Functional Characterization of NIPBL Physiological Splice Variants and Eight Splicing Mutations in Patients with Cornelia de Lange Syndrome
Abstract
:1. Introduction
2. Results
2.1. Clinical Findings
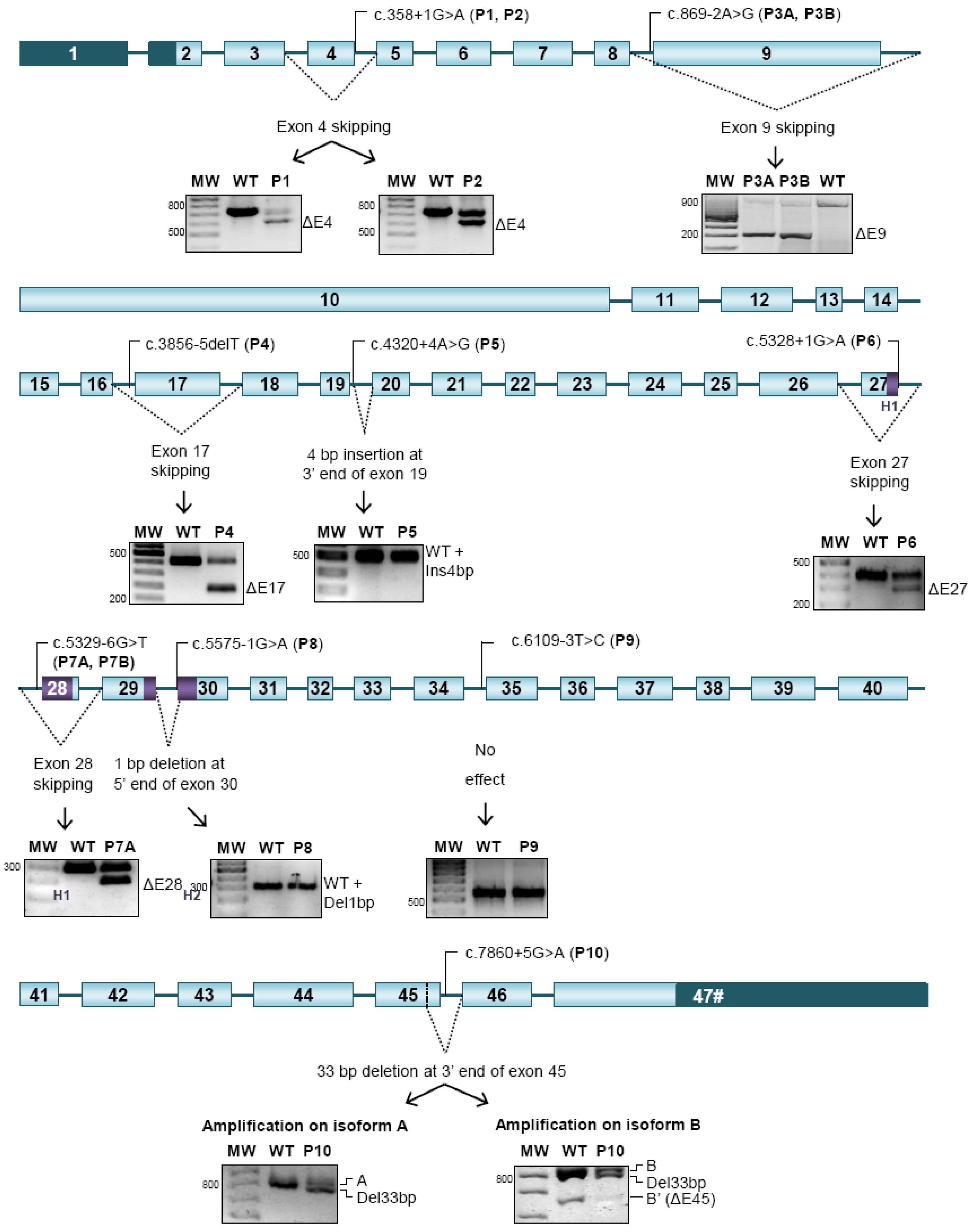
2.2. Aberrant Splicing Caused by NIPBL-Mutations
| Patient | Mutation | Intron | mRNA Change | In Silico Analysis | Predicted Protein Change | Clinical Data |
|---|---|---|---|---|---|---|
| P1 | c.358+1G>A | 4 | Exon 4 skipping (128 bp) | Donor site disruption | p.(Ile77Metfs*5) | 2-year-old boy with pre and postnatal growth retardation, developmental delay, typical facial features, heart valve stenosis and a feeding disorder. |
| P2 | c.358+1G>A | 4 | Exon 4 skipping (128 bp) | Donor site disruption | p.(Ile77Metfs*5) | 13-year-old boy with pre and postnatal growth retardation, intellectual disability with speech and motor delay, typical facial features, aortic stenosis and hirsutism. |
| P3A | c.869-2A>G | 8 | Exon 9 skipping (627 bp) | Acceptor site disruption | p.(Gly290_Lys498del) | 3-year-old boy with pre and postnatal growth retardation, intellectual disability with speech and motor delay, typical facial features, autistic spectrum disorders, a feeding disorder, cryptorchidism and hirsutism. |
| P3B | c.869-2A>G | 8 | Exon 9 skipping (627 bp) | Acceptor site disruption | p.(Gly290_Lys498del) | 36-year-old female with postnatal growth retardation, typical facial features, a feeding disorder, minor skeletal anomalies and hirsutism. |
| P4 | c.3856-5delT [18] | 16 | Exon 17 skipping (232 bp) | Acceptor site disruption | p.(Asn1286Glufs*3) | 23-year-old male with pre and postnatal growth retardation, typical features, intellectual disability, pyloric stenosis and a feeding disorder. |
| P5 | c.4320+4A>G | 19 | Insertion of 4 pb at 3' end of exon 19 | Donor site disruption and creation of a new cryptic donor site | p.(Phe1442Serfs*5) | 7-year-old boy with postnatal growth retardation, intellectual disability with speech delay, typical facial features, deafness, GERD with a feeding disorder and small hands. |
