G-Protein Coupled Receptors (GPCRs) in Insects—A Potential Target for New Insecticide Development
Abstract
1. Introduction
2. Whole Genome Sequencing and Transcriptome Analysis—Sequence Comparison and GPCR Characterization in Insects
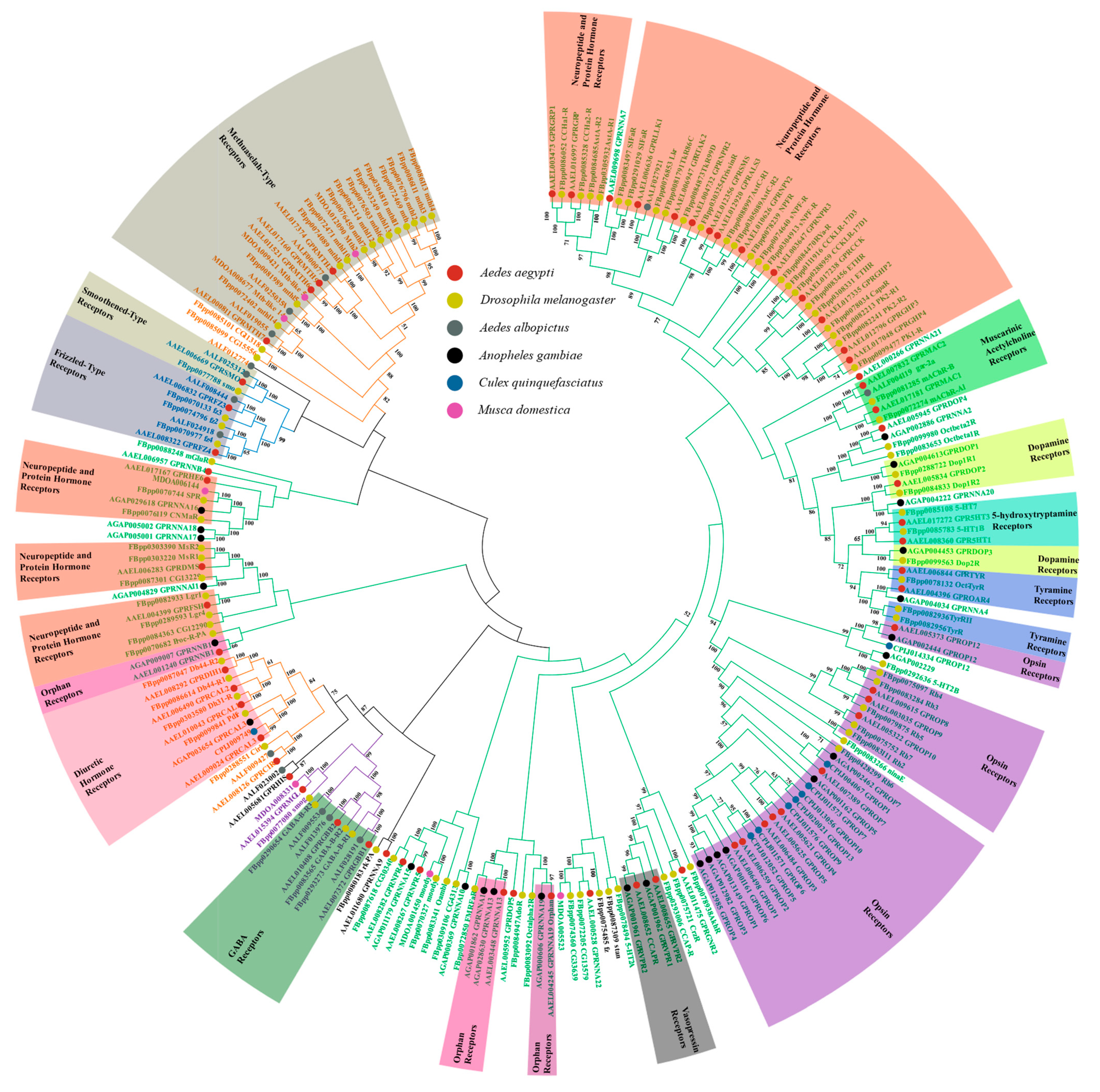
2.1. Classification Systems Used in Characterizing GPCRs
2.2. Receptors of GPCR Involved in Insect Physiology or Insecticide Resistance That Are Potential Targets for Insecticide Development
3. Tissue Specific Expression Analysis of GPCRs in Insects
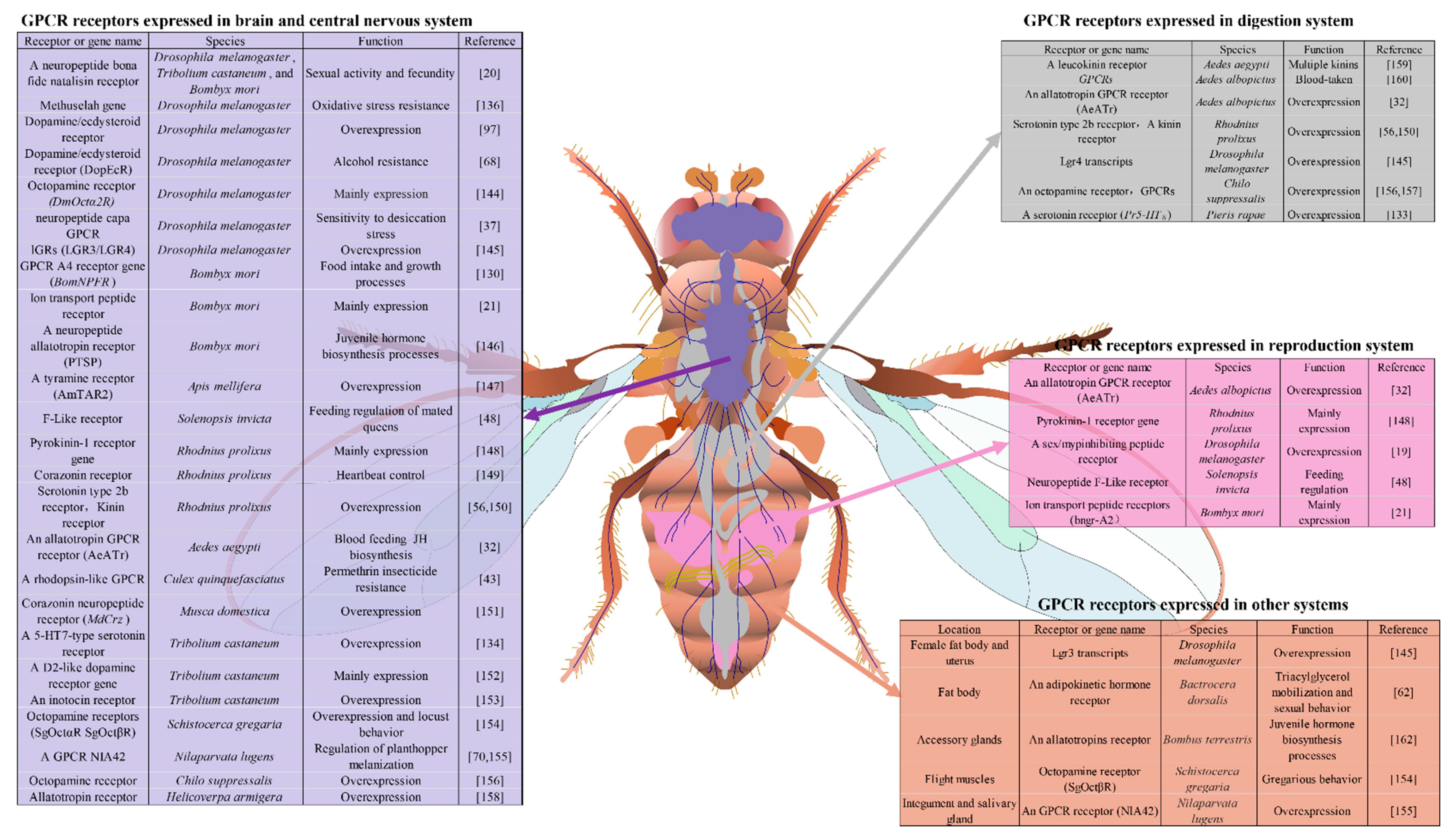
3.1. Brain Tissue and Central Nervous System
3.2. Digestion and Reproduction Systems
3.3. Other Insect Organs
4. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Hilger, D.; Masureel, M.; Kobilka, B.K. Structure and dynamics of GPCR signaling complexes. Nat. Struct. Mol. Biol. 2018, 25, 4–12. [Google Scholar] [CrossRef]
- Bockaert, J.; Pin, J.P. Molecular tinkering of G protein-coupled receptors: An evolutionary success. EMBO J. 1999, 18, 1723–1729. [Google Scholar] [CrossRef] [PubMed]
- Maudsley, S.; Martin, B.; Luttrell, L.M. The Origins of Diversity and Specificity in G Protein-Coupled Receptor Signaling. J. Pharmacol. Exp. Ther. 2005, 314, 485–494. [Google Scholar] [CrossRef]
- Lagerström, M.C.; Schiöth, H.B. Structural diversity of G protein-coupled receptors and significance for drug discovery. Nat. Rev. Drug Discov. 2008, 7, 339–357. [Google Scholar] [CrossRef]
- Spehr, M.; Munger, S.D. Olfactory receptors: G protein-coupled receptors and beyond. J. Neurochem. 2009, 109, 1570–1583. [Google Scholar] [CrossRef] [PubMed]
- Millar, R.P.; Newton, C.L. The Year in G Protein-Coupled Receptor Research. Mol. Endocrinol. 2010, 24, 261–274. [Google Scholar] [CrossRef]
- Eglen, R.M.; Reisine, T. GPCRs Revisited: New Insights Lead to Novel Drugs. Pharmaceuticals 2011, 4, 244–272. [Google Scholar] [CrossRef]
- Gether, U. Uncovering Molecular Mechanisms Involved in Activation of G Protein-Coupled Receptors. Endocr. Rev. 2000, 21, 90–113. [Google Scholar] [CrossRef] [PubMed]
- Goldsmith, Z.G.; Dhanasekaran, D.N. G Protein regulation of MAPK networks. Oncogene 2007, 26, 3122–3142. [Google Scholar] [CrossRef]
- Gerald, W.Z. Calcium channel signaling complexes with receptors and channels. Curr. Mol. Pharmacol. 2015, 8, 8–11. [Google Scholar]
- Flower, D.R. Modelling G-protein-coupled receptors for drug design. Biochim. Biophys. Acta (BBA)-Rev. Biomembr. 1999, 1422, 207–234. [Google Scholar] [CrossRef]
- Hopkins, A.L.; Groom, C.R. The druggable genome. Nat. Rev. Drug Discov. 2002, 1, 727–730. [Google Scholar] [CrossRef] [PubMed]
- Robas, N.; O’Reilly, M.; Katugampola, S.; Fidock, M. Maximizing serendipity: Strategies for identifying ligands for orphan G-protein-coupled receptors. Curr. Opin. Pharmacol. 2003, 3, 121–126. [Google Scholar] [CrossRef]
- Jacoby, E.; Bouhelal, R.; Gerspacher, M.; Seuwen, K. The 7 TM G-Protein-Coupled Receptor Target Family. ChemMedChem 2006, 1, 760–782. [Google Scholar] [CrossRef] [PubMed]
- Hauser, A.S.; Attwood, M.M.; Rask-Andersen, M.; Schiöth, H.B.; Gloriam, D.E. Trends in GPCR drug discovery: New agents, targets and indications. Nat. Rev. Drug Discov. 2017, 16, 829–842. [Google Scholar] [CrossRef] [PubMed]
- Congreve, M.; de Graaf, C.; Swain, N.A.; Tate, C.G. Impact of GPCR Structures on Drug Discovery. Cell 2020, 181, 81–91. [Google Scholar] [CrossRef] [PubMed]
