Structural Elucidation of an Atropisomeric Entcassiflavan-(4β→8)-Epicatechin Isolated from Dalbergia monetaria L.f. Based on NMR and ECD Calculations in Comparison to Experimental Data
Abstract
:1. Introduction
2. Results and Discussion
2.1. Compound Identification
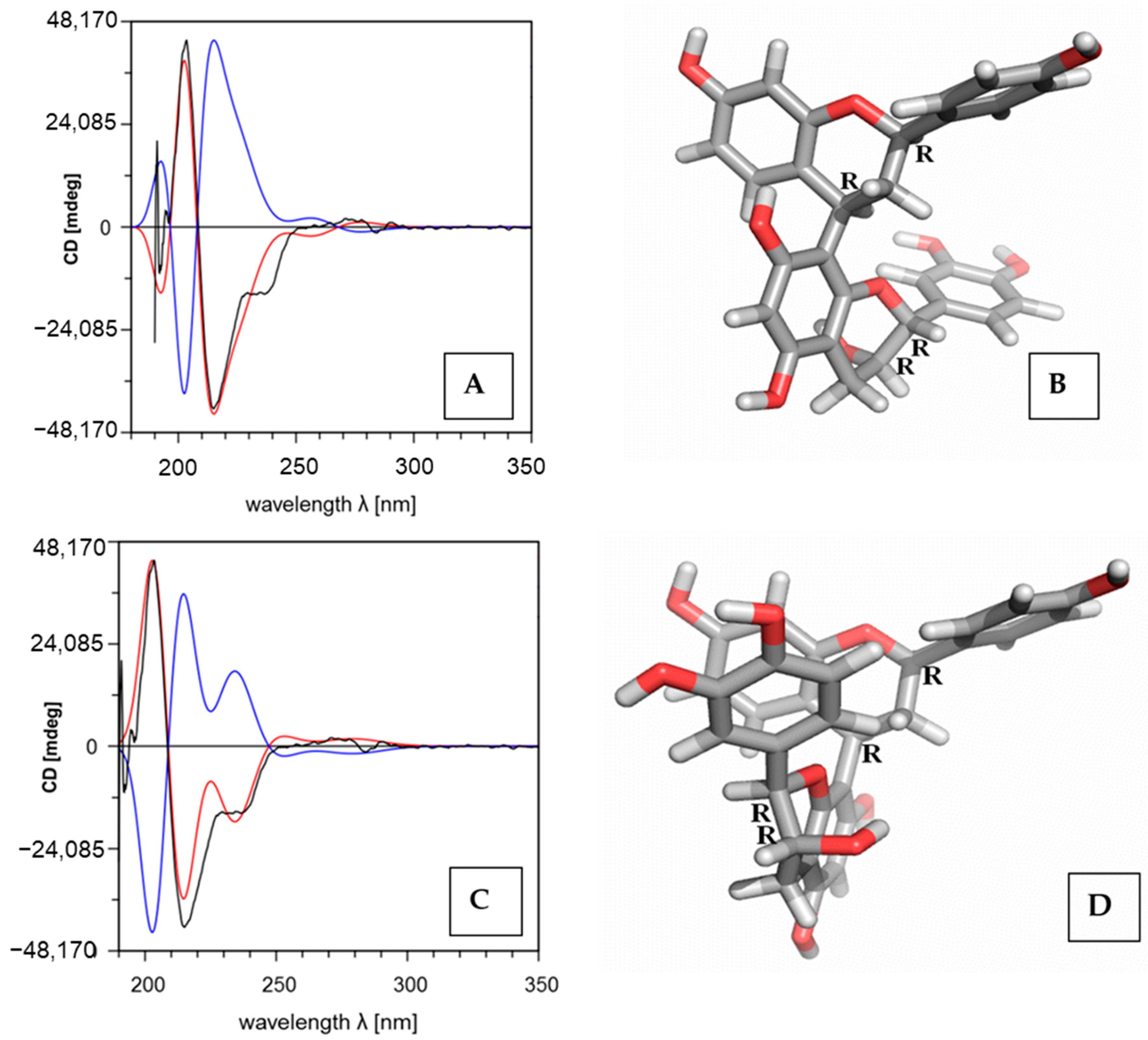
2.2. Determination of the Absolute Configuration and the Biaryl Position by Means of ECD Calculation
3. Materials and Methods
3.1. General Experimental Procedures
3.2. Extraction and Isolation of Compound
3.3. Computational Methods
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Sample Availability
References
- de Moura, P.H.B.; Lucas, F.C.A.; Lobato, G.D.J.M.; Tavares-Martins, A.C.C.; Gurgel, E.S.C. Etnobotânica de Chás Terapêuticos Em Rio Urubueua de Fátima, Abaetetuba—Pará, Brasil. Biotemas 2016, 29, 77. [Google Scholar] [CrossRef] [Green Version]
- Nunes, D.S.; Haag, A.; Bestmann, H.J. Two Proanthocyanidins from the Bark of Dalbergia Monetaria. Phytochemistry 1989, 28, 2183–2186. [Google Scholar] [CrossRef]
- Nunes, D.S.; Haag, A.; Hans, B.J. Inhaltsstoffe Der Rinde von Dalbergia Monetaria L. Drei Neue Isoflavon-C-glucoside. Liebigs Ann. Chem. 1989, 1989, 331–335. [Google Scholar] [CrossRef]
- Tang, C.; Xie, B.; Sun, Z. Antibacterial Activity and Mechanism of B-Type Oligomeric Procyanidins from Lotus Seedpod on Enterotoxigenic Escherichia Coli. J. Funct. Foods 2017, 38, 454–463. [Google Scholar] [CrossRef]
- Jekabsone, A.; Sile, I.; Cochis, A.; Makrecka-Kuka, M.; Lacautyte, G.; Makarova, E.; Rimondini, L.; Bernotiene, R.; Raudone, L.; Vedlugaite, E.; et al. Investigation of Antibacterial and Antiinflammatory Activities of Proanthocyanidins from Pelargonium Sidoides DC Root Extract. Nutrients 2019, 11, 2829. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Abou, P.J.; Momeni, J.; Adhikari, A.; Tsabang, N.; Tchinda, A.T.; Choudhary, M.I.; Nkengfack, A.E. New Coumestan and Coumaronochromone Derivatives from Dalbergia Boehmii Taub. (Fabaceae). Phytochem. Lett. 2017, 21, 109–113. [Google Scholar] [CrossRef]
