Synergistic Disruption of Foodborne Pathogen Biofilms by Oregano Essential Oil and Bacteriophage phiLLS: Atomic Force Microscopy Insights
Abstract
1. Introduction
2. Results
2.1. Composition of the Essential Oil
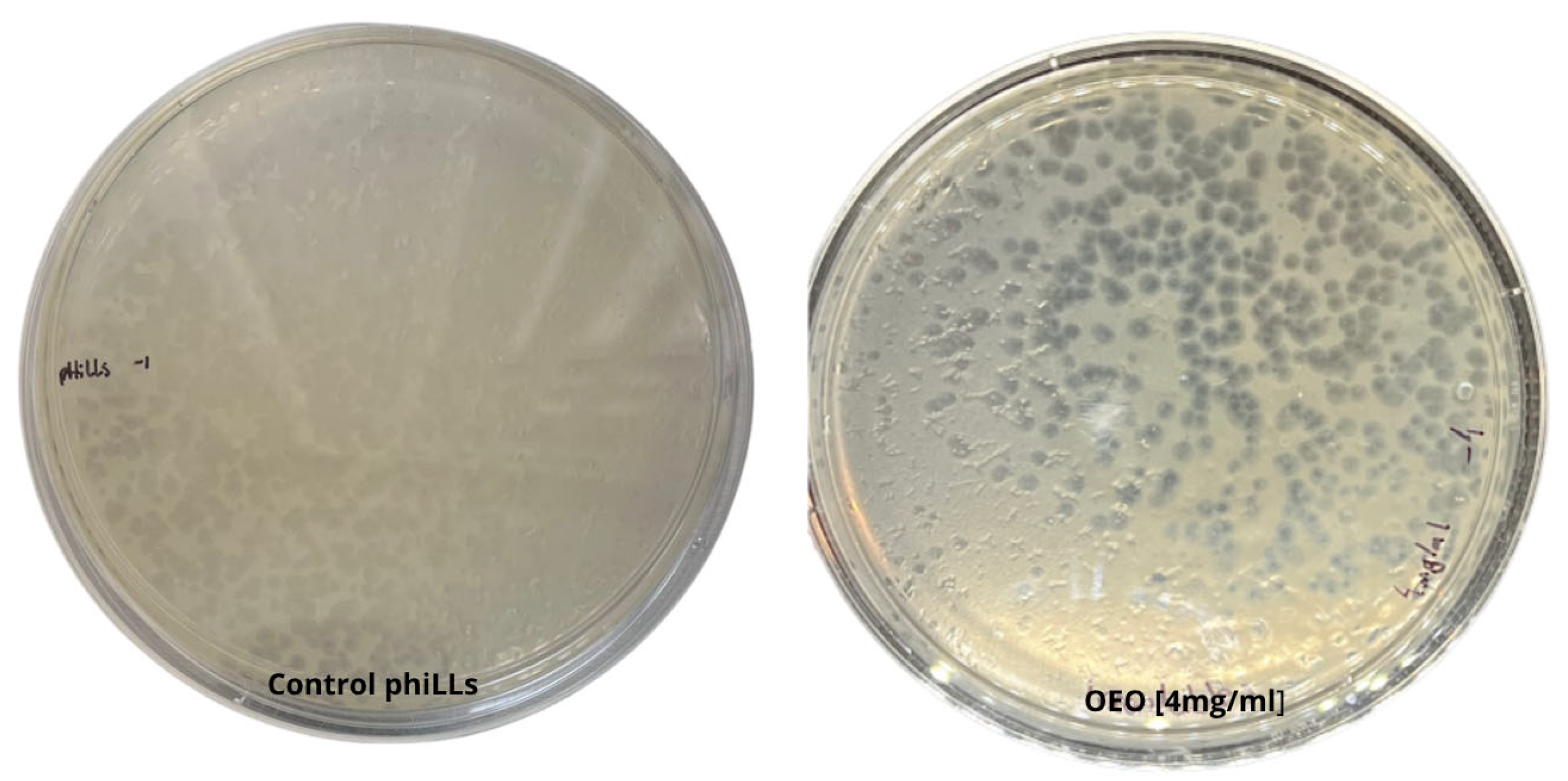
2.2. Phage Sensitivity to OEO
2.3. MIC and MBC Values of OEO
2.4. Combined Application of OEO and Phage
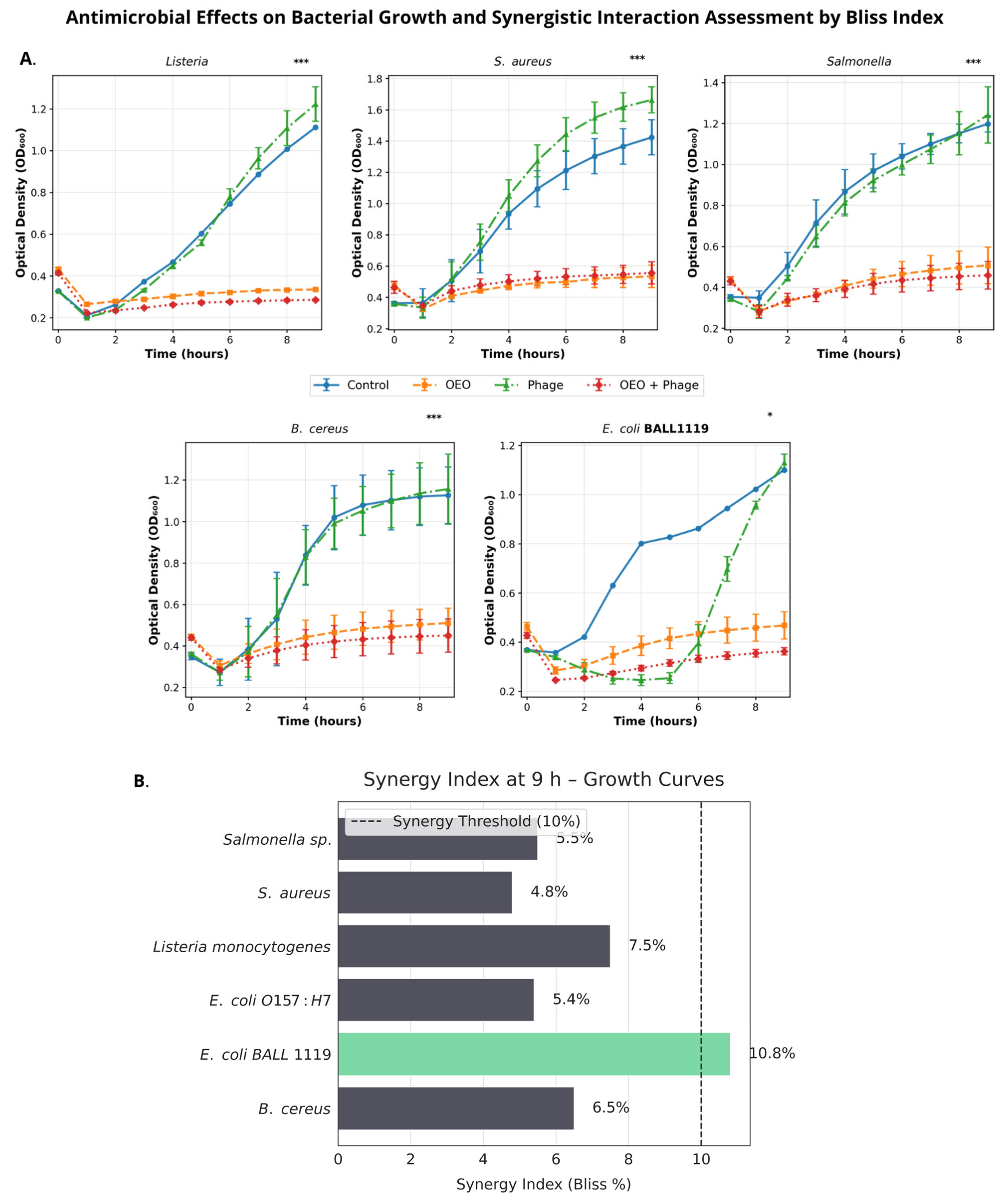
