Gram-Negative Flagella Glycosylation
Abstract
:1. Introduction
2. N-Glycosylation
3. O-Glycosylation
4. Flagellar Glycan Structures
4.1. Campylobacter spp
4.2. Helicobacter spp
4.3. Pseudomonas spp
4.4. Aeromonas spp
4.5. Burkholderia spp
4.6. Azospirillum brasilense
4.7. Shewanella oneidensis
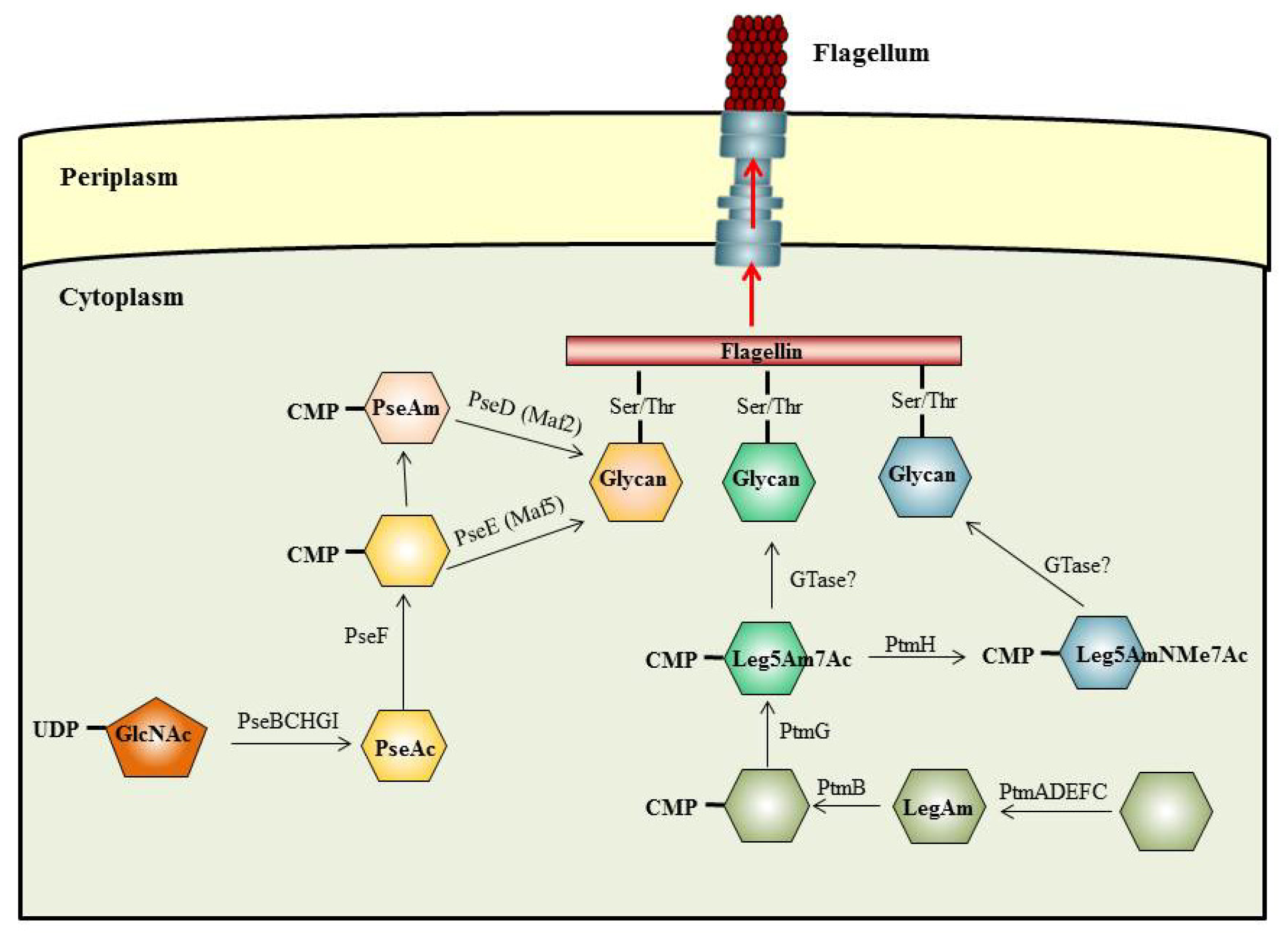
5. The O-Linked Pathway for Flagella Glycosylation
6. Biological Function of Flagellar Glycans
7. Conclusions
Acknowledgments
Conflicts of Interest
References
- Sleytr, U.B. Heterologous reattachment of regular arrays of glycoproteins on bacterial surfaces. Nature 1975, 257, 400–402. [Google Scholar]
- Mescher, M.F.; Strominger, J.L. Purification and characterization of a prokaryotic glucoprotein from the cell envelope of Halobacterium salinarium. J. Biol. Chem 1976, 251, 2005–2014. [Google Scholar]
- Calo, D.; Kaminski, L.; Eichler, J. Protein glycosylation in Archaea: Sweet and extreme. Glycobiology 2010, 20, 1065–1076. [Google Scholar]
- Nothaft, H.; Szymanski, C.M. Protein glycosylation in bacteria: Sweeter than ever. Nat. Rev. Microbiol 2010, 8, 765–778. [Google Scholar]
- Schirm, M.; Soo, E.C.; Aubry, A.J.; Austin, J.; Thibault, P.; Logan, S.M. Structural, genetic and functional characterization of the flagellin glycosylation process in Helicobacter pylori. Mol. Microbiol 2003, 48, 1579–1592. [Google Scholar]
- Young, N.M.; Brisson, J.-R.; Kelly, J.; Watson, D.C.; Tessier, L.; Lanthier, P.H.; Jarrell, H.C.; Cadotte, N.; Michael, F.; Aberg, E.; et al. Structure of the N-Linked glycan present on multiple glycoproteins in the Gram-negative bacterium Campylobacter jejuni. J. Biol. Chem 2002, 277, 42530–42539. [Google Scholar]
- Stimson, E.; Virji, M.; Makepeace, K.; Dell, A.; Morris, H.R.; Payne, G.; Saunder, J.R.; Jennings, M.P.; Barker, S.; Panico, M. Meningococcal pilin: A glycoprotein substituted with digalactosyl 2,4-diacetamido-2,4,6-trideoxyhexose. Mol. Microbiol 1995, 17, 1201–1214. [Google Scholar]
- Castric, P.; Cassels, F.J.; Carlson, R.W. Structural characterization of the Pseudomonas aeruginosa 1244 pilin glycan. J. Biol. Chem 2001, 276, 26479–26485. [Google Scholar]
- Gross, J.; Grass, S.; Davis, A.E.; Gilmore-Erdmann, P.; Townsend, R.R.; Geme, J.W.S. The Haemophilus influenzae HMW1 adhesin is a glycoprotein with an unusual N-linked carbohydrate modification. J. Biol. Chem 2008, 283, 26010–26015. [Google Scholar]
- Sekot, G.; Posch, G.; Messner, P.; Matejka, M.; Rausch-Fan, X.; Andrukhov, O.; Schaffer, C. Potential of the Tannerella forsythia S-layer to delay the immune response. J. Dent. Res 2011, 90, 109–114. [Google Scholar]
- Wacker, M.; Feldman, M.F.; Callewaert, N.; Kowarik, M.; Clarke, B.R.; Pohl, N.L.; Hernandez, M.; Vines, E.D.; Valvano, M.A.; Whitfield, C.; et al. Substrate specificity of bacterial oligosaccharyltransferase suggests acommontransfer mechanism for the bacterial and eukaryotic systems. Proc. Natl. Acad. Sci. USA 2006, 103, 7088–7093. [Google Scholar]
