Synthesis and Molecular Docking Study of Novel Pyrimidine Derivatives against COVID-19
Abstract
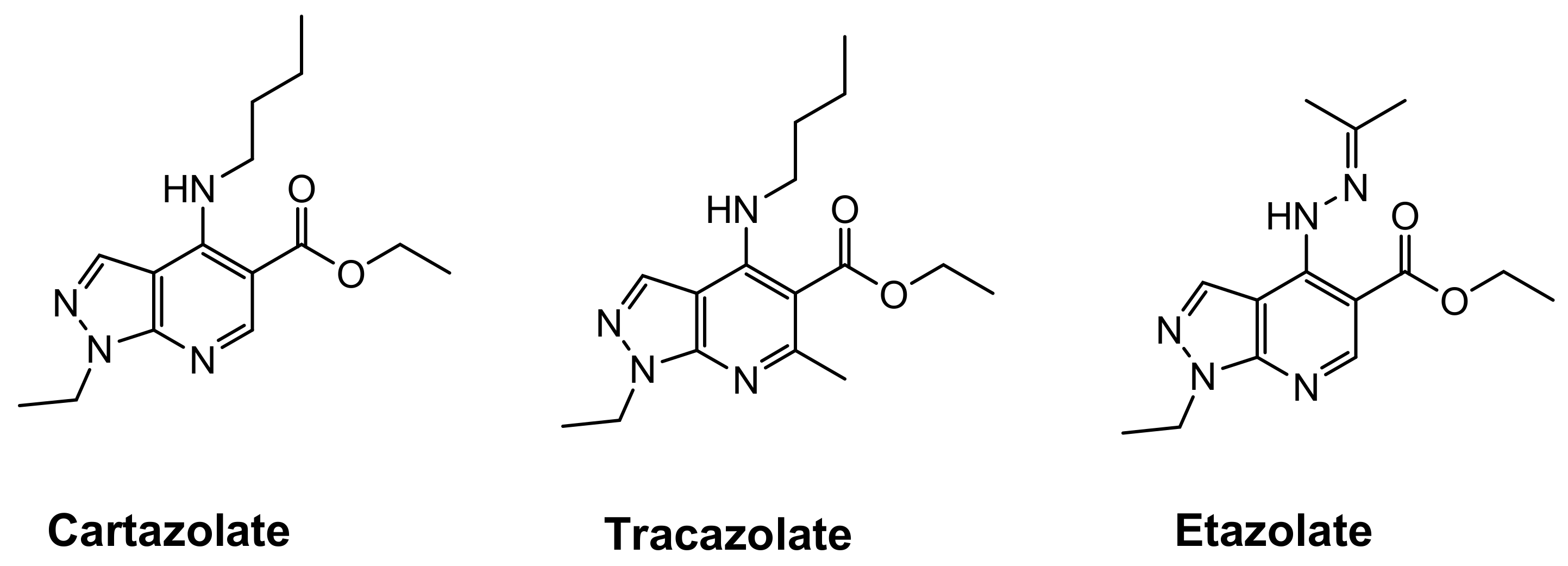
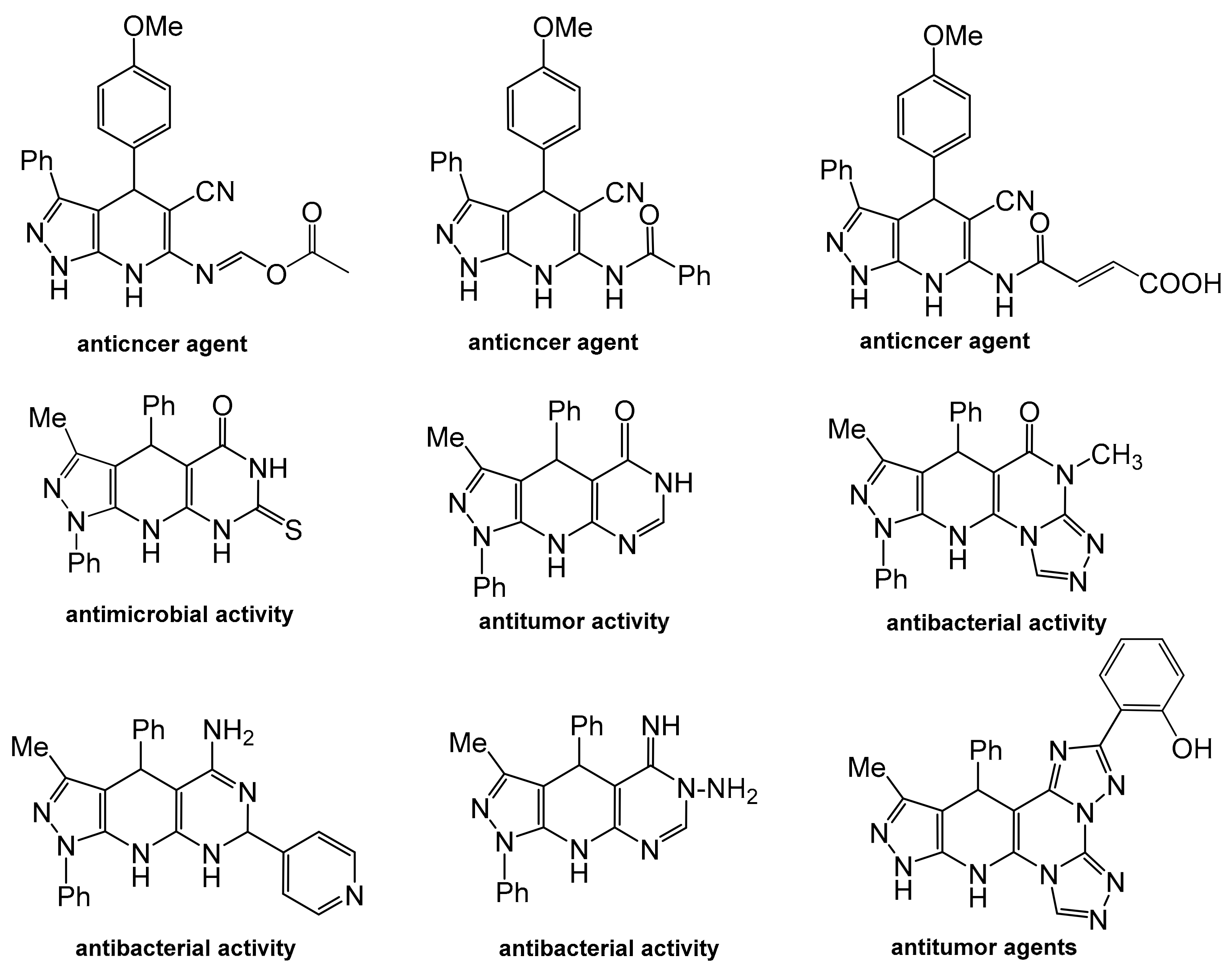
1. Introduction
2. Results and Discussions
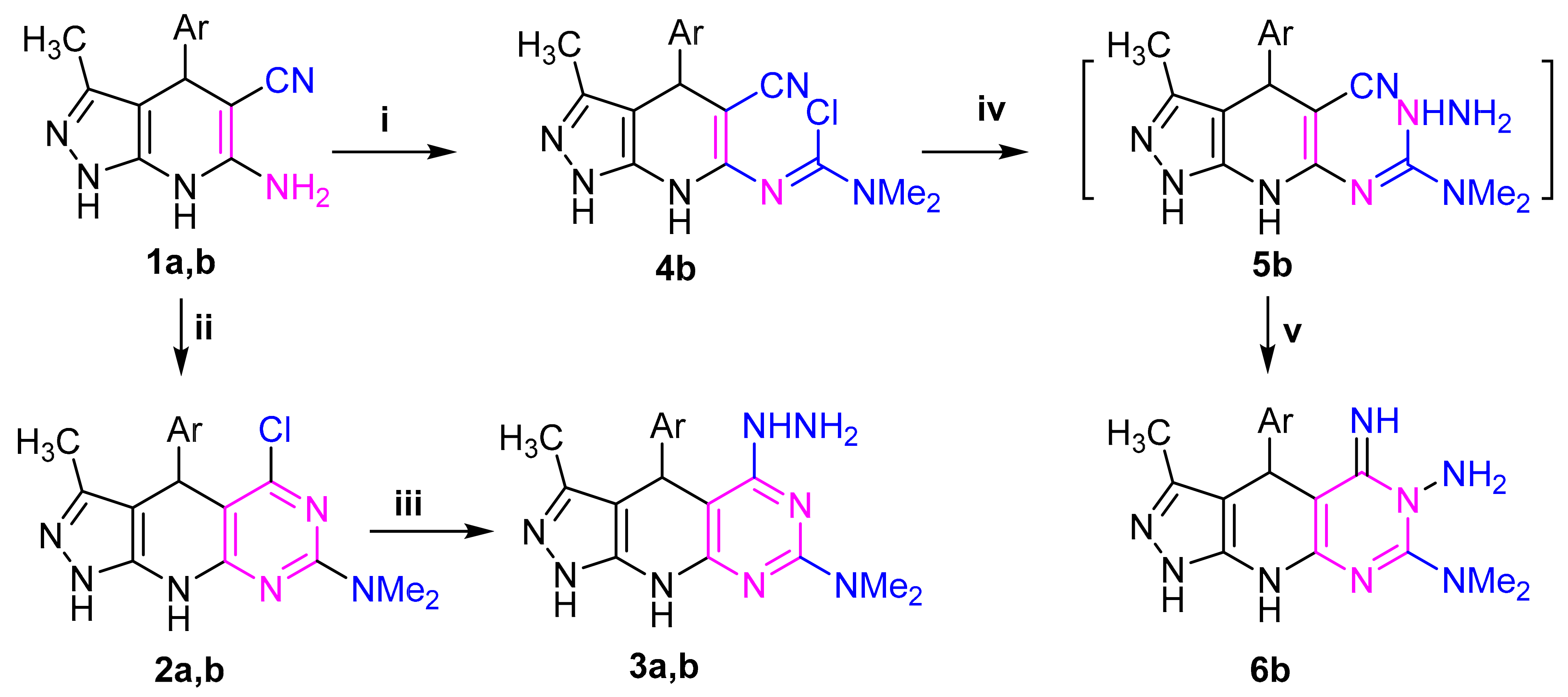
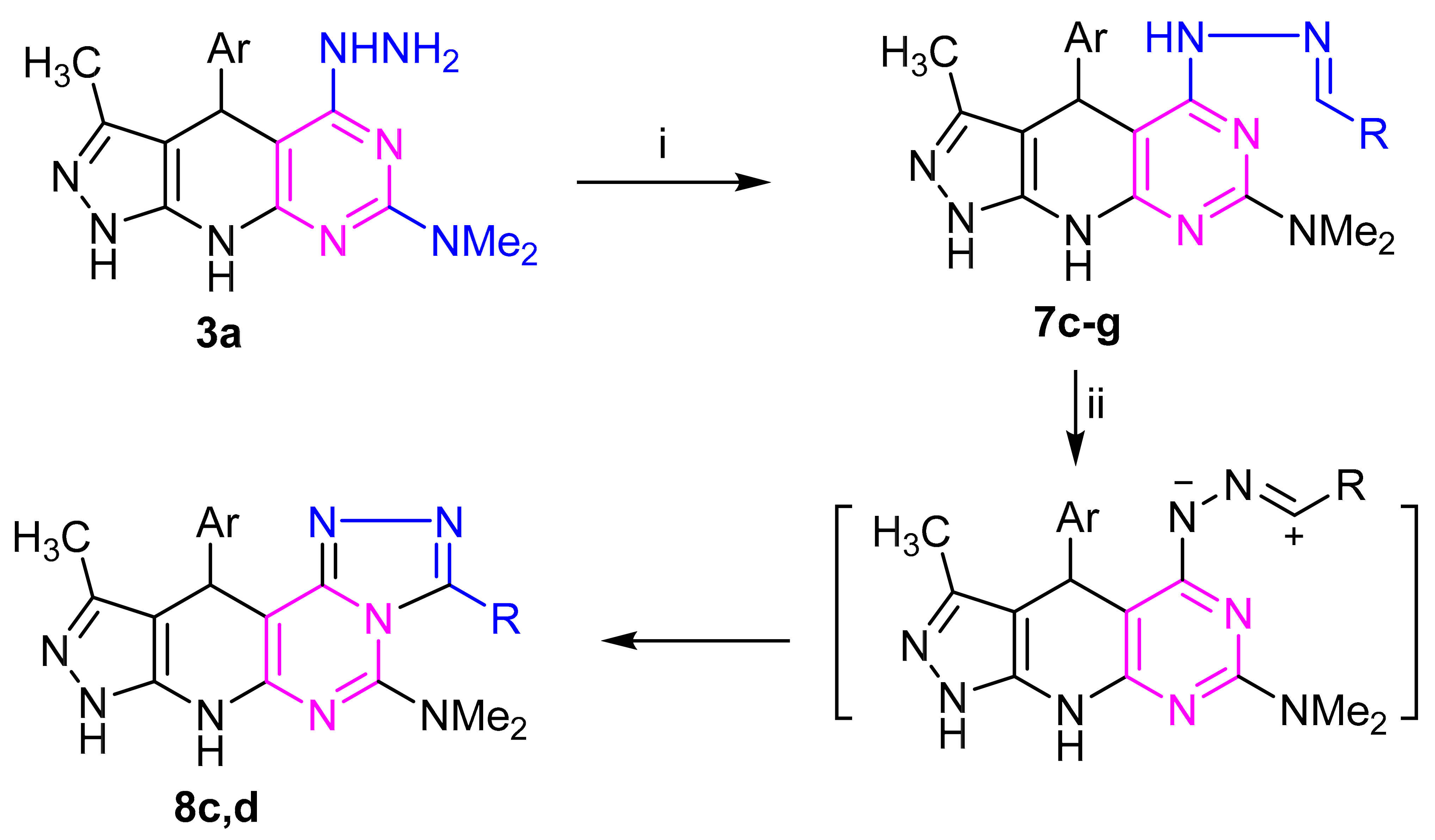
2.1. Chemistry
2.2. Antiviral Assay
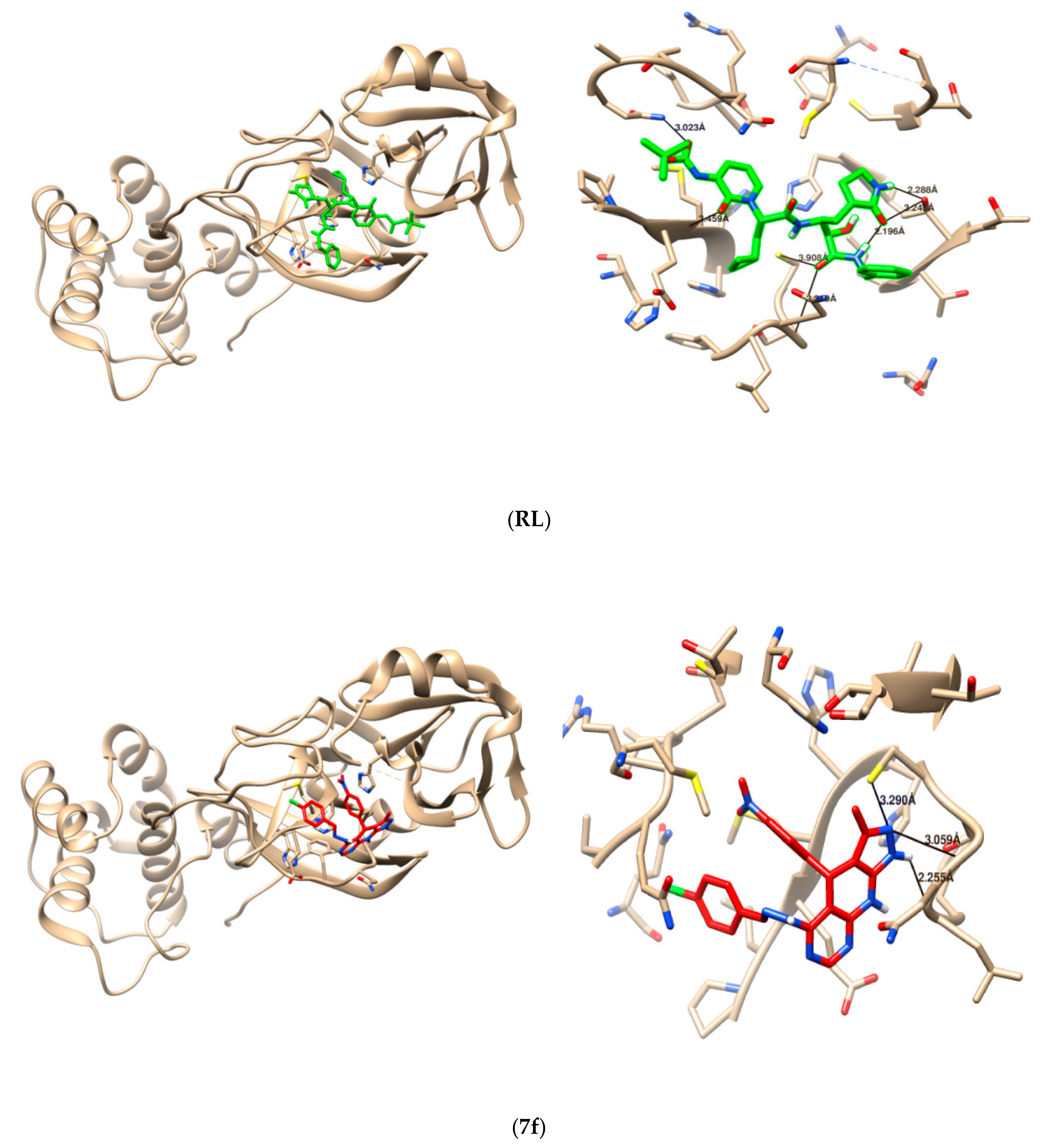
2.3. Molecular Docking
3. Structure Activity Relationships
4. Materials and Methods
4.1. General
4.2. Synthetic Procedures
4.2.1. General Procedure for Synthesis of Starting 6-Amino-3-Methyl-4-Aryl-Pyrazolo[3,4-b]Pyridine-5-Carbonitrile 1a,b
4.2.2. General Procedure for Synthesis of Compounds 2a, 2b, and 4b
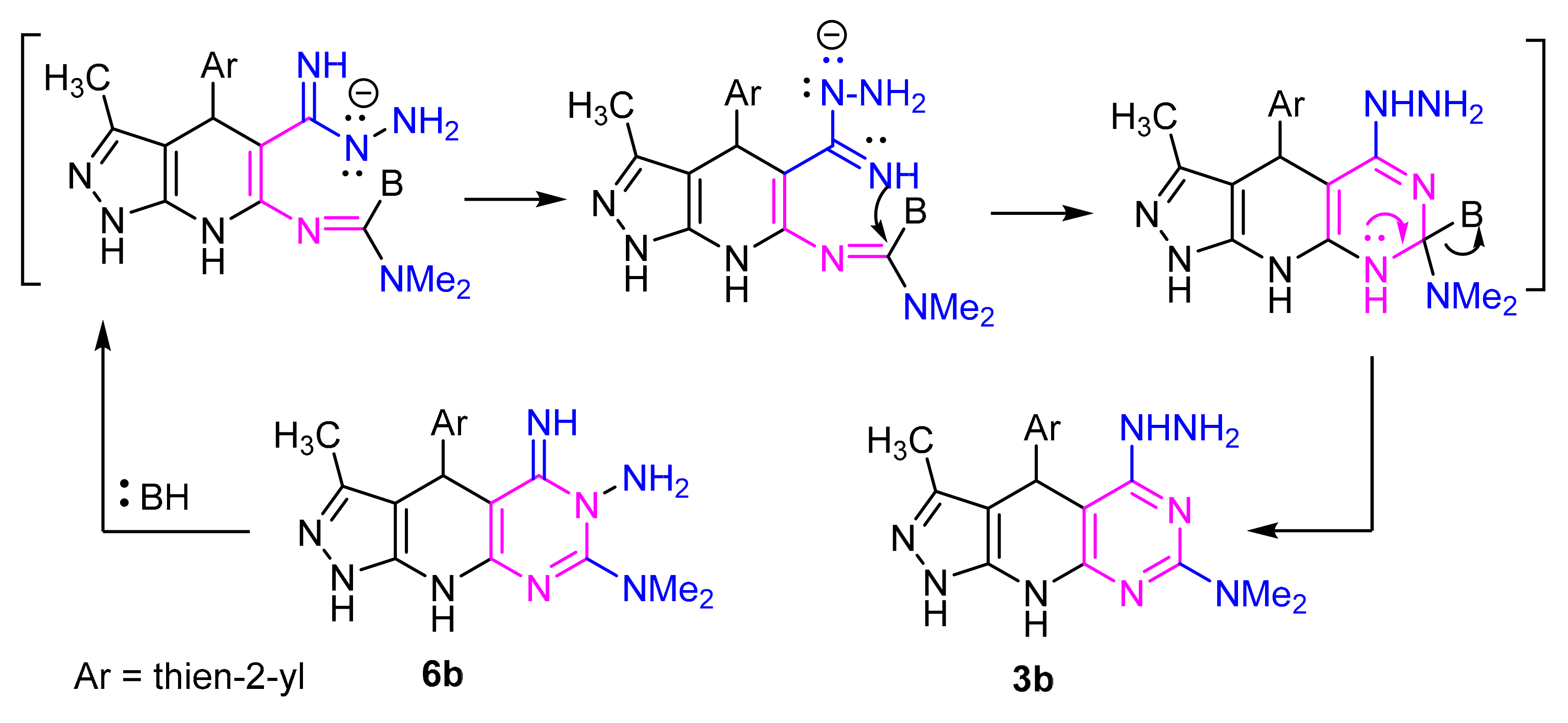
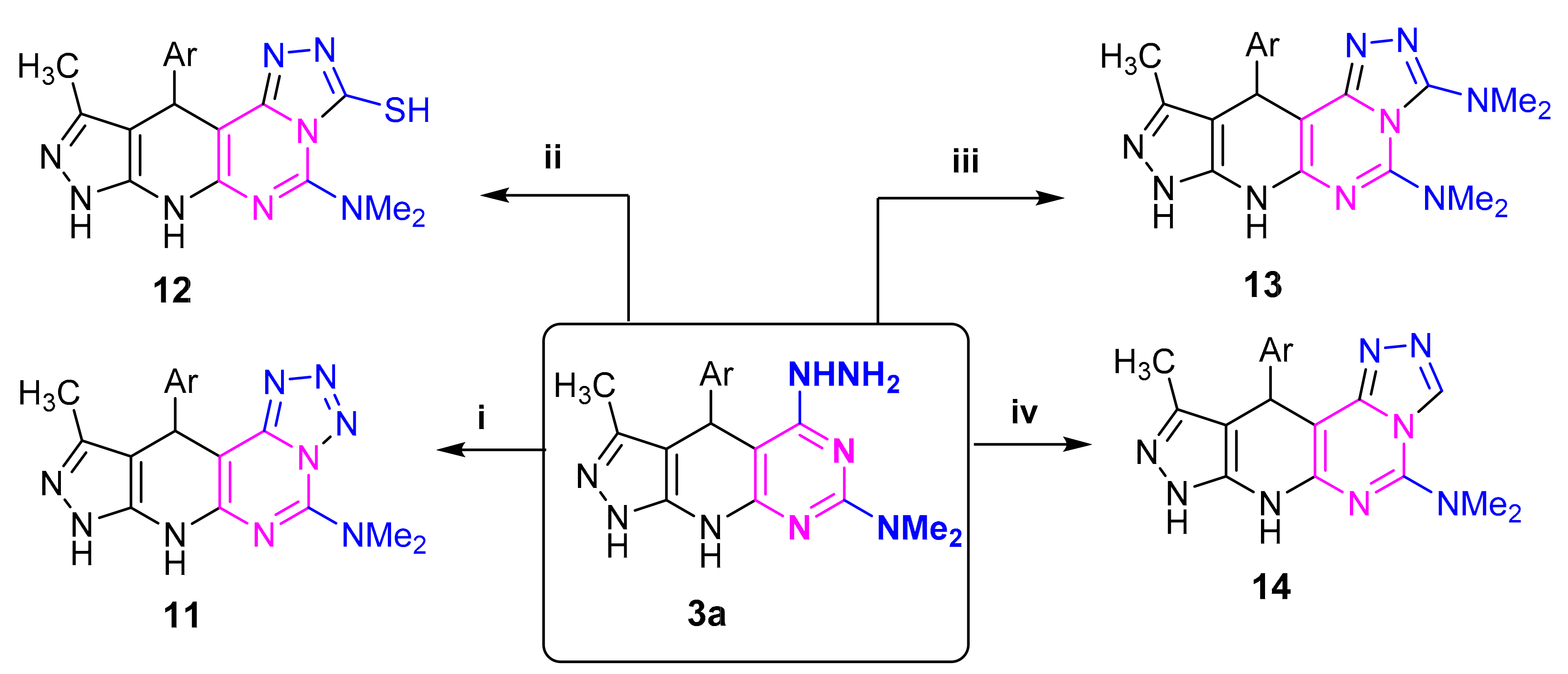
4.2.3. Synthesis of 3a, 3b, 6b
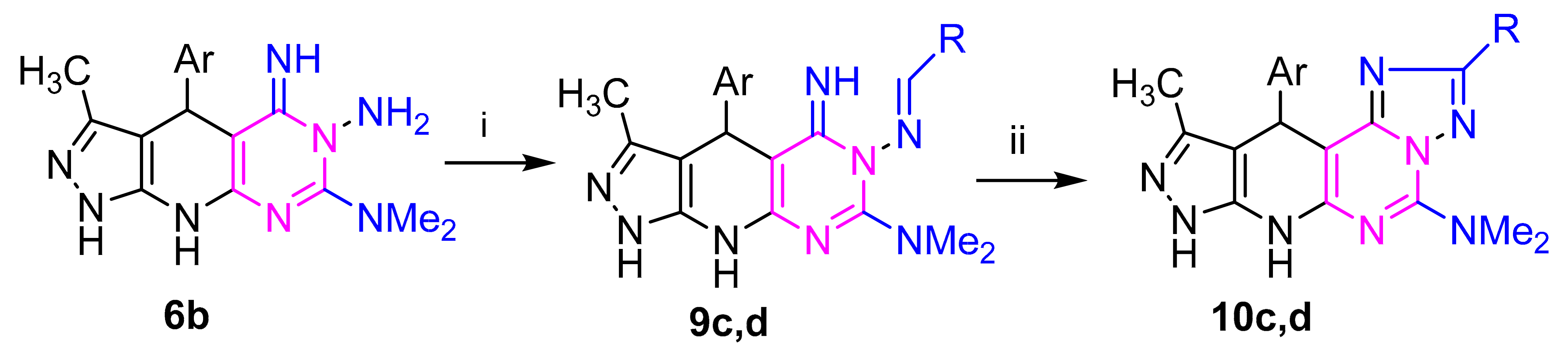
4.2.4. General Procedure for the Synthesis of 7c–g and 9c,d
4.2.5. General Procedure for the Synthesis of 8c,d and 10c,d
4.3. Cytotoxic Concentration 50 (CC50) and Viral Inhibitory Concentration 50 (IC50) Calculation
4.4. Molecular Docking Study
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Sample Availability
References
- Wang, C.; Wang, Z.; Wang, G.; Lau, J.Y.-N.; Zhang, K.; Li, W. COVID-19 in early 2021: Current status and looking forward. Signal Transduct Target Ther. 2021, 6, 114. [Google Scholar] [CrossRef] [PubMed]
- Wu, F.; Zhao, S.; Yu, B.; Chen, M.Y.; Wang, W.; Song, G.Z.; Hu, Y.; Tao, W.Z.; Tain, Y.Y.; Pei, Y.Y.; et al. A new coronavirus associated with human respiratory disease in China. Nature 2020, 579, 265–269. [Google Scholar] [CrossRef] [PubMed]
- Huang, C.; Wang, Y.; Li, X.; Ren, L.; Zhao, J.; Hu, Y.; Zhang, L.; Fan, G.; Xu, J.; Gu, X.; et al. Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet 2020, 395, 497–506. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Liu, J.; Cao, R.; Xu, M.; Wu, Y.; Shang, W.; Wang, X.; Zhang, H.; Jiang, X.; Sun, Y.; et al. Comparative Antiviral Efficacy of Viral Protease Inhibitors against the Novel SARS-CoV-2 In Vitro. Virol. Sin. 2020, 35, 776–784. [Google Scholar] [CrossRef]