2.4.1. Growth Kinetics: Combined vs. Individual Treatments
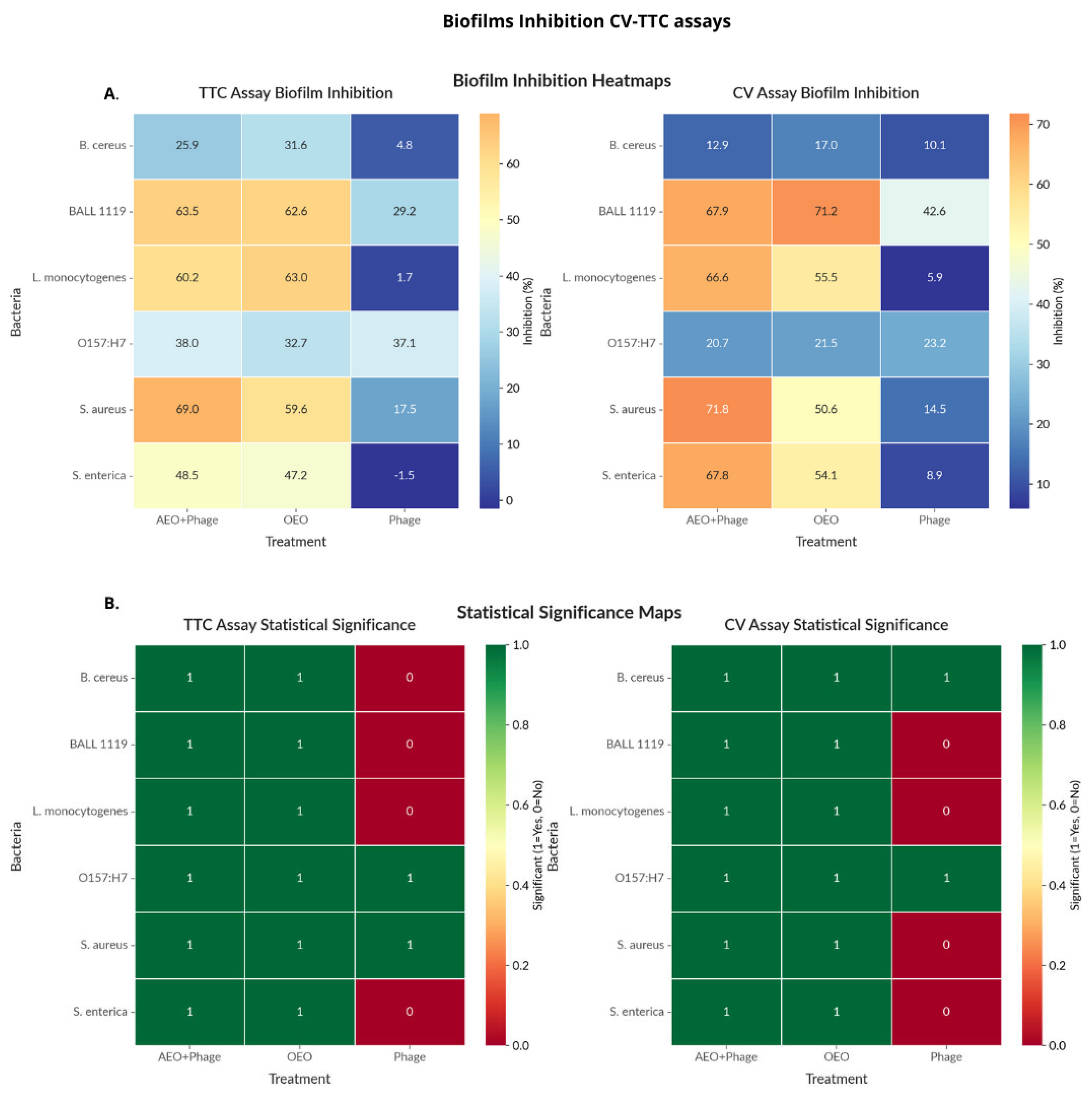
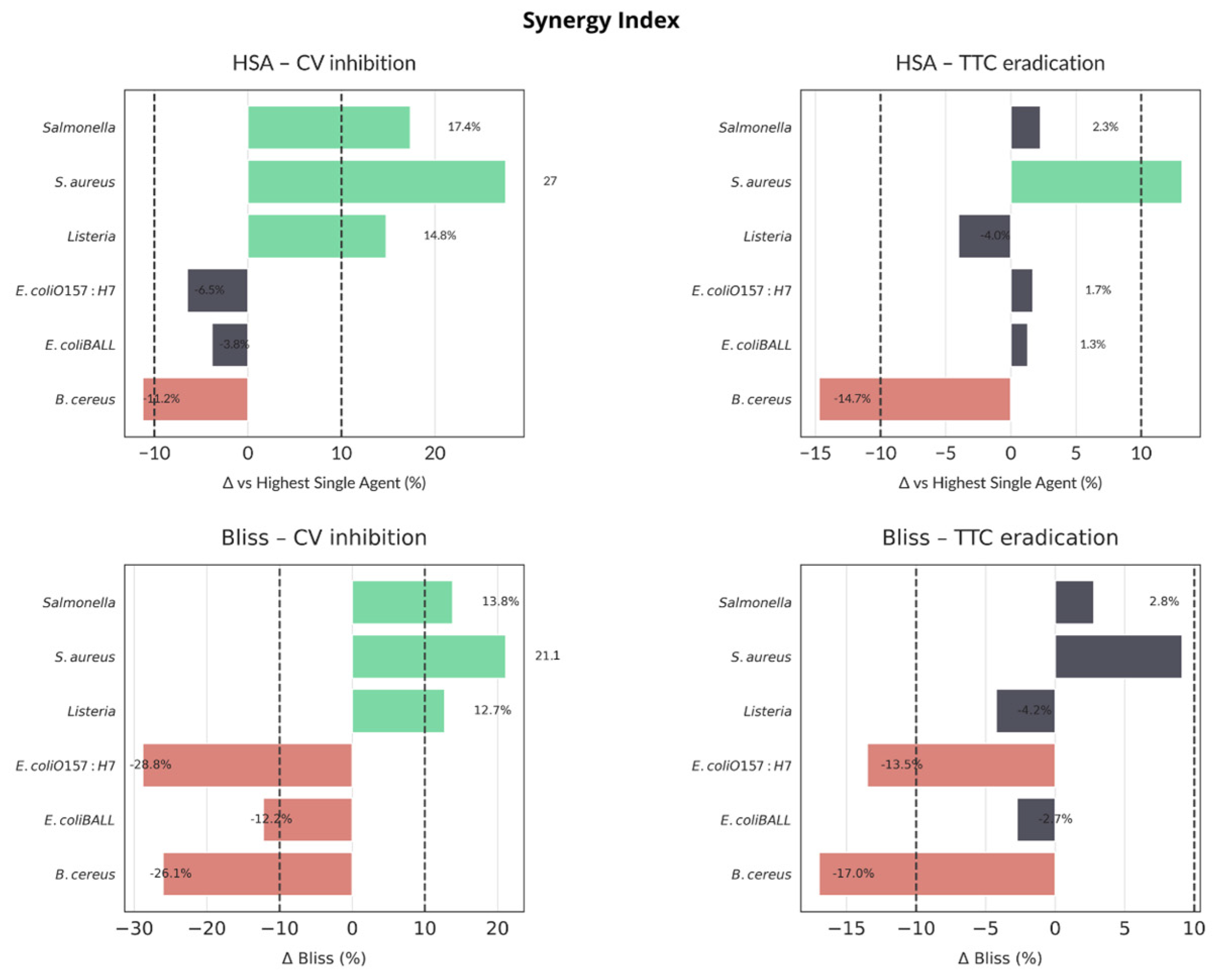
2.4.2. Biofilm Assays
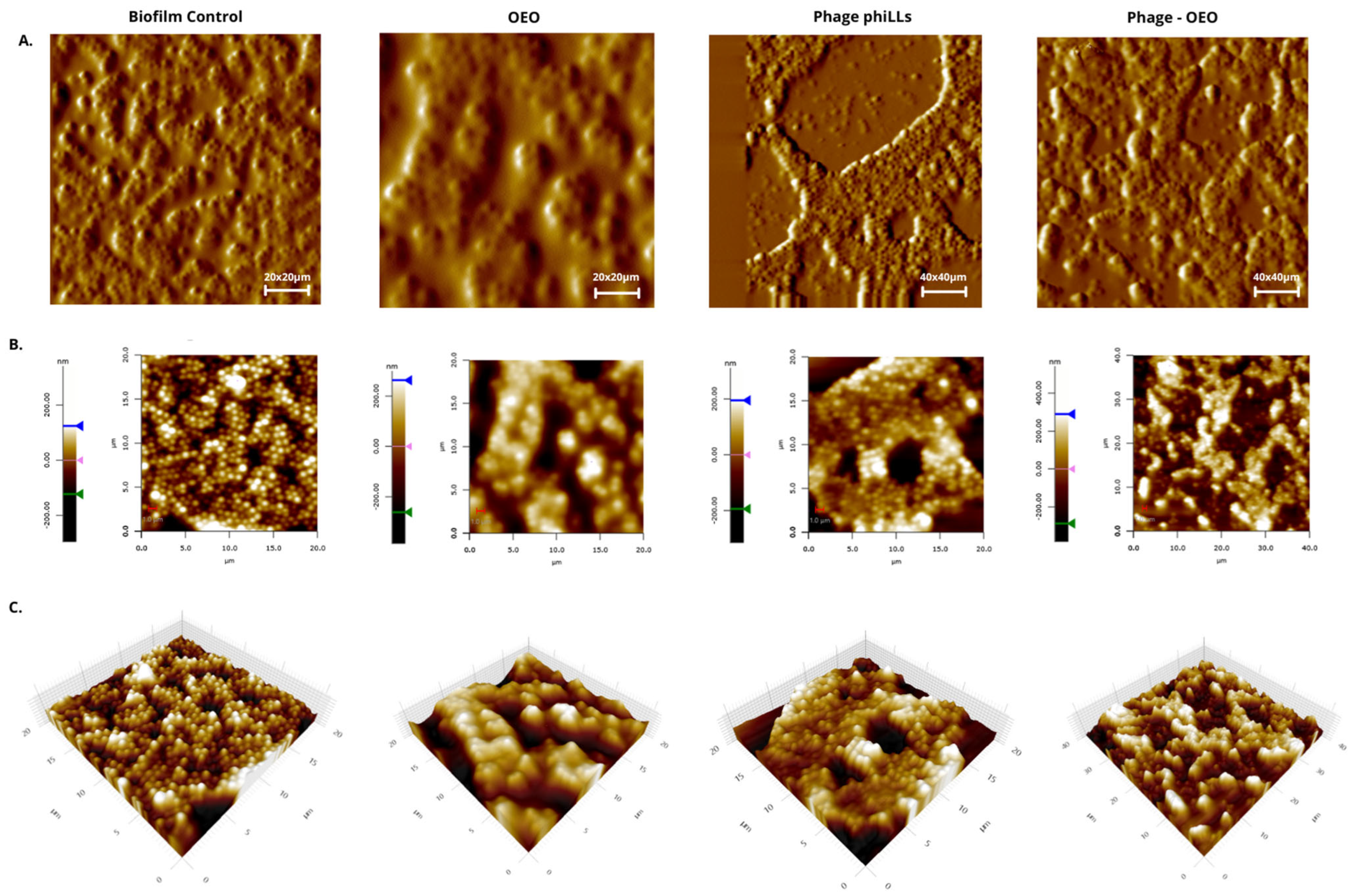
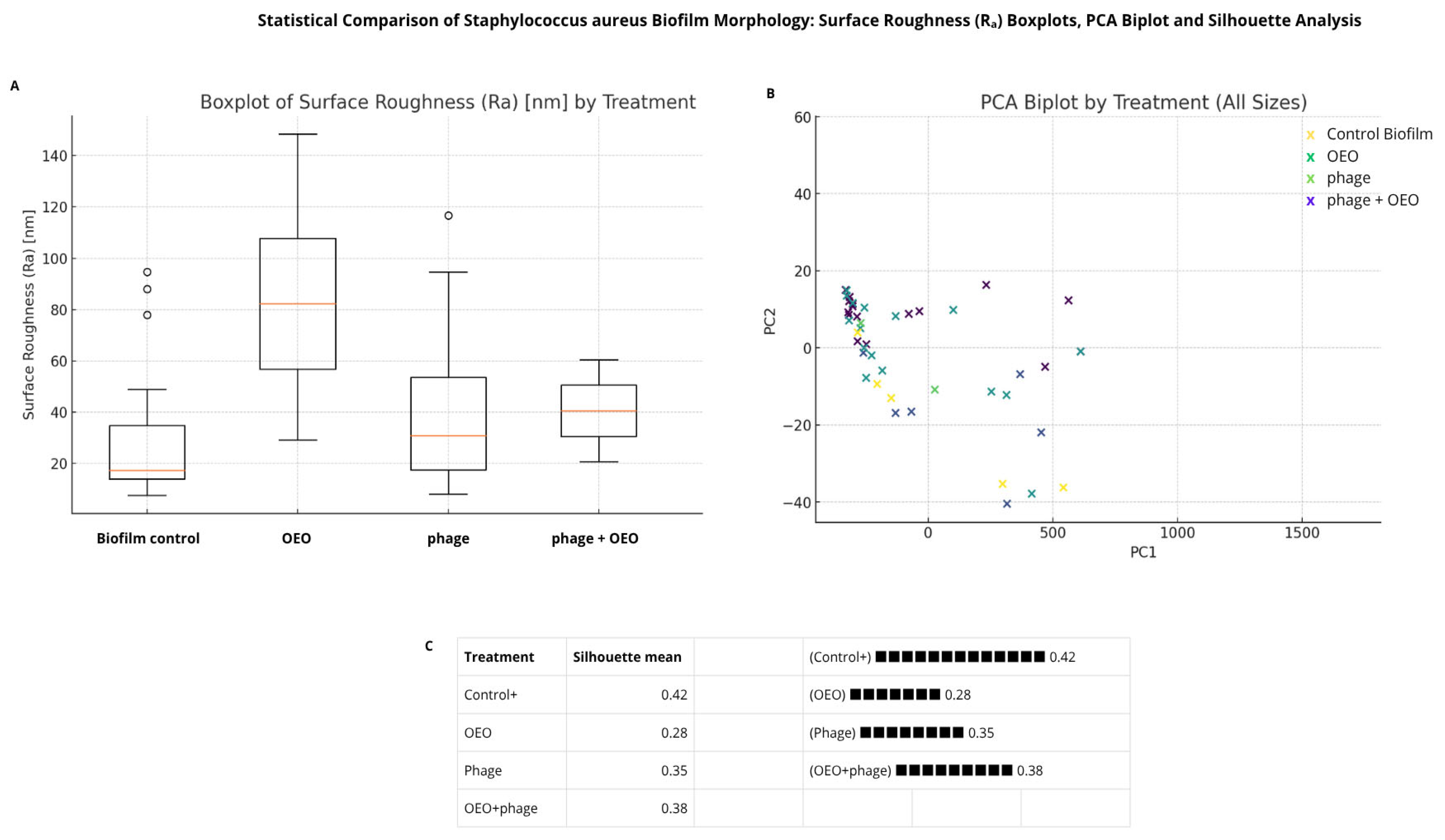
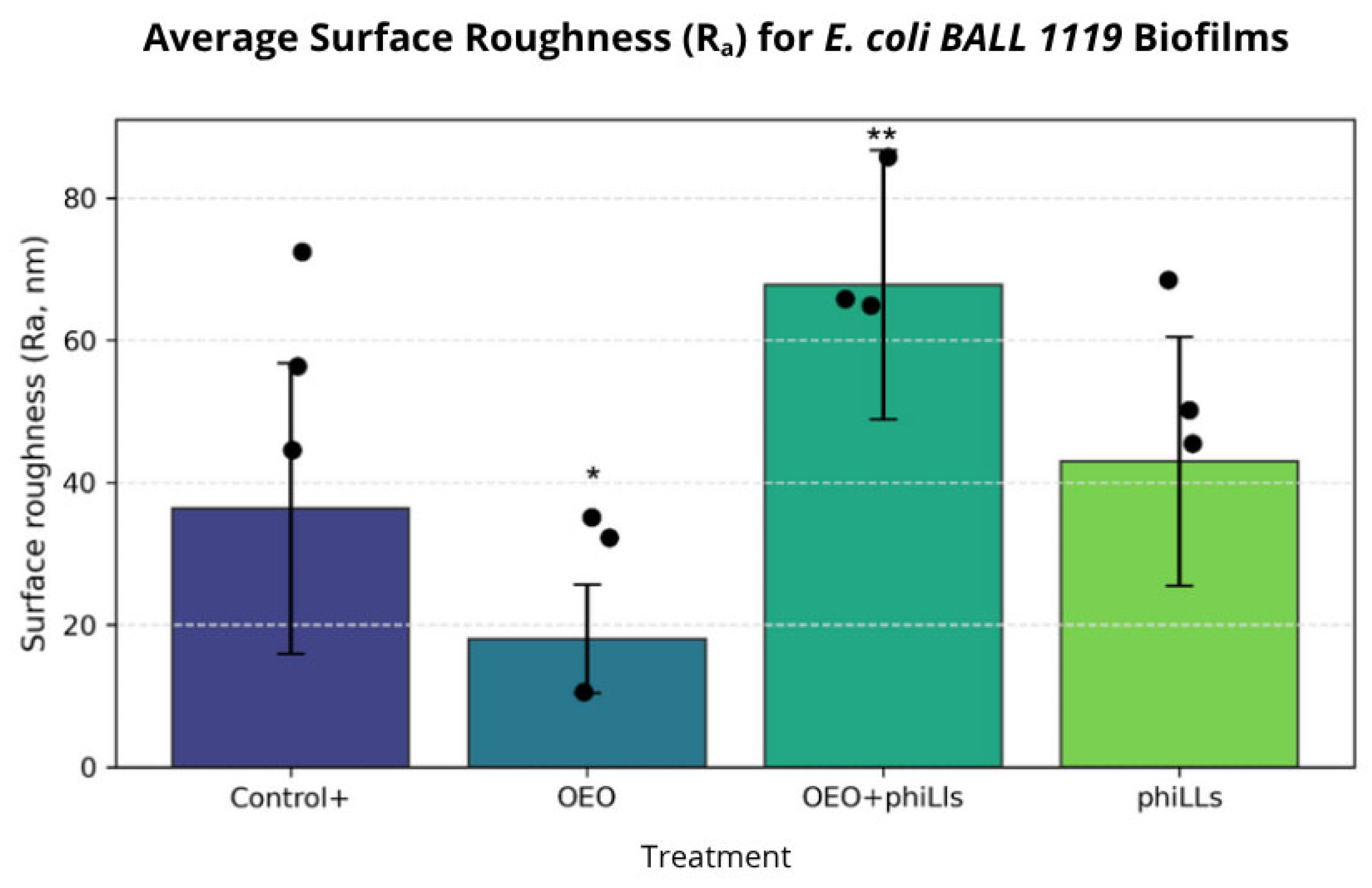
2.5. Evaluation of the Efficacy of the Combined phiLLS and OEO Under AFM for E. coli 1119 and S. aureus Biofilms
3. Discussion
3.1. Combined Application of OEO and Phage
3.2. Biofilm Inhibition Patterns Across Species
3.3. Methodological Considerations and Limitations
3.4. AFM-Based Insights on Biofilm Architecture Disruption
3.5. AFM-Based Insights into Antibiofilm Activity Against S. aureus and E. coli BALL 1119
3.6. Mechanistic Perspectives on Synergy and EPS Targeting
4. Materials and Methods
4.1. Bacterial Strains and Culture Conditions
4.2. Characterization of Oregano Essential Oil
Instrumentation and GC–MS Conditions
4.3. Phage Propagation
4.4. Determination of Phage Titer
4.5. Phage Sensitivity to OEO
4.6. Determination of MIC and MBC Values of OEO Against Foodborne Pathogens
4.7. Growth Kinetics Under Individual and Combined Treatments
– equal to the expected arithmetic/Bliss sum (additivity)
– smaller than expected (antagonism).