- Hauser, F.; Cazzamali, G.; Williamson, M.; Blenau, W.; Grimmelikhuijzen, C.J. A review of neurohormone GPCRs present in the fruitfly Drosophila melanogaster and the honey bee Apis mellifera. Prog. Neurobiol. 2006, 80, 1–19. [Google Scholar] [CrossRef]
- Hauser, F.; Cazzamali, G.; Williamson, M.; Park, Y.; Li, B.; Tanaka, Y.; Predel, R.; Neupert, S.; Schachtner, J.; Verleyen, P.; et al. A genome-wide inventory of neurohormone GPCRs in the red flour beetle Tribolium castaneum. Front. Neuroendocr. 2008, 29, 142–165. [Google Scholar] [CrossRef]
- Poels, J.; Van Loy, T.; Vandersmissen, H.P.; Van Hiel, B.; Van Soest, S.; Nachman, R.J.; Broeck, J.V. Myoinhibiting peptides are the ancestral ligands of the promiscuous Drosophila sex peptide receptor. Cell. Mol. Life Sci. 2010, 67, 3511–3522. [Google Scholar] [CrossRef]
- Jiang, H.; Lkhagva, A.; Park, Y.; Kim, Y.-J.; Daubnerová, I.; Chae, H.-S.; Šimo, L.; Jung, S.-H.; Yoon, Y.-K.; Lee, N.-R.; et al. Natalisin, a tachykinin-like signaling system, regulates sexual activity and fecundity in insects. Proc. Natl. Acad. Sci. USA 2013, 110, E3526–E3534. [Google Scholar] [CrossRef]
- Nagai, C.; Mabashi-Asazuma, H.; Nagasawa, H.; Nagata, S. Identification and Characterization of Receptors for Ion Transport Peptide (ITP) and ITP-like (ITPL) in the Silkworm Bombyx mori. J. Biol. Chem. 2014, 289, 32166–32177. [Google Scholar] [CrossRef]
- Bai, H.; Palli, S.R. Identification of G protein-coupled receptors required for vitellogenin uptake into the oocytes of the red flour beetle, Tribolium castaneum. Sci. Rep. 2016, 6, 27648. [Google Scholar] [CrossRef] [PubMed]
- Jing, Y.-P.; An, H.; Zhang, S.; Wang, N.; Zhou, S. Protein kinase C mediates juvenile hormone–dependent phosphorylation of Na+/K+-ATPase to induce ovarian follicular patency for yolk protein uptake. J. Biol. Chem. 2018, 293, 20112–20122. [Google Scholar] [CrossRef]
- Marciniak, P.; Urbański, A.; Lubawy, J.; Szymczak, M.; Pacholska-Bogalska, J.; Chowański, S.; Kuczer, M.; Rosiński, G. Short neuropeptide F signaling regulates functioning of male reproductive system in Tenebrio molitor beetle. J. Comp. Physiol. B 2020, 190, 521–534. [Google Scholar] [CrossRef] [PubMed]
- Mendive, F.M.; Van Loy, T.; Claeysen, S.; Poels, J.; Williamson, M.; Hauser, F.; Grimmelikhuijzen, C.J.; Vassart, G.; Broeck, J.V. Drosophila molting neurohormone bursicon is a heterodimer and the natural agonist of the orphan receptor DLGR2. FEBS Lett. 2005, 579, 2171–2176. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.-J.; Zitnan, D.; Cho, K.-H.; Schooley, D.A.; Mizoguchi, A.; Adams, M.E. Central peptidergic ensembles associated with organization of an innate behavior. Proc. Natl. Acad. Sci. USA 2006, 103, 14211–14216. [Google Scholar] [CrossRef] [PubMed]
- Žitňan, D.; Kim, Y.-J.; Žitňanová, I.; Roller, L.; Adams, M. Complex steroid–peptide–receptor cascade controls insect ecdysis. Gen. Comp. Endocrinol. 2007, 153, 88–96. [Google Scholar] [CrossRef] [PubMed]
- Yamanaka, N.; Hua, Y.-J.; Roller, L.; Spalovská-Valachová, I.; Mizoguchi, A.; Kataoka, H.; Tanaka, Y. Bombyx prothoracicostatic peptides activate the sex peptide receptor to regulate ecdysteroid biosynthesis. Proc. Natl. Acad. Sci. USA 2010, 107, 2060–2065. [Google Scholar] [CrossRef] [PubMed]
