CONSMI: Contrastive Learning in the Simplified Molecular Input Line Entry System Helps Generate Better Molecules
Abstract
1. Introduction
- 1.
- We propose CONSMI, a contrastive learning framework that learns representation from a large molecular dataset.
- 2.
- The CONSMI framework combined with a transformer decoder generates more successful molecules.
- 3.
- The CONSMI framework combined with a transformer encoder achieves SOTA results on multiple datasets of compound–protein interactions.
2. Results and Discussion
2.1. Molecular Generation Results
- Case Study We systematically examined all generated molecules with QED values exceeding 0.9 and assessed their similarity to the molecules in the Moses test set. As shown in Table 3 and Table 4, We found that although the model has not seen the molecules in the test set, a substantial number of the generated molecules exhibited a similarity of over 0.9 to those in the test set.
- Adjusting the Temperature We conducted an evaluation to understand how adjustments to the hyperparameter impact NT-Xent’s ability to distinguish effectively between positive and negative samples. This assessment involved exploring a set of values that are frequently used in practice, specifically 0.05, 0.1, 0.5, and 1, to identify the most suitable value. The results, encapsulated in Table 5, display the validation errors corresponding to each value during the model’s training process. We found that a of 0.1 yielded the most favorable results. Intriguingly, these results corroborate well with the existing literature, particularly concerning applications in molecular data [48].
2.2. Compound–Protein Interaction Results
3. Methods
3.1. CONSMI Framework
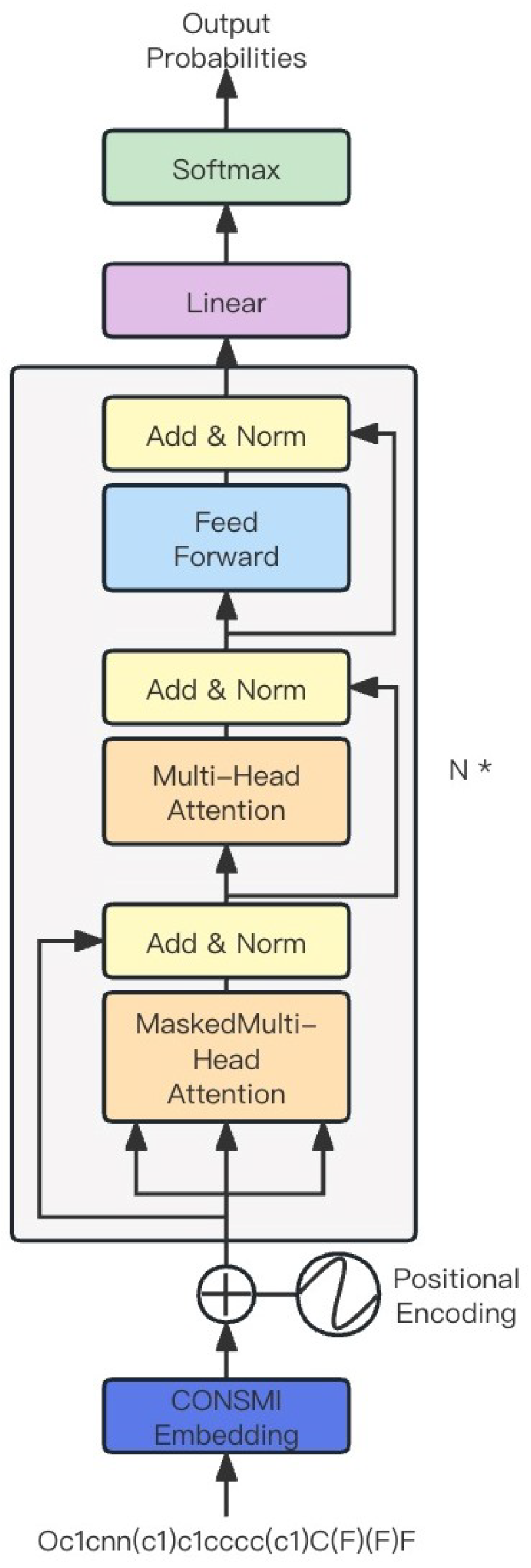
3.2. Generation Module
3.3. Classifier Module
4. Experiment Configuration
4.1. Datasets
4.2. Evaluation Metrics
4.3. Training Details
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Lee, S.I.; Celik, S.; Logsdon, B.A.; Lundberg, S.M.; Martins, T.J.; Oehler, V.G.; Estey, E.H.; Miller, C.P.; Chien, S.; Dai, J.a. A machine learning approach to integrate big data for precision medicine in acute myeloid leukemia. Nat. Commun. 2018, 9, 42. [Google Scholar] [CrossRef]
- Dimasi, J.A.; Grabowski, H.G.; Hansen, R.W. Innovation in the pharmaceutical industry: New estimates of R&D costs. J. Health Econ. 2016, 47, 20–33. [Google Scholar] [PubMed]
- Polishchuk, P.G.; Madzhidov, T.I.; Varnek, A. Estimation of the size of drug-like chemical space based on GDB-17 data. J. Comput.-Aided Mol. Des. 2013, 27, 675–679. [Google Scholar] [CrossRef]
- Sunghwan, K.; Thiessen, P.A.; Bolton, E.E.; Jie, C.; Gang, F.; Asta, G.; Lianyi, H.; Jane, H.; Siqian, H.; Shoemaker, B.A. PubChem Substance and Compound databases. Nucleic Acids Res. 2016, 44, D1202–D1213. [Google Scholar]
- Yoshikawa, N.; Terayama, K.; Sumita, M.; Homma, T.; Oono, K.; Tsuda, K. Population-based de novo molecule generation, using grammatical evolution. Chem. Lett. 2018, 47, 1431–1434. [Google Scholar] [CrossRef]