- Anand, K. Coronavirus Main Proteinase (3CLpro) Structure: Basis for Design of Anti-SARS Drugs. Science 2003, 300, 1763–1767. [Google Scholar] [CrossRef]
- Zhang, M.; Wei, W.; Peng, C.; Ma, X.; He, X.; Zhang, H.; Zhou, M. Discovery of novel pyrazolopyrimidine derivatives as potent mTOR/HDAC bi-functional inhibitors via pharmacophore-merging strategy. Bioorg. Med. Chem. Lett. 2021, 49, 128286. [Google Scholar] [CrossRef]
- Tuyen, N.T.; Maged, H. Synthesis and Applications of Nitrogen-Containing Heterocycles as Antiviral Agents. Molecules 2022, 27, 2700. [Google Scholar] [CrossRef]
- Ghasemi, L.; Hasanzadeh, E.M.; Abbasi, A.; Behzad, M. Synthesis and crystal structures of new mixed-ligand Schiff base complexes containing N-donor heterocyclic co-ligands: Molecular docking and pharmacophore modeling studies on the main proteases of SARS-CoV-2 virus (COVID-19 disease). Polyhedron 2022, 220, 115825. [Google Scholar] [CrossRef]
- Artem’ev, G.A.; Rusinov, V.L.; Kopchuk, D.S.; Savchuk, M.I.; Santra, S.; Ulomsky, E.N.; Zyryanov, G.V.; Majee, A.; Du, W.; Charushin, V.N.; et al. Synthetic approaches to 1,2,4-triazolo [5,1-c][1,2,4]triazin-7-ones as basic heterocyclic structures of the antiviral drug Riamilovir (“Triazavirin”) active against SARS-CoV-2 (COVID-19). Org. Biomol. Chem. 2022, 20, 1828–1837. [Google Scholar] [CrossRef]
- Horchani, M.; Heise, N.V.; Csuk, R.; Ben Jannet, H.; Harrath, A.; Romdhane, A. Synthesis and In Silico Docking Study towards M-Pro of Novel Heterocyclic Compounds Derived from Pyrazolopyrimidinone as Putative SARS-CoV-2 Inhibitors. Molecules 2022, 27, 5303. [Google Scholar] [CrossRef]
- Dorababu, A. Pyrazolopyrimidines as attractive pharmacophores in efficient drug design: A recent update. Arch. Pharm. 2022, 355, 2200154. [Google Scholar] [CrossRef] [PubMed]
