Metabolomics as a Tool to Investigate Abiotic Stress Tolerance in Plants
Abstract
:1. Metabolomics within the Context of Systems Biology
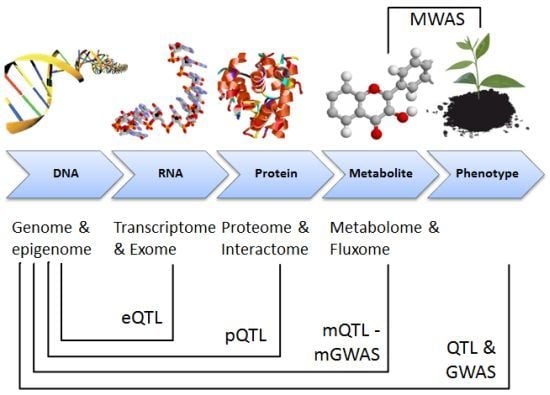
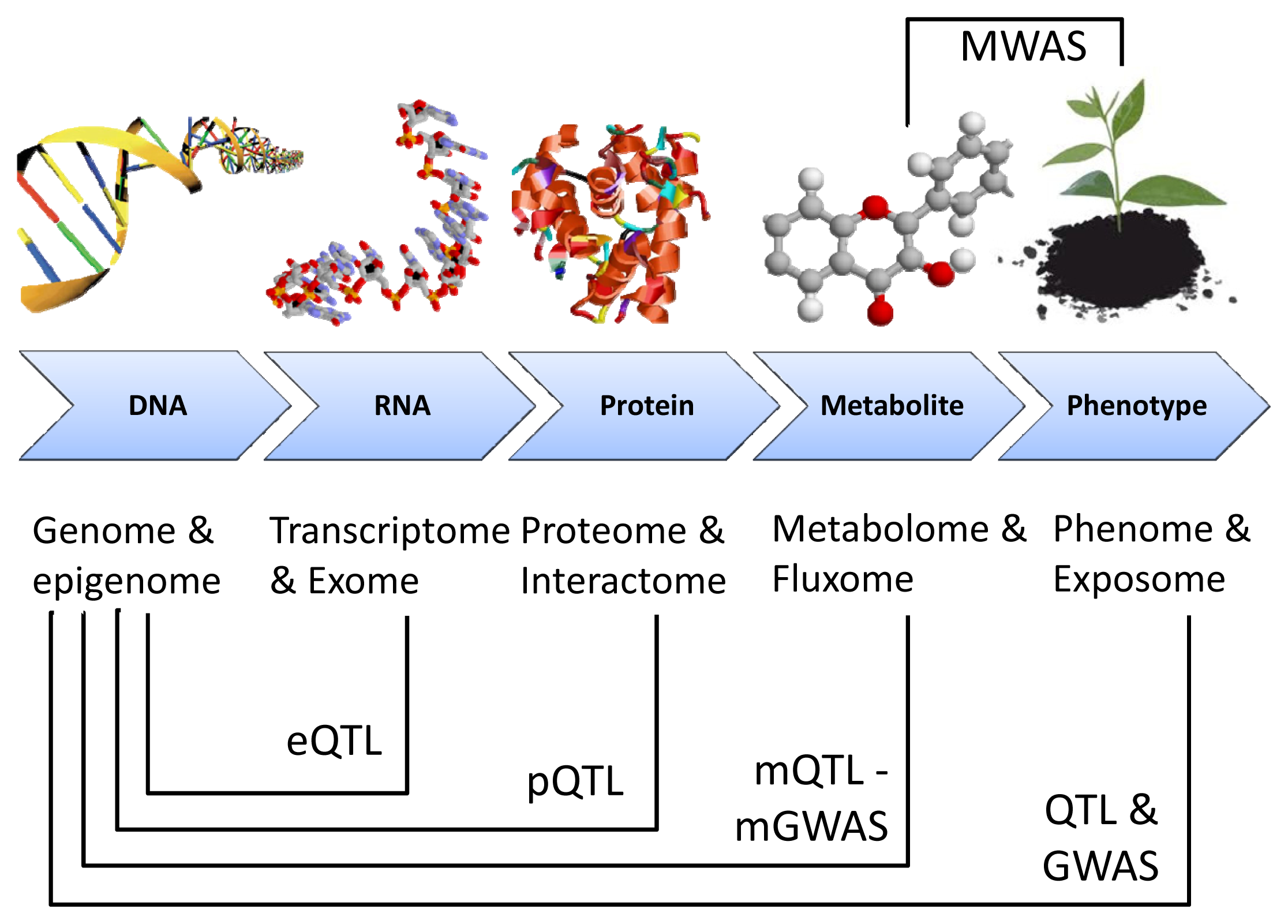
1.1. Omics Technologies: Transcriptomics, Proteomics and Metabolomics
1.2. Data Integration: Gene-Protein-Metabolite
1.2.1. Transcriptomic-Proteomic
1.2.2. Transcriptomic-Metabolomic
1.2.3. Metabolomic-Proteomic
2. Abiotic Stress, Causes and Physiological Responses
2.1. Drought
2.2. Salinity
2.3. Soil Flooding
2.4. Temperature Stress
3. Effect of Abiotic Stress on Plant Biochemistry: Metabolites as Effectors of Tolerance/Damage and Genes Involved
3.1. Primary Metabolism and Osmoprotectants
3.1.1. Carbohydrates
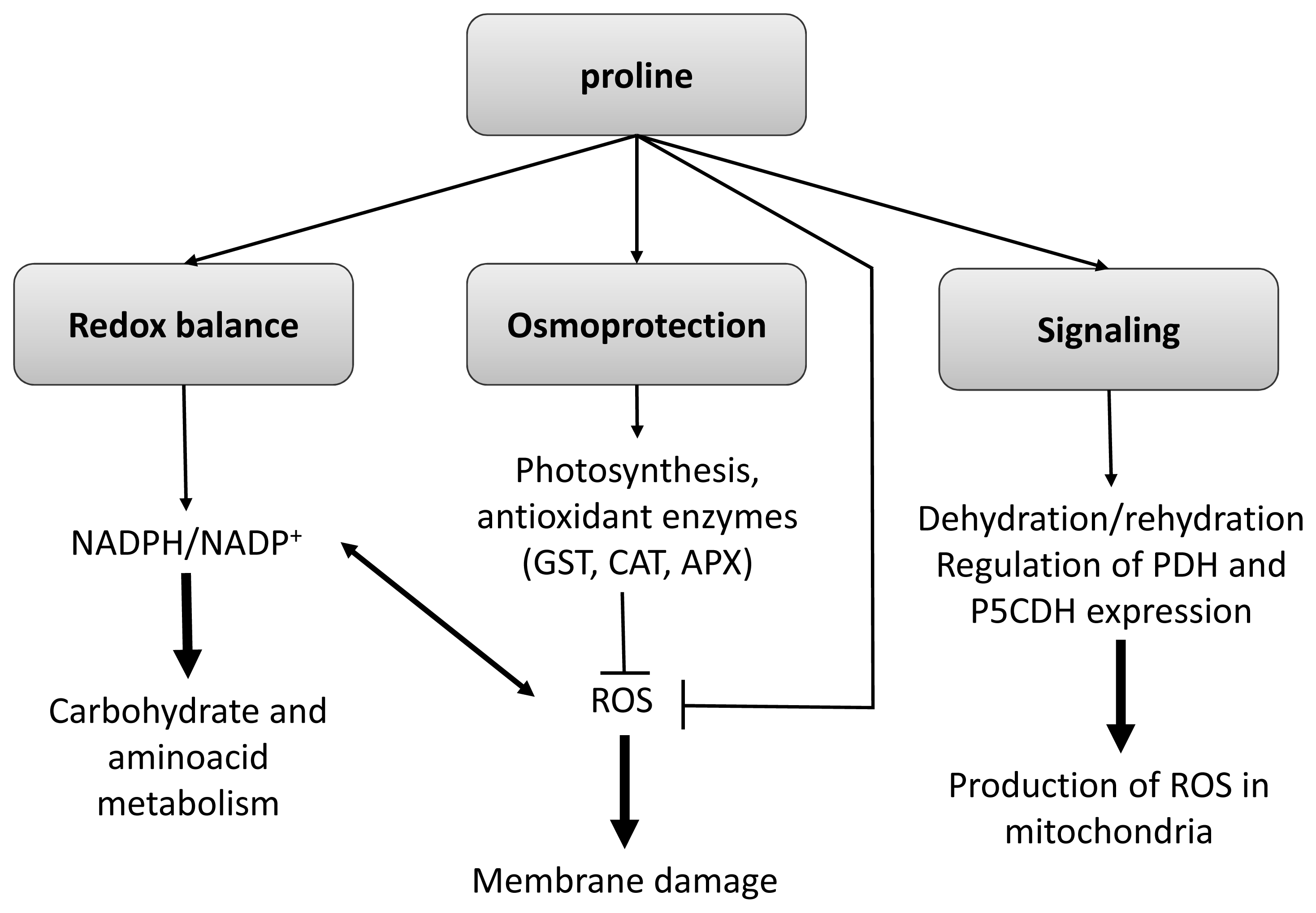
3.1.2. Aminoacids: Proline
3.1.3. Polyamines
3.1.4. Integration of Metabolites as Physiological Effectors
3.2. Secondary Metabolites: Antioxidants, Defense Compounds and Regulatory Metabolites
3.2.1. Phenolic Compounds
3.2.2. Glucosinolates
3.2.3. Carotenoids and Other Terpenoid Derivatives
3.2.4. Secondary Metabolites, Stress Tolerance and Fruit Quality
4. Development of Metabolic QTL for Improving Stress Tolerance
5. Conclusions and Future Prospects
Acknowledgments
Conflict of Interest
References
- Saito, K.; Matsuda, F. Metabolomics for functional genomics, systems biology, and biotechnology. Ann. Rev. Plant Biol 2010, 61, 463–489. [Google Scholar]
- Fukushima, A.; Kusano, M.; Redestig, H.; Arita, M.; Saito, K. Integrated omics approaches in plant systems biology. Curr. Opin. Chem. Biol 2009, 13, 532–538. [Google Scholar]
- Urano, K.; Kurihara, Y.; Seki, M.; Shinozaki, K. “Omics” analyses of regulatory networks in plant abiotic stress responses. Curr. Opin. Plant Biol 2010, 13, 132–138. [Google Scholar]
- Ma, Y.; Qin, F.; Tran, L.-S.P. Contribution of genomics to gene discovery in plant abiotic stress responses. Mol. Plant 2012. [Google Scholar] [CrossRef]
