene Name | Full Length (bp) | ORF (bp) | Amino Acid Sequence (aa) | MW (kDa) | pI |
|---|---|---|---|---|---|
| Tw DXS1 | 2556 | 2154 | 717 | 78.7 | 6.04 |
| Tw DXS2 | 2412 | 2148 | 715 | 77.0 | 7.10 |
| Tw DXR | 1816 | 1410 | 469 | 51.1 | 5.88 |
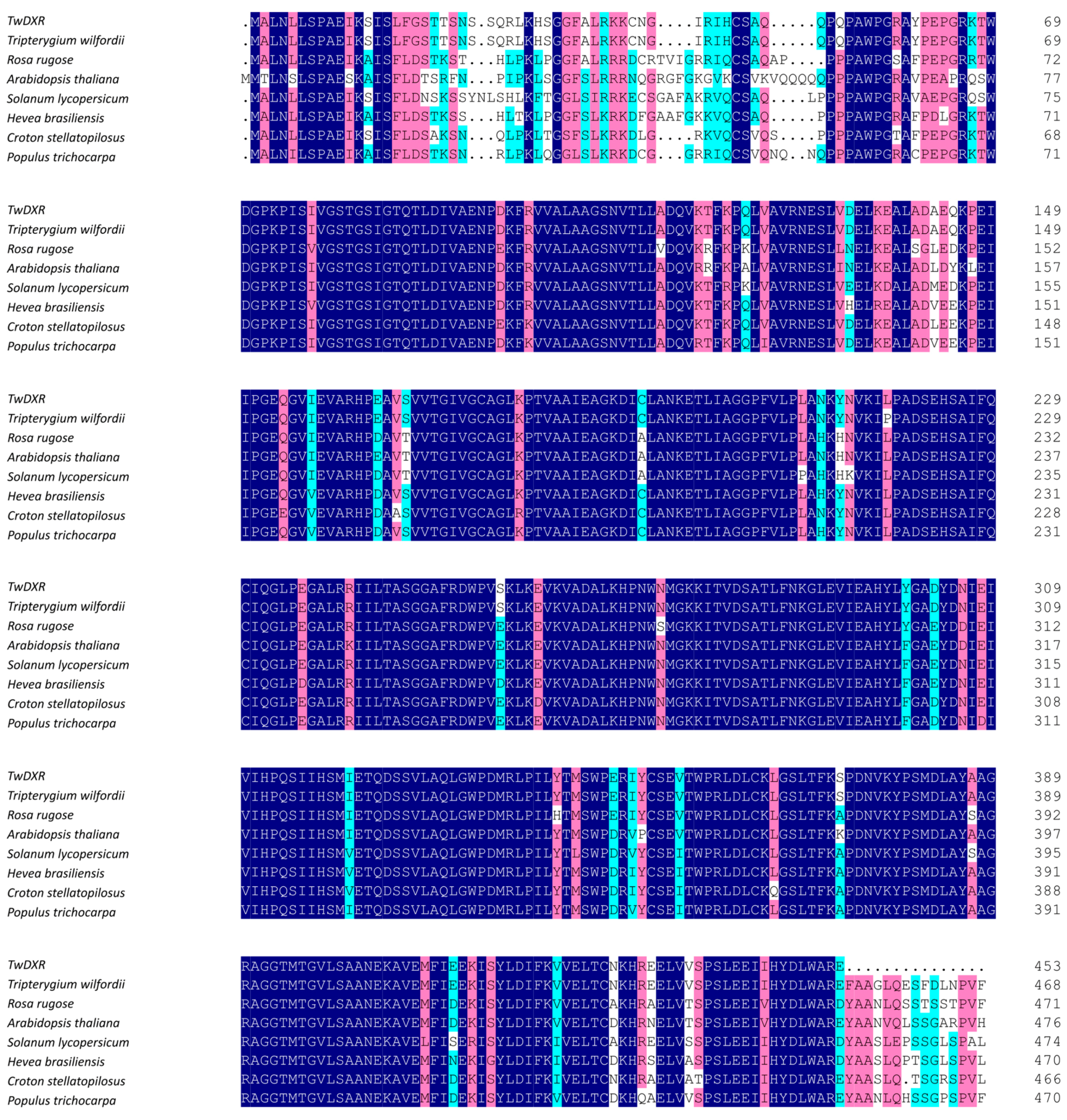
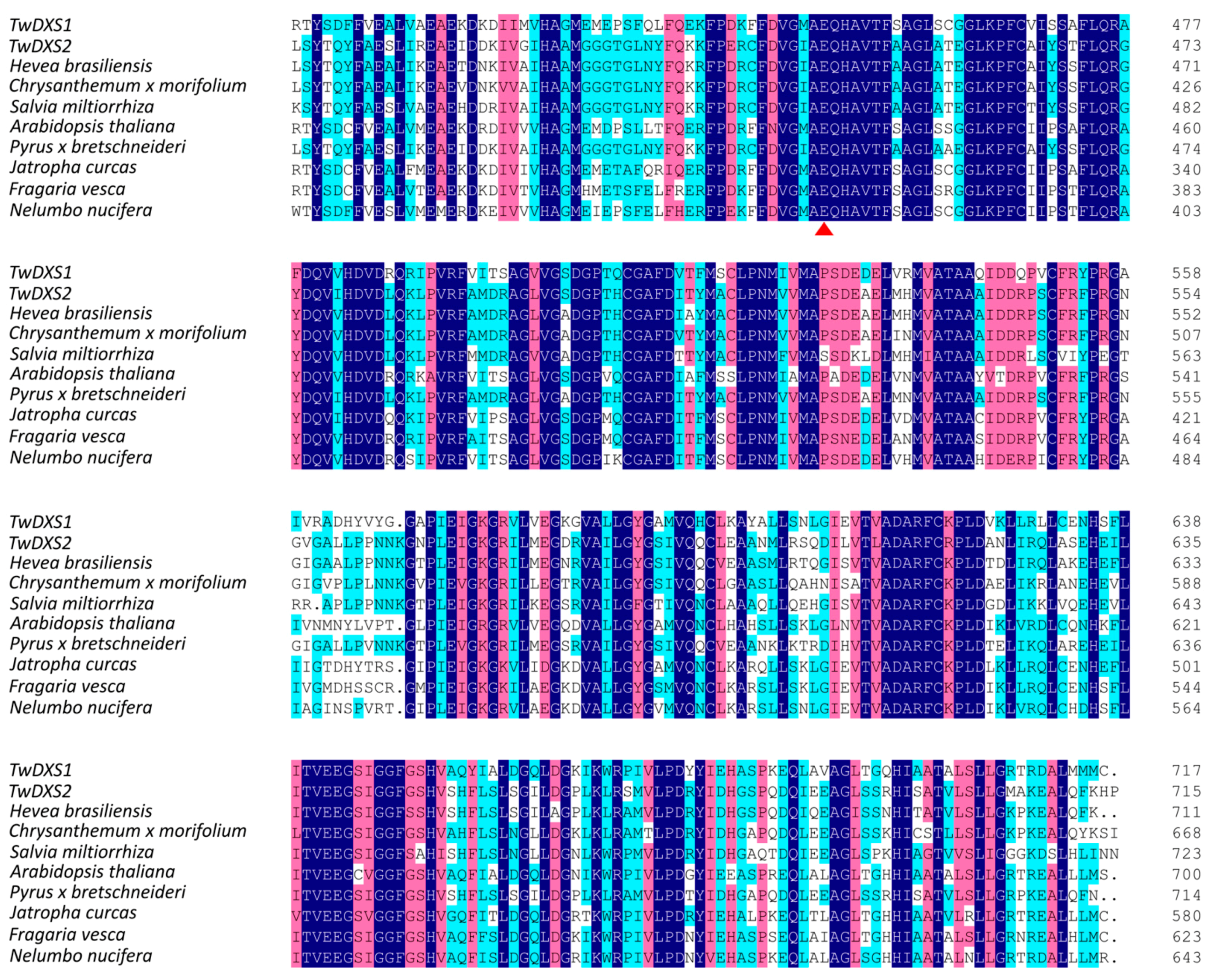
2.2. Bioinformatics Analysis
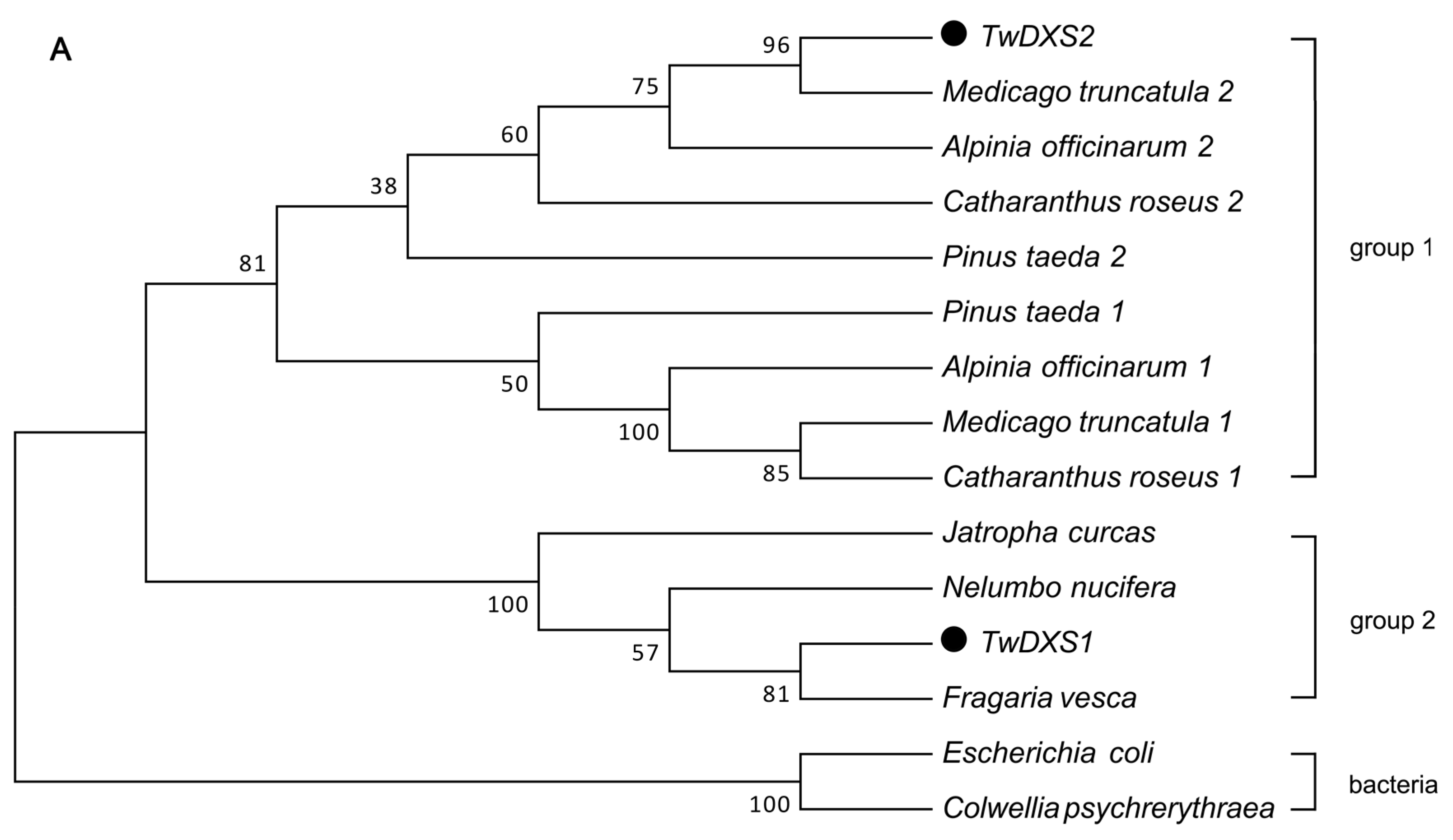
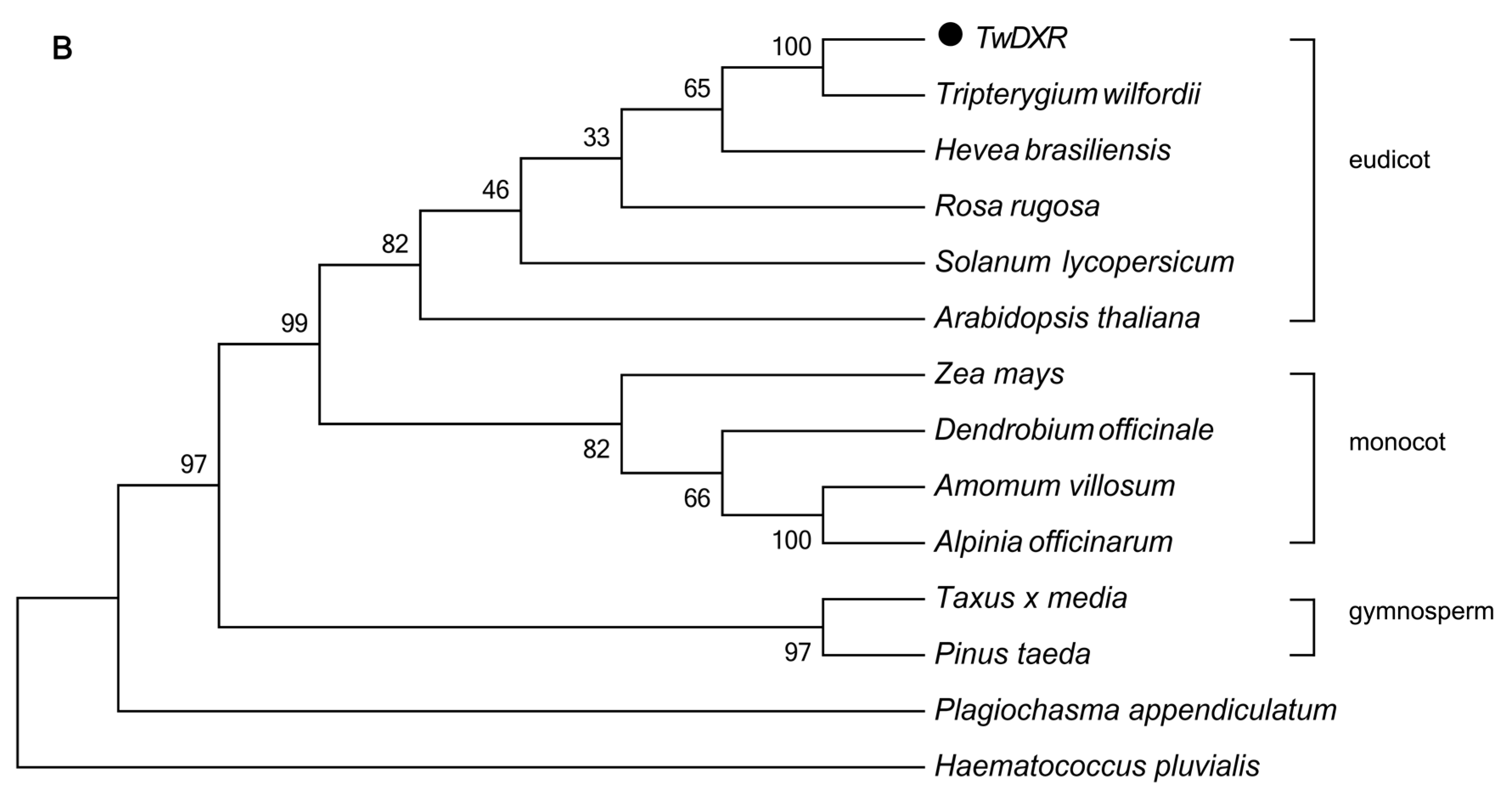
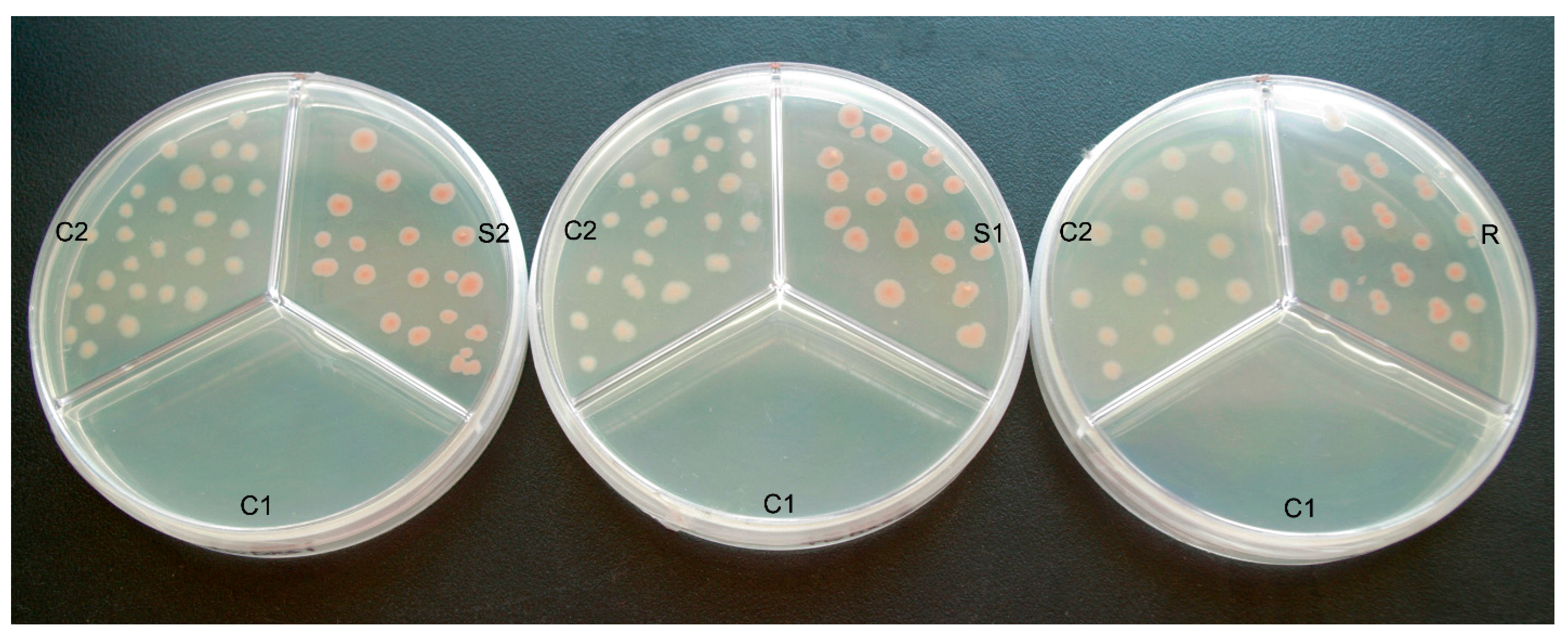
2.3. Color Complementation and Functional Analysis
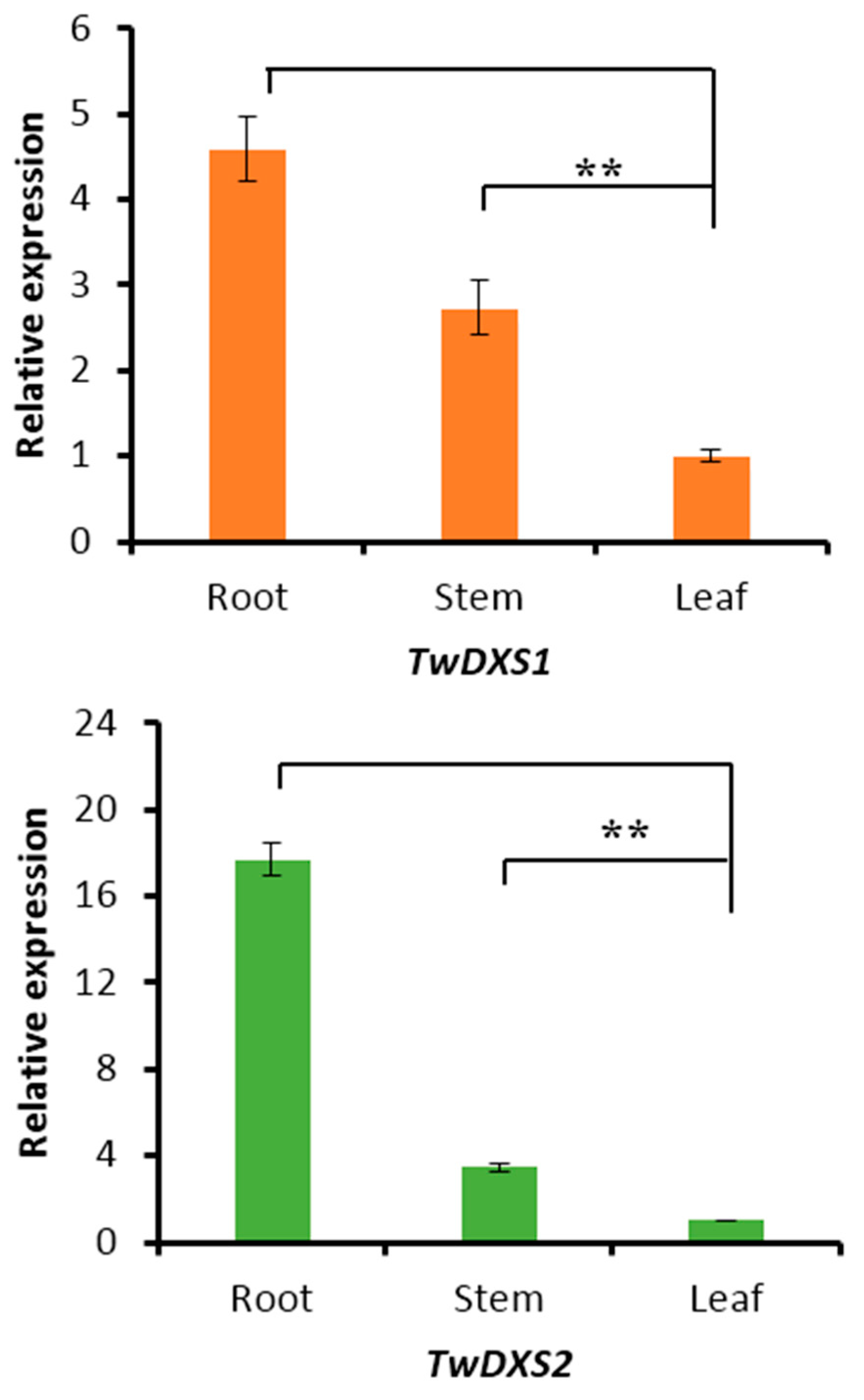
2.4. Tissue-Specific Expression Analysis
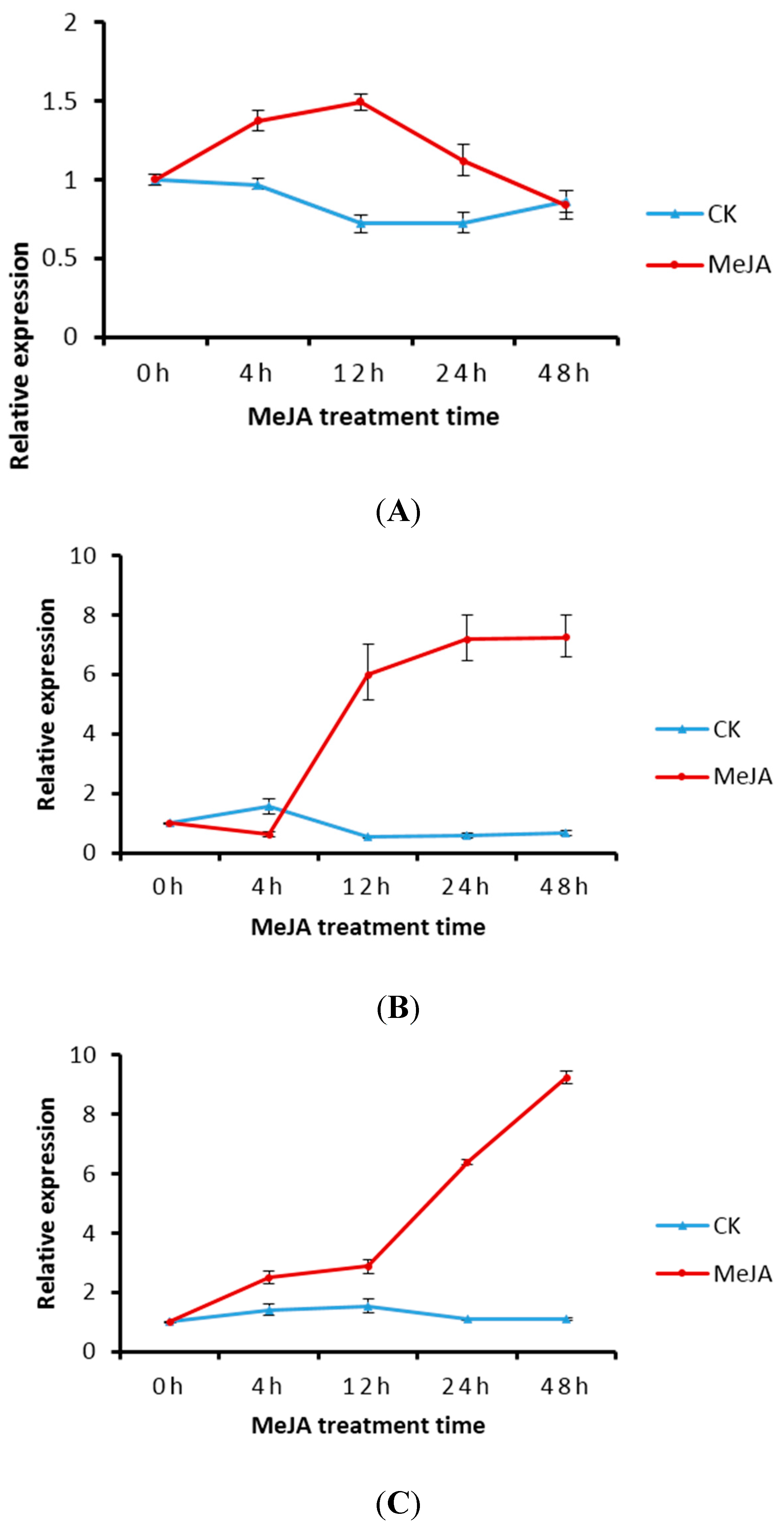
2.5. Transcript Level of DXS/DXR in Suspension Cells and Metabolite Analysis
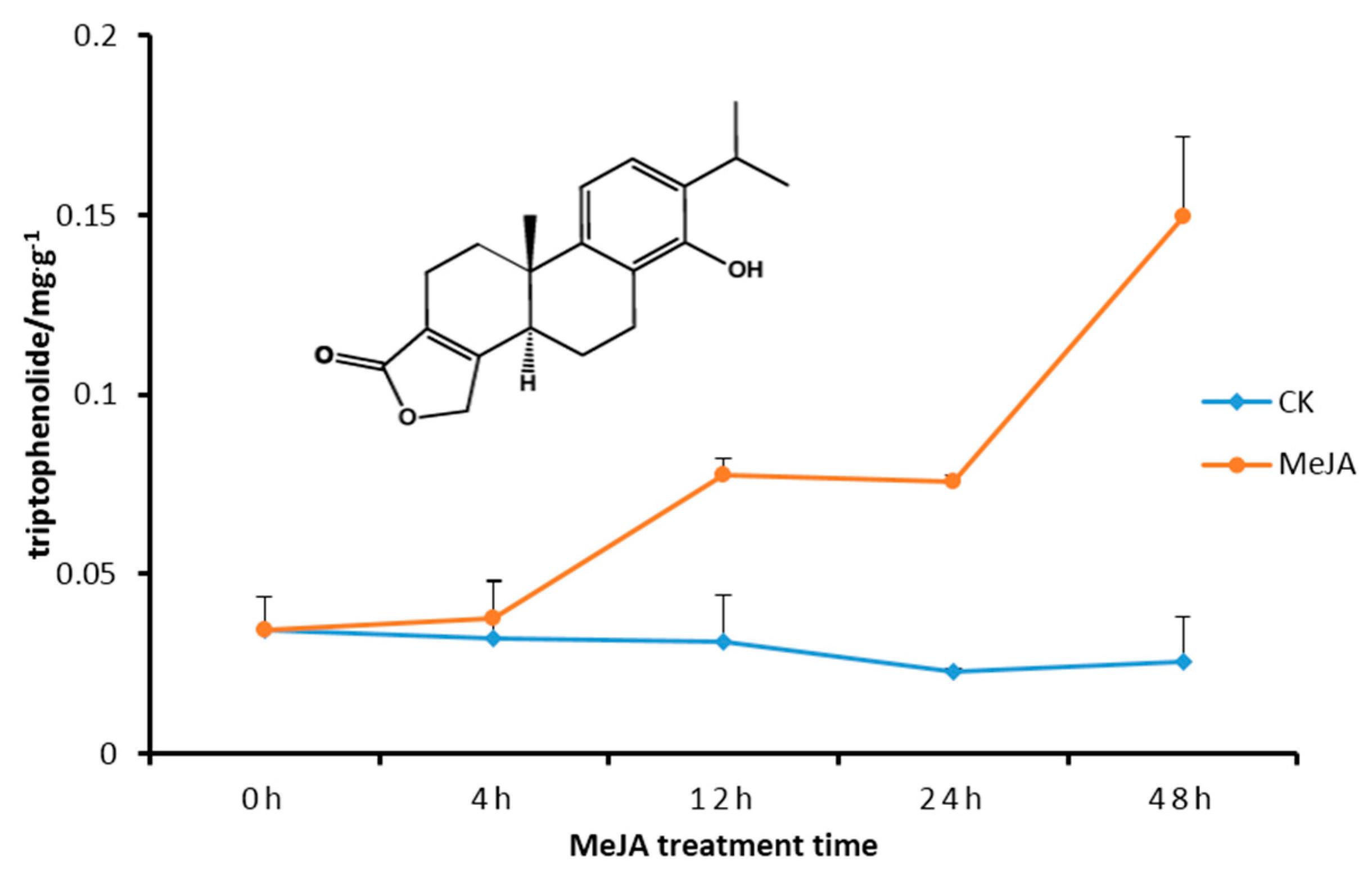
3. Discussion
4. Experimental Sections
4.1. Plant Materials
4.2. RNA Isolation and cDNA Cloning
| Usage | Primer Name | Primer Sequences 5′→3′ |
|---|---|---|
| RACE-PCR | DXR-3′ | GTGTATTCAAGGATTGCCAGAGGGTG |
| LC-PCR | DXS1-F | AGACTCGAATTATGCTCCGTTT |
| DXS1-R | GCAGCATTATACATTTTTCTACGG | |
| DXS2-F | AAGGGAGGATGATGATACGCA | |
| DXS2-R | CTCATAGCATGTATGGTGCTAGAGC | |
| DXR-F | CCCGAACCCAAACCACAC | |
| DXR-R | AGGAGTAGATTAACCGGGATGAT | |
| ORF-PCR | ORFDXS1-F | GGAAGATCTATGGGTACTGTTGGTATTCAGTGCTC |
| ORFDXS1-R | TTGCGGCCGCCTAGCACATCATC | |
| ORFDXS2-F | GGAAGATCTATGGGTACTGTTGGTATTCAGTGCTC | |
| ORFDXS2-R | CGGGGTACCATGGCGACTACTGGGTCTTTC | |
| ORFDXR-F | CGCGGATCCTTAGGGATGCTTAAACT | |
| ORFDXR-R | TTGCGGCCGCTCATGCAAAGACAGGATTTA | |
| qRT-PCR | RTDXS1-F | CTCCTTAGGCTGCTATGTGA |
| RTDXS1-R | GCTGTTCCTTGGGTGATGC | |
| RTDXS2-F | GGGCATCAGGCATACCCACACAAG | |
| RTDXS2-R | TGGCTCCATCTCCAATCAC | |
| RTDXR-F | TCAAGGATTGCCAGAGGG | |
| RTDXR-R | ATGAATGATAGACTGCGGATG | |
| Actin-F | AGGAACCACCGATCCAGACA | |
| Actin-R | GGTGCCCTGAGGTCCTGTT |
4.3. Data Analysis
4.4. Construction of Expression Vectors
4.5. Color Complementation
4.6. Tissue-Specific Expression Analysis
4.7. Elicitation of Suspension Cells with MeJA
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Tao, X.; Lipsky, P.E. The Chinese anti-inflammatory and immunosuppressive herbal remedy Tripterygium wilfordii Hook F. Rheum. Dis. Clin. N. Am. 2000, 26, 29–50. [Google Scholar] [CrossRef]
- Ushiro, S.; Ono, M.; Nakayama, J. New nortriterpenoid isolated from anti-rheumatoid arthritis plant, Tripterygium wilfordii, modulates tumor growth and neovascularization. Int. J. Cancer 1997, 72, 657–663. [Google Scholar] [CrossRef]
