MicroRNA Signatures as Biomarkers and Therapeutic Target for CNS Embryonal Tumors: The Pros and the Cons
Abstract
:1. Introduction
2. miRNAs
2.1. miRNA Detection Methods: Advantages and Concerns
2.1.1. miRNAs Detection by Microarray Approach
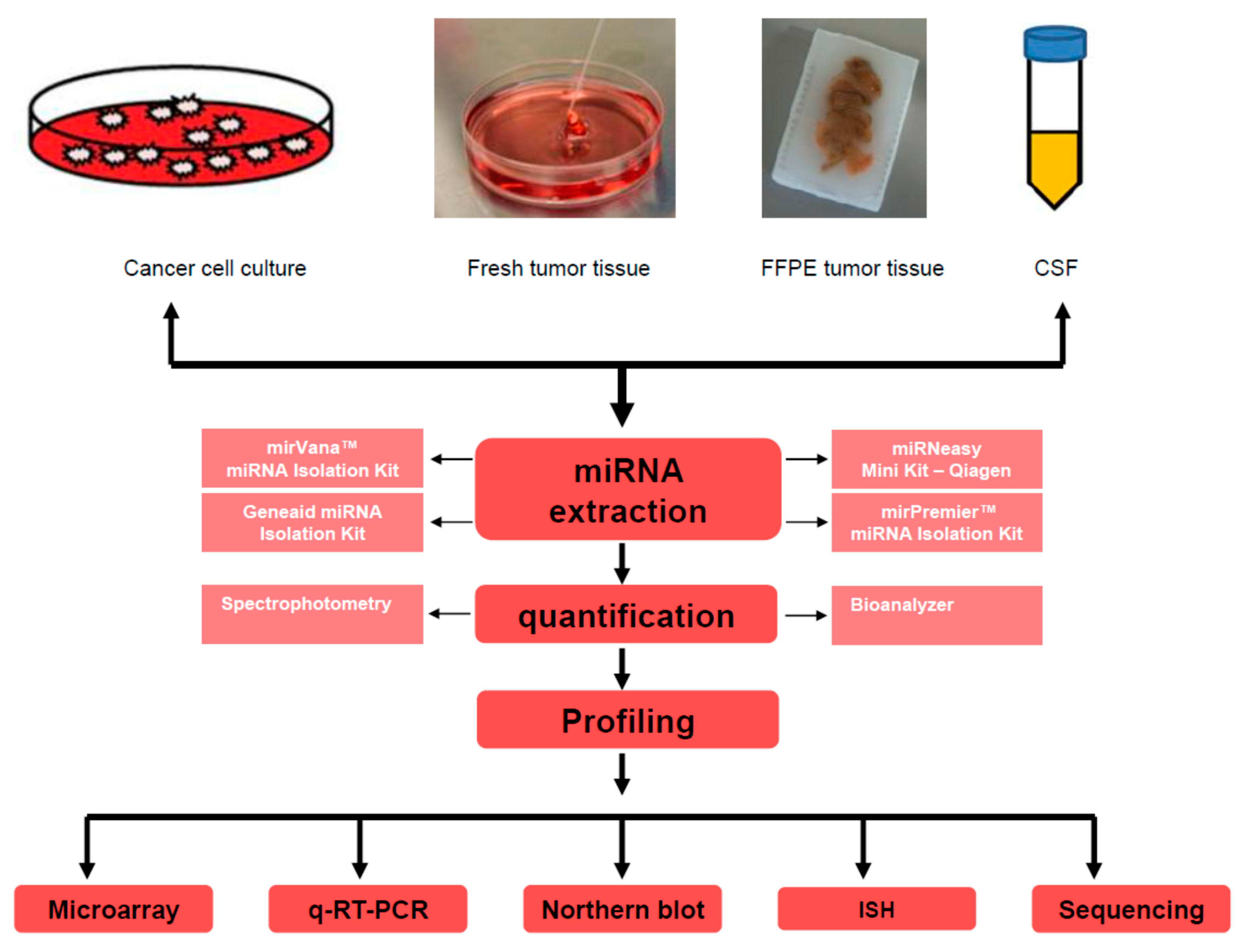
2.1.2. Real-Time-PCR-Based Detection of miRNAs
2.1.3. Northern Blot Analysis for miRNAs Detection
2.1.4. In Situ Hybridization (ISH) for miRNA Detection
2.1.5. miRNA Detection by High-Throughput Next-Generation Sequencing Platforms
2.2. miRNA Target Determination
3. miRNA Implications in CNS Embryonal Tumors
3.1. miRNAs Associated with Neuronal Development and Disorders
3.2. Biological Relevance of miRNAs in MBs
3.2.1. miRNAs Function as Oncogenes or Tumor Suppressors in MB
| miRNAs (miRs) | Targets | References |
|---|---|---|
| Oncomirs | ||
| miR-21 | PDCD4 | [78] |
| miR-517c, miR-520g | miR-517c WNT/JNK signaling, miR-520g ABCG2 | [79,80] |
| miR-221, miR-222 | p27Kip1 | [6] |
| miR-183-96–182 cluster | Undefined, however knockdown of the full miR-183-96–182 cluster results in enrichment of genes associated with apoptosis and dysregulation of the PI3K/AKT/mTOR signaling axis | [68] |
| miR-17/92 | TSP-1, Bmi-1, PTEN, PP2A | [63,81,82] |
| miR-214 | Gli1 | [4] |
| miR-30b, miR-30d | Undefined, Both miRNAs are part of an amplicon containing the KHDRBS3 gene on 8q24.22 in MB cell lines | [83] |
| miR-106b | p21 | [69,84] |
| Tumor Suppressors miRNAs | ||
| miR-125a | t-TrkC | [69] |
| miR-9 | REST/NRSF, Hes1 | [39,69] |
| miR-125b, miR-324-5p, miR-326 | PKM2, SMO, Notch | [4,85] |
| let-7 | RAS, STAT3 | [69,86] |
| miR-199b-5p | HES1, Notch pathway, ErbB2 | [87] |
| miR-124a | CDK6, SLC16A1, REST, BAF34a, RB1,t-TrkC | [35,58,60,88,89] |
| miR-218 | EGFR, Bcl-2, B-catenin and MAPK9 | [84,90] |
| miR-31, miR-153 | miR-153 is found in high ErbB2 expressing MB | [69,90] |
| miR-128a, miR-128b, miR-181b | Bmi-1 | [64,91] |
| let-7 miRNA family | HMGA2 | [7] |
3.2.2. miRNAs Associated with MB Metastasis
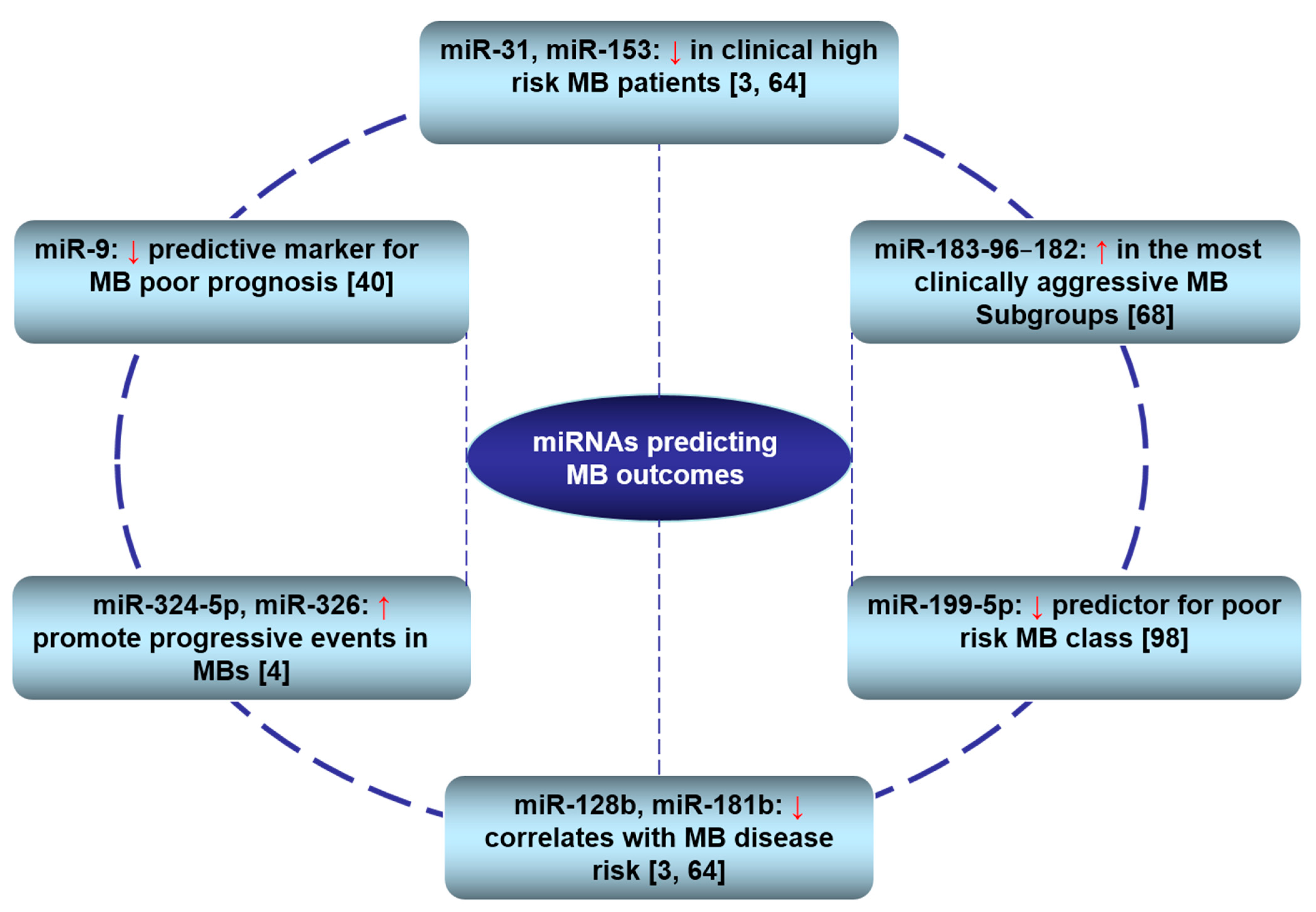
3.3. Clinical Relevance of miRNAs in MBs
3.4. Biological Relevance of miRNAs in CNS Atypical Teratoid/Rhabdoid Tumors
| miRNAs (miRs) | Targets | References |
|---|---|---|
| Oncomirs | ||
| miR-517c, miR-520g | miR-517c WNT/JNK signaling, miR-520g ABCG2 | [79,80] |
| miR-221, miR-222 | p27Kip1 | [6] |
| miR-520b, miR-629, miR-498, miR-373 | Undefined | [67] |
| Tumor Suppressors miRNAs | ||
| miR-140, let-7b, miR-139, miR-153, miR-376b | Undefined | [67] |
| let-7 miRNA family | HMGA2 | [7] |
| miR-9 | Undefined | [40,69] |
4. miRNAs as Reliable Clinical Markers: Advantages and Challenges
4.1. Process of Biomarker Development
4.2. miRNAs as Reliable Markers
4.3. Challenges
4.3.1. Lack of Collaborative Research Efforts
4.3.2. Availability of Tumor Tissue Samples
4.3.3. Reliability and Reproducibility of Results
4.3.4. Variability of Biomarker Assessment Methods
4.3.5. Expenses Involved with Assessing the Marker Status
4.3.6. The Need for Developing New miRNA Assessment Technologies
4.3.7. Lack of Cellular and Clinical Database Resources
5. Therapeutic Potential of miRNAs
5.1. miRNAs as Druggable Targets
5.2. miRNAs as Predictors and Modifiers of Chemo- and Radio-Therapy
5.3. Challenges of miRNA-Based Therapies
6. Concluding Remarks
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Louis, D.N.; Ohgaki, H.; Wiestler, O.D.; Cavenee, W.K.; Burger, P.C.; Jouvet, A.; Scheithauer, B.W.; Kleihues, P. The 2007 WHO classification of tumours of the central nervous system. Acta Neuropathol. 2007, 114, 97–109. [Google Scholar] [CrossRef] [PubMed]
- Legler, J.M.; Ries, L.A.; Smith, M.A.; Warren, J.L.; Heineman, E.F.; Kaplan, R.S.; Linet, M.S. Brain and other central nervous system cancers: Recent trends in incidence and mortality. J. Natl. Cancer Inst. 1999, 91, 1382–1390. [Google Scholar] [CrossRef] [PubMed]
- Ries, L.A.G.; Smith, M.A.; Gurney, J.G.; Linet, M.; Tamra, T.; Young, J.L.; Bunin, G.R. CNS and miscellaneous intracranial and intraspinal neoplasms. In Cancer Incidence and Survival Among Children and Adolescents: United States SEER Program, 1975–1995; Ries, L.A.G., Smith, M.A., et al., Eds.; National Institutes of Health: Bethesda, MD, USA, 1999; p. 179. [Google Scholar]
- Ferretti, E.; de Smaele, E.; Miele, E.; Laneve, P.; Po, A.; Pelloni, M.; Paganelli, A.; di Marcotullio, L.; Caffarelli, E.; Screpanti, I.; et al. Concerted microRNA control of Hedgehog signalling in cerebellar neuronal progenitor and tumour cells. EMBO J. 2008, 27, 2616–2627. [Google Scholar] [CrossRef] [PubMed]
- Rao, P.; Benito, E.; Fischer, A. MicroRNAs as biomarkers for CNS disease. Front Mol. Neurosci. 2013, 6, 39. [Google Scholar] [CrossRef] [PubMed]
- Sredni, S.T.; de Fátima Bonaldo, M.; Costa, F.F.; Huang, C.C.; Hamm, C.A.; Rajaram, V.; Tomita, T.; Goldman, S.; Bischof, J. M.; Soares, M.B. Upregulation of miR-221 and miR-222 in atypical teratoid/rhabdoid tumors: Potential therapeutic targets. Child’s Nerv. Syst. 2010, 26, 279–283. [Google Scholar] [CrossRef]
- Zhang, K.; Gao, H.; Wu, X.; Wang, J.; Zhou, W.; Sun, G.; Wang, J.; Wang, Y.; Mu, B.; Kim, C.; et al. Frequent overexpression of HMGA2 in human atypical teratoid/rhabdoid tumor and its correlation with let-7a3/let-7b miRNA. Clin. Cancer Res. 2014, 20, 1179–1189. [Google Scholar] [CrossRef] [PubMed]
- Vidal, D.O.; Marques, M.M.; Lopes, L.F.; Reis, R.M. The role of microRNAs in medulloblastoma. J. Pediatr. Hematol. Oncol. 2013, 30, 367–378. [Google Scholar] [CrossRef]
- To, K.K. MicroRNA: A prognostic biomarker and a possible druggable target for circumventing multidrug resistance in cancer chemotherapy. J. Biomed. Sci. 2013, 20, 99. [Google Scholar] [CrossRef] [PubMed]
- Bartel, D.P. MicroRNAs: Genomics, biogenesis, mechanism, and function. Cell 2004, 116, 281–297. [Google Scholar] [CrossRef] [PubMed]
- Bader, A.G.; Brown, D.; Stoudemire, J.; Lammers, P. Developing therapeutic microRNAs for cancer. Gene Ther. 2011, 18, 1121–1126. [Google Scholar] [PubMed]
- De Planell-Saguer, M.; Rodicio, M.C. Analytical aspects of microRNA in diagnostics: A review. Anal. Chim. Acta 2011, 699, 134–152. [Google Scholar] [CrossRef] [PubMed]