- Steibel, J.P.; Poletto, R.; Coussens, P.M.; Rosa, G.J.M. A powerful and flexible linear mixed model framework for the analysis of relative quantification RT-PCR data. Genomics 2009, 94, 146–152. [Google Scholar]
- Brady, S.M.; Orlando, D.A.; Lee, J.-Y.; Wang, J.Y.; Koch, J.; Dinneny, J.R.; Mace, D.; Ohler, U.; Benfey, P.N. A high-resolution root spatiotemporal map reveals dominant expression patterns. Science 2007, 318, 801–806. [Google Scholar]
- Filichkin, S.A.; Priest, H.D.; Givan, S.A.; Shen, R.; Bryant, D.W.; Fox, S.E.; Wong, W.-K.; Mockler, T.C. Genome-wide mapping of alternative splicing in Arabidopsis thaliana. Genome Res 2010, 20, 45–58. [Google Scholar]
- Yendrek, C.R.; Ainsworth, E.A.; Thimmapuram, J. The bench scientist’s guide to statistical analysis of RNA-Seq data. BMC Res. Not. 2012. [Google Scholar] [CrossRef]
- Birnbaum, K.; Shasha, D.E.; Wang, J.Y.; Jung, J.W.; Lambert, G.M.; Galbraith, D.W.; Benfey, P.N. A gene expression map of the Arabidopsis root. Science 2003, 302, 1956–1960. [Google Scholar]
- Iyer-Pascuzzi, A.S.; Jackson, T.; Cui, H.; Petricka, J.J.; Busch, W.; Tsukagoshi, H.; Benfey, P.N. Cell identity regulators link development and stress responses in the Arabidopsis root. Dev. Cell 2011, 21, 770–782. [Google Scholar]
- Schmid, M.; Davison, T.S.; Henz, S.R.; Pape, U.J.; Demar, M.; Vingron, M.; Schölkopf, B.; Weigel, D.; Lohmann, J.U. A gene expression map of Arabidopsis thaliana development. Nat. Genet 2005, 37, 501–506. [Google Scholar]
- Dinneny, J.R.; Long, T.A.; Wang, J.Y.; Jung, J.W.; Mace, D.; Pointer, S.; Barron, C.; Brady, S.M.; Schiefelbein, J.; Benfey, P.N. Cell identity mediates the response of Arabidopsis roots to abiotic stress. Science 2008, 320, 942–945. [Google Scholar]
- Genevestigator database. Available online: https://www.genevestigator.com/gv/index.jsp accessed on 22 Februrary 2013.
- Arabidopsis eFP browser. Available online: http://bar.utoronto.ca/efp/cgi-bin/efpWeb.cgi accessed on 22 February 2013.