- Marks, W.H. Tripterygium wilfordii Hook. f. versus sulfasalazine in the treatment of rheumatoid arthritis: A well-designed clinical trial of a botanical demonstrating effectiveness. Fitoterapia 2011, 82, 85–87. [Google Scholar] [PubMed]
- Manzo, S.G.; Zhou, Z.L.; Wang, Y.Q. Natural product triptolide mediates cancer cell death by triggering CDK7-dependent degradation of RNA polymerase II. Cancer Res. 2012, 72, 5363–5373. [Google Scholar] [CrossRef] [PubMed]
- Wong, K.F.; Yuan, Y.; Luk, J.M. Tripterygium wilfordii bioactive compounds as anticancer and anti-inflammatory agents. Clin. Exp. Pharmacol. Physiol. 2012, 39, 311–320. [Google Scholar] [CrossRef] [PubMed]
- Horiuch, M.; Murakami, C.; Fukamiya, N.; Yu, D.; Chen, T.H.; Bastow, K.F.; Zhang, D.C.; Takaishi, Y.; Imakura, Y.; Lee, K.H. Tripfordines A–C, sesquiterpene pyridine alkaloids from tripterygium wilfordii, and structure anti-HIV activity relationships of tripterygium alkaloids. J. Nat. Prod. 2006, 69, 1271–1274. [Google Scholar] [CrossRef] [PubMed]
- Cai, Y.; Luo, Q.; Sun, M.; Corke, H. Antioxidant activity and phenolic compounds of 112 traditional Chinese medicinal plants associated with anticancer. Life Sci. 2004, 74, 2157–2184. [Google Scholar] [CrossRef] [PubMed]
- Cai, M.Q.; Chen, X.H.; He, S.W.; Ouyang, X.K.; Jin, M.C. Determination of four pyridine alkaloids from Tripterygium wilfordii Hook. f. in human plasma by high-performance liquid chromatography coupled with mass spectrometry. J. Chromatogr. B 2011, 879, 3516–3522. [Google Scholar] [CrossRef] [PubMed]
- Ma, J.; Schmidt, B.M.; Poulev, A.; Raskin, I. Determination of tripdiolide in root extracts of Tripterygium wilfordii by solid-phase extraction and reversed-phase high-performance liquid chromatography. Phytochem. Anal. 2008, 19, 348–352. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Yu, D.F.; Hu, B.H.; Chen, G.P.; Yang, C.X.; Yang, J.; Xu, J.Y.; Li, L.Z. Structure revision of triptophenolide. Acta Pharm. Sin. 1989, 25, 929–931. [Google Scholar]
- Yang, D.; Ye, X.Y.; Xu, M. Enantioselective total synthesis of (−)-triptolide, (−)-triptonide, (+)-triptophenolide, and (+)-triptoquinonide. J. Org. Chem. 2000, 65, 2208–2217. [Google Scholar] [CrossRef] [PubMed]
