Strategies for Metabolic Engineering of Escherichia coli for β-Carotene Biosynthesis
Abstract
1. Introduction
| Producer Type | Strain | Compounds | Ref. |
|---|---|---|---|
| Natural producers | Haematococcus pluvialis | Astaxanthin | [8] |
| Chlorella vulgaris | Astaxanthin | [9] | |
| Dunaliella salina | β-carotene | [10] | |
| Blakeslea trispora | lycopene, β-carotene | [11] | |
| Rhodotorula mucilaginosa | β-carotene, astaxanthin | [12] | |
| Phaffia rhodozyma | β-carotene, astaxanthin | [13,14,15] | |
| Kluyveromyces marxianus | Astaxanthin | [16] | |
| Rhodococcus sp. | lycopene, γ-carotene | [17,18] | |
| Heterologous producers | Saccharomyces cerevisiae | lycopene, β-carotene, astaxanthin | [20,21,22] |
| Yarrowia lipolytica | lycopene, β-carotene, astaxanthin | [23,24,25,26] | |
| Komagataella phaffii | lycopene, β-carotene | [27,28] | |
| Escherichia coli | lycopene, β-carotene, astaxanthin | [29,30,31,32] |
2. Metabolic Engineering Strategies for β-Carotene Production in E. coli
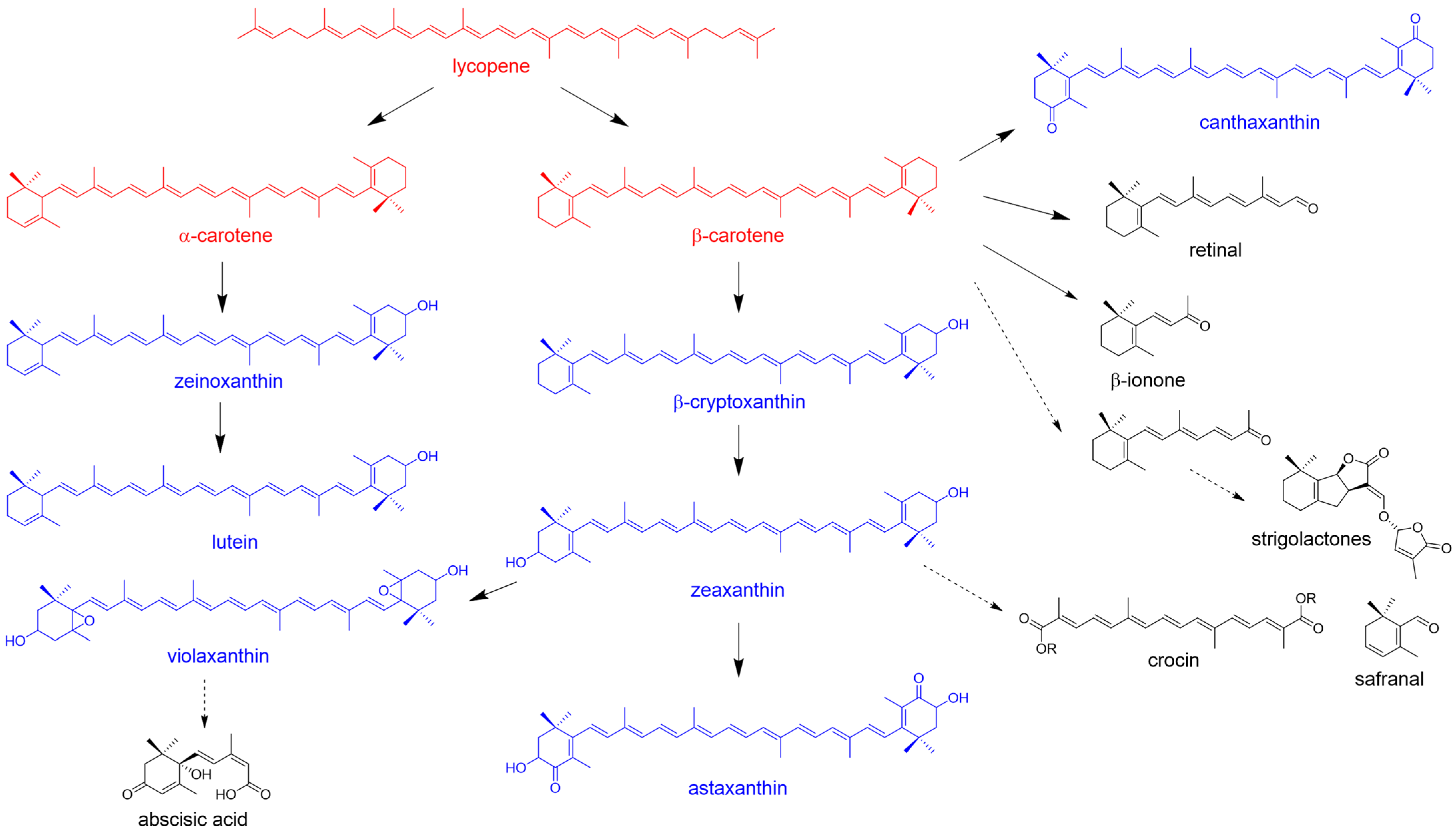
2.1. The β-Carotene Biosynthetic Pathway
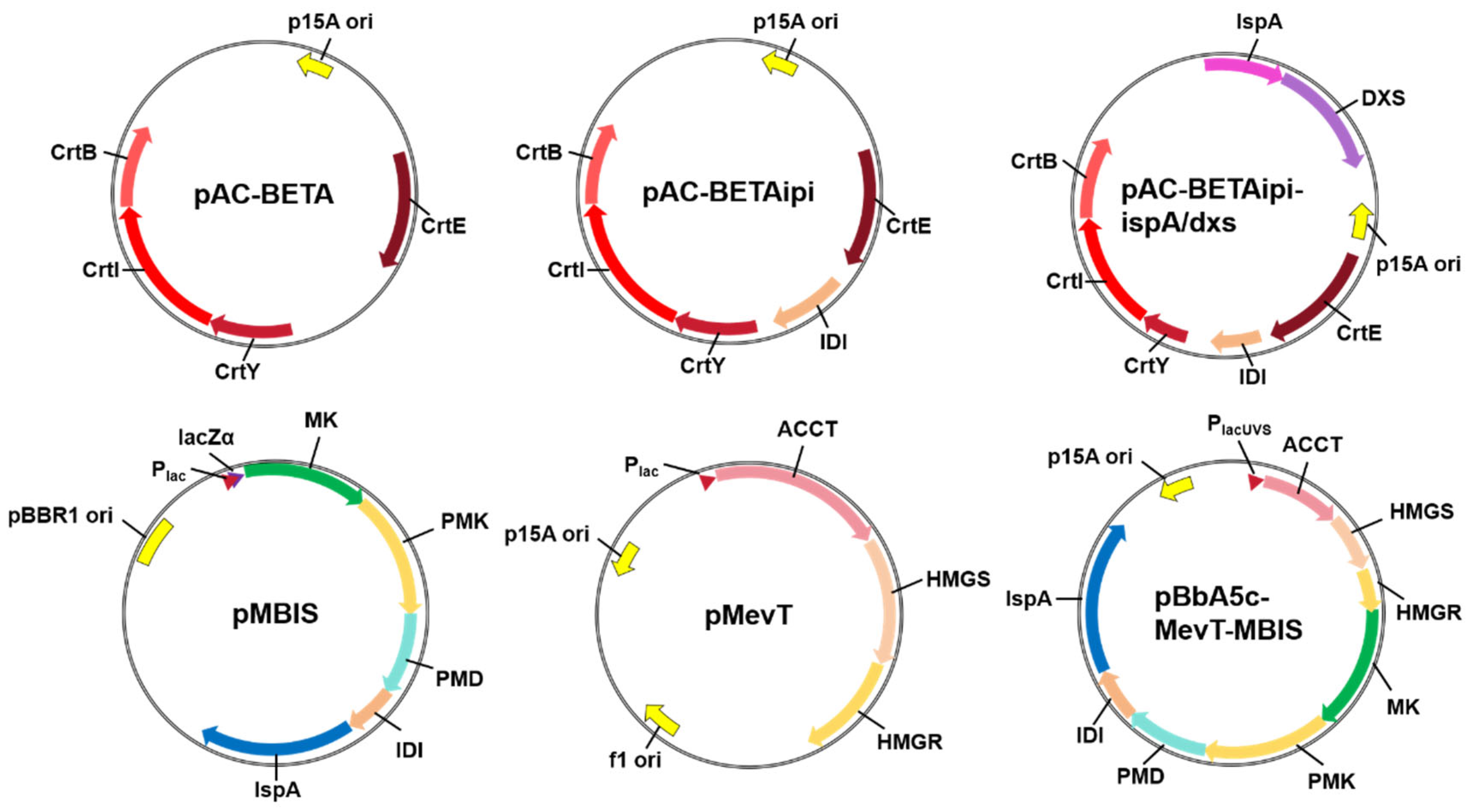
2.2. Introduction of β-Carotene Biosynthetic Genes
2.3. Manipulation of the Endogenous MEP Pathway
2.4. Introduction of Exogenous MVA Pathway
2.5. Modification of the Central Metabolic Module
2.6. Balancing the Gene Expression Levels
2.7. Engineering the Membrane Architecture
2.8. Other Factors Affecting β-Carotene Production
| Chassis | Bioreactor | Titer (g/L) | Primary Metabolic Engineering Strategies | Specific Fermentation Conditions | Ref. |
|---|---|---|---|---|---|
| MG1655 | 3-L fermenter | 3.93 | MVA pathway, membrane engineering, dynamic regulation of mvaE expression | Three-stage semi-continuous fermentation | [29] |
| BL21(DE3) | 5-L fermenter | 3.20 | Optimized MEP pathway with exogenous dxs and gpps2 genes, hybrid MVA pathway. | Glycerol feeding | [41] |
| MG1655 | 5-L fermenter | 2.58 | Deletion of gdhA, zwf, phtHIcrr, and yjgB, overexpression of nadK on a low-copy number plasmid. | Fed-batch fermentation, modified minimal medium with trace metal solution | [61] |
| DH5α | 7-L fermenter | 2.47 | Co-expression of pT-DHB (contains downstream genes for β-carotene biosynthesis) and pS-NA (contains the exogenous MVA pathway) | Chemically defined medium with amino acid supplementation, glycerol feeding | [80] |
| ATCC 8739 | 7-L fermenter | 2.1 | Combined engineering of the MEP, β-carotene biosynthesis, and central metabolic modules. | Feeding of glycerol, peptone, yeast extract, and MgSO4·7H2O | [31] |
| BW25113 | 5-L fermenter | 1.9 | Optimizing CrtY expression using an oligo-linker mediated DNA assembly (OLMA) library | Fed-batch fermentation | [81] |
| DH5α | 3.4-L fermenter | 0.66 | Co-expression of pT-DHB and pS-NA | Feeding glycerol, yeast extract, MgSO4·7H2O, and thiamine | [82] |
| DH5α | shake flask | 0.50 | Co-expression of pT-DHB and pSSN12Didi (contains downstream genes of the MVA pathway) | Fermentation at 29 °C in 2YT medium for 144 h, addition of 16.5 mM mevalonate and glycerol | [44] |
3. Significance of β-Carotene Production in E. coli
4. Conclusions and Prospect
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Giani, M.; Montoyo-Pujol, Y.G.; Peiró, G.; Martínez-Espinosa, R.M. Halophilic carotenoids and breast cancer: From salt marshes to biomedicine. Mar. Drugs 2021, 19, 564. [Google Scholar] [CrossRef]
- Dawson, M.I. The importance of vitamin A in nutrition. Curr. Pharm. Des. 2000, 6, 311–325. [Google Scholar] [CrossRef]
- Trisconi, M.J.; Campos-Gimenez, E.; Jaudzems, G.; Dowell, D. Determination of vitamin A in infant formula and adult nutritionals by UPLC-UV: First action 2011.07. J. AOAC Int. 2012, 95, 301–306. [Google Scholar] [CrossRef][Green Version]