- El-Gohary, S.N.; Gabr, T.M.; Shaaban, I.M. Synthesis, molecular modeling and biological evaluation of new pyrazolo [3,4-b]pyridine analogs as potential antimicrobial, antiquorum-sensing and anticancer agents. Bioorg. Chem. 2019, 89, 102976. [Google Scholar] [CrossRef] [PubMed]
- Shejale, S.R.; Rajput, C.G.; Bansode, S.S.; Kondawar, M.S. Synthesis, characterisation, evaluation of antimicrobial & antifungal activity of novel pyrazolopyrimidine & pyrazolopyridine derivatives. Int. J. Pharm. Chem. 2017, 7, 144–148. [Google Scholar] [CrossRef]
- El-Gohary, S.N.; Shaaban, I.M. Design, synthesis, antimicrobial, antiquorum-sensing and antitumor evaluation of new series of pyrazolopyridine derivatives. Eur. J. Med. Chem. 2018, 157, 729–742. [Google Scholar] [CrossRef]
- El-Gohary, S.N.; Shaaban, I.M. New pyrazolopyridine analogs: Synthesis, antimicrobial, antiquorum-sensing and antitumor screening. Eur. J. Med. Chem. 2018, 152, 126–136. [Google Scholar] [CrossRef]
- Salem, S.M.; Ali, M.A.M. Novel pyrazolo[3,4-b]pyridine derivatives: Synthesis, characterization, antimicrobial and antiproliferative profile. Biol. Pharm. Bull. 2016, 39, 473–483. [Google Scholar] [CrossRef]
- Eissa, H.I.; El-Naggar, M.A.; El-Hashash, A.M. Design, synthesis, molecular modeling and biological evaluation of novel-1H-pyrazolo [3,4-b]pyridine derivatives as potential anticancer agents. Bioorg. Chem. 2016, 67, 43–56. [Google Scholar] [CrossRef]
- Arias, A.D.; Montagut, A.M.; Puig de la Bellacasa, R.; Estrada-Tejedor, R.; Teixidó, J.; Borrell, J.I. 1H-Pyrazolo [3,4-b]pyridines: Synthesis and Biomedical Applications. Molecules 2022, 27, 2237. [Google Scholar] [CrossRef]
- Karrouchi, K.; Radi, S.; Ramli, Y.; Taoufik, J.; Mabkhot, Y.N.; Al-aizari, F.A.; Ansar, M. Synthesis and Pharmacological Activities of Pyrazole Derivatives. Molecules 2018, 23, 134. [Google Scholar] [CrossRef]
- Hamama, S.W.; El-Gohary, G.H.; Soliman, M.; Zoorob, H.H. A versatile synthesis, PM3-semiempirical, antibacterial, and antitumor evaluation of some bioactive pyrazoles. J. Heterocycl. Chem. 2012, 49, 543–554. [Google Scholar] [CrossRef]
