Current Updates on Naturally Occurring Compounds Recognizing SARS-CoV-2 Druggable Targets
Abstract
1. Introduction
2. Results
2.1. Natural Occurring Compounds Undergoing Clinical Trials
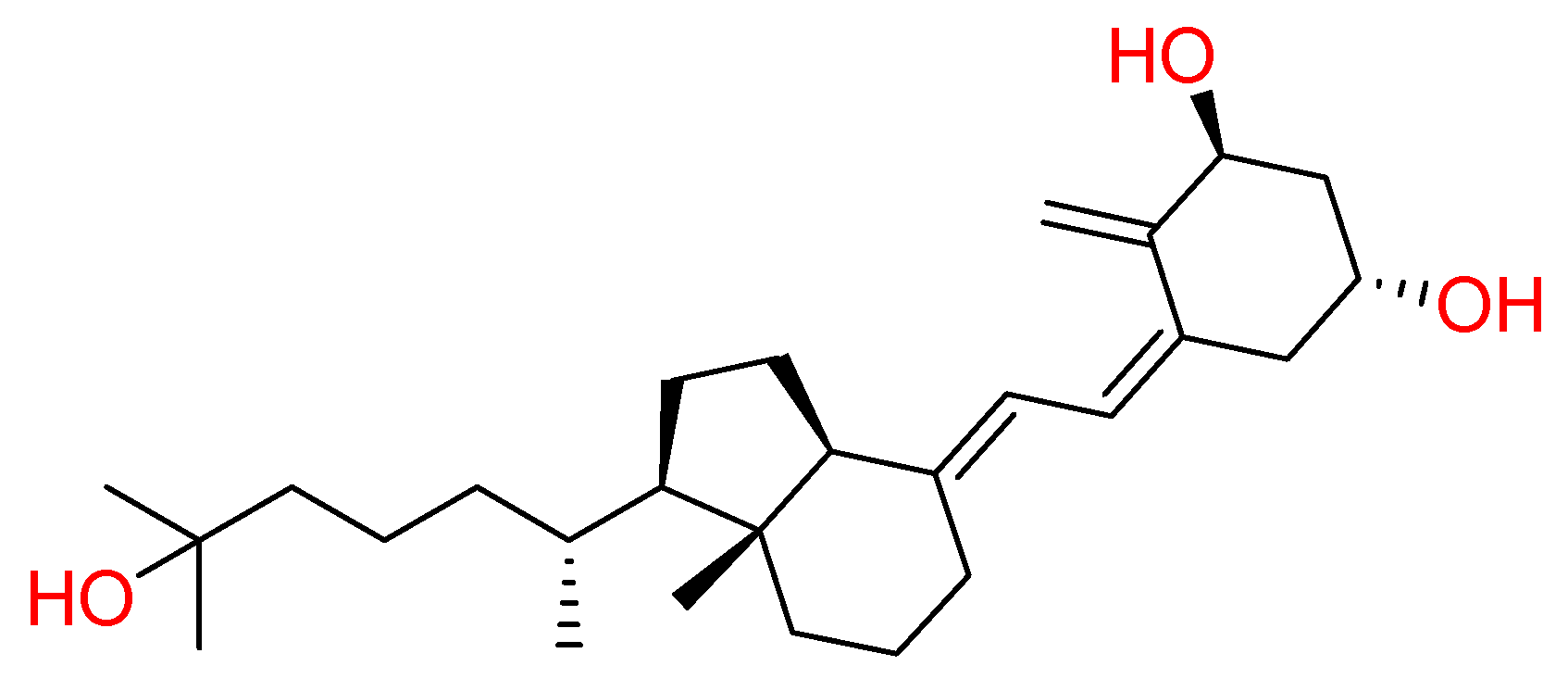
2.1.1. Vitamin D
2.1.2. Vitamin C
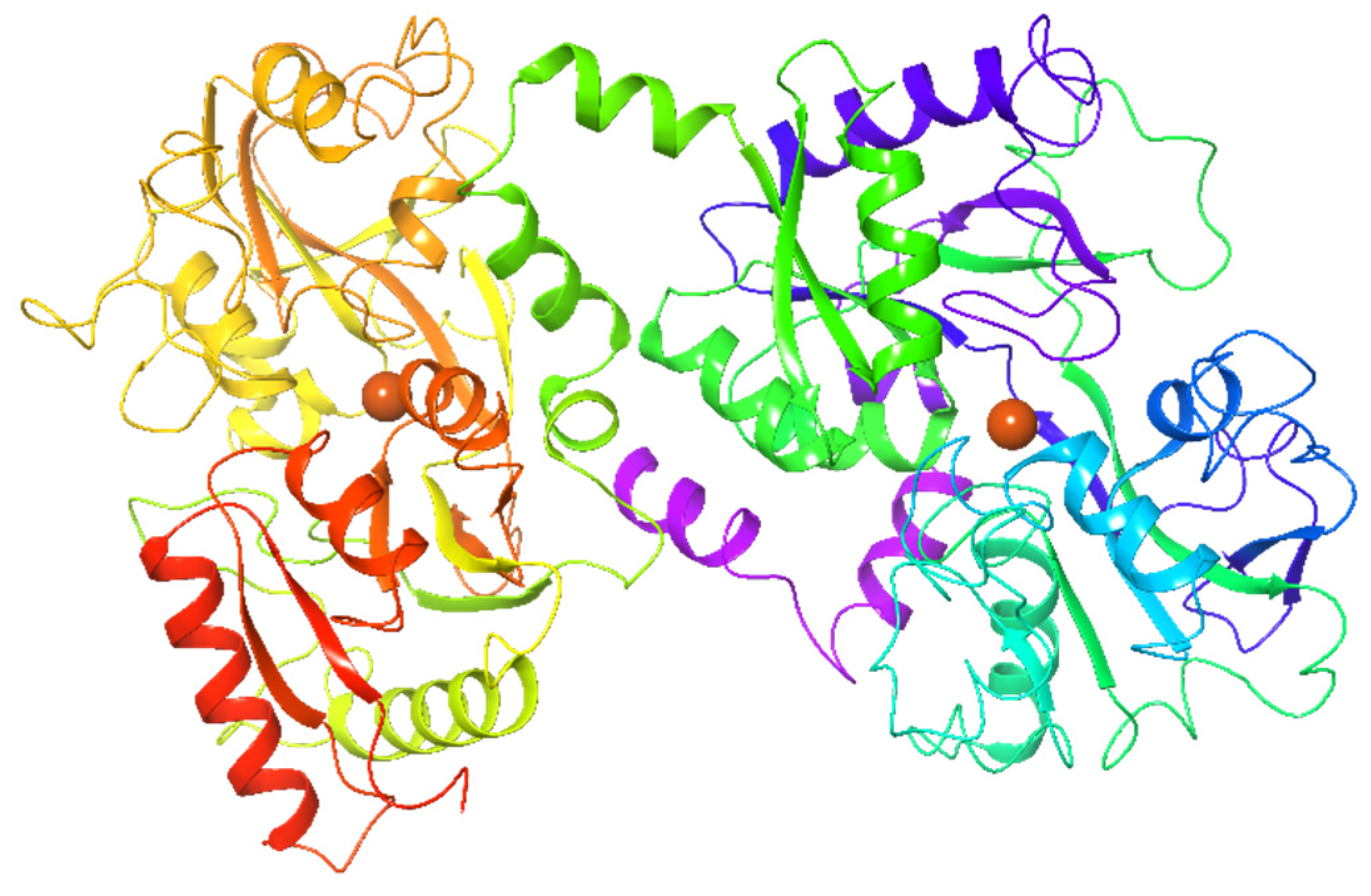
2.1.3. Lactoferrin
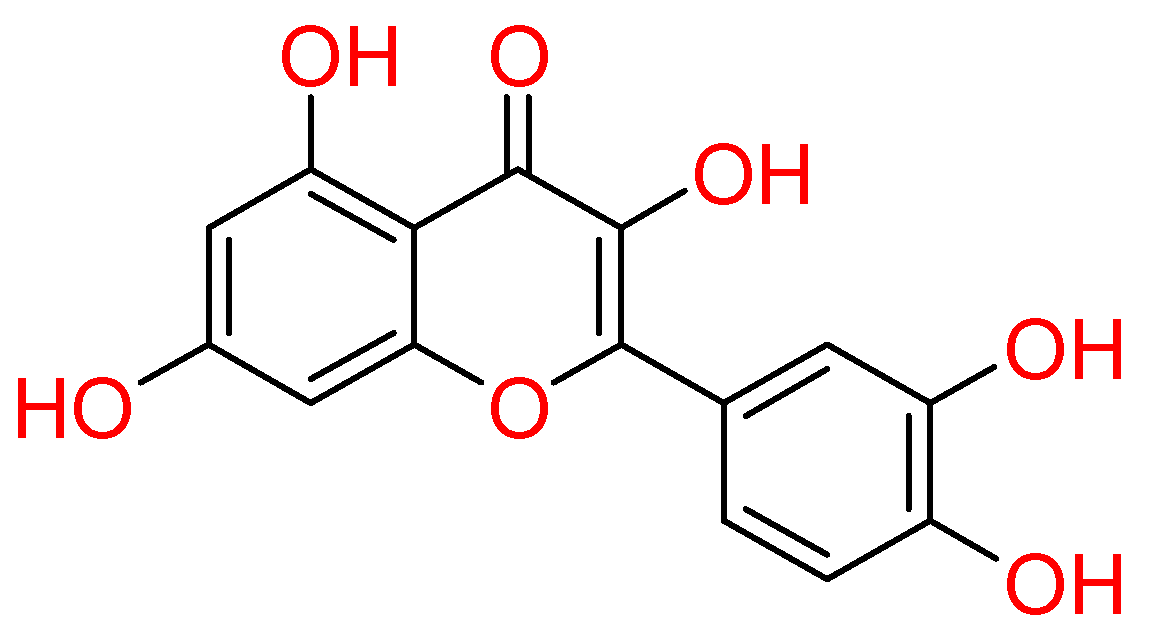
2.1.4. Quercetin
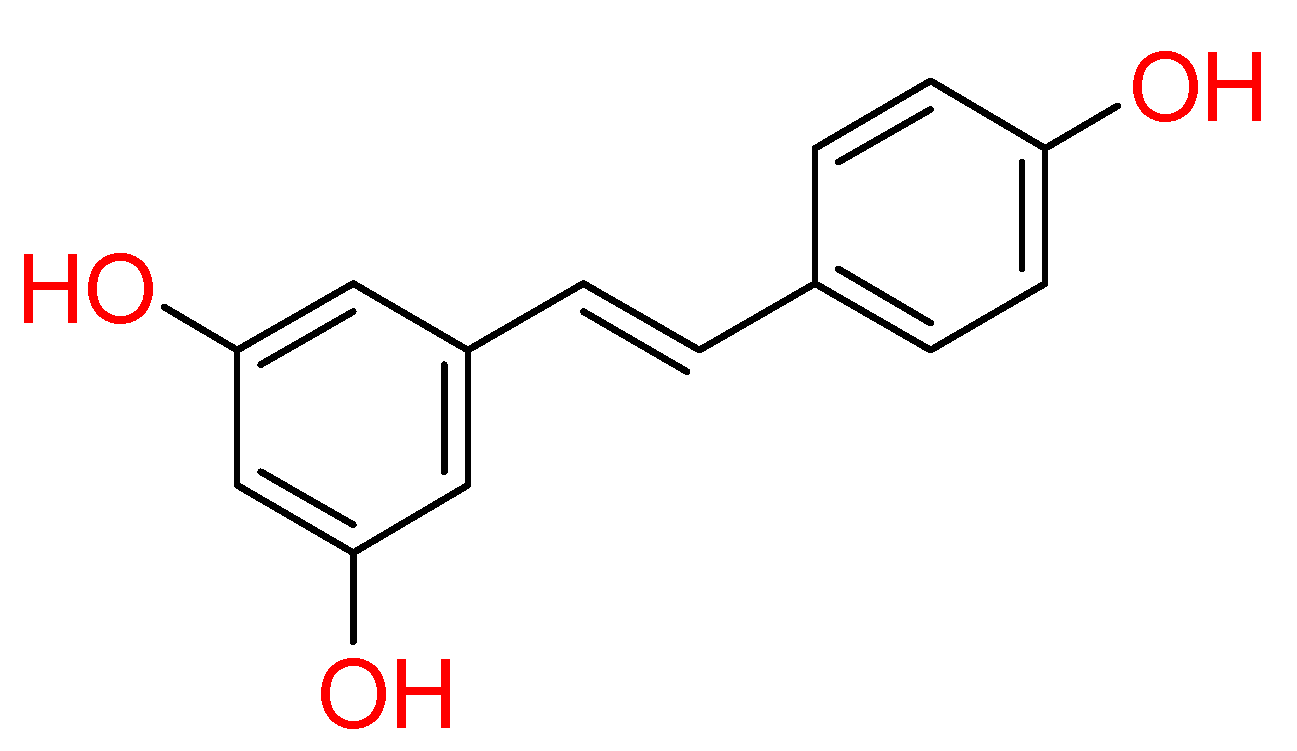
2.1.5. Resveratrol
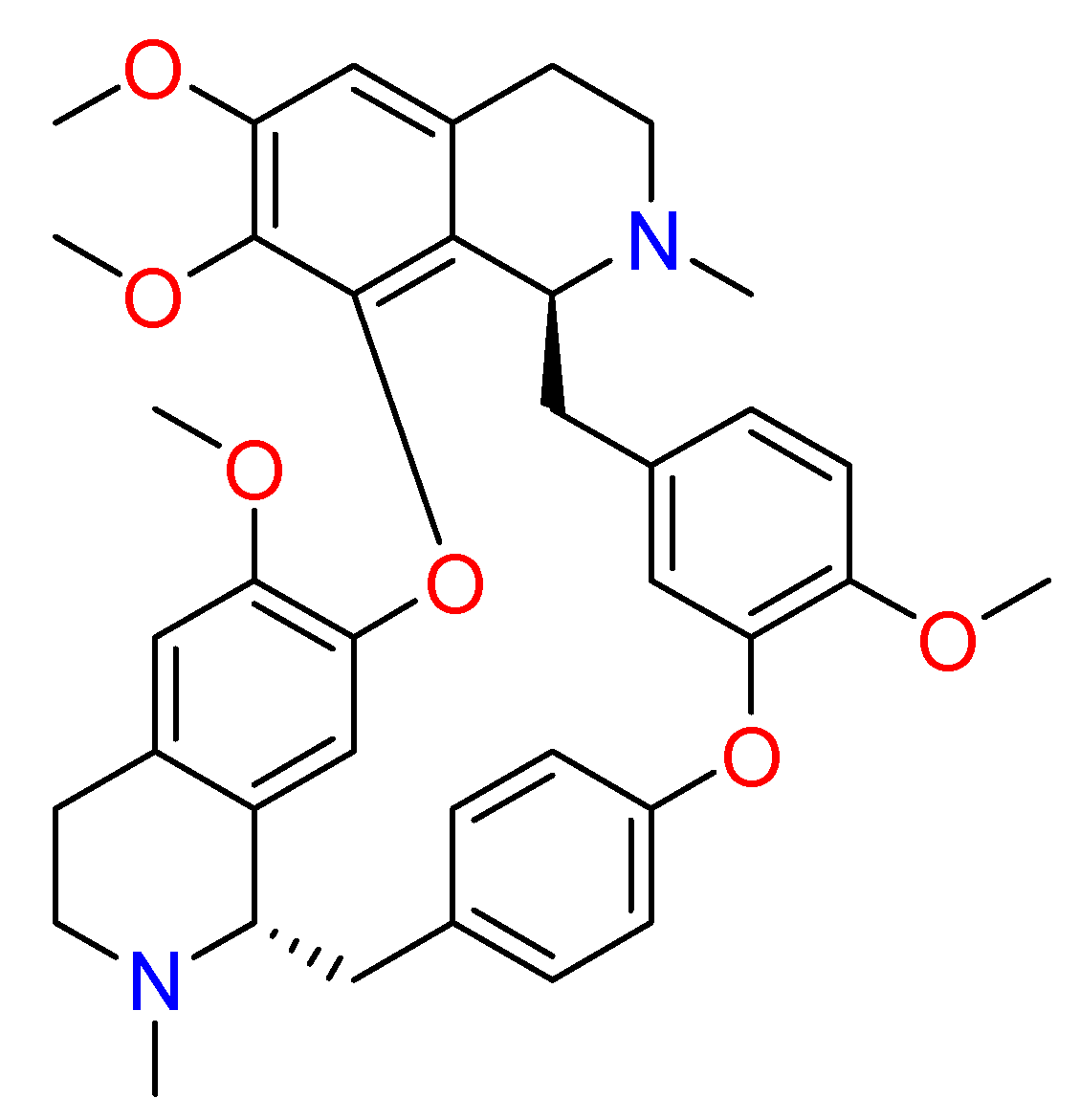
2.1.6. Hanfangchin A
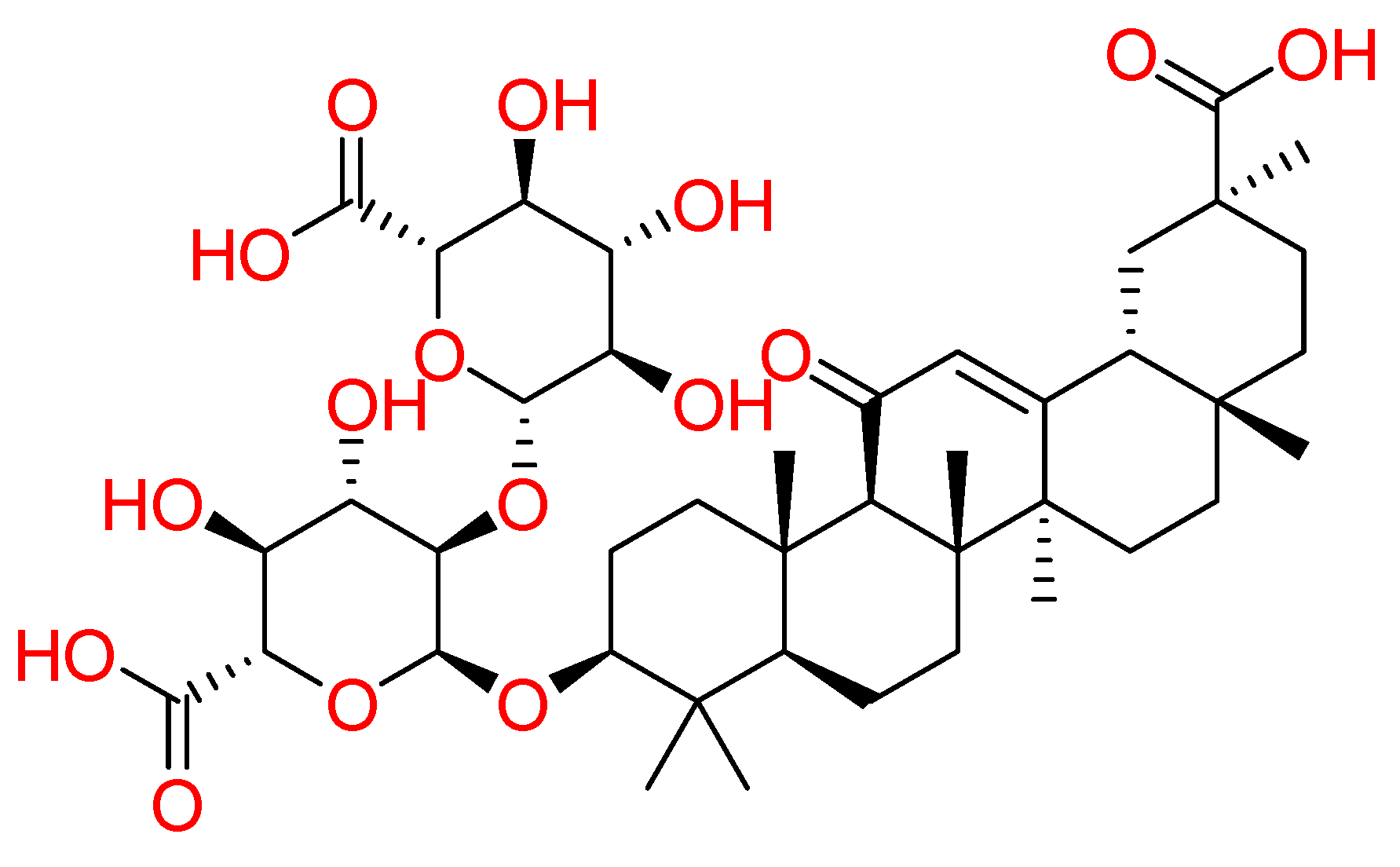
2.1.7. Glycyrrhizin
2.1.8. Artemisinin
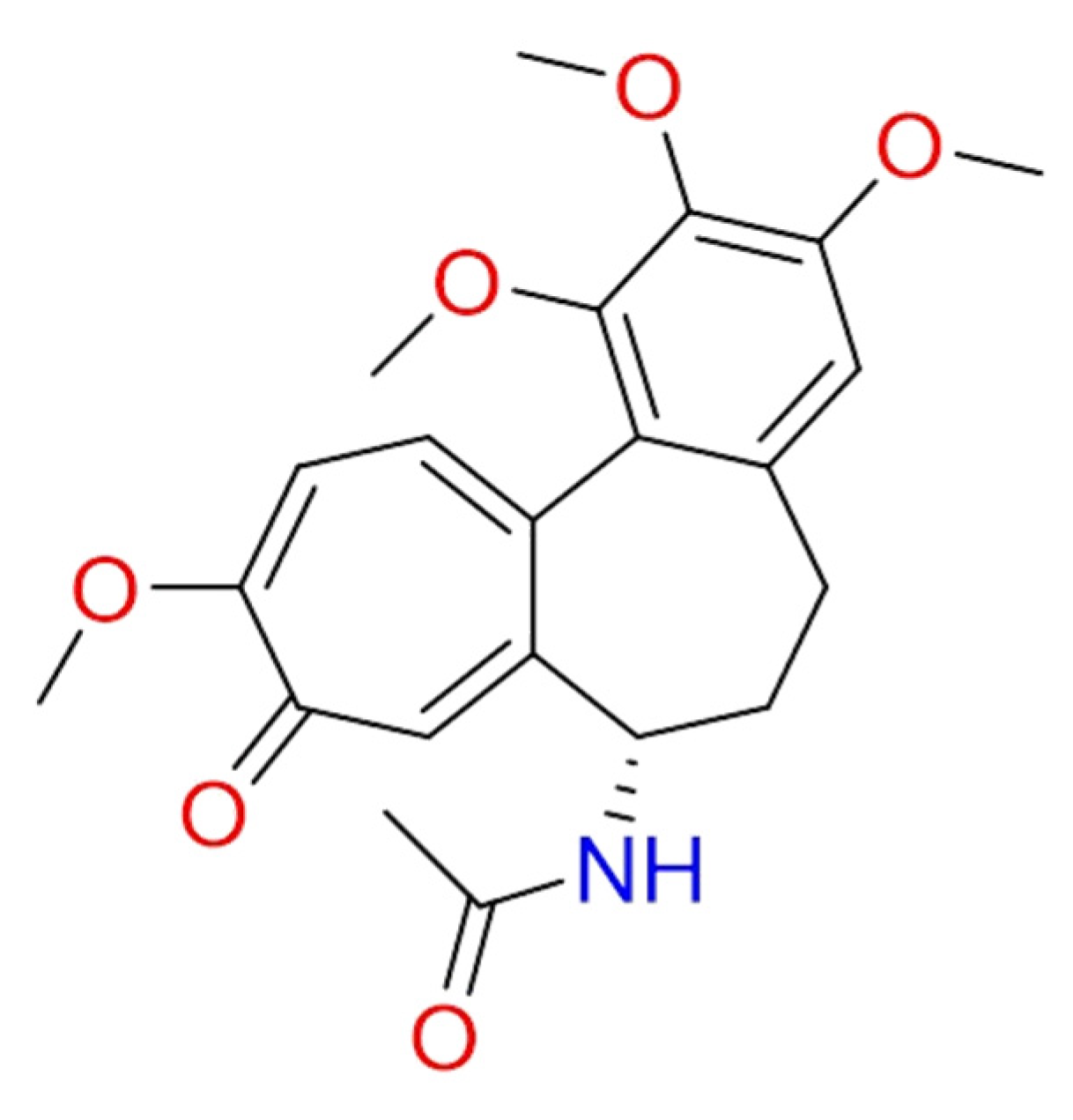
2.1.9. Colchicine
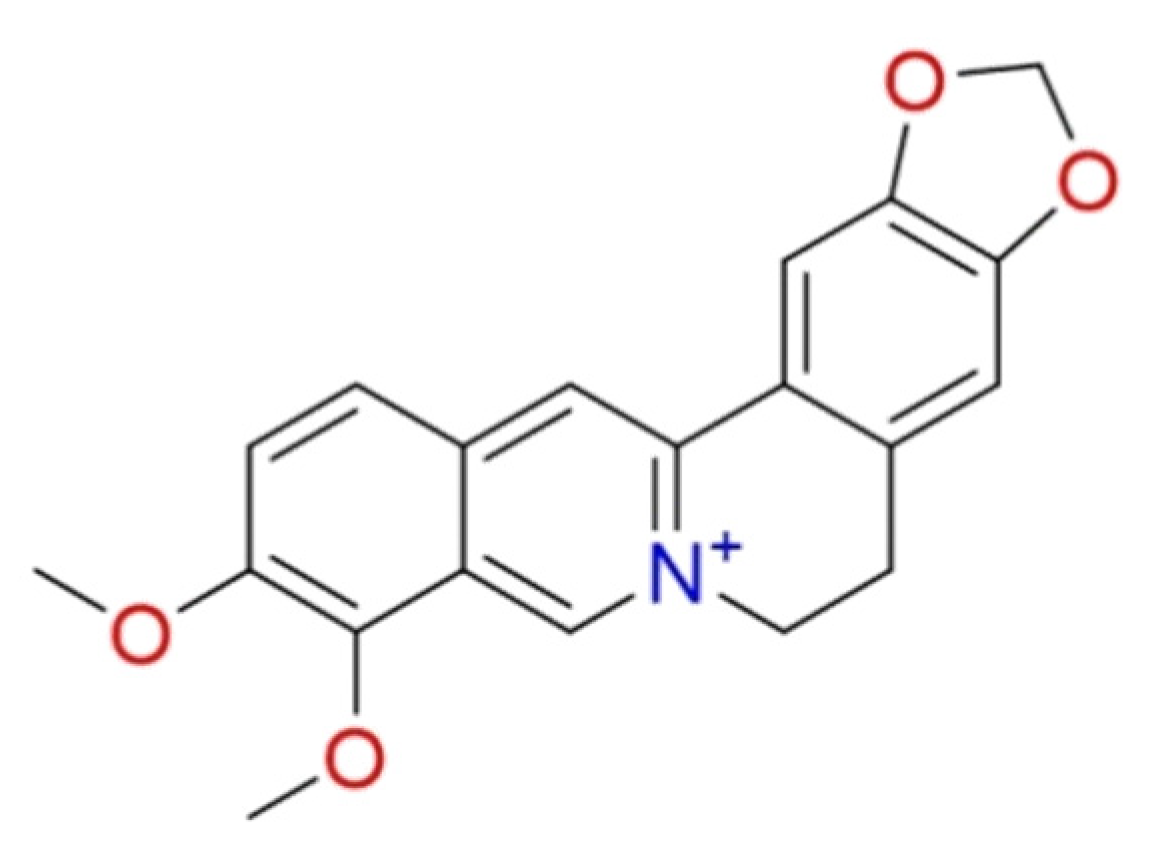
2.1.10. Berberine
2.2. Natural Compounds with Promising Activities Against SARS-CoV-2 Infection
2.2.1. Alkaloids
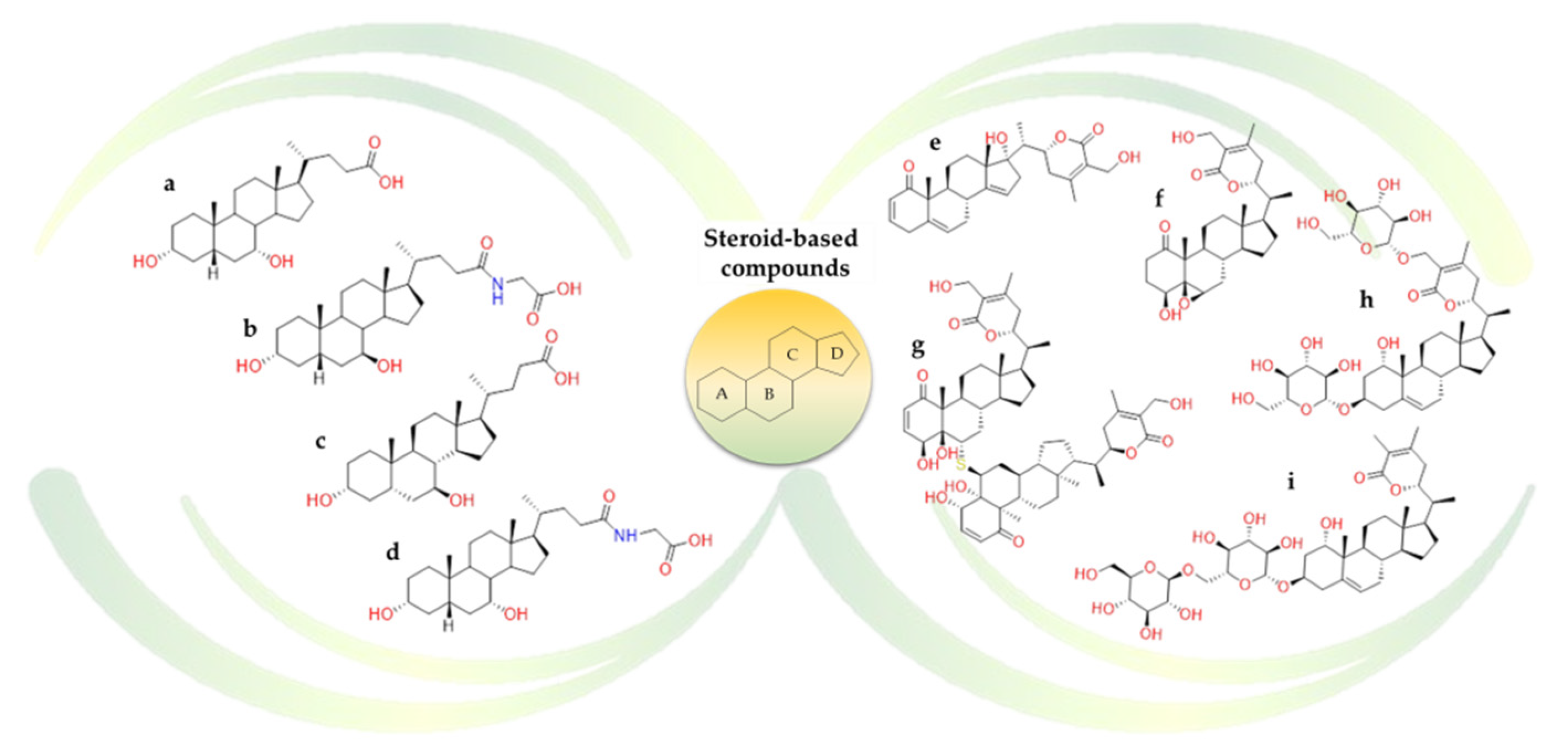
2.2.2. Terpenes
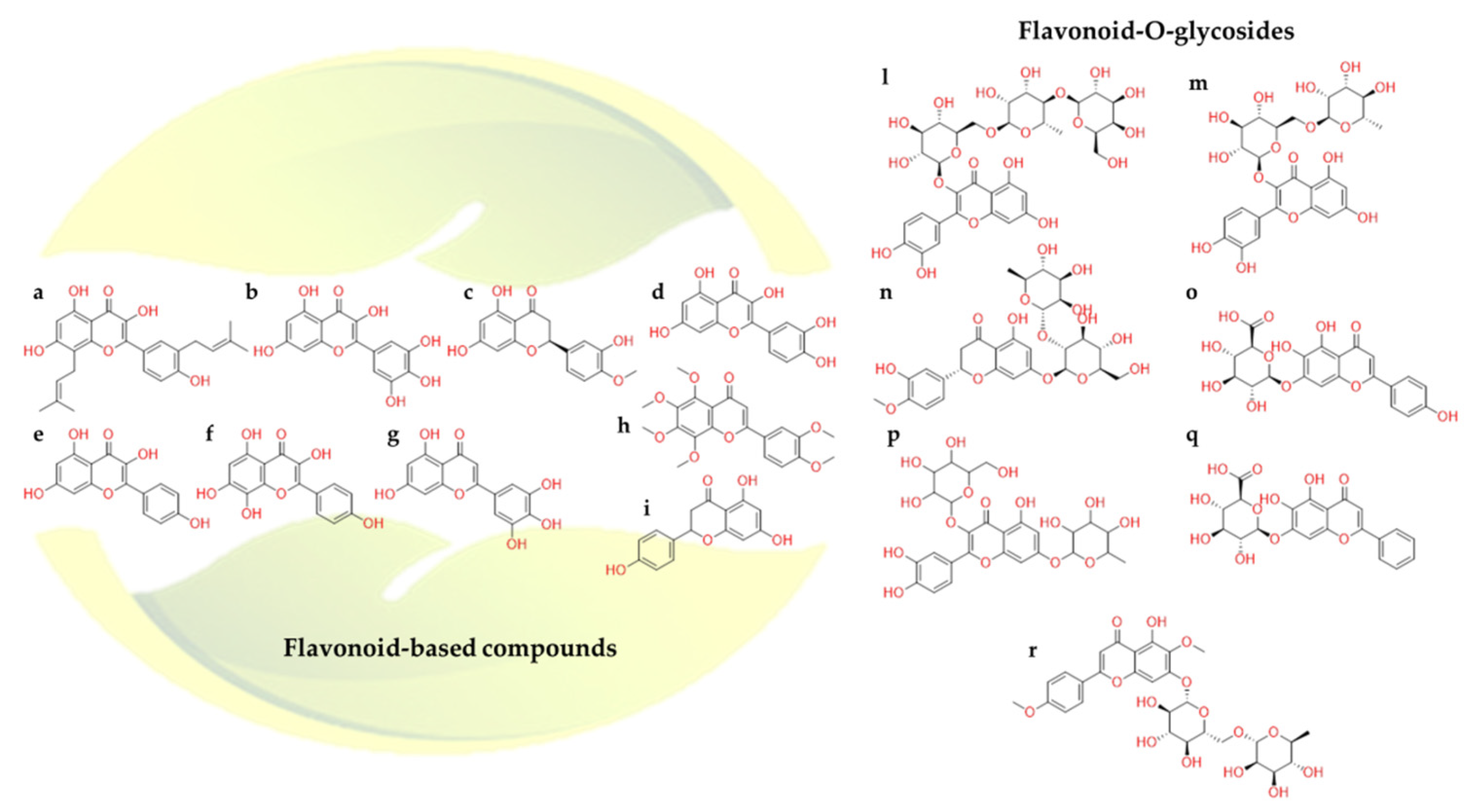
2.2.3. Flavonoids
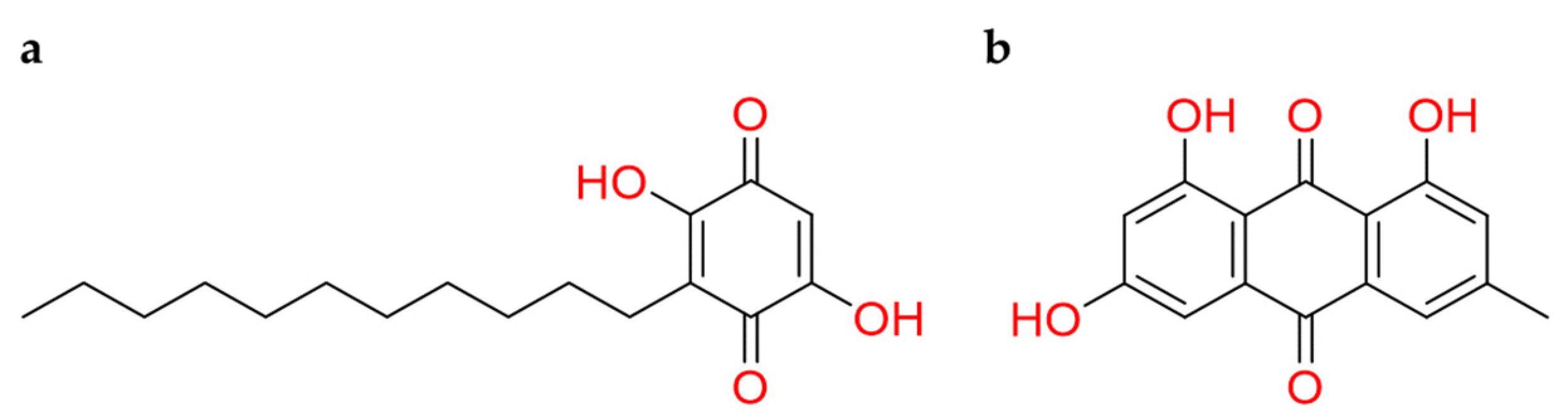
2.2.4. Benzoquinones
2.2.5. Other Natural Compounds
3. Discussion and Future Perspectives
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Zhou, P.; Yang, X.-L.; Wang, X.-G.; Hu, B.; Zhang, L.; Zhang, W.; Si, H.-R.; Zhu, Y.; Li, B.; Huang, C.-L.; et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 2020, 579, 270–273. [Google Scholar] [CrossRef] [PubMed]
- Borba, M.G.S.; Val, F.F.A.; Hajjar, L.A.; Pinto, R.C.; Balieiro, A.A.S.; Pacheco, A.G.F.; Santos, J.D.O.; Naveca, F.G.; Xavier, M.S.; Siqueira, A.M.; et al. Effect of High vs Low Doses of Chloroquine Diphosphate as Adjunctive Therapy for Patients Hospitalized With Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2) Infection: A Randomized Clinical Trial. JAMA Netw. Open 2020, 3, e208857. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.T.; Leung, K.; Leung, G.M. Nowcasting and forecasting the potential domestic and international spread of the 2019-nCoV outbreak originating in Wuhan, China: A modelling study. Lancet 2020, 395, 689–697. [Google Scholar] [CrossRef]
- Oliver, S.E.; Gargano, J.W.; Marin, M.; Wallace, M.; Curran, K.G.; Chamberland, M.; McClung, N.; Campos-Outcalt, D.; Morgan, R.L.; Mbaeyi, S.; et al. The Advisory Committee on Immunization Practices’ Interim Recommendation for Use of Pfizer-BioNTech COVID-19 Vaccine—United States, December 2020. MMWR. Morb. Mortal. Wkly. Rep. 2020, 69, 1922–1924. [Google Scholar] [CrossRef] [PubMed]
- Vasconcelos, M.H.; Alcaro, S.; Arechavala-Gomeza, V.; Baumbach, J.; Borges, F.; Brevini, T.A.; Rivas, J.D.L.; Devaux, Y.; Hozak, P.; Keinänen-Toivola, M.M.; et al. Joining European Scientific Forces to Face Pandemics. Trends Microbiol. 2020. [Google Scholar] [CrossRef]
- Salpini, R.; Alkhatib, M.; Costa, G.; Piermatteo, L.; Ambrosio, F.A.; Di Maio, V.C.; Scutari, R.; Duca, L.; Berno, G.; Fabeni, L.; et al. Key genetic elements, single and in clusters, underlying geographically dependent SARS-CoV-2 genetic adaptation and their impact on binding affinity for drugs and immune control. J. Antimicrob. Chemother. 2020. [Google Scholar] [CrossRef]
- Ortuso, F.; Mercatelli, D.; Guzzi, P.H.; Giorgi, F.M. Structural Genetics of circulating variants affecting the SARS CoV-2 Spike/human ACE2 complex. bioRxiv 2020. [Google Scholar] [CrossRef]
- Shen, Z.; Xiao, Y.; Kang, L.; Ma, W.; Shi, L.; Zhang, L.; Zhou, Z.; Yang, J.; Zhong, J.; Yang, D.; et al. Genomic Diversity of Severe Acute Respiratory Syndrome–Coronavirus 2 in Patients with Coronavirus Disease 2019. Clin. Infect. Dis. 2020, 71, 713–720. [Google Scholar] [CrossRef]
- Plante, J.A.; Liu, Y.; Liu, J.; Xia, H.; Johnson, B.A.; Lokugamage, K.G.; Zhang, X.; Muruato, A.E.; Zou, J.; Fontes-Garfias, C.R.; et al. Spike mutation D614G alters SARS-CoV-2 fitness. Nature 2020. [Google Scholar] [CrossRef]