- Coetzee, J.; Mciteka, L.; Elfranco, M.; Ferreira, D. Structure and Synthesis of the First Procassinidin Dimers Based on Epicatechin, and Gallo- and Epigallo-Catechin. Phytochemistry 2000, 53, 795–804. [Google Scholar] [CrossRef]
- Nakamura, S.; Xu, F.; Ninomiya, K.; Nakashima, S.; Oda, Y.; Morikawa, T.; Muraoka, O.; Yoshikawa, M.; Matsuda, H. Chemical Structures and Hepatoprotective Effects of Constituents from Cassia Auriculata Leaves. Chem. Pharm. Bull. 2014, 62, 1026–1031. [Google Scholar] [CrossRef] [Green Version]
- Sobeh, M.; Mahmoud, M.F.; Hasan, R.A.; Cheng, H.; El-shazly, A.M.; Wink, M. Senna Singueana: Antioxidant, Hepatoprotective, Antiapoptotic Properties and Phytochemical Profiling of a Methanol Bark Extract. Molecules 2017, 22, 1502. [Google Scholar] [CrossRef]
- Maia, I.R.D.O.; Teresa, M.; Trevisan, S.; Silva, M.G.D.V.; Breuer, A.; Owen, R.W. Characterization and Quantitation of Polyphenolic Compounds in Senna Macranthera Var Pudibunda From the Northeast of Brazil. Nat. Prod. Commun. 2019, 14, 1934578X19851704. [Google Scholar] [CrossRef] [Green Version]
- Farag, M.A.; El Senousy, A.S.; El-Ahmady, S.H.; Porzel, A.; Wessjohann, L.A. Comparative Metabolome-Based Classification of Senna Drugs: A Prospect for Phyto-equivalency of Its Different Commercial Products. Metabolomics 2019, 15, 80. [Google Scholar] [CrossRef] [PubMed]
- de Moura, P.H.B.; de Sousa, A.A.; Porzel, A.; Wessjohan, L.A.; Leal, I.C.R.; Martins, R.C.C. Characterization of Antibacterial Proanthocyanidins of Dalbergia Monetaria, an Amazonian Medicinal Plant, by UHPLC-HRMS/MS. Planta Med. 2020, 86, 858–866. [Google Scholar] [CrossRef]