- Martín, J.F.; Gudiña, E.; Barredo, J.L. Conversion of beta-carotene into astaxanthin: Two separate enzymes or a bifunctional hydroxylase-ketolase protein? Microb. Cell Fact. 2008, 7, 3. [Google Scholar] [CrossRef]
- Ruther, A.; Misawa, N.; Böger, P.; Sandmann, G. Production of zeaxanthin in Escherichia coli transformed with different carotenogenic plasmids. Appl. Microbiol. Biotechnol. 1997, 48, 162–167. [Google Scholar] [CrossRef]
- Zhou, A.; Kane, A.; Wu, S.; Wang, K.; Santiago, M.; Ishiguro, Y.; Yoneyama, K.; Palayam, M.; Shabek, N.; Xie, X.; et al. Evolution of interorganismal strigolactone biosynthesis in seed plants. Science 2025, 387, eadp0779. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.Y.; Chen, G.-T.E.; Braguy, J.; Al-Babili, S. Distinguishing the functions of canonical strigolactones as rhizospheric signals. Trends Plant Sci. 2024, 29, 925–936. [Google Scholar] [CrossRef]
- Mularczyk, M.; Michalak, I.; Marycz, K. Astaxanthin and other nutrients from Haematococcus pluvialis—Multifunctional applications. Mar. Drugs 2020, 18, 459. [Google Scholar] [CrossRef] [PubMed]
- Seyfabadi, J.; Ramezanpour, Z.; Amini Khoeyi, Z. Protein, fatty acid, and pigment content of Chlorella vulgaris under different light regimes. J. Appl. Phycol. 2011, 23, 721–726. [Google Scholar] [CrossRef]
- Hejazi, M.A.; Holwerda, E.; Wijffels, R.H. Milking microalga Dunaliella salina for β-carotene production in two-phase bioreactors. Biotechnol. Bioeng. 2004, 85, 475–481. [Google Scholar] [CrossRef]
- Mantzouridou, F.; Naziri, E.; Tsimidou, M.Z. Industrial glycerol as a supplementary carbon source in the production of β-carotene by Blakeslea trispora. J. Agric. Food. Chem. 2008, 56, 2668–2675. [Google Scholar] [CrossRef] [PubMed]
- Rodrigues, T.V.D.; Contessa, C.R.; Burkert, C.A.V.; de Medeiros Burkert, J.F. Evaluation of light-emitting diodes applied to Rhodotorula mucilaginosa to produce carotenoids and lipids. Food Bioprocess Technol. 2024, 17, 3267–3280. [Google Scholar] [CrossRef]
- Zheng, Y.G.; Hu, Z.C.; Wang, Z.; Shen, Y.C. Large-scale production of astaxanthin by Xanthophyllomyces dendrorhous. Food Bioprod. Process 2006, 84, 164–166. [Google Scholar] [CrossRef]
- Gupta, I.; Adin, S.N.; Panda, B.P.; Mujeeb, M. Β-carotene-production methods, biosynthesis from Phaffia rhodozyma, factors affecting its production during fermentation, pharmacological properties: A review. Biotechnol. Appl. Biochem. 2022, 69, 2517–2529. [Google Scholar] [CrossRef]
- Mussagy, C.U.; Farias, F.O.; Bila, N.M.; Giannini, M.J.S.M.; Pereira, J.F.B.; Santos-Ebinuma, V.C.; Pessoa, A., Jr. Recovery of β-carotene and astaxanthin from Phaffia rhodozyma biomass using aqueous solutions of cholinium-based ionic liquids. Sep. Purif. Technol. 2022, 290, 120852. [Google Scholar] [CrossRef]
- Lin, Y.-J.; Chang, J.-J.; Lin, H.-Y.; Thia, C.; Kao, Y.-Y.; Huang, C.-C.; Li, W.-H. Metabolic engineering a yeast to produce astaxanthin. Bioresour. Technol. 2017, 245, 899–905. [Google Scholar] [CrossRef]
- Ichiyama, S.; Shimokata, K.; Tsukamura, M. Carotenoid pigments of genus Rhodococcus. Microbiol. Immunol. 1989, 33, 503–508. [Google Scholar] [CrossRef] [PubMed]
- Tao, L.; Cheng, Q. Novel β-carotene ketolases from non-photosynthetic bacteria for canthaxanthin synthesis. Mol. Genet. Genom. 2004, 272, 530–537. [Google Scholar] [CrossRef]
- Zhou, T.; Park, Y.-K.; Ledesma-Amaro, R. Advances in the biosynthesis of β-carotene and its derivatives in yeast. Bioresour. Technol. 2025, 435, 132936. [Google Scholar] [CrossRef] [PubMed]