- Bai, H.; Zhu, F.; Shah, K.; Palli, S.R. Large-scale RNAi screen of G protein-coupled receptors involved in larval growth, molting and metamorphosis in the red flour beetle. BMC Genom. 2011, 12, 388. [Google Scholar] [CrossRef] [PubMed]
- Li, B.; Beeman, R.W.; Park, Y. Functions of duplicated genes encoding CCAP receptors in the red flour beetle, Tribolium castaneum. J. Insect Physiol. 2011, 57, 1190–1197. [Google Scholar] [CrossRef] [PubMed]
- Li, K.; Jia, Q.; Li, S. Juvenile hormone signaling—A mini review. Insect Sci. 2019, 26, 600–606. [Google Scholar] [CrossRef]
- Nouzova, M.; Brockhoff, A.; Mayoral, J.G.; Goodwin, M.; Meyerhof, W.; Noriega, F.G. Functional characterization of an allatotropin receptor expressed in the corpora allata of Mosquitoes. Peptides 2012, 34, 201–208. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Iga, M.; Nakaoka, T.; Suzuki, Y.; Kataoka, H. Pigment Dispersing Factor Regulates Ecdysone Biosynthesis via Bombyx Neuropeptide G Protein Coupled Receptor-B2 in the Prothoracic Glands of Bombyx mori. PLoS ONE 2014, 9, e103239. [Google Scholar] [CrossRef] [PubMed]
- Regna, K.; Kurshan, P.T.; Harwood, B.N.; Jenkins, A.M.; Lai, C.-Q.; Muskavitch, M.A.; Kopin, A.S.; Draper, I. A critical role for the Drosophila dopamine D1-like receptor Dop1R2 at the onset of metamorphosis. BMC Dev. Biol. 2016, 16, 15. [Google Scholar] [CrossRef]
- Kang, X.-L.; Zhang, J.-Y.; Wang, D.; Zhao, Y.-M.; Han, X.-L.; Wang, J.-X.; Zhao, X.-F. The steroid hormone 20-hydroxyecdysone binds to dopamine receptor to repress lepidopteran insect feeding and promote pupation. PLoS Genet. 2019, 15, e1008331. [Google Scholar] [CrossRef] [PubMed]
- Homma, T.; Watanabe, K.; Tsurumaru, S.; Kataoka, H.; Imai, K.; Kamba, M.; Niimi, T.; Yamashita, O.; Yaginuma, T. G protein-coupled receptor for diapause hormone, an inducer of Bombyx embryonic diapause. Biochem. Biophys. Res. Commun. 2006, 344, 386–393. [Google Scholar] [CrossRef]
- Terhzaz, S.; Cabrero, P.; Robben, J.H.; Radford, J.C.; Hudson, B.D.; Milligan, G.; Dow, J.A.T.; Davies, S.-A. Mechanism and Function of Drosophila capa GPCR: A Desiccation Stress-Responsive Receptor with Functional Homology to Human NeuromedinU Receptor. PLoS ONE 2012, 7, e29897. [Google Scholar] [CrossRef]
- Bryon, A.; Wybouw, N.; Dermauw, W.; Tirry, L.; Van Leeuwen, T. Genome wide gene-expression analysis of facultative reproductive diapause in the two-spotted spider mite Tetranychus urticae. BMC Genom. 2013, 14, 1–20. [Google Scholar] [CrossRef]
- Choi, M.-Y.; Estep, A.; Sanscrainte, N.; Becnel, J.; Meer, R.K.V. Identification and expression of PBAN/diapause hormone and GPCRs from Aedes aegypti. Mol. Cell. Endocrinol. 2013, 375, 113–120. [Google Scholar] [CrossRef]
- Devambez, I.; Agha, M.A.; Mitri, C.; Bockaert, J.; Parmentier, M.-L.; Marion-Poll, F.; Grau, Y.; Soustelle, L. Gαo Is Required for L-Canavanine Detection in Drosophila. PLoS ONE 2013, 8, e63484. [Google Scholar] [CrossRef]
- Jiang, H.; Wei, Z.; Nachman, R.J.; Park, Y. Molecular cloning and functional characterization of the diapause hormone receptor in the corn earworm Helicoverpa zea. Peptides 2014, 53, 243–249. [Google Scholar] [CrossRef] [PubMed]
- Li, T.; Liu, L.; Zhang, L.; Liu, N. Role of G-protein-coupled Receptor-related Genes in Insecticide Resistance of the Mosquito, Culex quinquefasciatus. Sci. Rep. 2015, 4, 6474. [Google Scholar] [CrossRef] [PubMed]
