Barnase-Barstar Pair: Contemporary Application in Cancer Research and Nanotechnology
Abstract
1. Introduction
2. Barnase as a Tool for Cancer Suicide Gene Therapy
3. The Use of Barnase Ribonuclease Activity in Cancer Therapy
4. The Use of Barnase-Barstar Interaction for Supramolecular Complexes Assembly
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Conflicts of Interest
References
- Hartley, R.W. Barnase and barstar: Two small proteins to fold and fit together. Trends Biochem. Sci. 1989, 14, 450–454. [Google Scholar] [CrossRef]
- Nishimura, S.; Nomura, M. Ribonuclease of Bacillus subtilis. BBA-Biochim. Biophys. Acta 1958, 30, 430–431. [Google Scholar] [CrossRef]
- Smeaton, J.R.; Elliott, W.H. Isolation and properties of a specific bacterial ribonuclease inhibitor. BBA Sect. Nucleic Acids Protein Synth. 1967, 145, 547–560. [Google Scholar] [CrossRef]
- Hartley, R.W.; Rogerson, D.L. Production and Purification of the Extracellular Ribonuclease of Bacillus Amyloliquefaciens (Barnase) and Its Intracellular Inhibitor (Barstar) I. Barnase. Prep. Biochem. 1972, 2, 229–242. [Google Scholar] [CrossRef] [PubMed]
- Hartley, R.W.; Rogerson, D.L.; Smeaton, J.R. Production and Purification of the Extracellular Ribonuclease of Bacillus Amyloliquefaciens (Barnase) and Its Intracellular Inhibitor (Barstar) II. Barstar. Prep. Biochem. 1972, 2, 243–250. [Google Scholar] [CrossRef]
- Hartley, R.W.; Barker, E.A. Amino-acid sequence of extracellular ribonuclease (barnase) of bacillus amyloliquefaciens. Nat. New Biol. 1972, 235, 15–16. [Google Scholar] [CrossRef]
- Hartley, R.T.; Smeaton, J.R. On the Reaction between the Extracellular Ribonuclease of Bacillus amyloliquefaciens (Barnase) and Its Intracellular Inhibitor (Barstar). J. Biol. Chem. 1973, 248, 5624–5626. [Google Scholar] [CrossRef]
- Hartley, R.W. Barnase-barstar interaction. In Methods in Enzymology; Academic Press Inc.: New York, NY, USA, 2001; Volume 341, pp. 599–611. [Google Scholar]
- Deyev, S.M.; Waibel, R.; Lebedenko, E.N.; Schubiger, A.P.; Plückthun, A. Design of multivalent complexes using the barnase·barstar module. Nat. Biotechnol. 2003, 21, 1486–1492. [Google Scholar] [CrossRef]
- Korchuganov, D.S.; Nolde, S.B.; Reibarkh, M.Y.; Orekhov, V.Y.; Schulga, A.A.; Ermolyuk, Y.S.; Kirpichnikov, M.P.; Arseniev, A.S. NMR study of monomer-dimer equilibrium of barstar in solution. J. Am. Chem. Soc. 2001, 123, 2068–2069. [Google Scholar] [CrossRef]
- Paddon, C.J.; Hartley, R.W. Cloning, sequencing and transcription of an inactivated copy of Bacillus amyloliquefaciens extracellular ribonuclease (barnase). Gene 1985, 40, 231–239. [Google Scholar] [CrossRef]
- Paddon, C.J.; Hartley, R.W. Expression of Bacillus amyloliquefaciens extracellular ribonuclease (barnase) in Escherichia coli following an inactivating mutation. Gene 1987, 53, 11–19. [Google Scholar] [CrossRef]
