In-Depth Exploration of Chemical Constituents from Justicia procumbens L. Through UHPLC-Q-Exactive Orbitrap Mass Spectrometry
Abstract
1. Introduction
2. Results and Discussion
2.1. Analytical Strategy
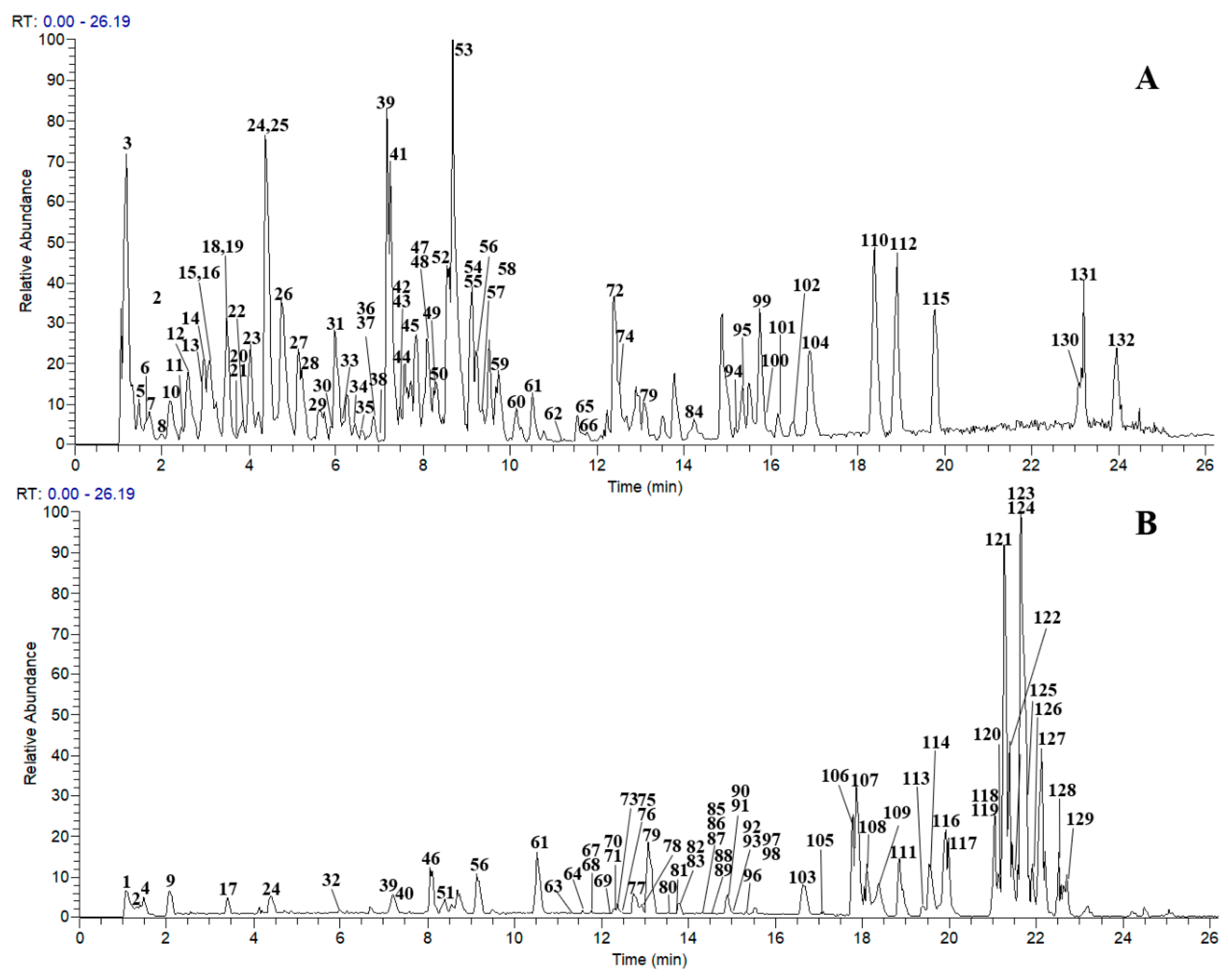
2.2. Characterization of the Chemical Composition in JP by LC-MS/MS
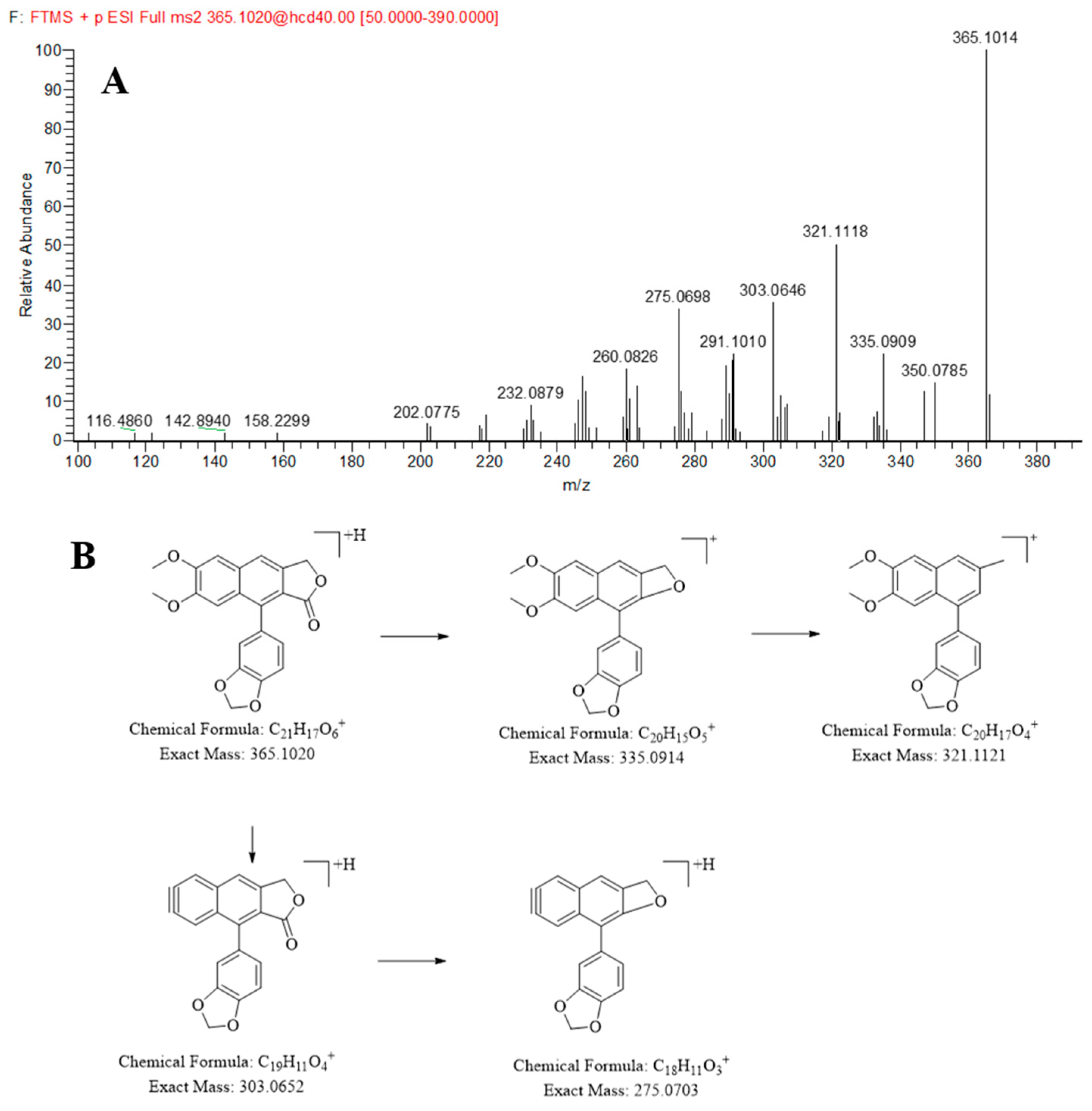
2.2.1. Identification of the Lignans in JP
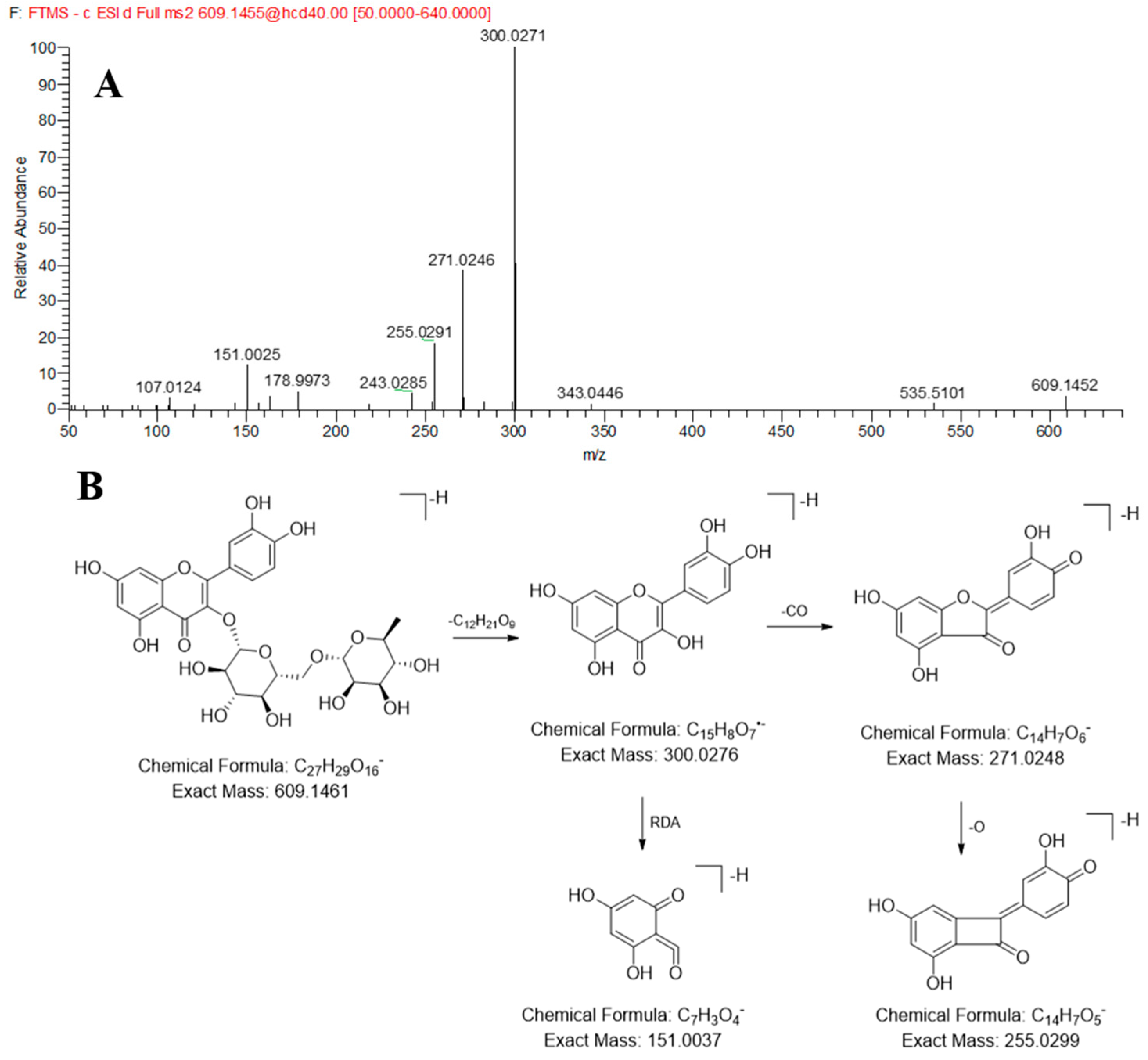
2.2.2. Identification of the Flavonoids in JP
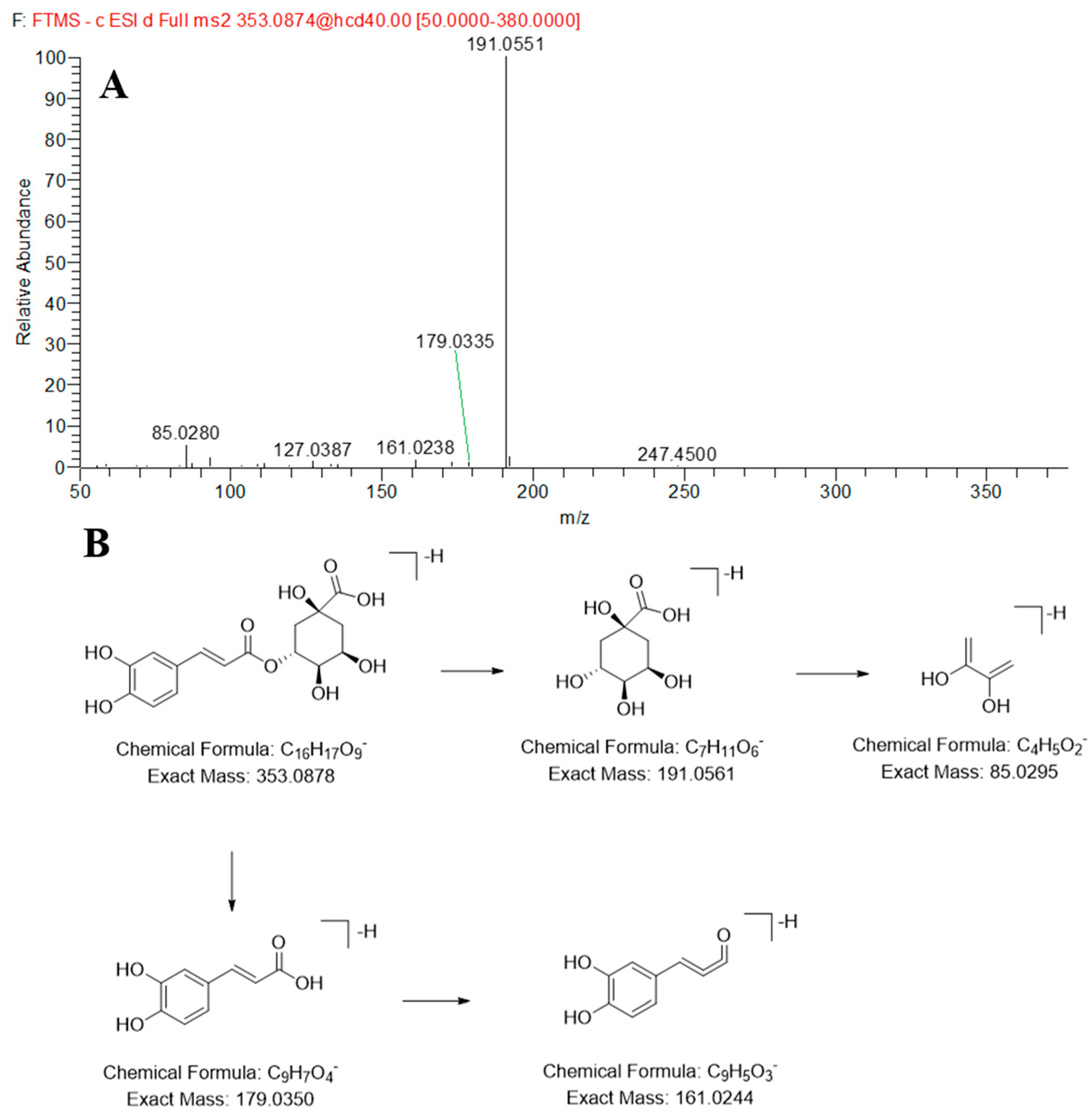
2.2.3. Identification of the Organic Acids in JP
2.2.4. Identification of the Alkaloids in JP
2.2.5. Other Chemical Constituents in JP
3. Materials and Methods
3.1. Chemicals, Reference Standard and Materials
3.2. Standard and Sample Preparation
3.3. Instruments and LC–MS/MS Conditions
3.4. Data Processing and Analysis
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Lee, H.; Jeon, J.; Yoon, J.; Kim, S.-H.; Choi, H.S.; Kang, J.S.; Lee, Y.S.; Lee, M.; Kim, Y.H.; Chang, H.B. Comparative Metabolite Profiling of Wild and Cultivated Justicia procumbens L. Based on 1H-NMR Spectroscopy and HPLC-DAD Analysis. Plants 2020, 9, 860. [Google Scholar] [CrossRef]
- Kim, D.; Lee, E.; Choi, P.G.; Kim, H.S.; Park, S.H.; Seo, H.D.; Hahm, J.H.; Ahn, J.; Jung, C.H. Justicia procumbens prevents hair loss in androgenic alopecia mice. Biomed. Pharmacother. 2024, 170, 115913. [Google Scholar] [CrossRef]