- Takada, S.; Mano, H. Profiling of microRNA expression by mRAP. Nat. Protoc. 2007, 2, 3136–3145. [Google Scholar] [CrossRef] [PubMed]
- Castoldi, M.; Benes, V.; Hentze, M.W.; Muckenthaler, M.U. miChip: A microarray platform for expression profiling of microRNAs based on locked nucleic acid (LNA) oligonucleotide capture probes. Methods 2007, 43, 146–152. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Ridzon, D.A.; Broomer, A.J.; Zhou, Z.; Lee, D.H.; Nguyen, J.T.; Barbisin, M.; Xu, N.L.; Mahuvakar, V.R.; Andersen, M.R.; et al. Real-time quantification of microRNAs by stem-loop RT-PCR. Nucleic Acids Res. 2005, 33, e179. [Google Scholar] [CrossRef] [PubMed]
- Válóczi, A.; Hornyik, C.; Varga, N.; Burgyan, J.; Kauppinen, S.; Havelda, Z. Sensitive and specific detection of microRNAs by northern blot analysis using LNA-modified oligonucleotide probes. Nucleic Acids Res. 2004, 32, e175. [Google Scholar] [CrossRef] [PubMed]
- De Planell-Saguer, M.; Rodicio, M.C.; Mourelatos, Z. Rapid in situ codetection of noncoding RNAs and proteins in cells and formalin-fixed paraffin-embedded tissue sections without protease treatment. Nat. Protoc. 2010, 5, 1061–1073. [Google Scholar] [CrossRef] [PubMed]
- Schulte, J.H.; Marschall, T.; Martin, M.; Rosenstiel, P.; Mestdagh, P.; Schlierf, S.; Thor, T.; Vandesompele, J.; Eggert, A.; Schreiber, S.; et al. Deep sequencing reveals differential expression of microRNAs in favorable vs. unfavorable neuroblastoma. Nucleic Acids Res. 2010, 38, 5919–5928. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Ruan, K. MicroRNA detection by microarray. Anal. Bioanal. Chem. 2009, 394, 1117–1124. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.G.; Calin, G.A.; Meloon, B.; Gamliel, N.; Sevignani, C.; Ferracin, M.; Dumitru, C.D.; Shimizu, M.; Zupo, S.; Dono, M.; et al. An oligonucleotide microchip for genome-wide microRNA profiling in human and mouse tissues. Pro. Natl. Acad. Sci. USA 2004, 101, 9740–9744. [Google Scholar] [CrossRef]
- Pritchard, C.C.; Cheng, H.H.; Tewari, M. MicroRNA profiling: Approaches and considerations. Nat. Rev. Genet. 2012, 13, 358–369. [Google Scholar] [CrossRef] [PubMed]
- Van Rooij, E. The art of microRNA research. Circ. Res. 2011, 108, 219–234. [Google Scholar] [CrossRef] [PubMed]
- Wightman, B.; Ha, I.; Ruvkun, G. Posttranscriptional regulation of the heterochronic gene lin-14 by lin-4 mediates temporal pattern formation in C. elegans. Cell 1993, 75, 855–862. [Google Scholar] [CrossRef] [PubMed]
- Sempere, L.F.; Freemantle, S.; Pitha-Rowe, I.; Moss, E.; Dmitrovsky, E.; Ambros, V. Expression profiling of mammalian microRNAs uncovers a subset of brain-expressed microRNAs with possible roles in murine and human neuronal differentiation. Genome Biol. 2004, 5, R13. [Google Scholar] [CrossRef] [PubMed]
- Marti, E.; Pantano, L.; Banez-Coronel, M.; Llorens, F.; Minones-Moyano, E.; Porta, S.; Sumoy, L.; Ferrer, I.; Estivill, X. A myriad of miRNA variants in control and Huntington’s disease brain regions detected by massively parallel sequencing. Nucleic Acids Res. 2010, 38, 7219–7235. [Google Scholar] [CrossRef] [PubMed]
- Linsen, S.E.; de Wit, E.; Janssens, G.; Heater, S.; Chapman, L.; Parkin, R.K.; Fritz, B.; Wyman, S.K.; de Bruijn, E.; Voest, E.E.; et al. Limitations and possibilities of small RNA digital gene expression profiling. Nat. Methods 2009, 6, 474–476. [Google Scholar] [CrossRef] [PubMed]
- Akbari Moqadam, F.; Pieters, R.; den Boer, M.L. The hunting of targets: Challenge in miRNA research. Leukemia 2013, 27, 16–23. [Google Scholar] [CrossRef] [PubMed]
- Bar, M.; Wyman, S.K.; Fritz, B.R.; Qi, J.; Garg, K.S.; Parkin, R.K.; Kroh, E.M.; Bendoraite, A.; Mitchell, P.S.; Nelson, A.M.; et al. MicroRNA discovery and profiling in human embryonic stem cells by deep sequencing of small RNA libraries. Stem Cells 2008, 26, 2496–2505. [Google Scholar] [CrossRef] [PubMed]
- Esquela-Kerscher, A.; Slack, F.J. Oncomirs—microRNAs with a role in cancer. Nat. Rev. Cancer. 2006, 6, 259–269. [Google Scholar] [CrossRef] [PubMed]
- He, L.; Thomson, J.M.; Hemann, M.T.; Hernando-Monge, E.; Mu, D.; Goodson, S.; Powers, S.; Cordon-Cardo, C.; Lowe, S.W.; Hannon, G.J.; et al. A microRNA polycistron as a potential human oncogene. Nature 2005, 435, 828–833. [Google Scholar] [CrossRef] [PubMed]
- Kunder, R.; Jalali, R.; Sridhar, E.; Moiyadi, A.; Goel, N.; Goel, A.; Gupta, T.; Krishnatry, R.; Kannan, S.; Kurkure, P.; et al. Real-time PCR assay based on the differential expression of microRNAs and protein-coding genes for molecular classification of formalin-fixed paraffin embedded medulloblastomas. Neuro. Oncol. 2013, 15, 1644–1651. [Google Scholar] [CrossRef] [PubMed]
- Fernandez, L.A.; Northcott, P.A.; Taylor, M.D.; Kenney, A.M. Normal and oncogenic roles for microRNAs in the developing brain. Cell Cycle 2009, 8, 4049–4054. [Google Scholar] [CrossRef] [PubMed]
- Zhao, C.; Deng, W.; Gage, F.H. Mechanisms and functional implications of adult neurogenesis. Cell 2008, 132, 645–660. [Google Scholar] [CrossRef] [PubMed]
- Yu, F.; Yao, H.; Zhu, P.; Zhang, X.; Pan, Q.; Gong, C.; Huang, Y.; Hu, X.; Su, F.; Lieberman, J.; et al. Let-7 regulates self renewal and tumorigenicity of breast cancer cells. Cell 2007, 131, 1109–1123. [Google Scholar] [CrossRef] [PubMed]
- Silber, J.; Lim, D.A.; Petritsch, C.; Persson, A.I.; Maunakea, A.K.; Yu, M.; Vandenberg, S.R.; Ginzinger, D.G.; James, C.D.; Costello, J.F.; et al. miR-124 and miR-137 inhibit proliferation of glioblastoma multiforme cells and induce differentiation of brain tumor stem cells. BMC Med. 2008, 6, 14. [Google Scholar] [CrossRef] [PubMed]
- DeSano, J.T.; Xu, L. MicroRNA regulation of cancer stem cells and therapeutic implications. AAPS J. 2009, 11, 682–692. [Google Scholar] [CrossRef] [PubMed]
- Hatfield, S.; Ruohola-Baker, H. MicroRNA and stem cell function. Cell Tissue Res. 2008, 331, 57–66. [Google Scholar] [CrossRef] [PubMed]
- Croce, C.M.; Calin, G.A. miRNAs, cancer, and stem cell division. Cell 2005, 122, 6–7. [Google Scholar] [CrossRef] [PubMed]