| P6 | c.5328+1G>A | 27 | Exon 27 skipping (103 bp) | Donor site disruption | p.(Met1743Serfs*16) | 4-year-old boy with pre and postnatal growth retardation, developmental delay, typical facial features, pulmonar stenosis, cryptorchidism, bilateral aplasia of the ulna, dysplasia of distal humerus and monodactyly. |
| P7A | c.5329-6T>G | 27 | Exon 28 skipping (99 bp) | Acceptor site disruption | p.(Ile1777_Arg1809del) | 9-year-old boy with has pre and postnatal growth retardation, intellectual disability with speech delay, typical facial features, hirsutism and small hands. |
| P7B | c.5329-6T>G | 27 | Exon 28 skipping (99 bp) | Acceptor site disruption | p.(Ile1777_Arg1809del) | 40-year-old male with typical features, intellectual disability, hearing loss, ventricular septal defect and crytorchidism. |
| P8 | c.5575-1G>A | 29 | Loss of 1 pb at 5' end of exon 30 | Acceptor site disruption and creation of a new acceptor site | p.(Asp1859Ilefs*9) | 1-year-old girl with pre and postnatal growth retardation, typical facial features, left ventricular hypertrophy, feeding disorder, horseshoe kidney, hirsutism, right amelia and small feet. |
| P9 | c.6109-3T>C [16,17,18] | 34 | No effect | Maintenance of acceptor site strengh | No effect | 26-year-old female with pre and postnatal growth retardation, intellectual disability with speech delay, typical facial features, deafness, feeding disorder, hirsutism, small hands and feet. |
| P10 | c.7860+5G>A | 45 | Loss of 33 pb at 3' end of exon 45 | Donor site disruption and activation of a cryptic donor site | p.(Val2610_Asp2620del) | 8-year-old girl with speech delay, typical facial features and a feeding disorder. |
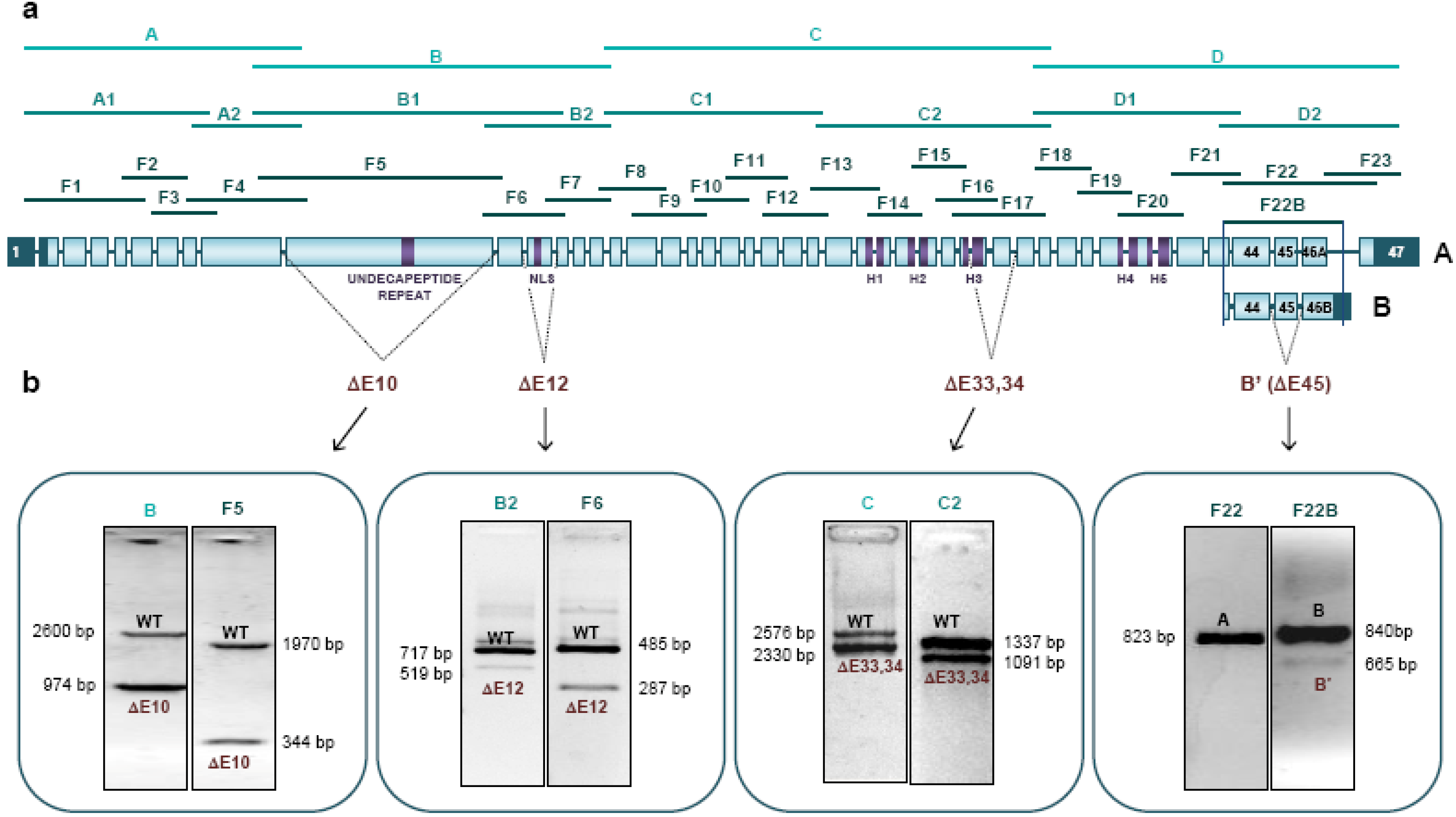
2.3. New Physiological Splicing Variants of NIPBL
| Splicing Variant | mRNA Change | In Silico Analysis | Predicted Protein Change | |