- Szymanski, C.M.; Yao, R.; Ewing, C.P.; Trust, T.J.; Guerry, P. Evidence for a system of general protein glycosylation in Campylobacter jejuni. Mol. Microbiol 1999, 32, 1022–1030. [Google Scholar]
- Linton, D.; Allan, E.; Karlyshev, A.V.; Cronshaw, A.D.; Wren, B.W. Identification of N- acetylgalactosamine-containing glycoproteins PEB3 and CgpA in Campylobacter jejuni. Mol. Microbiol 2002, 43, 497–508. [Google Scholar]
- Linton, D.; Dorrell, N.; Hitchen, P.G.; Amber, S.; Karlyshev, A.V.; Morris, H.R.; Dell, A.; Valvano, M.A.; Aebi, M.; Wren, B.W. Functional analysis of the Campylobacter jejuni N-linked protein glycosylation pathway. Mol. Microbiol 2005, 55, 1695–1703. [Google Scholar]
- Scott, N.E.; Parker, B.L.; Connolly, A.M.; Paulech, J.; Edwards, A.V.; Crossett, B.; Falconer, L.; Kolarich, D.; Djordjevic, S.P.; Højrup, P.; et al. Simultaneous glycan-peptide characterization using hydrophilic interaction chromatography and parallel fragmentation by CID, higher energy collisional dissociation, and electron transfer dissociation MS applied to the N-linked glycoproteome of Campylobacter jejuni. Mol. Cell Proteomics 2011, 10, M000031–MCP000201. [Google Scholar]
- Grass, S.; Lichti, C.F.; Townsend, R.R.; Gross, J.; St Geme, J.W. The Haemophilus influenza HMW1C protein is a glycosyltransferase that transfers hexose residues to asparagine sites in the HMW1 adhesin. PLoS Pathog 2010, 6, e1000919. [Google Scholar]
- Faridmoayer, A.; Fentabil, M.A.; Mills, D.C.; Klassen, J.S.; Feldman, M.F. Functional characterization of bacterial oligosaccharyltransferases involved in O-linked protein glycosylation. J. Bacteriol 2007, 189, 8088–8098. [Google Scholar]
- Hug, I.; Feldman, M.F. Analogies and homologies in lipopolysaccharide and glycoprotein biosynthesis in bacteria. Glycobiology 2011, 21, 138–151. [Google Scholar]
- Logan, S.M. Flagellar glycosylation—A new component of the motility repertoire? Microbiology 2006, 152, 1249–1262. [Google Scholar]
- Lindenthal, C.; Elsinghorst, E.A. Identification of a glycoprotein produced by enterotoxigenic Escherichia coli. Infect. Immun 1999, 67, 4084–4091. [Google Scholar]
- Schirm, M.; Kalmokoff, M.; Aubry, A.; Thibault, P.; Sandoz, M.; Logan, S.M. Flagellin from Listeria monocytogenes is glycosylated with beta-O-linked N-acetylglucosamine. J. Bacteriol 2004, 186, 6721–6727. [Google Scholar]
- Twine, S.M.; Paul, C.J.; Vinogradov, E.; McNally, D.J.; Brisson, J.R.; Mullen, J.A.; McMullin, D.R.; Jarrell, H.C.; Austin, J.W.; Kelly, J.F.; et al. Flagellar glycosylation in Clostridium botulinum. FEBS J 2008, 275, 4428–4444. [Google Scholar]
- Twine, S.M.; Reid, C.W.; Aubry, A.; McMullin, D.R.; Fulton, K.M.; Austin, J.; Logan, S.M. Motility and flagellar glycosylation in Clostridium difficile. J. Bacteriol 2009, 191, 7050–7062. [Google Scholar]