- Bian, S.; Hong, J.; Li, Q.; Schebelle, L.; Pollock, A.; Knauss, J.L.; Garg, V.; Sun, T. MicroRNA cluster miR-17-92 regulates neural stem cell expansion and transition to intermediate progenitors in the developing mouse neocortex. Cell Rep. 2013, 3, 1398–1406. [Google Scholar] [CrossRef] [PubMed]
- Fiaschetti, G.; Abela, L.; Nonoguchi, N.; Dubuc, A.M.; Remke, M.; Boro, A.; Grunder, E.; Siler, U.; Ohgaki, H.; Taylor, M.D.; et al. Epigenetic silencing of miRNA-9 is associated with HES1 oncogenic activity and poor prognosis of medulloblastoma. Br. J. Cancer 2014, 110, 636–647. [Google Scholar] [CrossRef]
- Spence, T.; Nguyen, J.; Bouffet, E.; Huang, A. MicroRNAs in brain tumors. In MicroRNAs in Cancer Translational Research; Cho, W.C.S., Ed.; Springer Netherlands: Berlin, Germany, 2011; pp. 343–371. [Google Scholar]
- Zollo, M.; Andolfo, I.; de Antonellis, P. MicroRNAs and cancer stem cells in medulloblastoma. In Cancer Stem Cells Therories and Practice; Shostak, S., Ed.; In Tech: Rijeka, Croatia, 2011; pp. 313–332. [Google Scholar]
- Lin, S.L.; Chang, D.C.; Chang-Lin, S.; Lin, C.H.; Wu, D.T.; Chen, D.T.; Ying, S.Y. Mir-302 reprograms human skin cancer cells into a pluripotent ES-cell-like state. RNA 2008, 14, 2115–2124. [Google Scholar] [CrossRef] [PubMed]
- Singh, S.K.; Hawkins, C.; Clarke, I.D.; Squire, J.A.; Bayani, J.; Hide, T.; Henkelman, R.M.; Cusimano, M.D.; Dirks, P.B. Identification of human brain tumour initiating cells. Nature 2004, 432, 396–401. [Google Scholar] [CrossRef] [PubMed]
- Singh, S.K.; Clarke, I.D.; Terasaki, M.; Bonn, V.E.; Hawkins, C.; Squire, J.; Dirks, P.B. Identification of a cancer stem cell in human brain tumors. Cancer Res. 2003, 63, 5821–5828. [Google Scholar] [PubMed]
- Yang, Z.J.; Ellis, T.; Markant, S.L.; Read, T.A.; Kessler, J.D.; Bourboulas, M.; Schuller, U.; Machold, R.; Fishell, G.; Rowitch, D.H.; et al. Medulloblastoma can be initiated by deletion of patched in lineage-restricted progenitors or stem cells. Cancer Cell 2008, 14, 135–145. [Google Scholar] [CrossRef] [PubMed]
- Fan, X.; Eberhart, C.G. Medulloblastoma stem cells. J. Clin. Oncol. 2008, 26, 2821–2827. [Google Scholar] [CrossRef]
- Mantamadiotis, T.; Taraviras, S. Self-renewal mechanisms in neural cancer stem cells. Front. Biosci. 2011, 16, 598–607. [Google Scholar] [CrossRef]
- Schroeder, K.; Gururangan, S. Molecular variants and mutations in medulloblastoma. Pharmacogenomics Pers. Med. 2014, 7, 43–51. [Google Scholar]
- Thompson, M.C.; Fuller, C.; Hogg, T.L.; Dalton, J.; Finkelstein, D.; Lau, C.C.; Chintagumpala, M.; Adesina, A.; Ashley, D.M.; Kellie, S.J.; et al. Genomics identifies medulloblastoma subgroups that are enriched for specific genetic alterations. J. Clin. Oncol. 2006, 24, 1924–1931. [Google Scholar] [CrossRef] [PubMed]
- Kool, M.; Koster, J.; Bunt, J.; Hasselt, N.E.; Lakeman, A.; van Sluis, P.; Troost, D.; Meeteren, N.S.; Caron, H.N.; Cloos, J.; et al. Integrated genomics identifies five medulloblastoma subtypes with distinct genetic profiles, pathway signatures and clinicopathological features. PLoS One 2008, 3, e3088. [Google Scholar] [CrossRef] [PubMed]
- Northcott, P.A.; Korshunov, A.; Witt, H.; Hielscher, T.; Eberhart, C.G.; Mack, S.; Bouffet, E.; Clifford, S.C.; Hawkins, C.E.; French, P.; et al. Medulloblastoma comprises four distinct molecular variants. J. Clin. Oncol. 2011, 29, 1408–1414. [Google Scholar] [CrossRef] [PubMed]
- Remke, M.; Hielscher, T.; Korshunov, A.; Northcott, P.A.; Bender, S.; Kool, M.; Westermann, F.; Benner, A.; Cin, H.; Ryzhova, M.; et al. FSTL5 is a marker of poor prognosis in non-WNT/non-SHH medulloblastoma. J. Clin. Oncol. 2011, 29, 3852–3861. [Google Scholar] [CrossRef] [PubMed]
- Taylor, M.D.; Northcott, P.A.; Korshunov, A.; Remke, M.; Cho, Y.J.; Clifford, S.C.; Eberhart, C.G.; Parsons, D.W.; Rutkowski, S.; Gajjar, A.; et al. Molecular subgroups of medulloblastoma: The current consensus. Acta Neuropathol. 2012, 123, 465–472. [Google Scholar] [CrossRef] [PubMed]
- Remke, M.; Ramaswamy, V.; Taylor, M.D. Medulloblastoma molecular dissection: The way toward targeted therapy. Curr. Opin. Oncol. 2013, 25, 674–681. [Google Scholar] [CrossRef] [PubMed]
- Ajeawung, N.F.; Li, B.; Kamnasaran, D. Translational applications of microRNA genes in medulloblastomas. Clin. Investig. Med. 2010, 33, E223–E233. [Google Scholar]
- Mei, H.; Lin, Z.Y.; Tong, Q.S. The roles of microRNAs in neuroblastoma. World J. Pediatr. 2014, 10, 10–16. [Google Scholar] [CrossRef]
- Pierson, J.; Hostager, B.; Fan, R.; Vibhakar, R. Regulation of cyclin dependent kinase 6 by microRNA 124 in medulloblastoma. J. Neurooncol. 2008, 90, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Cheng, L.C.; Pastrana, E.; Tavazoie, M.; Doetsch, F. miR-124 regulates adult neurogenesis in the subventricular zone stem cell niche. Nat. Neurosci. 2009, 12, 399–408. [Google Scholar] [CrossRef] [PubMed]
- Conaco, C.; Otto, S.; Han, J.J.; Mandel, G. Reciprocal actions of REST and a microRNA promote neuronal identity. Proc. Natl. Acad. Sci. USA 2006, 103, 2422–2427. [Google Scholar] [CrossRef] [PubMed]
- Makeyev, E.V.; Zhang, J.; Carrasco, M.A.; Maniatis, T. The microRNA miR-124 promotes neuronal differentiation by triggering brain-specific alternative pre-mRNA splicing. Mol. Cell 2007, 27, 435–448. [Google Scholar] [CrossRef] [PubMed]
- Mendrzyk, F.; Radlwimmer, B.; Joos, S.; Kokocinski, F.; Benner, A.; Stange, D.E.; Neben, K.; Fiegler, H.; Carter, N.P.; Reifenberger, G.; et al. Genomic and protein expression profiling identifies CDK6 as novel independent prognostic marker in medulloblastoma. J. Clin. Oncol. 2005, 23, 8853–8862. [Google Scholar] [CrossRef] [PubMed]
- Northcott, P.A.; Fernandez, L.A.; Hagan, J.P.; Ellison, D.W.; Grajkowska, W.; Gillespie, Y.; Grundy, R.; van Meter, T.; Rutka, J.T.; Croce, C.M.; et al. The miR-17/92 polycistron is up-regulated in sonic hedgehog-driven medulloblastomas and induced by N-myc in sonic hedgehog-treated cerebellar neural precursors. Cancer Res. 2009, 69, 3249–3255. [Google Scholar] [CrossRef]
- Pang, J.C.; Kwok, W.K.; Chen, Z.; Ng, H.K. Oncogenic role of microRNAs in brain tumors. Acta Neuropathol. 2009, 117, 599–611. [Google Scholar] [CrossRef]
