Antenna-Specific Glutathione S-Transferase in Male Silkmoth Bombyx mori
Abstract
:1. Introduction
2. Results
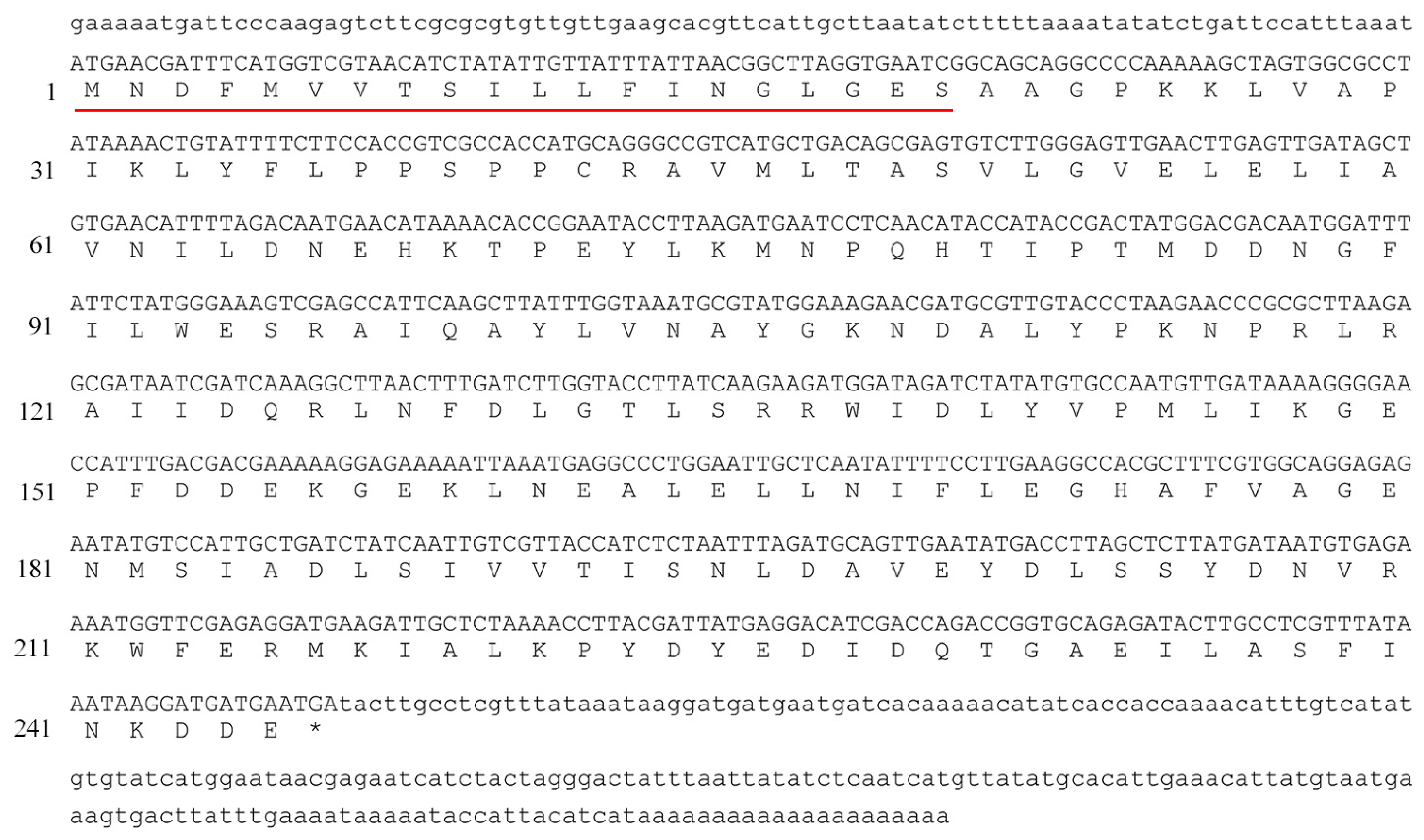
2.1. Cloning and Analysis of Bmgstd4
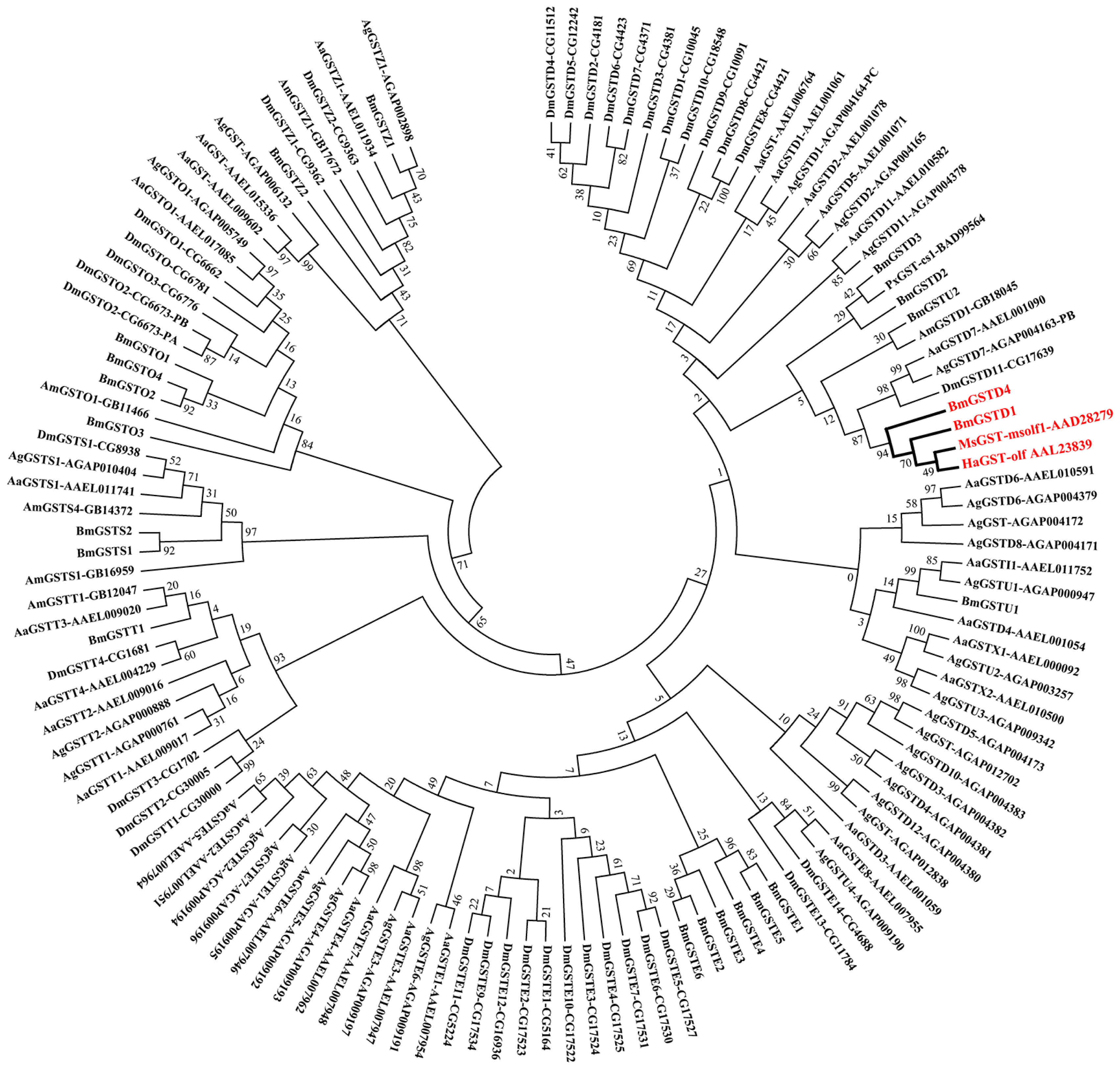
2.2. Phylogenetic Analysis of BmGSTD4 with Other Insect GSTs
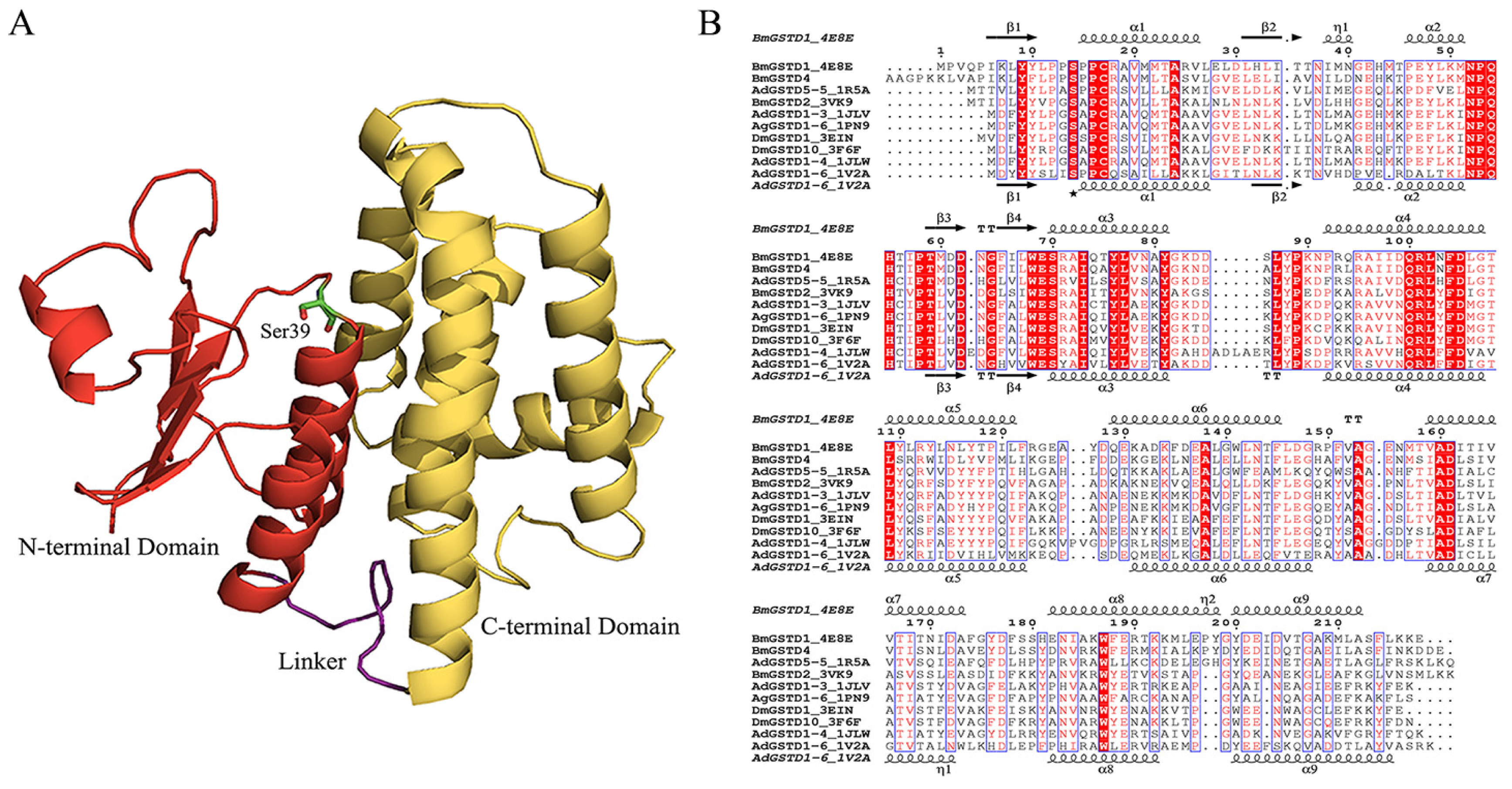
2.3. Structural Characterization of BmGSTD4
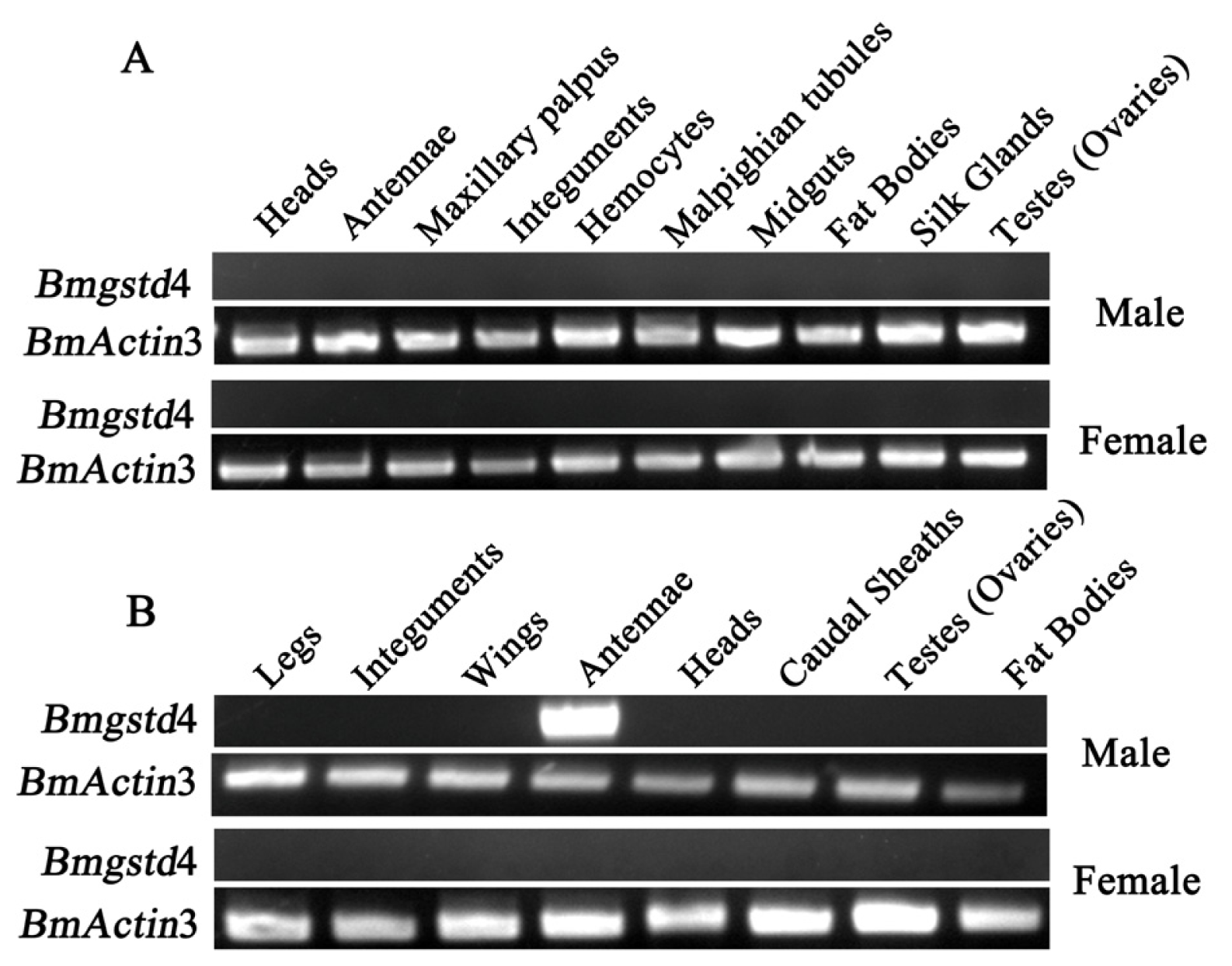
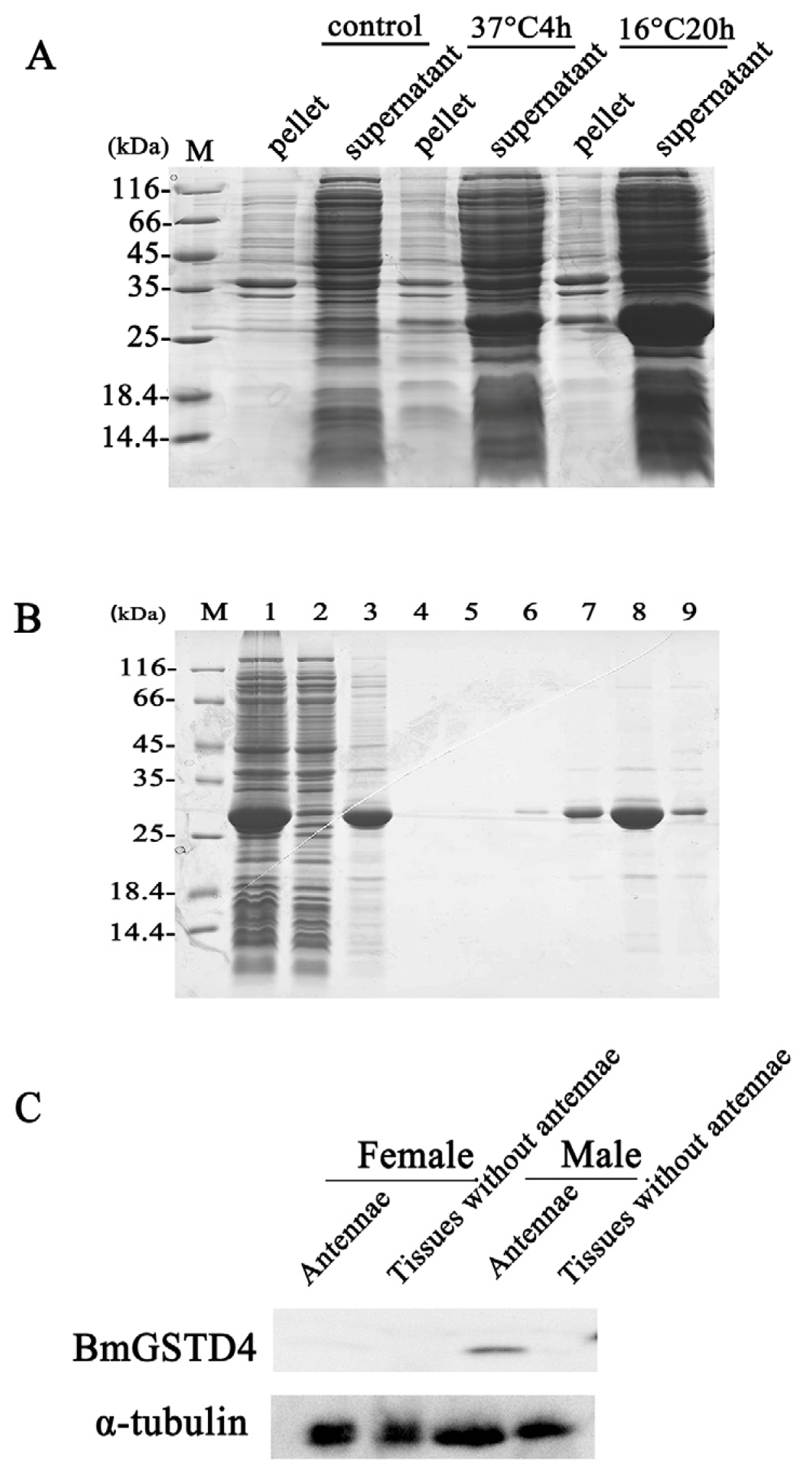
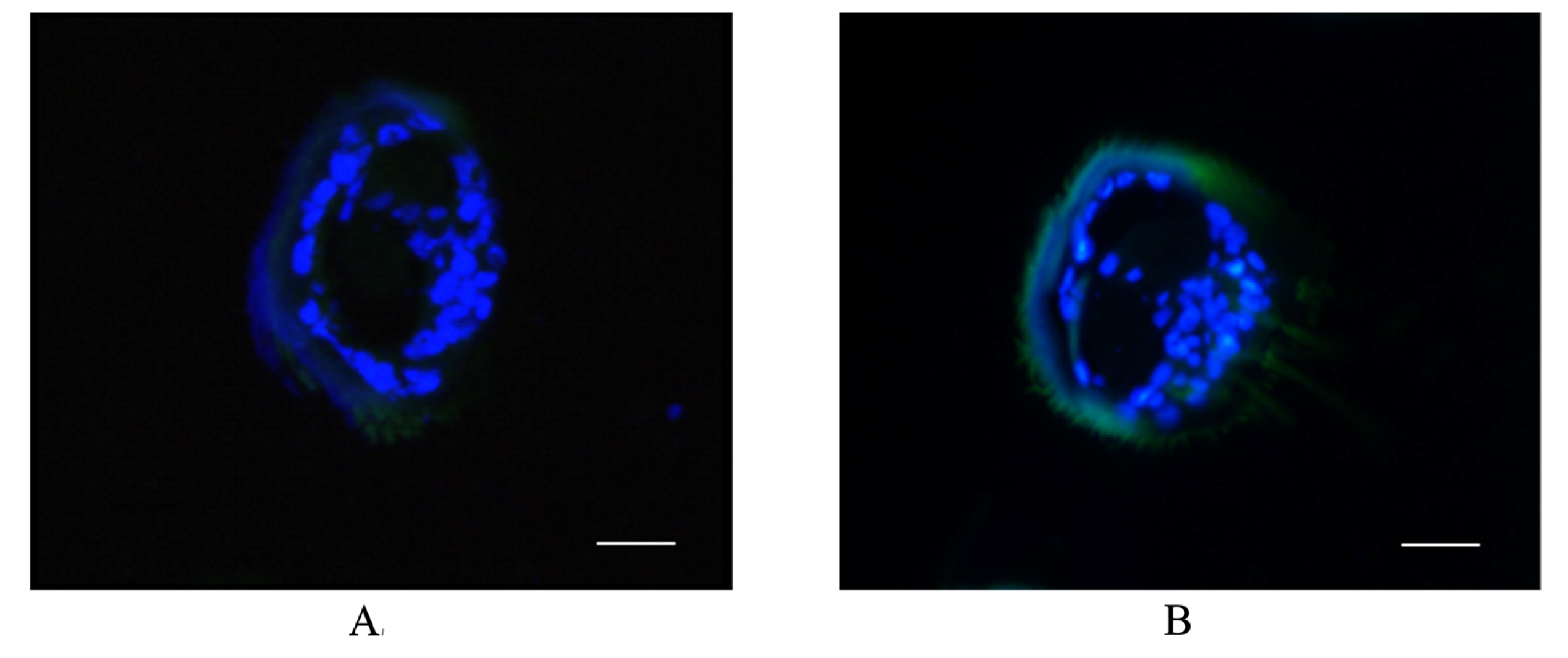
2.4. Antenna-Specific of BmGSTD4
2.5. Enzymatic Properties and GST Activity of BmGSTD4
3. Discussion
4. Experimental Section
4.1. Materials and Tissue Collection
4.2. Extraction of RNA and Synthesis of First-Strand cDNA
4.3. RT-PCR
4.4. Rapid Amplification of cDNA Ends of Bmgstd4
4.5. Sequence Analysis
4.6. Phylogenetic Analysis
4.7. Homology Modeling of BmGSTD4
4.8. Expression and Purification of Recombinant BmGSTD4
4.9. Western Blotting and Immunochemistry
4.10. Enzymatic Assays
5. Conclusions
Acknowledgments
Conflicts of Interest
References
- Hayes, J.D.; Flanagan, J.U.; Jowsey, I.R. Glutathione transferases. Annu. Rev. Pharmacol. Toxicol 2005, 45, 51–88. [Google Scholar]
- Armstrong, R.N. Structure, catalytic mechanism, and evolution of the glutathione transferases. Chem. Res. Toxicol 1997, 10, 2–18. [Google Scholar]
- Listowsky, I.; Abramovitz, M.; Homma, H.; Niitsu, Y. Intracellular binding and transport of hormones and xenobiotics by glutathione-S-transferases. Drug Metab. Rev 1988, 19, 305–318. [Google Scholar]