- Yadav, P.; Shah, K. An overview on synthetic and pharmaceutical prospective of pyrido [2,3-d]pyrimidines scaffold. Chem. Biol. Drug Des. 2020, 97, 633–648. [Google Scholar] [CrossRef] [PubMed]
- Yousif, M.N.M.; El-Gazzar, A.B.A.; El-Enany, M.M. Synthesis and Biological Evaluation of Pyrido(2,3-d)pyrimidines. Mini-Rev. Org. Chem. 2021, 18, 43–54. [Google Scholar] [CrossRef]
- Prachayasittikul, S.; Pingaew, R.; Worachartcheewan, A.; Sinthupoom, N.; Prachayasittikul, V.; Ruchirawat, S.; Prachayasittikul, V. Roles of Pyridine and Pyrimidine Derivatives as Privileged Scaffolds in Anticancer Agents. Mini-Rev. Med. Chem. 2017, 17, 869–901. [Google Scholar] [CrossRef] [PubMed]
- Sanghvi, Y.S.; Larson, S.B.; Willis, R.C.; Robins, R.K.; Revankar, G.R. Synthesis and biological evaluation of certain C-4 substituted pyrazolo[3,4-b]pyridine nucleosides. J. Med. Chem. 1989, 32, 945–951. [Google Scholar] [CrossRef] [PubMed]
- Monier, M.; Abdel-Latif, D.; El-Mekabaty, A.; Mert, B.D.; Elattar, K.M. Advances in the Chemistry of 6-6 Bicyclic Systems: Chemistry of Pyrido[3,4-d]pyrimidines. Curr. Org. Chem. 2019, 16, 812–854. [Google Scholar] [CrossRef]
- El-Remaily, A.A.M.; El Hady, M.O.; Abo Zaid, S.H.; Abd El-Raheem, M.M.E. Synthesis and in vitro antibacterial activity of some novel fused pyridopyrimidine derivatives. J. Heterocycl. Chem. 2016, 53, 1304–1309. [Google Scholar] [CrossRef]
- Abdelhameed, M.R.; Darwesh, M.O.; El-Shahat, M. Synthesis of arylidene hydrazinylpyrido[2,3-d]pyrimidin-4-ones as potent anti-microbial agents. Heliyon. 2020, 6, e04956. [Google Scholar] [CrossRef]
- Acosta, P.; Insuasty, B.; Ortiz, A.; Abonia, R.; Sortino, M.; Zacchino, A.S.; Quirog, J. Solvent-free microwave-assisted synthesis of novel pyrazolo[4′,3′:5,6]pyrido[2,3-d]pyrimidines with potential antifungal activity. Arab. J. Chem. 2016, 9, 481–492. [Google Scholar] [CrossRef]
