The Substrate Versatility of Δ1-Pyrroline-5-carboxylate Reductase (ProC) from Escherichia coli
Abstract
1. Introduction
2. Results
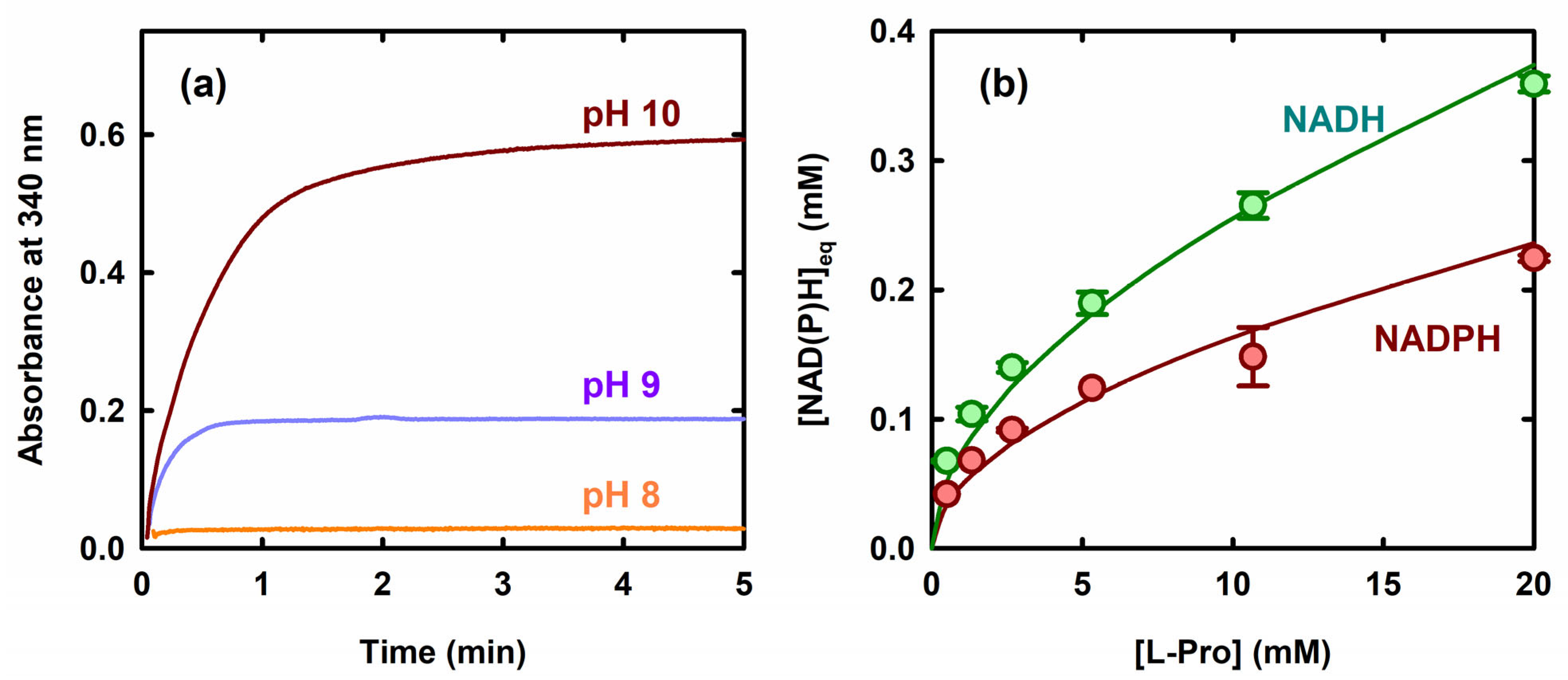
2.1. Reaction of ProC with L-Pro as a Function of pH and Coenzyme Type
2.2. Efficiency and Stereospecificity of the ProC-Catalyzed L-Pro Oxidation
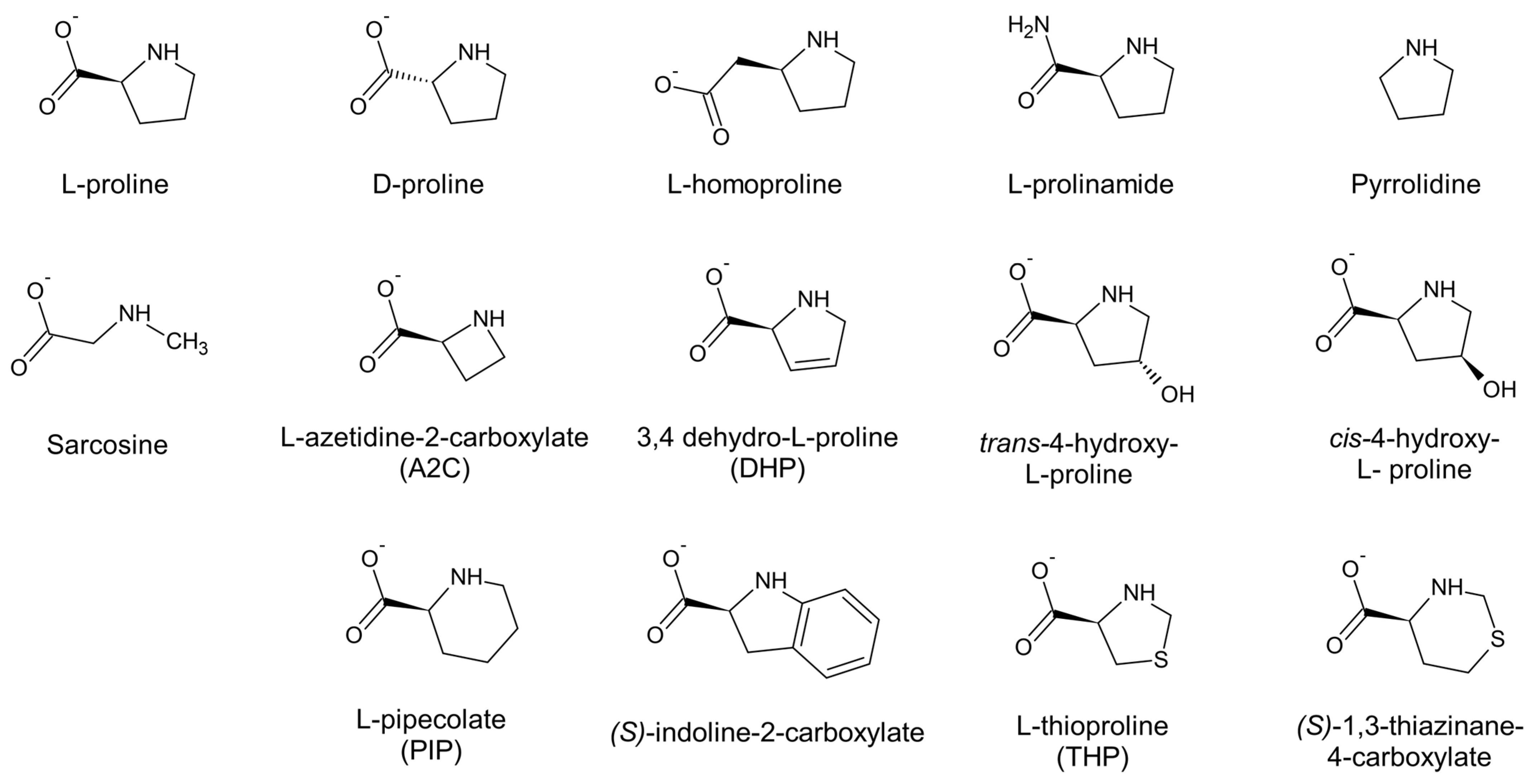
2.3. ProC Dehydrogenase Activity Towards L-Pro Analogs
2.4. Assessing the Ability of the Unreactive L-Pro Analogs to Inhibit ProC
2.5. Building a Structural Model for E. coli ProC
2.6. Docking of L-Pro in the Two Different Models
2.7. Docking Analysis of Different L-Pro Analogs Binding to ProC
2.8. Exploring the Genomic Context of proC in Bacteria
3. Discussion
3.1. Outlining the Substrate Preferences of E. coli ProC
3.2. Structural Basis of ProC Binding and Reactivity
3.3. On the Biological Function of ProC
3.4. Final Remarks
4. Materials and Methods
4.1. Materials
4.2. Kinetic Assays of ProC Activity
4.3. Calculating the Apparent Equilibrium Constant for L-Pro Oxidation at pH 10.0
4.4. Estimating Inhibition Constants for Unreactive L-Pro Analogs
4.5. ProC Structural Modeling
4.6. Docking Simulations
4.7. Genome Context Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| P5C | Δ1-Pyrroline-5-carboxylate |
| GSA | L-Glutamate γ-semialdehyde |
| ProC | Δ1-Pyrroline-5-carboxylate reductase (from E. coli, if not otherwise specified) |
| THP | L-thioproline |
| DHP | 3,4-dehydro-L-proline |
| A2C | L-azetidine-2-carboxylate |
| P6C | Δ1-piperideine-6-carboxylate |
| PIP | L-pipecolate |
References
- Morgan, A.A.; Rubenstein, E. Proline: The Distribution, Frequency, Positioning, and Common Functional Roles of Proline and Polyproline Sequences in the Human Proteome. PLoS ONE 2013, 8, e53785. [Google Scholar] [CrossRef]
- Adams, E.; Frank, L. Metabolism of Proline and the Hydroxyprolines. Annu. Rev. Biochem. 1980, 49, 1005–1061. [Google Scholar] [CrossRef]
- Fichman, Y.; Gerdes, S.Y.; Kovács, H.; Szabados, L.; Zilberstein, A.; Csonka, L.N. Evolution of Proline Biosynthesis: Enzymology, Bioinformatics, Genetics, and Transcriptional Regulation. Biol. Rev. 2015, 90, 1065–1099. [Google Scholar] [CrossRef] [PubMed]
- Tanner, J.J.; Fendt, S.M.; Becker, D.F. The Proline Cycle as a Potential Cancer Therapy Target. Biochemistry 2018, 57, 3433–3444. [Google Scholar] [CrossRef]
- Forlani, G.; Nocek, B.; Chakravarthy, S.; Joachimiak, A. Functional Characterization of Four Putative Δ1-Pyrroline-5-Carboxylate Reductases from Bacillus subtilis. Front. Microbiol. 2017, 8, 1442. [Google Scholar] [CrossRef] [PubMed]
- Smith, M.E.; Greenberg, D.M. Characterization of an Enzyme Reducing Pyrroline-5-Carboxylate to Proline. Nature 1956, 177, 1130. [Google Scholar] [CrossRef]