- Samatey, F.A.; Imada, K.; Vonderviszt, F.; Shirakihara, Y.; Namba, K. Crystallization of the F41 fragment of flagellin and data collection from extremely thin crystals. J. Struct. Biol 2000, 132, 106–111. [Google Scholar]
- Thibault, P.; Logan, S.M.; Kelly, J.F.; Brisson, J.-R.; Ewing, C.P.; Trust, T.J.; Guerry, P. Identification of the carbohydrate moieties and glycosylation motifs in Campylobacter jejuni flagellin. J. Biol. Chem 2001, 276, 34862–34870. [Google Scholar]
- Schirm, M.; Schoenhofen, I.C.; Logan, S.M.; Waldron, K.C.; Thibault, P. Identification of unusual bacterial glycosylation by tandem mass spectrometry analyses of intact proteins. Anal. Chem 2005, 77, 7774–7782. [Google Scholar]
- Josenhans, C.; Vossebein, L.; Friedrich, S.; Suerbaum, S. The neuA/flmD gene cluster of Helicobacter pylori is involved in flagellar biosynthesis and flagellin glycosylation. FEMS Microbiol. Lett 2002, 210, 165–172. [Google Scholar]
- Tabei, S.M.; Hitchen, P.G.; Day-Williams, M.J.; Merino, S.; Vart, R.; Pang, P.C.; Horsburgh, G.J.; Viches, S.; Wilhelms, M.; Tomás, J.M.; et al. An Aeromonas caviae genomic island is required for both O-antigen lipopolysaccharide biosynthesis and flagellin glycosylation. J. Bacteriol 2009, 191, 2851–2863. [Google Scholar]
- Scott, A.E.; Twine, S.M.; Fulton, K.M.; Titball, R.W.; Essex-Lopresti, A.E.; Atkins, T.P.; Prior, J.L. Flagellar glycosylation in Burkholderia pseudomallei and Burkholderia thailandensis. J. Bacteriol 2011, 193, 3577–3587. [Google Scholar]
- Belyakov, A.Y.; Burygin, G.L.; Arbatsky, N.P.; Shashkov, A.S.; Selivanov, N.Y.; Matora, L.Y.; Knirel, Y.A.; Shchyogolev, S.Y. Identification of an O-linked repetitive glycan chain of the polar flagellum flagellin of Azospirillum brasilense Sp7. Carbohydr. Res 2012, 361, 127–132. [Google Scholar]
- Harris, L.A.; Logan, S.M.; Guerry, P.; Trust, T.J. Antigenic variation of Campylobacter flagellin. J. Bacteriol 1987, 169, 5066–5071. [Google Scholar]
- Alm, R.A.; Guerry, P.; Power, M.E.; Trust, T.J. Variation in antigenicity and molecular weight of Campylobacter coli VC167 flagellin in different genetic backgrounds. J. Bacteriol 1992, 174, 4230–4238. [Google Scholar]
- Power, M.E.; Guerry, P.; McCubbin, W.B.; Kay, C.M.; Trust, T.J. Structural and antigenic characteristics of Campylobacter coli FlaA flagellin. J. Bacteriol 1994, 176, 3303–3313. [Google Scholar]
- Logan, S.M.; Schoenhofen, I.C.; Guerry, P. O- Linked Flagellar Glycosylation in Campylobacter. In Campylobacter, 3rd ed.; Nachamkin, I., Szymanski, C.M., Blaser, M.J., Eds.; ASM Press: Washington DC, USA, 2008; pp. 471–480. [Google Scholar]
- Wilhelms, M.; Fulton, K.M.; Twine, S.M.; Tomás, J.M.; Merino, S. Differential glycosylation of polar and lateral flagellins in Aeromonas hydrophila AH-3. J. Biol. Chem 2012, 287, 27851–27862. [Google Scholar]