- Breakfield, N.W.; Corcoran, D.L.; Petricka, J.J.; Shen, J.; Sae-Seaw, J.; Rubio-Somoza, I.; Weigel, D.; Ohler, U.; Benfey, P.N. High-resolution experimental and computational profiling of tissue-specific known and novel miRNAs in Arabidopsis. Genome Res 2012, 22, 163–176. [Google Scholar]
- Gao, L.; Yan, X.; Li, X.; Guo, G.; Hu, Y.; Ma, W.; Yan, Y. Proteome analysis of wheat leaf under salt stress by two-dimensional difference gel electrophoresis (2D-DIGE). Phytochemistry 2011, 72, 1180–1191. [Google Scholar]
- Baginsky, S.; Kleffmann, T.; von Zychlinski, A.; Gruissem, W. Analysis of shotgun proteomics and RNA profiling data from Arabidopsis thaliana chloroplasts. J. Proteome Res 2005, 4, 637–640. [Google Scholar]
- Beck, M.; Claassen, M.; Aebersold, R. Comprehensive proteomics. Curr. Opin. Biotechnol 2011, 22, 3–8. [Google Scholar]
- Arbona, V.; Iglesias, D.J.; Talón, M.; Gómez-Cadenas, A. Plant phenotype demarcation using nontargeted LC-MS and GC-MS metabolite profiling. J. Agric. Food Chem 2009, 57, 7338–7347. [Google Scholar]
- Fiehn, O. Combining genomics, metabolome analysis, and biochemical modelling to understand metabolic networks. Comp. Funct. Genomics 2001, 2, 155–168. [Google Scholar]
- Kim, H.K.; Verpoorte, R. Sample preparation for plant metabolomics. Phytochem. Anal 2010, 21, 4–13. [Google Scholar]
- Desbrosses, G.; Steinhauser, D.; Kopka, J. Metabolome analysis using GC-MS. 2005, 165–174. [Google Scholar]
- Cevallos-Cevallos, J.M.; Rouseff, R.; Reyes-De-Corcuera, J.I. Untargeted metabolite analysis of healthy and Huanglongbing-infected orange leaves by CE-DAD. Electrophoresis 2009, 30, 1240–1247. [Google Scholar]
- Grata, E.; Boccard, J.; Guillarme, D.; Glauser, G.; Carrupt, P.-A.; Farmer, E.E.; Wolfender, J.-L.; Rudaz, S. UPLC-TOF-MS for plant metabolomics: A sequential approach for wound marker analysis in Arabidopsis thaliana. J. Chromatogr. B 2008, 871, 261–270. [Google Scholar]
- Wiklund, S.; Johansson, E.; Sjöström, L.; Mellerowicz, E.J.; Edlund, U.; Shockcor, J.P.; Gottfries, J.; Moritz, T.; Trygg, J. Visualization of GC/TOF-MS-based metabolomics data for identification of biochemically interesting compounds using OPLS class models. Anal. Chem 2008, 80, 115–122. [Google Scholar]
- Zandalinas, S.I.; Vives-Peris, V.; Gómez-Cadenas, A.; Arbona, V. A fast and precise method to identify indolic glucosinolates and camalexin in plants by combining mass spectrometric and biological information. J. Agric. Food Chem 2012, 60, 8648–8658. [Google Scholar]
- Durgbanshi, A.; Arbona, V.; Pozo, O.; Miersch, O.; Sancho, J.V; Gómez-Cadenas, A. Simultaneous determination of multiple phytohormones in plant extracts by liquid chromatography-electrospray tandem mass spectrometry. J. Agric. Food Chem. 2005, 53, 8437–8442. [Google Scholar]
- Chiwocha, S.D.S.; Cutler, A.J.; Abrams, S.R.; Ambrose, S.J.; Yang, J.; Ross, A.R.S.; Kermode, A.R. The etr1-2 mutation in Arabidopsis thaliana affects the abscisic acid, auxin, cytokinin and gibberellin metabolic pathways during maintenance of seed dormancy, moist-chilling and germination. Plant J. 2005, 42, 35–48. [Google Scholar]
- Segarra, G.; Jáuregui, O.; Casanova, E.; Trillas, I. Simultaneous quantitative LC-ESI-MS/MS analyses of salicylic acid and jasmonic acid in crude extracts of Cucumis sativus under biotic stress. Phytochemistry 2006, 67, 395–401. [Google Scholar]
- Luedemann, A.; Strassburg, K.; Erban, A.; Kopka, J. TagFinder for the quantitative analysis of gas chromatography-mass spectrometry (GC-MS) based metabolite profiling experiments. Bioinformatics 2008, 24, 1–7. [Google Scholar]
- Matsuda, F.; Yonekura-Sakakibara, K.; Niida, R.; Kuromori, T.; Shinozaki, K.; Saito, K. MS/MS spectral tag-based annotation of non-targeted profile of plant secondary metabolites. Plant J 2009, 57, 555–577. [Google Scholar]
- Burton, L.; Ivosev, G.; Tate, S.; Impey, G.; Wingate, J.; Bonner, R. Instrumental and experimental effects in LC-MS-based metabolomics. J. Chromatogr. B 2008, 871, 227–35. [Google Scholar]
- Fiehn, O.; Wohlgemuth, G.; Scholz, M.; Kind, T.; Lee, D.Y.; Lu, Y.; Moon, S.; Nikolau, B. Quality control for plant metabolomics: Reporting MSI-compliant studies. Plant J 2008, 53, 691–704. [Google Scholar]
- Käll, L.; Vitek, O. Computational mass spectrometry-based proteomics. PLoS Comp. Biol 2011, 7, e1002277. [Google Scholar]
- Castellana, N.; Bafna, V. Proteogenomics to discover the full coding content of genomes: A computational perspective. J. Proteomics 2010, 73, 2124–2135. [Google Scholar]
- Hirai, M.Y.; Klein, M.; Fujikawa, Y.; Yano, M.; Goodenowe, D.B.; Yamazaki, Y.; Kanaya, S.; Nakamura, Y.; Kitayama, M.; Suzuki, H.; et al. Elucidation of gene-to-gene and metabolite-to-gene networks in arabidopsis by integration of metabolomics and transcriptomics. J. Biol. Chem 2005, 280, 25590–25595. [Google Scholar]
- Thimm, O.; Bläsing, O.; Gibon, Y.; Nagel, A.; Meyer, S.; Krüger, P.; Selbig, J.; Müller, L.A.; Rhee, S.Y.; Stitt, M. Mapman: A user-driven tool to display genomics data sets onto diagrams of metabolic pathways and other biological processes. Plant J. 2004, 37, 914–939. [Google Scholar]
- Joung, J.-G.; Corbett, A.M.; Fellman, S.M.; Tieman, D.M.; Klee, H.J.; Giovannoni, J.J.; Fei, Z. Plant MetGenMAP: An integrative analysis system for plant systems biology. Plant Physiol 2009, 151, 1758–1768. [Google Scholar]
