Optimization of Bifunctional Antisense Oligonucleotides for Regulation of Mutually Exclusive Alternative Splicing of PKM Gene
Abstract
1. Introduction
2. Results and Discussion
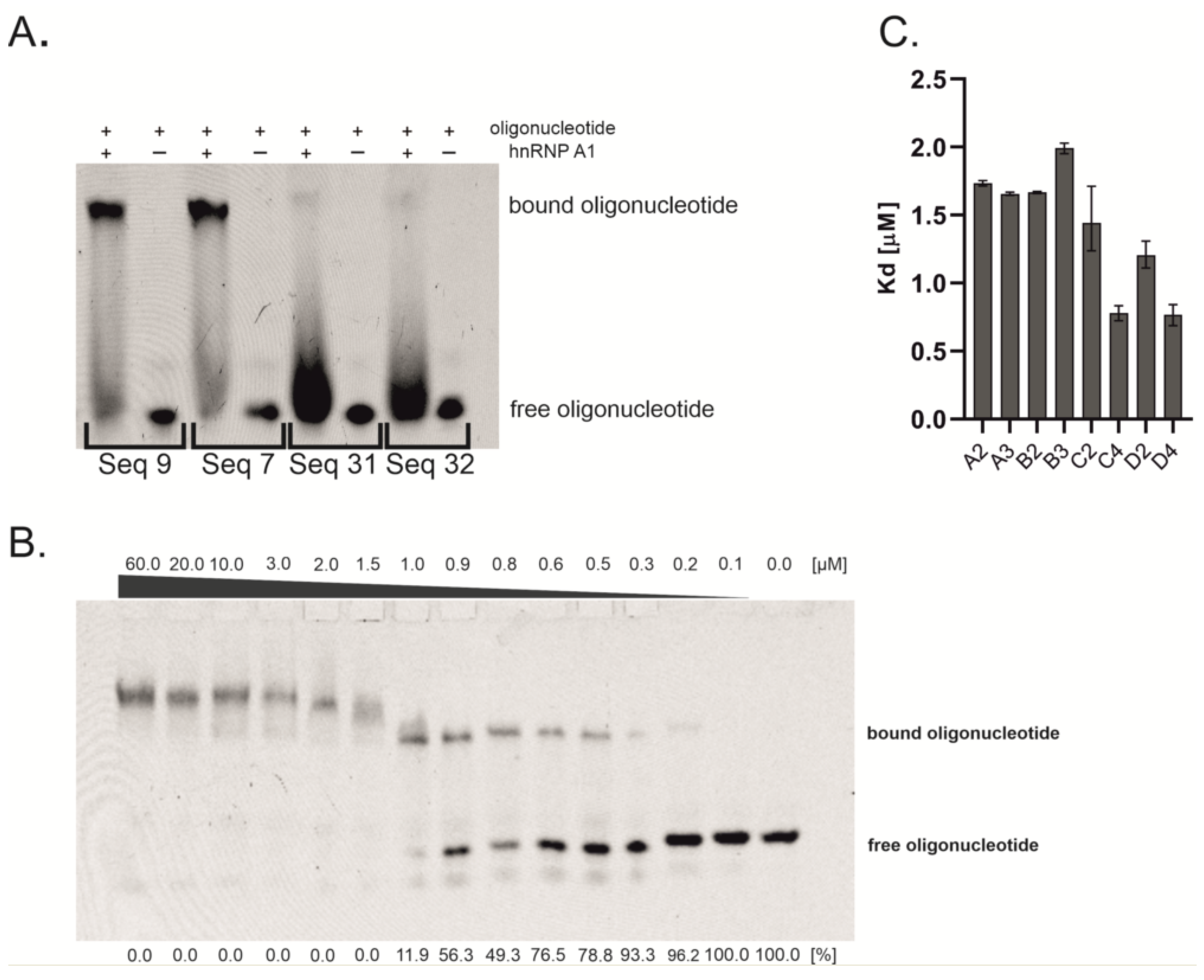
2.1. Oligonucleotides Screening and Binding Affinity Determination
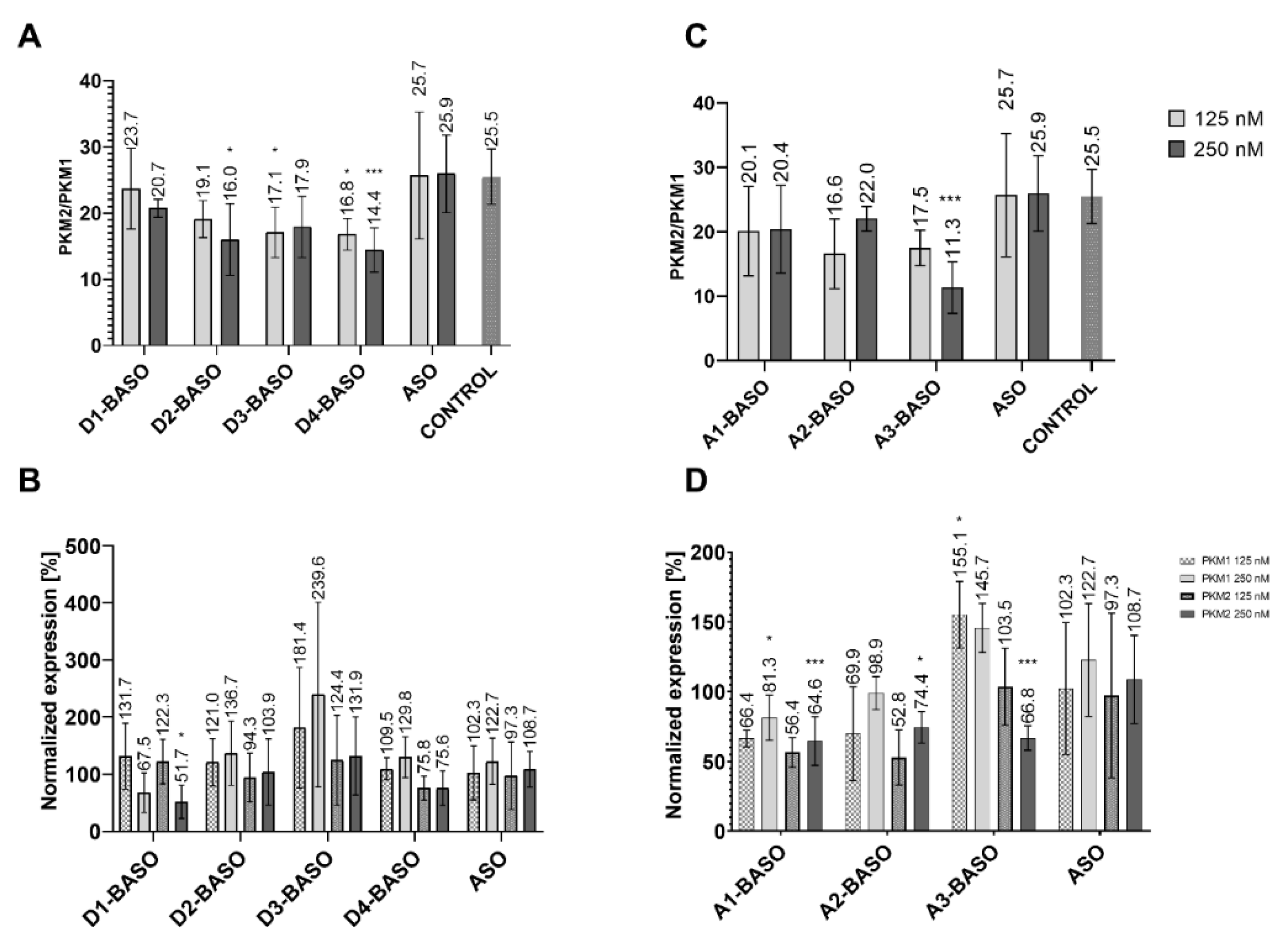
2.2. Cell Line Results
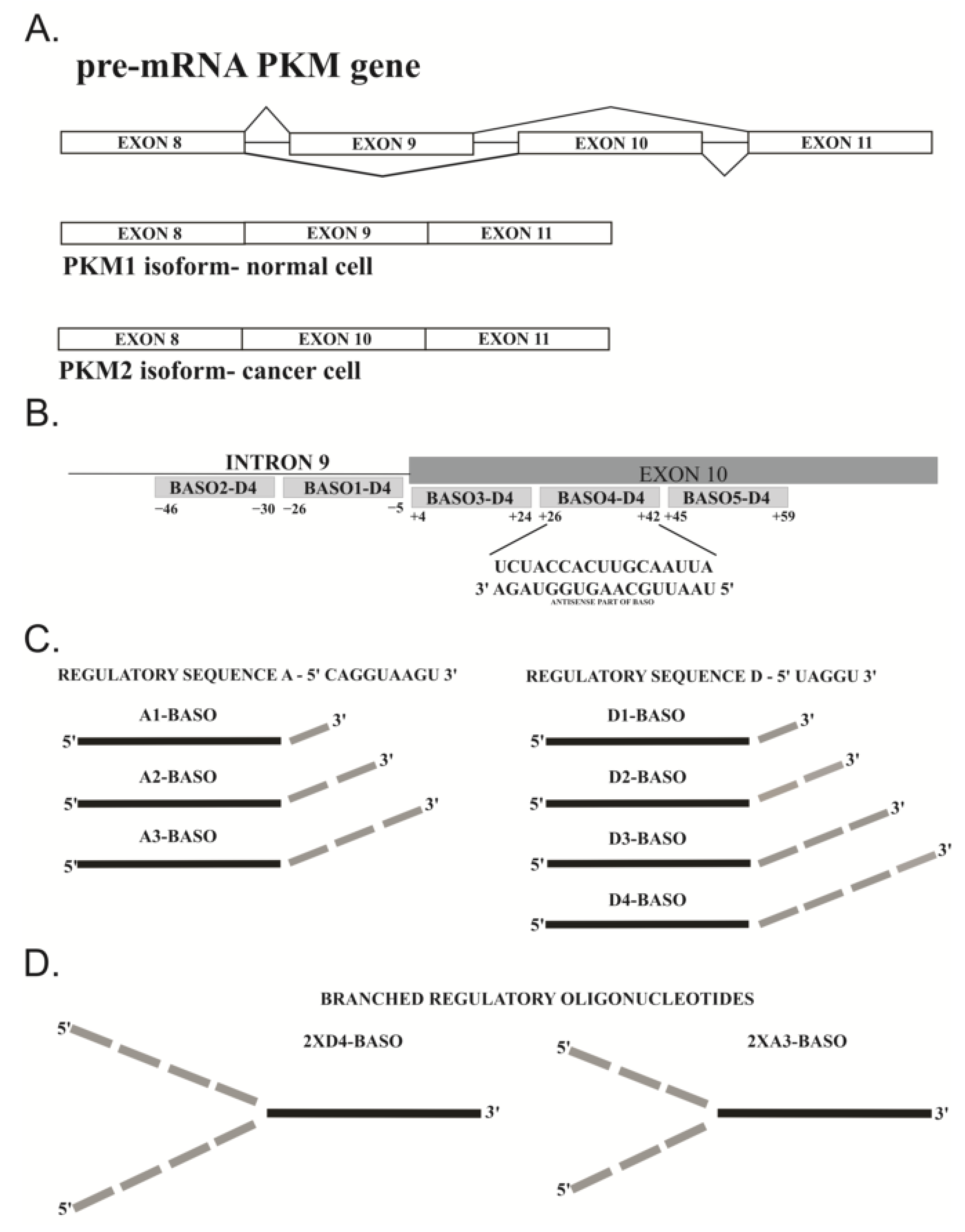
2.2.1. 5′ UAGGU 3′ Regulatory Sequence
2.2.2. 5′ CAGGUAAGU 3′ Regulatory Sequence
2.2.3. Branched BASOs
3. Material and Methods
3.1. Oligonucleotide Synthesis
3.2. hnRNP A1 Protein Production
3.3. Electrophoretic Mobility Shift Assay (EMSA)—Oligonucleotides Screening
3.4. Electrophoretic Mobility Shift Assay (EMSA)-Kd Determination
3.5. UV Melting Experiments
3.6. Cell Culture and Oligonucleotides Transfection
3.7. RT-qPCR Analysis
3.8. qPCR Statistical Analysis
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Sample Availability
References
- Pan, Q.; Shai, O.; Lee, L.J.; Frey, B.J.; Blencowe, B.J. Deep surveying of alternative splicing complexity in the human transcriptome by high-throughput sequencing. Nat. Genet. 2008, 40, 1413–1415. [Google Scholar] [CrossRef] [PubMed]
- Bartys, N.; Kierzek, R.; Lisowiec-Wąchnicka, J. The regulation properties of RNA secondary structure in alternative splicing. Biochim. et Biophys. Acta 2019, 1862, 194401. [Google Scholar] [CrossRef] [PubMed]
- Jiang, W.; Chen, L. Alternative splicing: Human disease and quantitative analysis from high-throughput sequencing. Comput. Struct. Biotechnol. J. 2020, 19, 183–195. [Google Scholar] [CrossRef]
- Scotti, M.M.; Swanson, M.S. RNA mis-splicing in disease. Nat. Rev. Genet. 2015, 17, 19–32. [Google Scholar] [CrossRef]