- Shoji, T.; Mutsugam, M.; Nakamura, T.; Nakamura, T.; Kanda, T.; Akiyama, H.; Goda, Y. Isolation and Structural Elucidation of Some Procyanidins from Apple by Low-Temperature Nuclear Magnetic Resonance. J. Agric. Food Chem. 2003, 51, 3806–3813. [Google Scholar] [CrossRef] [PubMed]
- Nam, J.; Phansalkar, R.S.; Lankin, D.C.; Bisson, J.; Mcalpine, J.B.; Leme, A.A.; Vidal, C.M.P.; Ramirez, B.; Niemitz, M.; Bedran-russo, A.; et al. Subtle Chemical Shifts Explain the NMR Fingerprints of Oligomeric Proanthocyanidins with High Dentin Biomodification Potency. J. Org. Chem. 2015, 80, 7495–7507. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nam, J.; Phansalkar, R.S.; Lankin, D.C.; Mcalpine, J.B.; Leme-kraus, A.A.; Vidal, C.M.P.; Gan, L.; Bedran-russo, A.; Chen, S.; Pauli, G.F. Absolute Configuration of Native Oligomeric Proanthocyanidins with Dentin Biomodification Potency. J. Org. Chem. 2017, 82, 1316–1329. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Phansalkar, R.S.; Nam, J.; Leme-kraus, A.A.; Gan, L.; Zhou, B.; Mcalpine, J.B.; Chen, S.; Bedran-russo, A.K.; Pauli, G.F. Proanthocyanidin Dimers and Trimers from Vitis Vinifera Provide Diverse Structural Motifs for the Evaluation of Dentin Biomodification. J. Nat. Prod. 2019, 82, 2387–2399. [Google Scholar] [CrossRef] [PubMed]
- Stephens, P.J.; Pan, J.J.; Devlin, F.J.; Urbanová, M.; Hájíček, J. Determination of the Absolute Configurations of Natural Products via Density Functional Theory Calculations of Vibrational Circular Dichroism, Electronic Circular Dichroism and Optical Rotation: The Schizozygane Alkaloid Schizozygine. J. Org. Chem. 2007, 72, 2508–2524. [Google Scholar] [CrossRef] [PubMed]
- Molecular Operating Environment (MOE), 2014.09; Chemical Computing Group Inc.: Montreal, QC, Canada, 2015.