- Ciafrè, S.A.; Galardi, S.; Mangiola, A.; Ferracin, M.; Liu, C.G.; Sabatino, G.; Negrini, M.; Maira, G.; Croce, C.M.; Farace, M.G. Extensive modulation of a set of microRNAs in primary glioblastoma. Biochem. Biophys. Res. Commun. 2005, 334, 1351–1358. [Google Scholar] [CrossRef] [PubMed]
- Turner, J.D.; Williamson, R.; Almefty, K.K.; Nakaji, P.; Porter, R.; Tse, V.; Kalani, M.Y. The many roles of microRNAs in brain tumor biology. Neurosurg. Focus. 2010, 28, E3. [Google Scholar] [CrossRef] [PubMed]
- Birks, D.K.; Barton, V.N.; Donson, A.M.; Handler, M.H.; Vibhakar, R.; Foreman, N.K. Survey of MicroRNA expression in pediatric brain tumors. Pediatr. Blood Cancer 2011, 56, 211–216. [Google Scholar] [CrossRef] [PubMed]
- Weeraratne, S.D.; Amani, V.; Teider, N.; Pierre-Francois, J.; Winter, D.; Kye, M.J.; Sengupta, S.; Archer, T.; Remke, M.; Bai, A.H.; et al. Pleiotropic effects of miR-183-96–182 converge to regulate cell survival, proliferation and migration in medulloblastoma. Acta Neuropathol. 2012, 123, 539–552. [Google Scholar] [CrossRef] [PubMed]
- Ferretti, E.; de Smaele, E.; Po, A.; di Marcotullio, L.; Tosi, E.; Espinola, M.S.; di Rocco, C.; Riccardi, R.; Giangaspero, F.; Farcomeni, A.; et al. MicroRNA profiling in human medulloblastoma. Int. J. Cancer 2009, 124, 568–577. [Google Scholar] [CrossRef] [PubMed]
- Chan, J.A.; Krichevsky, A.M.; Kosik, K.S. MicroRNA-21 is an antiapoptotic factor in human glioblastoma cells. Cancer Res. 2005, 65, 6029–6033. [Google Scholar] [CrossRef] [PubMed]
- Uziel, T.; Karginov, F.V.; Xie, S.; Parker, J.S.; Wang, Y.D.; Gajjar, A.; He, L.; Ellison, D.; Gilbertson, R.J.; Hannon, G.; et al. The miR-17-92 cluster collaborates with the Sonic Hedgehog pathway in medulloblastoma. Proc. Natl. Acad. Sci. USA 2009, 106, 2812–2817. [Google Scholar] [CrossRef] [PubMed]
- Northcott, P.A.; Nakahara, Y.; Wu, X.; Feuk, L.; Ellison, D.W.; Croul, S.; Mack, S.; Kongkham, P.N.; Peacock, J.; Dubuc, A.; et al. Multiple recurrent genetic events converge on control of histone lysine methylation in medulloblastoma. Nat. Genet. 2009, 41, 465–472. [Google Scholar] [CrossRef] [PubMed]
- Murphy, B.L.; Obad, S.; Bihannic, L.; Ayrault, O.; Zindy, F.; Kauppinen, S.; Roussel, M.F. Silencing of the miR-17-92 cluster family inhibits medulloblastoma progression. Cancer Res. 2013, 73, 7068–7078. [Google Scholar] [CrossRef] [PubMed]
- Roussel, M.F.; Robinson, G.W. Role of MYC in medulloblastoma. Cold Spring Harb. Perspect. Med. 2013, 3, a014308. [Google Scholar] [CrossRef] [PubMed]
- Bai, A.H.; Milde, T.; Remke, M.; Rolli, C.G.; Hielscher, T.; Cho, Y.J.; Kool, M.; Northcott, P.A.; Jugold, M.; Bazhin, A.V.; et al. MicroRNA-182 promotes leptomeningeal spread of non-sonic hedgehog-medulloblastoma. Acta Neuropathol. 2012, 123, 529–538. [Google Scholar] [CrossRef]
- Cho, Y.J.; Tsherniak, A.; Tamayo, P.; Santagata, S.; Ligon, A.; Greulich, H.; Berhoukim, R.; Amani, V.; Goumnerova, L.; Eberhart, C.G.; et al. Integrative genomic analysis of medulloblastoma identifies a molecular subgroup that drives poor clinical outcome. J. Clin. Oncol. 2011, 29, 1424–1430. [Google Scholar] [CrossRef] [PubMed]
- Gokhale, A.; Kunder, R.; Goel, A.; Sarin, R.; Moiyadi, A.; Shenoy, A.; Mamidipally, C.; Noronha, S.; Kannan, S.; Shirsat, N.V. Distinctive microRNA signature of medulloblastomas associated with the WNT signaling pathway. J. Cancer Res. Ther. 2010, 6, 521–529. [Google Scholar] [CrossRef] [PubMed]
- Grunder, E.; D’Ambrosio, R.; Fiaschetti, G.; Abela, L.; Arcaro, A.; Zuzak, T.; Ohgaki, H.; Lv, S.Q.; Shalaby, T.; Grotzer, M. MicroRNA-21 suppression impedes medulloblastoma cell migration. Eur. J. Cancer 2011, 47, 2479–2490. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Lee, K.F.; Lu, Y.; Clarke, I.; Shih, D.; Eberhart, C.; Collins, V.P.; van Meter, T.; Picard, D.; Zhou, L.; et al. Frequent amplification of a chr19q13.41 microRNA polycistron in aggressive primitive neuroectodermal brain tumors. Cancer Cell 2009, 16, 533–546. [Google Scholar] [CrossRef] [PubMed]
- Wang, F.; Xue, X.; Wei, J.; An, Y.; Yao, J.; Cai, H.; Wu, J.; Dai, C.; Qian, Z.; Xu, Z.; et al. Hsa-miR-520h downregulates ABCG2 in pancreatic cancer cells to inhibit migration, invasion, and side populations. Br. J. Cancer 2010, 103, 567–574. [Google Scholar] [CrossRef] [PubMed]
- Dews, M.; Homayouni, A.; Yu, D.; Murphy, D.; Sevignani, C.; Wentzel, E.; Furth, E.E.; Lee, W.M.; Enders, G.H.; Mendell, J.T.; et al. Augmentation of tumor angiogenesis by a Myc-activated microRNA cluster. Nat. Genet. 2006, 38, 1060–1065. [Google Scholar] [CrossRef] [PubMed]
- Mavrakis, K.J.; Wolfe, A.L.; Oricchio, E.; Palomero, T.; de Keersmaecker, K.; McJunkin, K.; Zuber, J.; James, T.; Khan, A.A.; Leslie, C.S.; et al. Genome-wide RNA-mediated interference screen identifies miR-19 targets in Notch-induced T-cell acute lymphoblastic leukaemia. Nat. Cell Biol. 2010, 12, 372–379. [Google Scholar] [CrossRef] [PubMed]
- Lu, Y.; Ryan, S.L.; Elliott, D.J.; Bignell, G.R.; Futreal, P.A.; Ellison, D.W.; Bailey, S.; Clifford, S.C. Amplification and overexpression of Hsa-miR-30b, Hsa-miR-30d and KHDRBS3 at 8q24.22-q24.23 in medulloblastoma. PLoS One 2009, 4, e6159. [Google Scholar] [CrossRef] [PubMed]
- Liu, W.; Gong, Y.H.; Chao, T.F.; Peng, X.Z.; Yuan, J.G.; Ma, Z.Y.; Jia, G.; Zhao, J.Z. Identification of differentially expressed microRNAs by microarray: A possible role for microRNAs gene in medulloblastomas. Chin. Med. J. 2009, 122, 2405–2411. [Google Scholar] [PubMed]
- Kefas, B.; Godlewski, J.; Comeau, L.; Li, Y.; Abounader, R.; Hawkinson, M.; Lee, J.; Fine, H.; Chiocca, E.A.; Lawler, S.; et al. MicroRNA-7 inhibits the epidermal growth factor receptor and the Akt pathway and is down-regulated in glioblastoma. Cancer Res. 2008, 68, 3566–3572. [Google Scholar] [CrossRef] [PubMed]
- Roush, S.; Slack, F.J. The let-7 family of microRNAs. Trends Cell Biol. 2008, 18, 505–516. [Google Scholar] [CrossRef]
- Garzia, L.; Andolfo, I.; Cusanelli, E.; Marino, N.; Petrosino, G.; de Martino, D.; Esposito, V.; Galeone, A.; Navas, L.; Esposito, S.; et al. MicroRNA-199b-5p impairs cancer stem cells through negative regulation of HES1 in medulloblastoma. PLoS One 2009, 4, e4998. [Google Scholar] [CrossRef]
