Chemoinformatics and Drug Discovery
Abstract
:Introduction
Traditional Drug Discovery Process
The Old Bottlenecks and HTS Technologies
Combinatorial Chemistry
Chemical Diversity and Cheminformatics
Drug-likeness and Lead-likeness
Paralleling Drug Discovery Process and Early ADMET Prediction
Challenges to Cheminformatics
2. The Achievements of Cheminformatics
The Origins of Cheminformatics
Descriptors and chemical structure database retrieval
Linear notations
Canonicalization
Screens and search keys
Bit-maps and fingerprints
Structure descriptors and profiling compound libraries
Dimension reduction and descriptor selection
Multidimensional scaling
Self-organising map
PCA and FA
Visualizing structures from graphed data points
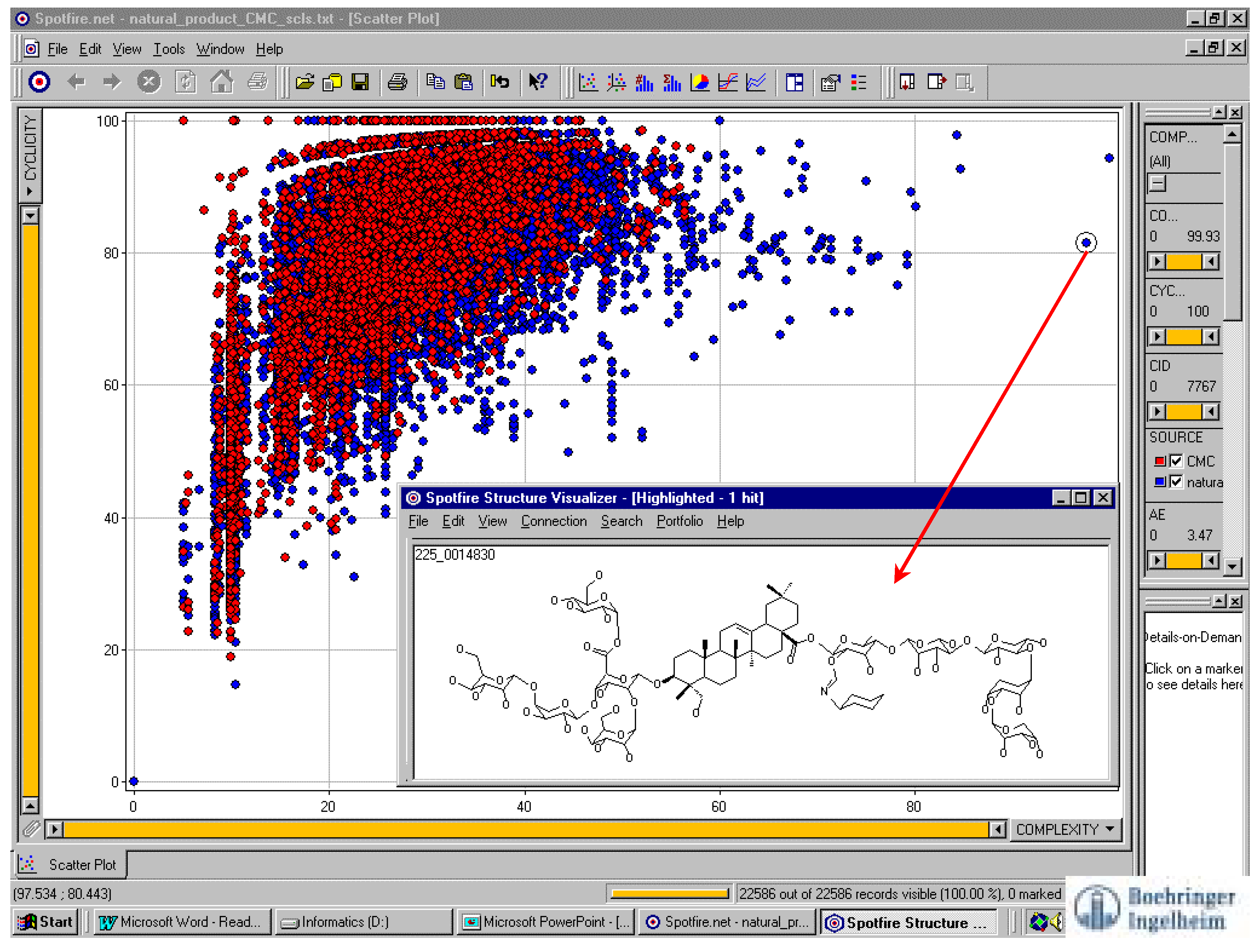
Descriptor selection
Classifications and pattern recognition
Patterns
| Pattern | Method | Remarks |
|---|---|---|
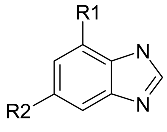 | Markush structure or generic structure | This is a topological pattern used by chemists for many years. It is determined by experience. It is an efficient way to represent an unlimited number of compounds with the same scaffold. Additional restrictions can be applied to make the pattern more specific. It is suitable for lead optimization and hit-to-lead efforts. |
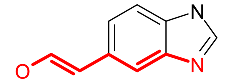 | Fingerprint | This is the topological pattern systematically generated from an algorithm. This pattern has no human bias, but can be meaningless to chemistry. It is used in HTS data mining. |
 | Three-dimensional pharmacophore | This pattern is derived, manually or computationally, from a three-dimensional molecular model. The pattern is based upon a physical model and binding mechanism. It is sensitive to conformation changes. Better results are obtained when supported by crystal or NMR structural data. It is suitable for lead optimization. |
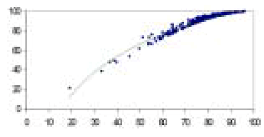 | Regression | Regression methods are the most traditional approaches for pattern recognition. These methods assume the variables are continuous and the curve shapes are pre-defined. For multidimensional data, curve patterns are not known and trying all possible curves is very time consuming. In these cases, genetic algorithms may be applied to partially solve the problem of identifying curve patterns. |
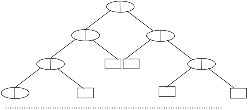 | Decision tree classification | This approach is applied when there are a great number of descriptors and, the descriptors have various value types and ranges. |
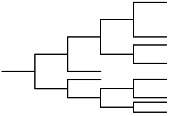 | Hierarchical clustering | This approach assumes the objects have hierarchical characters. The methods require similarity or distance matrices. The approach may produce multiple answers for users to explain or with which to experiment. |
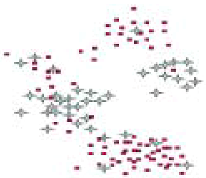 | Non-hierarchical clustering | The approach assumes the objects have non-hierarchical characters, and the number of clusters is known prior the computation. The method requires similarity or distance matrices. The approach may produce multiple answers for users to explain or with which to experiment. |
 | Self-Organization Map (SOM) | This is a neural network approach. The number of neurons, configuration of neurons, neighboring function, training rate and area, and monitoring parameters should be predefined. This method needs similarity or distance measurements [50]. |
Similarity or Distance metrics
Clustering
Partitioning
Applications in drug discovery
Compound selection
Virtual library generation
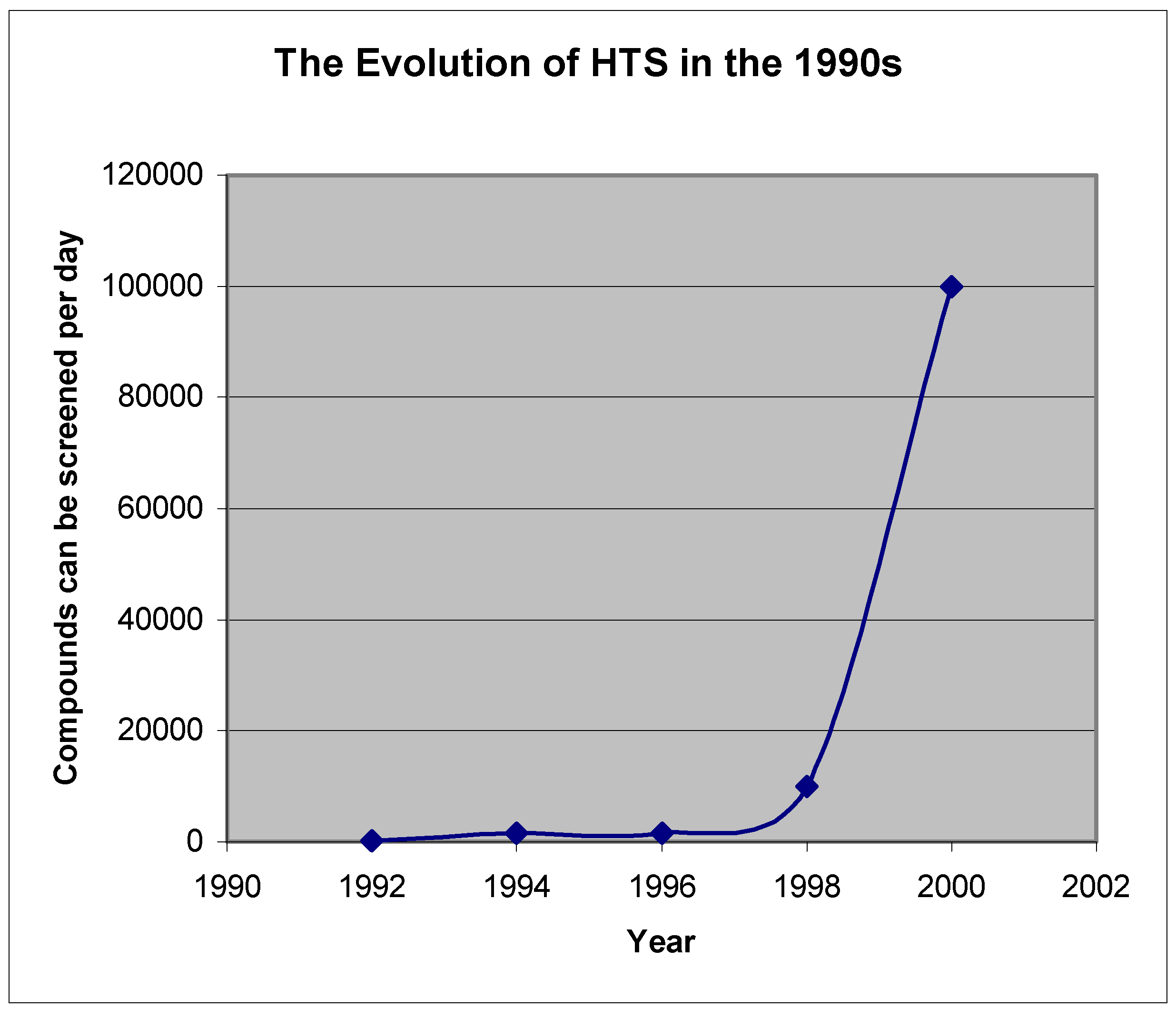
Virtual screening
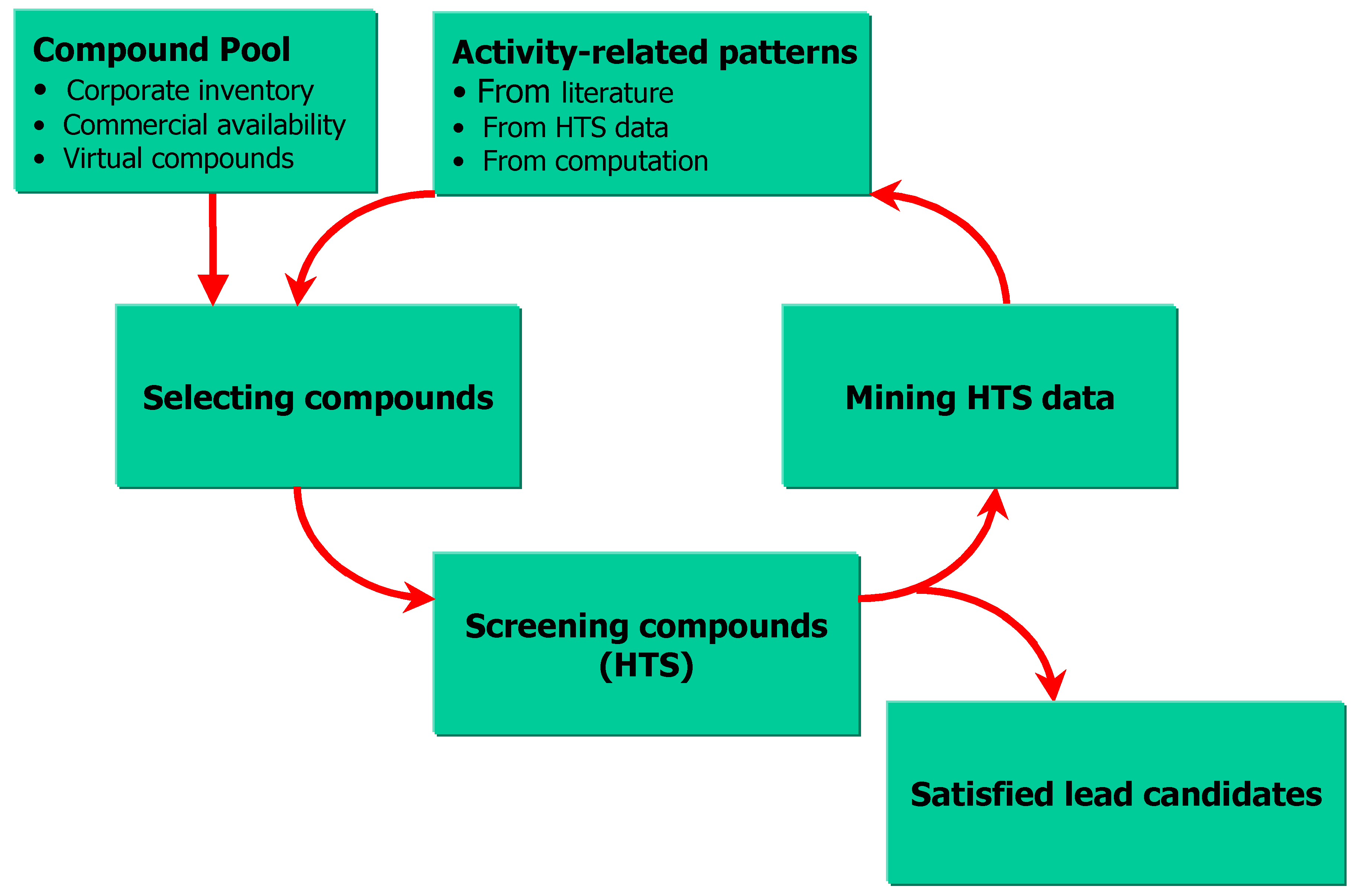
SAR on HTS data and sequential screening
In silico ADMET
- Absorption. Passive intestinal absorption (PIA) models have been studied by many groups, for years. The fluid mosaic model holds that the structure of a cell membrane is an interrupted phospholipid bilayer capable of both hydrophilic and hydrophobic interactions [106]. Trans-cellular passage through the membrane lipid/aqueous environment is the predominant pathway for passive absorption of lipophilic compounds, while low-molecular-weight (<200), hydrophilic compounds make use of the water-filled channels of the tight junctions between membrane cells (paracellular transport) [107]. Therefore, lipophilicity is considered a key property for activity in drug