- Rossi, J.J.; Vender, J.; Berg, C.M.; Coleman, W.H. Partial Purification and Some Properties of Δ1-Pyrroline-5-Carboxylate Reductase from Escherichia coli. J. Bacteriol. 1977, 129, 108–114. [Google Scholar] [CrossRef] [PubMed]
- Fujii, T.; Mukaihara, M.; Agematu, H.; Tsunekawa, H. Biotransformation of L-Lysine to L-Pipecolic Acid Catalyzed by L-Lysine 6-Aminotransferase and Pyrroline-5-Carboxylate Reductase. Biosci. Biotechnol. Biochem. 2002, 66, 622–627. [Google Scholar] [CrossRef]
- Niehaus, E.M.; Janevska, S.; Von Bargen, K.W.; Sieber, C.M.K.; Harrer, H.; Humpf, H.U.; Tudzynski, B. Apicidin F: Characterization and Genetic Manipulation of a New Secondary Metabolite Gene Cluster in the Rice Pathogen Fusarium fujikuroi. PLoS ONE 2014, 9, e103336. [Google Scholar] [CrossRef]
- Deutch, C.E.; Klarstrom, J.L.; Link, C.L.; Ricciardi, D.L. Oxidation of L-Thiazolidine-4-Carboxylate by Δ1-Pyrroline-5-Carboxylate Reductase in Escherichia coli. Curr. Microbiol. 2001, 42, 442–446. [Google Scholar] [CrossRef]
- Patel, S.M.; Seravalli, J.; Stiers, K.M.; Tanner, J.J.; Becker, D.F. Kinetics of Human Pyrroline-5-Carboxylate Reductase in L-Thioproline Metabolism. Amino Acids 2021, 53, 1863–1874. [Google Scholar] [CrossRef] [PubMed]
- Umansky, C.; Morellato, A.E.; Pontel, L.B. Illuminating Cellular Formaldehyde. Nat. Commun. 2021, 12, 580. [Google Scholar] [CrossRef]
- Klein, V.J.; Irla, M.; López, M.G.; Brautaset, T.; Brito, L.F. Unravelling Formaldehyde Metabolism in Bacteria: Road towards Synthetic Methylotrophy. Microorganisms 2022, 10, 220. [Google Scholar] [CrossRef]
- Franklin, M.C.; Cheung, J.; Rudolph, M.J.; Burshteyn, F.; Cassidy, M.; Gary, E.; Hillerich, B.; Yao, Z.K.; Carlier, P.R.; Totrov, M.; et al. Structural Genomics for Drug Design against the Pathogen Coxiella burnetii. Proteins Struct. Funct. Bioinform. 2015, 83, 2124–2136. [Google Scholar] [CrossRef] [PubMed]
- Nocek, B.; Chang, C.; Li, H.; Lezondra, L.; Holzle, D.; Collart, F.; Joachimiak, A. Crystal Structures of Δ1-Pyrroline-5-Carboxylate Reductase from Human Pathogens Neisseria meningitides and Streptococcus pyogenes. J. Mol. Biol. 2005, 354, 91–106. [Google Scholar] [CrossRef] [PubMed]
- Khersonsky, O.; Tawfik, D.S. Enzyme Promiscuity: A Mechanistic and Evolutionary Perspective. Annu. Rev. Biochem. 2010, 79, 471–505. [Google Scholar] [CrossRef]
- Copley, S.D. Shining a Light on Enzyme Promiscuity. Curr. Opin. Struct. Biol. 2017, 47, 167–175. [Google Scholar] [CrossRef]
- Ferla, M.P.; Brewster, J.L.; Hall, K.R.; Evans, G.B.; Patrick, W.M. Primordial-like Enzymes from Bacteria with Reduced Genomes. Mol. Microbiol. 2017, 105, 508–524. [Google Scholar] [CrossRef]
- Panikov, N.S. Genome-Scale Reconstruction of Microbial Dynamic Phenotype: Successes and Challenges. Microorganisms 2021, 9, 2352. [Google Scholar] [CrossRef]
- Chilson, O.P.; Kelly-Chilson, A.E.; Schneider, J.D. Pyrroline-5-Carboxylate Reductase in Soybean Nodules. Comparison of the Enzymes in Host Cytosol, Bradyrhizobium japonicum Bacteroids, and Cultures. Plant Physiol. 1992, 99, 119–123. [Google Scholar]
