Recombinase Polymerase Amplification (RPA) of CaMV-35S Promoter and nos Terminator for Rapid Detection of Genetically Modified Crops
Abstract
:1. Introduction
2. Results and Discussion
2.1. Primer Design and Screening
| Target Elements | No. of Primer Combinations Tested | No. of Primer Sets Amplified | Fluorescence Signal > 500 mV | Threshold Time of 6–10 min |
|---|---|---|---|---|
| P-35S | 24 | 3 | 2 | 1 |
| T-nos | 16 | 2 | 1 | 1 |
| Target | Primer/Probe | Sequence (5'–3') | Amplicon (bp) |
|---|---|---|---|
| P-35S | RPA-35S-F | TATCCGGAAACCTCCTCGGATTCCATTGCCCAGC | 266 |
| RPA-35S-R | GTGGGATTGTGCGTCATCCCTTACGTCAGTG | ||
| RPA-35S-P | TCGTTGAAGATGCCTCTGCCGACAG(FAM-dT)(dSpacer) G(BHQ1-dT)CCCAAAGATGG(phosphate) | ||
| T-nos | RPA-nos-F | TAAGATTGAATCCTGTTGCCGGTCTTGCGATGA | 183 |
| RPA-nos-R | CCTAGTTTGCGCGCTATATTTTGTTTTCTATCG | ||
| RPA-nos-P | CGTTATTTATGAGATGGGTTT(FAM-dT)(dSpacer) A(BHQ1-dT)GATTAGAGTCC(phosphate) |
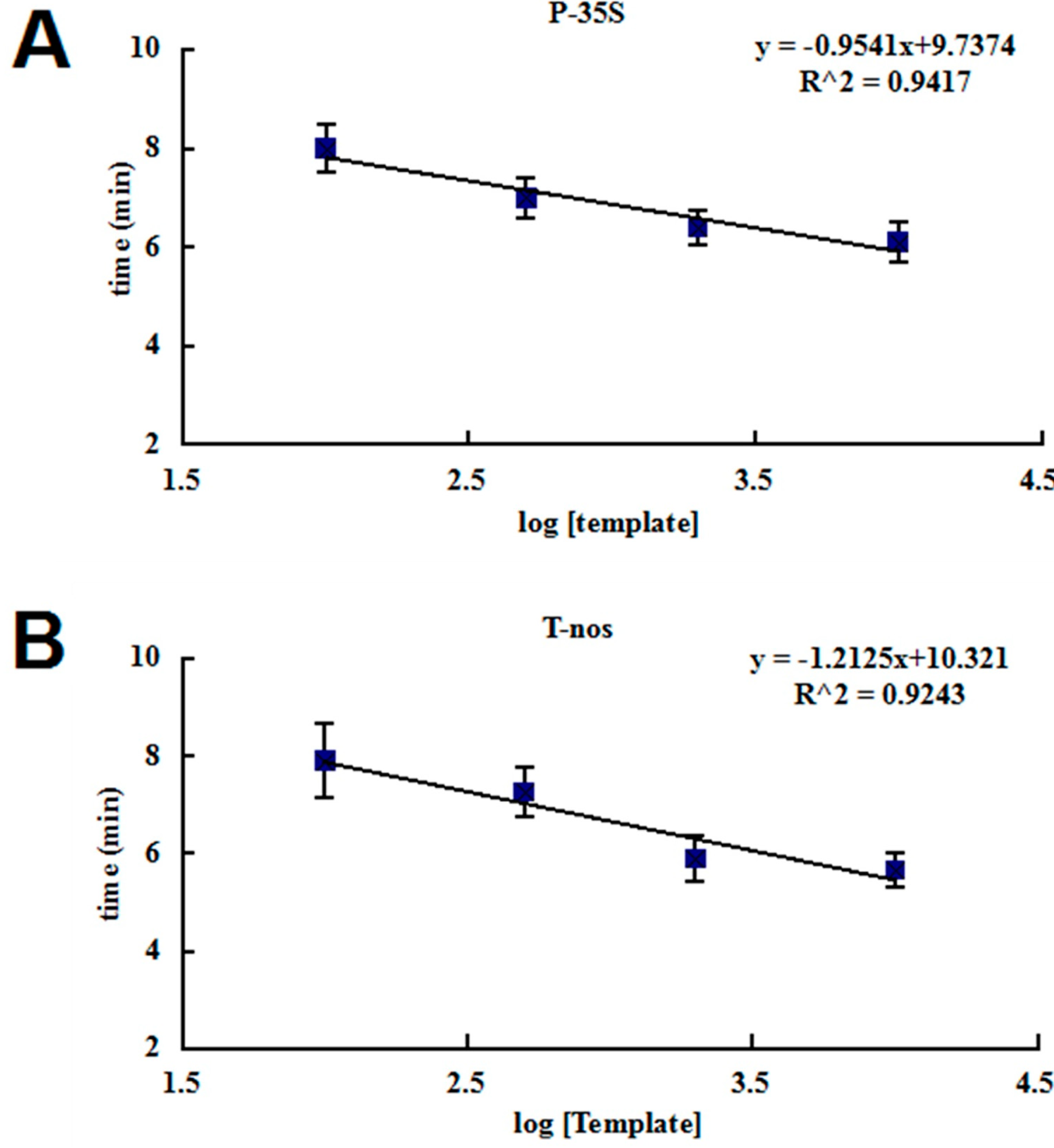
2.2. Sensitivity of the RPA Assays
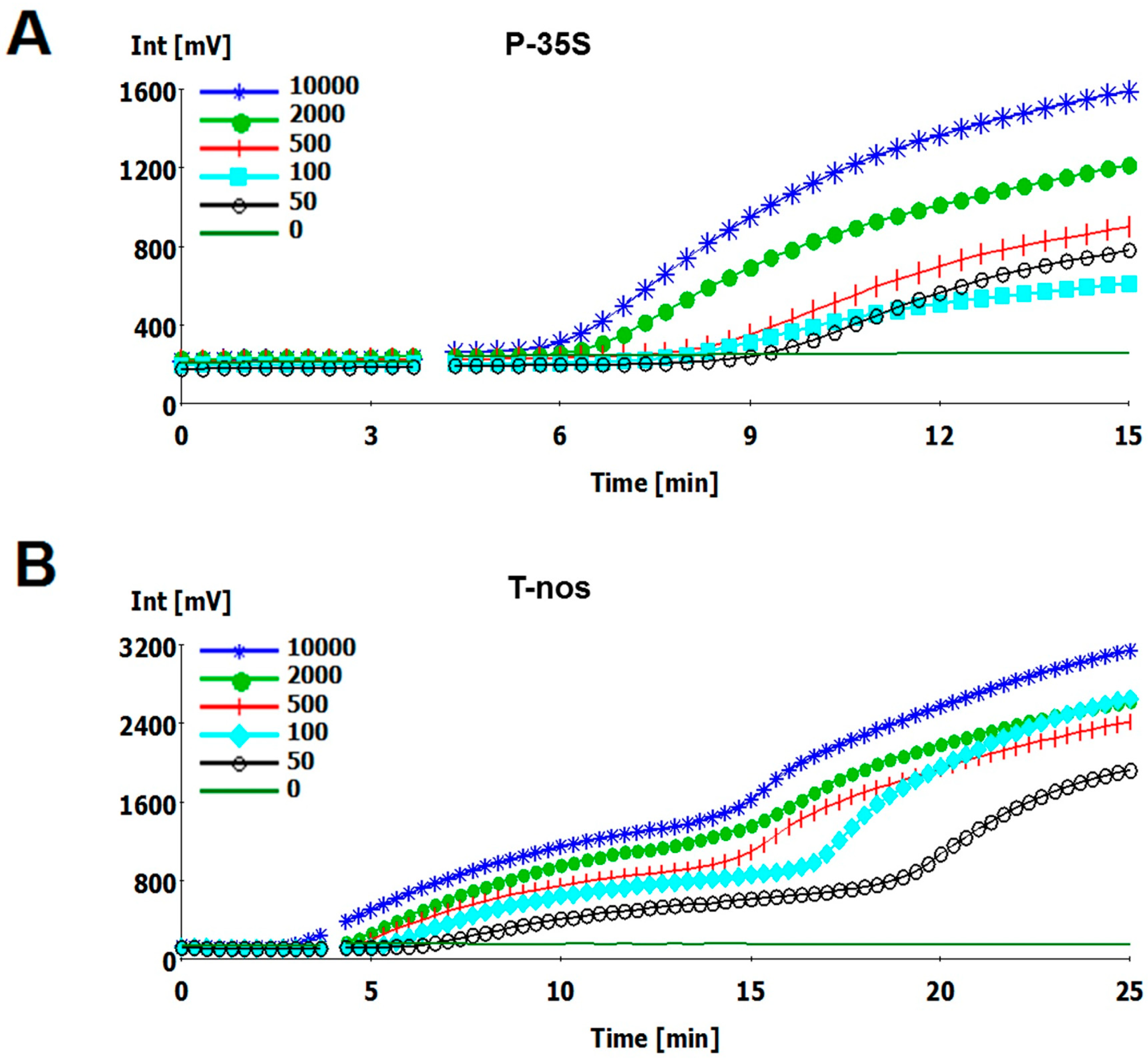
| Template Copy Number | P-35S | T-nos | ||
|---|---|---|---|---|
| Threshold Time Values | Positive Reactions/Total Reactions | Threshold Time Values | Positive Reactions/Total Reactions | |
| 10,000 | 6.1 | 6/6 | 5.5 | 6/6 |
| 2000 | 6.4 | 6/6 | 6.7 | 6/6 |
| 500 | 7 | 6/6 | 7.3 | 6/6 |
| 100 | 8 | 6/6 | 7.8 | 6/6 |
| 50 | ND | 5/6 | ND | 4/6 |
2.3. Application to Practical Sample Analysis
| Species | Event | Regulatory Elements Existing Status | RPA Detection Results | ||
|---|---|---|---|---|---|
| P-35S | T-nos | P-35S | T-nos | ||