|---|---|---|---|---|
| ΔE10 | Exon 10 skipping (1646 bp) c.1496_3121del | Exon 10 weak acceptor site | p.(Asp499_Lys1040del) | |
| ΔE12 | Exon 12 skipping (198 bp) c.3305_3502del | Exon 12 weak acceptor site | p.(Ser1102_Val1168delinsPhe) | |
| ΔE33,34 | Exons 33 and 34 skipping (246 bp) c.5863_6108del | Exon 33 weak acceptor site | p.(Leu1955_Ser2036del) | |
| B | Exon 47 skipping (365 bp)and introduction of 211 bp of intron 46 c.8050_8415delins42 | Exon 47 weak acceptor site | p.(Ser2864_Ser2904delinsValfs*13) | |
| B’(ΔE45) | Exons 45 (175 bp) and 47 (365 bp) skipping and introduction of 211 bp of intron 46 c.[7686_7861del(;)8050_8415delins42] | Exons 45 and 47 weak acceptor site | p.(Lys2563Serfs*63) | |
3. Discussion
4. Experimental Section
4.1. Patients and Controls
4.2. DNA Extraction and Sequence Analysis
4.3. RNA Extraction and cDNA Synthesis
4.4. Identification of Splicing Variants
4.5. In Silico Splicing Analysis
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Kline, A.D.; Krantz, I.D.; Sommer, A.; Kliewer, M.; Jackson, L.G.; FitzPatrick, D.R.; Levin, A.V.; Selicorni, A. Cornelia de Lange syndrome: Clinical review, diagnostic and scoring systems, and anticipatory guidance. Am. J. Med. Genet. A 2007, 143A, 1287–1296. [Google Scholar] [CrossRef]
- Krantz, I.D.; McCallum, J.; DeScipio, C.; Kaur, M.; Gillis, L.A.; Yaeger, D.; Jukofsky, L.; Wasserman, N.; Bottani, A.; Morris, C.A.; et al. Cornelia de Lange syndrome is caused by mutations in NIPBL, the human homolog of Drosophila melanogaster Nipped-B. Nat. Genet. 2004, 36, 631–635. [Google Scholar]
- Remeseiro, S.; Losada, A. Cohesin, a chromatin engagement ring. Curr. Opin. Cell. Biol. 2013, 25, 63–71. [Google Scholar] [CrossRef]
- Pié, J.; Gil-Rodríguez, M.C.; Ciero, M.; López-Viñas, E.; Ribate, M.P.; Arnedo, M.; Deardorff, M.A.; Puisac, B.; Legarreta, J.; de Karam, J.C.; et al. Mutations and variants in the cohesion factor genes NIPBL, SMC1A, and SMC3 in a cohort of 30 unrelated patients with Cornelia de Lange syndrome. Am. J. Med. Genet. A 2010, 152A, 924–929. [Google Scholar] [CrossRef]