- Hartley, R.W. Barnase and barstar. Expression of its cloned inhibitor permits expression of a cloned ribonuclease. J. Mol. Biol. 1988, 202, 913–915. [Google Scholar] [CrossRef]
- Kellis, J.T.; Nyberg, K.; Šail, D.; Fersht, A.R. Contribution of hydrophobic interactions to protein stability. Nature 1988, 333, 784–786. [Google Scholar] [CrossRef]
- Khurana, R.; Udgaonkar, J.B. Equilibrium Unfolding Studies of Barstar: Evidence for an Alternative Conformation Which Resembles a Molten Globule. Biochemistry 1994, 33, 106–115. [Google Scholar] [CrossRef] [PubMed]
- Mossakowska, D.E.; Nyberg, K.; Fersht, A.R. Kinetic Characterization of the Recombinant Ribonuclease from Bacillus amyloliquefaciens (Barnase) and Investigation of Key Residues in Catalysis by Site-Directed Mutagenesis. Biochemistry 1989, 28, 3843–3850. [Google Scholar] [CrossRef] [PubMed]
- Jones, D.N.M.; Bycroft, M.; Lubienski, M.J.; Fersht, A.R. Identification of the barstar binding site of barnase by NMR spectroscopy and hydrogen-deuterium exchange. FEBS Lett. 1993, 331, 165–172. [Google Scholar] [CrossRef]
- Schreiber, G.; Fersht, A.R. Interaction of Barnase with Its Polypeptide Inhibitor Barstar Studied by Protein Engineering. Biochemistry 1993, 32, 5145–5150. [Google Scholar] [CrossRef]
- Deyev, S.M.; Yazynin, S.A.; Kuznetsov, D.A.; Jukovich, M.; Hartley, R.W. Ribonuclease-charged vector for facile direct cloning with positive selection. Mol. Gen. Genet. 1998, 259, 379–382. [Google Scholar] [CrossRef]
- Yazynin, S.; Lange, H.; Mokros, T.; Deyev, S.; Lemke, H. A new phagemid vector for positive selection of recombinants based on a conditionally lethal barnase gene. FEBS Lett. 1999, 452, 351–354. [Google Scholar] [CrossRef]
- Mariani, C.; De Beuckeleer, M.; Truettner, J.; Leemans, J.; Goldberg, R.B. Induction of male sterility in plants by a chimaeric ribonuclease gene. Nature 1990, 347, 737–741. [Google Scholar] [CrossRef]
- Goldman, M.H.S.; Goldberg, R.B.; Mariani, C. Female sterile tobacco plants are produced by stigma-specific cell ablation. EMBO J. 1994, 13, 2976–2984. [Google Scholar] [CrossRef] [PubMed]
- Van Poucke, K.; Karimi, M.; Gheysen, G. Analysis of nematode-responsive promoters in sugar beet hairy roots. Meded. Rijksuniv. Gent. Fak. Landbouwkd Toegep. Biol. Wet 2001, 66, 591–598. [Google Scholar]
- Leuchtenberger, S.; Perz, A.; Gatz, C.; Bartsch, J.W. Conditional cell ablation by stringent tetracycline-dependent regulation of barnase in mammalian cells. Nucleic Acids Res. 2001, 29, 76. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Ilinskaya, O.N.; Shah Mahmud, R. Ribonucleases as antiviral agents. Mol. Biol. 2014, 48, 615–623. [Google Scholar] [CrossRef]