- Li, K.K.; Pang, J.C.; Ching, A.K.; Wong, C.K.; Kong, X.; Wang, Y.; Zhou, L.; Chen, Z.; Ng, H.K. MiR-124 is frequently down-regulated in medulloblastoma and is a negative regulator of SLC16A1. Hum. Pathol. 2009, 40, 1234–1243. [Google Scholar] [CrossRef] [PubMed]
- Yoo, A.S.; Staahl, B.T.; Chen, L.; Crabtree, G.R. MicroRNA-mediated switching of chromatin-remodelling complexes in neural development. Nature 2009, 460, 642–646. [Google Scholar] [PubMed]
- Wei, J.S.; Johansson, P.; Chen, Q.R.; Song, Y.K.; Durinck, S.; Wen, X.; Cheuk, A.T.; Smith, M.A.; Houghton, P.; Morton, C.; et al. MicroRNA profiling identifies cancer-specific and prognostic signatures in pediatric malignancies. Clin. Cancer Res. 2009, 15, 5560–5568. [Google Scholar] [CrossRef] [PubMed]
- Venkataraman, S.; Alimova, I.; Fan, R.; Harris, P.; Foreman, N.; Vibhakar, R. MicroRNA 128a increases intracellular ROS level by targeting Bmi-1 and inhibits medulloblastoma cancer cell growth by promoting senescence. PLoS One 2010, 5, e10748. [Google Scholar] [CrossRef]
- Ma, L.; Weinberg, R.A. Micromanagers of malignancy: Role of microRNAs in regulating metastasis. Trends Genet. 2008, 24, 448–456. [Google Scholar] [CrossRef]
- Ventura, A.; Jacks, T. MicroRNAs and cancer: Short RNAs go a long way. Cell 2009, 136, 586–591. [Google Scholar] [CrossRef] [PubMed]
- White, N.M.; Fatoohi, E.; Metias, M.; Jung, K.; Stephan, C.; Yousef, G.M. Metastamirs: A stepping stone towards improved cancer management. Nat. Rev. Clin. Oncol. 2010, 8, 75–84. [Google Scholar] [CrossRef] [PubMed]
- Asangani, I.A.; Rasheed, S.A.; Nikolova, D.A.; Leupold, J.H.; Colburn, N.H.; Post, S.; Allgayer, H. MicroRNA-21 (miR-21) post-transcriptionally downregulates tumor suppressor Pdcd4 and stimulates invasion, intravasation and metastasis in colorectal cancer. Oncogene 2008, 27, 2128–2136. [Google Scholar] [CrossRef]
- Li, K.K.; Xia, T.; Ma, F.M.; Zhang, R.; Mao, Y.; Wang, Y.; Zhou, L.; Lau, K.M.; Ng, H.K. MiR-106b is overexpressed in medulloblastomas and interacts directly with PTEN. Neuropathol. Appl. Neurobiol. 2014. [Google Scholar]
- Shi, J.A.; Lu, D.L.; Huang, X.; Tan, W. MiR-219 inhibits the proliferation, migration and invasion of medulloblastoma cells by targeting CD164. Int. J. Mol. Med. 2014, 34, 237–243. [Google Scholar] [PubMed]
- Lin, P.Y.; Yu, S.L.; Yang, P.C. MicroRNA in lung cancer. Br. J. Cancer 2010, 103, 1144–1148. [Google Scholar] [CrossRef] [PubMed]
- Rorke, L.B.; Packer, R.J.; Biegel, J.A. Central nervous system atypical teratoid/rhabdoid tumors of infancy and childhood: Definition of an entity. J. Neurosurg. 1996, 85, 56–65. [Google Scholar] [CrossRef] [PubMed]
- Reddy, A.T. Atypical teratoid/rhabdoid tumors of the central nervous system. J. Neurooncol. 2005, 75, 309–313. [Google Scholar] [CrossRef] [PubMed]
- Squire, S.E.; Chan, M.D.; Marcus, K.J. Atypical teratoid/rhabdoid tumor: The controversy behind radiation therapy. J. Neurooncol. 2007, 81, 97–111. [Google Scholar] [CrossRef] [PubMed]
- Oda, Y.; Tsuneyoshi, M. Extrarenal rhabdoid tumors of soft tissue: Clinicopathological and molecular genetic review and distinction from other soft-tissue sarcomas with rhabdoid features. Pathol. Int. 2006, 56, 287–295. [Google Scholar] [CrossRef] [PubMed]
- Versteege, I.; Sevenet, N.; Lange, J.; Rousseau-Merck, M.F.; Ambros, P.; Handgretinger, R.; Aurias, A.; Delattre, O. Truncating mutations of hSNF5/INI1 in aggressive paediatric cancer. Nature 1998, 394, 203–206. [Google Scholar] [CrossRef]
- Biegel, J.A.; Tan, L.; Zhang, F.; Wainwright, L.; Russo, P.; Rorke, L.B. Alterations of the hSNF5/INI1 gene in central nervous system atypical teratoid/rhabdoid tumors and renal and extrarenal rhabdoid tumors. Clin. Cancer Res. 2002, 8, 3461–3467. [Google Scholar] [PubMed]
- Bambakidis, N.C.; Robinson, S.; Cohen, M.; Cohen, A.R. Atypical teratoid/rhabdoid tumors of the central nervous system: Clinical, radiographic and pathologic features. Pediatr. Neurosurg. 2002, 37, 64–70. [Google Scholar] [PubMed]
- Hilden, J.M.; Meerbaum, S.; Burger, P.; Finlay, J.; Janss, A.; Scheithauer, B.W.; Walter, A.W.; Rorke, L.B.; Biegel, J.A. Central nervous system atypical teratoid/rhabdoid tumor: Results of therapy in children enrolled in a registry. J. Clin. Oncol. 2004, 22, 2877–2884. [Google Scholar] [CrossRef] [PubMed]
- Tekautz, T.M.; Fuller, C.E.; Blaney, S.; Fouladi, M.; Broniscer, A.; Merchant, T.E.; Krasin, M.; Dalton, J.; Hale, G.; Kun, L.E.; et al. Atypical teratoid/rhabdoid tumors (ATRT): Improved survival in children 3 years of age and older with radiation therapy and high-dose alkylator-based chemotherapy. J. Clin. Oncol. 2005, 23, 1491–1499. [Google Scholar] [CrossRef]
- Mitchell, P.S.; Parkin, R.K.; Kroh, E.M.; Fritz, B.R.; Wyman, S.K.; Pogosova-Agadjanyan, E.L.; Peterson, A.; Noteboom, J.; O’Briant, K.C.; Allen, A.; et al. Circulating microRNAs as stable blood-based markers for cancer detection. Proc. Natl. Acad. Sci. USA 2008, 105, 10513–10518. [Google Scholar] [CrossRef] [PubMed]
- White, N.M.; Bao, T.T.; Grigull, J.; Youssef, Y.M.; Girgis, A.; Diamandis, M.; Fatoohi, E.; Metias, M.; Honey, R.J.; Stewart, R.; et al. miRNA profiling for clear cell renal cell carcinoma: Biomarker discovery and identification of potential controls and consequences of miRNA dysregulation. J. Urol. 2011, 186, 1077–1083. [Google Scholar] [CrossRef] [PubMed]
- Heneghan, H.M.; Miller, N.; Kelly, R.; Newell, J.; Kerin, M.J. Systemic miRNA-195 differentiates breast cancer from other malignancies and is a potential biomarker for detecting noninvasive and early stage disease. Oncologist 2010, 15, 673–682. [Google Scholar] [CrossRef] [PubMed]
- Kapsimali, M.; Kloosterman, W.P.; de Bruijn, E.; Rosa, F.; Plasterk, R.H.; Wilson, S.W. MicroRNAs show a wide diversity of expression profiles in the developing and mature central nervous system. Genome Biol. 2007, 8, R173. [Google Scholar] [CrossRef] [PubMed]
- Nass, D.; Rosenwald, S.; Meiri, E.; Gilad, S.; Tabibian-Keissar, H.; Schlosberg, A.; Kuker, H.; Sion-Vardy, N.; Tobar, A.; Kharenko, O.; et al. miR-92b and miR-9/9* are specifically expressed in brain primary tumors and can be used to differentiate primary from metastatic brain tumors. Brain Pathol. 2009, 19, 375–383. [Google Scholar] [CrossRef] [PubMed]