- Lange, B.M.; Rujan, T.; Martin, W.; Croteau, R. Isoprenoid biosynthesis: The evolution of two ancient and distinct pathways across genomes. Proc. Natl. Acad. Sci. USA 2000, 97, 13172–13177. [Google Scholar] [CrossRef] [PubMed]
- Wölwer-Rieck, U.; May, B.; Lankes, C.; Wüst, M. Methylerythritol and mevalonate pathway contributions to biosynthesis of mono-, sesqui-, and diterpenes in glandular trichomes and leaves of Stevia rebaudiana bertoni. J. Agric. Food Chem. 2014, 62, 2428–2435. [Google Scholar] [CrossRef] [PubMed]
- Ge, X.; Wu, J. Induction and potentiation of diterpenoid tanshinone accumulation in Salvia miltiorrhiza hairy roots by aminobutyric acid. Appl. Microbiol. Biotechnol. 2005, 68, 183–188. [Google Scholar] [CrossRef] [PubMed]
- Lu, X.M.; Hu, X.J.; Zhao, Y.Z.; Song, W.B.; Zhang, M.; Chen, Z.L.; Chen, W.; Dong, Y.B.; Wang, Z.H.; Lai, J.S. Map-based cloning of zb7 encoding an IPP and DMAPP synthase in the MEP pathway of maize. Mol. Plant 2012, 5, 1100–1112. [Google Scholar] [CrossRef] [PubMed]
- Sprenger, G.A.; Schörken, U.; Wiegert, T.; Grolle, S.; de Graaf, A.A.; Taylor, S.V.; Begley, T.P.; Bringer-Meyer, S.; Sahm, H. Identification of a thiamin-dependent synthesis in Escherichia coli required for the formation of the 1-deoxy-d-xylulose-5-phosphate precursor to isoprenoids, thiamin, and pyridoxol. Proc. Natl. Acad. Sci. USA 1997, 94, 12857–12862. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, S.; Kuzuyama, T.; Watanabe, H.; Seto, H. Direct formation of 2-C-methyl-d-erythritol-4-phosphate from 1-deoxy-d-xylulose-5-phosphate by 1-deoxy-d-xylulose-5-phosphate reductoisomerase, a new enzyme in the non-mevalonate pathway to isopentenyl diphosphate. Proc. Natl. Acad. Sci. USA 1998, 39, 4509–4512. [Google Scholar]
- Tabata, S.; Kaneko, T.; Nakamura, Y. Sequence and analysis of chromosome 5 of the plant Arabidopsis thaliana. Nature 2000, 408, 823–826. [Google Scholar] [PubMed]
- Hans, J.; Hause, B.; Strack, D.; Walter, M.H. Cloning, characterization and immunolocalization of a mycorrhiza-inducible 1-deoxy-d-xylulose-5-phosphate reductoisomerase in arbuscule-containing cells of maize. Plant Physiol. 2004, 134, 614–624. [Google Scholar] [CrossRef] [PubMed]
- Wu, S.J.; Shi, M.; Wu, J.Y. Cloning and characterization of 1-deoxy-d-xylulose-5-phosphate reductoisomerase gene for diterpenoid tanshinone biosynthesis in Salvia miltiorrhiza hairy roots. Biotechnol. Appl. Bacteriol. 2009, 52, 89–95. [Google Scholar] [CrossRef] [PubMed]