- Logan, S.M.; Hui, J.P.M.; Vinogradov, E.; Aubrey, A.J.; Melanson, J.E.; Kelly, J.F.; Nothaft, H.; Soo, E.C. Identification of novel carbohydrate modifications on Campylobacter jejuni 11168 flagellin using metabolomicsbased approaches. FEBS J 2009, 276, 1014–1023. [Google Scholar]
- Logan, S.M.; Kelly, J.F.; Thibault, P.; Ewing, C.P.; Guerry, P. Structural heterogeneity of carbohydrate modifications affects serospecificity of Campylobacter flagellins. Mol. Microbiol 2002, 46, 587–597. [Google Scholar]
- Schirm, M.; Arora, K.; Verma, A.; Vinogradov, E.; Thibault, P.; Ramphal, R.; Logan, S.M. Structural and genetic characterization of glycosylation of type a flagellin in Pseudomonas aeruginosa. J. Bacteriol 2004, 186, 2523–2531. [Google Scholar]
- Verma, A.; Schirm, M.; Arora, S.K.; Thibault, P.; Logan, S.M.; Rampha, R. Glycosylation of b-type flagellin of Pseudomonas aeruginosa: Structural and genetic basis. J. Bacteriol 2006, 188, 4395–4403. [Google Scholar]
- Lindhout, T.; Lau, P.C.; Brewer, D.; Lam, J.S. Truncation in the core oligosaccharide of lipopolysaccharide affects flagella-mediated motility in Pseudomonas aeruginosa PAO1 via modulation of cell surface attachment. Microbiology 2009, 155, 3449–3460. [Google Scholar]
- Konishi, T.; Taguchi, F.; Iwaki, M.; Ohnishi-Kameyama, M.; Yamamoto, M.; Maeda, I.; Nishida, Y.; Ichinose, Y.; Yoshida, M.; Ishii, T. Structural characterization of an O-linked tetrasaccharide from Pseudomonas syringae pv. tabaci flagellin. Carbohydr. Res 2009, 344, 2250–2254. [Google Scholar]
- Takeuchi, K.; Ono, H.; Yoshida, M.; Ishii, T.; Katoh, E.; Taguchi, F.; Miki, R.; Murata, K.; Kaku, H.; Ichinose, Y. Flagellin glycans from two pathovars of Pseudomonas syringae contain rhamnose in D and L configurations in different ratios and modified 4-amino-4,6-dideoxyglucose. J. Bacteriol 2007, 189, 6945–6956. [Google Scholar]
- Sun, L.; Jin, M.; Ding, W.; Yuan, J.; Kelly, J.; Gao, H. Posttranslational Modification of Flagellin FlaB in Shewanella oneidensis. J. Bacteriol 2013, 195, 2550–2561. [Google Scholar]
- McNally, D.J.; Aubry, A.J.; Hui, J.P.; Khieu, N.H.; Whitfield, D.; Ewing, C.P.; Guerry, P.; Brisson, J.R.; Logan, S.M.; Soo, E.C. Targeted metabolomics analysis of Campylobacter coli VC167 reveals legionaminic acid derivatives as novel flagellar glycans. J. Biol. Chem 2007, 282, 14463–14475. [Google Scholar]
- Dorrell, N.; Mangan, J.A.; Laing, K.G.; Hinds, J.; Linton, D.; Al-Ghusein, H.; Barrell, B.; Parkhill, J.; Stoker, N.G.; Karylshev, A.V.; et al. Whole genome comparison of Campylobacter jejuni human isolates using a low-cost microarray reveals extensive genetic diversity. Genome Res 2001, 11, 1706–1715. [Google Scholar]
- Fouts, D.E.; Mongodin, E.F.; Mandrell, R.E.; Miller, W.G.; Rasko, D.A.; Ravel, J.; Brinkac, L.M.; DeBoy, R.T.; Parker, C.T.; Dougherty, S.C.; et al. Major structural differences and novel potential virulence mechanisms from the genomes of multiple Campylobacter species. PLoS Biol 2005, 3, e15. [Google Scholar]