- Bagetta, D.; Maruca, A.; Lupia, A.; Mesiti, F.; Catalano, R.; Romeo, I.; Moraca, F.; Ambrosio, F.A.; Costa, G.; Artese, A.; et al. Mediterranean products as promising source of multi-target agents in the treatment of metabolic syndrome. Eur. J. Med. Chem. 2020, 186, 111903. [Google Scholar] [CrossRef]
- Maruca, A.; Catalano, R.; Bagetta, D.; Mesiti, F.; Ambrosio, F.A.; Romeo, I.; Moraca, F.; Rocca, R.; Ortuso, F.; Artese, A.; et al. The Mediterranean Diet as source of bioactive compounds with multi-targeting anti-cancer profile. Eur. J. Med. Chem. 2019, 181, 111579. [Google Scholar] [CrossRef] [PubMed]
- Medicinal plants and primary health care: Part 2. Essent. Drugs Monit. 1991, 11, 15–17.
- Narkhede, R.R.; Pise, A.V.; Cheke, R.S.; Shinde, S.D. Recognition of Natural Products as Potential Inhibitors of COVID-19 Main Protease (Mpro): In-Silico Evidences. Nat. Prod. Bioprospect. 2020, 10, 297–306. [Google Scholar] [CrossRef] [PubMed]
- David, B.; Wolfender, J.-L.; Dias, D.A. The pharmaceutical industry and natural products: Historical status and new trends. Phytochem. Rev. 2015, 14, 299–315. [Google Scholar] [CrossRef]
- Firn, R.D.; Jones, C.G. Natural Products—A Simple Model to Explain Chemical Diversity. Nat. Prod. Rep. 2003, 20, 382–391. [Google Scholar] [CrossRef]
- Dias, D.A.; Urban, S.; Roessner, U. A Historical Overview of Natural Products in Drug Discovery. Metabolites 2012, 2, 303–336. [Google Scholar] [CrossRef]
- Catalogna, G.; Moraca, F.; D’Antona, L.; Dattilo, V.; Perrotti, G.; Lupia, A.; Costa, G.; Ortuso, F.; Iuliano, R.; Trapasso, F.; et al. Review about the multi-target profile of resveratrol and its implication in the SGK1 inhibition. Eur. J. Med. Chem. 2019, 183, 111675. [Google Scholar] [CrossRef]
- Deorukhkar, A.; Krishnan, S.; Sethi, G.; Aggarwal, B.B. Back to basics: How natural products can provide the basis for new therapeutics. Expert Opin. Investig. Drugs 2007, 16, 1753–1773. [Google Scholar] [CrossRef] [PubMed]
- Lani, R.; Hassandarvish, P.; Shu, M.-H.; Phoon, W.H.; Chu, J.J.H.; Higgs, S.; VanLandingham, D.; Abu Bakar, S.; Zandi, K. Antiviral activity of selected flavonoids against Chikungunya virus. Antivir. Res. 2016, 133, 50–61. [Google Scholar] [CrossRef] [PubMed]
- Zakaryan, H.; Arabyan, E.; Oo, A.; Zandi, K. Flavonoids: Promising natural compounds against viral infections. Arch. Virol. 2017, 162, 2539–2551. [Google Scholar] [CrossRef] [PubMed]
- Artese, A.; Svicher, V.; Costa, G.; Salpini, R.; Di Maio, V.C.; Alkhatib, M.; Ambrosio, F.A.; Santoro, M.M.; Assaraf, Y.G.; Alcaro, S.; et al. Current status of antivirals and druggable targets of SARS CoV-2 and other human pathogenic coronaviruses. Drug Resist. Updat. 2020, 53, 100721. [Google Scholar] [CrossRef] [PubMed]
- Berretta, A.A.; Silveira, M.A.D.; Capcha, J.M.C.; De Jong, D. Propolis and its potential against SARS-CoV-2 infection mechanisms and COVID-19 disease: Running title: Propolis against SARS-CoV-2 infection and COVID-19. Biomed. Pharmacother. 2020, 131, 110622. [Google Scholar] [CrossRef] [PubMed]
- Benskin, L.L. A basic review of the preliminary evidence that COVID-19 risk and severity is increased in vitamin D deficiency. Front. Public Health 2020, 8, 513. [Google Scholar] [CrossRef] [PubMed]
- Cannell, J.J.; Vieth, R.; Umhau, J.C.; Holick, M.F.; Grant, W.B.; Madronich, S.; Garland, C.F.; Giovannucci, E. Epidemic influenza and vitamin D. Epidemiol. Infect. 2006, 134, 1129–1140. [Google Scholar] [CrossRef] [PubMed]
- Bracale, R.; Vaccaro, C.M.; Coletta, V.; Cricelli, C.; Gamaleri, F.C.; Parazzini, F.; Carruba, M. Nutrition behaviour and compliance with the Mediterranean diet pyramid recommendations: An Italian survey-based study. Eat. Weight Disord. Stud. Anorex. Bulim. Obesity 2020, 25, 1789–1798. [Google Scholar] [CrossRef]
