Arabidopsis RIBA Proteins: Two out of Three Isoforms Have Lost Their Bifunctional Activity in Riboflavin Biosynthesis
Abstract
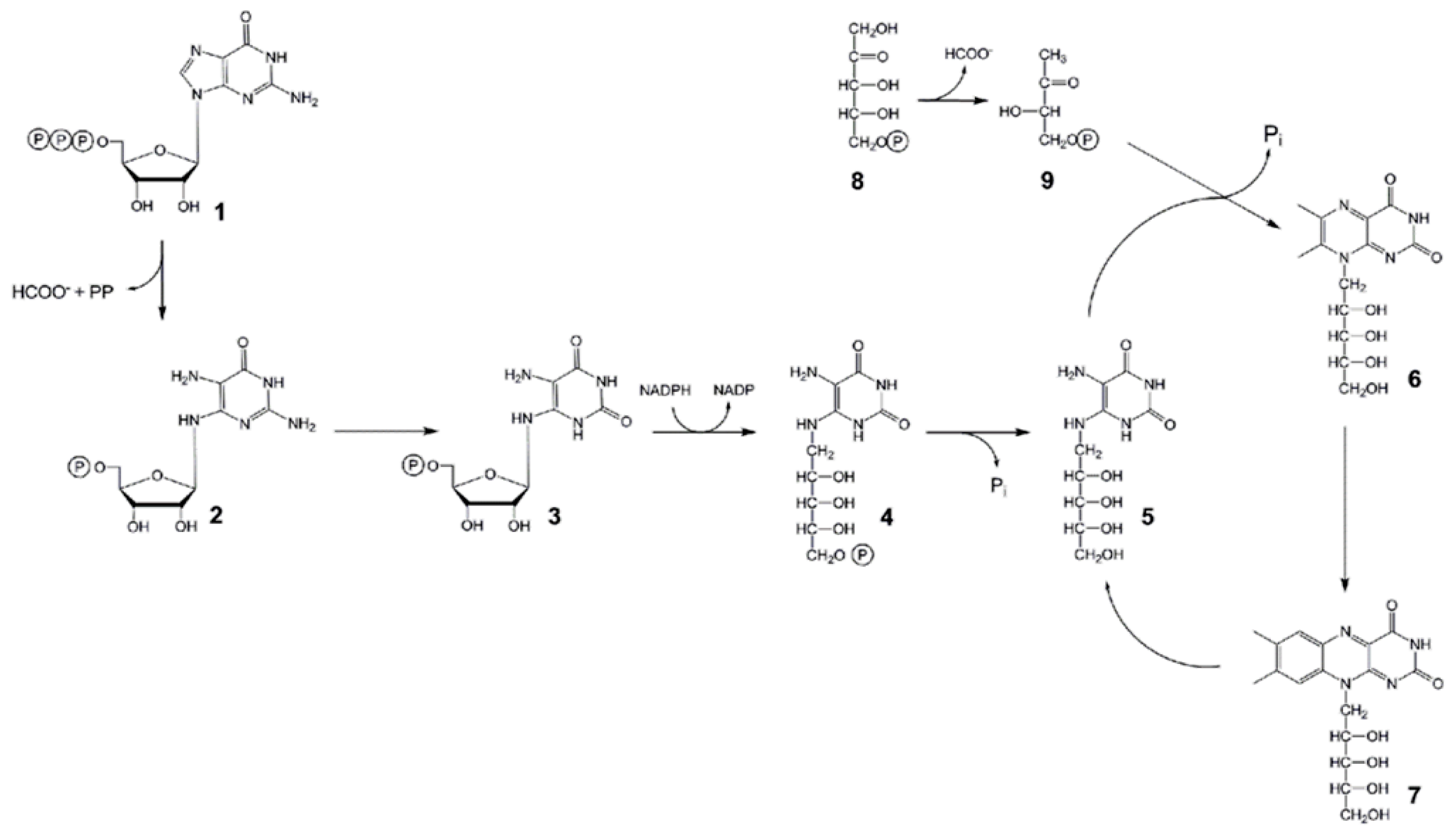
:1. Introduction
2. Results
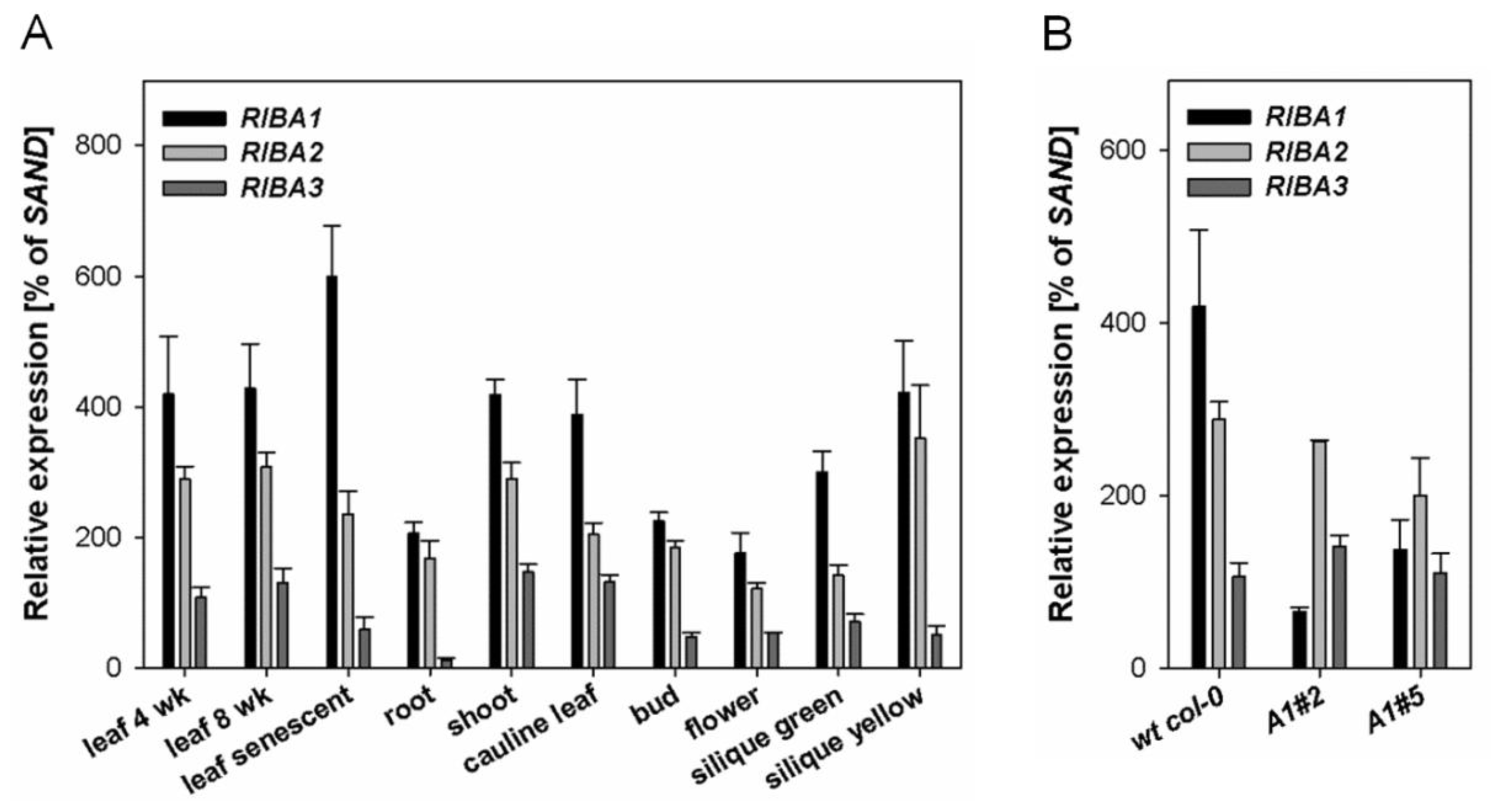
2.1. Expression Patterns of AtRIBA Genes
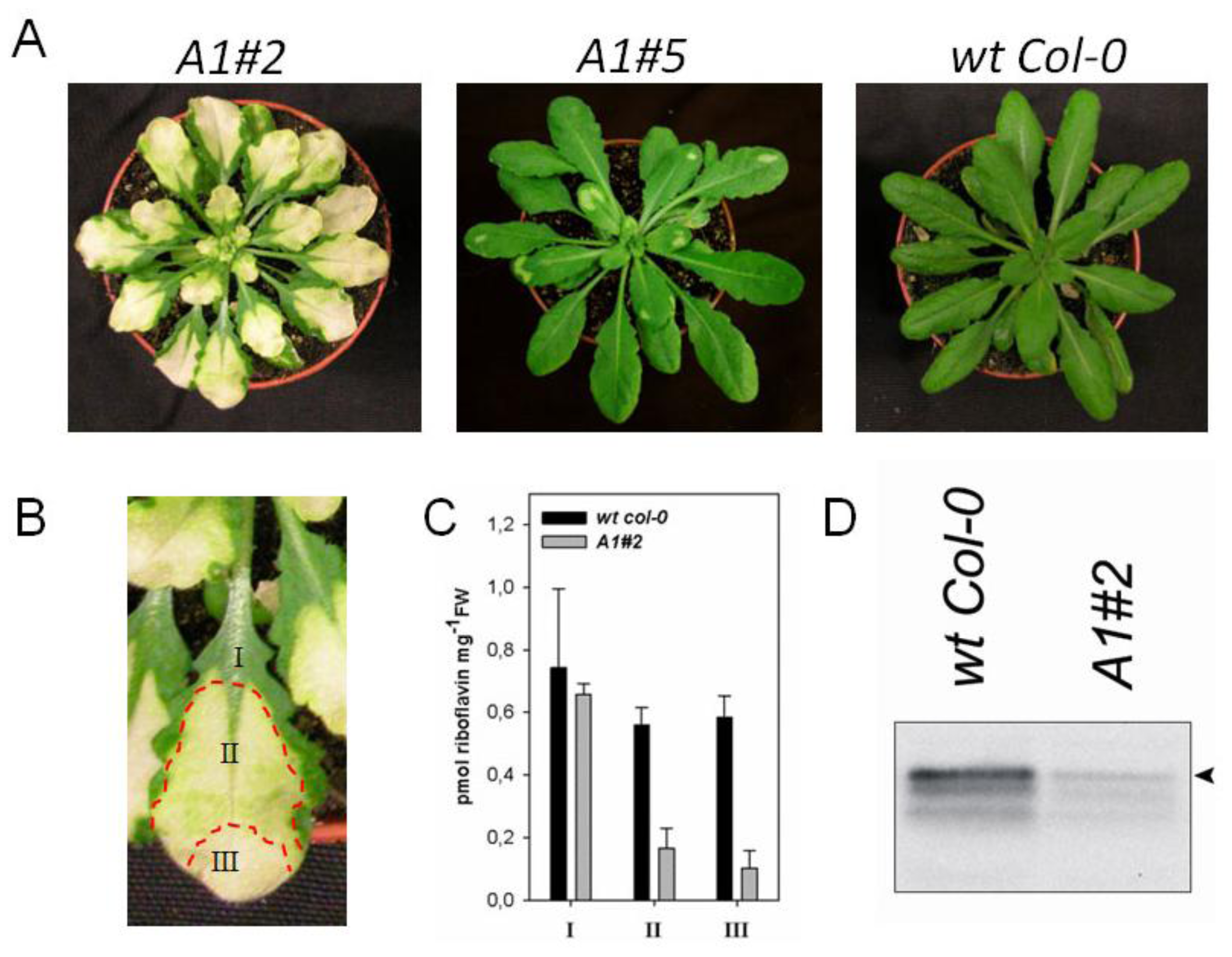
2.2. Downregulation of AtRIBA1 Causes a Bleached Phenotype
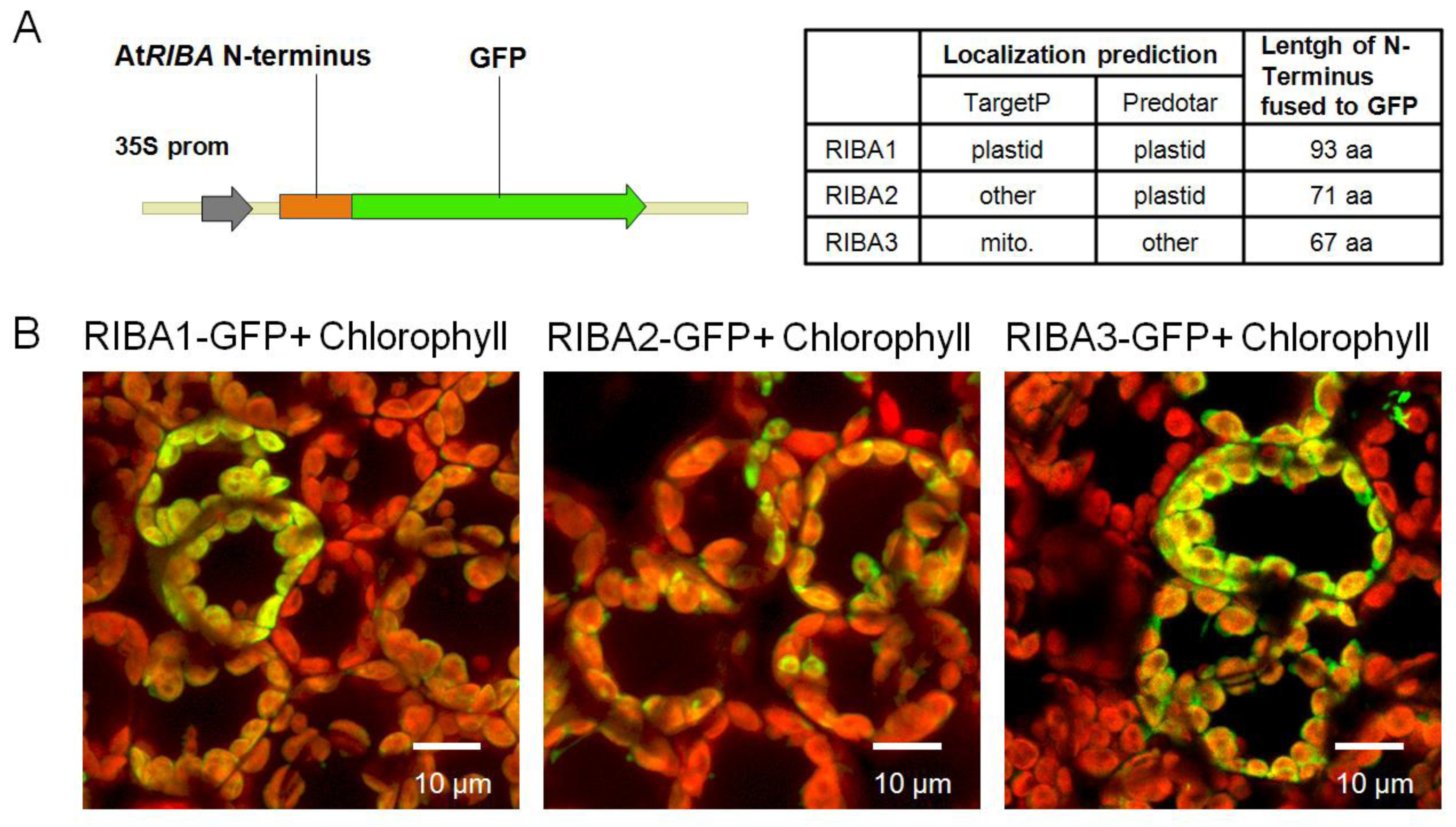
2.3. Subcellular Localization of AtRIBA Proteins
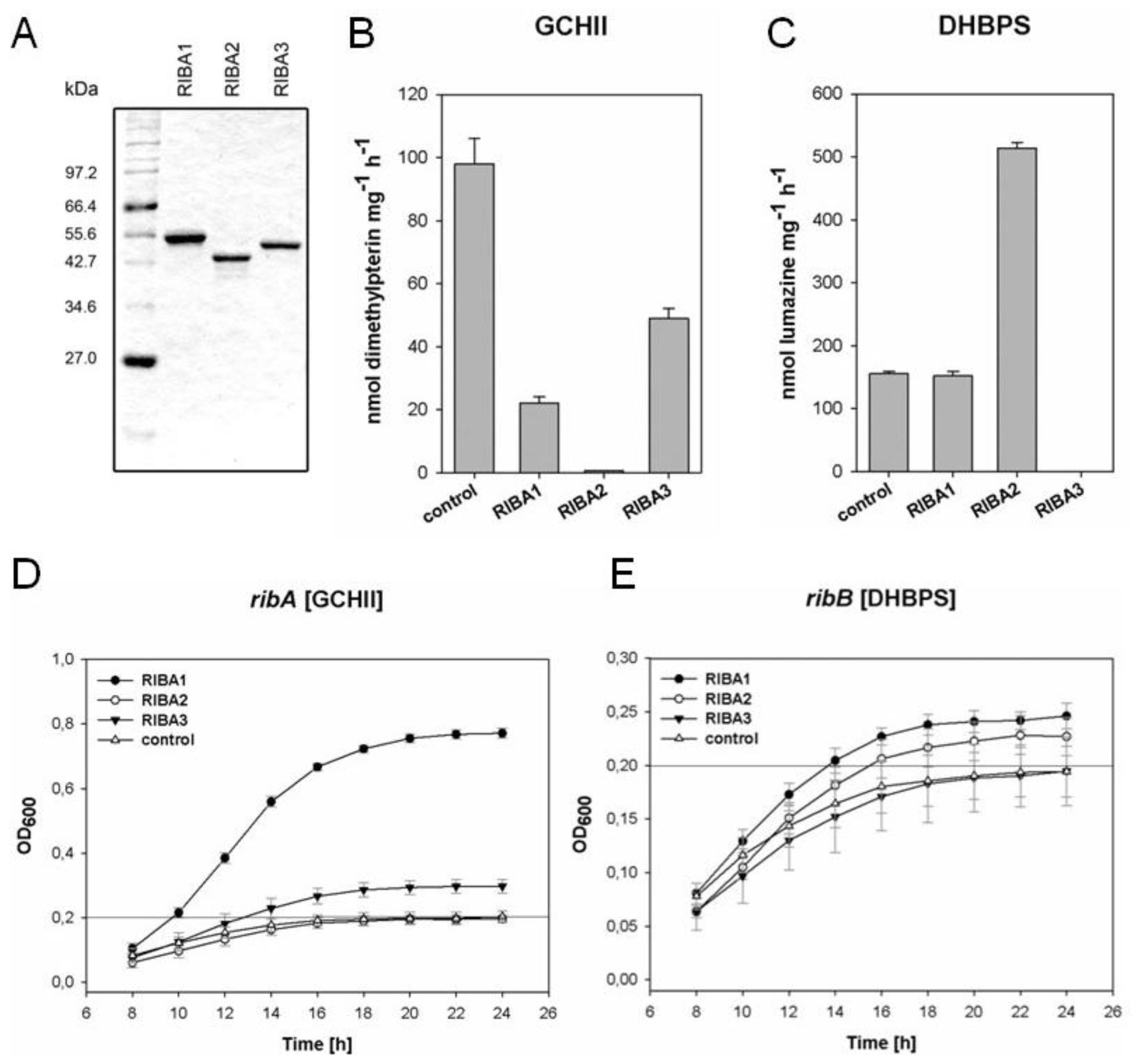
2.4. In vitro Enzyme Assays with Recombinant AtRIBA Proteins
2.5. Complementation of Bacterial Mutants
3. Discussion
3.1. Consequences of AtRIBA1 Deficiency in Arabidopsis
3.2. Analyses of Enzymatic Activities of AtRIBA Isoforms
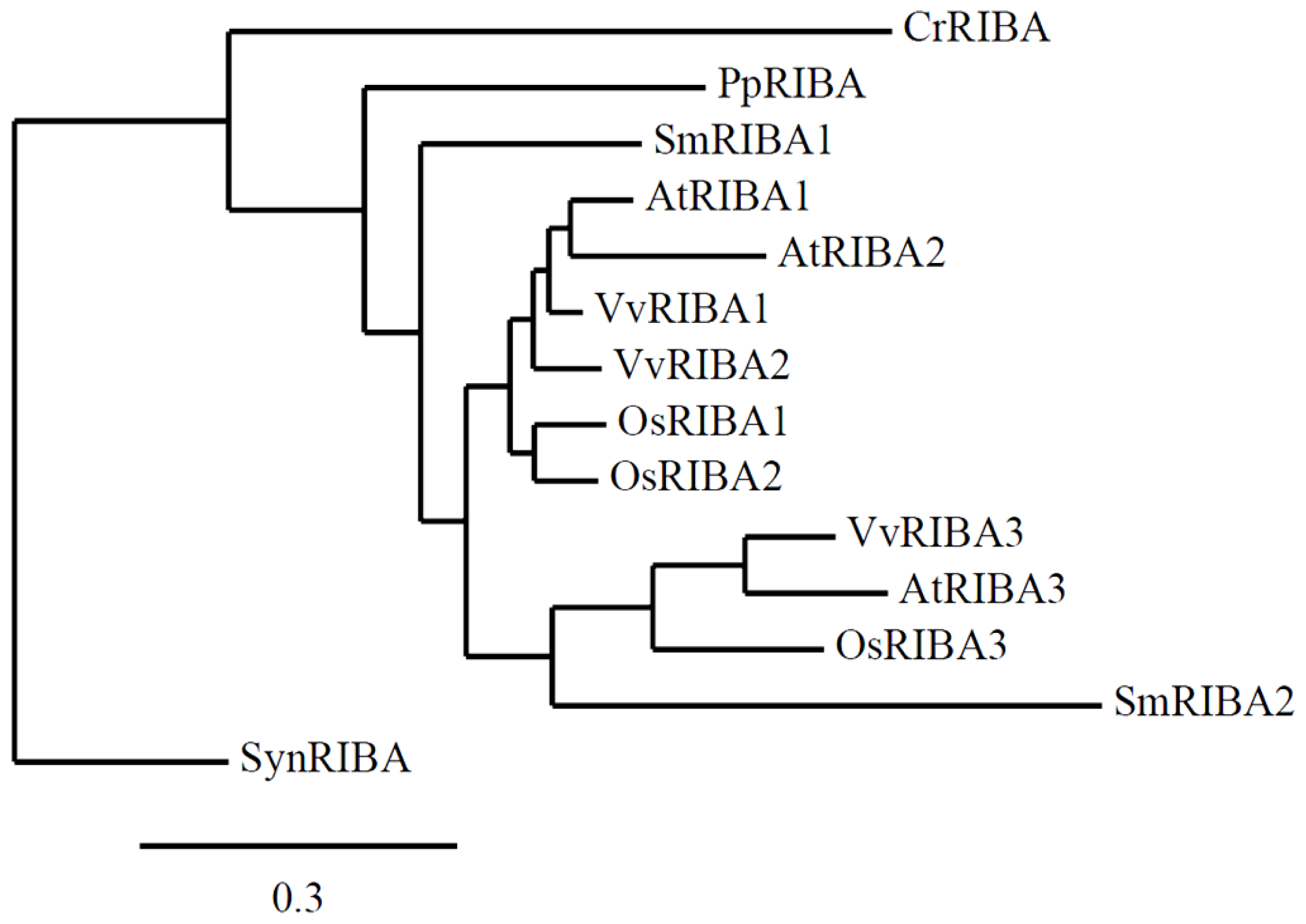
3.3. RIBA Genes Lacking DHBPS Activity Evolved Early in Vascular Plant Phylogeny
4. Experimental Section
4.1. Generation of Antisense Lines
4.2. Heterologous Overexpression
4.3. Assays of GCHII and DHBPS Activity
4.4. Complementation Assays
4.5. RNA Isolation and Quantification
4.6. HPLC Analysis
4.7. Subcellular Localization of RIBA-GFP Fusions
4.8. Protein Analysis
5. Conclusions
Supplementary Information
Figure S1
Figure S2
Data S1
| Nr. | Designation | Sequence 5′→3′ |
|---|---|---|
| 1 | RibA2_PsiI_fw | AATTATAACAGTCGACGGGCCCG |
| 2 | RibA2_PsiI_rev | GCTTATAATACCTCGCGAATGCATCT |
| 3 | RibA1_GFP_fw | ACCCGGGACAATGTCTTCCATCAATTTATCC |
| 4 | RibA1_GFP_rev | ACCCGGGATCTTCTCTAGAGATCACTGCAG |
| 5 | RibA2_GFP_fw | CAGGTACCAAAATGGCGTCGCTTACT |
| 6 | RibA2_GFP_rev | ACCCGGGTTCAGGAGAATCCATTGTTG |
| 7 | RibA3_GFP_fw | CAGGTACCACGATGATGGATTCTGCTTTA |
| 8 | RibA3_GFP_rev | ACCCGGGATCAAACAACGACCCGTC |