- Ren, P.; Lu, L.; Cai, S.; Chen, J.; Lin, W.; Han, F. Alternative Splicing: A New Cause and Potential Therapeutic Target in Au-toimmune Disease. Front. Immunol. 2021, 12, 3284. [Google Scholar] [CrossRef]
- Venables, J.P.; Klinck, R.; Koh, C.; Gervais-Bird, J.; Bramard, A.; Inkel, L.; Durand, M.; Couture, S.; Froehlich, U.; Lapointe, E.; et al. Cancer-associated regulation of alternative splicing. Nat. Struct. Mol. Biol. 2009, 16, 670–676. [Google Scholar] [CrossRef] [PubMed]
- Wang, F.; Zuroske, T.; Watts, J.K. RNA therapeutics on the rise. Nat. Rev. Drug Discov. 2020, 19, 441–442. [Google Scholar] [CrossRef] [PubMed]
- Skordis, L.A.; Dunckley, M.G.; Yue, B.; Eperon, I.C.; Muntoni, F. Bifunctional antisense oligonucleotides provide a trans-acting splicing enhancer that stimulates SMN2 gene expression in patient fibroblasts. Proc. Natl. Acad. Sci. USA 2003, 100, 4114–4119. [Google Scholar] [CrossRef]
- Villemaire, J.; Dion, I.; Elela, S.A.; Chabot, B. Reprogramming Alternative Pre-messenger RNA Splicing through the Use of Protein-binding Antisense Oligonucleotides. J. Biol. Chem. 2003, 278, 50031–50039. [Google Scholar] [CrossRef]
- Baughan, T.D.; Dickson, A.; Osman, E.Y.; Lorson, C.L. Delivery of bifunctional RNAs that target an intronic repressor and increase SMN levels in an animal model of spinal muscular atrophy. Hum. Mol. Genet. 2009, 18, 1600–1611. [Google Scholar] [CrossRef]
- Owen, N.; Zhou, H.; Malygin, A.; Sangha, J.; Smith, L.D.; Muntoni, F.; Eperon, I.C. Design principles for bifunctional targeted oligonucleotide enhancers of splicing. Nucleic Acids Res. 2011, 39, 7194–7208. [Google Scholar] [CrossRef] [PubMed]
- Dickson, A.; Osman, E.; Lorson, C.L. A Negatively Acting Bifunctional RNA Increases Survival Motor Neuron Both In Vitro and In Vivo. Hum. Gene Ther. 2008, 19, 1307–1315. [Google Scholar] [CrossRef] [PubMed]