| maize | Bt11 | + | + | P | P |
| DAS-59122-7 | + | − | P | N | |
| TC1507 | + | − | P | N | |
| MIR604 | − | + | N | P | |
| Non-GM | − | − | N | N | |
| rice | TT51-1 | − | + | N | P |
| Kefeng 6 | + | + | P | P | |
| Kemingdao 1 | + | + | P | P | |
| Non-GM | − | − | N | N | |
| cotton | MON15985 | + | + | P | P |
| MON531 | + | + | P | P | |
| Non-GM | − | − | N | N | |
| soybean | A5547-127 | + | − | P | N |
| Non-GM | − | − | N | N | |
2.4. Discussion
3. Experimental Section
3.1. Materials
3.2. Extraction of Genomic DNA
3.3. Oligonucleotide Primers and Probes
3.4. RPA Assays
4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- James, C. Global Status of Commercialized Biotech/GM Crops: 2013; ISAAA Brief No. 46; ISAAA: Ithaca, NY, USA, 2013. [Google Scholar]
- Marmiroli, N.; Maestri, E.; Gullì, M.; Malcevschi, A.; Peano, C.; Bordoni, R.; de Bellis, G. Methods for detection of GMOs in food and feed. Anal. Bioanal. Chem. 2008, 392, 369–384. [Google Scholar] [CrossRef] [PubMed]
- Holst-Jensen, A.; Rønning, S.B.; Løvseth, A.; Berdal, K.G. PCR technology for screening and quantification of genetically modified organisms (GMOs). Anal. Bioanal. Chem. 2003, 375, 985–993. [Google Scholar] [PubMed]
- Chen, L.; Guo, J.; Wang, Q.; Kai, G.; Yang, L. Development of the visual loop-mediated isothermal amplification assays for seven genetically modified maize events and their application in practical samples analysis. J. Agric. Food Chem. 2011, 59, 5914–5918. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Morrical, S. Assembly and dynamics of the bacteriophage T4 homologous recombination machinery. Virol. J. 2010, 7, 357. [Google Scholar] [CrossRef] [PubMed]