| 9 | qRT_At5g64300_fw2 | TTGTTACTTCTTGTTGTCGGG |
| 10 | qRT_At5g64300_rev2 | TGATGATCCACATTCCACAC |
| 11 | qRT_At2g22450_fw2 | GGTTCCACTCATTACTACTCCT |
| 12 | qRT_At2g22450_rev2 | AAACTAAGTCACTCAAGAAGCC |
| 13 | qRT_At5g59750_fw1 | AGACTAATGACGAATAACCCTG |
| 14 | qRT_At5g59750_rev1 | ATATCTTCTGTTCTCCTTGGTG |
| 15 | qRT_SAND_fw | AACTCTATGCAGCATTTGATCCACT |
| 16 | qRT_SAND_rev | TGATTGCATATCTTTATCGCCATC |
| 17 | AtRIBA1fw | ACCCGGGACAATGTCTTCCATCAATTTATCC |
| 18 | AtRIBA1rev | ACCCGGGTCAGGACTCAGATTCAGACTCAATC |
Abbreviations
| CLSM | confocal laser scanning microscopy |
| DHBPS | 3,4-dihydroxy-2-butanone-4- phosphate synthase |
| GCHII | GTP cyclohydrolase II |
| GFP | green fluorescent protein |
References
- Bacher, A.; Eberhardt, S.; Fischer, M.; Kis, K.; Richter, G. Biosynthesis of vitamin b2 (riboflavin). Annu. Rev. Nutr 2000, 20, 153–167. [Google Scholar]
- Fischer, M.; Bacher, A. Biosynthesis of vitamin B2: Structure and mechanism of riboflavin synthase. Arch. Biochem. Biophys 2008, 474, 252–265. [Google Scholar]
- Ouyang, M.; Ma, J.; Zou, M.; Guo, J.; Wang, L.; Lu, C.; Zhang, L. The photosensitive phs1 mutant is impaired in the riboflavin biogenesis pathway. J. Plant. Physiol 2010, 167, 1466–1476. [Google Scholar]
- Bacher, A.; Eberhardt, S.; Eisenreich, W.; Fischer, M.; Herz, S.; Illarionov, B.; Kis, K.; Richter, G. Biosynthesis of riboflavin. Vitam. Horm 2001, 61, 1–49. [Google Scholar]
- Mitsuda, H.; Suzuki, Y.; Kawai, F. Biogenesis of Riboflavin in Green Leaves. Vi. Non-Enzymatic Production of 6-Methyl-7-Hydroxy-8-Ribityllumazine from New Organic Reaction of 6,7-Dimethyl-8-Ribityllumazine with P-Quinone. J. Vitaminol. (Kyoto) 1963, 66, 125–135. [Google Scholar]