- Lin, P.; Zhang, L.; Du, G.; Chen, J.; Zhang, J.; Peng, Z. Construction of Saccharomyces cerevisiae platform strain for the biosynthesis of carotenoids and apocarotenoids. J. Agric. Food. Chem. 2025, 73, 9187–9196. [Google Scholar] [CrossRef]
- Shi, B.; Ma, T.; Ye, Z.; Li, X.; Huang, Y.; Zhou, Z.; Ding, Y.; Deng, Z.; Liu, T. Systematic metabolic engineering of Saccharomyces cerevisiae for lycopene overproduction. J. Agric. Food. Chem. 2019, 67, 11148–11157. [Google Scholar] [CrossRef]
- López, J.; Cataldo, V.F.; Peña, M.; Saa, P.A.; Saitua, F.; Ibaceta, M.; Agosin, E. Build your bioprocess on a solid strain—β-carotene production in recombinant Saccharomyces cerevisiae. Front. Bioeng. Biotechnol. 2019, 7, 171. [Google Scholar] [CrossRef]
- Ma, Y.; Liu, N.; Greisen, P.; Li, J.; Qiao, K.; Huang, S.; Stephanopoulos, G. Removal of lycopene substrate inhibition enables high carotenoid productivity in Yarrowia lipolytica. Nat. Commun. 2022, 13, 572. [Google Scholar] [CrossRef] [PubMed]
- Zhao, C.; Gao, Q.; Chen, J.; Wei, L.; Imanaka, T.; Hua, Q. Metabolomic changes and metabolic responses to expression of heterologous biosynthetic genes for lycopene production in Yarrowia lipolytica. J. Biotechnol. 2017, 251, 174–185. [Google Scholar] [CrossRef]
- Xu, S.; Zhang, X.; Zhang, Y.; Li, Q.; Ji, L.; Cheng, H. Concomitant production of erythritol and β-carotene by engineered Yarrowia lipolytica. J. Agric. Food. Chem. 2023, 71, 11567–11578. [Google Scholar] [CrossRef] [PubMed]
- Kildegaard, K.R.; Adiego-Pérez, B.; Doménech Belda, D.; Khangura, J.K.; Holkenbrink, C.; Borodina, I. Engineering of Yarrowia lipolytica for production of astaxanthin. Synth. Syst. Biotechnol. 2017, 2, 287–294. [Google Scholar] [CrossRef] [PubMed]
- Zhow, W.; Ling, R.-J.; Yang, Y.-C.; Hou, S.-T.; Wang, F.-Q.; Gao, B.; Wei, D.-Z. Engineering Komarataella phaffii to produce lycopepe sustainably from glucose or methanol. Metab. Eng. 2025, 90, 141–153. [Google Scholar] [CrossRef]
- Zhang, X.; Wang, D.; Duan, Y.; Zheng, X.; Lin, Y.; Liang, S. Production of lycopene by metabolically engineered Pichia pastoris. Biosci. Biotechnol. Biochem. 2020, 84, 463–470. [Google Scholar] [CrossRef]
- Ji, Y.; Yue, J.; Huang, Y.; Li, R.; Zhong, S.; Jiang, X.; Zhang, S.; Zou, Y.; Deng, A.; Qiao, S. A combinatorial stress-alleviation strategy for efficient β-carotene production in Escherichia coli. Bioresour. Technol. 2026, 439, 133336. [Google Scholar] [CrossRef]
- Kim, Y.-S.; Lee, J.-H.; Kim, N.-H.; Yeom, S.-J.; Kim, S.-W.; Oh, D.-K. Increase of lycopene production by supplementing auxiliary carbon sources in metabolically engineered Escherichia coli. Appl. Microbiol. Biotechnol. 2011, 90, 489–497. [Google Scholar] [CrossRef]
- Zhao, J.; Li, Q.Y.; Sun, T.; Zhu, X.N.; Xu, H.T.; Tang, J.L.; Zhang, X.L.; Ma, Y.H. Engineering central metabolic modules of Escherichia coli for improving β-carotene production. Metab. Eng. 2013, 17, 42–50. [Google Scholar] [CrossRef]
- Gong, Z.; Wang, H.; Tang, J.; Bi, C.; Li, Q.; Zhang, X. Coordinated expression of astaxanthin biosynthesis genes for improved astaxanthin production in Escherichia coli. J. Agric. Food. Chem. 2020, 68, 14917–14927. [Google Scholar] [CrossRef]
- Ridder, E.D.; Vandamme, P.; Willems, A. Carotenoid biosynthesis in bacteria: The crt gene products and their functional roles in the carotenogenic pathways. Crit. Rev. Microbiol. 2026, 52, 64–83. [Google Scholar] [CrossRef]
- Stra, A.; Almarwaey, L.O.; Alagoz, Y.; Moreno, J.C.; Al-Babili, S. Carotenoid metabolism: New insights and synthetic approaches. Front. Plant Sci. 2023, 13, 1072061. [Google Scholar] [CrossRef]