- Kenklies, J.; Ziehn, R.; Fritsche, K.; Pich, A.; Andreesen, J.R. Proline Biosynthesis from L-Ornithine in Clostridium sticklandii: Purification of Δ1-Pyrroline-5-Carboxylate Reductase, and Sequence and Expression of the Encoding Gene, proC. Microbiology 1999, 145, 819–826. [Google Scholar] [CrossRef][Green Version]
- Yang, Y.; Xu, S.; Zhang, M.; Jin, R.; Zhang, L.; Bao, J.; Wang, H. Purification and Characterization of a Functionally Active Mycobacterium tuberculosis Pyrroline-5-Carboxylate Reductase. Protein Expr. Purif. 2006, 45, 241–248. [Google Scholar] [CrossRef] [PubMed]
- Engel, P.C.; Dalziel, K. The Equilibrium Constants of the Glutamate Dehydrogenase Systems. Biochem. J. 1967, 105, 691. [Google Scholar] [CrossRef] [PubMed]
- Alberty, R.A. Thermodynamics of Reactions of Nicotinamide Adenine Dinucleotide and Nicotinamide Adenine Dinucleotide Phosphate. Arch. Biochem. Biophys. 1993, 307, 8–14. [Google Scholar] [CrossRef]
- Christensen, E.M.; Patel, S.M.; Korasick, D.A.; Campbell, A.C.; Krause, K.L.; Becker, D.F.; Tanner, J.J. Resolving the Cofactor-Binding Site in the Proline Biosynthetic Enzyme Human Pyrroline-5-Carboxylate Reductase 1. J. Biol. Chem. 2017, 292, 7233–7243. [Google Scholar] [CrossRef]
- Krueger, R.; Jager, H.-J.; Hintz, M.; Pahlich, E. Purification to Homogeneity of Pyrroline-5-Carboxylate Reductase of Barley. Plant Physiol. 1986, 80, 142–144. [Google Scholar] [CrossRef]
- Kubyshkin, V.; Rubini, M. Proline Analogues. Chem. Rev. 2024, 124, 8130–8232. [Google Scholar] [CrossRef]
- Peracchi, A.; Polverini, E. Using Steady-State Kinetics to Quantitate Substrate Selectivity and Specificity: A Case Study with Two Human Transaminases. Molecules 2022, 27, 1398. [Google Scholar] [CrossRef]
- Waterhouse, A.; Bertoni, M.; Bienert, S.; Studer, G.; Tauriello, G.; Gumienny, R.; Heer, F.T.; De Beer, T.A.P.; Rempfer, C.; Bordoli, L.; et al. SWISS-MODEL: Homology Modelling of Protein Structures and Complexes. Nucleic Acids Res. 2018, 46, W296–W303. [Google Scholar] [CrossRef]
- Studer, G.; Rempfer, C.; Waterhouse, A.M.; Gumienny, R.; Haas, J.; Schwede, T. QMEANDisCo—Distance Constraints Applied on Model Quality Estimation. Bioinformatics 2020, 36, 1765–1771. [Google Scholar] [CrossRef] [PubMed]
- Plonski, A.P.; Reed, S.M. Assessing Protein Homology Models with Docking Reproducibility. J. Mol. Graph. Model. 2023, 121, 108430. [Google Scholar] [CrossRef] [PubMed]
- Fersht, A. Structure and Mechanism in Protein Science: A Guide to Enzyme Catalysis and Protein Folding; W. H. Freeman & Co.: New York, NY, USA, 1999. [Google Scholar]
- Pastoriza, C.; Antelo, J.M.; Amoedo, F.A.; Parajó, M. N-Chlorination Rate of Five-Membered Heterocyclic Nitrogen Compounds. J. Phys. Org. Chem. 2016, 29, 382–393. [Google Scholar] [CrossRef]