- Halgren, T.A. MMFF VI. MMFF94s Option for Energy Minimization Studies. J. Comp. Chem. 1999, 20, 720–729. [Google Scholar] [CrossRef]
- Becke, A.D. Density-Functional Exchange-Energy Approximation with Correct Asymptotic Behavior. Phys. Rev. A 1988, 38, 3098–3100. [Google Scholar] [CrossRef]
- Perdew, J.P. Density-Functional Approximation for the Correlation Energy of the Inhomogeneous Electron Gas. Phys. Rev. B 1986, 33, 8822–8824. [Google Scholar] [CrossRef]
- Karton, A.; Tarnopolsky, A.; Lamère, J.F.; Schatz, G.C.; Martin, J.M.L. Highly Accurate First-Principles Benchmark Data Sets for the Parametrization and Validation of Density Functional and Other Approximate Methods. Derivation of a Robust, Generally Applicable, Double-Hybrid Functional for Thermochemistry and Thermochemical. J. Phys. Chem. A 2008, 112, 12868–12886. [Google Scholar] [CrossRef] [PubMed]
- Schäfer, A.; Horn, H.; Ahlrichs, R. Fully Optimized Contracted Gaussian-Basis Sets for Atoms Li to Kr. J. Chem. Phys. 1992, 97, 2571–2577. [Google Scholar] [CrossRef]
- Weigend, F.; Ahlrichs, R. Balanced Basis Sets of Split Valence, Triple Zeta Valence and Quadruple Zeta Valence Quality for H to Rn: Design and Assessment of Accuracy. Phys. Chem. Chem. Phys. 2005, 7, 3297–3305. [Google Scholar] [CrossRef] [PubMed]
- Neese, F.; Becker, U.; Ganyushin, D.; Hansen, A.; Liakos, D.; Kollmar, C.; Koßmann, S.; Petrenko, T.; Reimann, C.; Riplinger, C.; et al. ORCA—An Ab Initio, Density Functional and Semiempirical Program Package; Max Planck Institute for Bioinorganic Chemistry: Mühlheim, Germany, 2012. [Google Scholar]
- Sinnecker, S.; Rajendran, A.; Klamt, A.; Diedenhofen, M.; Neese, F. Calculation of Solvent Shifts on Electronic G-Tensors with the Conductor-like Screening Model (COSMO) and Its Self-Consistent Generalization to Real Solvents (Direct COSMO-RS). J. Phys. Chem. A 2006, 110, 2235–2245. [Google Scholar] [CrossRef] [PubMed]
- Bruhn, T.; Schaumlöffel, A.; Hemberger, Y.; Bringmann, G. SpecDis: Quantifying the Comparison of Calculated and Experimental Electronic Circular Dichroism Spectra. Chirality 2013, 25, 243–249. [Google Scholar] [CrossRef] [PubMed]
- Bruhn, T.; Schaumlöffel, A.; Hemberger, Y.; Bringmann, G. SpecDis Version 1.62; University of Würzburg: Würzburg, Germany, 2013. [Google Scholar]



| Pos. a | δ 13C (ppm) | δ 1H (ppm) m (J [Hz]) | Key HMBC (H→C) | Key NOE |
|---|---|---|---|---|
| C2 | 80.43 | 4.974 dd (11.6/2.0) ax | C4, B1′, B2′/6′ | C3 eq, C4, B2′/6′ |
| C3 | 36.71 | 2.640 ddd (13.2/12.0/11.6) ax; 1.836 ddd (13.2/5.8/2.0) eq | C2, C4, B1′, A10, D8; B4, A10 | B2′/6′; C2, C4 |
| C4 | 33.05 | 4.771 ddd (12.0/5.8/1.1) ax | B2, B3, A5, A9, A10, D7, D8, D9 | C2, C3 eq, A5 |
| A5 | 129.84 | 6.709 dd (8.4/1.2) | A7, A8, A9 | A6 |
| A6 | 108.99 | 6.262 dd (8.4/2.5) | A7, A8, A9, A10 | A5 |
| A7 | 156.75 | - | ||
| A8 | 103.97 | 6.311 d (2.5) | A6, A7, A9, A10 | |
| A9 | 157.26 | - | ||
| A10 | 120.12 | - | ||
| B1′ | 134.77 | - | ||
| B2′/6′ | 128.62 | 7.056 d-like (8.6) | B6′/2′, B4′ | B3′/5′ |
| B3′/5′ | 115.95 | 6.660 d-like (8.6) | B1′, B5′/3′, B4′ | B2′/6′ |
| B4′ | 157.99 | - | ||