- Ibrahim, S.R.M.; Mohamed, S.G.A.; Abdallah, H.M.; Mohamed, G.A. Ethnomedicinal uses, phytochemistry, and pharmacological relevance of Justicia procumbens (Oriental Water Willow)-A promising traditional plant. J. Ethnopharmacol. 2023, 317, 116819. [Google Scholar] [CrossRef] [PubMed]
- Liu, B.; Zhang, T.; Xie, Z.T.; Hong, Z.C.; Lu, Y.; Long, Y.M.; Ji, C.Z.; Liu, Y.T.; Yang, Y.F.; Wu, H.Z. Effective components and mechanism analysis of anti-platelet aggregation effect of Justicia procumbens L. J. Ethnopharmacol. 2022, 294, 115392. [Google Scholar] [CrossRef] [PubMed]
- Xu, X.Y.; Wang, D.Y.; Ku, C.F.; Zhao, Y.; Cheng, H.; Liu, K.L.; Rong, L.J.; Zhang, H.J. Anti-HIV lignans from Justicia procumbens. Chin. J. Nat. Med. 2019, 17, 945–952. [Google Scholar] [CrossRef] [PubMed]
- Youm, J.; Lee, H.; Chang, H.B.; Jeon, J.; Yoon, M.H.; Woo, J.Y.; Choi, M.S.; Hwang, Y.; Seong, S.; Na, K.; et al. Justicia procumbens Extract (DW2008) Selectively Suppresses Th2 Cytokines in Splenocytes and Ameliorates Ovalbumin-Induced Airway Inflammation in a Mouse Model of Asthma. Biol. Pharm. Bull. 2017, 40, 1416–1422. [Google Scholar] [CrossRef]
- Youm, J.; Lee, H.; Choi, Y.; Yoon, J. DW2008S and its major constituents from Justicia procumbens exert anti-asthmatic effect via multitargeting activity. J. Cell Mol. Med. 2018, 22, 2680–2691. [Google Scholar] [CrossRef]
- Jin, H.; Chen, L.; Tian, Y.; Li, B.; Dong, J.X. New cyclopeptide alkaloid and lignan glycoside fromJusticia procumbens. J. Asian Nat. Prod. Res. 2014, 17, 33–39. [Google Scholar] [CrossRef]
- Jin, H.; Yang, S.; Dong, J.X. New lignan glycosides from Justicia procumbens. J. Asian Nat. Prod. Res. 2017, 19, 1–8. [Google Scholar] [CrossRef]