– –10% to +10% → essentially additive
– –10% or less → antagonism
4.8. Biofilm Eradication Assays Based on Residual Biomass and Metabolic Activity
4.9. AFM Analysis of Biofilms
4.10. Statistical Analysis
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Correction Statement
Abbreviations
| OEO | Oregano essential oil |
| phage | Bacteriophage |
| AFM | Atomic force microscopy |
| CV | Crystal violet |
| TTC | 2,3,5-Triphenyltetrazolium chloride |
| EPS | Extracellular polymeric substance |
| MIC | Minimum inhibitory concentration |
| MBC | Minimum bactericidal concentration |
| MOI | Multiplicity of infection |
| Ra | Average surface roughness |
| Rz | Maximum height variation |
| ΣFIC | Sum of fractional inhibitory concentration |
| Bliss | Bliss independence model (ΔBliss) |
| HSA | Highest single agent model (ΔHSA) |
| PFU | Plaque-forming unit |
| CFU | Colony-forming unit |
| PBS | Phosphate-buffered saline |
| TSB | Tryptic soy broth |
| TSA | Tryptic soy agar |
| MHB | Mueller–Hinton broth |
| MHA | Mueller–Hinton agar |
| PCA | Principal component analysis |
| OD6 | Optical density |
References
- Flemming, H.C.; Baveye, P.; Neu, T.R.; Stoodley, P.; Szewzyk, U.; Wingender, J.; Wuertz, S. Who Put the Film in Biofilm? The Migration of a Term from Wastewater Engineering to Medicine and Beyond. npj Biofilms Microbiomes 2021, 7, 10. [Google Scholar] [CrossRef]
- Boudarel, H.; Mathias, J.D.; Blaysat, B.; Grédiac, M. Towards Standardized Mechanical Characterization of Microbial Biofilms: Analysis and Critical Review. npj Biofilms Microbiomes 2018, 4, 17. [Google Scholar] [CrossRef]
- Flemming, H.-C.; Wingender, J. The Biofilm Matrix. Nat. Rev. Microbiol. 2010, 8, 623–633. [Google Scholar] [CrossRef] [PubMed]
- Van Houdt, R.; Michiels, C.W. Biofilm Formation and the Food Industry, a Focus on the Bacterial Outer Surface. J. Appl. Microbiol. 2010, 109, 1117–1131. [Google Scholar] [CrossRef]
- Alves, D.; Cerqueira, M.A.; Pastrana, L.M.; Sillankorva, S. Entrapment of a Phage Cocktail and Cinnamaldehyde on Sodium Alginate Emulsion-Based Films to Fight Food Contamination by Escherichia Coli and Salmonella Enteritidis. Food Res. Int. 2020, 128, 108791. [Google Scholar] [CrossRef] [PubMed]
- Hoelzer, K.; Moreno Switt, A.I.; Wiedmann, M.; Boor, K.J. Emerging Needs and Opportunities in Foodborne Disease Detection and Prevention: From Tools to People. Food Microbiol. 2018, 75, 65–71. [Google Scholar] [CrossRef]
- Panda, S.K.; Buroni, S.; Swain, S.S.; Bonacorsi, A.; da Fonseca Amorim, E.A.; Kulshrestha, M.; da Silva, L.C.N.; Tiwari, V. Recent Advances to Combat ESKAPE Pathogens with Special Reference to Essential Oils. Front. Microbiol. 2022, 13, 1029098. [Google Scholar] [CrossRef]
- WHO. Antimicrobial Resistance: Global Report on Surveillance; World Health Organization: Geneva, Switzerland, 2024. [Google Scholar]
- FAO; WHO. Food Control System Assessment Tool: Introduction and Glossary; Food Safety and Quality Series No. 7/1; World Health Organization: Geneva, Switzerland, 2019; ISBN 9789251316306. [Google Scholar]
- Oliveira, A.; Dias, C.; Oliveira, R.; Almeida, C.; Fuciños, P.; Sillankorva, S.; Oliveira, H. Paving the Way Forward: Escherichia Coli Bacteriophages in a One Health Approach. Crit. Rev. Microbiol. 2023, 50, 87–104. [Google Scholar] [CrossRef] [PubMed]
- Souza, G.M.; de Oliveira Vieira, K.C.; Naldi, L.V.; Pereira, V.C.; Winkelstroter, L.K. Green Synthesized Nanoparticles as a Promising Strategy for Controlling Microbial Biofilm. In Nanotechnology for Advances in Medical Microbiology; Springer: Singapore, 2021. [Google Scholar]
- Carstens, C.K.; Salazar, J.K.; Darkoh, C. Multistate Outbreaks of Foodborne Illness in the United States Associated With Fresh Produce From 2010 to 2017. Front. Microbiol. 2019, 10, 2667. [Google Scholar] [CrossRef]
- Tack, D.M.; Ray, L.; Griffin, P.M.; Cieslak, P.R.; Dunn, J.; Rissman, T.; Jervis, R.; Lathrop, S.; Muse, A.; Duwell, M.; et al. Preliminary Incidence and Trends of Infections with Pathogens Transmitted Commonly Through Food—Foodborne Diseases Active Surveillance Network, 10 U.S. Sites, 2016–2019. MMWR Morb. Mortal. Wkly. Rep. 2020, 69, 509–514. [Google Scholar] [CrossRef]
- Crowe, S.J.; Mahon, B.E.; Vieira, A.R.; Gould, L.H. Vital Signs: Multistate Foodborne Outbreaks—United States, 2010–2014. MMWR Morb. Mortal. Wkly. Rep. 2015, 64, 1221–1225. [Google Scholar] [CrossRef] [PubMed]
- FAO. FAO Guide to Ranking Food Safety Risks at the National Level; Food and Agriculture Organization of the United Nations: Rome, Italy, 2020; ISBN 9789251332825. [Google Scholar]
- Jim, O. Tackling Drug-Resistant Infections Globally: Final Report and Recommendations; AMR Review: Sydney, Australia, 2016. [Google Scholar]
- Mah, T.F. Biofilm-Specific Antibiotic Resistance. Future Microbiol. 2012, 7, 1061–1072. [Google Scholar] [CrossRef]
- Moreau-Marquis, S.; Bomberger, J.M.; Anderson, G.G.; Swiatecka-Urban, A.; Ye, S.; O’TOole, G.A.; Stanton, B.A. The F508-CFTR Mutation Results in Increased Biofilm Formation by Pseudomonas Aeruginosa by Increasing Iron Availability. Am. J. Physiol. Lung Cell. Mol. Physiol. 2008, 295, L25–L37. [Google Scholar] [CrossRef] [PubMed]
- Bai, X.; Nakatsu, C.H.; Bhunia, A.K. Bacterial Biofilms and Their Implications in Pathogenesis and Food Safety. Foods 2021, 10, 2117. [Google Scholar] [CrossRef]