- Osaki, M.; Takeshita, F.; Ochiya, T. MicroRNAs as biomarkers and therapeutic drugs in human cancer. Biomarkers 2008, 13, 658–670. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Chen, J.; Chang, P.; LeBlanc, A.; Li, D.; Abbruzzesse, J.L.; Frazier, M.L.; Killary, A.M.; Sen, S. MicroRNAs in plasma of pancreatic ductal adenocarcinoma patients as novel blood-based biomarkers of disease. Cancer Prev. Res. (Phila) 2009, 2, 807–813. [Google Scholar] [CrossRef]
- Kanaan, Z.; Roberts, H.; Eichenberger, M.R.; Billeter, A.; Ocheretner, G.; Pan, J.; Rai, S.N.; Jorden, J.; Williford, A.; Galandiuk, S. A plasma microRNA panel for detection of colorectal adenomas: A step toward more precise screening for colorectal cancer. Ann. Surg. 2013, 258, 400–408. [Google Scholar] [CrossRef] [PubMed]
- Teplyuk, N.M.; Mollenhauer, B.; Gabriely, G.; Giese, A.; Kim, E.; Smolsky, M.; Kim, R.Y.; Saria, M.G.; Pastorino, S.; Kesari, S.; et al. MicroRNAs in cerebrospinal fluid identify glioblastoma and metastatic brain cancers and reflect disease activity. Neuro Oncol. 2012, 14, 689–700. [Google Scholar] [PubMed]
- Baraniskin, A.; Kuhnhenn, J.; Schlegel, U.; Maghnouj, A.; Zöllner, H.; Schmiegel, W.; Hahn, S.; Schroers, R. Identification of microRNAs in the cerebrospinal fluid as biomarker for the diagnosis of glioma. Neuro Oncol. 2012, 14, 29–33. [Google Scholar] [CrossRef] [PubMed]
- De Smaele, E.; Ferretti, E.; Gulino, A. MicroRNAs as biomarkers for CNS cancer and other disorders. Brain Res. 2010, 1338, 100–111. [Google Scholar] [CrossRef] [PubMed]
- Jin, X.F.; Wu, N.; Wang, L.; Li, J. Circulating microRNAs: A novel class of potential biomarkers for diagnosing and prognosing central nervous system diseases. Cell Mol. Neurobiol. 2013, 33, 601–613. [Google Scholar] [CrossRef] [PubMed]
- Wang, K.; Zhang, S.; Marzolf, B.; Troisch, P.; Brightman, A.; Hu, Z.; Hood, L.E.; Galas, D.J. Circulating microRNAs, potential biomarkers for drug-induced liver injury. Proc. Natl. Acad. Sci. USA 2009, 106, 4402–4407. [Google Scholar] [CrossRef] [PubMed]
- Cogswell, J.P.; Ward, J.; Taylor, I.A.; Waters, M.; Shi, Y.; Cannon, B.; Kelnar, K.; Kemppainen, J.; Brown, D.; Chen, C.; et al. Identification of miRNA changes in Alzheimer’s disease brain and CSF yields putative biomarkers and insights into disease pathways. J. Alzheimer’s Dis. 2008, 14, 27–41. [Google Scholar]
- Chen, X.; Liang, H.; Zhang, J.; Zen, K.; Zhang, C.Y. Secreted microRNAs: A new form of intercellular communication. Trends Cell Biol. 2012, 22, 125–132. [Google Scholar] [CrossRef] [PubMed]
- Sidransky, D. Emerging molecular markers of cancer. Nat. Rev. Cancer 2002, 2, 210–219. [Google Scholar] [CrossRef] [PubMed]
- Day, R.W. Future need for more cancer research. J. Am. Diet. Assoc. 1998, 98, 523. [Google Scholar] [CrossRef] [PubMed]
- Lewin, D.A.; Weiner, M.P. Molecular biomarkers in drug development. Drug Discov. Today 2004, 9, 976–983. [Google Scholar] [CrossRef] [PubMed]
- Barker, P.E. Cancer biomarker validation: Standards and process: Roles for the National Institute of Standards and Technology (NIST). Ann. N. Y. Acad. Sci. 2003, 983, 142–150. [Google Scholar] [CrossRef] [PubMed]
- Pepe, M.S.; Etzioni, R.; Feng, Z.; Potter, J.D.; Thompson, M.L.; Thornquist, M.; Winget, M.; Yasui, Y. Phases of biomarker development for early detection of cancer. J. Natl. Cancer Inst. 2001, 93, 1054–1061. [Google Scholar] [CrossRef] [PubMed]
- Srivastava, S. Cancer biomarker discovery and development in gastrointestinal cancers: Early detection research network—A collaborative approach. Gastrointest. Cancer Res. 2007, 1, S60–S63. [Google Scholar] [PubMed]
- Chen, S.Y.; Wang, Y.; Telen, M.J.; Chi, J.T. The genomic analysis of erythrocyte microRNA expression in sickle cell diseases. PLoS One 2008, 3, e2360. [Google Scholar] [CrossRef] [PubMed]
- Von Knebel-Doeberitz, M.; Syrjänen, K.J. Molecular markers: How to apply in practice. Gynecol. Oncol. 2006, 103, 18–20. [Google Scholar] [CrossRef] [PubMed]
- Martinkova, J.; Gadher, S.J.; Hajduch, M.; Kovarova, H. Challenges in cancer research and multifaceted approaches for cancer biomarker quest. FEBS Lett. 2009, 583, 1772–1784. [Google Scholar] [CrossRef] [PubMed]
- Etheridge, A.; Lee, I.; Hood, L.; Galas, D.; Wang, K. Extracellular microRNA: A new source of biomarkers. Mutat. Res. 2011, 717, 85–90. [Google Scholar] [CrossRef] [PubMed]
- Lu, J.; Getz, G.; Miska, E.A.; Alvarez-Saavedra, E.; Lamb, J.; Peck, D.; Sweet-Cordero, A.; Ebert, B.L.; Mak, R.H.; Ferrando, A.A.; et al. MicroRNA expression profiles classify human cancers. Nature 2005, 435, 834–838. [Google Scholar] [CrossRef] [PubMed]
- Lawrie, C.H.; Soneji, S.; Marafioti, T.; Cooper, C.D.; Palazzo, S.; Paterson, J.C.; Cattan, H.; Enver, T.; Mager, R.; Boultwood, J.; et al. MicroRNA expression distinguishes between germinal center B cell-like and activated B cell-like subtypes of diffuse large B cell lymphoma. Int. J. Cancer 2007, 121, 1156–1161. [Google Scholar] [CrossRef] [PubMed]
- Xi, Y.; Nakajima, G.; Gavin, E.; Morris, C.G.; Kudo, K.; Hayashi, K.; Ju, J. Systematic analysis of microRNA expression of RNA extracted from fresh frozen and formalin-fixed paraffin-embedded samples. RNA 2007, 13, 1668–1674. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Ba, Y.; Ma, L.; Cai, X.; Yin, Y.; Wang, K.; Guo, J.; Zhang, Y.; Chen, J.; Guo, X.; et al. Characterization of microRNAs in serum: A novel class of biomarkers for diagnosis of cancer and other diseases. Cell Res. 2008, 18, 997–1006. [Google Scholar] [CrossRef] [PubMed]
- Koshiol, J.; Wang, E.; Zhao, Y.; Marincola, F.; Landi, M.T. Strengths and limitations of laboratory procedures for microRNA detection. Cancer Epidemiol. Biomark. Prev. 2010, 19, 907–911. [Google Scholar]
- Jay, C.; Nemunaitis, J.; Chen, P.; Fulgham, P.; Tong, A.W. miRNA profiling for diagnosis and prognosis of human cancer. DNA Cell Biol. 2007, 26, 293–300. [Google Scholar] [CrossRef] [PubMed]
- Rosenfeld, N.; Aharonov, R.; Meiri, E.; Rosenwald, S.; Spector, Y.; Zepeniuk, M.; Benjamin, H.; Shabes, N.; Tabak, S.; Levy, A.; et al. MicroRNAs accurately identify cancer tissue origin. Nat. Biotechnol. 2008, 26, 462–469. [Google Scholar] [CrossRef] [PubMed]