- Mitsuda, H.; Kawai, F.; Suzuki, Y.; Yoshimoto, S. Biogenesis of riboflavin in green leaves. VII. Isolation and characterization of spinach riboflavin synthetase. J. Vitaminol. (Kyoto) 1970, 16, 285–292. [Google Scholar]
- Chatwell, L.; Krojer, T.; Fidler, A.; Romisch, W.; Eisenreich, W.; Bacher, A.; Huber, R.; Fischer, M. Biosynthesis of riboflavin: structure and properties of 2,5-diamino-6-ribosylamino- 4(3H)-pyrimidinone 5′-phosphate reductase of Methanocaldococcus jannaschii. J. Mol. Biol 2006, 359, 1334–1351. [Google Scholar]
- Fischer, M.; Haase, I.; Feicht, R.; Schramek, N.; Kohler, P.; Schieberle, P.; Bacher, A. Evolution of vitamin B2 biosynthesis: riboflavin synthase of Arabidopsis thaliana and its inhibition by riboflavin. Biol. Chem 2005, 386, 417–428. [Google Scholar]
- Fischer, M.; Romisch, W.; Saller, S.; Illarionov, B.; Richter, G.; Rohdich, F.; Eisenreich, W.; Bacher, A. Evolution of vitamin B2 biosynthesis: Structural and functional similarity between pyrimidine deaminases of eubacterial and plant origin. J. Biol. Chem 2004, 279, 36299–36308. [Google Scholar]
- Herz, S.; Eberhardt, S.; Bacher, A. Biosynthesis of riboflavin in plants. The ribA gene of Arabidopsis thaliana specifies a bifunctional GTP cyclohydrolase II/3,4-dihydroxy-2-butanone 4-phosphate synthase. Phytochemistry 2000, 53, 723–731. [Google Scholar]
- Jordan, D.B.; Bacot, K.O.; Carlson, T.J.; Kessel, M.; Viitanen, P.V. Plant riboflavin biosynthesis. Cloning, chloroplast localization, expression, purification, and partial characterization of spinach lumazine synthase. J. Biol. Chem 1999, 274, 22114–22121. [Google Scholar]