- Azrielant, S.; Shoenfeld, Y. Vitamin D and the immune system. Isr. Med. Assoc. J. 2017, 19, 510–511. [Google Scholar]
- Hughes, D.A.; Norton, R. Vitamin D and respiratory health. Clin. Exp. Immunol. 2009, 158, 20–25. [Google Scholar] [CrossRef]
- Martineau, A.R.; Forouhi, N.G. Vitamin D for COVID-19: A case to answer? Lancet Diabetes Endocrinol. 2020, 8, 735–736. [Google Scholar] [CrossRef]
- Charoute, H.; Saile, R.; Barakat, A. Computational Approach Revealed Potential Affinity of Antiasthmatics against Receptor Binding Domain of 2019n-Cov Spike Glycoprotein. ChemRxiv 2020. [Google Scholar] [CrossRef]
- Weir, E.K.; Thenappan, T.; Bhargava, M.; Chen, Y. Does vitamin D deficiency increase the severity of COVID-19? Clin. Med. 2020, 20, e107–e108. [Google Scholar] [CrossRef]
- Jain, A.; Chaurasia, R.; Sengar, N.S.; Singh, M.; Mahor, S.; Narain, S. Analysis of vitamin D level among asymptomatic and critically ill COVID-19 patients and its correlation with inflammatory markers. Sci. Rep. 2020, 10, 20191. [Google Scholar] [CrossRef] [PubMed]
- Torjesen, I. Evidence does not support vitamin D for reducing respiratory infections, reviews conclude. BMJ 2020, 369, m2629. [Google Scholar] [CrossRef] [PubMed]
- Kow, C.S.; Hadi, M.A.; Hasan, S.S. Vitamin D Supplementation in Influenza and COVID-19 Infections Comment on: “Evidence that Vitamin D Supplementation Could Reduce Risk of Influenza and COVID-19 Infections and Deaths” Nutrients. Nutrients 2020, 12, 1626. [Google Scholar] [CrossRef] [PubMed]
- Ali, N. Role of vitamin D in preventing of COVID-19 infection, progression and severity. J. Infect. Public Health 2020, 13, 1373–1380. [Google Scholar] [CrossRef]
- Lykkesfeldt, J.; Michels, A.J.; Frei, B. Vitamin C. Adv. Nutr. 2014, 5, 16–18. [Google Scholar] [CrossRef]
- Cheng, R.Z. Can early and high intravenous dose of vitamin C prevent and treat coronavirus disease 2019 (COVID-19)? Med. Drug Discov. 2020, 5, 100028. [Google Scholar] [CrossRef] [PubMed]
- Cerullo, G.; Negro, M.; Parimbelli, M.; Pecoraro, M.; Perna, S.; Liguori, G.; Rondanelli, M.; Cena, H.; D’Antona, G. The Long History of Vitamin C: From Prevention of the Common Cold to Potential Aid in the Treatment of COVID-19. Front. Immunol. 2020, 11, 574029. [Google Scholar] [CrossRef] [PubMed]
- Padayatty, S.J.; Katz, A.; Wang, Y.; Eck, P.; Kwon, O.; Lee, J.-H.; Chen, S.; Corpe, C.; Dutta, A.; Dutta, S.K.; et al. Vitamin C as an Antioxidant: Evaluation of Its Role in Disease Prevention. J. Am. Coll. Nutr. 2003, 22, 18–35. [Google Scholar] [CrossRef]
- Liu, F.; Zhu, Y.; Zhang, J.; Li, Y.; Peng, Z. Intravenous high-dose vitamin C for the treatment of severe COVID-19: Study protocol for a multicentre randomised controlled trial. BMJ Open 2020, 10, e039519. [Google Scholar] [CrossRef]
- Hemilä, H.; Chalker, E. Vitamin C for preventing and treating the common cold. Cochrane Database Syst. Rev. 2013. [Google Scholar] [CrossRef]
- Hemilä, H.; Chalker, E. Vitamin C as a Possible Therapy for COVID-19. Infect. Chemother. 2020, 52, 222–223. [Google Scholar] [CrossRef] [PubMed]
- Hemilä, H.; Chalker, E. Vitamin C Can Shorten the Length of Stay in the ICU: A Meta-Analysis. Nutrients 2019, 11, 708. [Google Scholar] [CrossRef] [PubMed]
- Hemilä, H.; Chalker, E. Vitamin C may reduce the duration of mechanical ventilation in critically ill patients: A meta-regression analysis. J. Intensive Care 2020, 8, 15. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Rao, X.; Li, Y.; Zhu, Y.; Liu, F.; Guo, G.; Luo, G.; Meng, Z.; De Backer, D.; Xiang, H.; et al. Pilot trial of high-dose vitamin C in critically ill COVID-19 patients. Ann. Intensiv. Care 2021, 11, 5. [Google Scholar] [CrossRef] [PubMed]