- Zhao, T.; Singhal, S.S.; Piper, J.T.; Cheng, J.; Pandya, U.; Clark-Wronski, J.; Awasthi, S.; Awasthi, Y.C. The role of human glutathione S-transferases hGSTA1–1 and hGSTA2–2 in protection against oxidative stress. Arch. Biochem. Biophys 1999, 367, 216–224. [Google Scholar]
- Johansson, A.S.; Mannervik, B. Human glutathione transferase A3–3, a highly efficient catalyst of double-bond isomerization in the biosynthetic pathway of steroid hormones. J. Biol. Chem 2001, 276, 33061–33065. [Google Scholar]
- Lo Bello, M.; Nuccetelli, M.; Caccuri, A.M.; Stella, L.; Parker, M.W.; Rossjohn, J.; McKinstry, W.J.; Mozzi, A.F.; Federici, G.; Polizio, F.; et al. Human glutathione transferase P1–1 and nitric oxide carriers; A new role for an old enzyme. J. Biol. Chem 2001, 276, 42138–42145. [Google Scholar]
- Dulhunty, A.; Gage, P.; Curtis, S.; Chelvanayagam, G.; Board, P. The glutathione transferase structural family includes a nuclear chloride channel and a ryanodine receptor calcium release channel modulator. J. Biol. Chem 2001, 276, 3319–3323. [Google Scholar]
- Bhargava, M.M.; Listowsky, I.; Arias, I.M. Ligandin. Bilirubin binding and glutathione-S-transferase activity are independent processes. J. Biol. Chem 1978, 253, 4112–4415. [Google Scholar]
- Mannervik, B.; Awasthi, Y.C.; Board, P.G.; Hayes, J.D.; di Ilio, C.; Ketterer, B.; Listowsky, I.; Morgenstern, R.; Muramatsu, M.; Pearson, W.R.; et al. Nomenclature for human glutathione transferases. Biochem. J 1992, 282 Pt 1, 305–306. [Google Scholar]