- Pearson, B.M.; Pin, C.; Wright, J.; I’Anson, K.; Humphrey, T.; Wells, J.M. Comparative genome analysis of Campylobacter jejuni using whole genome microarrays. FEBS Lett 2004, 554, 224–230. [Google Scholar]
- Guerry, P.; Ewing, C.P.; Schirm, M.; Lorenzo, M.; Kelly, J.; Pattarini, D.; Majam, G.; Thibault, P.; Logan, S. Changes in flagellin glycosylation affect Campylobacter autoagglutination and virulence. Mol. Microbiol 2006, 60, 299–311. [Google Scholar]
- Fox, J.G. The non-H. pylori helicobacters: Their expanding role in gastrointestinal and systemic diseases. Gut 2002, 50, 273–283. [Google Scholar]
- Schoenhofen, I.C.; McNally, D.J.; Brisson, J.R.; Logan, S.M. Elucidation of the CMP-pseudaminic acid pathway in Helicobacter pylori: Synthesis from UDP-N-acetylglucosamine by a single enzymatic reaction. Glycobiology 2006, 16, 8C–14C. [Google Scholar]
- Allison, J.S.; Dawson, M.; Drake, D.; Montie, T.C. Electrophoretic separation and molecular weight characterization of Pseudomonas aeruginosa H-antigen flagellins. Infect. Immun 1985, 49, 770–774. [Google Scholar]
- Lanyi, B. Serological properties of Pseudomonas aeruginosa. II. Type-specific thermolabile (flagellar) antigens. Acta Microbiol. Acad. Sci. Hung 1970, 17, 35–48. [Google Scholar]
- Arora, S.K.; Bangera, M.; Lory, S.; Ramphal, R. A genomic island in Pseudomonas aeruginosa carries the determinants of flagellin glycosylation. Proc. Natl. Acad. Sci. USA 2001, 98, 9342–9347. [Google Scholar]
- Brimer, C.D.; Montie, T.C. Cloning and comparison of fliC genes and identification of glycosylation in the flagellin of Pseudomonas aeruginosa a-type strains. J. Bacteriol 1998, 180, 3209–3217. [Google Scholar]
- Arora, S.K.; Wolfgang, M.C.; Lory, S.; Ramphal, R. Sequence polymorphism in the glycosylation island and flagellins of Pseudomonas aeruginosa. J. Bacteriol 2004, 186, 2115–2122. [Google Scholar]
- Miller, W.L.; Matewish, M.J.; McNally, D.J.; Ishiyama, N.; Anderson, E.M.; Brewer, D.; Brisson, J.R.; Berghuis, A.M.; Lam, J.S. Flagellin glycosylation in Pseudomonas aeruginosa PAK requires the O-antigen biosynthesis enzyme WbpO. J. Biol. Chem 2008, 283, 3507–3518. [Google Scholar]
- Janda, J.M.; Abbott, S.L. Evolving concepts regarding the genus Aeromonas: An expanding panorama of species, disease presentations, and unanswered questions. Clin. Infect. Dis 1998, 27, 332–344. [Google Scholar]
- Kirov, S.M.; Tassell, B.C.; Semmler, A.B.; O’Donovan, L.A.; Rabaan, A.A.; Shaw, J.G. Lateral flagella and swarming motility in Aeromonas species. J. Bacteriol 2002, 184, 547–555. [Google Scholar]
- Rabaan, A.A.; Gryllos, I.; Tomás, J.M.; Shaw, J.G. Motility and the polar flagellum are required for Aeromonas caviae adherence to HEp-2 cells. Infect. Immun 2001, 69, 4257–4267. [Google Scholar]
- Gavín, R.; Rabaan, A.A.; Merino, S.; Tomás, J.M.; Gryllos, I.; Shaw, J.G. Lateral flagella of Aeromonas species are essential for epithelial cell adherence and biofilm formation. Mol. Microbiol 2002, 43, 383–397. [Google Scholar]