design and is a common property used to estimate the membrane permeability of a molecule. Lipophilicity is measured as the log of the partition coefficient between n-octanol and water (logP). LogP prediction programs are available and results are reasonably good [108a-e]. But, the relationship between logP and permeability is not linear. Permeability drops at both low and high logP. It is theorized that These non-linearities due to: (1) the inability of weakly lipophilic compounds to penetrate the lipid portion of the membrane and (2) the excessive partitioning of strongly lipophilic compounds into the lipid portion of the membrane and their subsequent inability to pass through the aqueous portion of the membrane [108f]. A strong relationship between PIA and polar surface area (PSA) has been discovered by several groups [109,110,111,112,113]. However, the models usually do not take the effects of other descriptors into account. In addition, the datasets used to build the PSA models are small. Even though a wide range of PSA was covered, it is not necessarily true the models cover the entire chemical space. Therefore, linear and non-linear multivariate models have been introduced to model PIA based upon: logP, molecular weight, H-bonding, free energy, H-bond donor, H-bond acceptor, polarizability, numbers and strengths of H-bond acceptor nitrogen and oxygen atoms, number of H-bond donor atoms, and lipophilicity (log D at pH 7.4) on the Caco-2 cell permeability. To select the best descriptors for predictive models, a genetic algorithm has been used
- Distribution. CNS-active drugs (CNS, central nervous system) must cross the blood-brain barrier (BBB). The experimental determination of the brain-blood partition ratio is difficult and time-consuming to compute since it involves the direct measurement of the drug concentration in the brain and blood of laboratory animals. This obviously requires the synthesis of the compounds, often in radiolabeled form [120]. In vitro techniques to predict brain penetration are available [121], but they are experimentally cumbersome. The earlier work involved in correlating log(Cbrain/Cblood) or logBB and logP (octanol-cyclohexane), Pcyclohexane, or logPoct was based upon smaller (about 20 compounds) data sets [122,123,124]. More descriptors have been correlated with logBB [125,126,127], such as: excess molar refraction, solute polarizability, hydrogen bond acidity and basicity, and molecular volume. More recently a regression study on logBB and free energy G has been reported [122]. Descriptors derived from 3D molecular fields to estimate the BBB permeation on a larger set of compounds and to produce a simple mathematical model have been studied. The method used (VolSurf) transforms 3D fields into descriptors and correlates them to the experimental permeation by a discriminant partial least squares procedure [128]. Human serum albumin (HSA) protein is the major transporter of non-esterified fatty acids, as well as of different drugs and metabolites, to different tissues. HSA allows solubilization of hydrophobic compounds, contributes to a more homogeneous distribution of drugs in the body, and increases their biological lifetime. The binding strength of any drug to serum albumin is the main factor for availability of that drug to diffuse from the circulatory system to target tissues. All these factors cause the pharmacokinetics of almost any drug to be influenced and controlled by its binding to serum albumin [129]. Therefore, QSAR study on binding of drugs and metabolites to HSA is extremely important for the drug distribution. Biosensor analysis for prediction of HSA has been reported 130. In order to build an in silico predictive model for binding affinities to HSA, Colmenarejo and co-workers at GlaxoSmithKline used a genetic algorithm to exhaustively search and select for multivariate and non-linear equations, starting from a large pool of molecular descriptors. They found that hydrophobicity (as measured by the ClogP) is the most important variable for determining the binding extent to HSA. Binding to HSA turns out to be determined by a combination of hydrophobic forces together with some modulating shape factors [131]. This agrees with X-ray structures of HSA alone or, bound to ligands, where the binding pockets of both sites I and II are composed mainly of hydrophobic residues [132].