- Chelvanayagam, G.; Parker, M.W.; Board, P.G. Fly fishing for GSTs: A unified nomenclature for mammalian and insect glutathione transferases. Chem. Biol. Interact 2001, 133, 256–260. [Google Scholar]
- Enayati, A.A.; Ranson, H.; Hemingway, J. Insect glutathione transferases and insecticide resistance. Insect Mol. Biol 2005, 14, 3–8. [Google Scholar]
- Aceto, A.; Sacchetta, P.; Dragani, B.; Bucciarelli, T.; Angelucci, S.; Longo, V.; Gervasi, G.P.; Martini, F.; di Ilio, C. Glutathione transferase isoenzymes in olfactory and respiratory epithelium of cattle. Biochem. Pharmacol 1993, 46, 2127–2133. [Google Scholar]
- Banger, K.K.; Lock, E.A.; Reed, C.J. The characterization of glutathione S-transferases from rat olfactory epithelium. Biochem. J 1993, 290 Pt 1, 199–204. [Google Scholar]
- Ben-Arie, N.; Khen, M.; Lancet, D. Glutathione S-transferases in rat olfactory epithelium: Purification, molecular properties and odorant biotransformation. Biochem. J 1993, 292 Pt 2, 379–384. [Google Scholar]
- Rogers, M.E.; Jani, M.K.; Vogt, R.G. An olfactory-specific glutathione-S-transferase in the sphinx moth Manduca sexta. J. Exp. Biol 1999, 202, 1625–1637. [Google Scholar]
- Wang, G.-R.; Guo, Y.-Y.; Wu, K.-M. Cloning of a cDNA fragment of an antenna-specific gene in Helicoverpa armigera. Chin. J. Agric. Biotechnol 2004, 1, 37–43. [Google Scholar]
- Ono, H.; Ozaki, K.; Yoshikawa, H. Identification of cytochrome P450 and glutathione-S-transferase genes preferentially expressed in chemosensory organs of the swallowtail butterfly, Papilio xuthus L. Insect Biochem. Mol. Biol 2005, 35, 837–846. [Google Scholar]