- Morrone, D.; Lowry, L.; Determan, M.K.; Hershey, D.M.; Xu, M.; Peters, R.J. Increasing diterpene yield with a modular metabolic engineering system in E. coli: Comparison of MEV and MEP isoprenoid precursor pathway engineering. Appl. Microbiol. Biotechnol. 2010, 85, 1893–1906. [Google Scholar] [CrossRef] [PubMed]
- Peebles, C.A.; Sander, G.W.; Hughes, E.H.; Peacock, R.; Shanks, J.V.; San, K.Y. The expression of 1-deoxy-d-xylulose synthase and geraniol-10-hydroxylase or anthranilate synthase increases terpenoid indole alkaloid accumulation in Catharanthus roseus hairy roots. Metab. Eng. 2011, 13, 234–240. [Google Scholar] [CrossRef] [PubMed]
- Chang, K.; Qiu, F.; Chen, M.; Zeng, L.; Liu, X.; Yang, C.; Lan, X.; Wang, Q.; Liao, Z. Engineering the MEP pathway enhanced ajmalicine biosynthesis. Biotechnol. Appl. Biochem. 2014, 61, 249–255. [Google Scholar] [CrossRef] [PubMed]
- Xiang, S.; Usunow, G.; Lange, G.; Busch, M.; Tong, L. Crystal structure of 1-deoxy-d-xylulose-5-phosphate synthase, a crucial enzyme for isoprenoids biosynthesis. J. Biol. Chem. 2007, 282, 2676–2682. [Google Scholar] [CrossRef] [PubMed]
- Bouvier, F.; D’Harlingue, A.; Suire, C.; Backhaus, R.A.; Camara, B. Dedicated roles of plastid transketolases during the early onset of isoprenoid biosynthesis in pepper fruits. Plant Physiol. 1998, 117, 1423–1431. [Google Scholar] [CrossRef] [PubMed]
- Krushkal, J.; Pistilli, M.; Ferrell, K.M.; Souret, F.F.; Weathers, P.J. Computational analysis of the evolution of the structure and function of 1-deoxy-d-xylulose-5-phosphate synthase, a key regulator of the mevalonate-independent pathway in plants. Gene 2003, 313, 127–138. [Google Scholar] [CrossRef]
- Cunningham, F.X.; Sun, Z.; Chamovitz, D.; Hirschberg, J.; Gantt, E. Molecular structure and enzymatic function of lycopene cyclase from the Cyanobacterium Synechococcus sp strain PCC7942. Plant Cell 1994, 6, 1107–1121. [Google Scholar] [CrossRef] [PubMed]
- Sun, Z.; Gantt, E.; Cunningham, F.X. Cloning and functional analysis of the β-carotene hydroxylase of Arabidopsis thaliana. J. Biol. Chem. 1996, 271, 24349–24352. [Google Scholar] [CrossRef] [PubMed]
- Gallagher, C.E.; Cervantes, M.; Wurtzel, E.T. Surrogate biochemistry: use of Escherichia coli to identify plant cDNAs that impact metabolic engineering of carotenoid accumulation. Appl. Microbiol. Biotechnol. 2003, 60, 713–719. [Google Scholar] [CrossRef] [PubMed]