- Wierzba, J.; Gil-Rodríguez, M.C.; Polucha, A.; Puisac, B.; Arnedo, M.; Teresa-Rodrigo, M.E.; Winnicka, D.; Hegardt, F.G.; Ramos, F.J.; Limon, J.; et al. Cornelia de Lange syndrome with NIPBL mutation and mosaic Turner syndrome in the same individual. BMC Med. Genet. 2012. [Google Scholar] [CrossRef]
- Musio, A.; Selicorni, A.; Focarelli, M.L.; Gervasini, C.; Milani, D.; Russo, S.; Vezzoni, P.; Larizza, L. X-linked Cornelia de Lange syndrome owing to SMC1L1 mutations. Nat. Genet. 2006, 38, 528–530. [Google Scholar] [CrossRef]
- Deardorff, M.A.; Kaur, M.; Yaeger, D.; Rampuria, A.; Korolev, S.; Pie, J.; Gil-Rodríguez, C.; Arnedo, M.; Loeys, B.; Kline, A.D.; et al. Mutations in cohesin complex members SMC3 and SMC1A cause a mild variant of Cornelia de Lange syndrome with predominant mental retardation. Am. J. Hum. Genet. 2007, 80, 485–494. [Google Scholar] [CrossRef]
- Deardorff, M.A.; Wilde, J.J.; Albrecht, M.; Dickinson, E.; Tennstedt, S.; Braunholz, D.; Mönnich, M.; Yan, Y.; Xu, W.; Gil-Rodríguez, M.C.; et al. RAD21 mutations cause a human cohesinopathy. Am. J. Hum. Genet. 2012, 90, 1014–1027. [Google Scholar] [CrossRef]
- Deardorff, M.A.; Bando, M.; Nakato, R.; Watrin, E.; Itoh, T.; Minamino, M.; Saitoh, K.; Komata, M.; Katou, Y.; Clark, D.; et al. HDAC8 mutations in Cornelia de Lange syndrome affect the cohesin acetylation cycle. Nature 2012, 489, 313–317. [Google Scholar] [CrossRef]
- Vandenbroucke, I.I.; Vandesompele, J.; Paepe, A.D.; Messiaen, L. Quantification of splice variants using real-time PCR. Nucleic Acids Res. 2001, 29, e68. [Google Scholar] [CrossRef]
- Strachan, T. Cornelia de Lange syndrome and the link between chromosomal function, DNA repair and developmental gene regulation. Curr. Opin. Genet. Dev. 2005, 15, 258–264. [Google Scholar] [CrossRef]