- Li, M.; Hou, F.; Wu, T.; Jiang, X.; Li, F.; Liu, H.; Xian, M.; Zhang, H. Recent advances of metabolic engineering strategies in natural isoprenoid production using cell factories. Nat. Prod. Rep. 2020, 37, 80–99. [Google Scholar] [CrossRef]
- Misawa, N.; Shimada, H. Metabolic engineering for the production of carotenoids in non-carotenogenic bacteria and yeasts. J. Biotechnol. 1998, 59, 169–181. [Google Scholar] [CrossRef] [PubMed]
- Misawa, N.; Satomi, Y.; Kondo, K.; Yokoyama, A.; Kajiwara, S.; Saito, T.; Ohtani, T.; Miki, W. Structure and functional analysis of a marine bacterial carotenoid biosynthesis gene cluster and astaxanthin biosynthetic pathway proposed at the gene level. J. Bacteriol. 1995, 177, 6575–6584. [Google Scholar] [CrossRef] [PubMed]
- Nishizaki, T.; Tsuge, K.; Itaya, M.; Dio, N.; Yanagawa, H. Metabolic engineering of carotenoid biosynthesis in Escherichia coli by ordered gene assembly in Bacillus subtilis. Appl. Environ. Microbiol. 2006, 73, 1355–1361. [Google Scholar] [CrossRef]
- Xu, X.; Tian, L.; Xu, J.; Xie, C.; Jiang, L.; Huang, H. Analysis and expression of the carotenoid biosynthesis genes from Deinococcus wulumuqiensis R12 in engineered Escherichia coli. AMB Expr. 2018, 8, 94. [Google Scholar] [CrossRef] [PubMed]
- Cunningham, F.X., Jr.; Pogson, B.; Sun, Z.; McDonald, K.A.; DellaPenna, D.; Gantt, E. Functional analysis of the beta and epsilon lycopene cyclase enzymes of arabidopsis reveals a mechanism for control of cyclic carotenoid formation. Plant Cell 1996, 8, 1613–1626. [Google Scholar] [CrossRef]
- Yang, J.M.; Guo, L.Z. Biosynthesis of β-carotene in engineered E. coli using the MEP and MVA pathways. Microb. Cell Fact. 2014, 13, 160. [Google Scholar] [CrossRef] [PubMed]
- Cunningham, F.X., Jr.; Gantt, E. A study in scarlet: Enzymes of ketocarotenoid biosynthesis in the flowers of Adonis aestivalis. Plant J. 2005, 41, 478–492. [Google Scholar] [CrossRef]
- Wu, S.; Ma, X.; Zhou, A.; Valenzuela, A.; Zhou, K.; Li, Y. Establishment of strigolactone-producing bacterium-yeast consortium. Sci. Adv. 2021, 7, eabh4048. [Google Scholar] [CrossRef]
- Yoon, S.H.; Park, H.M.; Kim, J.E.; Lee, S.H.; Choi, M.S.; Kim, J.Y.; Oh, D.K.; Keasling, J.D.; Kim, S.W. Increased β-carotene production in recombinant Escherichia coli harboring an engineered isoprenoid precursor pathway with mevalonate addition. Biotechnol. Prog. 2007, 23, 599–605. [Google Scholar] [CrossRef]
- Yoon, S.H.; Lee, S.H.; Das, A.; Ryu, H.K.; Jang, H.J.; Kim, J.Y.; Oh, D.K.; Keasling, J.D.; Kim, S.W. Combinatorial expression of bacterial whole mevalonate pathway for the production of β-carotene in E. coli. J. Biotechnol. 2009, 140, 218–226. [Google Scholar] [CrossRef]
- Kim, S.W.; Kim, J.B.; Jung, W.H.; Kim, J.H.; Jung, J.K. Over-production of β-carotene from metabolically engineered Escherichia coli. Biotechnol. Lett. 2006, 28, 897–904. [Google Scholar] [CrossRef]
- Chen, H.-H.; Liang, M.-H.; Ye, Z.-W.; Zhu, Y.-H.; Jiang, J.-G. Engineering the β-carotene metabolic pathway of microalgae Dunaliella to confirm its carotenoid synthesis pattern in comparison to bacteria and plants. Microbiol. Spectr. 2023, 11, e04361-22. [Google Scholar] [CrossRef] [PubMed]
- Verdoes, J.C.; Krubasik, P.; Sandmann, G.; van Ooyen, A.J.J. Isolation and functional characterisation of a novel type of carotenoid biosynthetic gene from Xanthophyllomyces dendrorhous. Mol. Gen. Genet. 1999, 262, 453–461. [Google Scholar] [CrossRef] [PubMed]
- Albrecht, M.; Misawa, N.; Sandmann, G. Metabolic engineering of the terpenoid biosynthetic pathway of Escherichia coli for production of the carotenoids β-carotene and zeaxanthin. Biotechnol. Lett. 1999, 21, 791–795. [Google Scholar] [CrossRef]