- Giancaspero, T.A.; Locato, V.; de Pinto, M.C.; de Gara, L.; Barile, M. The occurrence of riboflavin kinase and FAD synthetase ensures FAD synthesis in tobacco mitochondria and maintenance of cellular redox status. FEBS J 2009, 276, 219–231. [Google Scholar]
- Sandoval, F.J.; Roje, S. An FMN hydrolase is fused to a riboflavin kinase homolog in plants. J. Biol. Chem 2005, 280, 38337–38345. [Google Scholar]
- Sandoval, F.J.; Zhang, Y.; Roje, S. Flavin nucleotide metabolism in plants: Monofunctional enzymes synthesize fad in plastids. J. Biol. Chem 2008, 283, 30890–30900. [Google Scholar]
- Richter, G.; Ritz, H.; Katzenmeier, G.; Volk, R.; Kohnle, A.; Lottspeich, F.; Allendorf, D.; Bacher, A. Biosynthesis of riboflavin: Cloning, sequencing, mapping, and expression of the gene coding for GTP cyclohydrolase II in Escherichia coli. J. Bacteriol 1993, 175, 4045–4051. [Google Scholar]
- Richter, G.; Volk, R.; Krieger, C.; Lahm, H.W.; Rothlisberger, U.; Bacher, A. Biosynthesis of riboflavin: Cloning, sequencing, and expression of the gene coding for 3,4-dihydroxy-2-butanone 4-phosphate synthase of Escherichia coli. J. Bacteriol 1992, 174, 4050–4056. [Google Scholar]
- Hedtke, B.; Alawady, A.; Albacete, A.; Kobayashi, K.; Melzer, M.; Roitsch, T.; Masuda, T.; Grimm, B. Deficiency in riboflavin biosynthesis affects tetrapyrrole biosynthesis in etiolated Arabidopsis tissue. Plant Mol. Biol 2012, 78, 77–93. [Google Scholar]
- Hedtke, B.; Grimm, B. Silencing of a plant gene by transcriptional interference. Nucleic Acids Res 2009, 37, 3739–3746. [Google Scholar]
- Bannai, H.; Tamada, Y.; Maruyama, O.; Nakai, K.; Miyano, S. Extensive feature detection of N-terminal protein sorting signals. Bioinformatics 2002, 18, 298–305. [Google Scholar]