- Urbanczyk-Wochniak, E.; Usadel, B.; Thimm, O.; Nunes-Nesi, A.; Carrari, F.; Davy, M.; Bläsing, O.; Kowalczyk, M.; Weicht, D.; Polinceusz, A.; et al. Conversion of MapMan to allow the analysis of transcript data from Solanaceous species: Effects of genetic and environmental alterations in energy metabolism in the leaf. Plant Mol. Biol 2006, 60, 773–792. [Google Scholar]
- Joosen, R.V.L.; Ligterink, W.; Dekkers, B.J.W.; Hilhorst, H.W.M. Visualization of molecular processes associated with seed dormancy and germination using MapMan. Seed Sci. Res 2011, 21, 143–152. [Google Scholar]
- Kempa, S.; Krasensky, J.; dal Santo, S.; Kopka, J.; Jonak, C. A central role of abscisic acid in stress-regulated carbohydrate metabolism. PLoS One 2008, 3, e3935. [Google Scholar]
- Luo, Z.-B.; Janz, D.; Jiang, X.; Göbel, C.; Wildhagen, H.; Tan, Y.; Rennenberg, H.; Feussner, I.; Polle, A. Upgrading root physiology for stress tolerance by ectomycorrhizas: Insights from metabolite and transcriptional profiling into reprogramming for stress anticipation. Plant Physiol 2009, 151, 1902–1917. [Google Scholar]
- Weckwerth, W.; Loureiro, M.E.; Wenzel, K.; Fiehn, O. Differential metabolic networks unravel the effects of silent plant phenotypes. Proc. Natl. Acad. Sci. USA 2004, 101, 7809–7814. [Google Scholar]
- Wienkoop, S.; Morgenthal, K.; Wolschin, F.; Scholz, M.; Selbig, J.; Weckwerth, W. Integration of metabolomic and proteomic phenotypes: Analysis of data covariance dissects starch and RFO metabolism from low and high temperature compensation response in Arabidopsis thaliana. Mol. Cell. Proteomics 2008, 1725–1736. [Google Scholar]
- Katari, M.S.; Nowicki, S.D.; Aceituno, F.F.; Nero, D.; Kelfer, J.; Thompson, L.P.; Cabello, J.M.; Davidson, R.S.; Goldberg, A.P.; Shasha, D.E.; et al. A VirtualPlant: A software platform to support systems biology research. Plant Physiol 2010, 152, 500–515. [Google Scholar]
- Mostafavi, S.; Ray, D.; Warde-Farley, D.; Grouios, C.; Morris, Q. GeneMANIA: A real-time multiple association network integration algorithm for predicting gene function. Genome Biol 2008, 9, S4. [Google Scholar]
- Ahuja, I.; de Vos, R.C.H.; Bones, A.M.; Hall, R.D. Plant molecular stress responses face climate change. Trends Plant Sci 2010, 15, 664–674. [Google Scholar]
- Des Marais, D.L.; Juenger, T.E. Pleiotropy, plasticity, and the evolution of plant abiotic stress tolerance. Ann. N.Y. Acad. Sci 2010, 1206, 56–79. [Google Scholar]
- Verslues, P.E.; Agarwal, M.; Katiyar-Agarwal, S.; Zhu, J.; Zhu, J.-K. Methods and concepts in quantifying resistance to drought, salt and freezing, abiotic stresses that affect plant water status. Plant J 2006, 45, 523–539. [Google Scholar]
- Cattivelli, L.; Rizza, F.; Badeck, F.-W.; Mazzucotelli, E.; Mastrangelo, A.M.; Francia, E.; Marè, C.; Tondelli, A.; Stanca, A.M. Drought tolerance improvement in crop plants: An integrated view from breeding to genomics. Field Crops Res 2008, 105, 1–14. [Google Scholar]
- Chaves, M.M.; Flexas, J.; Pinheiro, C. Photosynthesis under drought and salt stress: Regulation mechanisms from whole plant to cell. Ann. Bot 2009, 103, 551–560. [Google Scholar]
- Flexas, J. Drought-inhibition of photosynthesis in C3 plants: Stomatal and non-stomatal limitations revisited. Ann. Bot 2002, 89, 183–189. [Google Scholar]
- Hossain, Z.; López-Climent, M.F.; Arbona, V.; Pérez-Clemente, R.M.; Gómez-Cadenas, A. Modulation of the antioxidant system in Citrus under waterlogging and subsequent drainage. J. Plant Physiol 2009, 166, 1391–1404. [Google Scholar]
- Kirakosyan, A.; Kaufman, P.; Warber, S.; Zick, S.; Aaronson, K.; Bolling, S.; Chul Chang, S. Applied environmental stresses to enhance the levels of polyphenolics in leaves of hawthorn plants. Physiol. Plant 2004, 121, 182–186. [Google Scholar]
- Djoukeng, J.D.; Arbona, V.; Argamasilla, R.; Gomez-cadenas, A. Flavonoid profiling in leaves of citrus genotypes under different environmental situations flavonoid profiling in leaves of Citrus genotypes. J. Agric. Food Chem 2008, 56, 11087–11097. [Google Scholar]
- Urano, K.; Maruyama, K.; Ogata, Y.; Morishita, Y.; Takeda, M.; Sakurai, N.; Suzuki, H.; Saito, K.; Shibata, D.; Kobayashi, M.; et al. Characterization of the ABA-regulated global responses to dehydration in Arabidopsis by metabolomics. Plant J 2009, 57, 1065–1078. [Google Scholar]
- Munns, R.; Tester, M. Mechanisms of salinity tolerance. Ann. Rev. Plant Biol 2008, 59, 651–81. [Google Scholar]
- Breckle, S.W. Salinity, Halophytes and Salt-Affected Natural Ecosystems. In Salinity: Environment-Plant-Molecules; Läuchli, A., Lüttge, U., Eds.; Kluwer Academic Publishers: Dordrecht, NL, USA, 2002; pp. 53–77. [Google Scholar]
- Visser, E.J.W.; Voesenek, L.A.C.J. Acclimation to soil flooding-sensing and signal-transduction. Plant Soil 2005, 274, 197–214. [Google Scholar]
- Bailey-Serres, J.; Voesenek, L.A.C.J. Flooding stress: Acclimations and genetic diversity. Ann. Rev. Plant Biol 2008, 59, 313–339. [Google Scholar]
- Arbona, V.; López-Climent, M.F.; Pérez-Clemente, R.M.; Gómez-Cadenas, A. Maintenance of a high photosynthetic performance is linked to flooding tolerance in citrus. Environ. Exp. Bot 2009, 66, 135–142. [Google Scholar]
- Mittler, R.; Finka, A.; Goloubinoff, P. How do plants feel the heat? Trends Biochem. Sci 2012, 37, 118–125. [Google Scholar]