- Li, T.; Cao, C.; Yang, T.; Zhang, L.; He, L.; Xi, Z.; Bian, G.; Liu, N. A G-protein-coupled receptor regulation pathway in cytochrome P450-mediated permethrin-resistance in Mosquitoes, Culex quinquefasciatus. Sci. Rep. 2015, 5, 17772. [Google Scholar] [CrossRef] [PubMed]
- Li, T.; Liu, N. Regulation of P450-mediated permethrin resistance in Culex quinquefasciatus by the GPCR/Gαs/AC/cAMP/PKA signaling cascade. Biochem. Biophys. Rep. 2017, 12, 12–19. [Google Scholar] [CrossRef] [PubMed]
- Shen, Z.; Jiang, X.; Yan, L.; Chen, Y.; Wang, W.; Shi, Y.; Shi, L.; Liu, D.; Zhou, N. Structural basis for the interaction of diapause hormone with its receptor in the silkworm, Bombyx mori. FASEB J. 2018, 32, 1338–1353. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.-H.; Di, Y.-Q.; Shen, Q.-Y.; Wang, J.-X.; Zhao, X.-F. The steroid hormone 20-hydroxyecdysone induces phosphorylation and aggregation of stromal interacting molecule 1 for store-operated calcium entry. J. Biol. Chem. 2019, 294, 14922–14936. [Google Scholar] [CrossRef]
- Petruccelli, E.; Lark, A.; Mrkvicka, J.A.; Kitamoto, T. Significance of DopEcR, a G-protein coupled dopamine/ecdysteroid receptor, in physiological and behavioral response to stressors. J. Neurogenet. 2020, 34, 55–68. [Google Scholar] [CrossRef]
- Chen, M.-E.; Pietrantonio, P.V. The short neuropeptide F-like receptor from the red imported fire ant, Solenopsis invicta Buren (Hymenoptera: Formicidae). Arch. Insect Biochem. Physiol. 2006, 61, 195–208. [Google Scholar] [CrossRef]
- Kersch, C.N.; Pietrantonio, P.V. MosquitoAedes aegypti (L.) leucokinin receptor is critical forin vivofluid excretion post blood feeding. FEBS Lett. 2011, 585, 3507–3512. [Google Scholar] [CrossRef]
- Dillen, S.; Zels, S.; Verlinden, H.; Spit, J.; Van Wielendaele, P.; Broeck, J.V. Functional Characterization of the Short Neuropeptide F Receptor in the Desert Locust, Schistocerca gregaria. PLoS ONE 2013, 8, e53604. [Google Scholar] [CrossRef]
- Baumbach, J.; Xu, Y.; Hehlert, P.; Kühnlein, R.P. Gαq, Gγ1 and Plc21C Control Drosophila Body Fat Storage. J. Genet. Genom. 2014, 41, 283–292. [Google Scholar] [CrossRef]
- Lin, F.; Hossain, M.A.; Post, S.; Karashchuk, G.; Tatar, M.; De Meyts, P.; Wade, J.D. Total Solid-Phase Synthesis of Biologically Active Drosophila Insulin-Like Peptide 2 (DILP2). Aust. J. Chem. 2017, 70, 208–212. [Google Scholar] [CrossRef] [PubMed]
- Lu, K.; Zhang, X.; Chen, X.; Li, Y.; Li, W.; Cheng, Y.; Zhou, J.; You, K.; Zhou, Q. Adipokinetic Hormone Receptor Mediates Lipid Mobilization to Regulate Starvation Resistance in the Brown Planthopper, Nilaparvata lugens. Front. Physiol. 2018, 9, 1730. [Google Scholar] [CrossRef]
- Marchal, E.; Schellens, S.; Monjon, E.; Bruyninckx, E.; Marco, H.G.; Gäde, G.; Broeck, J.V.; Verlinden, H. Analysis of Peptide Ligand Specificity of Different Insect Adipokinetic Hormone Receptors. Int. J. Mol. Sci. 2018, 19, 542. [Google Scholar] [CrossRef] [PubMed]
- Yang, H.; Huang, J.; Liu, Y.; Li, J.; Luo, S.; Wu, J. Prediction of the post-translational modifications of adipokinetic hormone receptors from solitary to eusocial bees. Sociobiology 2018, 65, 271–279. [Google Scholar] [CrossRef]
- Sangha, V.; Lange, A.B.; Orchard, I. Identification and cloning of the kinin receptor in the Chagas disease vector, Rhodnius prolixus. Gen. Comp. Endocrinol. 2020, 289, 113380. [Google Scholar] [CrossRef]