- Emanuelsson, O.; Nielsen, H.; Brunak, S.; von Heijne, G. Predicting subcellular localization of proteins based on their N-terminal amino acid sequence. J. Mol. Biol 2000, 300, 1005–1016. [Google Scholar]
- Predotar 1.03—A Prediction Service for Identifying Putative N-Terminal Targeting Sequences. Available online: http://urgi.versailles.inra.fr/predotar/predotar.html accessed on 19 February 2012.
- Heazlewood, J.L.; Verboom, R.E.; Tonti-Filippini, J.; Small, I.; Millar, A.H. SUBA: The Arabidopsis Subcellular Database. Nucleic Acids Res 2007, 35, D213–D218. [Google Scholar]
- Fischer, M.; Romisch, W.; Schiffmann, S.; Kelly, M.; Oschkinat, H.; Steinbacher, S.; Huber, R.; Eisenreich, W.; Richter, G.; Bacher, A. Biosynthesis of riboflavin in archaea studies on the mechanism of 3,4-dihydroxy-2-butanone-4-phosphate synthase of Methanococcus jannaschii. J. Biol. Chem 2002, 277, 41410–41416. [Google Scholar]
- Kaiser, J.; Schramek, N.; Eberhardt, S.; Puttmer, S.; Schuster, M.; Bacher, A. Biosynthesis of vitamin B2. Eur. J. Biochem. 2002, 269, 5264–5270. [Google Scholar]
- Asai, S.; Mase, K.; Yoshioka, H. A key enzyme for flavin synthesis is required for nitric oxide and reactive oxygen species production in disease resistance. Plant J 2010, 62, 911–924. [Google Scholar]
- Moore, B. Bifunctional and moonlighting enzymes: Lighting the way to regulatory control. Trends Plant Sci 2004, 9, 221–228. [Google Scholar]
- Cobessi, D.; Dumas, R.; Pautre, V.; Meinguet, C.; Ferrer, J.L.; Alban, C. Biochemical and structural characterization of the Arabidopsis bifunctional enzyme dethiobiotin synthetase-diaminopelargonic acid aminotransferase: Evidence for substrate channeling in biotin synthesis. Plant Cell 2012, 24, 1608–1625. [Google Scholar]