- Allakhverdiev, S.I.; Kreslavski, V.D.; Klimov, V.V; Los, D.A.; Carpentier, R.; Mohanty, P. Heat stress: An overview of molecular responses in photosynthesis. Photosynth. Res. 2008, 98, 541–550. [Google Scholar]
- Theocharis, A.; Clément, C.; Barka, E.A. Physiological and molecular changes in plants grown at low temperatures. Planta 2012, 235, 1091–1105. [Google Scholar]
- Scholz, M.; Gatzek, S.; Sterling, A.; Fiehn, O.; Selbig, J. Metabolite fingerprinting: Detecting biological features by independent component analysis. Bioinformatics 2004, 20, 2447–2454. [Google Scholar]
- Johnson, H.E.; Broadhurst, D.; Goodacre, R.; Smith, A.R. Metabolic fingerprinting of salt-stressed tomatoes. Phytochemistry 2003, 62, 919–928. [Google Scholar]
- Hall, R.D. Plant metabolomics: From holistic hope, to hype, to hot topic. New Phytol 2006, 169, 453–468. [Google Scholar]
- Hall, R.; Beale, M.; Fiehn, O.; Hardy, N.; Sumner, L.; Bino, R. Plant metabolomics: The missing link in functional genomics strategies. Plant Cell 2002, 14, 1437–1440. [Google Scholar]
- Kim, J.K.; Bamba, T.; Harada, K.; Fukusaki, E.; Kobayashi, A. Time-course metabolic profiling in Arabidopsis thaliana cell cultures after salt stress treatment. J. Exp. Bot 2007, 58, 415–424. [Google Scholar]
- Kerchev, P.I.; Fenton, B.; Foyer, C.H.; Hancock, R.D. Plant responses to insect herbivory: Interactions between photosynthesis, reactive oxygen species and hormonal signalling pathways. Plant Cell Environ 2012, 35, 441–453. [Google Scholar]
- Krasensky, J.; Jonak, C. Drought, salt, and temperature stress-induced metabolic rearrangements and regulatory networks. J. Exp. Bot 2012, 63, 1593–1608. [Google Scholar]
- Valerio, C.; Costa, A.; Marri, L.; Issakidis-Bourguet, E.; Pupillo, P.; Trost, P.; Sparla, F. Thioredoxin-regulated beta-amylase (BAM1) triggers diurnal starch degradation in guard cells, and in mesophyll cells under osmotic stress. J. Exp. Bot 2011, 62, 545–555. [Google Scholar]
- Kaplan, F.; Guy, C.L. β-Amylase induction and the protective role of maltose during temperature shock. Plant Physiol 2004, 135, 1674–1684. [Google Scholar]
- Sicher, R.C.; Timlin, D.; Bailey, B. Responses of growth and primary metabolism of water-stressed barley roots to rehydration. J. Plant Physiol 2012, 169, 686–695. [Google Scholar]
- Sharp, R.E.; Poroyko, V.; Hejlek, L.G.; Spollen, W.G.; Springer, G.K.; Bohnert, H.J.; Nguyen, H.T. Root growth maintenance during water deficits: Physiology to functional genomics. J. Exp. Bot 2004, 55, 2343–2351. [Google Scholar]
- Iordachescu, M.; Imai, R. Trehalose biosynthesis in response to abiotic stresses. J. Integr. Plant Biol 2008, 50, 1223–1229. [Google Scholar]
- Nishizawa, A.; Yabuta, Y.; Shigeoka, S. Galactinol and raffinose constitute a novel function to protect plants from oxidative damage. Plant Physiol 2008, 147, 1251–1263. [Google Scholar]
- Xu, G.; Liu, D.; Chen, J.; Ye, X.; Shi, J. Composition of major flavanone glycosides and antioxidant capacity of three citrus varieties. J. Food Biochem 2009, 33, 453–469. [Google Scholar]
- Molinari, H.B.C.; Marur, C.J.; Filho, J.C.B.; Kobayashi, A.K.; Pileggi, M.; Júnior, R.P.L.; Pereira, L.F.P.; Vieira, L.G.E. Osmotic adjustment in transgenic citrus rootstock Carrizo citrange (Citrus sinensis Osb. × Poncirus trifoliata L. Raf.) overproducing proline. Plant Sci 2004, 167, 1375–1381. [Google Scholar]
- Szabados, L.; Savouré, A. Proline: A multifunctional amino acid. Trends Plant Sci 2010, 15, 89–97. [Google Scholar]
- Arbona, V.; Flors, V.; Jacas, J.; García-Agustín, P.; Gómez-Cadenas, A. Enzymatic and non-enzymatic antioxidant responses of Carrizo citrange, a salt-sensitive citrus rootstock, to different levels of salinity. Plant Cell Physiol 2003, 44, 388–394. [Google Scholar]
- Arbona, V.; Hossain, Z.; López-Climent, M.F.; Pérez-Clemente, R.M.; Gómez-Cadenas, A. Antioxidant enzymatic activity is linked to waterlogging stress tolerance in citrus. Physiol. Plant 2008, 132, 452–466. [Google Scholar]
- Yoshiba, Y.; Kiyosue, T.; Katagiri, T.; Ueda, H.; Mizoguchi, T.; Yamaguchi-Shinozaki, K.; Wada, K.; Harada, Y.; Shinozaki, K. Correlation between the induction of a gene for delta-pyrroline-5-carboxylate synthetase and the accumulation of proline in Arabidopsis thaliana under osmotic stress. Plant J 1995, 7, 751–760. [Google Scholar]
- Arbona, V.; Gómez-Cadenas, A. Hormonal modulation of citrus responses to flooding. J. Plant Growth Regul 2008, 27, 241–250. [Google Scholar]
- De Campos, M.K.F.; de Carvalho, K.; de Souza, F.S.; Marur, C.J.; Pereira, L.F.P.; Filho, J.C.B.; Vieira, L.G.E. Drought tolerance and antioxidant enzymatic activity in transgenic “Swingle” citrumelo plants over-accumulating proline. Environ. Exp. Bot 2011, 72, 242–250. [Google Scholar]
- Janská, A.; Marsík, P.; Zelenková, S.; Ovesná, J. Cold stress and acclimation—What is important for metabolic adjustment? Plant Biol 2010, 12, 395–405. [Google Scholar]
- Ghars, M.A.; Parre, E.; Debez, A.; Bordenave, M.; Richard, L.; Leport, L.; Bouchereau, A.; Savouré, A.; Abdelly, C. Comparative salt tolerance analysis between Arabidopsis thaliana and Thellungiella halophila, with special emphasis on K+/Na+ selectivity and proline accumulation. J. Plant Physiol 2008, 165, 588–599. [Google Scholar]
- Lv, W.-T.; Lin, B.; Zhang, M.; Hua, X.-J. Proline accumulation is inhibitory to Arabidopsis seedlings during heat stress. Plant Physiol 2011, 156, 1921–1933. [Google Scholar]