- Nagel, Z.D.; Klinman, J.P. Tunneling and Dynamics in Enzymatic Hydride Transfer. Chem. Rev. 2006, 106, 3095–3118. [Google Scholar] [CrossRef]
- Stojković, V.; Perissinotti, L.L.; Willmer, D.; Benkovic, S.J.; Kohen, A. Effects of the Donor–Acceptor Distance and Dynamics on Hydride Tunneling in the Dihydrofolate Reductase Catalyzed Reaction. J. Am. Chem. Soc. 2012, 134, 1738–1745. [Google Scholar] [CrossRef]
- Fitzpatrick, P.F. Conservation of Mechanism in Flavoprotein-Catalyzed Amine Oxidation. Arch. Biochem. Biophys. 2025, 764, 110242. [Google Scholar] [CrossRef]
- Overbeek, R.; Fonstein, M.; Pusch, G.D.; Maltsev, N. The Use of Gene Clusters to Infer Functional Coupling. Proc. Natl. Acad. Sci. USA 1999, 96, 2896–2901. [Google Scholar] [CrossRef]
- Price, M.N.; Huang, K.H.; Arkin, A.P.; Alm, E.J. Operon Formation Is Driven by Co-Regulation and Not by Horizontal Gene Transfer. Genome Res. 2005, 15, 809–819. [Google Scholar] [CrossRef] [PubMed]
- Newton, M.S.; Azadeh, A.L.; Morgenthaler, A.B.; Copley, S.D. Challenging a Decades-Old Paradigm: ProB and ProA Do Not Channel the Unstable Intermediate in Proline Synthesis after All. Proc. Natl. Acad. Sci. USA 2024, 121, e2413673121. [Google Scholar] [CrossRef]
- Deutch, C.E. Identification of Key Amino Acid Residues in L-Proline Catabolic Enzymes from Gram-Positive Bacteria. J. Adv. Microbiol. 2025, 25, 119–133. [Google Scholar] [CrossRef]
- Miller, G.; Honig, A.; Stein, H.; Suzuki, N.; Mittler, R.; Zilberstein, A. Unraveling Δ1-Pyrroline-5-Carboxylate-Proline Cycle in Plants by Uncoupled Expression of Proline Oxidation Enzymes. J. Biol. Chem. 2009, 284, 26482–26492. [Google Scholar] [CrossRef]
- Huang, Y.Y.; Martínez-Del Campo, A.; Balskus, E.P. Anaerobic 4-Hydroxyproline Utilization: Discovery of a New Glycyl Radical Enzyme in the Human Gut Microbiome Uncovers a Widespread Microbial Metabolic Activity. Gut Microbes 2018, 9, 437–451. [Google Scholar] [CrossRef]
- Savioz, A.; Jeenes, D.J.; Kocher, H.P.; Haas, D. Comparison of ProC and Other Housekeeping Genes of Pseudomonas aeruginosa with Their Counterparts in Escherichia coli. Gene 1990, 86, 107–111. [Google Scholar] [CrossRef]
- Ikegawa, S.; Isomura, M.; Koshizuka, Y.; Nakamura, Y. Cloning and Characterization of Human and Mouse PROSC (Proline Synthetase Co-Transcribed) Genes. J. Hum. Genet. 1999, 44, 337–342. [Google Scholar] [CrossRef][Green Version]
- Darin, N.; Reid, E.; Prunetti, L.; Samuelsson, L.; Husain, R.A.; Wilson, M.; El Yacoubi, B.; Footitt, E.; Chong, W.K.; Wilson, L.C.; et al. Mutations in PROSC Disrupt Cellular Pyridoxal Phosphate Homeostasis and Cause Vitamin-B6-Dependent Epilepsy. Am. J. Hum. Genet. 2016, 99, 1325–1337. [Google Scholar] [CrossRef]