- Jean-Philippe, J.; Paz, S.; Caputi, M. hnRNP A1: The Swiss Army Knife of Gene Expression. Int. J. Mol. Sci. 2013, 14, 18999–19024. [Google Scholar] [CrossRef]
- He, Y.; Smith, R. Nuclear functions of heterogeneous nuclear ribonucleoproteins A/B. Experientia 2008, 66, 1239–1256. [Google Scholar] [CrossRef] [PubMed]
- Gendron, D.; Carriero, S.; Garneau, D.; Villemaire, J.; Klinck, R.; Elela, S.A.; Damha, M.J.; Chabot, B. Modulation of 5’ splice site selection using tailed oligonucleotides carrying splicing signals. BMC Biotechnol. 2006, 6, 5. [Google Scholar] [CrossRef] [PubMed]
- Brosseau, J.-P.; Lucier, J.-F.; Lamarche, A.-A.; Shkreta, L.; Gendron, D.; Lapointe, E.; Thibault, P.; Paquet, E.; Perreault, J.-P.; Elela, S.A.; et al. Redirecting splicing with bifunctional oligonucleotides. Nucleic Acids Res. 2013, 42, e40. [Google Scholar] [CrossRef]
- Warburg, O. On the Origin of Cancer Cells. Science 1956, 123, 309–314. [Google Scholar] [CrossRef]
- Li, Z.; Yang, P.; Li, Z. The Multifaceted Regulation and Functions of PKM2 in Tumor Progression. Biochim. Biophys. Acta Rev. Cancer 2014, 1846, 285–296. [Google Scholar] [CrossRef]
- Guo, W.; Zhang, Z.; Li, G.; Lai, X.; Gu, R.; Xu, W.; Chen, H.; Xing, Z.; Chen, L.; Qian, J.; et al. Pyruvate Kinase M2 Promotes Prostate Cancer Metastasis Through Regulating ERK1/2-COX-2 Signaling. Front. Oncol. 2020, 10, 544288. [Google Scholar] [CrossRef]
- Yu, P.; Li, A.-X.; Chen, X.-S.; Tian, M.; Wang, H.-Y.; Wang, X.-L.; Zhang, Y.; Wang, K.-S.; Cheng, Y. PKM2–c-Myc–Survivin Cascade Regulates the Cell Proliferation, Migration, and Tamoxifen Resistance in Breast Cancer. Front. Pharmacol. 2020, 11, 469. [Google Scholar] [CrossRef]