- Gribencha, S.V.; Potselueva, L.A.; Barinskiĭ, I.F.; Balandin, T.G.; Deev, S.M.; Leshchinskaia, I.B. The antiviral activity of RNAse Bacillus intermedius in experiments with mice preinfected with street rabies virus. Vopr. Virusol. 2004, 49, 38–41. [Google Scholar] [PubMed]
- Gribencha, S.V.; Potselueva, L.A.; Barinskiĭ, I.F.; Deev, S.M.; Balandin, T.G.; Leshchinskaia, I.B. Antiviral activity of Bacillus intermedius RNAase in guinea-pigs and rabbits infected with outdoor rabies virus. Vopr. Virusol. 2006, 51, 41–43. [Google Scholar]
- Glinka, E.M. Killing of cancer cells through the use of eukaryotic expression vectors harbouring genes encoding nucleases and ribonuclease inhibitor. Tumor Biol. 2015, 36, 3147–3157. [Google Scholar] [CrossRef] [PubMed]
- Edelweiss, E.; Balandin, T.G.; Ivanova, J.L.; Lutsenko, G.V.; Leonova, O.G.; Popenko, V.I.; Sapozhnikov, A.M.; Deyev, S.M. Barnase as a new therapeutic agent triggering apoptosis in human cancer cells. PLoS ONE 2008, 3, e2434. [Google Scholar] [CrossRef] [PubMed]
- Ulyanova, V.; Dudkina, E.; Nadyrova, A.; Kalashnikov, V.; Surchenko, Y.; Ilinskaya, O. The cytotoxicity of rnase-derived peptides. Biomolecules 2021, 11, 16. [Google Scholar] [CrossRef]
- Garipov, A.R.; Nesmelov, A.A.; Cabrera-Fuentes, H.A.; Ilinskaya, O.N. Bacillus intermedius ribonuclease (BINASE) induces apoptosis in human ovarian cancer cells. Toxicon. 2014, 92, 54–59. [Google Scholar] [CrossRef]
- Malhotra, V.; Perry, M.C. Classical Chemotherapy: Mechanisms, Toxicities and the Therapeutc Window. Cancer Biol. Ther. 2003, 2, S2–S4. [Google Scholar] [CrossRef] [PubMed]
- Gotte, G.; Menegazzi, M. Biological Activities of Secretory RNases: Focus on Their Oligomerization to Design Antitumor Drugs. Front. Immunol. 2019, 10, 2626. [Google Scholar] [CrossRef]
- Guillet, V.; Lapthorn, A.; Hartley, R.; Mauguen, Y. Recognition between a bacterial ribonuclease, barnase, and its natural inhibitor, barstar. Structure 1993, 1, 165–176. [Google Scholar] [CrossRef]
- Buckle, A.M.; Schreiber, G.; Fersht, A.R. Protein-Protein Recognition: Crystal Structural Analysis of a Barnase-Barstar Complex at 2.0-Å Resolution. Biochemistry 1994, 33, 8878–8889. [Google Scholar] [CrossRef] [PubMed]
- Sheikh, S.; Ernst, D.; Keating, A. Prodrugs and prodrug-activated systems in gene therapy. Mol. Ther. 2021, 5, 1716–1728. [Google Scholar] [CrossRef] [PubMed]
- Castro, J.; Ribó, M.; Navarro, S.; Nogués, M.V.; Vilanova, M.; Benito, A. A human ribonuclease induces apoptosis associated with p21WAF1/CIP1induction and JNK inactivation. BMC Cancer 2011, 11, 9. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Mccadden, J.; Ferrer, F.; Kruszewski, M.; Carducci, M.; Simons, J.; Rodriguez, R. Prostate-specific Expression of the Diphtheria Toxin A Chain (DT-A): Studies of Inducibility and Specificity of Expression of Prostate-specific Antigen Promoter-driven DT-A Adenoviral-mediated Gene Transfer. Cancer Res. 2002, 62, 2576–2582. [Google Scholar] [PubMed]