- Tonkin, E.T.; Wang, T.J.; Lisgo, S.; Bamshad, M.J.; Strachan, T. NIPBL, encoding a homolog of fungal Scc2-type sister chromatid cohesion proteins and fly Nipped-B, is mutated in Cornelia de Lange syndrome. Nat. Genet. 2004, 36, 636–641. [Google Scholar] [CrossRef]
- Gillespie, P.J.; Hirano, T. Scc2 couples replication licensing to sister chromatid cohesion in Xenopus egg extracts. Curr. Biol. 2004, 14, 1598–1603. [Google Scholar] [CrossRef]
- Leiden Open Variant Database. Available online: http://www.dmd.nl (accessed on 10 April 2014).
- Mannini, L.; Cucco, F.; Quarantotti, V.; Krantz, I.D.; Musio, A. Mutation spectrum and genotype–phenotype correlation in Cornelia de Lange syndrome. Hum. Mutat. 2013, 34, 1589–1596. [Google Scholar] [CrossRef]
- Bhuiyan, Z.A.; Klein, M.; Hammond, P.; van Haeringen, A.; Mannens, M.M.; van Berckelaer-Onnes, I.; Hennekam, R.C. Genotype–phenotype correlations of 39 patients withCorneliade Lange syndrome: The Dutch experience. J. Med. Genet. 2006, 43, 568–575. [Google Scholar]
- Gillis, L.A.; McCallum, J.; Kaur, M.; DeScipio, C.; Yaeger, D.; Mariani, A.; Kline, A.D.; Li, H.H.; Devoto, M.; Jackson, L.G.; et al. NIPBL mutational analysis in 120 individuals with Cornelia de Lange syndrome and evaluation of genotype–phenotype correlations. Am. J. Hum. Genet. 2004, 75, 610–623. [Google Scholar] [CrossRef]
- Selicorni, A.; Russo, S.; Gervasini, C.; Castronovo, P.; Milani, D.; Cavalleri, F.; Bentivegna, A.; Masciadri, M.; Domi, A.; Divizia, M.T.; et al. Clinical score of 62 Italian patients with Cornelia de Lange syndrome and correlations with the presence and type of NIPBL mutation. Clin. Genet. 2007, 72, 98–108. [Google Scholar] [CrossRef]
- Splice Site Prediction by Neural Network. Available online: http//:www.fruitfly.org/seq_tools/splice.html (accessed on 10 April 2014).
- Human Splicing Finder. Available online: http//:www.umd.be/HSF/ (accessed on 10 April 2014).