- Tramonti, A.; Ghatge, M.S.; Babor, J.T.; Musayev, F.N.; di Salvo, M.L.; Barile, A.; Colotti, G.; Giorgi, A.; Paredes, S.D.; Donkor, A.K.; et al. Characterization of the Escherichia coli Pyridoxal 5′-Phosphate Homeostasis Protein (YggS): Role of Lysine Residues in PLP Binding and Protein Stability. Protein Sci. 2022, 31, e4471. [Google Scholar] [CrossRef]
- Serrano, H.; Blanchard, J.S. Kinetic and Isotopic Characterization of L-Proline Dehydrogenase from Mycobacterium tuberculosis. Biochemistry 2013, 52, 5009–5015. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Luesch, H.; Hoffmann, D.; Hevel, J.M.; Becker, J.E.; Golakoti, T.; Moore, R.E. Biosynthesis of 4-Methylproline in Cyanobacteria: Cloning of nosE and nosF Genes and Biochemical Characterization of the Encoded Dehydrogenase and Reductase Activities. J. Org. Chem. 2003, 68, 83–91. [Google Scholar] [CrossRef] [PubMed]
- Gómez, D.; Lucas-Elío, P.; Sanchez-Amat, A.; Solano, F. A Novel Type of Lysine Oxidase: L-Lysine-ε-Oxidase. Biochim. Biophys. Acta (BBA)—Proteins Proteom. 2006, 1764, 1577–1585. [Google Scholar] [CrossRef] [PubMed]
- Merx, J.; van Outersterp, R.E.; Engelke, U.F.H.; Hendriks, V.; Wevers, R.A.; Huigen, M.C.D.G.; Waterval, H.W.A.H.; Körver-Keularts, I.M.L.W.; Mecinović, J.; Rutjes, F.P.J.T.; et al. Identification of Δ1-Pyrroline-5-Carboxylate Derived Biomarkers for Hyperprolinemia Type II. Commun. Biol. 2022, 5, 997. [Google Scholar] [CrossRef]
- Newman, R.A.; Hacker, M.P.; McCormack, J.J.; Krakoff, I.H. Pharmacologic and Toxicologic Evaluation of Thioproline: A Proposed Nontoxic Inducer of Reverse Transformation. Cancer Treat. Rep. 1980, 64, 837–844. [Google Scholar]
- Backiam, A.D.S.; Duraisamy, S.; Karuppaiya, P.; Balakrishnan, S.; Sathyan, A.; Kumarasamy, A.; Raju, A. Analysis of the Main Bioactive Compounds from Ocimum basilicum for Their Antimicrobial and Antioxidant Activity. Biotechnol. Appl. Biochem. 2023, 70, 2038–2051. [Google Scholar] [CrossRef]
- Patil, M.; Jaisinghani, R.; Sharma, R.; Patil, R. Comparative Study of Quercetin & Lemon Peel Extract on Multiple Drug-Resistant Bacteria. J. Microbiol. Biotechnol. Food Sci. 2024, 14, e10099. [Google Scholar]
- Hormann, K.; Andreesen, J.R. A Flavin-Dependent Oxygenase Reaction Initiates the Degradation of Pyrrole-2-Carboxylate in Arthrobacter Strain Pyl (DSM 6386). Arch. Microbiol. 1991, 157, 43–48. [Google Scholar] [CrossRef]
- Omura, H.; Wieser, M.; Nagasawa, T. Pyrrole-2-Carboxylate Decarboxylase from Bacillus megaterium PYR2910, an Organic-Acid-Requiring Enzyme. Eur. J. Biochem. 1998, 253, 480–484. [Google Scholar] [CrossRef] [PubMed]
- Payne, K.A.P.; Marshall, S.A.; Fisher, K.; Rigby, S.E.J.; Cliff, M.J.; Spiess, R.; Cannas, D.M.; Larrosa, I.; Hay, S.; Leys, D. Structure and Mechanism of Pseudomonas aeruginosa PA0254/HudA, a PrFMN-Dependent Pyrrole-2-Carboxylic Acid Decarboxylase Linked to Virulence. ACS Catal. 2021, 11, 2865–2878. [Google Scholar] [CrossRef]