- Wang, C.Y.; Li, F.; Yang, Y.; Guo, H.Y.; Wu, C.X.; Wang, S. Recombinant baculovirus containing the Diphtheria toxin A gene for malignant glioma therapy. Cancer Res. 2006, 66, 5798–5806. [Google Scholar] [CrossRef]
- Chen, H. Exploiting the intron-splicing mechanism of insect cells to produce viral vectors harboring toxic genes for suicide gene therapy. Mol. Ther.-Nucleic Acids 2012, 1, e57. [Google Scholar] [CrossRef]
- Chen, H. Production of viral vectors with suicide genes by utilizing the intron-splicing mechanism of insect cells. In Methods in Molecular Biology; Humana Press Inc.: New York, NY, USA, 2019; Volume 1895, pp. 97–109. [Google Scholar] [CrossRef]
- Glinka, E.M.; Edelweiss, E.F.; Sapozhnikov, A.M.; Deyev, S.M. A new vector for controllable expression of an anti-HER2/neu mini-antibody-barnase fusion protein in HEK 293T cells. Gene 2006, 366, 97–103. [Google Scholar] [CrossRef]
- Agarwal, S.; Nikolai, B.; Yamaguchi, T.; Lech, P.; Somia, N.V. Construction and Use of Retroviral Vectors Encoding the Toxic Gene Barnase. Mol. Ther. 2006, 14, 555–563. [Google Scholar] [CrossRef] [PubMed]
- Voss, C.; Lindau, D.; Flaschel, E. Production of recombinant RNase Ba and its application in downstream processing of plasmid DNA for pharmaceutical use. Biotechnol. Prog. 2006, 22, 737–744. [Google Scholar] [CrossRef] [PubMed]
- Okorokov, A.L.; Hartley, R.W.; Panov, K.I. An improved system for ribonuclease Ba expression. Protein Expr. Purif. 1994, 5, 547–552. [Google Scholar] [CrossRef] [PubMed]
- Schulga, A.; Kurbanov, F.; Kirpichnikov, M.; Protasevich, V.; Lobachov, V.; Ranjbar, B.; Chekhov, V.; Polyakov, K.; Engelborghs, Y.; Makarov, A. Comparative study of binase and barnase: Experience in chimeric ribonucleases. Protein Eng. 1998, 11, 775–782. [Google Scholar] [CrossRef]
- Sommer, B.; Friehs, K.; Flaschel, E.; Reck, M.; Stahl, F.; Scheper, T. Extracellular production and affinity purification of recombinant proteins with Escherichia coli using the versatility of the maltose binding protein. J. Biotechnol. 2009, 140, 194–202. [Google Scholar] [CrossRef]
- Shilova, O.; Shramova, E.; Proshkina, G.; Deyev, S. Natural and designed toxins for precise therapy: Modern approaches in experimental oncology. Int. J. Mol. Sci. 2021, 22, 4975. [Google Scholar] [CrossRef]
- Prior, T.I.; FitzGerald, D.J.; Pastan, I. Barnase toxin: A new chimeric toxin composed of pseudomonas exotoxin A and barnase. Cell 1991, 64, 1017–1023. [Google Scholar] [CrossRef]
- Prior, T.I.; FitzGerald, D.J.; Pastan, I. Translocation Mediated by Domain II of Pseudomonas Exotoxin A: Transport of Barnase into the Cytosol. Biochemistry 1992, 31, 3555–3559. [Google Scholar] [CrossRef]