- Yuan, L.Z.; Rouvière, P.E.; LaRossa, R.A.; Suh, W. Chromosomal promoter replacement of the isoprenoid pathway for enhancing carotenoid production in E. coli. Metab. Eng. 2006, 8, 79–90. [Google Scholar] [CrossRef]
- Kim, S.W.; Keasling, J.D. Metabolic engineering of the nonmevalonate isopentenyl diphosphate synthesis pathway in Escherichia coli enhances lycopene production. Biotechnol. Bioeng. 2001, 72, 408–415. [Google Scholar] [CrossRef] [PubMed]
- Harker, M.; Bramley, P.M. Expression of prokaryotic 1-deoxy-d-xylulose-5-phosphatases in Escherichia coli increases carotenoid and ubiquinone biosynthesis. FEBS Lett. 1999, 448, 115–119. [Google Scholar] [CrossRef] [PubMed]
- Kim, M.J.; Noh, M.H.; Woo, S.; Lim, H.G.; Jung, G.Y. Enhanced lycopene production in Escherichia coli by expression of two mep pathway enzymes from Vibrio sp. Dhg. Catalysts 2019, 9, 1003. [Google Scholar] [CrossRef]
- Martin, V.J.J.; Pitera, D.J.; Withers, S.T.; Newman, J.D.; Keasling, J.D. Engineering a mevalonate pathway in Escherichia coli for production of terpenoids. Nat. Biotechnol. 2003, 21, 796–802. [Google Scholar] [CrossRef]
- Yoon, S.H.; Lee, Y.M.; Kim, J.E.; Lee, S.H.; Lee, J.H.; Kim, J.Y.; Jung, K.H.; Shin, Y.C.; Keasling, J.D.; Kim, S.W. Enhanced lycopene production in Escherichia coli engineered to synthesize isopentenyl diphosphate and dimethylallyl diphosphate from mevalonate. Biotechnol. Bioeng. 2006, 94, 1025–1032. [Google Scholar] [CrossRef]
- Yang, J.M.; Xian, M.; Su, S.Z.; Zhao, G.; Nie, Q.J.; Jiang, X.L.; Zheng, Y.N.; Liu, W. Enhancing production of bio-isoprene using hybrid MVA pathway and isoprene synthase in E. coli. PLoS ONE 2012, 7, e33509. [Google Scholar] [CrossRef] [PubMed]
- Duong, C. Enhanced β-carotene biosynthesis in recombinant Escherichia coli harboring the bottom portion of mevalonate pathway of Enterococcus faecium VTCC-B-935 isolated in Vietnam. J. Appl. Biotechnol. Bioeng. 2017, 3, 374–379. [Google Scholar]
- Ye, L.J.; Zhang, C.Z.; Bi, C.G.; Li, Q.Y.; Zhang, X.L. Combinatory optimization of chromosomal integrated mevalonate pathway for β-carotene production in Escherichia coli. Microb. Cell Fact. 2016, 15, 202. [Google Scholar] [CrossRef]
- Wei, Y.L.; Mohsin, A.; Hong, Q.; Guo, M.J.; Fang, H.Q. Enhanced production of biosynthesized lycopene via heterogenous mva pathway based on chromosomal multiple position integration strategy plus plasmid systems in Escherichia coli. Bioresour. Technol. 2018, 250, 382–389. [Google Scholar] [CrossRef]
- Zhou, Y.; Nambou, K.; Wei, L.J.; Cao, J.J.; Imanaka, T.; Hua, Q. Lycopene production in recombinant strains of Escherichia coli is improved by knockout of the central carbon metabolism gene coding for glucose-6-phosphate dehydrogenase. Biotechnol. Lett. 2013, 35, 2137–2145. [Google Scholar] [CrossRef]
- Wu, Y.; Yan, P.; Li, Y.; Liu, X.; Wang, Z.; Chen, T.; Zhao, X. Enhancing β-carotene production in Escherichia coli by perturbing central carbon metabolism and improving the NADPH supply. Front. Bioeng. Biotechnol. 2020, 8, 585. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Lin, Z.; Huang, C.; Zhang, Y.; Wang, Z.; Tang, Y.; Chen, T.; Zhao, X. Metabolic engineering of Escherichia coli using CRISPR-Cas9 mediated genome editing. Metab. Eng. 2015, 31, 13–21. [Google Scholar] [CrossRef]
- Toya, Y.; Ishii, N.; Nakahigashi, K.; Hirasawa, T.; Soga, T.; Tomita, M.; Shimizu, K. 13C-metabolic flux analysis for batch culture of Escherichia coli and its pyk and pgi gene knockout mutants based on mass isotopomer distribution of intracellular metabolites. Biotechnol. Prog. 2010, 26, 975–992. [Google Scholar] [CrossRef]
- Sun, T.; Miao, L.T.; Li, Q.Y.; Dai, G.P.; Lu, F.P.; Liu, T.; Zhang, X.L.; Ma, Y.H. Production of lycopene by metabolically-engineered Escherichia coli. Biotechnol. Lett. 2014, 36, 1515–1522. [Google Scholar] [CrossRef]