- Khan, S.G.; Muniz-Medina, V.; Shahlavi, T.; Baker, C.C.; Inui, H.; Ueda, T.; Emmert, S.; Schneider, T.D.; Kraemer, K.H. The human XPC DNA repair gene: Arrangement, splice site information content and influence of a single nucleotide polymorphism in a splice acceptor site on alternative splicing and function. Nucleic Acids Res. 2002, 30, 3624–3631. [Google Scholar] [CrossRef]
- Casals, N.; Pié, J.; Casale, C.H.; Zapater, N.; Ribes, A.; Castro-Gago, M.; Rodriguez-Segade, S.; Wanders, R.J.; Hegardt, F.G. A two-base deletion in exon 6 of the 3-hydroxy-3-methylglutaryl coenzyme A lyase (HL) gene producing the skipping of exons 5 and 6 determines 3-hydroxy-3-methylglutaric aciduria. J. Lipid Res. 1997, 38, 2303–2313. [Google Scholar]
- Puisac, B.; Teresa-Rodrigo, M.E.; Arnedo, M.; Gil-Rodríguez, M.C.; Pérez-Cerdá, C.; Ribes, A.; Pié, A.; Bueno, G.; Gómez-Puertas, P.; Pié, J. Analysis of aberrant splicing and nonsense-mediated decay of the stop codon mutations c.109G>T and c.504_505delCT in 7 patients with HMG-CoA lyase deficiency. Mol. Genet. Metab. 2013, 108, 232–240. [Google Scholar]
- Yan, J.; Saifi, G.M.; Wierzba, T.H.; Withers, M.; Bien-Willner, G.A.; Limon, J.; Stankiewicz, P.; Lupski, J.R.; Wierzba, J. Mutational and genotype-phenotype correlation analyses in 28 Polish patients with Cornelia de Lange syndrome. Am. J. Med. Genet. A 2006, 140, 1531–1541. [Google Scholar]
- Jahnke, P.; Xu, W.; Wülling, M.; Albrecht, M.; Gabriel, H.; Gillessen-Kaesbach, G.; Kaiser, F.J. The Cohesin loading factor NIPBL recruits histone deacetylases to mediate local chromatin modifications. Nucleic Acids. Res. 2008, 36, 6450–6458. [Google Scholar]
- Cartegni, L.; Chew, S.L.; Krainer, A.R. Listening to silence and understanding nonsense: Exonic mutations that affect splicing. Nat. Rev. Genet. 2002, 3, 285–298. [Google Scholar] [CrossRef]
- David, A.; Miraki-Moud, F.; Shaw, N.J.; Savage, M.O.; Clark, A.J.L.; Metherell, L.A. Identification and characterisation of a novel GHR defect disrupting the polypyrimidine tract and resulting in GH insensitivity. Eur. J. Endocrinol. 2010, 162, 37–42. [Google Scholar] [CrossRef]
- Baralle, D.; Baralle, M. Splicing in action: Assessing disease causing sequence changes. J. Med. Genet. 2005, 42, 737–748. [Google Scholar] [CrossRef]
- Krawczak, M.; Thomas, N.S.; Hundrieser, B.; Mort, M.; Wittig, M.; Hampe, J.; Cooper, D.N. Single base-pair substitutions in exon–intron junctions of human genes: Nature, distribution, and consequences for mRNA splicing. Hum. Mutat. 2007, 28, 150–158. [Google Scholar] [CrossRef]
- Athanasakis, E.; Fabretto, A.; Faletra, F.; Mocenigo, M.; Morgan, A.; Gasparini, P. Two novel COH1 mutations in an Italian patient with Cohen syndrome. Mol. Syndromol. 2012, 3, 30–33. [Google Scholar]
- Watanabe, T.; Hanawa, H.; Suzuki, T.; Jiao, S.; Yoshida, K.; Ogura, M.; Ohno, Y.; Hayashi, Y.; Ito, M.; Kashimura, T.; et al. A mutant mRNA expression in an endomyocardial biopsy sample obtained from a patient with a cardiac variant of fabry disease caused by a novel acceptor splice site mutation in the invariant AG of intron 5 of the α-galactosidase A gene. Intern. Med. 2013, 52, 777–780. [Google Scholar] [CrossRef]
- Schoumans, J.; Wincent, J.; Barbaro, M.; Djureinovic, T.; Maguire, P.; Forsberg, L.; Staaf, J.; Thuresson, A.C.; Borg, A.; Nordgren, A.; et al. Comprehensive mutational analysis of a cohort of Swedish Cornelia de Lange syndrome patients. Eur. J. Hum. Genet. 2007, 15, 143–149. [Google Scholar] [CrossRef]
- Oliveira, J.; Dias, C.; Redeker, E.; Costa, E.; Silva, J.; Reis Lima, M.; den Dunnen, J.T.; Santos, R. Development of NIPBL locus-specific database using LOVD: From novel mutations to further genotype–phenotype correlations in Cornelia de Lange syndrome. Hum. Mutat. 2010, 31, 1216–1222. [Google Scholar] [CrossRef]
- Human Genome Variation Society. Available online: http://www.hgvs.org/ (accessed on 10 April 2014).