- Prior, T.I.; Kunwar, S.; Pastan, I. Studies on the activity of barnase toxins in vitro and in vivo. Bioconjug. Chem. 1996, 7, 23–29. [Google Scholar] [CrossRef]
- Martsev, S.P.; Tsybovsky, Y.I.; Stremovskiy, O.A.; Odintsov, S.G.; Balandin, T.G.; Arosio, P.; Kravchuk, Z.I.; Deyev, S.M. Fusion of the antiferritin antibody VL domain to barnase results in enhanced solubility and altered pH stability. Protein Eng. Des. Sel. 2004, 17, 85–93. [Google Scholar] [CrossRef]
- Stepanov, A.V.; Belogurov, A.A.; Ponomarenko, N.A.; Stremovskiy, O.A.; Kozlov, L.V.; Bichucher, A.M.; Dmitriev, S.E.; Smirnov, I.V.; Shamborant, O.G.; Balabashin, D.S.; et al. Design of targeted B cell killing agents. PLoS ONE 2011, 6, e20991. [Google Scholar] [CrossRef] [PubMed]
- Balandin, T.G.; Edelweiss, E.; Andronova, N.V.; Treshalina, E.M.; Sapozhnikov, A.M.; Deyev, S.M. Antitumor activity and toxicity of anti-HER2 immunoRNase scFv 4D5-dibarnase in mice bearing human breast cancer xenografts. Investig. New Drugs 2011, 29, 22–32. [Google Scholar] [CrossRef]
- Shipunova, V.O.; Shramova, E.I.; Shulga, A.A.; Shilova, M.V.; Deyev, S.M.; Proshkina, G.M.; Schulga, A.A.; Shilova, M.V.; Deyev, S.M.; Proshkina, G.M. Delivery of barnase to cells in liposomes functionalized by HER2-specific DARPin module. Russ. J. Biorgan. Chem. 2020, 46, 1156–1161. [Google Scholar] [CrossRef]
- Shramova, E.; Proshkina, G.; Shipunova, V.; Ryabova, A.; Kamyshinsky, R.; Konevega, A.; Schulga, A.; Konovalova, E.; Telegin, G.; Deyev, S. Dual targeting of cancer cells with darpin-based toxins for overcoming tumor escape. Cancers 2020, 12, 3014. [Google Scholar] [CrossRef] [PubMed]
- Shilova, O.N.; Deyev, S.M. DARPins: Promising scaffolds for theranostics. Acta Nat. 2019, 11, 42–53. [Google Scholar] [CrossRef] [PubMed]
- Altai, M.; Membreno, R.; Cook, B.; Tolmachev, V.; Zeglis, B.M. Pretargeted imaging and therapy. J. Nucl. Med. 2017, 58, 1553–1559. [Google Scholar] [CrossRef]
- Hapuarachchige, S.; Artemov, D. Theranostic Pretargeting Drug Delivery and Imaging Platforms in Cancer Precision Medicine. Front. Oncol. 2020, 10, 1131. [Google Scholar] [CrossRef]
- Stéen, E.J.L.; Edem, P.E.; Nørregaard, K.; Jørgensen, J.T.; Shalgunov, V.; Kjaer, A.; Herth, M.M. Pretargeting in nuclear imaging and radionuclide therapy: Improving efficacy of theranostics and nanomedicines. Biomaterials 2018, 179, 209–245. [Google Scholar] [CrossRef]
- Lebedenko, E.N.; Balandin, T.G.; Edelweiss, E.F.; Georgiev, O.; Moiseeva, E.S.; Petrov, R.V.; Deyev, S.M. Visualization of cancer cells by means of the fluorescent EGFP-barnase protein. Dokl. Biochem. Biophys. 2007, 414, 120–123. [Google Scholar] [CrossRef]
- Shilova, M.; Shramova, E.I.; Ryabova, A.; Proshkina, G.; Deyev, S. Barnase-Barstar molecular pair in a system for the stepwise delivery of cytotoxins to HER2 positive tumors. In FEBS Open Bio; Wiley: Hoboken, NJ, USA, 2021; Volume 11, p. 405. [Google Scholar] [CrossRef]