- Bainton, R.J.; Tsai, L.T.-Y.; Schwabe, T.; DeSalvo, M.; Gaul, U.; Heberlein, U. moody Encodes Two GPCRs that Regulate Cocaine Behaviors and Blood-Brain Barrier Permeability in Drosophila. Cell 2005, 123, 145–156. [Google Scholar] [CrossRef] [PubMed]
- Hyun, S.; Lee, Y.; Hong, S.-T.; Bang, S.; Paik, D.; Kang, J.; Shin, J.; Lee, J.; Jeon, K.; Hwang, S.; et al. Drosophila GPCR Han Is a Receptor for the Circadian Clock Neuropeptide PDF. Neuron 2005, 48, 267–278. [Google Scholar] [CrossRef] [PubMed]
- Thamm, M.; Balfanz, S.; Scheiner, R.; Baumann, A.; Blenau, W. Characterization of the 5-HT1A receptor of the honeybee (Apis mellifera) and involvement of serotonin in phototactic behavior. Cell. Mol. Life Sci. 2010, 67, 2467–2479. [Google Scholar] [CrossRef]
- Abrieux, A.; Debernard, S.; Maria, A.; Gaertner, C.; Anton, S.; Gadenne, C.; Duportets, L. Involvement of the G-Protein-Coupled Dopamine/Ecdysteroid Receptor DopEcR in the Behavioral Response to Sex Pheromone in an Insect. PLoS ONE 2013, 8, e72785. [Google Scholar] [CrossRef]
- Kwon, H.; Agha, M.A.; Smith, R.C.; Nachman, R.J.; Marion-Poll, F.; Pietrantonio, P.V. Leucokinin mimetic elicits aversive behavior in Mosquito Aedes aegypti (L.) and inhibits the sugar taste neuron. Proc. Natl. Acad. Sci. USA 2016, 113, 6880–6885. [Google Scholar] [CrossRef] [PubMed]
- Hou, Q.-L.; Chen, E.-H.; Jiang, H.-B.; Wei, D.-D.; Gui, S.-H.; Wang, J.-J.; Smagghe, G. Adipokinetic hormone receptor gene identification and its role in triacylglycerol mobilization and sexual behavior in the oriental fruit fly (Bactrocera dorsalis). Insect Biochem. Mol. Biol. 2017, 90, 1–13. [Google Scholar] [CrossRef]
- Grohmann, L.; Blenau, W.; Erber, J.; Ebert, P.R.; Strünker, T.; Baumann, A. Molecular and functional characterization of an octopamine receptor from honeybee (Apis mellifera) brain. J. Neurochem. 2003, 86, 725–735. [Google Scholar] [CrossRef]
- Johnson, E.C.; Bohn, L.M.; Taghert, P.H. Drosophila CG8422 encodes a functional diuretic hormone receptor. J. Exp. Biol. 2004, 207, 743–748. [Google Scholar] [CrossRef]
- Kwon, H.; Lu, H.-L.; Longnecker, M.T.; Pietrantonio, P.V. Role in Diuresis of a Calcitonin Receptor (GPRCAL1) Expressed in a Distal-Proximal Gradient in Renal Organs of the Mosquito Aedes aegypti (L.). PLoS ONE 2012, 7, e50374. [Google Scholar] [CrossRef]
- Kwon, H.; Pietrantonio, P.V. Calcitonin receptor 1 (AedaeGPCRCAL1) hindgut expression and direct role in myotropic action in females of the Mosquito Aedes aegypti (L.). Insect Biochem. Mol. Biol. 2013, 43, 588–593. [Google Scholar] [CrossRef] [PubMed]
- Lee, D.; Broeck, J.V.; Lange, A.B. Identification and Expression of the CCAP Receptor in the Chagas’ Disease Vector, Rhodnius prolixus, and Its Involvement in Cardiac Control. PLoS ONE 2013, 8, e68897. [Google Scholar] [CrossRef] [PubMed]
- Petruccelli, E.; Li, Q.; Rao, Y.; Kitamoto, T. The Unique Dopamine/Ecdysteroid Receptor Modulates Ethanol-Induced Sedation in Drosophila. J. Neurosci. 2016, 36, 4647–4657. [Google Scholar] [CrossRef] [PubMed]
- Pietrantonio, P.V.; Xiong, C.; Nachman, R.J.; Shen, Y. G protein-coupled receptors in arthropod vectors: Omics and pharmacological approaches to elucidate ligand-receptor interactions and novel organismal functions. Curr. Opin. Insect Sci. 2018, 29, 12–20. [Google Scholar] [CrossRef] [PubMed]
- Uchiyama, H.; Maehara, S.; Ohta, H.; Seki, T.; Tanaka, Y. Elevenin regulates the body color through a G protein-coupled receptor NlA42 in the brown planthopper Nilaparvata lugens. Gen. Comp. Endocrinol. 2018, 258, 33–38. [Google Scholar] [CrossRef]