- Spoonamore, J.E.; Dahlgran, A.L.; Jacobsen, N.E.; Bandarian, V. Evolution of new function in the GTP cyclohydrolase II proteins of Streptomyces coelicolor. Biochemistry 2006, 45, 12144–12155. [Google Scholar]
- Dereeper, A.; Guignon, V.; Blanc, G.; Audic, S.; Buffet, S.; Chevenet, F.; Dufayard, J.F.; Guindon, S.; Lefort, V.; Lescot, M.; et al. Phylogeny.fr: Robust phylogenetic analysis for the non-specialist. Nucleic Acids Res 2008, 36, W465–W469. [Google Scholar]
- Becker, D.; Kemper, E.; Schell, J.; Masterson, R. New plant binary vectors with selectable markers located proximal to the left T-DNA border. Plant Mol. Biol 1992, 20, 1195–1197. [Google Scholar]
- Bacher, A.; Richter, G.; Ritz, H.; Eberhardt, S.; Fischer, M.; Krieger, C. Biosynthesis of riboflavin: GTP cyclohydrolase II, deaminase, and reductase. Methods Enzymol 1997, 280, 382–389. [Google Scholar]
- Bandrin, S.V.; Rabinovich, P.M.; Stepanov, A.I. 3 linkage groups of the genes of riboflavin biosynthesis in Escherichia coli. Genetika 1983, 19, 1419–1425. [Google Scholar]
- Sambrook, J.; Russell, D.W. Molecular Cloning: A Laboratory Manual, 3rd ed; Cold Spring Harbor Laboratory Press: Cold Spring Harbor, NY, USA, 2001. [Google Scholar]
- Czechowski, T.; Bari, R.P.; Stitt, M.; Scheible, W.R.; Udvardi, M.K. Real-time RT-PCR profiling of over 1400 Arabidopsis transcription factors: Unprecedented sensitivity reveals novel root- and shoot-specific genes. Plant J 2004, 38, 366–379. [Google Scholar]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 2001, 25, 402–408. [Google Scholar]