- Bazgir, A.; Khanaposhtani, M.M.; Soorki, A.A. One-pot synthesis and antibacterial activities of pyrazolo[4′,3′:5,6]pyrido[2,3-d]pyrimidine-dione derivatives. Bioorg. Med. Chem. Lett. 2008, 18, 5800–5803. [Google Scholar] [CrossRef]
- El-Emary, I.T.; Abd-El-Mohsen, A.S. Multi-component one-pot synthesis and antimicrobial activities 3-methyl-1,4-diphenyl-7-thioxo-4,6,8,9-tetrahydropyrazolo [5,4-b]pyrimidino [5,4-e]pyridine-5-one and related derivatives. Molecules 2012, 17, 14464–14483. [Google Scholar] [CrossRef]
- Javahershenas, R.; Khalafy, J. One-pot, four component synthesis of pyrazolo[4’,3’:5,6]pyrido[2,3-d]pyrimidines derivatives. Asian J. Green Chem. 2018, 2, 318–329. [Google Scholar] [CrossRef]
- El-Gohary, S.N.; Hawasa, S.S.; Gabra, M.T.; Shaaband, M.I.; El-Ashmawy, M.B. New series of fused pyrazolopyridines: Synthesis, molecular modeling, antimicrobial, antiquorum-sensing and antitumor activities. Bioorg. Chem. 2019, 92, 103109. [Google Scholar] [CrossRef]
- Mohamed, R.N.; Khaireldin, Y.N.; Fahmy, F.A.; El-Sayed, A.A. Facile synthesis of fused nitrogen containing heterocycles as anticancer agents. Der Pharma Chem. 2010, 2, 400–417. [Google Scholar]
- El-Sayed, A.A.; Pedersen, B.E.; Khaireldin, Y.N. Thermal Stability of Modified i-Motif Oligonucleotides with Naphthalimide Intercalating Nucleic Acids. Helv. Chim. Acta. 2016, 99, 14–19. [Google Scholar] [CrossRef]
- El-Sayed, A.A.; Pedersen, B.E.; Khaireldin, A.N. Studying the influence of the pyrene intercalator TINA on the stability of DNA i-motifs. Nucleosides Nucleotides Nucleic Acids 2012, 31, 872–879. [Google Scholar] [CrossRef]
- El-Sayed, A.A.; El-Saidi, M.M.T.; Reham, R.K. Unexpected Reactions of Azido-p-Benzoquinone Derivatives towards Lawesson’s Reagent & Molecular Docking Study as a promising anticancer agent. Egypt. J. Chem. 2019, 62, 315–326. [Google Scholar] [CrossRef]
- Khattab, R.R.; Hassan, A.A.; Kutkatd, O.M.; Abuzeid, K.M.; Hassan, N.A. Synthesis and Antiviral Activity of Novel Thieno [2,3-d]pyrimidine Hydrazones and Their C-Nucleosides. Russ. J. Gen. Chem. 2019, 89, 1707–1717. [Google Scholar] [CrossRef]