- Carr, A.C.; Rowe, S. The Emerging Role of Vitamin C in the Prevention and Treatment of COVID-19. Nutrients 2020, 12, 3286. [Google Scholar] [CrossRef]
- Bae, M.; Kim, H. Mini-Review on the Roles of Vitamin C, Vitamin D, and Selenium in the Immune System against COVID-19. Molecules 2020, 25, 5346. [Google Scholar] [CrossRef]
- Andersen, B.F.; Baker, H.M.; Morris, G.E.; Rumball, S.V.; Baker, E.N. Apolactoferrin structure demonstrates ligand-induced conformational change in transferrins. Nature 1990, 344, 784–787. [Google Scholar] [CrossRef]
- Vogel, H.J. Lactoferrin, a bird’s eye view. Biochem. Cell Biol. 2012, 90, 233–244. [Google Scholar] [CrossRef]
- Karav, S.; German, J.B.; Rouquié, C.; Le Parc, A.; Barile, D. Studying Lactoferrin N-Glycosylation. Int. J. Mol. Sci. 2017, 18, 870. [Google Scholar] [CrossRef]
- Sun, X.-L.; Baker, H.M.; Shewry, S.C.; Jameson, G.B.; Baker, E.N. Structure of recombinant human lactoferrin expressed in Aspergillus awamori. Acta Crystallogr. Sect. D Biol. Crystallogr. 1999, 55, 403–407. [Google Scholar] [CrossRef]
- Wang, B.; Timilsena, Y.P.; Blanch, E.; Adhikari, B. Lactoferrin: Structure, function, denaturation and digestion. Crit. Rev. Food Sci. Nutr. 2019, 59, 580–596. [Google Scholar] [CrossRef] [PubMed]
- Kell, D.B.; Heyden, E.L.; Pretorius, E. The Biology of Lactoferrin, an Iron-Binding Protein That Can Help Defend Against Viruses and Bacteria. Front. Immunol. 2020, 11, 1221. [Google Scholar] [CrossRef] [PubMed]
- Ganz, T. Iron and infection. Int. J. Hematol. 2018, 107, 7–15. [Google Scholar] [CrossRef] [PubMed]
- Siqueiros-Cendón, T.; Arévalo-Gallegos, S.; Iglesias-Figueroa, B.F.; García-Montoya, I.A.; Salazar-Martínez, J.; Rascón-Cruz, Q. Immunomodulatory effects of lactoferrin. Acta Pharmacol. Sin. 2014, 35, 557–566. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, Y.A.; Wong, H.; Ashida, K.-Y.; Schryvers, A.B.; Lönnerdal, B. The N1 Domain of Human Lactoferrin Is Required for Internalization by Caco-2 Cells and Targeting to the Nucleus. Biochemistry 2008, 47, 10915–10920. [Google Scholar] [CrossRef] [PubMed]
- Cutone, A.; Rosa, L.; Lepanto, M.S.; Scotti, M.J.; Berlutti, F.; Di Patti, M.C.B.; Musci, G.; Valenti, P. Lactoferrin Efficiently Counteracts the Inflammation-Induced Changes of the Iron Homeostasis System in Macrophages. Front. Immunol. 2017, 8, 705. [Google Scholar] [CrossRef] [PubMed]
- Valenti, P.; Frioni, A.; Rossi, A.; Ranucci, S.; De Fino, I.; Cutone, A.; Rosa, L.; Bragonzi, A.; Berlutti, F. Aerosolized bovine lactoferrin reduces neutrophils and pro-inflammatory cytokines in mouse models of Pseudomonas aeruginosa lung infections. Biochem. Cell Biol. 2017, 95, 41–47. [Google Scholar] [CrossRef] [PubMed]
- Lang, J.; Yang, N.; Deng, J.; Liu, K.; Yang, P.; Zhang, G.; Jiang, C. Inhibition of SARS Pseudovirus Cell Entry by Lactoferrin Binding to Heparan Sulfate Proteoglycans. PLoS ONE 2011, 6, e23710. [Google Scholar] [CrossRef]
- Berlutti, F.; Pantanella, F.; Natalizi, T.; Frioni, A.; Paesano, R.; Polimeni, A.; Valenti, P. Antiviral Properties of Lactoferrin—A Natural Immunity Molecule. Molecules 2011, 16, 6992–7018. [Google Scholar] [CrossRef]
- Redwan, E.M.; Uversky, V.N.; El-Fakharany, E.M.; Al-Mehdar, H. Potential lactoferrin activity against pathogenic viruses. C. R. Biol. 2014, 337, 581–595. [Google Scholar] [CrossRef]
- Milanetti, E.; Miotto, M.; Di Rienzo, L.; Monti, M.; Gosti, G.; Ruocco, G. In-Silico evidence for two receptors based strategy of SARS-CoV-2. arXiv 2020, arXiv:2003.11107. [Google Scholar]
- Vandelli, A.; Monti, M.; Milanetti, E.; Ponti, R.D.; Tartaglia, G.G. Structural analysis of SARS-CoV-2 and prediction of the human interactome. arXiv 2020, arXiv:2003.13655. [Google Scholar] [CrossRef]