- Verhellen, J. Graph-based molecular Pareto optimisation. Chem. Sci. 2022, 13, 7526–7535. [Google Scholar] [CrossRef] [PubMed]
- Lamanna, G.; Delre, P.; Marcou, G.; Saviano, M.; Varnek, A.; Horvath, D.; Mangiatordi, G.F. GENERA: A combined genetic/deep-learning algorithm for multiobjective target-oriented de novo design. J. Chem. Inf. Model. 2023, 63, 5107–5119. [Google Scholar] [CrossRef]
- Creanza, T.M.; Lamanna, G.; Delre, P.; Contino, M.; Corriero, N.; Saviano, M.; Mangiatordi, G.F.; Ancona, N. DeLA-Drug: A deep learning algorithm for automated design of druglike analogues. J. Chem. Inf. Model. 2022, 62, 1411–1424. [Google Scholar] [CrossRef]
- Goodfellow, I.; Pouget-Abadie, J.; Mirza, M.; Xu, B.; Warde-Farley, D.; Ozair, S.; Courville, A.; Bengio, Y. Generative adversarial nets Advances in neural information processing systems. arXiv 2014, arXiv:1406.2661. [Google Scholar]
- Vaswani, A.; Shazeer, N.; Parmar, N.; Uszkoreit, J.; Jones, L.; Gomez, A.N.; Kaiser, Ł.; Polosukhin, I. Attention is all you need. Adv. Neural Inf. Process. Syst. 2017, 30, 1–11. [Google Scholar]
- Krenn, M.; Häse, F.; Nigam, A.; Friederich, P.; Aspuru-Guzik, A. Self-referencing embedded strings (SELFIES): A 100% robust molecular string representation. Mach. Learn. Sci. Technol. 2020, 1, 045024. [Google Scholar] [CrossRef]
- Lim, J.; Hwang, S.Y.; Kim, S.; Moon, S.; Kim, W.Y. Scaffold-based molecular design using graph generative model. Chem. Sci. 2020, 11, 1153–1164. [Google Scholar] [CrossRef] [PubMed]
- Jin, W.; Barzilay, R.; Jaakkola, T. Junction tree variational autoencoder for molecular graph generation. In Proceedings of the International Conference on Machine Learning, PMLR, Stockholm, Sweden, 10–15 July 2018; pp. 2323–2332. [Google Scholar]
- Yamada, M.; Sugiyama, M. Molecular Graph Generation by Decomposition and Reassembling. ACS Omega 2023, 8, 19575–19586. [Google Scholar] [CrossRef] [PubMed]
- Yu, C.; Yongshun, G.; Yuansheng, L.; Bosheng, S.; Quan, Z. Molecular design in drug discovery: A comprehensive review of deep generative models. Briefings Bioinform. 2021, 22, bbab344. [Google Scholar]
- Weininger, D. SMILES, a chemical language and information system. 1. Introduction to methodology and encoding rules. J. Chem. Inf. Comput. Sci. 1988, 28, 31–36. [Google Scholar] [CrossRef]
- Pathak, Y.; Laghuvarapu, S.; Mehta, S.; Priyakumar, U.D. Chemically interpretable graph interaction network for prediction of pharmacokinetic properties of drug-like molecules. In Proceedings of the AAAI Conference on Artificial Intelligence, New York, NY, USA, 7–12 February 2020; Volume 34, pp. 873–880. [Google Scholar]
- Chen, H.; Engkvist, O.; Wang, Y.; Olivecrona, M.; Blaschke, T. The rise of deep learning in drug discovery. Drug Discov. Today 2018, 23, 1241–1250. [Google Scholar] [CrossRef] [PubMed]
- Jordan, M.I. Serial Order: A Parallel Distributed Processing Approach. Adv. Psychol. 1997, 121, 471–495. [Google Scholar]