- Metabolism. Drug metabolism is another barrier to overcome. Metabolism is studied, by in vitro, in vivo and in silico approaches. HTS has been used for metabolism and pharmacokinetics [133,134]. In vitro approaches determine metabolic stability, screening for inhibitors of specific cytochrome P450 isozymes and, identifying the most important metabolites. In vivo approaches measure hepatic metabolic clearance, volume of distribution, bioavailability, and, identify major metabolites. In silico approaches are categorized into three classes [135]: QSAR and pharmacophore models, protein models, and expert systems. QSAR and pharmacophore models predict substrates and inhibitors of a specific cytochrome P450 isozyme [136,137]. Protein models rationalize metabolite formations and identify possible substrates, potential metabolites or, inhibitors by means of docking algorithms [138,139]. Stereoelectronic factors involved in metabolic transformations can be taken into account using quantum chemical calculations. Expert systems are predictive databases that attempt to identify potential metabolites of a compound as determined by knowledge based rules defining the most likely products [140,141]. Testa advised that in structure-metabolism relationship (SMR) studies, the greater the chemical diversity of the investigated compounds, the smaller the chance that SMRs exist and can be uncovered. On the other hand, the information content of an SMR (if it exists) will increase as the boundaries of the chemical space increases and as the diversity of the compounds under investigation increases [142]. This paradox may limit the capacity of SMR, no matter which approach is used. Keseru and Molnar [135] think efficient PK optimization requires metabolic diversity within the focused library that cannot be achieved by the application of a simple SMR with limited information content. The high degree of structural similarity (especially in combinatorial libraries with a common core) prevents the application in metabolic diversity analysis. Therefore, they introduced a metabolic fingerprint concept, METAPRINT, for the assessment of metabolic similarity and diversity in combinatorial chemical libraries. Their metabolic fingerprint was developed by predicting metabolic pathways and corresponding potential metabolites.
- Excretion/Elimination. Drugs such as the non-steroidal anti-inflammatory drugs (NSAIDs), are used in long term treatment. The accumulation of these drugs in the body may lead to serious side effects. Therefore, the prediction of half-life, which determines the length of time a drug will persist in the body, is important in order to reduce subsequent drug failures. Prediction of half-life is difficult, due to the multi-faceted nature of drug elimination. Distribution of drug in fat and major organs, excretion by kidneys and metabolism by liver all contribute to the rate at which a drug is eliminated from the body. On the other hand, it may be possible to make use of qualitative predictions of half-life. Such information can be used, for example, to predict whether a drug is likely to accumulate to a significant extent when used for prolonged treatment [143].
- Toxicity. Many drugs are withdrawn for safety reasons and there are many reasons, including metabolism and excretion/elimination that cause toxicity. Current toxicity prediction approaches use either mechanistic or correlative methods. Correlative systems take molecular descriptors, biological data, and chemical structures and, by use of statistical analysis of data sets, represent them in mathematical models. The models describe the relationships between structure and activity and can be used to predict toxicity. The mechanistic approach involves human experts who make a considered assessment of the mechanism of interaction with a biological system, taking the molecular properties, biological data, and chemical structures into account [144]. The correlative approach uses an unbiased assessment of the data to generate relationships and predict toxicity. It is capable of discovering potentially new SARs [145] and, can lead to new ideas in the human assessment of mechanisms by which chemicals interact with biological systems. It is most useful for congeneric data sets or when one has a large amount of good data but little mechanistic knowledge. However, it can also generate relationships that have little chemical or biological plausibility. Results obtained are heavily dependent upon the quality of the data used to build the model. For these reasons careful validation is required for effective use of the correlative approach. The mechanistic method is based upon an understanding or hypothesis of the mechanisms of molecular interactions that determine the activity, i.e., there is some human input into the system of SAR generation. However, systems using this approach are restricted to human knowledge, being incapable of discovering new relationships automatically. As a consequence, they also have a tendency to be biased toward current ideas about mechanisms of action [144]. The early toxicity models were based on QSAR models and were used to predict LD50 [144], based upon various descriptors [146,147,148]. It was also reported that QSAR models (partial least-squares (PLS), Bayesian regularized neural network) correlating IGC50 [149] with the hydrophobicity, the logarithm of the 1-octanol/water partition coefficient, the molecular orbital properties, the lowest unoccupied molecular orbital energy (Elumo) and, maximum acceptor super-delocalizability (Amax) [150,151]. More QSAR models are still coming forth [152,153]. A representative mechanistic toxicity prediction approach was reported by Sanderson and co-workers [144,154,155,156]. The program is now commercially available [157]. Artificial neural networks (ANN) have recently been applied in toxicity predictions [158,159,160]; these include: back-propagation neural network, Bayesian-Regularized Neural Networks, and self-organization map (SOM). The organizations providing ADMET solutions are listed in reference [161].
3. Future Directions
Parallel optimization.