- Jiang, H.; Haltli, B.; Feng, X.; Cai, P.; Summers, M.; Lotvin, J.; He, M. Investigation of the Biosynthesis of the Pipecolate Moiety of Neuroprotective Polyketide Meridamycin. J. Antibiot. 2011, 64, 533–538. [Google Scholar] [CrossRef]
- Liras, P.; Martín, J.F. Interconnected Set of Enzymes Provide Lysine Biosynthetic Intermediates and Ornithine Derivatives as Key Precursors for the Biosynthesis of Bioactive Secondary Metabolites. Antibiotics 2023, 12, 159. [Google Scholar] [CrossRef] [PubMed]
- Blaby-Haas, C.E.; de Crécy-Lagard, V. Mining High-Throughput Experimental Data to Link Gene and Function. Trends Biotechnol. 2011, 29, 174–182. [Google Scholar] [CrossRef]
- Ling, M.; Allen, S.W.; Wood, J.M. Sequence Analysis Identifies the Proline Dehydrogenase and Δ1-Pyrroline-5-Carboxylate Dehydrogenase Domains of the Multifunctional Escherichia coli PutA Protein. J. Mol. Biol. 1994, 243, 950–956. [Google Scholar] [CrossRef]
- Moxley, M.A.; Tanner, J.J.; Becker, D.F. Steady-State Kinetic Mechanism of the Proline:Ubiquinone Oxidoreductase Activity of Proline Utilization A (PutA) from Escherichia coli. Arch. Biochem. Biophys. 2011, 516, 113–120. [Google Scholar] [CrossRef] [PubMed]
- Mao, Y.; Seravalli, J.; Smith, T.G.; Morton, M.; Tanner, J.J.; Becker, D.F. Evidence for Proline Catabolic Enzymes in the Metabolism of Thiazolidine Carboxylates. Biochemistry 2021, 60, 3610–3620. [Google Scholar] [CrossRef] [PubMed]
- Kitagawa, M.; Ara, T.; Arifuzzaman, M.; Ioka-Nakamichi, T.; Inamoto, E.; Toyonaga, H.; Mori, H. Complete Set of ORF Clones of Escherichia coli ASKA Library (A Complete Set of E. coli K-12 ORF Archive): Unique Resources for Biological Research. DNA Res. 2005, 12, 291–299. [Google Scholar] [CrossRef]
- Zangelmi, E.; Ronda, L.; Castagna, C.; Campanini, B.; Veiga-da-Cunha, M.; Van Schaftingen, E.; Peracchi, A. Off to a Slow Start: Analyzing Lag Phases and Accelerating Rates in Steady-State Enzyme Kinetics. Anal. Biochem. 2020, 593, 113595. [Google Scholar] [CrossRef]
- Deutch, A.H.; Smith, C.J.; Rushlow, K.E.; Kretschmer, P.J. Escherichia coli Δ1-Pyrroline-5-Carboxylate Reductase: Gene Sequence, Protein Overproduction and Purification. Nucleic Acids Res. 1982, 10, 7701–7714. [Google Scholar] [CrossRef]
- Berman, H.M.; Westbrook, J.; Feng, Z.; Gilliland, G.; Bhat, T.N.; Weissig, H.; Shindyalov, I.N.; Bourne, P.E. The Protein Data Bank. Nucleic Acids Res. 2000, 28, 235–242. [Google Scholar] [CrossRef]
- Abramson, J.; Adler, J.; Dunger, J.; Evans, R.; Green, T.; Pritzel, A.; Ronneberger, O.; Willmore, L.; Ballard, A.J.; Bambrick, J.; et al. Accurate Structure Prediction of Biomolecular Interactions with AlphaFold 3. Nature 2024, 630, 493–500. [Google Scholar] [CrossRef]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic Local Alignment Search Tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