- Gupta, A.; Müller, A.T.; Huisman, B.J.H.; Fuchs, J.A.; Schneider, P.; Schneider, G. Erratum: Generative Recurrent Networks for De Novo Drug Design. Mol. Inform. 2018, 37, 1880141. [Google Scholar] [CrossRef]
- Segler, M.H.; Kogej, T.; Tyrchan, C.; Waller, M.P. Generating focused molecule libraries for drug discovery with recurrent neural networks. ACS Cent. Sci. 2018, 4, 120–131. [Google Scholar] [CrossRef]
- Popova, M.; Isayev, O.; Tropsha, A. Deep reinforcement learning for de novo drug design. Sci. Adv. 2018, 4, eaap7885. [Google Scholar] [CrossRef]
- Olivecrona, M.; Blaschke, T.; Engkvist, O.; Chen, H. Molecular de novo design through deep reinforcement learning. J. Cheminform. 2017, 9, 1–14. [Google Scholar] [CrossRef]
- Simonovsky, M.; Komodakis, N. GraphVAE: Towards Generation of Small Graphs Using Variational Autoencoders. In Artificial Neural Networks and Machine Learning–ICANN 2018: 27th International Conference on Artificial Neural Networks, Rhodes, Greece, 4–7 October 2018; Springer: Cham, Switzerland, 2018. [Google Scholar]
- Jaechang, L.; Seongok, R.; Woo, K.J.; Youn, K.W. Molecular generative model based on conditional variational autoencoder for de novo molecular design. J. Cheminform. 2018, 10, 31. [Google Scholar]
- Makhzani, A.; Shlens, J.; Jaitly, N.; Goodfellow, I.; Frey, B. Adversarial autoencoders. arXiv 2015, arXiv:1511.05644. [Google Scholar]
- Hong, S.H.; Lim, J.; Ryu, S.; Kim, W.Y. Molecular Generative Model Based On Adversarially Regularized Autoencoder. J. Chem. Inf. Model. 2019, 60, 29–36. [Google Scholar] [CrossRef] [PubMed]
- Creswell, A.; White, T.; Dumoulin, V.; Arulkumaran, K.; Sengupta, B.; Bharath, A.A. Generative adversarial networks: An overview. IEEE Signal Process. Mag. 2018, 35, 53–65. [Google Scholar] [CrossRef]
- Guimaraes, G.L.; Sanchez-Lengeling, B.; Outeiral, C.; Farias, P.L.C.; Aspuru-Guzik, A. Objective-reinforced generative adversarial networks (organ) for sequence generation models. arXiv 2017, arXiv:1705.10843. [Google Scholar]
- Prykhodko, O.; Johansson, S.V.; Kotsias, P.C.; Arús-Pous, J.; Bjerrum, E.J.; Engkvist, O.; Chen, H. A de novo molecular generation method using latent vector based generative adversarial network. J. Cheminform. 2019, 11, 1–13. [Google Scholar] [CrossRef]
- Shen, C.; Krenn, M.; Eppel, S.; Aspuru-Guzik, A. Deep molecular dreaming: Inverse machine learning for de novo molecular design and interpretability with surjective representations. Mach. Learn. Sci. Technol. 2021, 2, 03LT02. [Google Scholar] [CrossRef]
- Nigam, A.; Pollice, R.; Krenn, M.; dos Passos Gomes, G.; Aspuru-Guzik, A. Beyond generative models: Superfast traversal, optimization, novelty, exploration and discovery (STONED) algorithm for molecules using SELFIES. Chem. Sci. 2021, 12, 7079–7090. [Google Scholar] [CrossRef]
- Grechishnikova, D. Transformer neural network for protein-specific de novo drug generation as a machine translation problem. Sci. Rep. 2021, 11, 321. [Google Scholar] [CrossRef]
- Zheng, S.; Lei, Z.; Ai, H.; Chen, H.; Deng, D.; Yang, Y. Deep scaffold hopping with multimodal transformer neural networks. J. Cheminform. 2021, 13, 1–15. [Google Scholar] [CrossRef]