- Lambert, R.J.W.; Skandamis, P.N.; Coote, P.J.; Nychas, G.-J.E. A Study of the Minimum Inhibitory Concentration and Mode of Action of Oregano Essential Oil, Thymol and Carvacrol. J. Appl. Microbiol. 2001, 91, 453–462. [Google Scholar] [CrossRef]
- Ultee, A.; Bennik, M.H.J.; Moezelaar, R. The Phenolic Hydroxyl Group of Carvacrol Is Essential for Action against the Food-Borne Pathogen Bacillus Cereus. Appl. Environ. Microbiol. 2002, 68, 1561–1568. [Google Scholar] [CrossRef]
- Chouhan, S.; Sharma, K.; Guleria, S. Antimicrobial Activity of Some Essential Oils—Present Status and Future Perspectives. Medicines 2017, 4, 58. [Google Scholar] [CrossRef]
- Guo, Z.; Liu, M.; Zhang, D. Potential of Phage Depolymerase for the Treatment of Bacterial Biofilms. Virulence 2023, 14, 2273567. [Google Scholar] [CrossRef] [PubMed]
- Wintachai, P.; Voravuthikunchai, S.P. Characterization of Novel Lytic Myoviridae Phage Infecting Multidrug-Resistant Acinetobacter Baumannii and Synergistic Antimicrobial Efficacy between Phage and Sacha Inchi Oil. Pharmaceuticals 2022, 15, 291. [Google Scholar] [CrossRef]
- Amarillas, L.; Rubí-Rangel, L.; Chaidez, C.; González-Robles, A.; Lightbourn-Rojas, L.; León-Félix, J. Isolation and Characterization of PhiLLS, a Novel Phage with Potential Biocontrol Agent against Multidrug-Resistant Escherichia Coli. Front. Microbiol. 2017, 8, 1355. [Google Scholar] [CrossRef]
- Topka-Bielecka, G.; Dydecka, A.; Necel, A.; Bloch, S.; Nejman-Faleńczyk, B.; Węgrzyn, G.; Węgrzyn, A. Bacteriophage-Derived Depolymerases against Bacterial Biofilm. Antibiotics 2021, 10, 175. [Google Scholar] [CrossRef]
- Amankwah, S.; Abdella, K.; Kassa, T. Bacterial Biofilm Destruction: A Focused Review on the Recent Use of Phage-Based Strategies with Other Antibiofilm Agents. Nanotechnol. Sci. Appl. 2021, 14, 161–177. [Google Scholar] [CrossRef]
- Pei, R.; Lamas-Samanamud, G.R. Inhibition of Biofilm Formation by T7 Bacteriophages Producing Quorum-Quenching Enzymes. Appl. Environ. Microbiol. 2014, 80, 5340–5348. [Google Scholar] [CrossRef]
- Ali, S.; Karaynir, A.; Doğan, H.S.; Nachimuthu, R.; Badu, K.; Bozdoğan, B. Degradation of Preformed Gram-Positive and Gram-Negative Bacterial Biofilms Using Disintegrated and Intact Phages. PHAGE 2025, 6, 20–31. [Google Scholar] [CrossRef]
- Ni, P.; Wang, L.; Deng, B.; Jiu, S.; Ma, C.; Zhang, C.; Almeida, A.; Wang, D.; Xu, W.; Wang, S. Combined Application of Bacteriophages and Carvacrol in the Control of Pseudomonas Syringae Pv. Actinidiae Planktonic and Biofilm Forms. Microorganisms 2020, 8, 837. [Google Scholar] [CrossRef]
- Ben Said, M.; Trabelsi, D.; Achouri, F.; Saad, M.B.; Bousselmi, L.; Ghrabi, A. Application of Bacteriophage and Essential Oil to Monitor Bacterial Biofilm Formation. In Recent Advances in Environmental Science from the Euro-Mediterranean and Surrounding Regions; Springer: Cham, Switzerland, 2018; pp. 273–274. [Google Scholar]
- Haney, E.F.; Trimble, M.J.; Hancock, R.E.W. Microtiter Plate Assays to Assess Antibiofilm Activity against Bacteria. Nat. Protoc. 2021, 16, 2615–2632. [Google Scholar] [CrossRef] [PubMed]
- Villalobos-delgado, L.H.; Caro, I.; Quinto, E.J.; Mateo, J. Natural Antimicrobial Agents to Improve Foods Shelf Life; Elsevier Inc.: Amsterdam, The Netherlands, 2019; ISBN 9780128171905. [Google Scholar]
- Alexander, F.; Agudelo, G.; Ospina, J.N.; Mary, L.; Pulido, S. Innovaciones En La Terapia Antimicrobiana Innovations in Antimicrobial Therapy. Nova 2020, 18, 9–25. [Google Scholar]
- Luna Guevara, M.L.; Luna Guevara, J.J.; Espinosa, H.R.; Abascal, L.L.; Díaz González, C.B. Eficiencia de La Desinfección Con Aceites Esenciales y Ultrasonido Sobre Escherichia Coli Inoculada En Frutos de Tomate y El Impacto Sobre La Actividad Antioxidante. Rev. Argent. Microbiol. 2015, 47, 251–255. [Google Scholar] [CrossRef] [PubMed]
- Yang, S.; Huang, Q.; Cai, P. Application of Atomic Force Microscopy (AFM) to Study Bacterial Biofilms. Shengwu Gongcheng Xuebao/Chin. J. Biotechnol. 2017, 33, 1399–1410. [Google Scholar] [CrossRef]
- Wang, X.; Xue, J.; Zhu, H.; Chen, S.; Wang, Y.; Xiao, Z.; Luo, Y. Advances in Biofilm Characterization: Utilizing Rheology and Atomic Force Microscopy in Foods and Related Fields. Adv. Compos. Hybrid. Mater. 2024, 7, 143. [Google Scholar] [CrossRef]
- Kropinski, A.M.; Mazzocco, A.; Waddell, T.E.; Lingohr, E.; Johnson, R.P. Enumeration of Bacteriophages by Double Agar Overlay Plaque Assay. In Bacteriophages: Methods and Protocols, Volume 1: Isolation, Characterization, and Interactions; Clokie, M.R.J., Kropinski, A.M., Eds.; Humana Press: Totowa, NJ, USA, 2009; pp. 69–76. ISBN 978-1-60327-164-6. [Google Scholar]
- Lehár, J.; Zimmermann, G.R.; Krueger, A.S.; Molnar, R.A.; Ledell, J.T.; Heilbut, A.M.; Short, G.F.; Giusti, L.C.; Nolan, G.P.; Magid, O.A.; et al. Chemical Combination Effects Predict Connectivity in Biological Systems. Mol. Syst. Biol. 2007, 3, 80. [Google Scholar] [CrossRef]
- Bliss, C.I. The Toxicity of Poisons Applied Jointly. Ann. Appl. Biol. 1939, 26, 585–615. [Google Scholar] [CrossRef]
- Eaton, P.; Fernandes, J.C.; Pereira, E.; Pintado, M.E.; Xavier Malcata, F. Atomic Force Microscopy Study of the Antibacterial Effects of Chitosans on Escherichia Coli and Staphylococcus Aureus. Ultramicroscopy 2008, 108, 1128–1134. [Google Scholar] [CrossRef]