- Beusch, I.; Barraud, P.; Moursy, A.; Cléry, A.; Allain, F.H.-T. Tandem hnRNP A1 RNA recognition motifs act in concert to repress the splicing of survival motor neuron exon 7. eLife 2017, 6, e25736. [Google Scholar] [CrossRef] [PubMed]
- Burd, C.G.; Dreyfuss, G. RNA Binding Specificity of HnRNP A1: Significance of HnRNP A1 High-Affinity Binding Sites in Pre-MRNA Splicing. EMBO J. 1994, 1, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Hamilton, B.; Nagy, E.; Malter, J.; Arrick, B.; Rigby, W. Association of heterogeneous nuclear ribonucleoprotein A1 and C proteins with reiterated AUUUA sequences. J. Biol. Chem. 1993, 268, 8881–8887. [Google Scholar] [CrossRef]
- Dodon, M.D.; Hamaia, S.; Martin, J.; Gazzolo, L. Heterogeneous Nuclear Ribonucleoprotein A1 Interferes with the Binding of the Human T Cell Leukemia Virus Type 1 Rex Regulatory Protein to Its Response Element. J. Biol. Chem. 2002, 277, 18744–18752. [Google Scholar] [CrossRef] [PubMed]
- Morgan, C.E.; Meagher, J.L.; Levengood, J.D.; Delproposto, J.; Rollins, C.; Stuckey, J.A.; Tolbert, B.S. The First Crystal Structure of the UP1 Domain of hnRNP A1 Bound to RNA Reveals a New Look for an Old RNA Binding Protein. J. Mol. Biol. 2015, 427, 3241–3257. [Google Scholar] [CrossRef] [PubMed]
- De Conti, L.; Skoko, N.; Buratti, E.; Baralle, M. Complexities of 5’splice site definition: Implications in clinical analyses. RNA Biol. 2012, 9, 911–923. [Google Scholar] [CrossRef]
- Liu, X.; Ishizuka, T.; Bao, H.-L.; Wada, K.; Takeda, Y.; Iida, K.; Nagasawa, K.; Yang, D.; Xu, Y. Structure-Dependent Binding of hnRNPA1 to Telomere RNA. J. Am. Chem. Soc. 2017, 139, 7533–7539. [Google Scholar] [CrossRef]