- Moehs, C.P.; Tian, L.; Osteryoung, K.W.; Dellapenna, D. Analysis of carotenoid biosynthetic gene expression during marigold petal development. Plant Mol. Biol. 2001, 45, 281–293. [Google Scholar] [CrossRef] [PubMed]
- Cunningham, F.X.; Gantt, E. A portfolio of plasmids for identification and analysis of carotenoid pathway enzymes: Adonis aestivalis as a case study. Photosynth. Res. 2007, 92, 245–259. [Google Scholar] [CrossRef] [PubMed]
- Yan, X.; Zhang, L.; Wang, J.; Liao, P.; Zhang, Y.; Zhang, Y.; Zhang, R.; Kai, G. Molecular characterization and expression of 1-deoxy-d-xylulose 5-phosphate reductoisomerase (DXR) gene from Salvia miltiorrhiza. Acta Physiol. Plant 2009, 31, 1015–1022. [Google Scholar] [CrossRef]
- Sun, Z.; Cunningham, F.X.; Gantt, E. Differential expression of two isopentenyl pyrophosphate isomerases and enhanced carotenoid accumulation in a unicellular chlorophyte. Proc. Natl. Acad. Sci. USA 1998, 95, 11482–11488. [Google Scholar] [CrossRef] [PubMed]
- Estévez, J.M.; Cantero, A.; Romero, C.; Kawaide, H.; Jiménez, L.F.; Kuzuyama, T.; Seto, H.; Kamiya, Y.; León, P. Analysis of the expression of CLA1, a gene that encodes the 1-deoxy-xylulose-5-phosphate synthase of the 2-C-methyl-d-erythritol-4-phosphate pathway in Arabidopsis. Plant. Physiol. 2000, 124, 95–104. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Adhikari, M.N.; Liu, H.; Xu, H.; He, G.; Zhan, R.; Wei, J.; Chen, W. Characterization and functional analysis of the genes encoding 1-deoxy-d-xylulose-5-phosphate reductoisomerase and 1-deoxy-d-xylulose-5-phosphate synthase, the two enzymes in the MEP pathway, from Amomum villosum Lour. Mol. Biol. Rep. 2012, 39, 8287–8296. [Google Scholar] [CrossRef] [PubMed]
- Xu, Y.; Liu, J.; Liang, L.; Yang, X.; Zhang, Z.; Gao, Z.; Sui, C.; Wei, J. Molecular cloning and characterization of three cDNAs encoding 1-deoxy-d-xylulose-5-phosphate synthase in Aquilaria sinensis (Lour.) Gilg. Plant Physiol. Biochem. 2014, 82, 133–141. [Google Scholar] [CrossRef] [PubMed]
- Gupta, P.; Agarwal, A.V.; Akhtar, N.; Sangwan, R.S.; Singh, S.P.; Trivedi, P.K. Cloning and characterization of 2-C-methyl-d-erythritol-4-phosphate pathway genes for isoprenoid biosynthesis from Indian ginseng, Withania somnifera. Protoplasma 2013, 250, 285–295. [Google Scholar] [CrossRef] [PubMed]