- Jiang, J.; Dong, H.; Wang, T.; Zhao, R.; Mu, Y.; Geng, Y.; Zheng, Z.; Wang, X. A Strategy for Preparative Separation of 10 Lignans from Justicia procumbens L. by High-Speed Counter-Current Chromatography. Molecules 2017, 22, 2024. [Google Scholar] [CrossRef]
- Nasiri, A.; Jahani, R.; Mokhtari, S.; Yazdanpanah, H.; Daraei, B.; Faizi, M.; Kobarfard, F. Overview, consequences, and strategies for overcoming matrix effects in LC-MS analysis: A critical review. Analyst 2021, 146, 6049–6063. [Google Scholar] [CrossRef]
- D’Ovidio, C.; Locatelli, M.; Perrucci, M.; Ciriolo, L.; Furton, K.G.; Gazioglu, I.; Kabir, A.; Merone, G.M.; de Grazia, U.; Ali, I.; et al. LC-MS/MS Application in Pharmacotoxicological Field: Current State and New Applications. Molecules 2023, 28, 2127. [Google Scholar] [CrossRef]
- Bylda, C.; Thiele, R.; Kobold, U.; Volmer, DA. Recent advances in sample preparation techniques to overcome difficulties encountered during quantitative analysis of small molecules from biofluids using LC-MS/MS. Analyst 2014, 139, 2265–2276. [Google Scholar] [CrossRef] [PubMed]
- Lin, L.F.; Lin, H.M.; Zhang, M.; Dong, X.; Yin, X.B.; Qu, C.H.; Ni, J. Types, principle, and characteristics of tandem high-resolution mass spectrometry and its applications. RSC Adv. 2015, 5, 107623–107636. [Google Scholar] [CrossRef]
- Beccaria, M.; Cabooter, D. Current developments in LC-MS for pharmaceutical analysis. Analyst 2020, 145, 1129–1157. [Google Scholar] [CrossRef]