- Espinoza, H.M.; Shireman, L.M.; McClain, V.; Atkins, W.; Gallagher, E.P. Cloning, expression and analysis of the olfactory glutathione S-transferases in coho salmon. Biochem. Pharmacol 2013, 85, 839–848. [Google Scholar]
- Tan, X.; Guo, P.; Hu, X.; Chen, Q.; Zhao, P.; Xia, Q.; Zhou, C. Southwest University, Chongqing, China. Structural and biochemical characterisation of Bombyx mori glutathione S-transferase BmGSTD1 preferentially expressed in the olfactory organs. Unpublished work. 2014. [Google Scholar]
- Kiefer, F.; Arnold, K.; Kunzli, M.; Bordoli, L.; Schwede, T. The SWISS-MODEL repository and associated resources. Nucleic Acids Res 2009, 37, D387–D392. [Google Scholar]
- Arnold, K.; Bordoli, L.; Kopp, J.; Schwede, T. The SWISS-MODEL workspace: A web-based environment for protein structure homology modelling. Bioinformatics 2006, 22, 195–201. [Google Scholar]
- Schwede, T.; Kopp, J.; Guex, N.; Peitsch, M.C. SWISS-MODEL: An automated protein homology-modeling server. Nucleic Acids Res 2003, 31, 3381–3385. [Google Scholar]
- Benkert, P.; Biasini, M.; Schwede, T. Toward the estimation of the absolute quality of individual protein structure models. Bioinformatics 2011, 27, 343–350. [Google Scholar]
- Allocati, N.; Federici, L.; Masulli, M.; di Ilio, C. Glutathione transferases in bacteria. FEBS J 2009, 276, 58–75. [Google Scholar]