- Perry, C.C.; Weatherly, M.; Beale, T.; Randriamahefa, A. Atomic Forcemicroscopy Study of the Antimicrobial Activity of Aqueous Garlic versus Ampicillin against Escherichia Coli and Staphylococcus Aureus. J. Sci. Food Agric. 2009, 89, 958–964. [Google Scholar] [CrossRef]
- Pei, Z.J.; Li, C.; Dai, W.; Lou, Z.; Sun, X.; Wang, H.; Khan, A.A.; Wan, C. The Anti-Biofilm Activity and Mechanism of Apigenin-7-O-Glucoside Against Staphylococcus Aureus and Escherichia Coli. Infect. Drug Resist. 2023, 16, 2129–2140. [Google Scholar] [CrossRef]
- Liu, C.G.; Green, S.I.; Min, L.; Clark, J.R.; Salazar, K.C.; Terwilliger, A.L.; Kaplan, H.B.; Trautner, B.W.; Ramig, R.F.; Maresso, A.W. Phage-Antibiotic Synergy Is Driven by a Unique Combination of Antibacterial Mechanism of Action and Stoichiometry. mBio 2020, 11, e01462-20. [Google Scholar] [CrossRef]
- Alavi, M.; Karimi, N.; Salimikia, I. Phytosynthesis of Zinc Oxide Nanoparticles and Its Antibacterial, Antiquorum Sensing, Antimotility, and Antioxidant Capacities against Multidrug Resistant Bacteria. J. Ind. Eng. Chem. 2019, 72, 457–473. [Google Scholar] [CrossRef]
- AL-Dujaily, A.H.; Mahmood, A.K. Evaluation of Antibacterial and Antibiofilm Activity of Biogenic Silver Nanoparticles and Gentamicin Against Staphylococcus Aureus Isolated from Caprine Mastitis. Iraqi J. Vet. Med. 2022, 46, 10–16. [Google Scholar] [CrossRef]
- Mahmoudi-Qashqay, S.; Zamani-Meymian, M.R.; Sadati, S.J. Improving Antibacterial Ability of Ti-Cu Thin Films with Co-Sputtering Method. Sci. Rep. 2023, 13, 16593. [Google Scholar] [CrossRef]
- Odds, F.C. Synergy, Antagonism, and What the Chequerboard Puts between Them. J. Antimicrob. Chemother. 2003, 52, 1. [Google Scholar] [CrossRef] [PubMed]
- Khong, E.; Oh, J.J.; Jimenez, J.M.; Liu, R.; Dunham, S.; Monsibais, A.; Rhoads, A.; Ghatbale, P.; Garcia, A.; Cobián Güemes, A.G.; et al. A Simple Solid Media Assay for Detection of Synergy between Bacteriophages and Antibiotics. Microbiol. Spectr. 2024, 12, e0322123. [Google Scholar] [CrossRef] [PubMed]
- Valério, N.; Oliveira, C.; Jesus, V.; Branco, T.; Pereira, C.; Moreirinha, C.; Almeida, A. Effects of Single and Combined Use of Bacteriophages and Antibiotics to Inactivate Escherichia Coli. Virus Res. 2017, 240, 8–17. [Google Scholar] [CrossRef]
- Azeredo, J.; García, P.; Drulis-Kawa, Z. Targeting Biofilms Using Phages and Their Enzymes. Curr. Opin. Biotechnol. 2021, 68, 251–261. [Google Scholar] [CrossRef]
- Knecht, L.E.; Veljkovic, M.; Fieseler, L. Diversity and Function of Phage Encoded Depolymerases. Front. Microbiol. 2020, 10, 2949. [Google Scholar] [CrossRef]
- Janesomboon, S.; Sawaengwong, T.; Muangsombut, V.; Vanaporn, M.; Santanirand, P.; Kritsiriwuthinan, K.; Gundogdu, O.; Chantratita, N.; Nale, J.Y.; Korbsrisate, S.; et al. Synergistic Antibacterial Activity of Curcumin and Phage against Multidrug-Resistant Acinetobacter Baumannii. Sci. Rep. 2025, 15, 8959. [Google Scholar] [CrossRef]
- Gunn, J.; Bakaletz, L.; Wozniak, D. What’s on the Outside Matters: The Role of the Extracellular Polymeric Substance of Gram-Negative Biofilms in Evading Host Immunity and as a Target for Therapeutic Intervention. J. Biol. Chem. 2016, 291, 12538–12546. [Google Scholar] [CrossRef]
- Ruhal, R.; Kataria, R. Biofilm Patterns in Gram-Positive and Gram-Negative Bacteria. Microbiol. Res. 2021, 251, 126829. [Google Scholar] [CrossRef]
- Cheeseman, S.; Shaw, Z.; Vongsvivut, J.; Crawford, R.; Dupont, M.; Boyce, K.; Gangadoo, S.; Bryant, S.; Bryant, G.; Cozzolino, D.; et al. Analysis of Pathogenic Bacterial and Yeast Biofilms Using the Combination of Synchrotron ATR-FTIR Microspectroscopy and Chemometric Approaches. Molecules 2021, 26, 3890. [Google Scholar] [CrossRef]
- Mahdhi, A.; Leban, N.; Chakroun, I.; Chaouch, M.A.; Hafsa, J.; Fdhila, K.; Mahdouani, K.; Majdoub, H. Extracellular Polysaccharide Derived from Potential Probiotic Strain with Antioxidant and Antibacterial Activities as a Prebiotic Agent to Control Pathogenic Bacterial Biofilm Formation. Microb. Pathog. 2017, 109, 214–220. [Google Scholar] [CrossRef] [PubMed]
- Harimawan, A.; Ting, Y.P. Investigation of Extracellular Polymeric Substances (EPS) Properties of P. Aeruginosa and B. Subtilis and Their Role in Bacterial Adhesion. Colloids Surf. B Biointerfaces 2016, 146, 459–467. [Google Scholar] [CrossRef]
- Misba, L.; Zaidi, S.; Khan, A. A Comparison of Antibacterial and Antibiofilm Efficacy of Phenothiazinium Dyes between Gram Positive and Gram Negative Bacterial Biofilm. Photodiagnosis Photodyn. Ther. 2017, 18, 24–33. [Google Scholar] [CrossRef]
- Sinha, S.; Choudhuri, M.; Basu, T.; Gupta, D.; Datta, A. Decisive Role of Polymer-Bovine Serum Albumin Interactions in Biofilm Substrates on “Philicity” and Extracellular Polymeric Substances Composition. Langmuir 2022, 38, 1966–1976. [Google Scholar] [CrossRef] [PubMed]
- Fulaz, S.; Vitale, S.; Quinn, L.; Casey, E. Nanoparticle-Biofilm Interactions: The Role of the EPS Matrix. Trends Microbiol. 2019, 27, 915–926. [Google Scholar] [CrossRef]
- Pinto, R.; Soares, F.; Reis, S.; Nunes, C.; Van Dijck, P. Innovative Strategies Toward the Disassembly of the EPS Matrix in Bacterial Biofilms. Front. Microbiol. 2020, 11, 952. [Google Scholar] [CrossRef] [PubMed]