- Sievers, F.; Higgins, D.G. Clustal Omega for Making Accurate Alignments of Many Protein Sequences. Protein Sci. 2018, 27, 135–145. [Google Scholar] [CrossRef]
- Bertoni, M.; Kiefer, F.; Biasini, M.; Bordoli, L.; Schwede, T. Modeling Protein Quaternary Structure of Homo- and Hetero-Oligomers beyond Binary Interactions by Homology. Sci. Rep. 2017, 7, 10480. [Google Scholar] [CrossRef] [PubMed]
- Guex, N.; Peitsch, M.C. SWISS-MODEL and the Swiss-PdbViewer: An Environment for Comparative Protein Modeling. Electrophoresis 1997, 18, 2714–2723. [Google Scholar] [CrossRef] [PubMed]
- Pettersen, E.F.; Goddard, T.D.; Huang, C.C.; Couch, G.S.; Greenblatt, D.M.; Meng, E.C.; Ferrin, T.E. UCSF Chimera—A Visualization System for Exploratory Research and Analysis. J. Comput. Chem. 2004, 25, 1605–1612. [Google Scholar] [CrossRef]
- Gasteiger, J.; Marsili, M. Iterative Partial Equalization of Orbital Electronegativity—A Rapid Access to Atomic Charges. Tetrahedron 1980, 36, 3219–3228. [Google Scholar] [CrossRef]
- Morris, G.M.; Huey, R.; Lindstrom, W.; Sanner, M.F.; Belew, R.K.; Goodsell, D.S.; Olson, A.J. AutoDock4 and AutoDockTools4: Automated Docking with with Selective Receptor Flexibility. J. Comput. Chem. 2009, 30, 2785–2791. [Google Scholar] [CrossRef] [PubMed]
- Stroet, M.; Caron, B.; Visscher, K.M.; Geerke, D.P.; Malde, A.K.; Mark, A.E. Automated Topology Builder Version 3.0: Prediction of Solvation Free Enthalpies in Water and Hexane. J. Chem. Theory Comput. 2018, 14, 5834–5845. [Google Scholar] [CrossRef]
- Morris, G.M.; Goodsell, D.S.; Halliday, R.S.; Huey, R.; Hart, W.E.; Belew, R.K.; Olson, A.J. Automated Docking Using a Lamarckian Genetic Algorithm and an Empirical Binding Free Energy Function. J. Comput. Chem. 1998, 19, 1639–1662. [Google Scholar]
- Humphrey, W.; Dalke, A.; Schulten, K. VMD: Visual Molecular Dynamics. J. Mol. Graph. 1996, 14, 33–38. [Google Scholar] [CrossRef] [PubMed]
- Szklarczyk, D.; Kirsch, R.; Koutrouli, M.; Nastou, K.; Mehryary, F.; Hachilif, R.; Gable, A.L.; Fang, T.; Doncheva, N.T.; Pyysalo, S.; et al. The STRING Database in 2023: Protein-Protein Association Networks and Functional Enrichment Analyses for Any Sequenced Genome of Interest. Nucleic Acids Res. 2023, 51, D638–D646. [Google Scholar] [CrossRef]
- Zangelmi, E.; Stanković, T.; Malatesta, M.; Acquotti, D.; Pallitsch, K.; Peracchi, A. Discovery of a New, Recurrent Enzyme in Bacterial Phosphonate Degradation: (R)-1-Hydroxy-2-Aminoethylphosphonate Ammonia-Lyase. Biochemistry 2021, 60, 1214–1225. [Google Scholar] [CrossRef] [PubMed]
- Zangelmi, E.; Ruffolo, F.; Dinhof, T.; Gerdol, M.; Malatesta, M.; Chin, J.P.; Rivetti, C.; Secchi, A.; Pallitsch, K.; Peracchi, A. Deciphering the Role of Recurrent FAD-Dependent Enzymes in Bacterial Phosphonate Catabolism. iScience 2023, 26, 108108. [Google Scholar] [CrossRef]