- Liu, L.; Chopra, P.; Li, X.; Wolfert, M.A.; Tompkins, S.M.; Boons, G.J. SARS-CoV-2 spike protein binds heparan sulfate in a length- and sequence-dependent manner. bioRxiv 2020. [Google Scholar] [CrossRef]
- Robson, B. The use of knowledge management tools in viroinformatics. Example study of a highly conserved sequence motif in Nsp3 of SARS-CoV-2 as a therapeutic target. Comput. Biol. Med. 2020, 125, 103963. [Google Scholar] [CrossRef]
- Miotto, M.; Di Rienzo, L.; Bò, L.; Boffi, A.; Ruocco, G.; Milanetti, E. Molecular mechanisms behind anti SARS-CoV-2 action of lactoferrin. arXiv 2020, arXiv:2007.07341. [Google Scholar]
- Campione, E.; Lanna, C.; Cosio, T.; Rosa, L.; Conte, M.P.; Iacovelli, F.; Romeo, A.; Falconi, M.; Del Vecchio, C.; Franchin, E. Pleiotropic effect of Lactoferrin in the prevention and treatment of COVID-19 infection: In vivo, in silico and in vitro preliminary evidences. bioRxiv 2020. [Google Scholar] [CrossRef]
- Campione, E.; Cosio, T.; Rosa, L.; Lanna, C.; Di Girolamo, S.; Gaziano, R.; Valenti, P.; Bianchi, L. Lactoferrin as Protective Natural Barrier of Respiratory and Intestinal Mucosa against Coronavirus Infection and Inflammation. Int. J. Mol. Sci. 2020, 21, 4903. [Google Scholar] [CrossRef]
- Wang, Y.; Wang, P.; Wang, H.; Luo, Y.; Wan, L.; Jiang, M.; Chu, Y. Lactoferrin for the treatment of COVID-19 (Review). Exp. Ther. Med. 2020, 20, 272. [Google Scholar] [CrossRef]
- Chimenti, F.; Cottiglia, F.; Bonsignore, L.; Casu, L.; Casu, M.; Floris, C.; Secci, D.; Bolasco, A.; Chimenti, P.; Granese, A.; et al. Quercetin as the active principle of Hypericum hircinum exerts a selective inhibitory activity against MAO-A: Extraction, Biological analysis, and computational study. J. Nat. Prod. 2006, 69, 945–949. [Google Scholar] [CrossRef]
- Wu, W.; Li, R.; Li, X.; He, J.; Jiang, S.; Liu, S.; Yang, J. Quercetin as an Antiviral Agent Inhibits Influenza A Virus (IAV) Entry. Viruses 2016, 8, 6. [Google Scholar] [CrossRef]
- Abian, O.; Ortega-Alarcon, D.; Jimenez-Alesanco, A.; Ceballos-Laita, L.; Vega, S.; Reyburn, H.T.; Rizzuti, B.; Velázquez-Campoy, A. Structural stability of SARS-CoV-2 3CLpro and identification of quercetin as an inhibitor by experimental screening. Int. J. Biol. Macromol. 2020, 164, 1693–1703. [Google Scholar] [CrossRef] [PubMed]
- DeRosa, G.; Maffioli, P.; D’Angelo, A.; Di Pierro, F. A role for quercetin in coronavirus disease 2019 (COVID-19). Phytother. Res. 2020. [Google Scholar] [CrossRef] [PubMed]
- Smith, M.; Smith, J.C. Repurposing therapeutics for COVID-19: Supercomputer-based docking to the SARS-CoV-2 viral spike protein and viral spike protein-human ACE2 interface. ChemRxiv 2020. [Google Scholar] [CrossRef]
- Liu, X.; Raghuvanshi, R.; Ceylan, F.D.; Bolling, B.W. Quercetin and Its Metabolites Inhibit Recombinant Human Angiotensin-Converting Enzyme 2 (ACE2) Activity. J. Agric. Food Chem. 2020, 68, 13982–13989. [Google Scholar] [CrossRef] [PubMed]
- Oz, M.; Lorke, D.E.; Kabbani, N. A comprehensive guide to the pharmacologic regulation of angiotensin converting enzyme 2 (ACE2), the SARS-CoV-2 entry receptor. Pharmacol. Ther. 2020, 107750. [Google Scholar] [CrossRef]
- Arslan, B.; Ergun, N.U.; Topuz, S.; Semerci, S.Y.; Suner, N.; Kocatas, A.; Onal, H. Synergistic Effect of Quercetin and Vitamin C Against COVID-19: Is a Possible Guard for Front Liners. SSRN Electron. J. 2020. [Google Scholar] [CrossRef]
- Ahmed, A.K.; Albalawi, Y.S.; Shora, H.A.; Abdelseed, H.K.; Al-Kattan, A.N. Effects of quadruple therapy: Zinc, quercetin, bromelain and vitamin C on the clinical outcomes of patients infected with COVID-19. Res. Int. J. Endocrinol. Diabetes 2020, 1, 18–21. [Google Scholar]
- Onal, H.; Semerci, S. Effect of Quercetin on Prophylaxis and Treatment of COVID-19. 2020. NCT04377789. Available online: https://clinicaltrials.gov/ct2/home (accessed on 28 December 2020).