- Sheehan, D.; Meade, G.; Foley, V.M.; Dowd, C.A. Structure, function and evolution of glutathione transferases: Implications for classification of non-mammalian members of an ancient enzyme superfamily. Biochem. J 2001, 360, 1–16. [Google Scholar]
- Armstrong, R.N. Mechanistic diversity in a metalloenzyme superfamily. Biochemistry 2000, 39, 13625–13632. [Google Scholar]
- Bernstein, F.C.; Koetzle, T.F.; Williams, G.J.; Meyer, E.F., Jr.; Brice, M.D.; Rodgers, J.R.; Kennard, O.; Shimanouchi, T.; Tasumi, M. The protein data bank: A computer-based archival file for macromolecular structures. J. Mol. Biol 1977, 112, 535–542. [Google Scholar]
- Yamamoto, K.; Ichinose, H.; Aso, Y.; Banno, Y.; Kimura, M.; Nakashima, T. Molecular characterization of an insecticide-induced novel glutathione transferase in silkworm. Biochim. Biophys. Acta 2011, 1810, 420–426. [Google Scholar]
- Yamamoto, K.; Shigeoka, Y.; Aso, Y.; Banno, Y.; Kimura, M.; Nakashima, T. Molecular and biochemical characterization of a Zeta-class glutathione S-transferase of the silkmoth. Pestic. Biochem. Phys 2009, 94, 30–35. [Google Scholar]
- Yamamoto, K.; Nagaoka, S.; Banno, Y.; Aso, Y. Biochemical properties of an omega-class glutathione S-transferase of the silkmoth Bombyx mori. Comp. Biochem. Physiol. C Toxicol. Pharmacol 2009, 149, 461–467. [Google Scholar]
- Yamamoto, K.; Zhang, P.B.; Banno, Y.; Fujii, H. Identification of a sigma-class glutathione-S-transferase from the silkworm Bombyx mori. J. Appl. Entomol 2006, 130, 515–522. [Google Scholar]