- Parker, J.L.; Day-Williams, M.J.; Tomás, J.M.; Stafford, G.P.; Shaw, J.G. Identification of a putative glycosyltransferase responsible for the transfer of pseudaminic acid onto the polar flagellin of Aeromonas caviae Sch3N. Microbiol. Open 2012, 1, 149–160. [Google Scholar]
- Knirel, Y.A.; Shashkov, A.S.; Senchenkova, S.N.; Merino, S.; Tomás, J.M. Structure of the O-polysaccharide of Aeromonas hydrophila O:34; A case of random O-acetylation of 6-deoxy-L-talose. Carbohydr. Res 2002, 337, 1381–1386. [Google Scholar]
- Canals, R.; Altarriba, M.; Vilches, S.; Horsburgh, G.; Shaw, J.G.; Tomás, J.M.; Merino, S. Analysis of the lateral flagellar gene system of Aeromonas hydrophila AH-3. J. Bacteriol 2006, 188, 852–862. [Google Scholar]
- Canals, R.; Ramirez, S.; Vilches, S.; Horsburgh, G.; Shaw, J.G.; Tomás, J.M.; Merino, S. Polar flagellum biogenesis in Aeromonas hydrophila. J. Bacteriol 2006, 188, 542–555. [Google Scholar]
- Zhulin, I.B.; Armitage, J.P. Motility, chemokinesis, and methylation-independent chemotaxis in Azospirillum brasilense. J. Bacteriol 1993, 175, 952–958. [Google Scholar]
- Croes, C.; van Bastelaere, E.; DeClercq, E.; Eyers, M.; Vanderleyden, J.; Michiels, K. Identification and mapping of loci involved in motility, adsorption to wheat roots, colony morphology, and growth in minimal medium on the Azospirillum brasilense Sp7 90-MDa plasmid. Plasmid 1991, 26, 83–93. [Google Scholar]
- Moens, S.; Michiels, K.; Vanderleyden, J. Glycosylation of the flagellin of the polar flagellum of Azospirillum brasilense, a Gram-negative nitrogen-fixing bacterium. J. Microbiol 1995, 141, 2651–2657. [Google Scholar]
- Wang, F.; Wang, J.; Jian, H.; Zhang, B.; Li, S.; Wang, F.; Zeng, X.; Gao, L.; Bartlett, D.H.; Yu, J.; et al. Environmental adaptation: Genomic analysis of the piezotolerant and psychrotolerant deep-sea iron reducing bacterium Shewanella piezotolerans. PLoS One 2008, 3, e1937. [Google Scholar]
- Bubendorfer, S.; Held, S.; Windel, N.; Paulick, A.; Klingl, A.; Thormann, K.M. Specificity of motor components in the dual flagellar system of Shewanella putrefaciens CN-32. Mol. Microbiol 2012, 83, 335–350. [Google Scholar]
- Wu, L.; Wang, J.; Tang, P.; Chen, H.; Gao, H. Genetic and molecular characterization of flagellar assembly in Shewanella oneidensis. PLoS One 2011, 6, e21479. [Google Scholar]
- Macnab, R.M. Type III flagellar protein export and flagellar assembly. Biochim. Biophys. Acta 2004, 1694, 207–217. [Google Scholar]
- Evans, L.D.; Poulter, S.; Terentjev, E.M.; Hughes, C.; Fraser, G.M. A chain mechanism for flagellum growth. Nature 2013, 504, 287–290. [Google Scholar]
- Evdokimov, A.G.; Phan, J.; Tropea, J.E.; Routzahn, K.M.; Peters, H.K.; Pokross, M.; Waugh, D.S. Similar modes of polypeptide recognition by export chaperones in flagellar biosynthesis and type III secretion. Nat. Struct. Biol 2003, 10, 789–793. [Google Scholar]