| F2 | 79.30 | 4.731 br s-like ax | F3, F4,E1′, E2′, E6′ | F3, F4ax, E2′, E6′ |
| F3 | 67.44 | 4.067 ddd (4.6/2.3/1.2) eq | F2, D10 | F2, F4ax, F4eq, E2′, E6′ (w) b |
| F4 | 29.35 | 2.846 dd (16.8/4.6) ax; 2.757 ddd (16.8/2.3/0.9) eq | F2, F3, D5, D9, D10; F2, F3, D5, D9, D10 | F2 ax, F3; F3 |
| D5 | 156.03 | - | ||
| D6 | 96.09 | 6.081 s | D5, D7, D8, D10 | |
| D7 | 155.74 | - | ||
| D8 | 110.05 | - | ||
| D9 | 155.50 | - | ||
| D10 | 100.78 | - | ||
| E1′ | 131.89 | - | ||
| E2′ | 114.14 | 6.540 dd (2.1/0.6) | E3′, E4′, E6′ | |
| E3′ | 145.72 | - | ||
| E4′ | 145.43 | - | ||
| E5′ | 115.95 | 6.638 d (8.2) | E1′, E3′, E4′ | E6′ |
| E6′ | 119.76 | 6.132 ddd (8.2/2.1/0.6) | E4′ | E5′ |
| Biaryl Bond | Energy 1 (kcal/mol) | Config. 2- Atropoisomer | S 3 | Shift | Figure | Config. 2- Atropisomer | S 3 | Shift | Figure |
|---|---|---|---|---|---|---|---|---|---|
| 4–8 | 0.0 | RRRR-P | 0.9674 | +19 | Figure 1A | SSSS-M | 0.6071 | −30 | Figure S8 |
| 4–8 | 1.7 | RRRR-M | 0.9642 | +30 | Figure 1C | SSSS-P | 0.8631 | −3 | Figure S15 |
| 4–8 | 0.0 | SSRR-P | 0.7511 | −26 | Figure S9 | RRSS-M | 0.6300 | +16 | Figure S10 |
| 4–8 | 2.4 | SSRR-M | 0.7720 | +20 | Figure S16 | RRSS-P | 0.7557 | −21 | Figure S17 |
| 4–6 | 0.0 | RRRR-M | 0.6709 | −5 | Figure S11 | SSSS-P | 0.7564 | −26 | Figure S18 |
| 4–6 | 2.3 | RRRR-P | 0.9278 | +27 | Figure 2 | SSSS-M | 0.8379 | −1 | Figure S12 |
| 4–6 | 0.0 | SSRR-P | 0.7730 | −30 | Figure S13 | RRSS-M | 0.8427 | +29 | Figure S20 |
| 4–6 | 1.3 | SSRR-M | 0.8580 | −2 | Figure S19 | RRSS-P | 0.7863 | −27 | Figure S14 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
de Moura, P.H.B.; Brandt, W.; Porzel, A.; Martins, R.C.C.; Leal, I.C.R.; Wessjohann, L.A. Structural Elucidation of an Atropisomeric Entcassiflavan-(4β→8)-Epicatechin Isolated from Dalbergia monetaria L.f. Based on NMR and ECD Calculations in Comparison to Experimental Data. Molecules 2022, 27, 2512. https://doi.org/10.3390/molecules27082512
de Moura PHB, Brandt W, Porzel A, Martins RCC, Leal ICR, Wessjohann LA. Structural Elucidation of an Atropisomeric Entcassiflavan-(4β→8)-Epicatechin Isolated from Dalbergia monetaria L.f. Based on NMR and ECD Calculations in Comparison to Experimental Data. Molecules. 2022; 27(8):2512. https://doi.org/10.3390/molecules27082512
Chicago/Turabian Stylede Moura, Patrícia Homobono Brito, Wolfgang Brandt, Andrea Porzel, Roberto Carlos Campos Martins, Ivana Correa Ramos Leal, and Ludger A. Wessjohann. 2022. "Structural Elucidation of an Atropisomeric Entcassiflavan-(4β→8)-Epicatechin Isolated from Dalbergia monetaria L.f. Based on NMR and ECD Calculations in Comparison to Experimental Data" Molecules 27, no. 8: 2512. https://doi.org/10.3390/molecules27082512
APA Stylede Moura, P. H. B., Brandt, W., Porzel, A., Martins, R. C. C., Leal, I. C. R., & Wessjohann, L. A. (2022). Structural Elucidation of an Atropisomeric Entcassiflavan-(4β→8)-Epicatechin Isolated from Dalbergia monetaria L.f. Based on NMR and ECD Calculations in Comparison to Experimental Data. Molecules, 27(8), 2512. https://doi.org/10.3390/molecules27082512