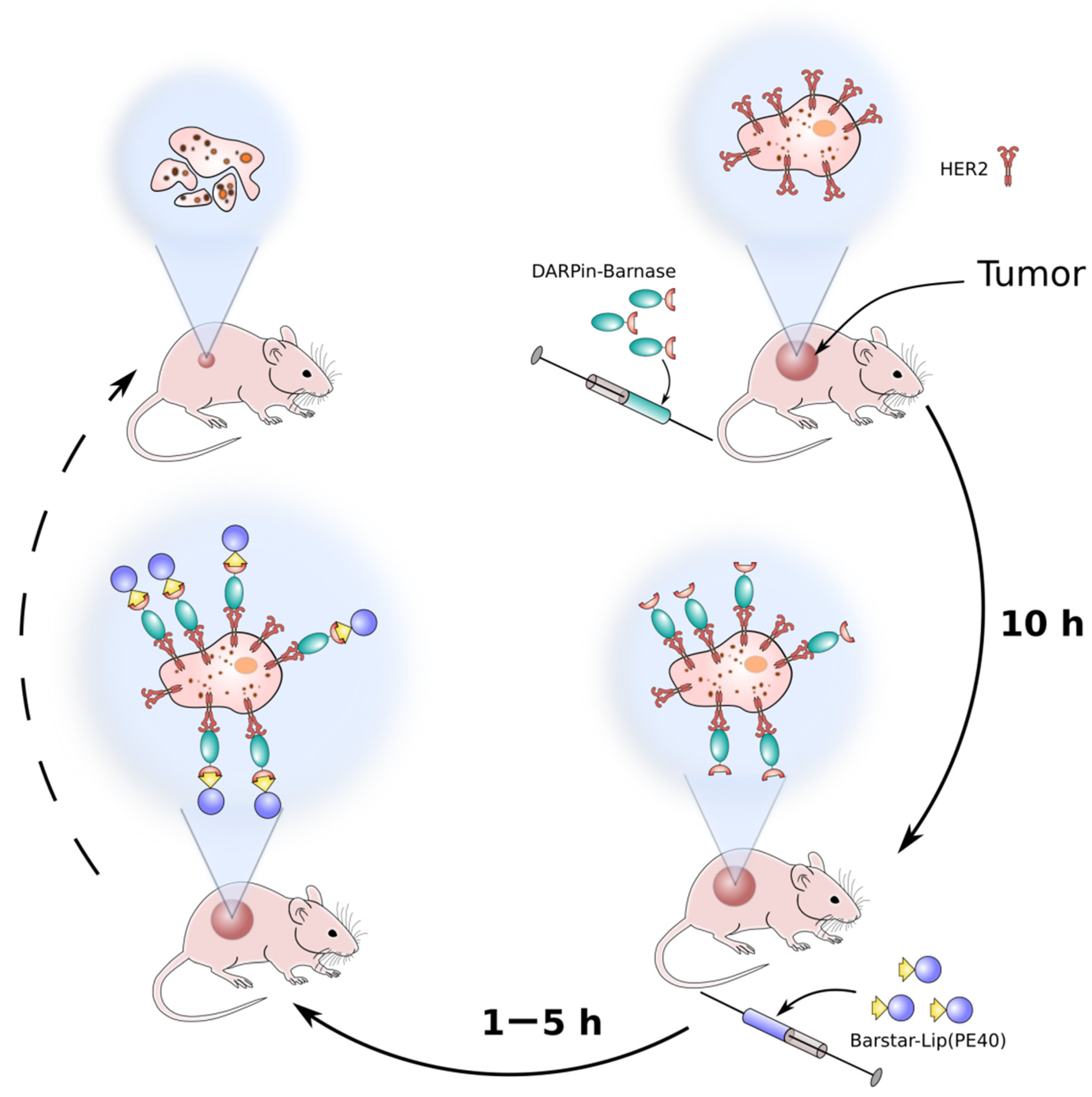
- Shramova, E.I.; Shilova, M.V.; Ryabova, A.V.; Dzhalilova, D.S.; Zolotova, N.A.; Telegin, G.B.; Deyev, S.M.; Proshkina, G.M. Barnase*Barstar-guided two-step targeting approach for drug delivery to tumor cells in vivo. J. Control. Release 2021, 340, 200–208. [Google Scholar] [CrossRef]
- Aghayeva, U.F.; Nikitin, M.P.; Lukash, S.V.; Deyev, S.M. Denaturation-resistant bifunctional colloidal superstructures assembled via the proteinaceous barnase-barstar interface. ACS Nano 2013, 7, 950–961. [Google Scholar] [CrossRef]
- Spång, H.C.; Braathen, R.; Bogen, B. Heterodimeric barnase-barstar vaccine molecules: Influence of one versus two targeting units specific for antigen presenting cells. PLoS ONE 2012, 7, e45393. [Google Scholar] [CrossRef]
- Braathen, R.; Spång, H.C.L.; Hinke, D.M.; Blazevski, J.; Bobic, S.; Fossum, E.; Bogen, B. A DNA vaccine that encodes an antigen-presenting cell-specific heterodimeric protein protects against cancer and influenza. Mol. Ther. Methods Clin. Dev. 2020, 17, 378–392. [Google Scholar] [CrossRef] [PubMed]
- Sapozhnikov, A.M.; Klinkova, A.V.; Shustova, O.A.; Grechikhina, M.V.; Kilyachus, M.S.; Stremovskiy, O.A.; Kovalenko, E.I.; Deyev, S.M. A Novel Approach to Anticancer Therapy: Molecular Modules Based on the Barnase:Barstar Pair for Targeted Delivery of HSP70 to Tumor Cells. Acta Nat. 2018, 10, 85–91. [Google Scholar] [CrossRef]
- Kalinin, R.; Shipunova, V.; Chernikov, I.; Volkov, D.; Shulga, A.; Deyev, S.; Stepanov, A.; Gabibov, A. Modular approach to CAR-T regulation based on the barnase-barstar complex for therapy of oncological diseases. In FEBS Open Bio; WILEY: Hoboken, NJ, USA, 2021; Volume 11, p. 93. [Google Scholar] [CrossRef]
- Zdobnova, T.A.; Stremovskiy, O.A.; Lebedenko, E.N.; Deyev, S.M. Self-Assembling Complexes of Quantum Dots and scFv Antibodies for Cancer Cell Targeting and Imaging. PLoS ONE 2012, 7, e48248. [Google Scholar] [CrossRef]
- Sreenivasan, V.K.A.; Ivukina, E.A.; Deng, W.; Kelf, T.A.; Zdobnova, T.A.; Lukash, S.V.; Veryugin, B.V.; Stremovskiy, O.A.; Zvyagin, A.V.; Deyev, S.M. Barstar:barnase—A versatile platform for colloidal diamond bioconjugation. J. Mater. Chem. 2011, 21, 65–68. [Google Scholar] [CrossRef]
- Grebenik, E.A.; Nadort, A.; Generalova, A.N.; Nechaev, A.V.; Sreenivasan, V.K.A.; Khaydukov, E.V.; Semchishen, V.A.; Popov, A.P.; Sokolov, V.I.; Akhmanov, A.S.; et al. Feasibility study of the optical imaging of a breast cancer lesion labeled with upconversion nanoparticle biocomplexes. J. Biomed. Opt. 2013, 18, 076004. [Google Scholar] [CrossRef] [PubMed]
- Kotelnikova, P.A.; Shipunova, V.O.; Aghayeva, U.F.; Stremovskiy, O.A.; Nikitin, M.P.; Novikov, I.A.; Schulga, A.A.; Deyev, S.M.; Petrov, R.V. Synthesis of Magnetic Nanoparticles Stabilized by Magnetite-Binding Protein for Targeted Delivery to Cancer Cells. Dokl. Biochem. Biophys. 2018, 481, 198–200. [Google Scholar] [CrossRef]