- Cléry, A.; Sinha, R.; Anczuków, O.; Corrionero, A.; Moursy, A.; Daubner, G.M.; Valcárcel, J.; Krainer, A.R.; Allain, F.H.-T. Isolated pseudo–RNA-recognition motifs of SR proteins can regulate splicing using a noncanonical mode of RNA recognition. Proc. Natl. Acad. Sci. USA 2013, 110, E2802–E2811. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.-X.; Zhang, M.; Krainer, A.R. Identification of functional exonic splicing enhancer motifs recognized by individual SR proteins. Genes Dev. 1998, 12, 1998–2012. [Google Scholar] [CrossRef]
- Bluemlein, K.; Grüning, N.-M.; Feichtinger, R.G.; Lehrach, H.; Kofler, B.; Ralser, M. No evidence for a shift in pyruvate kinase PKM1 to PKM2 expression during tumorigenesis. Oncotarget 2011, 2, 393–400. [Google Scholar] [CrossRef]
- Havens, M.A.; Hastings, M.L. Splice-switching antisense oligonucleotides as therapeutic drugs. Nucleic Acids Res. 2016, 44, 6549–6563. [Google Scholar] [CrossRef]
- Wang, Z.; Chatterjee, D.; Jeon, H.Y.; Akerman, M.; Heiden, M.G.V.; Cantley, L.C.; Krainer, A.R. Exon-centric regulation of pyruvate kinase M alternative splicing via mutually exclusive exons. J. Mol. Cell Biol. 2011, 4, 79–87. [Google Scholar] [CrossRef]
- Osman, E.Y.; Yen, P.-F.; Lorson, C.L. Bifunctional RNAs Targeting the Intronic Splicing Silencer N1 Increase SMN Levels and Reduce Disease Severity in an Animal Model of Spinal Muscular Atrophy. Mol. Ther. 2012, 20, 119–126. [Google Scholar] [CrossRef]
- Carriero, S. Inhibition of pre-mRNA splicing by synthetic branched nucleic acids. Nucleic Acids Res. 2003, 31, 6157–6167. [Google Scholar] [CrossRef] [PubMed]
- Carriero, S.; Braich, R.S.; Hudson, R.H.E.; Anglin, D.; Friesen, J.D.; Damha, M.J. Inhibition ofin vitropre-mrna splicing ins. cerevisiaeby branched oligonucleotides. Nucleosides Nucleotides Nucleic Acids 2001, 20, 873–877. [Google Scholar] [CrossRef] [PubMed]