The paradox of predictivity versus diversity
From data mining to knowledge discovery
Acknowledgments
References and Notes
- Augen, J. The evolving role of information technology in the drug discovery process. Drug Discov. Today 2002, 7, 315–323. [Google Scholar]
- Gallop, M. A.; Barrett, R. W.; Dower, W. J.; Fodor, S. P. A.; Gordon, E. M. Applications of Combinatorial Technologies to Drug Discovery. 1. Background and Peptide Combinatorial Libraries. J. Med. Chem. 1994, 37, 1233–1251. [Google Scholar]
- Hecht, P. High-throughput screening: beating the odds with informatics-driven chemistry. Curr. Drug Discov. 2002, 21–24. [Google Scholar]
- Hall, D. G.; Manku, S.; Wang, F. Solution- and Solid-Phase Strategies for the Design, Synthesis, and Screening of Libraries Based on Natural Product Templates: A Comprehensive Survey. J. Comb. Chem. 2001, 3, 125–150. [Google Scholar]
- Bemis, G. W.; Murcko, M. A. The properties of known drugs. 1. Molecular Frameworks. J. Med. Chem. 1996, 39, 2887–2893. [Google Scholar] [CrossRef] Bemis, G. W.; Murcko, M. A. The properties of known drugs. 2. Side Chains. J. Med. Chem. 1999, 42, 5095–5099. [Google Scholar] [CrossRef]
- Ajay; Walters, W. P.; Murcko, M. A. Can we learn to distinguish between “drug-like” and “non-drug-like” molecules? J. Med. Chem. 1998, 41, 3314–3324. [Google Scholar]
- Sadowski, J.; Kubinyi, H. A scoring scheme for discriminating between drugs and non-drugs. J. Med. Chem. 1998, 41, 3325–3329. [Google Scholar] [CrossRef]
- Xu, J.; Stevenson, J. Drug-like Index: A New Approach To Measure Drug-like Compounds and Their Diversity. J. Chem. Inf. Comput. Sci. 2000, 40, 1177–1187. [Google Scholar] [CrossRef]
- Lipinski, C.A.; Lombardo, F.; Dominy, B.W.; Feeney, P.J. Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv. Drug Deliv. Rev. 1997, 23, 3–25. [Google Scholar] [CrossRef]
- Clark, D. E.; Pickett, S. D. Computational methods for the prediction of ‘drug-likeness’. Drug Discov. Today 2000, 5, 49–58. [Google Scholar]
- Matter, H.; Baringhaus, K.-H.; Naumann, T.; Klabunde, T.; Pirard, B. Computational approaches towards the rational design of drug-like compound libraries. Comb. Chem. High T. Scr. 2001, 4, 453–475. [Google Scholar]
- Oprea, T. I.; Davis, A. M.; Teague, S. J.; Leeson, P. D. Is There a Difference between Leads and Drugs? A Historical Perspective. J. Chem. Inf. Comput. Sci. 2001, 41, 1308–1315. [Google Scholar]
- Proudfoot, J. R. Drugs, Leads, and Drug-Likeness: An Analysis of Some Recently Launched Drugs. Bioorg. Med. Chem. Lett. 2002, (in press). [Google Scholar]
- Stewart, L.; Clark, R.; Behnke, C. High-throughput crystallization and structure determination in drug discovery. Drug Discov. Today 2002, 7, 187–196. [Google Scholar]
- Luft, J. R.; Wolfley, J.; Collins, R.; Bianc, M.; Weeks, D.; Jurisica, I.; Rogers, P.; Glasgow, J.; Fortier, S.; DeTitta, G. T. High Throughput Protein Crystallization: Keeping up with the Genomics. 2002. www.imca.aps.anl.gov/~ahoward/luft_ab.html.
- Kennedy, T. Drug Discov. Today 1997, 2, 436–444. Start-Up: Windhover’s Review of Emerging Medical Ventures. July 2000, p. page 34. www.windhoverinfo.com/contents/monthly/exex/e_2000900126.htm.
- Manly, C. J.; Louise-May, S.; Hammer, J. D. The impact of informatics and computational chemistry on synthesis and screening. Drug Discov. Today 2001, 6, 1101–1110. [Google Scholar]
- Baxter, A. D.; Lockey, P. ‘Hit’ to ‘lead’ and ‘lead’ to ‘candidate’ optimization using multi-parametric principles. Drug Discov. World 2001, 2, 9–15. [Google Scholar]
- Wilson, E. K. Picking the winners. Chem. Eng. News 2002, 35–39. [Google Scholar]
- http://pubs.acs.org/archives/percent.html.
- Xu, J. GMA: A Generic Match Algorithm for structural Homomorphism, Isomorphism, Maximal Common Substructure Match and Its Applications. J. Chem. Inf. Comput. Sci. 1996, 36, 25–34. [Google Scholar] [CrossRef]
- http://www.asis.org/Features/Pioneers/wiswess.htm.
- Weininger, D. SMILES, a chemical language and information system. 1. Introduction to methodology and encoding rules. J. Chem. Inf. Comput. Sci. 1988, 28, 31–6. [Google Scholar] [CrossRef]
- http://esc.syrres.com/interkow/docsmile.htm.
- Wiener, H. Structural Determination of Paraffin Boiling Points. J. Am. Chem. Soc. 1947, 69, 17–20. [Google Scholar] [CrossRef]
- Hu, C.; Xu, L. On Highly Discriminating Molecular Topological Index. J. Chem. Inf. Comput. Sci. 1996, 36, 82–90. [Google Scholar]
- The definitions of MDL’s 166 MACCS search keys can be found from ISIS/Base Help file under “Remote QB in a Molecule Database: Searching Concepts/Examples” at the section 49.2.4: Specifying Searchable Keys as a Query.
- http://www.daylight.com/about/f_search.html.
- Rhodes, N.; Willett, P. Bit-String Methods for Selective Compound Acquisition. J. Chem. Inf. Comput. Sci. 2000, 40, 210–214. [Google Scholar]
- Kier, L. B.; Hall, L. H. Molecular Connectivity in Structure-Activity Analysis. Research Studies Press: Lectchworth, Hertfordshire, England, 1986. [Google Scholar]
- http://www.disat.unimib.it/chm/.
- Chemical Computing Group, Inc., 1010 Sherbrooke Street West, Suite 910, Montreal, Quebec, Canada, H3A 2R7, Tel: (514) 393-1055 Fax: (514) 874-9538.
- Hall, L. H. Computational Aspects of Molecular Connectivity and its Role in Structure-Property Modeling. In Computational Chemical Graph Theory; Rouvray, D. H., Ed.; Nova Press: New York, 1990; Chap. 8; pp. 202–233. [Google Scholar]
- Chemical Computing Group, Inc., 1010 Sherbrooke Street West, Suite 910, Montreal, Quebec, Canada, H3A 2R7, Tel:(514) 393-1055 Fax: (514) 874-9538.
- Accelrys Inc. a subsidiary of Pharmacopeia Inc.
- Cox, T.F.; Cox, M. A. A. Multidimensional Scaling. 2000. [Google Scholar]
- http://www.statsoft.com/textbook/stmulsca.html#general.
- Kohonen, T.; Kangas, J.; Laaksonen, J. SOM_PAK, The Self-Organizing Map Program Package available for anonymous ftp user at Internet site cochlea.hut.fi, version 1.2, November 1992.
- Zupan, J.; Gasteiger, J. Neural Networks for Chemists. VCH: Weinheim, 1993. [Google Scholar]
- Bernard, P.; Golbraikh, A.; Kireev, D.; Chrétien, J. R.; Rozhkova, N. Comparison of chemical databases: Analysis of molecular diversity with Self Organising Maps (SOM). Analusis 1998, 26, 333–346. [Google Scholar]
- http://www.statsoft.com/textbook/stfacan.html.
- Joliffe, I.T. Principal Component Analysis; Springer-Verlag: New York, 1986. [Google Scholar]
- Malinowski, E.H.; Howery, D.G. Factor Analysis in Chemistry; John Wiley & Sons: New York, 1980. [Google Scholar]
- http://www.spotfire.com/.
- Xu, J. SCA: New Cluster Algorithm for Structural Diversity Analysis and Applications. The First Spotfire Users Conference. Philadelphia, May 30, 2001.
- Brown, R. D.; Martin, Y. C. Use of Structure-Activity Data To Compare Structure-Based Clustering Methods and Descriptors for Use in Compound Selection. J. Chem. Inf. Comput. Sci. 1996, 36, 572–584. [Google Scholar]
- Matter, H.; Pötter, T. Comparing 3D Pharmacophore Triplets and 2D Fingerprints for Selecting Diverse Compound Subsets. J. Chem. Inf. Comput. Sci. 1999, 39, 1211–1225. [Google Scholar] [CrossRef]
- Estrada, E.; Molina, E.; Perdomo-Lopez, I. Can 3D Structural Parameters Be Predicted from 2D (Topological) Molecular Descriptors? J. Chem. Inf. Comput. Sci. 2001, 41, 1015–1021. [Google Scholar]
- Xue, L.; Stahura, F. L.; Godden, J. W.; Bajorath, J. Mini-fingerprints Detect Similar Activity of Receptor Ligands Previously Recognized Only by Three-Dimensional Pharmacophore-Based Methods. J. Chem. Inf. Comput. Sci. 2001, 41, 394–401. [Google Scholar]
- 2002. http://spheroid.ncifcrf.gov/scripts/mapviewer.cfm.
- 2001. http://www.daylight.com/about/f_search.html.
- Tryon, R. C. J. Chronic Dis. 1939, 20, 511–524. http://www.statsoftinc.com/textbook/stcluan.html.