- Bagal, V.; Aggarwal, R.; Vinod, P.; Priyakumar, U.D. MolGPT: Molecular generation using a transformer-decoder model. J. Chem. Inf. Model. 2021, 62, 2064–2076. [Google Scholar] [CrossRef]
- Lifan, C.; Xiaoqin, T.; Dingyan, W.; Feisheng, Z.; Xiaohong, L.; Tianbiao, Y.; Xiaomin, L.; Kaixian, C.; Hualiang, J.; Mingyue, Z. TransformerCPI: Improving compound–protein interaction prediction by sequence-based deep learning with self-attention mechanism and label reversal experiments. Bioinformatics 2020, 36, 4406–4414. [Google Scholar]
- Huang, K.; Xiao, C.; Glass, L.; Sun, J. MolTrans: Molecular Interaction Transformer for Drug Target Interaction Prediction. Bioinformatics 2021, 37, 830–836. [Google Scholar] [CrossRef] [PubMed]
- Bjerrum, E.J. SMILES enumeration as data augmentation for neural network modeling of molecules. arXiv 2017, arXiv:1703.07076. [Google Scholar]
- Arús-Pous, J.; Johansson, S.V.; Prykhodko, O.; Bjerrum, E.J.; Tyrchan, C.; Reymond, J.L.; Chen, H.; Engkvist, O. Randomized SMILES strings improve the quality of molecular generative models. J. Cheminform. 2019, 11, 1–13. [Google Scholar] [CrossRef]
- Wu, C.K.; Zhang, X.C.; Yang, Z.J.; Lu, A.P.; Hou, T.J.; Cao, D.S. Learning to SMILES: BAN-based strategies to improve latent representation learning from molecules. Briefings Bioinform. 2021, 22, bbab327. [Google Scholar] [CrossRef]
- Hjelm, R.D.; Fedorov, A.; Lavoie-Marchildon, S.; Grewal, K.; Bachman, P.; Trischler, A.; Bengio, Y. Learning deep representations by mutual information estimation and maximization. arXiv 2018, arXiv:1808.06670. [Google Scholar]
- Chen, T.; Kornblith, S.; Norouzi, M.; Hinton, G. A simple framework for contrastive learning of visual representations. In Proceedings of the International Conference on Machine Learning (PMLR), Virtual Event, 13–18 July 2020; pp. 1597–1607. [Google Scholar]
- Zhang, Y.; He, R.; Liu, Z.; Lim, K.H.; Bing, L. An unsupervised sentence embedding method by mutual information maximization. arXiv 2020, arXiv:2009.12061. [Google Scholar]
- Fang, H.; Wang, S.; Zhou, M.; Ding, J.; Xie, P. Cert: Contrastive self-supervised learning for language understanding. arXiv 2020, arXiv:2005.12766. [Google Scholar]
- Gao, T.; Yao, X.; Chen, D. Simcse: Simple contrastive learning of sentence embeddings. arXiv 2021, arXiv:2104.08821. [Google Scholar]
- Wang, Y.; Wang, J.; Cao, Z.; Barati Farimani, A. Molecular contrastive learning of representations via graph neural networks. Nat. Mach. Intell. 2022, 4, 279–287. [Google Scholar] [CrossRef]
- Sun, M.; Xing, J.; Wang, H.; Chen, B.; Zhou, J. MoCL: Data-driven molecular fingerprint via knowledge-aware contrastive learning from molecular graph. In Proceedings of the 27th ACM SIGKDD Conference on Knowledge Discovery & Data Mining, Virtual Event, 14–18 August 2021; pp. 3585–3594. [Google Scholar]