- Chen, I.M.A.; Chu, K.; Palaniappan, K.; Ratner, A.; Huang, J.; Huntemann, M.; Hajek, P.; Ritter, S.J.; Webb, C.; Wu, D.; et al. The IMG/M Data Management and Analysis System v.7: Content Updates and New Features. Nucleic Acids Res. 2023, 51, D723–D732. [Google Scholar] [CrossRef]





| Substrate | kcat (s−1) | KM (mM) | kcat/KM (M−1s−1) | K′ |
|---|---|---|---|---|
| L-Proline | 2.8 ± 0.1 | 0.33 ± 0.05 | 8500 ± 1400 | (1.6 ± 0.1) × 10−3 |
| L-Prolinamide | 4.3 ± 0.6 | 5.2 ± 2 | 830 ± 220 | (1.2 ± 0.1) × 10−4 |
| 3,4-dehydro-L-proline (DHP) | 9.1 ± 0.9 | 0.54 ± 0.31 | 16,900 ± 7500 | - |
| trans-4-hydroxy L-proline | 1.1 ± 0.1 | 20 ± 3 | 55 ± 9 | (1.0 ± 0.01) × 10−4 |
| cis-4-hydroxy L-proline | 1.5 ± 0.7 | 57 ± 35 | 27± 4 | (1.1 ± 0.01) × 10−4 |
| L-Pipecolate (PIP) * | 0.6 ± 0.001 | 4.4 ± 0.6 | 140 ± 22 | (1.0 ± 0.02) × 10−4 |
| L-Thioproline (THP) | 4.7 ± 0.4 | 2.3 ± 0.7 | 2100 ± 650 | - |
| (S)-1,3-Thiazinane 4-carboxylate ** | 0.14 ± 0.01 | 2.5 ± 0.6 | 56 ± 14 | - |
| Ligand | pKa of the Amino Group | Prevalent Ionization State of the Amino Group (pH 10) | Binding Energy (kcal/mol) |
|---|---|---|---|
| L-Proline | 10.65 | protonated | −4.6 |
| D-proline | 10.65 | protonated | −4.2 |
| L-homoproline | 11.3 | protonated | −5.0 |
| L-prolinamide | 8.9 | neutral | −3.7 |
| Pyrrolidine | 11.31 | protonated | −4.0 |
| Sarcosine | 10.01 | protonated/neutral | −3.4/−2.7 |
| L-azetidine-2-carboxylate (A2C) | 10.7 | protonated | −4.1 |
| 3,4-dehydro-L-proline (DHP) | 9.57 | protonated/neutral | −4.5/−3.5 |
| trans-4-hydroxy-L-proline | 9.4 | neutral | −3.4 |
| cis-4-hydroxy-L-proline | 10.3 | protonated/neutral | −4.6/−3.6 |
| L-pipecolate (PIP) * | 10.77 | protonated | −4.7 |
| Indoline-2-carboxylate | 4.9 | neutral | −3.7 |
| L-thioproline (THP) | 6.74 | neutral | −3.2 |
| (S)-1,3-Thiazinane 4-carboxylate | 8.3 | neutral | −3.6 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Polverini, E.; Vecchi, A.; Capra, G.; Pastori, A.; Peracchi, A. The Substrate Versatility of Δ1-Pyrroline-5-carboxylate Reductase (ProC) from Escherichia coli. Molecules 2026, 31, 501. https://doi.org/10.3390/molecules31030501
Polverini E, Vecchi A, Capra G, Pastori A, Peracchi A. The Substrate Versatility of Δ1-Pyrroline-5-carboxylate Reductase (ProC) from Escherichia coli. Molecules. 2026; 31(3):501. https://doi.org/10.3390/molecules31030501
Chicago/Turabian StylePolverini, Eugenia, Alessandro Vecchi, Giulia Capra, Alessia Pastori, and Alessio Peracchi. 2026. "The Substrate Versatility of Δ1-Pyrroline-5-carboxylate Reductase (ProC) from Escherichia coli" Molecules 31, no. 3: 501. https://doi.org/10.3390/molecules31030501
APA StylePolverini, E., Vecchi, A., Capra, G., Pastori, A., & Peracchi, A. (2026). The Substrate Versatility of Δ1-Pyrroline-5-carboxylate Reductase (ProC) from Escherichia coli. Molecules, 31(3), 501. https://doi.org/10.3390/molecules31030501