- Damha, M.J.; Ganeshan, K.; Hudson, R.H.; Zabarylo, S.V. Solid-phase synthesis of branched oligoribonucleotides related to messenger RNA splicing intermediates. Nucleic Acids Res. 1992, 20, 6565–6573. [Google Scholar] [CrossRef] [PubMed]


| Name | Sequence (5′ -> 3′) |
|---|---|
| A2 | CAGGUAAGU CAGGUAAGU |
| A3 | CAGGUAAGU CAGGUAAGU CAGGUAAGU |
| B2 | CAGGUGAGU CAGGUGAGU |
| B3 | CAGGUGAGU CAGGUGAGU CAGGUGAGU |
| C2 | UAGGA UAGGA |
| C4 | UAGGA UAGGA UAGGA UAGGA |
| D2 | UAGGU UAGGU |
| D4 | UAGGU UAGGU UAGGU UAGGU |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bartyś, N.; Pasternak, A.; Lisowiec-Wąchnicka, J. Optimization of Bifunctional Antisense Oligonucleotides for Regulation of Mutually Exclusive Alternative Splicing of PKM Gene. Molecules 2022, 27, 5682. https://doi.org/10.3390/molecules27175682
Bartyś N, Pasternak A, Lisowiec-Wąchnicka J. Optimization of Bifunctional Antisense Oligonucleotides for Regulation of Mutually Exclusive Alternative Splicing of PKM Gene. Molecules. 2022; 27(17):5682. https://doi.org/10.3390/molecules27175682
Chicago/Turabian StyleBartyś, Natalia, Anna Pasternak, and Jolanta Lisowiec-Wąchnicka. 2022. "Optimization of Bifunctional Antisense Oligonucleotides for Regulation of Mutually Exclusive Alternative Splicing of PKM Gene" Molecules 27, no. 17: 5682. https://doi.org/10.3390/molecules27175682
APA StyleBartyś, N., Pasternak, A., & Lisowiec-Wąchnicka, J. (2022). Optimization of Bifunctional Antisense Oligonucleotides for Regulation of Mutually Exclusive Alternative Splicing of PKM Gene. Molecules, 27(17), 5682. https://doi.org/10.3390/molecules27175682