- Jarvis, R.A.; Patrick, E.A. Clustering Using a Similarity Measure Based on Shared Near Neighbors. 1973, C22, 1025–1034. [Google Scholar]
- Hierarchical cluster methods are implemented in agglomerative (bottom-up) or divisive (top-down) procedure. The hierarchical clustering approach finds a hierarchy of objects represented by a number of descriptors. There are three methods to merge objects into clusters: the centroid method, Ward's method and average linkage. For an agglomerative procedure, each object begins in a cluster by itself. The two closest clusters are merged to form a new cluster replacing the two old clusters. Merging of the two closest clusters is repeated until only one cluster remains. The different hierarchical clustering methods differ in how the distance between two clusters is computed. In the centroid method, the distance between two clusters is defined as the distance between their centroids or means. The centroid method is more robust than most other hierarchical methods but, in many other respects, does not perform as well as Ward's method or, average linkage. In Ward's method, the distance between two clusters is the sum of squares between the two clusters added up over all of the variables. At each generation, the within-cluster sum of squares is minimized over all partitions obtainable by merging two clusters from the previous generation. This method tends to join clusters with a small number of objects and, is biased toward producing clusters with roughly the same number of objects. The average linkage distance between two clusters is defined as the average distance (squared Euclidean) between pairs of objects, one in each cluster. Average linkage tends to join clusters with small variances and, is biased toward producing clusters with roughly the same variance. Studies suggest that Ward's method and average linkage method are among the better hierarchical clustering algorithms. Intrinsically, hierarchical clustering approaches ignore the fact that scientific data may have many outliers. They average all objects eventually to one cluster. However, the outliers should statistically be left alone.
- Most popular partitional cluster algorithms are K-mean algorithms and Javis-Patrick (K-nearest neighbor, Knn) algorithms. K-mean clustering algorithms use an interchange (or switching) method to divide n data points into K groups (clusters) so that the sum of distances/dissimilarities among the objects within the same cluster is minimized. The K-mean approach requires that K (the number of clusters) is known before clustering. In the most of cases, however, the number of clusters may be not known. The K-mean clustering result depends on the order of the rows in the input data, the options of K-bins initialization, and number of iterations for minimizing distances. Even if there is a best guess for K, the K-mean approach involves a NP problem (combinatorial explosion). The number of combinations of partitioning N objects into K groups is an astronomical high figure. It will force a program to abort after a given number of iterations in order to produce result in a feasible period of time. Javis-Patrick requires the user specifies the number of nearest neighbors, and the number of neighbors in common to merge to objects. Javis-Patrick is a deterministic algorithm, it doesn’t require number of iterations for computations. Both K-mean and Javis-Patrick algorithms do not directly give the answer for the number of clusters.
- Willett, P. Similarity and Clustering in Chemical Information Systems. Research Studies Press, Wiley: New York, 1987. [Google Scholar]
- Rusinko, A., III; Farmen, M. W.; Lambert, C. G.; Brown, P. L.; Young, S. S. Analysis of a Large Structure/Biological Activity Data Set Using Recursive Partitioning. J. Chem. Inf. Comput. Sci. 1999, 39, 1017–1026. [Google Scholar]
- Rusinko, A., III; Young, S. S.; Drewry, D. H.; Gerritz, S. W. Optimization of Focused Chemical Libraries Using Recursive Partitioning. Comb. Chem. High T. Scr. 2002, 5, 125–133. [Google Scholar]
- Wikel, J. H.; Higgs, R. E. Applications of molecular diversity analysis in high throughput screening. J. Biomol. Screen. 1997, 2, 65–67. [Google Scholar] [CrossRef]
- Sadowski, J.; Wagener, M.; Gasteiger, J. Assessing similarity and diversity of combinatorial libraries by spatial autocorrelation functions and neural networks. Angew. Chem. Int. Ed. Engl. 1995, 34, 2674–2677. [Google Scholar] [CrossRef]
- Sheridan, R. P.; Kearsley, S. K. Using a genetic algorithm to suggest combinatorial libraries. J. Chem. Inf. Comput. Sci. 1995, 35, 310–320. [Google Scholar] [CrossRef]
- Brown, R. D.; Martin, Y. C. Use of Structure-Activity Data To Compare Structure-Based Clustering Methods and Descriptors for Use in Compound Selection. J. Chem. Inf. Comput. Sci. 1996, 36, 572–584. [Google Scholar]
- Gillet, V. J.; Willett, P.; Bradshaw, J. The Effectiveness of Reactant Pools for Generating Structurally-Diverse Combinatorial Libraries. J. Chem. Inf. Comput. Sci. 1997, 37, 731–740. [Google Scholar]
- Agrafiotis, D. K. Stochastic Algorithms for Maximizing Molecular Diversity. J. Chem. Inf. Comput. Sci. 1997, 37, 841–851. [Google Scholar]
- Agrafiotis, D. K.; Lobanov, V. S. An Efficient Implementation of Distance-Based Diversity Measures Based on k-d Trees. J. Chem. Inf. Comput. Sci. 1999, 39, 51–58. [Google Scholar]
- Clark, R. D. OptiSim: An Extended Dissimilarity Selection Method for Finding Diverse Representative Subsets. J. Chem. Inf. Comput. Sci. 1997, 37, 1181–1188. [Google Scholar]
- Clark, R. D.; Langton, W. J. Balancing Representativeness Against Diversity using Optimizable K-Dissimilarity and Hierarchical Clustering. J. Chem. Inf. Comput. Sci. 1998, 38, 1079–1086. [Google Scholar]
- Pötter, T.; Matter, H. Random or Rational Design? Evaluation of Diverse Compound Subsets from Chemical Structure Databases. J. Med. Chem. 1998, 41, 478–488. [Google Scholar] [CrossRef]
- Pearlman, R. S.; Smith, K. M. Metric Validation and the Receptor-Relevant Subspace Concept. J. Chem. Inf. Comput. Sci. 1999, 39, 28–35. [Google Scholar]
- Bayada, D. M.; Hamersma, H.; van Geerestein, V. J. Molecular Diversity and Representativity in Chemical Databases. J. Inf. Comput. Sci. 1999, 39, 1–10. [Google Scholar]
- Xue, L.; Godden, J.; Gao, H.; Bajorath, J. Identification of a Preferred Set of Molecular Descriptors for Compound Classification Based on Principal Component Analysis. J. Info. Comput. Sci. 1999, 39, 699–704. [Google Scholar]
- Munk Jörgensen, A. M.; Pedersen, J. T. Structural Diversity of Small Molecule Libraries. J. Chem. Inf. Comput. Sci. 2001, 41, 338–445, This paper reported a method for assessing structural diversity based upon maximum common sub-graph identity as the measure of similarity between two chemical structures. A conditional probability treatment of similarity distributions for libraries of chemical structures is used to define diversity. [Google Scholar]
- Mount, J.; Ruppert, J.; Welch, W.; Jain, A. N. IcePick: a flexible suface-based system for molecular diversity. J. Med. Chem. 1999, 42, 60–66. [Google Scholar] [CrossRef]
- Zheng, W.; Cho, S. J.; Waller, C. L.; Tropsha, A. J. J. Chem. Inf. Comput. Sci. 1999, 39, 738–746.