- Piepenburg, O.; Williams, C.H.; Stemple, D.L.; Armes, N.A. DNA detection using recombination proteins. PLoS Biol. 2006, 4, e204. [Google Scholar] [CrossRef]
- Crannell, Z.A.; Castellanos-Gonzalez, A.; Irani, A.; Rohrman, B.; White, A.C.; Richards-Kortum, R. Nucleic acid test to diagnose Cryptosporidiosis: Lab assessment in animal and patient specimens. Anal. Chem. 2014, 86, 2565–2571. [Google Scholar] [CrossRef] [PubMed]
- Euler, M.; Wang, Y.; Otto, P.; Tomaso, H.; Escudero, R.; Anda, P.; Hufert, F.T.; Weidmann, M. Recombinase polymerase amplification assay for rapid detection of Francisella tularensis. J. Clin. Microbiol. 2012, 50, 2234–2238. [Google Scholar] [CrossRef] [PubMed]
- Euler, M.; Wang, Y.; Nentwich, O.; Piepenburg, O.; Hufert, F.T.; Weidmann, M. Recombinase polymerase amplification assay for rapid detection of Rift Valley fever virus. J. Clin. Virol. 2012, 54, 308–312. [Google Scholar] [CrossRef] [PubMed]
- Boyle, D.S.; Lehman, D.A.; Lillis, L.; Peterson, D.; Singhal, M.; Armes, N.; Parker, M.; Piepenburg, O.; Overbaugh, J. Rapid detection of HIV-1 proviral DNA for early infant diagnosis using recombinase polymerase amplification. MBio 2013, 4. [Google Scholar] [CrossRef]
- Reiting, R.; Broll, H.; Waiblinger, H.U.; Grohmann, L. Collaborative study of a T-nos real-time PCR method for screening of genetically modified organisms in food products. J. Consum. Prot. Food Saf. 2007, 2, 116–121. [Google Scholar]
- AGBIOS GM Crop Database. Available online: http://www.cera-gmc.org/GMCropDatabase (accessed on 20 April 2014).