- Wasfy, A.F.A.; Hassan, A.A.; Khattab, R.R.; Abu-Zied, M.K.; Awad, M.H.; Al Otaibi, F.; Hassan, A.N. Synthesis of Some New Thioglycosides Derived from Thieno[2,3-d]pyrimidine Derivatives and Their Anticancer and Antioxidant Activity. Res. J. Pharm. Biol. Chem. Sci. 2018, 9, 77–88. [Google Scholar]
- Hassan, N.A.; Abu-Zeid, K.M.; El-Gazzar, A.B. New routes to pyrano[2,3-d]pyrimidine derivatives from β-enamino nitrile and phosgene iminium chloride. Heterocycl. Comm. 2006, 12, 73–78. [Google Scholar] [CrossRef]
- El-Sayed, H.A.; Moustafa, A.H.; Hassan, A.A.; El-Seadawy, N.A.M.; Pasha, S.H.; Shmiess, N.A.M.; Awad, H.M.; Hassan, N.A. Microwave synthesis, anti-oxidant and anti-tumor activity of some nucleosides derived 2-oxonicotinonitrile. Synth. Commun. 2019, 49, 3465–3474. [Google Scholar] [CrossRef]
- Khattab, R.R.; Hassan, A.A.; Elganzor, H.H.; Tashkandi, N.Y.; Awad, H.M.; Nossier, E.S.; El-Sayed, W.A.; Hassan, N.A. Click Chemistry Based Synthesis, Cytotoxic Activity and Molecular Docking of Novel Triazole-Thienopyrimidine Hybride Glycosides Targeting EGFR. J. Enzyme Inhib. Med. Chem. 2020, 36, 405. [Google Scholar] [CrossRef]
- Basiony, E.A.; Hassan, A.A.; Al-Amshany, Z.M.; Abd-Rabou, A.A.; Abdel-Rahman, A.A.H.; Hassan, N.A.; El-Sayed, W.A. Synthesis and Cytotoxic Activity of New Thiazolopyrimidine Sugar Hydrazones and Their Derived Acyclic Nucleoside Analogues. Molecules 2020, 25, 399. [Google Scholar] [CrossRef] [PubMed]
- Rashad, A.E.; Shamroukh, A.H.; Abdel-Megeid, R.E.; Ali, H.S. Synthesis and Isomerization of Some Novel Pyrazolopyrimidine and Pyrazolotriazolopyrimidine Derivatives. Molecules 2014, 19, 5459–5469. [Google Scholar] [CrossRef] [PubMed]
- Shawali, A.S.; Hassaneen, H.M.; Shurrab, N.K. A new strategy for the synthesis of pyrazolo[4,3-e][1,2,4]triazolo[4,3-c]pyrimidines and pyrazolo[4,3-e][1,2,4]triazolo[1,5-c]pyrimidines. Tetrahedron 2008, 64, 10339–10343. [Google Scholar] [CrossRef]