- Yamamoto, K.; Zhang, P.; Miake, F.; Kashige, N.; Aso, Y.; Banno, Y.; Fujii, H. Cloning, expression and characterization of theta-class glutathione S-transferase from the silkworm Bombyx mori. Comp. Biochem. Physiol. B Biochem. Mol. Biol 2005, 141, 340–346. [Google Scholar]
- Yamamoto, K.; Usuda, K.; Kakuta, Y.; Kimura, M.; Higashiura, A.; Nakagawa, A.; Aso, Y.; Suzuki, M. Structural basis for catalytic activity of a silkworm Delta-class glutathione transferase. Biochim. Biophys. Acta 2012, 1820, 1469–1474. [Google Scholar]
- Rybczynski, R.; Vogt, R.G.; Lerner, M.R. Antennal-specific pheromone-degrading aldehyde oxidases from the moths Antheraea polyphemus and Bombyx mori. J. Biol. Chem 1990, 265, 19712–19715. [Google Scholar]
- Raza, H. Dual localization of glutathione S-transferase in the cytosol and mitochondria: Implications in oxidative stress, toxicity and disease. FEBS J 2011, 278, 4243–4251. [Google Scholar]
- Zimniak, P.; Singh, S.P. Families of glutathione transferases. In Toxicology of Glutathione Transferases; Taylor & Francis CRC Press: Boca Raton, FL, USA, 2006; pp. 11–26. [Google Scholar]
- Litwack, G.; Ketterer, B.; Arias, I.M. Ligandin: A hepatic protein which binds steroids, bilirubin, carcinogens and a number of exogenous organic anions. Nature 1971, 234, 466–467. [Google Scholar]
- Yu, Q.; Lu, C.; Li, B.; Fang, S.; Zuo, W.; Dai, F.; Zhang, Z.; Xiang, Z. Identification, genomic organization and expression pattern of glutathione S-transferase in the silkworm Bombyx mori. Insect Biochem. Mol. Biol 2008, 38, 1158–1164. [Google Scholar]