- Yanaihara, N.; Caplen, N.; Bowman, E.; Seike, M.; Kumamoto, K.; Yi, M.; Stephens, R.M.; Okamoto, A.; Yokota, J.; Tanaka, T.; et al. Unique microRNA molecular profiles in lung cancer diagnosis and prognosis. Cancer Cell 2006, 9, 189–198. [Google Scholar] [CrossRef]
- Kim, W.T.; Kim, W.J. MicroRNAs in prostate cancer. Prostate Int. 2013, 1, 3–9. [Google Scholar] [CrossRef]
- Wang, Y.; Gao, X.; Wei, F.; Zhang, X.; Yu, J.; Zhao, H.; Sun, Q.; Yan, F.; Yan, C.; Li, H.; et al. Diagnostic and prognostic value of circulating miR-21 for cancer: A systematic review and meta-analysis. Gene 2014, 533, 389–397. [Google Scholar] [CrossRef] [PubMed]
- Yu, S.L.; Chen, H.Y.; Yang, P.C.; Chen, J.J. Unique microRNA signature and clinical outcome of cancers. DNA Cell Biol. 2007, 26, 283–292. [Google Scholar] [CrossRef] [PubMed]
- Izquierdo, L.; Ingelmo-Torres, M.; Mallofré, C.; Lozano, J.J.; Verhasselt-Crinquette, M.; Leroy, X.; Colin, P.; Comperat, E.; Roupret, M.; Alcaraz, A.; et al. Prognostic value of microRNA expression pattern in upper tract urothelial carcinoma. BJU Int. 2014, 113, 813–821. [Google Scholar] [CrossRef] [PubMed]
- Hotchi, M.; Shimada, M.; Kurita, N.; Iwata, T.; Sato, H.; Morimoto, S.; Yoshikawa, K.; Higashijima, J.; Miyatani, T. MicroRNA expression is able to predict response to chemoradiotherapy in rectal cancer. Mol. Clin. Oncol. 2013, 1, 137–142. [Google Scholar] [PubMed]
- Besse, A.; Sana, J.; Fadrus, P.; Slaby, O. MicroRNAs involved in chemo- and radioresistance of high-grade gliomas. Tumour Biol. 2013, 34, 1969–1978. [Google Scholar] [CrossRef] [PubMed]
- Dedeoğlu, B.G. High-throughput approaches for microRNA expression analysis. Methods Mol. Biol. 2014, 1107, 91–103. [Google Scholar] [PubMed]
- Hoffman, R.M. Orthotopic metastatic mouse models for anticancer drug discovery and evaluation: A bridge to the clinic. Investig. New Drugs 1999, 17, 343–359. [Google Scholar] [CrossRef]
- Fowler, B.A. Molecular biomarkers: Challenges and prospects for the future. Toxicol. Appl. Pharmacol. 2005, 206, 97. [Google Scholar] [CrossRef] [PubMed]
- Colburn, W.A.; Lee, J.W. Biomarkers, validation and pharmacokinetic-pharmacodynamic modelling. Clin. Pharmacokinet. 2003, 42, 997–1022. [Google Scholar] [CrossRef] [PubMed]
- Miller, K.J.; Bowsher, R.R.; Celniker, A.; Gibbons, J.; Gupta, S.; Lee, J.W.; Swanson, S.J.; Smith, W.C.; Weiner, R.S. Workshop on bioanalytical methods validation for macromolecules: Summary report. Pharm. Res. 2001, 18, 1373–1383. [Google Scholar] [CrossRef] [PubMed]
- Chau, C.H.; Rixe, O.; McLeod, H.; Figg, W.D. Validation of analytic methods for biomarkers used in drug development. Clin. Cancer Res. 2008, 14, 5967–5976. [Google Scholar] [CrossRef] [PubMed]
- Arsanious, A.; Bjarnason, G.A.; Yousef, G.M. From bench to bedside: Current and future applications of molecular profiling in renal cell carcinoma. Mol. Cancer 2009, 8, 20. [Google Scholar] [CrossRef] [PubMed]
- Adams, V.; Möbius-Winkler, S.; Schuler, G. Basic and translational research: from molecule, to mouse, to man. Eur. J. Cardiovasc. Prev. Rehabil. 2009, 16, S48–S52. [Google Scholar] [CrossRef] [PubMed]
- Clermont, G.; Auffray, C.; Moreau, Y.; Rocke, D.M.; Dalevi, D.; Dubhashi, D.; Marshall, D.R.; Raasch, P.; Dehne, F.; Provero, P.; et al. Bridging the gap between systems biology and medicine. Genome Eed. 2009, 1, 88. [Google Scholar] [CrossRef]
- Hawkins, N.; de Vries, J.; Boddington, P.; Kaye, J.; Heeney, C. Planning for translational research in genomics. Genome Eed. 2009, 1, 87. [Google Scholar] [CrossRef]
- Payne, P.R.O.; Embi, P.J.; Sen, C.K. Translational informatics: Enabling high-throughput research paradigms. Physiol. Genomics 2009, 39, 131–140. [Google Scholar] [CrossRef] [PubMed]
- Arbab, A.S.; Janic, B.; Haller, J.; Pawelczyk, E.; Liu, W.; Frank, J.A. In vivo cellular imaging for translational medical research. Curr. Med. Imaging Rev. 2009, 5, 19–38. [Google Scholar] [CrossRef] [PubMed]
- Ioannidis, J.P.; Panagiotou, O.A. Comparison of effect sizes associated with biomarkers reported in highly cited individual articles and in subsequent meta-analyses. JAMA 2011, 305, 2200–2210. [Google Scholar] [CrossRef] [PubMed]
- Adam, D. Online tumour bank aims to offer ready route to tissues. Nature 2002, 416, 464. [Google Scholar] [CrossRef] [PubMed]
- Herpel, E.; Koleganova, N.; Schirmacher, P. Tissue bank of the National Centre for Tumour Disease: An innovative platform for translational tumour. Pathologe 2008, 29, 204–209. [Google Scholar] [CrossRef] [PubMed]
- Qualman, S.J.; France, M.; Grizzle, W.E.; LiVolsi, V.A.; Moskaluk, C.A.; Ramirez, N.C.; Washington, M.K. Establishing a tumour bank: Banking, informatics and ethics. Br. J. Cancer 2004, 90, 1115–1119. [Google Scholar] [CrossRef]
- Elger, B.S.; Caplan, A.L. Consent and anonymization in research involving biobanks: Differing terms and norms present serious barriers to an international framework. EMBO Rep. 2006, 7, 661–666. [Google Scholar] [CrossRef] [PubMed]
- Chadwick, R.; Berg, K. Solidarity and equity: New ethical frameworks for genetic databases. Nat. Rev. Genet. 2001, 2, 318–321. [Google Scholar] [CrossRef] [PubMed]
- Everett, M. The social life of genes: Privacy, property and the new genetics. Soc. Sci. Med. 2003, 56, 53–65. [Google Scholar] [CrossRef] [PubMed]
- Gilbertson, R.J.; Gajjar, A. Molecular biology of medulloblastoma: Will it ever make a difference to clinical management? J. Neurooncol. 2005, 75, 273–278. [Google Scholar] [CrossRef] [PubMed]
- Ioannidis, J.P.; Allison, D.B.; Ball, C.A.; Coulibaly, I.; Cui, X.; Culhane, A.C.; Falchi, M.; Furlanello, C.; Game, L.; Jurman, G.; et al. Repeatability of published microarray gene expression analyses. Nat. Genet. 2009, 41, 149–155. [Google Scholar] [CrossRef] [PubMed]
- Nasedkin, J.N. The evaluation of occlusal dysfunctional therapy. J. Prosthet. Dent. 1980, 44, 4–9. [Google Scholar] [CrossRef] [PubMed]
- Lee, E.J.; Gusev, Y.; Jiang, J.; Nuovo, G.J.; Lerner, M.R.; Frankel, W.L.; Morgan, D.L.; Postier, R.G.; Brackett, D.J.; Schmittgen, T.D. Expression profiling identifies microRNA signature in pancreatic cancer. Int. J. Cancer 2007, 120, 1046–1054. [Google Scholar] [CrossRef] [PubMed]