- Weiskirchen, S.; Weiskirchen, R. Resveratrol: How Much Wine Do You Have to Drink to Stay Healthy? Adv. Nutr. 2016, 7, 706–718. [Google Scholar] [CrossRef]
- Wahedi, H.M.; Ahmad, S.; Abbasi, S.W. Stilbene-based natural compounds as promising drug candidates against COVID-19. J. Biomol. Struct. Dyn. 2020, 1–10. [Google Scholar] [CrossRef]
- ter Ellen, B.; Kumar, N.D.; Bouma, E.; Troost, B.; van de Pol, D.; van der Ende-Metselaar, H.; Apperloo, L.; van Gosliga, D.; van den Berge, M.; Nawijn, M. Resveratrol and pterostilbene potently inhibit SARS-CoV-2 infection in vitro. bioRxiv 2020. [Google Scholar] [CrossRef]
- King, V.F.; Garcia, M.L.; Himmel, D.; Reuben, J.P.; Lam, Y.K.; Pan, J.X.; Han, G.Q.; Kaczorowski, G.J. Interaction of tetrandrine with slowly inactivating calcium channels. Characterization of calcium channel modulation by an alkaloid of Chinese medicinal herb origin. J. Biol. Chem. 1988, 263, 2238–2244. [Google Scholar] [CrossRef]
- Heister, P.M.S.; Poston, R.N. Pharmacological hypothesis: TPC2 antagonist tetrandrine as a potential therapeutic agent for COVID-19. Pharmacol. Res. Perspect. 2020, 8, e00653. [Google Scholar] [CrossRef] [PubMed]
- Sun, Y.F.; Wink, M. Tetrandrine and fangchinoline, bisbenzylisoquinoline alkaloids from Stephania tetrandra can reverse multidrug resistance by inhibiting P-glycoprotein activity in multidrug resistant human cancer cells. Phytomedicine 2014, 21, 1110–1119. [Google Scholar] [CrossRef] [PubMed]
- Lan, J.; Huang, L.; Lou, H.; Chen, C.; Liu, T.; Hu, S.; Yao, Y.; Song, J.; Luo, J.; Liu, Y.; et al. Design and synthesis of novel C14-urea-tetrandrine derivatives with potent anti-cancer activity. Eur. J. Med. Chem. 2018, 143, 1968–1980. [Google Scholar] [CrossRef] [PubMed]
- Leeson, P.D.; Springthorpe, B. The influence of drug-like concepts on decision-making in medicinal chemistry. Nat. Rev. Drug Discov. 2007, 6, 881–890. [Google Scholar] [CrossRef] [PubMed]
- Garg, S.; Roy, A. In silico analysis of selected alkaloids against main protease (Mpro) of SARS-CoV-2. Chem. Biol. Interact. 2020, 332, 109309. [Google Scholar] [CrossRef]
- Riva, L.; Yuan, S.; Yin, X.; Martin-Sancho, L.; Matsunaga, N.; Pache, L.; Burgstaller-Muehlbacher, S.; De Jesus, P.D.; Teriete, P.; Hull, M.V.; et al. Discovery of SARS-CoV-2 antiviral drugs through large-scale compound repurposing. Nature 2020, 586, 113–119. [Google Scholar] [CrossRef]
- Muhseen, Z.T.; Hameed, A.R.; Al-Hasani, H.M.; Qamar, M.T.U.; Li, G. Promising terpenes as SARS-CoV-2 spike receptor-binding domain (RBD) attachment inhibitors to the human ACE2 receptor: Integrated computational approach. J. Mol. Liq. 2020, 320, 114493. [Google Scholar] [CrossRef]
- Luo, P.; Liu, D.; Li, J. Pharmacological perspective: Glycyrrhizin may be an efficacious therapeutic agent for COVID-19. Int. J. Antimicrob. Agents 2020, 55, 105995. [Google Scholar] [CrossRef]
- D’Alessandro, S.; Scaccabarozzi, D.; Signorini, L.; Perego, F.; Ilboudo, D.P.; Ferrante, P.; Delbue, S. The Use of Antimalarial Drugs against Viral Infection. Microorganisms 2020, 8, 85. [Google Scholar] [CrossRef]
- Reiter, C.; Fröhlich, T.; Gruber, L.; Hutterer, C.; Marschall, M.; Voigtländer, C.; Friedrich, O.; Kappes, B.; Efferth, T.; Tsogoeva, S.B. Highly potent artemisinin-derived dimers and trimers: Synthesis and evaluation of their antimalarial, antileukemia and antiviral activities. Bioorg. Med. Chem. 2015, 23, 5452–5458. [Google Scholar] [CrossRef] [PubMed]
- Sehailia, M.; Chemat, S. Antimalarial-agent artemisinin and derivatives portray more potent binding to Lys353 and Lys31-binding hotspots of SARS-CoV-2 spike protein than hydroxychloroquine: Potential repurposing of artenimol for COVID-19. J. Biomol. Struct. Dyn. 2020, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Cao, R.; Hu, H.; Li, Y.; Wang, X.; Xu, M.; Liu, J.; Zhang, H.; Yan, Y.; Zhao, L.; Li, W.; et al. Anti-SARS-CoV-2 Potential of Artemisinins In Vitro. ACS Infect. Dis. 2020, 6, 2524–2531. [Google Scholar] [CrossRef] [PubMed]
- Slobodnick, A.; Shah, B.; Pillinger, M.H.; Krasnokutsky, S. Colchicine: Old and new. Am. J. Med. 2015, 128, 461–470. [Google Scholar] [CrossRef] [PubMed]
- Ito, M.; Yanagi, Y.; Ichinohe, T. Encephalomyocarditis Virus Viroporin 2B Activates NLRP3 Inflammasome. PLOS Pathog. 2012, 8, e1002857. [Google Scholar] [CrossRef]
- Leung, Y.Y.; Hui, L.L.Y.; Kraus, V.B. Colchicine—Update on mechanisms of action and therapeutic uses. Semin. Arthritis Rheum. 2015, 45, 341–350. [Google Scholar] [CrossRef]
- Sohag, A.A.M.; Hannan, A.; Rahman, S.; Hossain, M.; Hasan, M.; Khan, K.; Khatun, A.; Dash, R.; Uddin, J. Revisiting potential druggable targets against SARS-CoV-2 and repurposing therapeutics under preclinical study and clinical trials: A comprehensive review. Drug Dev. Res. 2020, 81, 919–941. [Google Scholar] [CrossRef]
- Scarsi, M.; Piantoni, S.; Colombo, E.; Airó, P.; Richini, D.; Miclini, M.; Bertasi, V.; Bianchi, M.; Bottone, D.; Civelli, P.; et al. Association between treatment with colchicine and improved survival in a single-centre cohort of adult hospitalised patients with COVID-19 pneumonia and acute respiratory distress syndrome. Ann. Rheum. Dis. 2020, 79, 1286–1289. [Google Scholar] [CrossRef]
- Gendelman, O.; Amital, H.; Bragazzi, N.L.; Watad, A.; Chodick, G. Continuous hydroxychloroquine or colchicine therapy does not prevent infection with SARS-CoV-2: Insights from a large healthcare database analysis. Autoimmun. Rev. 2020, 19, 102566. [Google Scholar] [CrossRef]
- Rocca, R.; Moraca, F.; Costa, G.; Alcaro, S.; Distinto, S.; Maccioni, E.; Ortuso, F.; Artese, A.; Parrotta, L. Structure-Based Virtual Screening of Novel Natural Alkaloid Derivatives as Potential Binders of h-telo and c-myc DNA G-Quadruplex Conformations. Molecules 2014, 20, 206–223. [Google Scholar] [CrossRef]
- Moraca, F.; Amato, J.; Ortuso, F.; Artese, A.; Pagano, B.; Novellino, E.; Alcaro, S.; Parrinello, M.; Limongelli, V. Ligand binding to telomeric G-quadruplex DNA investigated by funnel-metadynamics simulations. Proc. Natl. Acad. Sci. USA 2017, 114, E2136–E2145. [Google Scholar] [CrossRef] [PubMed]
- Kaserer, T.; Rigo, R.; Schuster, P.; Alcaro, S.; Sissi, C.; Schuster, D. Optimized Virtual Screening Workflow for the Identification of Novel G-Quadruplex Ligands. J. Chem. Inf. Model. 2016, 56, 484–500. [Google Scholar] [CrossRef] [PubMed]
- Varghese, F.S.; Kaukinen, P.; Gläsker, S.; Bespalov, M.; Hanski, L.; Wennerberg, K.; Kümmerer, B.M.; Ahola, T. Discovery of berberine, abamectin and ivermectin as antivirals against chikungunya and other alphaviruses. Antivir. Res. 2016, 126, 117–124. [Google Scholar] [CrossRef] [PubMed]
- Pizzorno, A.; Padey, B.; Dubois, J.; Julien, T.; Traversier, A.; Dulière, V.; Brun, P.; Lina, B.; Rosa-Calatrava, M.; Terrier, O. In vitro evaluation of antiviral activity of single and combined repurposable drugs against SARS-CoV-2. Antivir. Res. 2020, 181, 104878. [Google Scholar] [CrossRef] [PubMed]
- Zhang, B.; Chen, M.; Chen, X.; Cao, K.; You, Y.; Qian, Y.; Yu, W. Berberine reduces circulating inflammatory mediators in patients with severe COVID-19. Br. J. Surg. 2020. [Google Scholar] [CrossRef]
- Varghese, F.S.; van Woudenbergh, E.; Overheul, G.J.; Eleveld, M.J.; Kurver, L.; van Heerbeek, N.; van Laarhoven, A.; Miesen, P.; den Hartog, G.; de Jonge, M.I. Berberine and obatoclax inhibit SARS-CoV-2 replication in primary human nasal epithelial cells in vitro. bioRxiv 2020. [Google Scholar] [CrossRef]
- Krupanidhi, S.; Abraham Peele, K.; Venkateswarulu, T.; Ayyagari, V.S.; Nazneen Bobby, M.; John Babu, D.; Venkata Narayana, A.; Aishwarya, G. Screening of phytochemical compounds of Tinospora cordifolia for their inhibitory activity on SARS-CoV-2: An in silico study. J. Biomol. Struct. Dyn. 2020, 1–5. [Google Scholar] [CrossRef]
- Debnath, B.; Singh, W.S.; Das, M.; Goswami, S.; Singh, M.K.; Maiti, D.; Manna, K. Role of plant alkaloids on human health: A review of biological activities. Mater. Today Chem. 2018, 9, 56–72. [Google Scholar] [CrossRef]
- Borquaye, L.S.; Gasu, E.N.; Ampomah, G.B.; Kyei, L.K.; Amarh, M.A.; Mensah, C.N.; Nartey, D.; Commodore, M.; Adomako, A.K.; Acheampong, P.; et al. Alkaloids from Cryptolepis sanguinolenta as Potential Inhibitors of SARS-CoV-2 Viral Proteins: An In Silico Study. BioMed Res. Int. 2020, 2020, 5324560. [Google Scholar] [CrossRef]
- Gyebi, G.A.; Adegunloye, A.P.; Ibrahim, I.M.; Ogunyemi, O.M.; Afolabi, S.O.; Ogunro, O.B. Prevention of SARS-CoV-2 cell entry: Insight from in silico interaction of drug-like alkaloids with spike glycoprotein, human ACE2, and TMPRSS2. J. Biomol. Struct. Dyn. 2020, 1–25. [Google Scholar] [CrossRef]
- Alfaro, M.; Alfaro, I.; Angel, C. Identification of potential inhibitors of SARS-CoV-2 papain-like protease from tropane alkaloids from Schizanthus porrigens: A molecular docking study. Chem. Phys. Lett. 2020, 761, 138068. [Google Scholar] [CrossRef] [PubMed]
- Ghosh, R.; Chakraborty, A.; Biswas, A.; Chowdhuri, S. Identification of alkaloids from Justicia adhatoda as potent SARS CoV-2 main protease inhibitors: An in silico perspective. J. Mol. Struct. 2020, 129489. [Google Scholar] [CrossRef] [PubMed]
- Quimque, M.T.J.; Notarte, K.I.R.; Fernandez, R.A.T.; Mendoza, M.A.O.; Liman, R.A.D.; Lim, J.A.K.; Pilapil, L.A.E.; Ong, J.K.H.; Pastrana, A.M.; Khan, A.; et al. Virtual screening-driven drug discovery of SARS-CoV2 enzyme inhibitors targeting viral attachment, replication, post-translational modification and host immunity evasion infection mechanisms. J. Biomol. Struct. Dyn. 2020, 1–18. [Google Scholar] [CrossRef] [PubMed]
- Diniz, L.R.L.; Pérez-Castillo, Y.; Elshabrawy, H.A.; Filho, C.D.S.M.B.; De Sousa, D.P. Bioactive Terpenes and Their Derivatives as Potential SARS-CoV-2 Proteases Inhibitors from Molecular Modeling Studies. Biomolecules 2021, 11, 74. [Google Scholar] [CrossRef] [PubMed]
- Sepay, N.; Sekar, A.; Halder, U.C.; Alarifi, A.; Afzal, M. Anti-COVID-19 terpenoid from marine sources: A docking, admet and molecular dynamics study. J. Mol. Struct. 2021, 1228, 129433. [Google Scholar] [CrossRef] [PubMed]