- Shaban, M.A.; Nasr, A.Z. Synthesis of condensed 1,2,4-triazol[3,4-z] heterocycles. Adv. Heterocycl. Chem. 1999, 49, 277. [Google Scholar]
- Shawali, A.S.; Hassaneen, H.M.; Shurrab, N.K. Aconvenient synthesis of novel series of 4-cyclohexyl-2-substituted [1,2,4] triazolo [1,5-a] quinazolin-5(4H)-ones. Novel isoomers of H1 antihistaminic acitve agents. Heterocycles 2008, 75, 1479. [Google Scholar] [CrossRef]
- Kandeil, A.; Mostafa, A.; El-Shesheny, R.; Shehata, M.; Roshdy, W.H.; Ahmed, S.S.; Gomaa, M.; Taweel, A.E.; Kayed, A.E.; Mahmoud, S.H. Coding-Complete Genome Sequences of Two SARS-CoV-2 Isolates from Egypt. Microbiol. Resour. Announc. 2020, 9, e00489-20. [Google Scholar] [CrossRef]
- Andrés, P.; Blandine, P.; Julia, D.; Thomas, J.; Aurélien, T.; Victoria, D.; Pauline, B.; Bruno, L.; Manuel, R.; Olivier, T. In vitro evaluation of antiviral activity of single and combined repurposable drugs against SARS-CoV-2. Antiviral Res. 2020, 181, 104878. [Google Scholar] [CrossRef]
- Trott, O.; Olson, A.J. AutoDock Vina: Improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Comput. Chem. 2010, 31, 455–461. [Google Scholar] [CrossRef]
- Pettersen, E.F.; Goddard, T.D.; Huang, C.C.; Couch, G.S.; Greenblatt, D.M.; Meng, E.C.; Ferrin, T.E. UCSF Chimera-A visualization system for exploratory research and analysis. J. Comput. Chem. 2004, 25, 1605–1612. [Google Scholar] [CrossRef]
- Pedretti, A.; Villa, L.; Vistoli, G. VEGA—An open platform to develop chemo-bio-informatics applications, using plug-in architecture and script programming. J. Comput. Aided Mol. Des. 2004, 18, 167–173. [Google Scholar] [CrossRef] [PubMed]
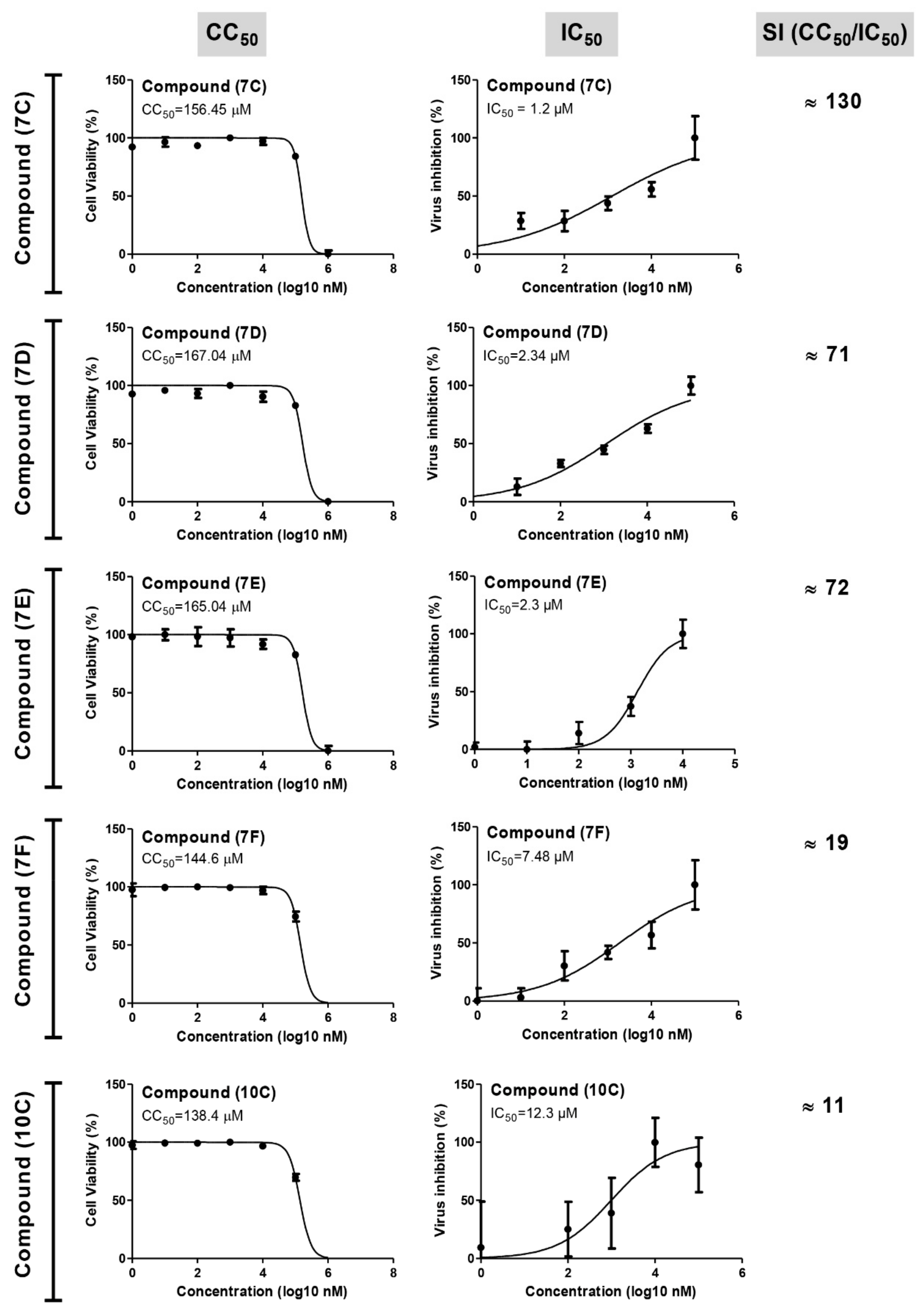
| Compound Code | Cytotoxicity (CC50, µM) | Antiviral Activity (IC50, µM) | Selectivity Index (SI) |
|---|---|---|---|
| 7c | 156.45 | 1.2 | 130 |
| 7d | 167.04 | 2.34 | 71 |
| 7e | 165.04 | 2.3 | 72 |
| 7f | 144.6 | 7.48 | 19 |
| 10c | 138.4 | 12.3 | 11 |
| Lopinavir | 44.95 | 5.246 | 8.57 |
| Compound | Free Energy of Binding (Kcal/mol) | Compound | Free energy of Binding (Kcal/mol) |