- Schmittgen, T.D.; Livak, K.J. Analyzing real-time PCR data by the comparative C(T) method. Nat. Protoc 2008, 3, 1101–1108. [Google Scholar]
- Chincinska, I.A.; Liesche, J.; Krugel, U.; Michalska, J.; Geigenberger, P.; Grimm, B.; Kuhn, C. Sucrose transporter StSUT4 from potato affects flowering, tuberization, and shade avoidance response. Plant Physiol 2008, 146, 515–528. [Google Scholar]
- Bendahmane, A.; Querci, M.; Kanyuka, K.; Baulcombe, D.C. Agrobacterium transient expression system as a tool for the isolation of disease resistance genes: Application to the Rx2 locus in potato. Plant J 2000, 21, 73–81. [Google Scholar]
- Laemmli, U.K. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 1970, 227, 680–685. [Google Scholar]
- Fischer, M.; Romisch, W.; Schiffmann, S.; Kelly, M.; Oschkinat, H.; Steinbacher, S.; Huber, R.; Eisenreich, W.; Richter, G.; Bacher, A. Biosynthesis of riboflavin in archaea studies on the mechanism of 3,4-dihydroxy-2-butanone-4-phosphate synthase of Methanococcus jannaschii. J. Biol. Chem 2002, 277, 41410–41416. [Google Scholar]
- Kaiser, J.; Schramek, N.; Eberhardt, S.; Puttmer, S.; Schuster, M.; Bacher, A. Biosynthesis of vitamin B2. Eur. J. Biochem 2002, 269, 5264–5270. [Google Scholar]
- Steinbacher, S.; Schiffmann, S.; Richter, G.; Huber, R.; Bacher, A.; Fischer, M. Structure of 3,4-dihydroxy-2-butanone 4-phosphate synthase from Methanococcus jannaschii in complex with divalent metal ions and the substrate ribulose 5-phosphate: implications for the catalytic mechanism. J. Biol. Chem 2003, 278, 42256–42265. [Google Scholar]