- Gasteiger, E.; Gattiker, A.; Hoogland, C.; Ivanyi, I.; Appel, R.D.; Bairoch, A. ExPASy: The proteomics server for in-depth protein knowledge and analysis. Nucleic Acids Res 2003, 31, 3784–3788. [Google Scholar]
- Petersen, T.N.; Brunak, S.; von Heijne, G.; Nielsen, H. SignalP 4.0: Discriminating signal peptides from transmembrane regions. Nat. Methods 2011, 8, 785–786. [Google Scholar]
- Letunic, I.; Doerks, T.; Bork, P. SMART 7: Recent updates to the protein domain annotation resource. Nucleic Acids Res 2012, 40, D302–D305. [Google Scholar]
- Xia, Q.; Zhou, Z.; Lu, C.; Cheng, D.; Dai, F.; Li, B.; Zhao, P.; Zha, X.; Cheng, T.; Chai, C.; et al. A draft sequence for the genome of the domesticated silkworm (Bombyx mori). Science 2004, 306, 1937–1940. [Google Scholar]
- Duan, J.; Li, R.; Cheng, D.; Fan, W.; Zha, X.; Cheng, T.; Wu, Y.; Wang, J.; Mita, K.; Xiang, Z.; et al. SilkDB v2.0: A platform for silkworm (Bombyx mori) genome biology. Nucleic Acids Res 2010, 38, D453–D456. [Google Scholar]
- Friedman, R. Genomic organization of the glutathione S-transferase family in insects. Mol. Phylogenet Evol 2011, 61, 924–932. [Google Scholar]
- Tamura, K.; Stecher, G.; Peterson, D.; Filipski, A.; Kumar, S. MEGA6: Molecular evolutionary genetics analysis version 6.0. Mol. Biol. Evol 2013, 30, 2725–2729. [Google Scholar]
- DeLano, W.L. The PyMOL Molecular Graphic System; DeLano Scientific: San Carlos, CA, USA, 2002. [Google Scholar]
- Corpet, F. Multiple sequence alignment with hierarchical clustering. Nucleic Acids Res 1988, 16, 10881–10890. [Google Scholar]