- Cao, Y.; Wu, G.; Wu, Y.; Nie, S.; Zhang, L.; Lu, C. Characterization of the transgenic rice event TT51-1 and construction of a reference plasmid. J. Agric. Food Chem. 2011, 59, 8550–8559. [Google Scholar] [CrossRef] [PubMed]
- Su, C.; Xie, J.; Wang, X.; Peng, Y. Integrated structure and event-specific real-time detection of transgenic cry1Ac/SCK rice Kefeng 6. Eur. Food Res. Technol. 2011, 232, 351–359. [Google Scholar] [CrossRef]
- Babekova, R.; Funk, T.; Pecoraro, S.; Engel, K.H.; Busch, U. Development of an event-specific real-time PCR detection method for the transgenic Bt rice line KMD1. Eur. Food Res. Technol. 2009, 228, 707–716. [Google Scholar] [CrossRef]
- Gill, P.; Ghaemi, A. Nucleic acid isothermal amplification technologies: A review. Nucleosides Nucleotides Nucleic Acids 2008, 27, 224–243. [Google Scholar] [CrossRef] [PubMed]
- Morisset, D.; Dobnik, D.; Hamels, S.; Žel, J.; Gruden, K. NAIMA: Target amplification strategy allowing quantitative on-chip detection of GMOs. Nucleic Acids Res. 2008, 36, e118. [Google Scholar] [CrossRef]
- Fukuta, S.; Mizukami, Y.; Ishida, A.; Ueda, J.; Hasegawa, M.; Hayashi, I.; Hashimoto, M.; Kanbe, M. Real-time loop-mediated isothermal amplification for the CaMV-35S promoter as a screening method for genetically modified organisms. Eur. Food Res. Technol. 2004, 218, 496–500. [Google Scholar] [CrossRef]
- Lee, D.; La Mura, M.; Allnutt, T.; Powell, W. Detection of genetically modified organisms (GMOs) using isothermal amplification of target DNA sequences. BMC Biotechnol. 2009, 9, 7. [Google Scholar] [PubMed]
- Li, Q.; Fang, J.; Liu, X.; Xi, X.; Li, M.; Gong, Y.; Zhang, M. Loop-mediated isothermal amplification (LAMP) method for rapid detection of cry1Ab gene in transgenic rice (Oryza sativa L.). Eur. Food Res. Technol. 2013, 236, 589–598. [Google Scholar] [CrossRef]
- Höhne, M.; Santisi, C.; Meyer, R. Real-time multiplex PCR: An accurate method for the detection and quantification of 35S-CaMV promoter in genetically modified maize-containing food. Eur. Food Res. Technol. 2002, 215, 59–64. [Google Scholar] [CrossRef]
- Piepenburg, O.; Williams, C.H.; Armes, N.A.; Stemple, D.L. Recombinase polymerase amplification. U.S. Patent 7,399,590, 2008. [Google Scholar]
- TwistDx. Appendix to the TwistAmpTM reaction kit manuals. Available online: http://www.twistdx.co.uk/images/uploads/docs/Appendix.pdf (accessed on 14 August 2014).
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Xu, C.; Li, L.; Jin, W.; Wan, Y. Recombinase Polymerase Amplification (RPA) of CaMV-35S Promoter and nos Terminator for Rapid Detection of Genetically Modified Crops. Int. J. Mol. Sci. 2014, 15, 18197-18205. https://doi.org/10.3390/ijms151018197
Xu C, Li L, Jin W, Wan Y. Recombinase Polymerase Amplification (RPA) of CaMV-35S Promoter and nos Terminator for Rapid Detection of Genetically Modified Crops. International Journal of Molecular Sciences. 2014; 15(10):18197-18205. https://doi.org/10.3390/ijms151018197
Chicago/Turabian StyleXu, Chao, Liang Li, Wujun Jin, and Yusong Wan. 2014. "Recombinase Polymerase Amplification (RPA) of CaMV-35S Promoter and nos Terminator for Rapid Detection of Genetically Modified Crops" International Journal of Molecular Sciences 15, no. 10: 18197-18205. https://doi.org/10.3390/ijms151018197
APA StyleXu, C., Li, L., Jin, W., & Wan, Y. (2014). Recombinase Polymerase Amplification (RPA) of CaMV-35S Promoter and nos Terminator for Rapid Detection of Genetically Modified Crops. International Journal of Molecular Sciences, 15(10), 18197-18205. https://doi.org/10.3390/ijms151018197