- Li, M.; Reid, W.R.; Zhang, L.; Scott, J.G.; Gao, X.; Kristensen, M.; Liu, N. A whole transcriptomal linkage analysis of gene co-regulation in insecticide resistant house flies, Musca domestica. BMC Genom. 2013, 14, 803. [Google Scholar] [CrossRef] [PubMed]
- Ma, Z.; Zhang, Y.; You, C.; Zeng, X.; Gao, X. The role of G protein-coupled receptor-related genes in cytochrome P450-mediated resistance of the house fly, Musca domestica (Diptera: Muscidae), to imidacloprid. Insect Mol. Biol. 2019, 29, 92–103. [Google Scholar] [CrossRef] [PubMed]
- Liu, N. Insecticide Resistance in Mosquitoes: Impact, Mechanisms, and Research Directions. Annu. Rev. Ѐntomol. 2015, 60, 537–559. [Google Scholar] [CrossRef]
- Attwood, T.K.; Findlay, J.B.C. Fingerprinting G-protein-coupled receptors. Protein Eng. Des. Sel. 1994, 7, 195–203. [Google Scholar] [CrossRef] [PubMed]
- Kolakowski, L.F., Jr. GCRDb: A G-protein-coupled receptor database. Recept. Channels 1994, 2, 1–7. [Google Scholar] [PubMed]
- Schiöth, H.B.; Fredriksson, R. The GRAFS classification system of G-protein coupled receptors in comparative perspective. Gen. Comp. Endocrinol. 2005, 142, 94–101. [Google Scholar] [CrossRef] [PubMed]
- Hu, G.-M.; Mai, T.-L.; Chen, C.-M. Visualizing the GPCR Network: Classification and Evolution. Sci. Rep. 2017, 7, 1–15. [Google Scholar] [CrossRef]
- Adams, M.D.; Celniker, S.E.; Holt, R.A.; Evans, C.A.; Gocayne, J.D.; Amanatides, P.G.; Scherer, S.E.; Li, P.W.; Hoskins, R.A.; Galle, R.F.; et al. The Genome Sequence of Drosophila melanogaster. Science 2000, 287, 2185–2195. [Google Scholar] [CrossRef]
- Holt, R.A.; Subramanian, G.M.; Halpern, A.; Sutton, G.G.; Charlab, R.; Nusskern, D.R.; Wincker, P.; Clark, A.G.; Ribeiro, J.C.; Wides, R.; et al. The Genome Sequence of the Malaria Mosquito Anopheles gambiae. Science 2002, 298, 129–149. [Google Scholar] [CrossRef] [PubMed]
- Nene, V.; Wortman, J.R.; Lawson, D.; Haas, B.; Kodira, C.; Tu, Z.; Loftus, B.; Xi, Z.; Megy, K.; Grabherr, M.; et al. Genome Sequence of Aedes aegypti, a Major Arbovirus Vector. Science 2007, 316, 1718–1723. [Google Scholar] [CrossRef]
- Arensburger, P.; Megy, K.; Waterhouse, R.M.; Abrudan, J.; Amedeo, P.; Antelo, B.; Bartholomay, L.; Bidwell, S.; Caler, E.; Camara, F.; et al. Sequencing of Culex quinquefasciatus Establishes a Platform for Mosquito Comparative Genomics. Science 2010, 330, 86–88. [Google Scholar] [CrossRef] [PubMed]
- Scott, J.G.; Warren, W.C.; Beukeboom, L.W.; Bopp, D.; Clark, A.G.; Giers, S.D.; Hediger, M.; Jones, A.K.; Kasai, S.; A Leichter, C.; et al. Genome of the house fly, Musca domestica L., a global vector of diseases with adaptations to a septic environment. Genome Biol. 2014, 15, 1–17. [Google Scholar] [CrossRef] [PubMed]
- Li, F.; Zhao, X.; Li, M.; He, K.; Huang, C.; Zhou, Y.; Li, Z.; Walters, J.R. Insect genomes: Progress and challenges. Insect Mol. Biol. 2019, 28, 739–758. [Google Scholar] [CrossRef] [PubMed]
- Brody, T.; Cravchik, A. Drosophila melanogaster G Protein–Coupled Receptors. J. Cell Biol. 2000, 150, F83–F88. [Google Scholar] [CrossRef]
- Hanlon, C.D.; Andrew, D.J. Outside-in signaling–A brief review of GPCR signaling with a focus on the Drosophila GPCR family. J. Cell Sci. 2015, 128, 3533–3542. [Google Scholar] [CrossRef] [PubMed]