- Ozin, A.J.; Claret, L.; Auvray, F.; Hughes, C. The FliS chaperone selectively binds the disordered flagellin C-terminal D0 domain central to polymerisation. FEMS Microbiol. Lett 2003, 219, 219–224. [Google Scholar]
- Schoenhofen, I.C.; Vinogradov, E.; Whitfield, D.M.; Brisson Logan, J-R.S.M. The CMP-legionaminic acid pathway in Campylobacter: Biosynthesis involving novel GDP-linked precursors. Glycobiology 2009, 19, 715–725. [Google Scholar]
- Yamamoto, M.; Ohnishi-Kameyama, M.; Nguyen, C.L.; Taguchi, F.; Chiku, K.; Ishii, T.; Ono, H.; Yoshida, M.; Ichinose, Y. Identification of genes involved in the glycosylation of modified viosamine of flagellins in Pseudomonas syringae by mass spectrometry. Genes 2011, 2, 788–803. [Google Scholar]
- Ewing, C.P.; Andreishcheva, E.; Guerry, P. Functional characterization of flagellin glycosylation in Campylobacter jejuni. J. Bacteriol 2009, 191. [Google Scholar]
- Takeuchi, K.; Taguchi, F.; Inagaki, Y.; Toyoda, K.; Shiraishi, T.; Ichinose, Y. Flagellin glycosylation island in Pseudomonas syringae pv. glycinea and its role in host specificity. J. Bacteriol 2003, 185, 6658–6665. [Google Scholar]
- Guerry, P. Campylobacter flagella: Not just for motility. Trends Microbiol 2007, 15, 456–461. [Google Scholar]
- Guerry, P.; Szymanski, C.M. Campylobacter sugars sticking out. Trends Microbiol 2008, 16, 428–435. [Google Scholar]
- Howard, S.L.; Jagannathan, A.; Soo, E.C.; Hui, J.P.M.; Aubry, A.J.; Ahmed, I.; Karlyshev, A.; Kelly, J.F.; Jones, M.A.; Stevens, M.P.; et al. Campylobacter jejuni glycosylation island important in cell charge, legionaminic acid biosynthesis, and colonization of chickens. Infect. Immun 2009, 77, 2544–2556. [Google Scholar]
- Goon, S.; Kelly, J.F.; Logan, S.M.; Ewing, C.P.; Guerry, P. Pseudaminic acid, the major modification on Campylobacte r flagellin, is synthesized via the Cj1293 gene. Mol. Microbiol 2003, 50, 659–671. [Google Scholar]
- Asakura, H.; Churin, Y.; Bauer, B.; Boettcher, J.P.; Bartfeld, S.; Hashii, N.; Kawasaki, N.; Mollenkopf, H.J.; Jungblut, P.R.; Brinkmann, V.; et al. Helicobacter pylori HP0518 affects flagellin glycosylation to alter bacterial motility. Mol. Microbiol 2010, 78, 1130–1144. [Google Scholar]
- Arora, S.K.; Neely, A.N.; Blair, B.; Lory, S.; Ramphal, R. Role of motility and flagellin glycosylation in the pathogenesis of Pseudomonas aeruginosa burn wound infections. Infect. Immun 2005, 73, 4395–4398. [Google Scholar]
- Verma, A.; Arora, S.K.; Kuravi, S.K.; Ramphal, R. Roles of specific amino acids in the N-terminus of Pseudomonas aeruginosa flagellin and of flagellin glycosylation in the innate immune response. Infect. Immun 2005, 73, 8237–8246. [Google Scholar]
- Taguchi, F.; Shimizu, R.; Inagaki, Y.; Toyoda, K.; Shiraishi, T.; Ichinose, Y. Post-translational modification of flagellin determines the specificity of HR induction. Plant Cell Physiol 2003, 44, 342–349. [Google Scholar]