- Kiyama, R. Estrogenic terpenes and terpenoids: Pathways, functions and applications. Eur. J. Pharmacol. 2017, 815, 405–415. [Google Scholar] [CrossRef]
- Gershenzon, J.; Dudareva, N. The function of terpene natural products in the natural world. Nat. Chem. Biol. 2007, 3, 408–414. [Google Scholar] [CrossRef]
- Kumar, K.J.S.; Vani, M.G.; Wang, C.-S.; Chen, C.-C.; Chen, Y.-C.; Lu, L.-P.; Huang, C.-H.; Lai, C.-S.; Wang, S.-Y. Geranium and Lemon Essential Oils and Their Active Compounds Downregulate Angiotensin-Converting Enzyme 2 (ACE2), a SARS-CoV-2 Spike Receptor-Binding Domain, in Epithelial Cells. Plants 2020, 9, 770. [Google Scholar] [CrossRef]
- Kulkarni, S.A.; Nagarajan, S.K.; Ramesh, V.; Palaniyandi, V.; Sellamuthu, P.S.; Madhavan, T. Computational evaluation of major components from plant essential oils as potent inhibitors of SARS-CoV-2 spike protein. J. Mol. Struct. 2020, 1221, 128823. [Google Scholar] [CrossRef]
- Vardhan, S.; Sahoo, S.K. Searching inhibitors for three important proteins of COVID-19 through molecular docking studies. arXiv 2020, arXiv:2004.08095. [Google Scholar]
- Carino, A.; Moraca, F.; Fiorillo, B.; Marchianò, S.; Sepe, V.; Biagioli, M.; Finamore, C.; Bozza, S.; Francisci, D.; Distrutti, E.; et al. Hijacking SARS-CoV-2/ACE2 Receptor Interaction by Natural and Semi-synthetic Steroidal Agents Acting on Functional Pockets on the Receptor Binding Domain. Front. Chem. 2020, 8, 572885. [Google Scholar] [CrossRef] [PubMed]
- Singh, S.; Florez, H. Bioinformatic study to discover natural molecules with activity against COVID-19. F1000Research 2020, 9, 1203. [Google Scholar] [CrossRef] [PubMed]
- Kumar, A.; Choudhir, G.; Shukla, S.K.; Sharma, M.; Tyagi, P.; Bhushan, A.; Rathore, M. Identification of phytochemical inhibitors against main protease of COVID-19 using molecular modeling approaches. J. Biomol. Struct. Dyn. 2020, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Poochi, S.P.; Easwaran, M.; Balasubramanian, B.; Anbuselvam, M.; Meyyazhagan, A.; Park, S.; Bhotla, H.K.; Anbuselvam, J.; Arumugam, V.A.; Keshavarao, S.; et al. Employing bioactive compounds derived from Ipomoea obscura (L.) to evaluate potential inhibitor for SARS-CoV-2 main protease and ACE2 protein. Food Front. 2020, 1, 168–179. [Google Scholar] [CrossRef] [PubMed]
- Tripathi, M.K.; Singh, P.; Sharma, S.; Singh, T.P.; Ethayathulla, A.S.; Kaur, P. Identification of bioactive molecule from Withania somnifera (Ashwagandha) as SARS-CoV-2 main protease inhibitor. J. Biomol. Struct. Dyn. 2020, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Chikhale, R.V.; Gurav, S.S.; Patil, R.B.; Sinha, S.K.; Prasad, S.; Shakya, A.; Shrivastava, S.K.; Gurav, N.; Prasad, R.S. Sars-cov-2 host entry and replication inhibitors from Indian ginseng: An in-silico approach. J. Biomol. Struct. Dyn. 2020, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Siddiqui, S.; Upadhyay, S.; Ahmad, R.; Gupta, A.; Srivastava, A.; Trivedi, A.; Husain, I.; Ahmad, B.; Ahamed, M.; Khan, M.A. Virtual screening of phytoconstituents from miracle herb nigella sativa targeting nucleocapsid protein and papain-like protease of SARS-CoV-2 for COVID-19 treatment. J. Biomol. Struct. Dyn. 2020, 1–21. [Google Scholar] [CrossRef]
- Gidaro, M.C.; Astorino, C.; Petzer, A.; Carradori, S.; Alcaro, F.; Costa, G.; Artese, A.; Rafele, G.; Russo, F.M.; Petzer, J.P.; et al. Kaempferol as Selective Human MAO-A Inhibitor: Analytical Detection in Calabrian Red Wines, Biological and Molecular Modeling Studies. J. Agric. Food Chem. 2016, 64, 1394–1400. [Google Scholar] [CrossRef]
- Carradori, S.; Gidaro, M.C.; Petzer, A.; Costa, G.; Guglielmi, P.; Chimenti, P.; Alcaro, S.; Petzer, J.P. Inhibition of Human Monoamine Oxidase: Biological and Molecular Modeling Studies on Selected Natural Flavonoids. J. Agric. Food Chem. 2016, 64, 9004–9011. [Google Scholar] [CrossRef]
- Qamar, M.T.U.; Alqahtani, S.M.; Alamri, M.A.; Chen, L.-L. Structural basis of SARS-CoV-2 3CLpro and anti-COVID-19 drug discovery from medicinal plants. J. Pharm. Anal. 2020, 10, 313–319. [Google Scholar] [CrossRef]
- Sen, D.; Debnath, P.; Debnath, B.; Bhaumik, S.; Debnath, S. Identification of potential inhibitors of SARS-CoV-2 main protease and spike receptor from 10 important spices through structure-based virtual screening and molecular dynamic study. J. Biomol. Struct. Dyn. 2020, 1–22. [Google Scholar] [CrossRef] [PubMed]
- Cherrak, S.A.; Merzouk, H.; Mokhtari-Soulimane, N. Potential bioactive glycosylated flavonoids as SARS-CoV-2 main protease inhibitors: A molecular docking and simulation studies. PLoS ONE 2020, 15, e0240653. [Google Scholar] [CrossRef] [PubMed]
- Park, J.-Y.; Yuk, H.J.; Ryu, H.W.; Lim, S.H.; Kim, K.S.; Park, K.H.; Ryu, Y.B.; Lee, W.S. Evaluation of polyphenols from Broussonetia papyrifera as coronavirus protease inhibitors. J. Enzym. Inhib. Med. Chem. 2017, 32, 504–512. [Google Scholar] [CrossRef] [PubMed]
- Jo, S.; Kim, S.; Kim, D.Y.; Kim, M.-S.; Shin, D.H. Flavonoids with inhibitory activity against SARS-CoV-2 3CLpro. J. Enzym. Inhib. Med. Chem. 2020, 35, 1539–1544. [Google Scholar] [CrossRef]
- Chen, L.; Li, J.; Luo, C.; Liu, H.; Xu, W.; Chen, G.; Liew, O.W.; Zhu, W.; Puah, C.; Shen, X.; et al. Binding interaction of quercetin-3-β-galactoside and its synthetic derivatives with SARS-CoV 3CLpro: Structure-activity relationship studies reveal salient pharmacophore features. Bioorg. Med. Chem. 2006, 14, 8295–8306. [Google Scholar] [CrossRef]
- Sahlan, M.; Irdiani, R.; Flamandita, D.; Aditama, R.; Alfarraj, S.; Ansari, M.J.; Khayrani, A.C.; Pratami, D.K.; Lischer, K. Molecular interaction analysis of Sulawesi propolis compounds with SARS-CoV-2 main protease as preliminary study for COVID-19 drug discovery. J. King Saud Univ. Sci. 2021, 33, 101234. [Google Scholar] [CrossRef]
- Muchtaridi, M.; Fauzi, M.; Ikram, N.K.K.; Gazzali, A.M.; Wahab, H.A. Natural Flavonoids as Potential Angiotensin-Converting Enzyme 2 Inhibitors for Anti-SARS-CoV-2. Molecules 2020, 25, 3980. [Google Scholar] [CrossRef]
- Güler, H.I.; Tatar, G.; Yildiz, O.; Belduz, A.O.; Kolayli, S. An investigation of ethanolic propolis extracts: Their potential inhibitor properties against ACE-II receptors for COVID-19 treatment by Molecular Docking Study. ScienceOpen Prepr. 2020. [Google Scholar] [CrossRef]
- Brown, A.S.; Ackerley, D.F.; Calcott, M.J. High-Throughput Screening for Inhibitors of the SARS-CoV-2 Protease Using a FRET-Biosensor. Molecules 2020, 25, 4666. [Google Scholar] [CrossRef]
- Seadawy, M.G.; Gad, A.F.; Shamel, M.; Elharty, B.; Mohamed, M.F.; Elfiky, A.A.; Ahmed, A.; Zekri, A.R.N. In vitro: Natural Compounds (Thymol, Carvacrol, Hesperidine, and Thymoquinone) against SARS-CoV2 Strain Isolated From Egyptian Patients. bioRxiv 2020. [Google Scholar] [CrossRef]
- Thomson, R. Naturally Occurring Quinones; Academic Press: London, UK, 2012. [Google Scholar]
- Shuveksh, K.A.P.S.; Ahmed, K.; Padhye, S.; Schobert, R.; Biersack, B. Chemical and Biological Aspects of the Natural 1,4-Benzoquinone Embelin and its (semi-)Synthetic Derivatives. Curr. Med. Chem. 2017, 24, 1998–2009. [Google Scholar] [CrossRef] [PubMed]
- Radhakrishnan, N.; Gnanamani, A. 2,5-dihydroxy-3-undecyl-1, 4-benzoquinone (Embelin)-A second solid gold of India-A Review. Int. J. Pharm. Pharm. Sci. 2014, 6, 23–30. [Google Scholar]
- Bittner, S. When quinones meet amino acids: Chemical, physical and biological consequences. Amino Acids 2006, 30, 205–224. [Google Scholar] [CrossRef] [PubMed]
- Caruso, F.; Rossi, M.; Pedersen, J.Z.; Incerpi, S. Computational studies reveal mechanism by which quinone derivatives can inhibit SARS-CoV-2. Study of embelin and two therapeutic compounds of interest, methyl prednisolone and dexamethasone. J. Infect. Public Health 2020, 13. [Google Scholar] [CrossRef] [PubMed]
- Basu, A.; Sarkar, A.; Maulik, U. Computational approach for the design of potential spike protein binding natural compounds in SARS-CoV2. Pharmacodynamics 2020. [Google Scholar] [CrossRef]
- Ho, T.-Y.; Wu, S.-L.; Chen, J.-C.; Li, C.-C.; Hsiang, C.-Y. Emodin blocks the SARS coronavirus spike protein and angiotensin-converting enzyme 2 interaction. Antivir. Res. 2007, 74, 92–101. [Google Scholar] [CrossRef]
- Thuy, B.T.P.; My, T.T.A.; Hai, N.T.T.; Hieu, L.T.; Hoa, T.T.; Loan, H.T.P.; Triet, N.T.; Van Anh, T.T.; Quy, P.T.; Van Tat, P.; et al. Investigation into SARS-CoV-2 Resistance of Compounds in Garlic Essential Oil. ACS Omega 2020, 5, 8312–8320. [Google Scholar] [CrossRef]
- My, T.T.A.; Loan, H.T.P.; Hai, N.T.T.; Hieu, L.T.; Hoa, T.T.; Thuy, B.T.P.; Quang, D.T.; Triet, N.T.; Van Anh, T.T.; Dieu, N.T.X.; et al. Evaluation of the Inhibitory Activities of COVID-19 of Melaleuca cajuputi Oil Using Docking Simulation. ChemistrySelect 2020, 5, 6312–6320. [Google Scholar] [CrossRef]
- Shaldam, M.A.; Yahya, G.; Mohamed, N.H.; Abdel-Daim, M.M.; Al Naggar, Y. In silico screening of potent bioactive compounds from honey bee products against COVID-19 target enzymes. ChemRxiv Prepr. 2020. [Google Scholar] [CrossRef]
- Poduri, R.; Joshi, G.; Jagadeesh, G. Drugs targeting various stages of the SARS-CoV-2 life cycle: Exploring promising drugs for the treatment of Covid-19. Cell. Signal. 2020, 74, 109721. [Google Scholar] [CrossRef]
- Peele, K.A.; Chandrasai, P.; Srihansa, T.; Krupanidhi, S.; Sai, A.V.; Babu, D.J.; Indira, M.; Reddy, A.R.; Venkateswarulu, T. Molecular docking and dynamic simulations for antiviral compounds against SARS-CoV-2: A computational study. Inform. Med. Unlocked 2020, 19, 100345. [Google Scholar] [CrossRef] [PubMed]
- Verma, S.; Twilley, D.; Esmear, T.; Oosthuizen, C.B.; Reid, A.-M.; Nel, M.; Lall, N. Anti-SARS-CoV Natural Products With the Potential to Inhibit SARS-CoV-2 (COVID-19). Front. Pharmacol. 2020, 11, 561334. [Google Scholar] [CrossRef] [PubMed]