- Shipunova, V.O.; Zelepukin, I.V.; Stremovskiy, O.A.; Nikitin, M.P.; Care, A.; Sunna, A.; Zvyagin, A.V.; Deyev, S.M. Versatile Platform for Nanoparticle Surface Bioengineering Based on SiO2-Binding Peptide and Proteinaceous Barnase*Barstar Interface. ACS Appl. Mater. Interfaces 2018, 10, 17437–17447. [Google Scholar] [CrossRef]
- Balalaeva, I.V.; Zdobnova, T.A.; Sokolova, E.A.; Deyev, S.M. Targeted delivery of quantum dots to the HER2-expressing tumor using recombinant antibodies. Russ. J. Bioorg. Chem. 2015, 41, 536–542. [Google Scholar] [CrossRef]
- Nikitin, M.P.; Zdobnova, T.A.; Lukash, S.V.; Stremovskiy, O.A.; Deyev, S.M. Protein-assisted self-assembly of multifunctional nanoparticles. Proc. Natl. Acad. Sci. USA 2010, 107, 5827–5832. [Google Scholar] [CrossRef] [PubMed]
- Shipunova, V.O.; Kotelnikova, P.A.; Aghayeva, U.F.; Stremovskiy, O.A.; Novikov, I.A.; Schulga, A.A.; Nikitin, M.P.; Deyev, S.M. Self-assembling nanoparticles biofunctionalized with magnetite-binding protein for the targeted delivery to HER2/neu overexpressing cancer cells. J. Magn. Magn. Mater. 2019, 469, 450–455. [Google Scholar] [CrossRef]
- Makarov, A.A.; Kolchinsky, A.; Ilinskaya, O.N. Binase and other microbial RNases as potential anticancer agents. Bioessays 2008, 30, 781–790. [Google Scholar] [CrossRef] [PubMed]
- Fang, E.F.; Ng, T.B. Ribonucleases of different origins with a wide spectrum of medicinal applications. Biochim. Biophys. Acta 2011, 1815, 65–74. [Google Scholar] [CrossRef] [PubMed]
- Ulyanova, V.; Vershinina, V.; Ilinskaya, O. Barnase and binase: Twins with distinct fates. FEBS J. 2011, 278, 3633–3643. [Google Scholar] [CrossRef] [PubMed]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Shilova, O.; Kotelnikova, P.; Proshkina, G.; Shramova, E.; Deyev, S. Barnase-Barstar Pair: Contemporary Application in Cancer Research and Nanotechnology. Molecules 2021, 26, 6785. https://doi.org/10.3390/molecules26226785
Shilova O, Kotelnikova P, Proshkina G, Shramova E, Deyev S. Barnase-Barstar Pair: Contemporary Application in Cancer Research and Nanotechnology. Molecules. 2021; 26(22):6785. https://doi.org/10.3390/molecules26226785
Chicago/Turabian StyleShilova, Olga, Polina Kotelnikova, Galina Proshkina, Elena Shramova, and Sergey Deyev. 2021. "Barnase-Barstar Pair: Contemporary Application in Cancer Research and Nanotechnology" Molecules 26, no. 22: 6785. https://doi.org/10.3390/molecules26226785
APA StyleShilova, O., Kotelnikova, P., Proshkina, G., Shramova, E., & Deyev, S. (2021). Barnase-Barstar Pair: Contemporary Application in Cancer Research and Nanotechnology. Molecules, 26(22), 6785. https://doi.org/10.3390/molecules26226785