- Taguchi, F.; Shibata, S.; Suzuki, T.; Ogawa, Y.; Aizawa, S.; Takeuchi, K.; Ichinose, Y. Effects of glycosylation on swimming ability and flagellar polymorphic transformation in Pseudomonas syringae pv. tabaci 6605. J. Bacteriol 2008, 190, 764–768. [Google Scholar]
- Taguchi, F.; Yamamoto, M.; Ohnishi-Kameyama, M.; Iwaki, M.; Yoshida, M.; Ishii, T.; Konishi, T.; Ichinose, Y. Defects in flagellin glycosylation affect the virulence of Pseudomonas syringae pv. tabaci 6605. Microbiology 2010, 156, 72–80. [Google Scholar]
| Organism | Glycoprotein | Number of sites | Glycan structure | Reference |
|---|---|---|---|---|
| Aeromonas caviae Sch3N | FlaA/FlaB | 6/7 | Pse5Ac7Ac | [28] |
| Aeromonas caviae UU51 | FlaA/FlaB | 6/7 | Pse5Ac7Ac8Ac | [26] |
| Aeromonas hydrophila AH3 | FlaA/FlaB | 6FlaB | PseAc derivative-Hex-Hex-HexNAc-HexNAc-HexNAc-102 Da | [35] |
| LafA | 3 | PseAc derivative | [35] | |
| Azospirillum brasilense Sp7 | Laf1 | ND | (→3)-α-l-Rhap-(1→3)-β-d-Galp-(1 →3)-β-d-GlcpNAc-(1→)n = 3–4 | [30] |
| Burkholderia pseudomallei K96243 | FliC | 1 | 582.4 Da (2 × 291 Da) | [29] |
| Burkholderia thailandensis E264 | FliC | 1 | 342 Da | [29] |
| Campylobacter jejuni 81–176 | FlaA | 19 | Pse5Ac7Ac, Pse5Am7Ac, Pse5Ac7Ac8OAc, Pse5Am7Ac8GlnAc, Pse5Pr7Pr | [25,26] |
| Campylobacter jejuni 11168 | FlaA | ND | Pse5Ac7Ac, Leg5Am7Ac, Leg5AmNMe7Ac | [36] |
| Campylobacter coli VC167 | FlaA | 16 | Pse5Ac7Ac, PseAm, PseAc/LegAm-deoxypentose, Leg5Ac7Ac, Leg5Am7Ac, Leg5AmNMe7Ac | [37] |
| Helicobacter pylori | FlaA/FlaB | 7/10 | Pse5Ac7Ac | [5,26] |
| Pseudomonas aeruginosa PAK | FliC | 2 | 11 residues (pentose, hexose, deoxyhexose, hexuronic) attached via l-rhamnose | [38] |
| Pseudomonas aeruginosa JJ692 | FliC | 2 | l-rhamnose [38] Pseudomonas aeruginosa PAO1 FliC 2 l-rhamnose and 209 Da phosphate | [39,40] |
| Pseudomonas syringae pv. tabaci | FliC | 6 | β-d-Quip4N(3-hydroxy-1-oxobutyl)2Me-(133)-α-l-Rhap- (132)-α-l-Rhap | [41,42] |
| Shewanella oneidensis | FlaB | 5 | 274 Da and 274 ± 14 Da | [43] |
© 2014 by the authors; licensee MDPI, Basel, Switzerland This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Merino, S.; Tomás, J.M. Gram-Negative Flagella Glycosylation. Int. J. Mol. Sci. 2014, 15, 2840-2857. https://doi.org/10.3390/ijms15022840
Merino S, Tomás JM. Gram-Negative Flagella Glycosylation. International Journal of Molecular Sciences. 2014; 15(2):2840-2857. https://doi.org/10.3390/ijms15022840
Chicago/Turabian StyleMerino, Susana, and Juan M. Tomás. 2014. "Gram-Negative Flagella Glycosylation" International Journal of Molecular Sciences 15, no. 2: 2840-2857. https://doi.org/10.3390/ijms15022840
APA StyleMerino, S., & Tomás, J. M. (2014). Gram-Negative Flagella Glycosylation. International Journal of Molecular Sciences, 15(2), 2840-2857. https://doi.org/10.3390/ijms15022840