- Raghavan, I.; Juman, R.; Wang, Z.Q. The non-mevalonate pathway requires a delicate balance of intermediates to maximize terpene production. Appl. Microbiol. Biotechnol. 2024, 108, 245. [Google Scholar] [CrossRef] [PubMed]
- Bongers, M.; Chrysanthopoulos, P.K.; Behrendorff, J.B.Y.H.; Hodson, M.P.; Vickers, C.E.; Nielsen, L.K. Systems analysis of methylerythritol-phosphate pathway flux in E. coli: Insights into the role of oxidative stress and the validity of lycopene as an isoprenoid reporter metabolite. Microb. Cell Fact. 2015, 14, 193. [Google Scholar] [CrossRef] [PubMed]
- Rinaldi, M.A.; Ferraz, C.A.; Scrutton, N.S. Alternative metabolic pathways and strategies to high-titre terpenoid production in Escherichia coli. Nat. Prod. Rep. 2022, 39, 90–118. [Google Scholar] [CrossRef]
- Li, Q.Y.; Fan, F.Y.; Gao, X.; Yang, C.; Bi, C.H.; Tang, J.L.; Liu, T.; Zhang, X.L. Balanced activation of IspG and IspH to eliminate MEP intermediate accumulation and improve isoprenoids production in Escherichia coli. Metab. Eng. 2017, 44, 13–21. [Google Scholar] [CrossRef]
- You, S.P.; Yin, Q.D.A.; Zhang, J.Y.; Zhang, C.Y.; Qi, W.; Gao, L.; Tao, Z.P.; Su, R.X.; He, Z.M. Utilization of biodiesel by-product as substrate for high-production of β-farnesene via relatively balanced mevalonate pathway in Escherichia coli. Bioresour. Technol. 2017, 243, 228–236. [Google Scholar] [CrossRef] [PubMed]
- Kang, W.; Ma, T.; Liu, M.; Qu, J.L.; Liu, Z.J.; Zhang, H.W.; Shi, B.; Fu, S.; Ma, J.C.; Lai, L.T.F.; et al. Modular enzyme assembly for enhanced cascade biocatalysis and metabolic flux. Nat. Commun. 2019, 10, 4248. [Google Scholar] [CrossRef]
- Li, D.; Li, Y.; Xu, J.Y.; Li, Q.Y.; Tang, J.L.; Jia, S.R.; Bi, C.H.; Dai, Z.B.; Zhu, X.N.; Zhang, X.L. Engineering Crtw and Crtz for improving biosynthesis of astaxanthin in Escherichia coli. Chin. J. Nat. Med. 2020, 18, 666–676. [Google Scholar] [CrossRef]
- Yang, D.; Park, S.Y.; Lee, S.Y. Production of rainbow colorants by metabolically engineered Escherichia coli. Adv. Sci. 2021, 8, e2100743. [Google Scholar] [CrossRef]
- Ye, L.J.; He, P.; Li, Q.Y.; Zhang, X.L.; Bi, C.H. Type IIs restriction based combinatory modulation technique for metabolic pathway optimization. Microb. Cell Fact. 2017, 16, 47. [Google Scholar] [CrossRef]
- Wu, T.; Jiang, J.; Zhang, H.; Liu, J.; Ruan, H. Transcending membrane barriers: Advances in membrane engineering to enhance the production capacity of microbial cell factories. Microb. Cell Fact. 2024, 23, 154. [Google Scholar] [CrossRef]
- Wu, T.; Ye, L.J.; Zhao, D.D.; Li, S.W.; Li, Q.Y.; Zhang, B.L.; Bi, C.H.; Zhang, X.L. Membrane engineering—A novel strategy to enhance the production and accumulation of β-carotene in Escherichia coli. Metab. Eng. 2017, 43, 85–91. [Google Scholar] [CrossRef] [PubMed]
- Wu, T.; Ye, L.; Zhao, D.; Li, S.; Li, Q.; Zhang, B.; Bi, C. Engineering membrane morphology and manipulating synthesis for increased lycopene accumulation in Escherichia coli cell factories. 3 Biotech 2018, 8, 269. [Google Scholar] [CrossRef] [PubMed]
- Wu, T.; Li, S.W.; Ye, L.J.; Zhao, D.D.; Fan, F.Y.; Li, Q.Y.; Zhang, B.L.; Bi, C.H.; Zhang, X.L. Engineering an artificial membrane vesicle trafficking system (AMVTS) for the excretion of β-carotene in Escherichia coli. ACS Synth. Biol. 2019, 8, 1037–1046. [Google Scholar] [CrossRef]
- Kim, J.; Kong, M.K.; Lee, S.Y.; Lee, P.C. Carbon sources-dependent carotenoid production in metabolically engineered Escherichia coli. World J. Microbiol. Biotechnol. 2010, 26, 2231–2239. [Google Scholar] [CrossRef]