- Mutalyzer. Available online: https://mutalyzer.nl (accessed on 10 April 2014).
- Reese, M.G.; Eeckman, F.H.; Kulp, D.; Haussler, D. Improvedsplice site detection in Genie. J. Comput. Biol. 1997, 4, 311–323. [Google Scholar] [CrossRef]
- Desmet, F.O.; Hamroun, D.; Lalande, M.; Collod-Béroud, G.; Claustres, M.; Béroud, C. Human Splicing Finder: An online bioinformatics tool to predict splicing signals. Nucleic Acids Res. 2009, 37, e67. [Google Scholar] [CrossRef]
- Yeo, G.; Burge, C.B. Maximum entropy modeling of short sequence motifs with applications to RNA splicing signals. J. Comput. Biol. 2004, 11, 377–394. [Google Scholar] [CrossRef]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Teresa-Rodrigo, M.E.; Eckhold, J.; Puisac, B.; Dalski, A.; Gil-Rodríguez, M.C.; Braunholz, D.; Baquero, C.; Hernández-Marcos, M.; De Karam, J.C.; Ciero, M.; et al. Functional Characterization of NIPBL Physiological Splice Variants and Eight Splicing Mutations in Patients with Cornelia de Lange Syndrome. Int. J. Mol. Sci. 2014, 15, 10350-10364. https://doi.org/10.3390/ijms150610350
Teresa-Rodrigo ME, Eckhold J, Puisac B, Dalski A, Gil-Rodríguez MC, Braunholz D, Baquero C, Hernández-Marcos M, De Karam JC, Ciero M, et al. Functional Characterization of NIPBL Physiological Splice Variants and Eight Splicing Mutations in Patients with Cornelia de Lange Syndrome. International Journal of Molecular Sciences. 2014; 15(6):10350-10364. https://doi.org/10.3390/ijms150610350
Chicago/Turabian StyleTeresa-Rodrigo, María E., Juliane Eckhold, Beatriz Puisac, Andreas Dalski, María C. Gil-Rodríguez, Diana Braunholz, Carolina Baquero, María Hernández-Marcos, Juan C. De Karam, Milagros Ciero, and et al. 2014. "Functional Characterization of NIPBL Physiological Splice Variants and Eight Splicing Mutations in Patients with Cornelia de Lange Syndrome" International Journal of Molecular Sciences 15, no. 6: 10350-10364. https://doi.org/10.3390/ijms150610350
APA StyleTeresa-Rodrigo, M. E., Eckhold, J., Puisac, B., Dalski, A., Gil-Rodríguez, M. C., Braunholz, D., Baquero, C., Hernández-Marcos, M., De Karam, J. C., Ciero, M., Santos-Simarro, F., Lapunzina, P., Wierzba, J., Casale, C. H., Ramos, F. J., Gillessen-Kaesbach, G., Kaiser, F. J., & Pié, J. (2014). Functional Characterization of NIPBL Physiological Splice Variants and Eight Splicing Mutations in Patients with Cornelia de Lange Syndrome. International Journal of Molecular Sciences, 15(6), 10350-10364. https://doi.org/10.3390/ijms150610350