- Hill, C.A.; Fox, A.N.; Pitts, R.J.; Kent, L.B.; Tan, P.L.; Chrystal, M.A.; Cravchik, A.; Collins, F.H.; Robertson, H.M.; Zwiebel, L.J. G Protein-Coupled Receptors inAnopheles gambiae. Science 2002, 298, 176–178. [Google Scholar] [CrossRef]
- Fan, Y.; Sun, P.; Wang, Y.; He, X.; Deng, X.; Chen, X.; Zhang, G.; Chen, X.; Zhou, N. The G protein-coupled receptors in the silkworm, Bombyx mori. Insect Biochem. Mol. Biol. 2010, 40, 581–591. [Google Scholar] [CrossRef]
- Anstead, C.A.; Korhonen, P.K.; Young, N.D.; Hall, R.S.; Jex, A.R.; Murali, S.C.; Hughes, D.S.; Lee, S.F.; Perry, T.; Stroehlein, A.J.; et al. Lucilia cuprina genome unlocks parasitic fly biology to underpin future interventions. Nat. Commun. 2015, 6, 7344. [Google Scholar] [CrossRef]
- Şahbaz, B.D. Prediction and expression analysis of G protein-coupled receptors in the laboratory stick insect, Carausius morosus. Turk. J. Boil. 2019, 43, 77–88. [Google Scholar] [CrossRef]
- Calkins, T.L.; Tamborindeguy, C.; Pietrantonio, P.V. GPCR annotation, G proteins, and transcriptomics of fire ant (Solenopsis invicta) queen and worker brain: An improved view of signaling in an invasive superorganism. Gen. Comp. Endocrinol. 2019, 278, 89–103. [Google Scholar] [CrossRef]
- Veenstra, J.A.; Rombauts, S.; Grbić, M. In silico cloning of genes encoding neuropeptides, neurohormones and their putative G-protein coupled receptors in a spider mite. Insect Biochem. Mol. Biol. 2012, 42, 277–295. [Google Scholar] [CrossRef]
- Aikins, M.J.; Schooley, D.A.; Begum, K.; Detheux, M.; Beeman, R.W.; Park, Y. Vasopressin-like peptide and its receptor function in an indirect diuretic signaling pathway in the red flour beetle. Insect Biochem. Mol. Biol. 2008, 38, 740–748. [Google Scholar] [CrossRef] [PubMed]
- Caers, J.; Verlinden, H.; Zels, S.; Vandersmissen, H.P.; Vuerinckx, K.; Schoofs, L. More than two decades of research on insect neuropeptide GPCRs: An overview. Front. Endocrinol. 2012, 3, 151. [Google Scholar] [CrossRef]
- Garczynski, S.F.; Brown, M.R.; Shen, P.; Murray, T.F.; Crim, J.W. Characterization of a functional neuropeptide F receptor from Drosophila melanogaster. Peptides 2002, 23, 773–780. [Google Scholar] [CrossRef]
- Vogel, K.J.; Brown, M.R.; Strand, M.R. Phylogenetic Investigation of Peptide Hormone and Growth Factor Receptors in Five Dipteran Genomes. Front. Endocrinol. 2013, 4, 193. [Google Scholar] [CrossRef] [PubMed]
- Xia, R.; Li, M.; Wu, Y.; Qi, Y.; Ye, G.; Huang, J. A new family of insect muscarinic acetylcholine receptors. Insect Mol. Biol. 2016, 25, 362–369. [Google Scholar] [CrossRef]
- Srivastava, D.P.; Yu, E.J.; Kennedy, K.; Chatwin, H.; Reale, V.; Hamon, M.; Smith, T.; Evans, P.D. Rapid, Nongenomic Responses to Ecdysteroids and Catecholamines Mediated by a Novel Drosophila G-Protein-Coupled Receptor. J. Neurosci. 2005, 25, 6145–6155. [Google Scholar] [CrossRef]
- Deveci, D.; Martin, F.A.; Leopold, P.; Romero, N.M. AstA Signaling Functions as an Evolutionary Conserved Mechanism Timing Juvenile to Adult Transition. Curr. Biol. 2019, 29, 813–822. [Google Scholar] [CrossRef]
- Chen, J.; Reiher, W.; Hermann-Luibl, C.; Sellami, A.; Cognigni, P.; Kondo, S.; Helfrich-Förster, C.; Veenstra, J.A.; Wegener, C. Allatostatin A Signalling in Drosophila Regulates Feeding and Sleep and Is Modulated by PDF. PLoS Genet. 2016, 12, e1006346. [Google Scholar] [CrossRef]