- Kavanaugh, J.; Flack, C.; Lister, J.; Ricker, E.; Ibberson, C.; Jenul, C.; Moormeier, D.; Delmain, E.; Bayles, K.; Horswill, A. Identification of Extracellular DNA-Binding Proteins in the Biofilm Matrix. mBio 2019, 10, e01137-19. [Google Scholar] [CrossRef]
- Karygianni, L.; Ren, Z.; Koo, H.; Thurnheer, T. Biofilm Matrixome: Extracellular Components in Structured Microbial Communities. Trends Microbiol. 2020, 28, 668–681. [Google Scholar] [CrossRef]
- Tomlinson, K.; Lung, T.W.F.; Dach, F.; Annavajhala, M.; Gabryszewski, S.; Groves, R.; Drikic, M.; Francoeur, N.; Sridhar, S.; Smith, M.; et al. Staphylococcus Aureus Induces an Itaconate-Dominated Immunometabolic Response That Drives Biofilm Formation. Nat. Commun. 2021, 12, 1399. [Google Scholar] [CrossRef]
- Salim, S.; Jadoon, V.; Batool, M.; Ali, I.; Irshad, U.; Ahmed, R.; Qayum, S.; Noor, A.; Naqvi, T. A DNase from Halophilic Bacterium Bacillus Pacificus Targets Two Notorious Biofilms of Pseudomonas Aeruginosa PAO1 and Staphylococcus Aureus. Biocatal. Agric. Biotechnol. 2025, 64, 103515. [Google Scholar] [CrossRef]
- Bai, X.; Shen, Y.; Zhang, T.; Meng, R.; Zhang, Y.; Deng, Y.; Guo, N. Anti-Biofilm Activity of Biochanin A against Staphylococcus Aureus. Appl. Microbiol. Biotechnol. 2022, 107, 867–879. [Google Scholar] [CrossRef]
- Gupta, A.; Guptasarma, P.E. Coli Cells Advance into Phase-Separated (Biofilm-Simulating) Extracellular Polymeric Substance Containing DNA, HU, and Lipopolysaccharide. J. Bacteriol. 2024, 206, e0030924. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.; Oh, S.; Kim, S. Released Exopolysaccharide (r-EPS) Produced from Probiotic Bacteria Reduce Biofilm Formation of Enterohemorrhagic Escherichia Coli O157:H7. Biochem. Biophys. Res. Commun. 2009, 379, 324–329. [Google Scholar] [CrossRef] [PubMed]
- Hu, X.; Zhang, Y.; Chen, Z.; Gao, Y.; Teppen, B.; Boyd, S.; Zhang, W.; Tiedje, J.; Li, H. Tetracycline Accumulation in Biofilms Enhances the Selection Pressure on Escherichia Coli for Expression of Antibiotic Resistance. Sci. Total Environ. 2022, 857, 159441. [Google Scholar] [CrossRef] [PubMed]
- Park, D.W.; Lee, J.H.; Park, J.H. Thymol and Eugenol in Essential Oils Enhance Phage Endolysin LysECP26-Mediated Cell Wall Disruption of Escherichia Coli O157:H7. Korean J. Food Sci. Technol. 2021, 53, 819–822. [Google Scholar] [CrossRef]
- Park, D.W.; Park, J.H. Characterization of a Novel Phage Depolymerase Specific to Escherichia Coli O157:H7 and Biofilm Control on Abiotic Surfaces. J. Microbiol. 2021, 59, 1002–1009. [Google Scholar] [CrossRef] [PubMed]

| Compound | Chemical Class | RT (min) | R-Match | % Composition |
|---|---|---|---|---|
| α-Pinene | Monoterpene hydrocarbons | 3.906 | 849 | 0.870 |
| β-Myrcene | Monoterpene hydrocarbons | 4.196 | 841 | 1.750 |
| p-Cymene | Monoterpene hydrocarbons | 4.364 | 816 | 28.616 |
| γ-Terpinene | Monoterpene hydrocarbons | 4.589 | 798 | 2.885 |
| Terpinen-4-ol | Oxygenated monoterpenes | 5.657 | 717 | 0.739 |
| Methylthymol | Phenolic derivatives/esters | 5.926 | 910 | 0.295 |
| Isothymol methyl ester | Phenolic derivatives/esters | 6.006 | 915 | 0.198 |
| Thymol | Phenolic monoterpenes | 6.865 | 750 | 0.174 |
| Carvacrol | Phenolic monoterpenes | 6.520 | 750 | 58.897 |
| Caryophyllene | Sesquiterpene hydrocarbons | 7.350 | 783 | 2.593 |
| Humulene | Sesquiterpene hydrocarbons | 7.615 | 700 | 2.065 |
| Aromadendrene | Sesquiterpene hydrocarbons | 7.831 | 846 | 0.323 |
| Caryophyllene oxide | Oxygenated sesquiterpenes | 8.702 | 843 | 0.597 |
| 48 h | OEO 2 mg/mL | OEO 1 mg/mL | OEO 0.5 mg/mL | OEO 0.25 mg/mL |
|---|---|---|---|---|
| E. coli BALL 1119 | − | + | + | + |
| L. monocytogenes | − | − | + | + |
| Salmonella spp. | − | − | + | + |
| S. aureus | − | − | + | + |
| E. coli O157:H7 | − | − | + | + |
| B. cereus | − | − | + | + |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kao Godínez, A.K.; Regalado-González, C.; Villicaña, C.; Heredia, J.B.; Valdez-Torres, J.B.; Muy-Rangel, M.; Escamilla-García, M.; León-Félix, J. Synergistic Disruption of Foodborne Pathogen Biofilms by Oregano Essential Oil and Bacteriophage phiLLS: Atomic Force Microscopy Insights. Molecules 2025, 30, 3552. https://doi.org/10.3390/molecules30173552
Kao Godínez AK, Regalado-González C, Villicaña C, Heredia JB, Valdez-Torres JB, Muy-Rangel M, Escamilla-García M, León-Félix J. Synergistic Disruption of Foodborne Pathogen Biofilms by Oregano Essential Oil and Bacteriophage phiLLS: Atomic Force Microscopy Insights. Molecules. 2025; 30(17):3552. https://doi.org/10.3390/molecules30173552
Chicago/Turabian StyleKao Godínez, Ana Karina, Carlos Regalado-González, Claudia Villicaña, José Basilio Heredia, José Benigno Valdez-Torres, María Muy-Rangel, Monserrat Escamilla-García, and Josefina León-Félix. 2025. "Synergistic Disruption of Foodborne Pathogen Biofilms by Oregano Essential Oil and Bacteriophage phiLLS: Atomic Force Microscopy Insights" Molecules 30, no. 17: 3552. https://doi.org/10.3390/molecules30173552
APA StyleKao Godínez, A. K., Regalado-González, C., Villicaña, C., Heredia, J. B., Valdez-Torres, J. B., Muy-Rangel, M., Escamilla-García, M., & León-Félix, J. (2025). Synergistic Disruption of Foodborne Pathogen Biofilms by Oregano Essential Oil and Bacteriophage phiLLS: Atomic Force Microscopy Insights. Molecules, 30(17), 3552. https://doi.org/10.3390/molecules30173552