- Gouet, P.; Courcelle, E.; Stuart, D.I.; Metoz, F. ESPript: Analysis of multiple sequence alignments in PostScript. Bioinformatics 1999, 15, 305–308. [Google Scholar]
- Habig, W.H.; Pabst, M.J.; Jakoby, W.B. Glutathione S-transferases. The first enzymatic step in mercapturic acid formation. J. Biol. Chem 1974, 249, 7130–7139. [Google Scholar]

© 2014 by the authors; licensee MDPI, Basel, Switzerland This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Tan, X.; Hu, X.-M.; Zhong, X.-W.; Chen, Q.-M.; Xia, Q.-Y.; Zhao, P. Antenna-Specific Glutathione S-Transferase in Male Silkmoth Bombyx mori. Int. J. Mol. Sci. 2014, 15, 7429-7443. https://doi.org/10.3390/ijms15057429
Tan X, Hu X-M, Zhong X-W, Chen Q-M, Xia Q-Y, Zhao P. Antenna-Specific Glutathione S-Transferase in Male Silkmoth Bombyx mori. International Journal of Molecular Sciences. 2014; 15(5):7429-7443. https://doi.org/10.3390/ijms15057429
Chicago/Turabian StyleTan, Xiang, Xiao-Ming Hu, Xiao-Wu Zhong, Quan-Mei Chen, Qing-You Xia, and Ping Zhao. 2014. "Antenna-Specific Glutathione S-Transferase in Male Silkmoth Bombyx mori" International Journal of Molecular Sciences 15, no. 5: 7429-7443. https://doi.org/10.3390/ijms15057429
APA StyleTan, X., Hu, X.-M., Zhong, X.-W., Chen, Q.-M., Xia, Q.-Y., & Zhao, P. (2014). Antenna-Specific Glutathione S-Transferase in Male Silkmoth Bombyx mori. International Journal of Molecular Sciences, 15(5), 7429-7443. https://doi.org/10.3390/ijms15057429