- Reynolds, C. H.; Druker, R.; Pfahler, L. B. Lead Discovery Using Stochastic Cluster Analysis (SCA): A New Method for Clustering Structurally Similar Compounds. J. Chem. Inf. Comput. Sci. 1998, 38, 305–312. [Google Scholar] [CrossRef]
- Reynolds, C. H.; Tropsha, A.; Pfahler, L. B.; Druker, R.; Chakravorty, S.; Ethiraj, G.; Zheng, W. Diversity and Coverage of Structural Sublibraries Selected Using the SAGE and SCA Algorithms. J. Chem. Inf. Comput. Sci. 2001, 41, 1470–1477, This paper discussed rational approaches to selecting representative subsets of virtual libraries that help direct experimental synthetic efforts for diverse library design. The authors compared the performance of two stochastic sampling algorithms, Simulating Annealing Guided Evaluation (SAGE) and Stochastic Cluster Analysis (SCA) for their ability to select both diverse and representative subsets of the entire chemical library space. Tests were carried out using simulated two-dimensional data sets and a 27,000 compound proprietary structural library as represented by computed Molconn-Z descriptors. The algorithmically simple SCA method is capable of selecting subsets that are comparable to the more computationally intensive SAGE method. [Google Scholar] [CrossRef]
- Agrafiotis, D. K.; Rassokhin, D. N. A Fractal Approach for Selecting an Appropriate Bin Size for Cell-Based Diversity Estimation. J. Chem. Inf. Comput. Sci. 2002, 42, 117–122, This paper reported an approach for selecting an appropriate bin size for cell-based diversity assessment. The method measures the sensitivity of the diversity index as a function of grid resolution, using a box-counting algorithm that is reminiscent of those used in fractal analysis. It is shown that the relative variance of the diversity score (sum of squared cell occupancies) of several commonly used molecular descriptor sets exhibits a bell-shaped distribution, whose exact characteristics depend on the distribution of the data set, the number of points considered, and the dimensionality of the feature space. The peak of this distribution represents the optimal bin size for a given data set and sample size. Although box counting can be performed in an algorithmically efficient manner, the ability of cell-based methods to distinguish between subsets of different spread falls sharply with dimensionality, and the method becomes useless beyond a few dimensions. [Google Scholar]
- Trepalin, S. V.; Gerasimenko, V. A.; Kozyukov, A.V; Savchuk, N. Ph.; Ivaschenko, A. A. New Diversity Calculations Algorithms Used for Compound Selection. J. Chem. Inf. Comput. Sci. 2002, 42, 249–258. [Google Scholar]
- Hamprecht, F. A.; Thiel, W.; van Gunsteren, W. F. Chemical Library Subset Selection Algorithms: A Unified Derivation Using Spatial Statistics. J. Chem. Inf. Comput. Sci. 2002, 42, 414–428, The authors modeled activity in a bioassay as realization of a stochastic process and use the best linear unbiased estimator to construct spatial sampling designs that optimize the integrated mean square prediction error, the maximum mean square prediction error, or the entropy. Author’s approach constitutes a unifying framework encompassing most proposed techniques as limiting cases and sheds light on their underlying assumptions. In particular, vector quantization is obtained, in dimensions up to eight, in the limiting case of very smooth response surfaces for the integrated mean square error criterion. Closest packing is obtained for very rough surfaces under the integrated mean square error and entropy criteria. The paper suggested using either the integrated mean square prediction error or the entropy as optimization criteria rather than approximations thereof and proposing a scheme for direct iterative minimization of the integrated mean square prediction error.. [Google Scholar]
- Bajorath, J. Selected Concepts and Investigations in Compound Classification, Molecular Descriptor Analysis, and Virtual Screening. J. Chem. Inf. Comput. Sci. 2001, 41, 233–245. [Google Scholar]
- Mander, T. Beyond uHTS: ridiculously HTS? Drug Discov. Today 2000, 5, 223–225. [Google Scholar]
- Valler, M. J.; Green, D. Diversity screening versus focused screening in drug discovery. Drug Discov. Today 2000, 5, 286–293. [Google Scholar]
- Walters, W. P.; Stahl, M. T.; Murcko, M. A. Virtual screening – an overview. Drug Discov. Today 1998, 3, 160–178. [Google Scholar]
- Joseph-McCarthy, D. An overview of in silico design and screening: Toward efficient drug discovery. Curr. Drug Discov. 2002, 20–23. [Google Scholar]
- Bajorath, J. Virtual screening in drug discovery: Methods, expectations and reality. Curr. Drug Discov. 2002, 24–27. [Google Scholar]
- Downs, G. M.; Barnard, J. M. Techniques for Generating Descriptive Fingerprints in Combinatorial Libraries. J. Chem. Inf. Comput. Sci. 1997, 37, 59–61. [Google Scholar] [CrossRef]
- Lobanov, V. S.; Agrafiotis, D. K. Scalable Methods for the Construction and Analysis of Virtual Combinatorial Libraries. Comb. Chem. High T. Scr. 2002, 5, 167–178. [Google Scholar] [Green Version]
- Lipinski, C. A.; Lombardo, F.; Dominy, B. W.; Feeney, P. J. Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv. Drug Deliver. Rev. 1997, 23, 3–25. [Google Scholar] [CrossRef]
- Huuskonen, J.; Rantanen, J.; Livingstone, D. Prediction of aqueous solubility for a diverse set of organic compounds based on atom-type electrotopological state indices. Eur. J. Med. Chem. 2000, 35, 1081–1088. [Google Scholar] [CrossRef]
- Zuegge, J.; Schneider, G.; Coassolo, P.; Lave, T. Prediction of hepatic metabolic clearance-comparison and assessment of prediction models. Clin. Pharmacokinet. 2001, 40, 553–563. [Google Scholar] [CrossRef]
- Roche, O.; Schneider, P.; Zuegge, J.; Guba, W.; Kansy, M.; Alanine, A.; Bleicher, K.; Danel, F.; Gutknecht, E. M.; Rogers-Evans, M.; Neidhart, W.; Stalder, H.; Dillon, M.; Sjogren, E.; Fotouhi, N.; Gillespie, P.; Goodnow, R.; Harris, W.; Jones, P.; Taniguchi, M.; Tsujii, S.; von der Saal, W.; Zimmermann, G.; Schneider, G. Development of a Virtual Screening Method for Identification of ‘Frequent Hitters’ in Compound Libraries. J. Med. Chem. 2002, 45, 137–142. [Google Scholar] [CrossRef]
- Abagyan, R.; Totrov, M. High-throughput docking for lead generation. Curr. Opin. Chem. Biol. 2001, 5, 375–382. [Google Scholar] [CrossRef]
- Diller, D. J.; Merz, K. M., Jr. High throughput docking for library design and library prioritization. Proteins 2001, 43, 113–124. [Google Scholar]
- Willett, P. Chemoinformatics – similarity and diversity in chemical libraries. Curr. Opin. Biotech. 2000, 11, 85–88. [Google Scholar] [CrossRef]
- Hopfinger, A. J.; Duca, J. S. Estimation of molecular similarity based on 4D-QSAR analysis: formalism and validation. J. Chem. Inf. Comput. Sci. 2001, 41, 1367–1387. [Google Scholar] [CrossRef]
- Makara, G. M. Measuring molecular similarity and diversity: total pharmacophore diversity. J. Med. Chem. 2001, 44, 3563–3571. [Google Scholar] [CrossRef]
- Hopfinger, A. J.; Duca, J. Extraction of pharmacophore information from high-throughput screens. Curr. Opin. Biotech. 2000, 11, 97–103. [Google Scholar] [CrossRef]
- Roberts, G.; Myatt, G. J.; Johnson, W. P.; Cross, K. P.; Blower, P. E., Jr. LeadScope: Software for Exploring Large Sets of Screening Data. J. Chem. Inf. Comput. Sci. 2000, 40, 1302–1314. [Google Scholar]
- Willet, P.; Gedeck, P. Visual and computational analysis of structure-activity relationships in high-throughput screening data. Curr. Opin. Chem. Biol 2001, 5, 389–395. [Google Scholar] [CrossRef]
- Hopfinger, A. J.; Reaka, A.; Venkatarangan, P.; Duca, J. S.; Wang, S. Construction of a Virtual High Throughput Screen by 4D-QSAR Analysis: Application to a Combinatorial Library of Glucose Inhibitors of Glycogen Phosphorylase b. J. Chem. Inf. Comput. Sci. 1999, 39, 1151–1160. [Google Scholar]
- Good, A. C.; Krystek, S. R.; Mason, J. S. High-througput and virtual screening: core lead discovery technologies move towards integration. Drug Discov. Today 2001, 5 (suppl.). [Google Scholar]
- Hawkins, D.M.; Young, S.S.; Rusinko, A., III. Analysis of a large structure-activity data set using recursive partitioning. Quant. Struct.-Act. Relat. 1997, 16, 296–302. [Google Scholar]
- Young, S. S. Sequential Screening. ScreenTech 2002 2002. [Google Scholar]
- Tropsha, A.; Zheng, W. Rational Principles of Focused Chemical Libraries Using Recursive Partitioning. Comb. Chem. High T. Scr. 2002, 5, 111–123. [Google Scholar]
- Lipinski, C. A. Poor aqueous solubility – an industry wide problem in ADME screening. 2002. Spotfire Users Europe Conference. http://www.spotfire.com/images/pdf/presentations2002/Chris_Lipinski_Lead_Identification_Europe.pdf.