- Bloomston, M.; Frankel, W.L.; Petrocca, F.; Volinia, S.; Alder, H.; Hagan, J.P.; Liu, C.G.; Bhatt, D.; Taccioli, C.; Croce, C.M. MicroRNA expression patterns to differentiate pancreatic adenocarcinoma from normal pancreas and chronic pancreatitis. JAMA 2007, 297, 1901–1908. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Z.; Moley, K.H.; Gronowski, A.M. Diagnostic potential for miRNAs as biomarkers for pregnancy-specific diseases. Clin. Biochem. 2013, 46, 953–960. [Google Scholar] [CrossRef] [PubMed]
- Ptolemy, A.S.; Rifai, N. What is a biomarker? Research investments and lack of clinical integration necessitate a review of biomarker terminology and validation schema. Scand. J. Clin. Lab. Investig. 2010, 70, 6–14. [Google Scholar]
- Wallow, I. Necrotizing retinitis in severe general diseases. Ber. Zusammenkunft Dtsch. Ophthalmol. Ges. 1970, 70, 55–58. [Google Scholar] [PubMed]
- Aebersold, R.; Auffray, C.; Baney, E.; Barillot, E.; Brazma, A.; Brett, C.; Brunak, S.; Butte, A.; Califano, A.; Celis, J.; et al. Report on EU-USA workshop: How systems biology can advance cancer research (27 October 2008). Mol. Oncol. 2009, 3, 9–17. [Google Scholar] [CrossRef] [PubMed]
- Tricoli, J.V.; Jacobson, J.W. MicroRNA: Potential for cancer dcetection, dciagnosis, and prognosis. Cancer Res. 2007, 67, 4553–4555. [Google Scholar] [CrossRef]
- George, G.P.; Mittal, R.D. MicroRNAs: Potential biomarkers in cancer. Indian J. Clin. Biochem. 2010, 25, 4–14. [Google Scholar] [CrossRef] [PubMed]
- Budhu, A.; Ji, J.; Wang, X.W. The clinical potential of microRNAs. J. Hematol. Oncol. 2010, 3, 37. [Google Scholar] [CrossRef] [PubMed]
- Shalaby, T.; Fiaschetti, G.; Baumgartner, M.; Grotzer, M.A. Significance and therapeutic value of miRNAs in embryonal neural tumors. Molecules 2014, 19, 5821–5862. [Google Scholar] [CrossRef]
- Wang, V.; Wu, W. MicroRNA-based therapeutics for cancer. Biol. Drugs 2009, 23, 15–23. [Google Scholar]
- Si, M.L.; Zhu, S.; Wu, H.; Lu, Z.; Wu, F.; Mo, Y.Y. MiR-21-mediated tumor growth. Oncogene 2007, 26, 2799–2803. [Google Scholar] [CrossRef] [PubMed]
- Gandellini, P.; Profumo, V.; Folini, M.; Zaffaroni, N. MicroRNAs as new therapeutic targets and tools in cancer. Expert Opin. Ther. Targets 2011, 15, 265–279. [Google Scholar] [CrossRef] [PubMed]
- Van Rooij, E.; Purcell, A.L.; Levin, A.A. Developing microRNA therapeutics. Circ. Res. 2010, 110, 496–507. [Google Scholar] [CrossRef]
- Hummel, R.; Hussey, D.J.; Haier, J. MicroRNAs: Predictors and modifiers of chemo- and radio-therapy in different tumour types. Eur. J. Cancer 2010, 46, 298–311. [Google Scholar] [CrossRef] [PubMed]
- Gong, C.; Yao, Y.; Wang, Y.; Liu, B.; Wu, W.; Chen, J.; Su, F.; Yao, H.; Song, E. Up-regulation of miR-21 mediates resistance to trastuzumab therapy for breast cancer. J. Biol. Chem. 2011, 286, 19127–19137. [Google Scholar] [CrossRef] [PubMed]
- Nasser, M.W.; Datta, J.; Nuovo, G.; Kutay, H.; Motiwala, T.; Majumder, S.; Wang, B.; Suster, S.; Jacob, S.T.; Ghoshal, K. Down-regulation of micro-RNA-1 (miR-1) in lung cancer: Suppression of tumorigenic property of lung cancer cells and their sensitization to doxorubicin-induced apoptosis by miR-1. J. Biol. Chem. 2008, 283, 33394–33405. [Google Scholar] [CrossRef] [PubMed]
- Nana-Sinkam, S.P.; Croce, C.M. Clinical applications for microRNAs in cancer. Clin. Pharmacol. Ther. 2012, 93, 98–104. [Google Scholar] [CrossRef] [PubMed]
- Blower, P.E.; Chung, J.H.; Verducci, J.S.; Lin, S.; Park, J.K.; Dai, Z.; Liu, C.G.; Schmittgen, T.D.; Reinhold, W.C.; Croce, C.M.; et al. MicroRNAs modulate the chemosensitivity of tumor cells. Mol. Cancer Ther. 2008, 7, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; D’Andrade, P.; Fulmer-Smentek, S.; Lorenzi, P.; Kohn, K.W.; Weinstein, J.N.; Pommier, Y.; Reinhold, W.C. mRNA and microRNA expression profiles of the NCI-60 integrated with drug activities. Mol. Cancer Ther. 2010, 9, 1080–1091. [Google Scholar] [CrossRef] [PubMed]
- Schetter, A.J.; Leung, S.Y.; Sohn, J.J.; Zanetti, K.A.; Bowman, E.D.; Yanaihara, N.; Yuen, S.T.; Chan, T.L.; Kwong, D.L.; Au, G.K.; et al. MicroRNA expression profiles associated with prognosis and therapeutic outcome in colon adenocarcinoma. JAMA 2008, 299, 425–436. [Google Scholar] [PubMed]
- Halimi, M.; Asghari, S. M.; Sariri, R.; Moslemi, D.; Parsian, H. Cellular response to ionizing radiation: A microRNA story. Int. J. Mol. Cell Med. 2012, 1, 178–184. [Google Scholar] [PubMed]
- Metheetrairut, C.; Slack, F.J. MicroRNAs in the ionizing radiation response and in radiotherapy. Curr. Opin. Genet. Dev. 2013, 23, 12–19. [Google Scholar] [CrossRef]
- Garzon, R.; Marcucci, G.; Croce, C.M. Targeting microRNAs in cancer: Rationale, strategies and challenges. Nat. Rev. Drug Discov. 2010, 9, 775–789. [Google Scholar] [CrossRef] [PubMed]
- Deeken, J.F.; Loscher, W. The blood-brain barrier and cancer: Transporters, treatment, and Trojan horses. Clin. Cancer Res. 2007, 6, 1663–1674. [Google Scholar] [CrossRef]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Shalaby, T.; Fiaschetti, G.; Baumgartner, M.; Grotzer, M.A. MicroRNA Signatures as Biomarkers and Therapeutic Target for CNS Embryonal Tumors: The Pros and the Cons. Int. J. Mol. Sci. 2014, 15, 21554-21586. https://doi.org/10.3390/ijms151121554
Shalaby T, Fiaschetti G, Baumgartner M, Grotzer MA. MicroRNA Signatures as Biomarkers and Therapeutic Target for CNS Embryonal Tumors: The Pros and the Cons. International Journal of Molecular Sciences. 2014; 15(11):21554-21586. https://doi.org/10.3390/ijms151121554
Chicago/Turabian StyleShalaby, Tarek, Giulio Fiaschetti, Martin Baumgartner, and Michael A. Grotzer. 2014. "MicroRNA Signatures as Biomarkers and Therapeutic Target for CNS Embryonal Tumors: The Pros and the Cons" International Journal of Molecular Sciences 15, no. 11: 21554-21586. https://doi.org/10.3390/ijms151121554
APA StyleShalaby, T., Fiaschetti, G., Baumgartner, M., & Grotzer, M. A. (2014). MicroRNA Signatures as Biomarkers and Therapeutic Target for CNS Embryonal Tumors: The Pros and the Cons. International Journal of Molecular Sciences, 15(11), 21554-21586. https://doi.org/10.3390/ijms151121554