- Ren, J.; Kotaka, M.; Lockyer, M.; Lamb, H.K.; Hawkins, A.R.; Stammers, D.K. GTP Cyclohydrolase II Structure and Mechanism. J. Biol. Chem 2005, 280, 36912–36919. [Google Scholar]
- Corpet, F. Multiple sequence alignment with hierarchical clustering. Nucl. Acids Res 1988, 16, 10881–10890. [Google Scholar]
- Nicholas, K.B.; Nicholas, H.B., Jr; Deerfield, D.W., II. GeneDoc: Analysis and Visualization of Genetic Variation. EMBnet News 1997, 4, 14. [Google Scholar]

© 2012 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Hiltunen, H.-M.; Illarionov, B.; Hedtke, B.; Fischer, M.; Grimm, B. Arabidopsis RIBA Proteins: Two out of Three Isoforms Have Lost Their Bifunctional Activity in Riboflavin Biosynthesis. Int. J. Mol. Sci. 2012, 13, 14086-14105. https://doi.org/10.3390/ijms131114086
Hiltunen H-M, Illarionov B, Hedtke B, Fischer M, Grimm B. Arabidopsis RIBA Proteins: Two out of Three Isoforms Have Lost Their Bifunctional Activity in Riboflavin Biosynthesis. International Journal of Molecular Sciences. 2012; 13(11):14086-14105. https://doi.org/10.3390/ijms131114086
Chicago/Turabian StyleHiltunen, Hanna-Maija, Boris Illarionov, Boris Hedtke, Markus Fischer, and Bernhard Grimm. 2012. "Arabidopsis RIBA Proteins: Two out of Three Isoforms Have Lost Their Bifunctional Activity in Riboflavin Biosynthesis" International Journal of Molecular Sciences 13, no. 11: 14086-14105. https://doi.org/10.3390/ijms131114086
APA StyleHiltunen, H.-M., Illarionov, B., Hedtke, B., Fischer, M., & Grimm, B. (2012). Arabidopsis RIBA Proteins: Two out of Three Isoforms Have Lost Their Bifunctional Activity in Riboflavin Biosynthesis. International Journal of Molecular Sciences, 13(11), 14086-14105. https://doi.org/10.3390/ijms131114086