- Yao, H.; Gong, Y.; Zuo, K.; Ling, H.; Qiu, C.; Zhang, F.; Wang, Y.; Pi, Y.; Liu, X.; Sun, X.; Tang, K. Molecular cloning, expression profiling and functional analysis of a DXR gene encoding 1-deoxy-d-xylulose-5-phosphate reductoisomerase from Camptotheca acuminate. J. Plant. Physiol. 2008, 165, 203–213. [Google Scholar] [CrossRef] [PubMed]
- Blas, A.L.; Ming, R.; Liu, Z.; Veatch, O.J.; Paull, R.E.; Moore, P.H.; Yu, Q. Cloning of the papaya chromoplast-specific lycopene β-Cyclase, CpCYC-b, controlling fruit flesh color reveals conserved microsynteny and a recombination hot spot. Plant Physiol. 2010, 152, 2013–2022. [Google Scholar] [CrossRef] [PubMed]
- Gong, Y.; Liao, Z.; Guo, B. Molecular cloning and expression profile analysis of Ginkgo biloba DXS gene encoding 1-deoxy-d-xylulose 5-phosphate synthase, the first committed enzyme of the 2-C-methyl-d-erythritol 4-phosphate pathway. Planta Med. 2006, 72, 329–335. [Google Scholar] [CrossRef] [PubMed]
- Enfissi, E.; Fraser, P.D.; Lois, L.M. Metabolic engineering of the mevalonate and non-mevalonate isopentenyl diphosphate-forming pathways for the production of health-promoting isoprenoids in tomato. Plant Biotechnol. J. 2005, 3, 17–27. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez-Concepción, M.; Boronat, A. Elucidation of the methylerythritol phosphate pathway for isoprenoid biosynthesis in bacteria and plastids. A metabolic milestoneachieved through genomics. Plant Physiol. 2002, 130, 1079–1089. [Google Scholar] [CrossRef] [PubMed]
- Paetzold, H.; Garms, S.; Bartram, S. The isogene 1-deoxy-d-xylulose- 5-phosphate synthase 2 controls isoprenoid profiles, precursor pathway allocation, and density of tomato trichomes. Mol. Plant 2010, 3, 904–916. [Google Scholar] [CrossRef] [PubMed]
- Cordoba, E.; Porta, H.; Arroyo, A.; San Román, C.; Medina, L.; Rodríguez-Concepción, M.; León, P. Functional characterization of the three genes encoding 1-deoxy-d-xylulose 5-phosphate synthase in maize. J. Exp. Bot. 2011, 62, 2023–2038. [Google Scholar] [CrossRef] [PubMed]
- Hasunuma, T.; Takeno, S.; Hayashi, S. Overexpression of 1-deoxy-d-xylulose-5-phosphate reductoisomerase gene in chloroplast contributes to increment of isoprenoid production. J. Biosci. Bioeng. 2008, 105, 518–526. [Google Scholar] [CrossRef] [PubMed]
- Rohmer, M. Mevalonate-independent methylerythritol phosphate pathway for isoprenoid biosynthesis. Elucidation and distribution. Pure Appl. Chem. 2003, 75, 375–388. [Google Scholar] [CrossRef]