|---|---|---|---|
| PDB: 6Y2F | PDB: 6Y2F | ||
| Ref. ligand * | −7.5 | 8c | −7.6 |
| 2a | −7.1 | 8d | −7.5 |
| 3b | −6.5 | 9c | −7.8 |
| 5b | −6.8 | 9d | −7.7 |
| 6a | −6.7 | 10c | −8.1 |
| 7c | −8.4 | 10d | −8.1 |
| 7d | −8.3 | 11 | −7.7 |
| 7e | −8.5 | 12 | −6.9 |
| 7f | −8.0 | 13 | −7.8 |
| 7g | −7.5 | 14 | −7.0 |
| Compound | No. of H- Bonds | Length of H-Bonds | Formed Amino Acids with H-Bonds |
|---|---|---|---|
| Reference ligand | 5 | 3.139 Å 3.299 Å 2.890 Å 2.959 Å 3.023 Å | THR26 GLU166 CSY145 CYS145 SER144 |
| 7c | 2 | 3.290 Å 3.059 Å | CYC145 GLY143 |
| 7d | 2 | 3.548 Å 3.310 Å | CYC145 GLU166 |
| 7e | 1 | 2.475 Å | HIS146 |
| 7F | 5 | 3.249 Å 3.614 Å 3.080 Å 2.973 Å 2.252 Å | CYC145 CYC145 SER144 SER144 GLU166 |
| 10c | 2 | 3.270 Å 4.025 Å | GLU166 CYS145 |
| 10d | 2 | 4.135 Å 2.590 Å | GLN189 ARG188 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Alamshany, Z.M.; Khattab, R.R.; Hassan, N.A.; El-Sayed, A.A.; Tantawy, M.A.; Mostafa, A.; Hassan, A.A. Synthesis and Molecular Docking Study of Novel Pyrimidine Derivatives against COVID-19. Molecules 2023, 28, 739. https://doi.org/10.3390/molecules28020739
Alamshany ZM, Khattab RR, Hassan NA, El-Sayed AA, Tantawy MA, Mostafa A, Hassan AA. Synthesis and Molecular Docking Study of Novel Pyrimidine Derivatives against COVID-19. Molecules. 2023; 28(2):739. https://doi.org/10.3390/molecules28020739
Chicago/Turabian StyleAlamshany, Zahra M., Reham R. Khattab, Nasser A. Hassan, Ahmed A. El-Sayed, Mohamed A. Tantawy, Ahmed Mostafa, and Allam A. Hassan. 2023. "Synthesis and Molecular Docking Study of Novel Pyrimidine Derivatives against COVID-19" Molecules 28, no. 2: 739. https://doi.org/10.3390/molecules28020739
APA StyleAlamshany, Z. M., Khattab, R. R., Hassan, N. A., El-Sayed, A. A., Tantawy, M. A., Mostafa, A., & Hassan, A. A. (2023). Synthesis and Molecular Docking Study of Novel Pyrimidine Derivatives against COVID-19. Molecules, 28(2), 739. https://doi.org/10.3390/molecules28020739