- Liu, N.; Liu, B.; Wang, G.Y.; Soong, Y.H.V.; Tao, Y.; Liu, W.F.; Xie, D.M. Lycopene production from glucose, fatty acid and waste cooking oil by metabolically engineered Escherichia coli. Biochem. Eng. J. 2020, 155, 107488. [Google Scholar] [CrossRef]
- Nam, H.-K.; Choi, J.-G.; Lee, J.H.; Kim, S.-W.; Oh, D.-K. Increase in the production of β-carotene in recombinant Escherichia coli cultured in a chemically defined medium supplemented with amino acids. Biotechnol. Lett. 2013, 35, 265–271. [Google Scholar] [CrossRef]
- Jin, Y.; Han, L.; Zhang, S.; Li, S.; Liu, W.; Tao, Y. Construction of high-yield strain by optimizing lycopene cyclase for β-carotene production. Chin. J. Biotechnol. 2017, 33, 1814–1826. [Google Scholar]
- Kim, J.H.; Kim, S.W.; Nguyen, D.Q.A.; Li, H.; Kim, S.B.; Seo, Y.G.; Yang, J.K.; Chung, I.Y.; Kim, D.H.; Kim, C.J. Production of β-carotene by recombinant Escherichia coli with engineered whole mevalonate pathway in batch and fed-batch cultures. Biotechnol. Bioprocess Eng. 2009, 14, 559–564. [Google Scholar] [CrossRef]
- Ben-Amotz, A.; Avron, M. The wavelength dependence of massive carotene synthesis in Dunaliella bardawil (chlorophyceae). J. Phycol. 1989, 25, 175–178. [Google Scholar] [CrossRef]
- Bruno, M.; Al-Babili, S. On the substrate specificity of the rice strigolactone biosynthesis enzyme DWARF27. Planta 2016, 243, 1429–1440. [Google Scholar] [CrossRef]
- Zhang, M.; Gong, Z.K.; Tang, J.L.; Lu, F.P.; Li, Q.Y.; Zhang, X.L. Improving astaxanthin production in Escherichia coli by co-utilizing Crtz enzymes with different substrate preference. Microb. Cell Fact. 2022, 21, 71. [Google Scholar] [CrossRef]
- Tao, L.; Wilczek, J.; Odom, J.M.; Cheng, Q. Engineering a beta-carotene ketolase for astaxanthin production. Metab. Eng. 2006, 8, 526–531. [Google Scholar] [CrossRef]
- Hussain, M.H.; Hong, Q.; Zaman, W.Q.; Mohsin, A.; Wei, Y.; Zhang, N.; Fang, H.; Wang, Z.; Hang, H.; Zhuang, Y.; et al. Rationally optimized generation of integrated Escherichia coli with stable and high yield lycopene biosynthesis from heterologous mevalonate (MVA) and lycopene expression pathways. Synth. Syst. Biotechnol. 2021, 6, 85–94. [Google Scholar] [CrossRef]
- Roth, J.H.; Ward, V.C.A. Production of astaxanthin using CBFD1/HFBD1 from Adonis aestivalis and the isopentenol utilization pathway in Escherichia coli. Bioengineering 2023, 10, 1033. [Google Scholar] [CrossRef] [PubMed]
- Li, X.-R.; Tian, G.-Q.; Shen, H.-J.; Liu, J.-Z. Metabolic engineering of Escherichia coli to produce zeaxanthin. J. Ind. Microbiol. Biotechnol. 2015, 42, 627–636. [Google Scholar] [CrossRef]
- Neuman, H.; Galpaz, N.; Cunningham, F.X.; Zamir, D.; Hirschberg, J. The tomato mutation nxd1 reveals a gene necessary forneoxanthin biosynthesis and demonstrates that violaxanthinis a sufficient precursor for abscisic acid biosynthesis. Plant J. 2014, 78, 80–93. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.-L.; Xu, G.; Fordjour, E.; Zhao, L.; Yang, Y.; Liu, X.; Bai, Z. Efficient biosynthesis of retinal from Escherichia coli by metabolic engineering. Bioresour. Technol. 2026, 440, 133501. [Google Scholar] [CrossRef]
- Lee, J.H.; Lee, S.R.; Lee, S.Y.; Lee, P.Y. Complete microbial synthesis of crocetin and crocins from glycerol in Escherichia coli. Microb. Cell Fact. 2024, 23, 10. [Google Scholar] [CrossRef] [PubMed]
- Qi, Z.; Tong, X.; Zhang, Y.; Jia, S.; Fang, X.; Zhao, L. Carotenoid cleavage dioxygenase 1 and its application for the production of C13-apocarotenoids in microbial cell factories: A review. J. Agric. Food Chem. 2023, 71, 19240–19254. [Google Scholar] [CrossRef]