- Pinheiro, G.A.; Da Silva, J.L.; Quiles, M.G. Smiclr: Contrastive learning on multiple molecular representations for semisupervised and unsupervised representation learning. J. Chem. Inf. Model. 2022, 62, 3948–3960. [Google Scholar] [CrossRef] [PubMed]
- Landrum, G. Rdkit documentation. Release 2013, 1, 4. [Google Scholar]
- Singh, A.; Hu, R. UniT: Multimodal Multitask Learning with a Unified Transformer. In Proceedings of the IEEE/CVF International Conference on Computer Vision, Virtual Conference, 11–17 October 2021. [Google Scholar]
- Masashi, T.; Kentaro, T.; Jun, S. Compound-protein Interaction Prediction with End-to-end Learning of Neural Networks for Graphs and Sequences. Bioinformatics 2019, 35, 309–318. [Google Scholar]
- Dou, L.; Zhang, Z.; Qian, Y.; Zhang, Q. BCM-DTI: A fragment-oriented method for drug–target interaction prediction using deep learning. Comput. Biol. Chem. 2023, 104, 107844. [Google Scholar] [CrossRef]
- Davis, M.I.; Hunt, J.P.; Herrgard, S.; Ciceri, P.; Wodicka, L.M.; Pallares, G.; Hocker, M.; Treiber, D.K.; Zarrinkar, P.P. Comprehensive analysis of kinase inhibitor selectivity. Nat. Biotechnol. 2011, 29, 1046–1051. [Google Scholar] [CrossRef] [PubMed]
- Radford, A.; Wu, J.; Child, R.; Luan, D.; Amodei, D.; Sutskever, I. Language models are unsupervised multitask learners. OpenAI Blog 2019, 1, 9. [Google Scholar]
- Hendrycks, D.; Gimpel, K. Gaussian error linear units (gelus). arXiv 2016, arXiv:1606.08415. [Google Scholar]
- Sterling, T.; Irwin, J.J. ZINC 15–ligand discovery for everyone. J. Chem. Inf. Model. 2015, 55, 2324–2337. [Google Scholar] [CrossRef]
- Polykovskiy, D.; Zhebrak, A.; Sanchez-Lengeling, B.; Golovanov, S.; Tatanov, O.; Belyaev, S.; Kurbanov, R.; Artamonov, A.; Aladinskiy, V.; Veselov, M.; et al. Molecular sets (MOSES): A benchmarking platform for molecular generation models. Front. Pharmacol. 2020, 11, 565644. [Google Scholar] [CrossRef] [PubMed]
- Brown, N.; Fiscato, M.; Segler, M.H.; Vaucher, A.C. GuacaMol: Benchmarking models for de novo molecular design. J. Chem. Inf. Model. 2019, 59, 1096–1108. [Google Scholar] [CrossRef] [PubMed]
- Gaulton, A.; Bellis, L.J.; Bento, A.P.; Chambers, J.; Davies, M.; Hersey, A.; Light, Y.; McGlinchey, S.; Michalovich, D.; Al-Lazikani, B.; et al. ChEMBL: A large-scale bioactivity database for drug discovery. Nucleic Acids Res. 2012, 40, D1100–D1107. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Sun, J.; Guan, J.; Zheng, J.; Zhou, S. Improving compound–protein interaction prediction by building up highly credible negative samples. Bioinformatics 2015, 31, i221–i229. [Google Scholar] [CrossRef]
- Loshchilov, I.; Hutter, F. Fixing weight decay regularization in adam. arXiv 2017, arXiv:1711.05101. [Google Scholar]





| Models | Validity | Unique@10k | Novelty | Success Rate | |
|---|---|---|---|---|---|
| CharRNN | 0.975 | 0.999 | 0.842 | 0.820 | 0.856 |
| VAE | 0.977 | 0.998 | 0.695 | 0.678 | 0.856 |
| AAE | 0.937 | 0.997 | 0.793 | 0.741 | 0.856 |
| LatentGAN | 0.897 | 0.997 | 0.949 | 0.849 | 0.857 |
| JT-VAE | 1.0 | 0.999 | 0.914 | 0.913 | 0.855 |