- Alanazi, S. Recent Advances in Liquid Chromatography-Mass Spectrometry (LC-MS) Applications in Biological and Applied Sciences. Anal. Sci. Adv. 2025, 6, e70024. [Google Scholar] [CrossRef]
- Li, J.; Chen, Y.; Yu, K.; Zhang, M.; Li, Q.; Tang, S.; Liu, Y.; Li, H.; Zhang, Z. Rapid chemical characterization and pharmacological mechanism of Fining Granules in the treatment of chronic bronchitis based on UHPLC–Q-exactive orbitrap mass spectrometer and network pharmacology. Heliyon 2024, 10, e31804. [Google Scholar] [CrossRef]
- Zhao, X.; Ren, D.; Jin, R.; Chen, W.; Xu, L.; Guo, D.; Zhang, Q.; Luo, Z. Development of UHPLC-Q-Exactive Orbitrap/MS Technique for Determination of Proanthocyanidins (PAs) Monomer Composition Content in Persimmon. Plants 2024, 13, 1440. [Google Scholar] [CrossRef] [PubMed]
- Huo, Y.; Li, K.; Yang, S.; Yi, B.; Chai, Z.; Fan, L.; Shu, L.; Gao, B.; Li, H.; Cai, W. A Systematic Methodology for the Identification of the Chemical Composition of the Mongolian Drug Erdun-Uril Compound Utilizing UHPLC-Q-Exactive Orbitrap Mass Spectrometry. Molecules 2024, 29, 4349. [Google Scholar] [CrossRef]
- Zhang, J.; Cai, W.; Zhou, Y.; Liu, Y.; Wu, X.; Li, Y.; Lu, J.; Qiao, Y. Profiling and identification of the metabolites of baicalin and study on their tissue distribution in rats by ultra-high-performance liquid chromatography with linear ion trap-Orbitrap mass spectrometer. J. Chromatogr. B 2015, 985, 91–102. [Google Scholar] [CrossRef]