- Groppa, M.D.; Benavides, M.P. Polyamines and abiotic stress: Recent advances. Aminoacids 2008, 34, 35–45. [Google Scholar]
- Alet, A.I.; Sánchez, D.H.; Cuevas, J.C.; Marina, M.; Carrasco, P.; Altabella, T.; Tiburcio, A.F.; Ruiz, O.A. New insights into the role of spermine in Arabidopsis thaliana under long-term salt stress. Plant Sci 2012, 182, 94–100. [Google Scholar]
- Martin-Tanguy, J. Conjugated polyamines and reproductive development: Biochemical, molecular and physiological approaches. Physiol. Plant 1997, 100, 675–688. [Google Scholar]
- Bitrián, M.; Zarza, X.; Altabella, T.; Tiburcio, A.F.; Alcázar, R. Polyamines under abiotic stress: Metabolic crossroads and hormonal crosstalks in plants. Metabolites 2012, 2, 516–528. [Google Scholar]
- Hussain, S.S.; Ali, M.; Ahmad, M.; Siddique, K.H.M. Polyamines: Natural and engineered abiotic and biotic stress tolerance in plants. Biotechnol. Adv 2011, 29, 300–311. [Google Scholar]
- Nayyar, H.; Chander, S. Protective effects of polyamines against oxidative stress induced by water and cold stress in chickpea. J. Agron. Crop Sci 2004, 190, 355–365. [Google Scholar]
- Verma, S.; Mishra, S.N. Putrescine alleviation of growth in salt stressed Brassica juncea by inducing antioxidative defense system. J. Plant Physiol 2005, 162, 669–677. [Google Scholar]
- Alcázar, R.; Planas, J.; Saxena, T.; Zarza, X.; Bortolotti, C.; Cuevas, J.; Bitrián, M.; Tiburcio, A.F.; Altabella, T. Putrescine accumulation confers drought tolerance in transgenic Arabidopsis plants over-expressing the homologous Arginine decarboxylase 2 gene. Plant Physiol. Biochem 2010, 48, 547–552. [Google Scholar]
- Cuevas, J.C.; López-Cobollo, R.; Alcázar, R.; Zarza, X.; Koncz, C.; Altabella, T.; Salinas, J.; Tiburcio, A.F.; Ferrando, A. Putrescine as a signal to modulate the indispensable ABA increase under cold stress. Plant Signal. Behav 2009, 4, 219–220. [Google Scholar]
- Tun, N.N.; Santa-Catarina, C.; Begum, T.; Silveira, V.; Handro, W.; Segal Floh, E.I.; Scherer, G.F.E. Polyamines induce rapid biosynthesis of nitric oxide (NO) in Arabidopsis thaliana seedlings. Plant Cell Physiol 2006, 47, 346–354. [Google Scholar]
- Gill, S.S.; Tuteja, N. Reactive oxygen species and antioxidant machinery in abiotic stress tolerance in crop plants. Plant Physiol. Biochem 2010, 48, 909–930. [Google Scholar]
- Verslues, P.E.; Juenger, T.E. Drought, metabolites, and Arabidopsis natural variation: A promising combination for understanding adaptation to water-limited environments. Curr. Opin. Plant Biol 2011, 14, 240–245. [Google Scholar]
- Sanchez, D.H.; Schwabe, F.; Erban, A.; Udvardi, M.K.; Kopka, J. Comparative metabolomics of drought acclimation in model and forage legumes. Plant Cell Environ 2012, 35, 136–149. [Google Scholar]
- Sanchez, D.H.; Pieckenstain, F.L.; Escaray, F.; Erban, A.; Kraemer, U.; Udvardi, M.K.; Kopka, J. Comparative ionomics and metabolomics in extremophile and glycophytic Lotus species under salt stress challenge the metabolic pre-adaptation hypothesis. Plant Cell Environ 2011, 34, 605–617. [Google Scholar]
- Arbona, V.; Argamasilla, R.; Gómez-Cadenas, A. Common and divergent physiological, hormonal and metabolic responses of Arabidopsis thaliana and Thellungiella halophila to water and salt stress. J. Plant Physiol 2010, 167, 1342–1350. [Google Scholar]
- Wong, C.E.; Li, Y.; Whitty, B.R.; Díaz-Camino, C.; Akhter, S.R.; Brandle, J.E.; Golding, G.B.; Weretilnyk, E.A.; Moffatt, B.A.; Griffith, M. Expressed sequence tags from the Yukon ecotype of Thellungiella reveal that gene expression in response to cold, drought and salinity shows little overlap. Plant Mol. Biol 2005, 58, 561–574. [Google Scholar]
- Gavaghan, C.L.; Li, J.V.; Hadfield, S.T.; Hole, S.; Nicholson, J.K.; Wilson, I.D.; Howe, P.W.A.; Stanley, P.D.; Holmes, E. Application of NMR-based metabolomics to the investigation of salt stress in maize (Zea mays). Phytochem. Anal 2011, 22, 214–224. [Google Scholar]
- Lee, Y.P.; Babakov, A.; de Boer, B.; Zuther, E.; Hincha, D.K. Comparison of freezing tolerance, compatible solutes and polyamines in geographically diverse collections of Thellungiella sp. and Arabidopsis thaliana accessions. BMC Plant Biol 2012, 12, 131. [Google Scholar]
- Colinet, H.; Larvor, V.; Laparie, M.; Renault, D. Exploring the plastic response to cold acclimation through metabolomics. Funct. Ecol 2012, 26, 711–722. [Google Scholar]
- Korn, M.; Gärtner, T.; Erban, A.; Kopka, J.; Selbig, J.; Hincha, D.K. Predicting Arabidopsis freezing tolerance and heterosis in freezing tolerance from metabolite composition. Mol. Plant 2010, 3, 224–235. [Google Scholar]
- Guy, C.; Kaplan, F.; Kopka, J.; Selbig, J.; Hincha, D.K. Metabolomics of temperature stress. Physiol. Plant 2008, 132, 220–235. [Google Scholar]
- Witt, S.; Galicia, L.; Lisec, J.; Cairns, J.; Tiessen, A.; Araus, J.L.; Palacios-Rojas, N.; Fernie, A.R. Metabolic and phenotypic responses of greenhouse-grown maize hybrids to experimentally controlled drought stress. Mol. Plant 2012, 5, 401–417. [Google Scholar]
- Widodo Patterson, J.H.; Newbigin, E.; Tester, M.; Bacic, A.; Roessner, U. Metabolic responses to salt stress of barley (Hordeum vulgare L.) cultivars, Sahara and Clipper, which differ in salinity tolerance. J. Exp. Bot. 2009, 60, 4089–4103. [Google Scholar]