- Banerjee, A.; Sharkey, T.D. Methylerythritol 4-phosphate (MEP) pathway metabolic regulation. Nat. Prod. Rep. 2014, 31, 1043–1055. [Google Scholar] [CrossRef] [PubMed]
- Banerjee, A.; Wu, Y.; Banerjee, R.; Li, Y.; Yan, H.; Sharkey, T.D. Feedback inhibition of deoxy-d-xylulose-5-phosphate synthase regulates the methylerythritol-4-phosphate pathway. J. Biol. Chem. 2013, 288, 16926–16936. [Google Scholar] [CrossRef] [PubMed]
- Ghirardo, A.; Wright, L.P.; Bi, Z.; Rosenkranz, M.; Pulido, P.; Rodriguez-Concepcion, M.; Niinemets, U.; Bruggemann, N.; Gershenzon, J.; Schnitzler, J.P. Metabolic flux analysis of plastidic isoprenoid biosynthesis in poplar leaves emitting and nonemitting isoprene. Plant Physiol. 2014, 165, 37–51. [Google Scholar] [CrossRef] [PubMed]
- Patel, H.; Nemeria, N.S.; Brammer, L.A.; Meyers, C.L.; Jordan, F. Observation of thiamin-bound intermediates and microscopic rate constants for their interconversion on 1-deoxy-d-xylulose-5-phosphate synthase: 600-Fold rate acceleration of pyruvate decarboxylation by d-glyceraldehyde-3-phosphate. J. Am. Chem. Soc. 2012, 134, 18374–18379. [Google Scholar] [CrossRef] [PubMed]
- Wright, L.P.; Rohwer, J.M.; Ghirardo, A.; Hammerbacher, A.; Ortiz-Alcaide, M.; Raguschke, B.; Schnitzler, J.P.; Gershenzon, J.; Phillips, M.A. Deoxyxylulose 5-phosphate synthase controls flux through the methylerythritol 4-phosphate pathway in Arabidopsis. Plant Physiol. 2014, 165, 1488–1504. [Google Scholar] [CrossRef] [PubMed]
- Wu, C.M. Studies on Chemical Constituents and Muti-Components Content Determination of Tripterygium wilfordii. Doctoral thesis, The Second Military Medical University, Shanghai, China, May 2010. [Google Scholar]
- Shi, M.; Luo, X.; Ju, G.; Yu, X.; Hao, X.; Huang, Q.; Xiao, J.; Cui, L.; Kai, G. Increased accumulation of the cardio-cerebrovascular disease treatment drug tanshinone in Salvia miltiorrhiza hairy roots by the enzymes 3-hydroxy-3-methylglutaryl CoA reductase and 1-deoxy-d-xylulose 5-phosphate reductoisomerase. Funct. Integr. Genom. 2014, 14, 603–615. [Google Scholar] [CrossRef] [PubMed]
- Brückner, K.; Tissier, A. High-level diterpene production by transient expression in Nicotiana benthamiana. Plant Methods 2013, 9, 46–55. [Google Scholar] [CrossRef] [PubMed]