- Singer, S. J.; Nicolson, G. L. The Fluid Mosaic Model of the Structure of Cell Membranes. Science 1972, 175, 720–731. [Google Scholar]
- Conradi, R. A.; Burton, P. S.; Borchardt, R. T. Physicochemical and Biological Factors that Influence a Drug's Cellular Permeability by Passive Diffusion. Methods. Princ. Med. Chem. 1996, 4, 233–252. [Google Scholar]
- (a) CLogP program was developed BioByte Corp., Claremont, CA.Viswanadhan, V. N.; Reddy, M. R.; Bacquet, R. J.; Erion, D. M. Assessment of Methods Used for Predicting Lipophilicity: Application to Nucleosides and Nucleoside Bases. J. Comput. Chem. 1993, 9, 1019–1026. [Google Scholar] [CrossRef] Klopman, G.; Li, J.-Y.; Wang, S.; Dimayuga, M. Computer Automated logP Calculations Based on an Extended Group Contribution Approach. J. Chem. Inf. Comput. Sci. 1994, 34, 752–781. [Google Scholar] [CrossRef] Wang, R.; Fu, Y.; Lai, L. A New Atom-Additive Method for Calculating Partition Coefficients. J. Chem. Inf. Comput. Sci. 1997, 37, 615–621. [Google Scholar] [CrossRef] Beck, B.; Breindl, A.; Clark, T. QM/NN QSPR Models with Error Estimation: Vapor Pressure and LogP. Chem. Inf. Comput. Sci. 2000, 40, 1046–1051. [Google Scholar] Egan, W. J.; Merz, K. M., Jr.; Baldwin, J. J. Prediction of Drug Absorption Using Multivariate Statistics. J. Med. Chem. 2000, 43, 3867–3877. [Google Scholar]
- Palm, K.; Stenberg, P.; Luthman, K.; Artursson, P. Polar Molecular Surface Properties Predict the Intestinal Absorption of Drugs in Humans. Pharm. Res. 1997, 14, 568–571. [Google Scholar]
- Palm, K.; Luthman, K.; Ungell, A.; Strandlund, G.; Beigi, F.; Lundahl, P.; Artursson, P. Evaluation of Dynamic Polar Molecular Surface Area as Predictor of Drug Absorption: Comparison with Other Computational and Experimental Predictors. J. Med. Chem. 1998, 41, 5382–5392. [Google Scholar] [CrossRef]
- Clark, D. E. Rapid Calculation of Polar Molecular Surface Area and Its Application to the Prediction of Transport Phenomena. 1. Prediction of Intestinal Absorption. J. Pharm. Sci. 1999, 88, 807–814. [Google Scholar]
- Kelder, J.; Grootenhuis, P. D. J.; Bayada, D. M.; Delbressine, L. P. C.; Ploemen, J. Polar Molecular Surface as a Dominating Determinant for Oral Absorption and Brain Penetration of Drugs. Pharm. Res. 1999, 16, 1514–1519. [Google Scholar]
- Stenberg, P.; Luthman, K.; Artursson, P. Prediction of Membrane Permeability to Peptides from Calculated Dynamic Molecular Surface Properties. Pharm. Res. 1999, 16, 205–212. [Google Scholar]
- Camenisch, G.; Alsenz, J.; van de Waterbeemd, H.; Folkers, G. Estimation of Permeability by Passive Diffusion through Caco-2 Cell Monolayers Using Drugs' Lipophilicity and Molecular Weight. Eur. J. Pharm. Sci. 1998, 6, 313–319. [Google Scholar]
- Camenisch, G.; Folkers, G.; van de Waterbeemd, H. Shape of Membrane Permeability-Lipophilicity Curves: Extension of Theoretical Models with an Aqueous Pore Pathway. Eur. J. Pharm. Sci. 1998, 6, 321–329. [Google Scholar]
- van de Waterbeemd, H.; Camenisch, G.; Folkers, G.; Raevsky, O. A. Estimation of Caco-2 Cell Permeability Using Calculated Molecular Descriptors. Quant. Struct.-Act. Relat. 1996, 15, 480–490. [Google Scholar] [CrossRef]
- Norinder, U.; Osterberg, T.; Artursson, P. Theoretical Calculation and Prediction of Caco-2 Cell Permeability Using MolSurf Parametrization and PLS Statistics. Pharm. Res. 1997, 14, 1786–1791. [Google Scholar]
- Norinder, U.; Osterberg, T.; Artursson, P. Theoretical Calculation and Prediction of Intestinal Absorption of Drugs in Humans Using MolSurf Parametrization and PLS Statistics. Eur. J. Pharm. Sci. 1999, 8, 49–56. [Google Scholar]
- Wessel, M. D.; Jurs, P. C.; Tolan, J. W.; Muskal, S. M. Prediction of Human Intestinal Absorption of Drug Compounds from Molecular Structure. J. Chem. Inf. Comput. Sci. 1998, 38, 726–735. [Google Scholar] [CrossRef]
- Lombardo, F.; Blake, J. F.; Curatolo, W. J. Computation of Brain-Blood Partitioning of Organic Solutes via Free Energy Calculations. J. Med. Chem. 1996, 39, 4750–4755. [Google Scholar] [CrossRef]
- Chikhale, E. G.; Ng, K.-Y.; Burton, P. S.; Borchardt, R. T. Hydrogen Bonding Potential as a Determinant of the in Vitro and in Situ Blood-Brain Barrier Permeability of Peptides. Pharm. Res. 1994, 11, 412–419. [Google Scholar]
- Young, R. C.; Mitchell, R. C.; Brown, T. H.; Ganellin, C. R.; Griffith, R.; Jones, M.; Rana, K. K.; Saunders, D.; Smith, I. R.; Sore, N. E.; Wilks, T. J. Development of a New Physicochemical Model for Brain Penetration and Its Application to the Design of Centrally acting H2 Receptor Histamine Antagonists. J. Med. Chem. 1988, 31, 656–671. [Google Scholar] [CrossRef]
- Seiler, P. Interconversion of Lipophilicities from Hydrocarbon/Water Systems into the Octanol/Water System. Eur. J. Med. Chem. 1974, 9, 473–479. [Google Scholar]
- van de Waterbeemd, H.; Kansy, M. Hydrogen-bonding Capacity and Brain Penetration. Chimia 1992, 46, 299–303. [Google Scholar]
- Abraham, M. H.; Chadha, H. S.; Mitchell, R. C. Hydrogen Bonding Factors that Influence the Distribution of Solutes between Blood and Brain. J. Pharm. Sci. 1994, 83, 1257–1268. [Google Scholar]
- Chadha, H. S.; Abraham, M. H.; Mitchell, R. C. Physicochemical analysis of the factors Governing Distribution of Solutes Between Blood and Brain. Bioorg. Med. Chem. Lett. 1994, 4, 2511–2516. [Google Scholar] [CrossRef]
- Abraham, M. H. Scales of Solutes Hydrogen-Bonding: Their Construction and Application to Physicochemical and Biochemical Processes. Chem. Soc. Rev. 1993, 22, 73–83. [Google Scholar] [CrossRef]
- Crivori, P.; Cruciani, G.; Carrupt, P. -A.; Testa, B. Predicting Blood-Brain Barrier Permeation from Three-Dimensional Molecular Structure. J. Med. Chem. 2000, 43, 11, 2204 -2216. [Google Scholar]
- Herve, F.; Urien, S.; Albengres, E.; Duche, J.-C.; Tillement, J. Drug Binding in Plasma. A Summary of Recent Trends in the Study of Drug and Hormone Binding. Clin. Pharmacokinet. 1994, 26, 44–58. [Google Scholar]
- Frostell-Karlsson, Å.; Remaeus, A.; Roos, H.; Andersson, K.; Borg, P.; Hamalainen, M.; Karlsson, R. Biosensor Analysis of the Interaction between Immobilized Human Serum Albumin and Drug Compounds for Prediction of Human Serum Albumin Binding Levels. J. Med. Chem. 2000, 43, 1986–1992. [Google Scholar]
- Colmenarejo, G.; Alvarez-Pedraglio, A.; Lavandera, J. –L. Cheminformatic Models To Predict Binding Affinities to Human Serum Albumin. J. Med. Chem. 2001, 44, 4370–4378. [Google Scholar]
- Carter, D. C.; He, X.-M. Structure of Human Serum Albumin. Science 1990, 249, 302–303. [Google Scholar]
- Roberts, S. A. High-throughput screening approaches for investigating drug metabolism and pharmacokinetics. Xenobiotica 2001, 31, 557–589. [Google Scholar]
- Watt, A. P.; Morrison, D.; Evans, D. C. Approaches to higher-throughput pharmacokinetics (HTPK) in drug discovery. Drug Discov. Today 2001, 5, 17–24. [Google Scholar]
- Keseruu, G. M.; Molnar, L. METAPRINT: A Metabolic Fingerprint. Application to Cassette Design for High-Throughput ADME Screening. J. Chem. Inf. Comput. Sci. 2002, 42, 437–444. [Google Scholar]
- Ekins, S.; Bravi, G.; Blinkley, S.; Gillespie, J. S.; Ring, B. J.; Wikel, J. H.; Wrighton, S. A. Three- and four-dimensional quantitative structure activity relationship analyses of cytochrome P-450 3A4 inhibitors. J. Pharm. Exp. Ther. 1999, 290, 429–438. [Google Scholar]
- Ekins, S.; Bravi, G.; Blinkley, S.; Gillespie, J. S.; Ring, B. J.; Wikel, J. H.; Wrighton, S. A. Three and four dimensional-quantitative structure activity relationship (3D/4D-QSAR) analyses of CYP2D6 inhibitors. Pharmacogenetics 1999, 9, 477–489. [Google Scholar]
- De Groot, M. J.; Vermeulen, N. P. Modeling the active sites of cytochrome P450s and glutathione S-transferases, two of the most important biotransformation enzymes. Drug Metab. Rev. 1997, 29, 747–799. [Google Scholar]
- Keseru, G. M. A. Virtual high throughput screen for high affinity cytochrome P450cam substrates. Implication for in silico prediction of drug metabolism. J. Comput.-Aided Mol. Des. 2001, 15, 649–657. [Google Scholar] [CrossRef]
- Darvas, F.; Marokhazi, S.; Kormos, P.; Kulkarni, P.; Kalasz, H.; Papp, Á. Drug Metabolism, Databases and High Throughput Testing During Drug Design and Development; Erhardt, P. W., Ed.; Blackwell Science: Cambridge, MA, 1999; pp. 237–270. [Google Scholar]
- Klopman, G.; Tu, M. Drug Metabolism, Databases and High Throughput Testing During Drug Design and Development; Erhardt, P. W., Ed.; Blackwell Science: Cambridge, MA, 1999; pp. 271–276. [Google Scholar]
- Testa, B.; Cruciani, G. Pharmacokinetic Optimization in Drug Research: Biological, Physicochemical and Computational Strategies; Testa, B., van de Waterbeemd, H., Folkers, G., Eds.; Verlag Helvetica Chimica Acta (VHCA); Wiley-VCH: Zurich; Weinheim, Germany, 2001; pp. 65–84. [Google Scholar]
- Duffy, J. C.; Cronin, M. T. D. Prediction of Half-Life of Non Steroidal Anti-Inflammatory Drugs. School of Pharmacy & Chemistry, Liverpool John Moores University, Byrom Street, Liverpool, L3 3AF, UK. http://www.pharm.uni-duesseldorf.de/QSAR/068.htm.