- Liu, B.; Yang, Y.; Liu, H.; Xie, Z.; Li, Q.; Deng, M.; Li, F.; Peng, J.; Wu, H. Screening for cytotoxic chemical constituents from Justicia procumbens by HPLC–DAD–ESI–MS and NMR. Chem. Cent. J. 2018, 12, 6. [Google Scholar] [CrossRef]
- Zhao, Y.; Ku, C.F.; Xu, X.Y.; Tsang, N.Y.; Zhu, Y.; Zhao, C.L.; Liu, K.L.; Li, C.C.; Rong, L.; Zhang, H.J. Stable Axially Chiral Isomers of Arylnaphthalene Lignan Glycosides with Antiviral Potential Discovered from Justicia procumbens. J. Org. Che 2021, 86, 5568–5583. [Google Scholar] [CrossRef]
- Chen, C.C.; Hsin, W.C.; Huang, Y.L. Six New Diarylbutane Lignans from Justicia procumbens. J. Nat. Prod. 1998, 61, 227–229. [Google Scholar] [CrossRef]
- Gao, S.Q.; Liu, W.K.; Wang, L.N.; Luo, Y.M.; Yang, M.H. HPLC simultaneous determination of 6′-hydroxy-justicidin B and 6′-hydroxy- justicidin A in Herba Justiciae, Chin. J. Pharm. Anal. 2010, 10, 1420–1422. [Google Scholar]
- Weng, J.R.; Ko, H.H.; Yeh, T.L.; Lin, H.C.; Lin, C.N. Two New Arylnaphthalide Lignans and Antiplatelet Constituents from Justicia procumbens. Archiv der Pharmazie 2004, 337, 207–212. [Google Scholar] [CrossRef]
- Asano, J.; Chiba, K.; Tada, M.; Yoshii, T. Antiviral activity of lignans and their glycosides from Justicia procumbens. Phytochemistry 1996, 42, 713–717. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.; Wang, X.; Cheng, Y.; Gao, H.; Chen, X. A Review of Classification, Biosynthesis, Biological Activities and Potential Applications of Flavonoids. Molecules 2023, 28, 4982. [Google Scholar] [CrossRef] [PubMed]