- Huang, C.-N.; Lim, X.; Ong, L.; Lim, C.; Chen, X.; Zhang, C. Mediating oxidative stress enhances α-ionone biosynthesis and strain robustness during process scaling up. Microb. Cell Fact. 2022, 21, 246. [Google Scholar] [CrossRef]
- Tong, Y.; Jørgensen, T.S.; Whitford, C.M.; Weber, T.; Lee, S.Y. A versatile genetic engineering toolkit for E. coli based on CRISPR-prime editing. Nat. Commun. 2021, 12, 5206. [Google Scholar] [CrossRef]
- Mao, J.; Zhang, H.; Chen, Y.; Wei, L.; Liu, J.; Nielsen, J.; Chen, Y.; Xu, N. Relieving metabolic burden to improve robustness and bioproduction by industrial microorganisms. Biotechnol. Adv. 2024, 74, 108401. [Google Scholar] [CrossRef]
- da Silva, G.P.; Mack, M.; Contiero, J. Glycerol: A promising and abundant carbon source for industrial microbiology. Biotechnol. Adv. 2009, 27, 30–39. [Google Scholar] [CrossRef] [PubMed]
- Hibbert, E.G.; Dalby, P.A. Directed evolution strategies for improved enzymatic performance. Microb. Cell Fact. 2005, 4, 29. [Google Scholar] [CrossRef] [PubMed]
- Meng, J.; Ke, J.; Zhu, Y.; Zhang, T.; Wan, L.; Mu, W. Spatial organization of the metabolic pathway for enhancing L-fucose biosynthesis in engineered Escherichia coli. J. Agric. Food Chem. 2025, 73, 22626–22632. [Google Scholar] [CrossRef]
- Lin, P.; Du, G.; Chen, J.; Zhang, J.; Peng, Z. Engineered microbial production of carotenoids and their cleavage products: Recent advances and prospects. Biotechnol. Adv. 2025, 85, 108708. [Google Scholar] [CrossRef]
- Seok, J.; Styczynski, M.P. Whole-cell and cell-free biosensor-driven metabolic engineering. Curr. Opin. Biotechnol. 2025, 96, 103385. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Lee, T.S. Enhancing isoprenol production by systematically tuning metabolic pathways using CRISPR interference in E. coli. Front. Bioeng. Biotechnol. 2023, 11, 1296132. [Google Scholar] [CrossRef]
- Stohr, A.M.; Hansen, H.; Richards, B.; Park, H.; Goncalves, A.G.; Agrawal, A.; Blenner, M.; Chen, W. Metabolite-responsive scaffold RNAs for dynamic CRISPR transcriptional regulation. Nucleic Acids Res. 2025, 53, gkaf1290. [Google Scholar] [CrossRef]
- Elizondo, B.R.; Saa, P.A. Complex kinetic models predict β-carotene production and reveal flux limitations in recombinant Saccharomyces cerevisiae strains. ACS Synth. Biol. 2025, 14, 3457–3472. [Google Scholar] [CrossRef]
- Merzbacher, C.; Aodha, O.M.; Oyarzún, D.A. Modeling host-pathway dynamics at the genome scale with machine learning. Metab. Eng. 2025, 91, 480–491. [Google Scholar] [CrossRef] [PubMed]
- Tu, W.; Xu, J.; Ma, Y.; Katsimpouras, C.; Stephanopoulos, G.S. Machine learning-driven optimization of metabolic balance for β-carotene production. Metab. Eng. 2026, 94, 192–201. [Google Scholar] [CrossRef] [PubMed]


Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Liu, J.; Shi, Y.; Zhao, D.; Lin, M.; Wang, P.; Zhou, Y.; Yan, X. Strategies for Metabolic Engineering of Escherichia coli for β-Carotene Biosynthesis. Molecules 2026, 31, 611. https://doi.org/10.3390/molecules31040611
Liu J, Shi Y, Zhao D, Lin M, Wang P, Zhou Y, Yan X. Strategies for Metabolic Engineering of Escherichia coli for β-Carotene Biosynthesis. Molecules. 2026; 31(4):611. https://doi.org/10.3390/molecules31040611
Chicago/Turabian StyleLiu, Jiali, Yilin Shi, Daxin Zhao, Minghao Lin, Ping Wang, Ying Zhou, and Xiaohui Yan. 2026. "Strategies for Metabolic Engineering of Escherichia coli for β-Carotene Biosynthesis" Molecules 31, no. 4: 611. https://doi.org/10.3390/molecules31040611
APA StyleLiu, J., Shi, Y., Zhao, D., Lin, M., Wang, P., Zhou, Y., & Yan, X. (2026). Strategies for Metabolic Engineering of Escherichia coli for β-Carotene Biosynthesis. Molecules, 31(4), 611. https://doi.org/10.3390/molecules31040611