- Greene, N. Computer Software for Risk Assessment. J. Chem. Inf. Comput. Sci. 1997, 37, 148–150. [Google Scholar]
- Richard, A. M. Application of SAR methods to non-congeneric databases associated with carcinogenicity and mutagenicity: issues and approaches. Mutation Res. 1994, 305, 73–97. [Google Scholar]
- The Lethal Dose 50 (LD50) test involves the administration of a substance to a group of animals at increasing doses in order to determine the dose that kills 50 percent of the test subjects within a set time frame.
- Hall, L.; Kier, L.; Phipps, G. Structure-Activity Relationship Studies on the Toxicities of Benzene Derivatives I an Additivity Model. Environ. Toxicol. Chem. 1984, 3, 355–365. [Google Scholar] [CrossRef]
- Gute, B.; Basak, S. Predicting Acute Toxicity (LC50) of Benzene Derivatives Using Theoretical Molecular Descriptors: A Hierarchical QSAR Approach. SAR QSAR Environ. Res. 1997, 7, 117–131. [Google Scholar]
- IGC50 is the fifty percent growth inhibitory concentration against Tetrahymena pyriformis
- Cronin, M. T. D.; Gregory, B. W.; Schultz, T. W. Quantitative Structure-Activity Analyses of Nitrobenzene Toxicity to Tetrahymena pyriformis. Chem. Res. Toxicol. 1998, 11, 902–908. [Google Scholar] [CrossRef]
- Cronin, M. T. D.; Schultz, T. W. Development of Quantitative Structure-Activity Relationships for the Toxicity of Aromatic Compounds to Tetrahymena pyriformis: Comparative Assessment of the Methodologies. Chem. Res. Toxicol. 2001, 14, 1284–1295. [Google Scholar]
- Schultz, T. W.; Cronin, M. T. D. Response-Surface Analyses for Toxicity to Tetrahymena pyriformis: Reactive Carbonyl-Containing Aliphatic Chemicals. J. Chem. Inf. Comput. Sci. 1999, 39, 304–309. [Google Scholar] [CrossRef]
- Katritzky, A. R.; Tatham, D. B. Theoretical Descriptors for the Correlation of Aquatic Toxicity of Environmental Pollutants by Quantitative Structure-Toxicity Relationships. J. Chem. Inf. Comput. Sci. 2001, 41, 1162–1176. [Google Scholar]
- Sanderson, D. M.; Earnshaw, C. G.; Judson, P. N. Computer prediction of possible toxic action from chemical structure; the DEREK system. Human Experim. Toxicol. 1991, 10, 261–273. [Google Scholar]
- Ridings, J. E.; Barratt, M. D.; Cary, R.; Earnshaw, C. G.; Eggington, C. E.; Ellis, M. K.; Judson, P. N.; Langowski, J. J.; Marchant, C. A.; Payne, M. P.; Watson, W. P.; Yih, T. D. Computer prediction of possible toxic action from chemical structure: an update on the DEREK system. Toxicology 1996, 106, 267–279. [Google Scholar]
- Tonnelier, C. A. G.; Fox, J.; Judson, P.; Krause, P.; Pappas, N.; Patel, M. Representation of Chemical Structures in Knowledge-Based Systems: The StAR System. J. Chem. Inf. Comput. Sci. 1997, 37, 117–123. [Google Scholar] [CrossRef]
- http://www.chem.leeds.ac.uk/luk/derek/index.html.
- Benfenati, E.; Grasso, P.; Bruschi, M. Predictive Carcinogenicity: A Model for Aromatic Compounds, with Nitrogen-Containing Substituents, Based on Molecular Descriptors Using an Artificial Neural Network. J. Chem. Inf. Comput. Sci. 1999, 39, 1076–1080. [Google Scholar]
- Arenas, G. E.A.; Giralt, F. An Integrated SOM-Fuzzy ARTMAP Neural System for the Evaluation of Toxicity. J. Chem. Inf. Comput. Sci. 2002, 42, 343–359. [Google Scholar]
- Burden, F. R.; Winkler, D. A. A Quantitative Structure-Activity Relationships Model for the Acute Toxicity of Substituted Benzenes to Tetrahymena pyriformis Using Bayesian-Regularized Neural Networks. Chem. Res. Toxicol. 2000, 13, 436–440. [Google Scholar] [CrossRef]
- The companies providing in silico ADMET programs are: Advanced Chemistry Development ; Amedis Pharmaceuticals ; Accelrys ; ArQule ; Bioreason ; Chemical Computing Group ; Lhasa;Leadscope; Lion Bioscience ; Multicase ; Simulations Plus ; Tripos;
- Frawley, W.J.; Piatetsky-Shapiro, G.; Matheus, C. Knowledge Discovery. In Databases: An Overview. In Knowledge Discovery In Databases; Piatetsky-Shapiro, G., Frawley, W. J., Eds.; AAAI Press/MIT Press: Cambridge, MA, 1991; pp. 1–30. [Google Scholar]
- Wright, P. Knowledge Discovery In Databases: Tools and Techniques. http://www.acm.org/crossroads/xrds5-2/kdd.html.
- Brown, M. P. S.; Grundy, W. N.; Lin, D.; Cristianini, N.; Sugnet, C. W.; Furey, T. S.; Ares, M. J.; Haussler, D. Knowledge-based analysis of microarray gene expression data by using support vector machines. Proc. Nat. Acad. Sci. U.S.A. 2000, 97, 262–267. [Google Scholar] [CrossRef]
© 2002 by MDPI (http://www.mdpi.org). Reproduction is permitted for noncommercial purposes.
Share and Cite
Xu, J.; Hagler, A. Chemoinformatics and Drug Discovery. Molecules 2002, 7, 566-600. https://doi.org/10.3390/70800566
Xu J, Hagler A. Chemoinformatics and Drug Discovery. Molecules. 2002; 7(8):566-600. https://doi.org/10.3390/70800566
Chicago/Turabian StyleXu, Jun, and Arnold Hagler. 2002. "Chemoinformatics and Drug Discovery" Molecules 7, no. 8: 566-600. https://doi.org/10.3390/70800566
APA StyleXu, J., & Hagler, A. (2002). Chemoinformatics and Drug Discovery. Molecules, 7(8), 566-600. https://doi.org/10.3390/70800566