- Chiorcea Paquim, A.M. Electrochemistry of Flavonoids: A Comprehensive Review. Int. J. Mol. Sci. 2023, 24, 15667. [Google Scholar] [CrossRef]
- Schmidt, S.; Zietz, M.; Schreiner, M.; Rohn, S.; Kroh, L.W.; Krumbein, A. Identification of complex, naturally occurring flavonoid glycosides in kale (Brassica oleracea var. sabellica) by high-performance liquid chromatography diode-array detection/electrospray ionization multi-stage mass spectrometry. Rapid Commun. Mass. Spectrom. 2010, 24, 2009–2022. [Google Scholar] [CrossRef]
- Sun, Y.; Qin, Y.; Li, H.; Peng, H.; Chen, H.; Xie, H.R.; Deng, Z. Rapid characterization of chemical constituents in Radix Tetrastigma, a functional herbal mixture, before and after metabolism and their antioxidant/antiproliferative activities. J. Funct. Foods 2015, 18, 300–318. [Google Scholar] [CrossRef]
- Ayoub, I.M.; Abdel-Aziz, M.M.; Elhady, S.S.; Bagalagel, A.A.; Malatani, R.T.; Elkady, W.M. Valorization of Pimenta racemosa Essential Oils and Extracts: GC-MS and LC-MS Phytochemical Profiling and Evaluation of Helicobacter pylori Inhibitory Activity. Molecules 2022, 27, 7965. [Google Scholar] [CrossRef]
- Lv, J.P.; Yang, S.; Dong, J.X.; Jin, H. New cyclopeptide alkaloids from the whole plant of Justicia procumbens L. Nat. Prod. Res. 2021, 35, 4032–4040. [Google Scholar] [CrossRef]
- Lee, T.H.; Lu, C.K.; Wang, G.J.; Chang, Y.C.; Yang, W.B.; Ju, Y.M. Sesquiterpene glycosides from Cosmospora joca. J. Nat. Prod. 2011, 74, 1561–1567. [Google Scholar] [CrossRef] [PubMed]