- Moya, J.L.; Gómez-Cadenas, A.; Primo-Millo, E.; Talón, M. Chloride absorption in salt-sensitive Carrizo citrange and salt-tolerant Cleopatra mandarin citrus rootstocks is linked to water use. J. Exp. Bot 2003, 54, 825–833. [Google Scholar]
- López-Climent, M.F.; Arbona, V.; Pérez-Clemente, R.M.; Gómez-Cadenas, A. Effects of cadmium on gas exchange and phytohormone contents in citrus. Biol. Plant 2011, 55, 187–190. [Google Scholar]
- Callahan, D.L.; Roessner, U.; Dumontet, V.; de Livera, A.M.; Doronila, A.; Baker, A.J.M.; Kolev, S.D. Elemental and metabolite profiling of nickel hyperaccumulators from New Caledonia. Phytochemistry 2012, 81, 80–89. [Google Scholar]
- Forner-Giner, M.A.; Llosá, M.J.; Carrasco, J.L.; Perez-Amador, M.A.; Navarro, L.; Ancillo, G. Differential gene expression analysis provides new insights into the molecular basis of iron deficiency stress response in the citrus rootstock Poncirus trifoliata (L.) Raf. J. Exp. Bot 2010, 61, 483–490. [Google Scholar]
- Kabir, A.H.; Paltridge, N.G.; Roessner, U.; Stangoulis, J.C.R. Mechanisms associated with Fe-deficiency tolerance and signaling in shoots of Pisum sativum. Physiol. Plant 2012, 147, 381–395. [Google Scholar]
- Kliebenstein, D.J. Secondary metabolites and plant/environment interactions: A view through Arabidopsis thaliana tinged glasses. Plant Cell Environ 2004, 27, 675–684. [Google Scholar]
- Sánchez-Rodríguez, E.; Ruiz, J.M.; Ferreres, F.; Moreno, D.A. Phenolic profiles of cherry tomatoes as influenced by hydric stress and rootstock technique. Food Chem 2012, 134, 775–782. [Google Scholar]
- Karowe, D.N.; Grubb, C. Elevated CO2 increases constitutive phenolics and trichomes, but decreases inducibility of phenolics in Brassica rapa (Brassicaceae). J. Chem. Ecol 2011, 37, 1332–1340. [Google Scholar]
- Wahid, A.; Gelani, S.; Ashraf, M.; Foolad, M. Heat tolerance in plants: An overview. Environ. Exp. Bot 2007, 61, 199–223. [Google Scholar]
- Schenke, D.; Böttcher, C.; Scheel, D. Crosstalk between abiotic ultraviolet-B stress and biotic (flg22) stress signalling in Arabidopsis prevents flavonol accumulation in favor of pathogen defence compound production. Plant Cell Environ 2011, 34, 1849–1864. [Google Scholar]
- Demkura, P.V.; Ballaré, C.L. UVR8 mediates UV-B-induced Arabidopsis defense responses against Botrytis cinerea by controlling sinapate accumulation. Mol. Plant 2012, 5, 116–126. [Google Scholar]
- Gallego-Giraldo, L.; Jikumaru, Y.; Kamiya, Y.; Tang, Y.; Dixon, R.A. Selective lignin downregulation leads to constitutive defense response expression in alfalfa (Medicago sativa L.). New Phytol 2011, 190, 627–639. [Google Scholar]
- Moura, J.C.M.S.; Bonine, C.A.V.; de Oliveira Fernandes Viana, J.; Dornelas, M.C.; Mazzafera, P. Abiotic and biotic stresses and changes in the lignin content and composition in plants. J. Integr. Plant Biol 2010, 52, 360–376. [Google Scholar]
- D’Auria, J.C.; Gershenzon, J. The secondary metabolism of Arabidopsis thaliana: Growing like a weed. Curr. Opin. Plant Biol 2005, 8, 308–316. [Google Scholar]
- Van Poecke, R.M.P.; Posthumus, M.A.; Dicke, M. Herbivore-induced volatile production by Arabidopsis thaliana leads to attraction of the parasitoid Cotesia rubecula: Chemical, behavioral, and gene-expression analysis. J. Chem. Ecol 2001, 27, 1911–1928. [Google Scholar]
- Zhao, N.; Guan, J.; Ferrer, J.-L.; Engle, N.; Chern, M.; Ronald, P.; Tschaplinski, T.J.; Chen, F. Biosynthesis and emission of insect-induced methyl salicylate and methyl benzoate from rice. Plant Physiol. Biochem 2010, 48, 279–287. [Google Scholar]
- Roetschi, A.; Si-Ammour, A.; Belbahri, L.; Mauch, F.; Mauch-Mani, B. Characterization of an Arabidopsis-Phytophthora pathosystem: Resistance requires a functional PAD2 gene and is independent of salicylic acid, ethylene and jasmonic acid signalling. Plant J 2001, 28, 293–305. [Google Scholar]
- Kang, Y.; Udvardi, M. Global regulation of reactive oxygen species scavenging genes in alfalfa root and shoot under gradual drought stress and recovery. Plant Signal. Behav 2012, 7, 539–543. [Google Scholar]
- Hara, M.; Harazaki, A.; Tabata, K. Administration of isothiocyanates enhances heat tolerance in Arabidopsis thaliana. Plant Growth Regul 2012, 69, 71–77. [Google Scholar]
- Noriega, G.; Cruz, D.S.; Batlle, A.; Tomaro, M.; Balestrasse, K. Heme oxygenase is involved in the protection exerted by jasmonic acid against cadmium stress in soybean roots. J. Plant Growth Regul 2012, 31, 79–89. [Google Scholar]
- Shi, Q.; Bao, Z.; Zhu, Z.; Ying, Q.; Qian, Q. Effects of different treatments of salicylic acid on heat tolerance, chlorophyll fluorescence, and antioxidant enzyme activity in seedlings of Cucumis sativa L. Plant Growth Regul 2006, 48, 127–135. [Google Scholar]
- Grubb, C.D.; Abel, S. Glucosinolate metabolism and its control. Trends Plant Sci 2006, 11, 89–100. [Google Scholar]
- Mewis, I.; Khan, M.A.M.; Glawischnig, E.; Schreiner, M.; Ulrichs, C. Water stress and aphid feeding differentially influence metabolite composition in Arabidopsis thaliana (L.). PLoS One 2012, 7, e48661. [Google Scholar]
- Matsumoto, H.; Ikoma, Y.; Kato, M.; Kuniga, T.; Nakajima, N.; Yoshida, T. Quantification of carotenoids in citrus fruit by LC-MS and comparison of patterns of seasonal changes for carotenoids among citrus varieties. J. Agric. Food Chem 2007, 55, 2356–2368. [Google Scholar]