- Su, P.; Cheng, Q.; Wang, X.; Cheng, X.; Zhang, M.; Tong, Y.; Li, F.; Gao, W.; Huang, L. Characterization of eight terpenoids from tissue cultures of the Chinese herbal plant, Tripterygium wilfordii, by high-performance liquid chromatography coupled with electrosprayionization tandem mass spectrometry. Biomed. Chromatogr. 2014, 28, 1183–1192. [Google Scholar] [CrossRef] [PubMed]
- Zhu, C.; Chen, X.; Guo, J.; Miao, G.; Feng, J.; Zhang, X. Cloning and expression analysis of 1-deoxy-d-xylulose 5-phosphate reductoisomerase gene (DXR) in Tripterygiun wilfordii. Agric. Biotechnol. 2014, 22, 298–308. [Google Scholar]
- Cunningham, F.X.; Pogson, B.; Sun, Z.; McDonald, K.A.; DellaPenna, D.; Gantt, E. Functional analysis of the β and ε lycopene cyclase enzymes of arabidopsis reveals a mechanism for control of cyclic carotenoid formation. Plant Cell 1996, 8, 1613–1626. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Cunningham, F.X.; Lafond, T.P.; Gantt, E. Evidence of a role for lytB in the nonmevalonate pathway of isoprenoid biosynthesis. J. Bacteriol. 2000, 182, 5841–5848. [Google Scholar] [CrossRef] [PubMed]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−∆∆Ct method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tong, Y.; Su, P.; Zhao, Y.; Zhang, M.; Wang, X.; Liu, Y.; Zhang, X.; Gao, W.; Huang, L. Molecular Cloning and Characterization of DXS and DXR Genes in the Terpenoid Biosynthetic Pathway of Tripterygium wilfordii. Int. J. Mol. Sci. 2015, 16, 25516-25535. https://doi.org/10.3390/ijms161025516
Tong Y, Su P, Zhao Y, Zhang M, Wang X, Liu Y, Zhang X, Gao W, Huang L. Molecular Cloning and Characterization of DXS and DXR Genes in the Terpenoid Biosynthetic Pathway of Tripterygium wilfordii. International Journal of Molecular Sciences. 2015; 16(10):25516-25535. https://doi.org/10.3390/ijms161025516
Chicago/Turabian StyleTong, Yuru, Ping Su, Yujun Zhao, Meng Zhang, Xiujuan Wang, Yujia Liu, Xianan Zhang, Wei Gao, and Luqi Huang. 2015. "Molecular Cloning and Characterization of DXS and DXR Genes in the Terpenoid Biosynthetic Pathway of Tripterygium wilfordii" International Journal of Molecular Sciences 16, no. 10: 25516-25535. https://doi.org/10.3390/ijms161025516
APA StyleTong, Y., Su, P., Zhao, Y., Zhang, M., Wang, X., Liu, Y., Zhang, X., Gao, W., & Huang, L. (2015). Molecular Cloning and Characterization of DXS and DXR Genes in the Terpenoid Biosynthetic Pathway of Tripterygium wilfordii. International Journal of Molecular Sciences, 16(10), 25516-25535. https://doi.org/10.3390/ijms161025516