- Gyebi, G.A.; Ogunro, O.B.; Adegunloye, A.P.; Ogunyemi, O.M.; Afolabi, S.O. Potential inhibitors of coronavirus 3-chymotrypsin-like protease (3CLpro): An in silico screening of alkaloids and terpenoids from African medicinal plants. J. Biomol. Struct. Dyn. 2020, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Yang, H.; Bartlam, M.; Rao, Z. Drug Design Targeting the Main Protease, the Achilles Heel of Coronaviruses. Curr. Pharm. Des. 2006, 12, 4573–4590. [Google Scholar] [CrossRef] [PubMed]
- da Silva Antonio, A.; Wiedemann, L.S.M.; Veiga-Junior, V.F. Natural products’ role against COVID-19. RSC Adv. 2020, 10, 23379–23393. [Google Scholar] [CrossRef]
- Gendrot, M.; Andreani, J.; Boxberger, M.; Jardot, P.; Fonta, I.; Le Bideau, M.; Duflot, I.; Mosnier, J.; Rolland, C.; Bogreau, H. Antimalarial drugs inhibit the replication of SARS-CoV-2: An in vitro evaluation. Travel Med. Infect. Dis. 2020, 37, 101873. [Google Scholar] [CrossRef]
- Jiang, Y.; Liu, M.; Liu, H.; Liu, S. A critical review: Traditional uses, phytochemistry, pharmacology and toxicology of Stephania tetrandra S. Moore (Fen Fang Ji). Phytochem. Rev. 2020, 19, 449–489. [Google Scholar] [CrossRef]
- Xu, Y.; Baylink, D.J.; Chen, C.-S.; Reeves, M.E.; Xiao, J.; Lacy, C.; Lau, E.; Cao, H. The importance of vitamin d metabolism as a potential prophylactic, immunoregulatory and neuroprotective treatment for COVID-19. J. Transl. Med. 2020, 18, 322. [Google Scholar] [CrossRef]
- Ginde, A.A.; Blatchford, P.; Breese, K.; Zarrabi, L.; Linnebur, S.A.; Wallace, J.I.; Schwartz, R.S. High-Dose Monthly Vitamin D for Prevention of Acute Respiratory Infection in Older Long-Term Care Residents: A Randomized Clinical Trial. J. Am. Geriatr. Soc. 2017, 65, 496–503. [Google Scholar] [CrossRef]
- Wang, R.; DeGruttola, V.; Lei, Q.; Mayer, K.H.; Redline, S.; Hazra, A.; Mora, S.; Willett, W.C.; Ganmaa, D.; Manson, J.E. The vitamin D for COVID-19 (VIVID) trial: A pragmatic cluster-randomized design. Contemp. Clin. Trials 2021, 100, 106176. [Google Scholar] [CrossRef]
- Gautret, P.; Lagier, J.C.; Parola, P.; Hoang, V.T.; Meddeb, L.; Mailhe, M.; Doudier, B.; Courjon, J.; Giordanengo, V.; Vieira, V.E.; et al. Hydroxychloroquine and azithromycin as a treatment of COVID-19: Results of an open-label non-randomized clinical trial. Int. J. Antimicrob. Agents 2020, 56, 105949. [Google Scholar] [CrossRef] [PubMed]
- Molina, J.-M.; Delaugerre, C.; Le Goff, J.; Mela-Lima, B.; Ponscarme, D.; Goldwirt, L.; De Castro, N. No evidence of rapid antiviral clearance or clinical benefit with the combination of hydroxychloroquine and azithromycin in patients with severe COVID-19 infection. Med. Mal. Infect. 2020, 50, 384. [Google Scholar] [CrossRef] [PubMed]
- Magagnoli, J.; Narendran, S.; Pereira, F.; Cummings, T.H.; Hardin, J.W.; Sutton, S.S.; Ambati, J. Outcomes of Hydroxychloroquine Usage in United States Veterans Hospitalized with COVID-19. Med 2020, 1, 114.e3–127.e3. [Google Scholar] [CrossRef] [PubMed]
- Million, M.; Gautret, P.; Colson, P.; Roussel, Y.; Dubourg, G.; Chabriere, E.; Honore, S.; Rolain, J.-M.; Fenollar, F.; Fournier, P.-E.; et al. Clinical efficacy of chloroquine derivatives in COVID-19 infection: Comparative meta-analysis between the big data and the real world. New Microbes New Infect. 2020, 38, 100709. [Google Scholar] [CrossRef] [PubMed]
- Tang, W.; Cao, Z.; Han, M.; Wang, Z.; Chen, J.; Sun, W.; Wu, Y.; Xiao, W.; Liu, S.; Chen, E.; et al. Hydroxychloroquine in patients with mainly mild to moderate coronavirus disease 2019: Open label, randomised controlled trial. BMJ 2020, 369, m1849. [Google Scholar] [CrossRef] [PubMed]
- Gao, J.; Tian, Z.; Yang, X. Breakthrough: Chloroquine phosphate has shown apparent efficacy in treatment of COVID-19 associated pneumonia in clinical studies. Biosci. Trends 2020, 14, 72–73. [Google Scholar] [CrossRef] [PubMed]
- Mahevas, M.; Tran, V.-T.; Roumier, M.; Chabrol, A.; Paule, R.; Guillaud, C.; Gallien, S.; Lepeule, R.; Szwebel, T.-A.; Lescure, X. No evidence of clinical efficacy of hydroxychloroquine in patients hospitalized for COVID-19 infection with oxygen requirement: Results of a study using routinely collected data to emulate a target trial. medRxiv 2020. [Google Scholar] [CrossRef]
- Chorin, E.; Dai, M.; Shulman, E.; Wadhwani, L.; Bar Cohen, R.; Barbhaiya, C.; Aizer, A.; Holmes, D.; Bernstein, S.; Soinelli, M. The QT Interval in Patients with SARS-CoV-2 Infection Treated with Hydroxychloroquine/Azithromycin. medRxiv 2020. [Google Scholar] [CrossRef]



| Compound Name | Natural Sources | In Silico | In Vitro | Clinical Trials | Ref |
|---|---|---|---|---|---|
| Alpha-spinasterol | Nigella sativa | • | [128] | ||
| Anethole | Fennel, anise, Star anise, Essential oils | • | [120] | ||
| Anisotine | Justicia adhatoda | • | [113] | ||
| Aronadendin | Onion | • | [132] | ||
| Arteannuin B | Artemisia annua | • | [94] | ||
| Artemisinin | Artemisia Annua | • | • | • | [93,94,158] |
| Artepillin C | Honey and propolis | • | [151] | ||
| Ashwagandhanolide | Withania somnifera | • | [127] | ||
| Baicalin | Scutellaria baicalensis | • | • | [135,138] | |
| Berberine | Berberis vulgaris L. | • | • | • | [101,102,104,105,106,107,108] |
| Beta-sitosterol | Nigella sativa | • | [128] | ||
| Betulinic Acid | Betula pubescens | • | [123] | ||
| Biscryptolepine | Cryptolepis sanguinolenta | • | [110] | ||
| Broussoflavonol F | Macaranga indica | • | [137] | ||
| Campesterol | Nigella sativa | • | [128] | ||
| Carvacrol | Origanum vulgare | • | • | [120] | |
| Chenodeoxycholic acid | Bile Acids | • | • | [122] | |
| Cinnamaldehyde | Cinnamomum sp. | • | [120] | ||
| Cinnamyl Acetate | Essential oils | • | [120] | ||
| Citronellol | Cymbopogon winterianus, Geranium | • | • | [119] | |
| Colchicine | Colchicum autumnale | • | [99,100] | ||
| Cryptomisrine | Cryptolepis sanguinolenta | • | [110] | ||
| Cryptoquindoline | Cryptolepis sanguinolenta | • | [110] | ||
| Cryptospirolepine | Cryptolepis sanguinolenta | • | [110] | ||
| Cycloeucalenol | Nigella sativa | • | [128] | ||
| Dihydrowithaferin A | Withania somnifera | • | [127] | ||
| Ellagic acid | Honey and propolis | • | [151] | ||
| Embelin | Embelia ribes plant | • | [146] | ||
| Emodin | Rheum officinale and Reynoutria multiflora | • | • | [147,148] | |
| Geraniol | Essential oils | • | • | [119,120] | |
| Glico-chenodeoxycholic acid | Bile Acids | • | [122] | ||
| Glyco-ursodeoxycholic acid | Bile Acids | • | • | [122] | |
| Glycyrrhetinic Acid | Glycyrrhiza glabra | • | [123] | ||
| Glycyrrhizin | Glycyrrhiza glabra | • | • | • | [89,90] |
| Hanfangchin A | Stephania tetrandra | • | • | • | [82,83,84,88,159] |
| Herbacetin | Rhodiola rosea | • | • | [135] | |
| Hesperetin | Citrus sp. | • | [140,141] | ||
| Kaempferol | Aloe vera | • | [134] | ||
| L-4-terpineol | Essential oils | • | [120] | ||
| Lactoferrin | Mammalian and Animal milk | • | • | [56,57,67] | |
| Leucopelargonidin | Onion | • | [111,132] | ||
| Limonene | Essential oils | • | • | [119] | |
| Limonin | Euodia rutaecarpa | • | [119,120,121] | ||
| Linalool | Essential oils | • | • | [119] | |
| Myricetin | Onion, Chilli | • | [132,139,140] | ||
| Naringenin | Citrus sp. | • | [138,139] | ||
| Neohesperidin | Citrus sp. | • | • | [138,139,141] | |
| Neryl Acetate | Citrus oils | • | • | [119] | |
| Nobiletin | Citrus sp. | • | [138,139] | ||
| Oleanolic Acid | Olea europaea L. | • | • | [122] | |
| Pectolinarin | Cirsium setidens | • | • | [135] | |
| Pulegone | Essential oils | • | [120] | ||
| Quercetin | Onion, Garlic, Peppermint, Fenugreek, Origan Capers, Onions, Elderberries, Kale, Okra, Apple Peels, Aronia Berries, Cranberries, Asparagus, Goji Berries | • | • | • | [10,71,132,139] |
| Quercetin-3-O-galactosyl-rhamnosyl-glucoside | Withania somnifera | • | [127] | ||
| Quercetin-3-β-galactoside | Artemisia capillaris | • | [136] | ||
| Quinadoline B | Fungi | • | [114] | ||
| Resveratrol | Red wine, Red grape juice, Peanuts, Fresh grapes, Pistachios, Peanut butter, Cocoa powder, Dark chocolate, Milk chocolate, Strawberries, Jackfruit skin, Blueberries, Bilberries, Red currants, Cranberries, Lingonberries, Mulberries | • | • | • | [80,81] |
| Rutin | Citrus sp. | • | [139] | ||
| Scedapin C | Fungi | • | [114] | ||
| Schizanthine Y | Schizanthus porrigens | • | [112] | ||
| Schizanthine Z | Schizanthus porrigens | • | [112] | ||
| Scopadulcic Acid B | Scoparia dulcis | • | |||
| Scutellarin | Scutellaria barbata | • | [139] | ||
| Sophaline | Sophora alopecuriodes | • | [86] | ||
| Thalimonine | Thalictrum simplex | • | [86] | ||
| Thymol | Essential oils | • | [120] | ||
| Tricetin | Ginger, Onion, flowers of pomegranate. | • | [132] | ||
| Ursodeoxycholic acid | Bile Acids | • | • | [122] | |
| Ursolic acid | Bile Acids | • | • | [122] | |
| Vitamin C | Citrus Fruit, Tropical Fruit, Peppers, Cruciferous Vegetables, Leafy Vegetables | • | [36,40,41,42,43,44,45,46] | ||
| Vitamin D | Human and Jay Jasmine plant | • | • | [32,33,34] | |
| Withanolide N | Withania somnifera | • | [127] | ||
| Withanoside V | Withania somnifera | • | [127] | ||
| Withanoside X | Withania somnifera | • | [127] | ||
| 5S-5-Hydroxy-1,7-bis(4-hydroxy-3-methoxyphenyl)-3-heptanone | Ginger | • | [132] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Romeo, I.; Mesiti, F.; Lupia, A.; Alcaro, S. Current Updates on Naturally Occurring Compounds Recognizing SARS-CoV-2 Druggable Targets. Molecules 2021, 26, 632. https://doi.org/10.3390/molecules26030632
Romeo I, Mesiti F, Lupia A, Alcaro S. Current Updates on Naturally Occurring Compounds Recognizing SARS-CoV-2 Druggable Targets. Molecules. 2021; 26(3):632. https://doi.org/10.3390/molecules26030632
Chicago/Turabian StyleRomeo, Isabella, Francesco Mesiti, Antonio Lupia, and Stefano Alcaro. 2021. "Current Updates on Naturally Occurring Compounds Recognizing SARS-CoV-2 Druggable Targets" Molecules 26, no. 3: 632. https://doi.org/10.3390/molecules26030632
APA StyleRomeo, I., Mesiti, F., Lupia, A., & Alcaro, S. (2021). Current Updates on Naturally Occurring Compounds Recognizing SARS-CoV-2 Druggable Targets. Molecules, 26(3), 632. https://doi.org/10.3390/molecules26030632