| MolGPT | 0.995 | 1.0 | 0.781 | 0.777 | 0.850 |
| CON-GPT (unfrozen) | 0.992 | 1.0 | 0.791 | 0.785 | 0.850 |
| CON-GPT (frozen) | 0.991 | 1.0 | 0.834 | 0.826 | 0.850 |
| Models | Validity | Unique | Novelty | Success Rate |
|---|---|---|---|---|
| SMILES LSTM | 0.959 | 1.0 | 0.912 | 0.875 |
| VAE | 0.870 | 0.999 | 0.974 | 0.847 |
| AAE | 0.822 | 1.0 | 0.998 | 0.820 |
| MolGPT | 0.979 | 0.998 | 0.958 | 0.936 |
| CON-GPT (unfrozen) | 0.968 | 0.999 | 0.968 | 0.936 |
| CON-GPT (frozen) | 0.961 | 0.999 | 0.975 | 0.936 |
| Number | Molecules Generated | Molecules in the Test Set |
|---|---|---|
| 1 | COc1ccc(NC(=O)N2CCN(C(=O)C3 CCCCC3)CC2)cc1 | COc1ccc(NC(=O)N2CCN(C(=O)C3 CCCC3)CC2)cc1 |
| 2 | Cc1nc2cc3c(cc2n1CC(=O)NC1CCCCC1) OCCO3 | Cc1nc2cc3c(cc2n1CC(=O)NC1CCCC1) OCCO3 |
| 3 | Cn1ccc(C(=O)Nc2cc(F)ccc2N2CCCC2)cc1=O | Cn1ccc(C(=O)Nc2cc(F)ccc2N2CCCCC2)cc1=O |
| 4 | Cc1cc(CN(C)C(=O)Nc2ccccc2N2CCCC2)no1 | Cc1cc(CN(C)C(=O)Nc2ccccc2N2CCCCC2)no1 |
| 5 | COc1ccccc1-c1noc(C(=O)N2CCCC2)c1N | COc1ccccc1-c1noc(C(=O)N2CCCCC2)c1N |
| 6 | CC(CC#N)N(C)C(=O)Nc1ccccc1N1CCCC1 | CC(CC#N)N(C)C(=O)Nc1ccccc1N1CCCCC1 |
| Number | Tanimoto Similarity | QED (Generated) | SAScore (Generated) | LogP (Generated) |
|---|---|---|---|---|
| 1 | 0.958 | 0.916 | 1.839 | 2.952 |
| 2 | 0.957 | 0.940 | 2.318 | 2.565 |
| 3 | 0.957 | 0.941 | 2.184 | 2.377 |
| 4 | 0.956 | 0.950 | 2.102 | 3.247 |
| 5 | 0.952 | 0.935 | 2.284 | 2.168 |
| 6 | 0.952 | 0.925 | 2.685 | 3.053 |
| 0.05 | 0.10 | 0.50 | 1.00 | |
| Loss | 0.16411 | 0.16326 | 0.16379 | 0.17299 |
| Models | F1 | Precision | Recall |
|---|---|---|---|
| GNN-CPI | 0.933 | 0.938 | 0.929 |
| TransformerCPI | 0.952 | 0.952 | 0.953 |
| Moltrans | 0.954 | 0.947 | 0.962 |
| BCM-DTI | 0.969 | 0.967 | 0.971 |
| UniT | 0.964 | 0.966 | 0.961 |
| CON-UniT | 0.969 | 0.972 | 0.966 |
| Models | F1 | Precision | Recall |
|---|---|---|---|
| GNN-CPI | 0.658 | 0.647 | 0.669 |
| TransformerCPI | 0.584 | 0.46 | 0.8 |
| Moltrans | 0.306 | 0.185 | 0.884 |
| BCM-DTI | 0.611 | 0.853 | 0.476 |
| UniT | 0.841 | 0.844 | 0.837 |
| CON-UniT | 0.868 | 0.874 | 0.862 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Qian, Y.; Shi, M.; Zhang, Q. CONSMI: Contrastive Learning in the Simplified Molecular Input Line Entry System Helps Generate Better Molecules. Molecules 2024, 29, 495. https://doi.org/10.3390/molecules29020495
Qian Y, Shi M, Zhang Q. CONSMI: Contrastive Learning in the Simplified Molecular Input Line Entry System Helps Generate Better Molecules. Molecules. 2024; 29(2):495. https://doi.org/10.3390/molecules29020495
Chicago/Turabian StyleQian, Ying, Minghua Shi, and Qian Zhang. 2024. "CONSMI: Contrastive Learning in the Simplified Molecular Input Line Entry System Helps Generate Better Molecules" Molecules 29, no. 2: 495. https://doi.org/10.3390/molecules29020495
APA StyleQian, Y., Shi, M., & Zhang, Q. (2024). CONSMI: Contrastive Learning in the Simplified Molecular Input Line Entry System Helps Generate Better Molecules. Molecules, 29(2), 495. https://doi.org/10.3390/molecules29020495