- Cidade, L.C.; de Oliveira, T.M.; Mendes, A.F.S.; Macedo, A.F.; Floh, E.I.S.; Gesteira, A.S.; Soares-Filho, W.S.; Costa, M.G.C. Ectopic expression of a fruit phytoene synthase from Citrus paradisi Macf. promotes abiotic stress tolerance in transgenic tobacco. Mol. Biol. Rep 2012, 39, 10201–10209. [Google Scholar]
- Dhuique-Mayer, C.; Fanciullino, A.-L.; Dubois, C.; Ollitrault, P. Effect of genotype and environment on citrus juice carotenoid content. J. Agric. Food Chem 2009, 57, 9160–9168. [Google Scholar]
- Manners, G.D. Citrus limonoids: Analysis, bioactivity, and biomedical prospects. J. Agric. Food Chem 2007, 55, 8285–8294. [Google Scholar]
- Rasmussen, S.; Parsons, A.J.; Jones, C.S. Metabolomics of forage plants: A review. Ann. Bot 2012, 110, 1281–1290. [Google Scholar]
- Lugan, R.; Niogret, M.-F.; Kervazo, L.; Larher, F.R.; Kopka, J.; Bouchereau, A. Metabolome and water status phenotyping of Arabidopsis under abiotic stress cues reveals new insight into ESK1 function. Plant Cell Environ 2009, 32, 95–108. [Google Scholar]
- Wentzell, A.M.; Rowe, H.C.; Hansen, B.G.; Ticconi, C.; Halkier, B.A.; Kliebenstein, D.J. Linking metabolic QTLs with network and cis-eQTLs controlling biosynthetic pathways. PLoS Genet 2007, 3, 1687–1701. [Google Scholar]
- Qiu, F.; Zheng, Y.; Zhang, Z.; Xu, S. Mapping of QTL associated with waterlogging tolerance during the seedling stage in maize. Ann. Bot 2007, 99, 1067–1081. [Google Scholar]
- Messmer, R.; Fracheboud, Y.; Bänziger, M.; Vargas, M.; Stamp, P.; Ribaut, J.-M. Drought stress and tropical maize: QTL-by-environment interactions and stability of QTLs across environments for yield components and secondary traits. Theor. Appl. Genet 2009, 119, 913–930. [Google Scholar]
- Neeraja, C.N.; Maghirang-Rodriguez, R.; Pamplona, A.; Heuer, S.; Collard, B.C.Y.; Septiningsih, E.M.; Vergara, G.; Sanchez, D.; Xu, K.; Ismail, A.M.; et al. A marker-assisted backcross approach for developing submergence-tolerant rice cultivars. Theor. Appl. Genet 2007, 115, 767–776. [Google Scholar]
- Frei, M.; Tanaka, J.P.; Chen, C.P.; Wissuwa, M. Mechanisms of ozone tolerance in rice: Characterization of two QTLs affecting leaf bronzing by gene expression profiling and biochemical analyses. J. Exp. Bot 2010, 61, 1405–1417. [Google Scholar]
- Fernie, A.R.; Schauer, N. Metabolomics-assisted breeding: A viable option for crop improvement? Trends Genet 2009, 25, 39–48. [Google Scholar]
- Dumas, M.-E. Metabolome 2.0: Quantitative genetics and network biology of metabolic phenotypes. Mol. Biosyst 2012, 8, 2494–2502. [Google Scholar]
- Steinfath, M.; Strehmel, N.; Peters, R.; Schauer, N.; Groth, D.; Hummel, J.; Steup, M.; Selbig, J.; Kopka, J.; Geigenberger, P.; et al. Discovering plant metabolic biomarkers for phenotype prediction using an untargeted approach. Plant Biotechnol. J 2010, 8, 900–911. [Google Scholar]
- Schauer, N.; Semel, Y.; Roessner, U.; Gur, A.; Balbo, I.; Carrari, F.; Pleban, T.; Perez-Melis, A.; Bruedigam, C.; Kopka, J.; et al. Comprehensive metabolic profiling and phenotyping of interspecific introgression lines for tomato improvement. Nat. Biotechnol 2006, 24, 447–454. [Google Scholar]
- Meyer, R.C.; Steinfath, M.; Lisec, J.; Becher, M.; Witucka-Wall, H.; Törjék, O.; Fiehn, O.; Eckardt, A.; Willmitzer, L.; Selbig, J.; et al. The metabolic signature related to high plant growth rate in Arabidopsis thaliana. Proc. Natl. Acad. Sci. USA 2007, 104, 4759–4764. [Google Scholar]
- Lisec, J.; Steinfath, M.; Meyer, R.C.; Selbig, J.; Melchinger, A.E.; Willmitzer, L.; Altmann, T. Identification of heterotic metabolite QTL in Arabidopsis thaliana RIL and IL populations. Plant J 2009, 59, 777–788. [Google Scholar]
- Lisec, J.; Römisch-Margl, L.; Nikoloski, Z.; Piepho, H.-P.; Giavalisco, P.; Selbig, J.; Gierl, A.; Willmitzer, L. Corn hybrids display lower metabolite variability and complex metabolite inheritance patterns. Plant J 2011, 68, 326–336. [Google Scholar]
- Hannah, M.A.; Wiese, D.; Freund, S.; Fiehn, O.; Heyer, A.G.; Hincha, D.K. Natural genetic variation of freezing tolerance in Arabidopsis. Plant Physiol 2006, 142, 98–112. [Google Scholar]
- The Tomato Genomic Consortium. The tomato genome sequence provides insights into fleshy fruit evolution. Nature 2012, 485, 635–641.[Green Version]
- Xu, Q.; Chen, L.-L.; Ruan, X.; Chen, D.; Zhu, A.; Chen, C.; Bertrand, D.; Jiao, W.-B.; Hao, B.-H.; Lyon, M.P.; et al. The draft genome of sweet orange (Citrus sinensis). Nat. Genet 2013, 45, 59–66. [Google Scholar]
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Arbona, V.; Manzi, M.; Ollas, C.D.; Gómez-Cadenas, A. Metabolomics as a Tool to Investigate Abiotic Stress Tolerance in Plants. Int. J. Mol. Sci. 2013, 14, 4885-4911. https://doi.org/10.3390/ijms14034885
Arbona V, Manzi M, Ollas CD, Gómez-Cadenas A. Metabolomics as a Tool to Investigate Abiotic Stress Tolerance in Plants. International Journal of Molecular Sciences. 2013; 14(3):4885-4911. https://doi.org/10.3390/ijms14034885
Chicago/Turabian StyleArbona, Vicent, Matías Manzi, Carlos De Ollas, and Aurelio Gómez-Cadenas. 2013. "Metabolomics as a Tool to Investigate Abiotic Stress Tolerance in Plants" International Journal of Molecular Sciences 14, no. 3: 4885-4911. https://doi.org/10.3390/ijms14034885
APA StyleArbona, V., Manzi, M., Ollas, C. D., & Gómez-Cadenas, A. (2013). Metabolomics as a Tool to Investigate Abiotic Stress Tolerance in Plants. International Journal of Molecular Sciences, 14(3), 4885-4911. https://doi.org/10.3390/ijms14034885