- Atanasov, A.G.; Zotchev, S.B.; Dirsch, V.M.; International Natural Product Sciences Taskforce; Supuran, C.T. Natural products in drug discovery: Advances and opportunities. Nat. Rev. Drug Discov. 2021, 20, 200–216. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.; Hu, J.S.; Xu, J.L.; Shao, C.L.; Wang, G.Y. Biological and chemical diversity of ascidian-associated microorganisms. Mar. Drugs 2018, 16, 362. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Guan, L.; Luo, H.; Liu, S.; Ming, X.; Hu, M.; Luo, L.; Tan, J.; Xiao, S. In-Depth Exploration of Chemical Constituents from Justicia procumbens L. Through UHPLC-Q-Exactive Orbitrap Mass Spectrometry. Molecules 2025, 30, 3554. https://doi.org/10.3390/molecules30173554
Guan L, Luo H, Liu S, Ming X, Hu M, Luo L, Tan J, Xiao S. In-Depth Exploration of Chemical Constituents from Justicia procumbens L. Through UHPLC-Q-Exactive Orbitrap Mass Spectrometry. Molecules. 2025; 30(17):3554. https://doi.org/10.3390/molecules30173554
Chicago/Turabian StyleGuan, Liangjun, Huibin Luo, Siqiong Liu, Xinrong Ming, Mengdie Hu, Lan Luo, Jingyi Tan, and Shunli Xiao. 2025. "In-Depth Exploration of Chemical Constituents from Justicia procumbens L. Through UHPLC-Q-Exactive Orbitrap Mass Spectrometry" Molecules 30, no. 17: 3554. https://doi.org/10.3390/molecules30173554
APA StyleGuan, L., Luo, H., Liu, S., Ming, X., Hu, M., Luo, L., Tan, J., & Xiao, S. (2025). In-Depth Exploration of Chemical Constituents from Justicia procumbens L. Through UHPLC-Q-Exactive Orbitrap Mass Spectrometry. Molecules, 30(17), 3554. https://doi.org/10.3390/molecules30173554