- Zandawala, M.; Yurgel, M.E.; Texada, M.J.; Liao, S.; Rewitz, K.F.; Keene, A.C.; Nässel, D.R. Modulation of Drosophila post-feeding physiology and behavior by the neuropeptide leucokinin. PLoS Genet. 2018, 14, e1007767. [Google Scholar] [CrossRef]
- Yapici, N.; Kim, Y.-J.; Ribeiro, C.; Dickson, B.J. A receptor that mediates the post-mating switch in Drosophila reproductive behaviour. Nat. Cell Biol. 2007, 451, 33–37. [Google Scholar] [CrossRef] [PubMed]
- Sellami, A.; Veenstra, J.A. SIFamide acts on fruitless neurons to modulate sexual behavior in Drosophila melanogaster. Peptides 2015, 74, 50–56. [Google Scholar] [CrossRef] [PubMed]
- Yurgel, M.E.; Kakad, P.; Zandawala, M.; Nässel, D.R.; Godenschwege, T.A.; Keene, A.C. A single pair of leucokinin neurons are modulated by feeding state and regulate sleep–metabolism interactions. PLoS Biol. 2019, 17, e2006409. [Google Scholar] [CrossRef] [PubMed]
- Wu, F.; Deng, B.; Xiao, N.; Wang, T.; Li, Y.; Wang, R.; Shi, K.; Luo, D.-G.; Rao, Y.; Zhou, C. A neuropeptide regulates fighting behavior in Drosophila melanogaster. eLife 2020, 9. [Google Scholar] [CrossRef] [PubMed]
- Sano, H.; Nakamura, A.; Texada, M.J.; Truman, J.W.; Ishimoto, H.; Kamikouchi, A.; Nibu, Y.; Kume, K.; Ida, T.; Kojima, M. The Nutrient-Responsive Hormone CCHamide-2 Controls Growth by Regulating Insulin-like Peptides in the Brain of Drosophila melanogaster. PLoS Genet. 2015, 11, e1005209. [Google Scholar] [CrossRef]
- Iyison, N.B.; Sinmaz, M.G.; Sahbaz, B.D.; Shahraki, A.; Aksoydan, B.; Durdagi, S. In silico characterization of adipokinetic hormone receptor and screening for pesticide candidates against stick insect, Carausius morosus. J. Mol. Graph. Model. 2020, 101, 107720. [Google Scholar] [CrossRef]
- Posnien, N.; Hopfen, C.; Hilbrant, M.; Ramos-Womack, M.; Murat, S.; Schönauer, A.; Herbert, S.L.; Nunes, M.D.S.; Arif, S.; Breuker, C.J.; et al. Evolution of Eye Morphology and Rhodopsin Expression in the Drosophila melanogaster Species Subgroup. PLoS ONE 2012, 7, e37346. [Google Scholar] [CrossRef]
- Egerod, K.; Reynisson, E.; Hauser, F.; Cazzamali, G.; Williamson, M.; Grimmelikhuijzen, C.J.P. Molecular cloning and functional expression of the first two specific insect myosuppressin receptors. Proc. Natl. Acad. Sci. USA 2003, 100, 9808–9813. [Google Scholar] [CrossRef]
- Hauser, F.; Williamson, M.; Cazzamali, G.; Grimmelikhuijzen, C.J.P. Identifying neuropeptide and protein hormone receptors in Drosophila melanogaster by exploiting genomic data. Briefings Funct. Genom. Proteom. 2006, 4, 321–330. [Google Scholar] [CrossRef]
- Bayliss, A.; Roselli, G.; Evans, P.D. A comparison of the signalling properties of two tyramine receptors from Drosophila. J. Neurochem. 2013, 125, 37–48. [Google Scholar] [CrossRef]
- Li, C.; Chen, M.; Sang, M.; Liu, X.; Wu, W.; Li, B. Comparative genomic analysis and evolution of family-B G protein-coupled receptors from six model insect species. Gene 2013, 519, 1–12. [Google Scholar] [CrossRef] [PubMed]
- De Mendoza, A.; Jones, J.W.; Friedrich, M. Methuselah/Methuselah-like G protein-coupled receptors constitute an ancient metazoan gene family. Sci. Rep. 2016, 6, 21801. [Google Scholar] [CrossRef] [PubMed]
- Hector, C.E.; Bretz, C.A.; Zhao, Y.; Johnson, E.C. Functional differences between two CRF-related diuretic hormone receptors in Drosophila. J. Exp. Biol. 2009, 212, 3142–3147. [Google Scholar] [CrossRef] [PubMed]
- Goda, T.; Doi, M.; Umezaki, Y.; Murai, I.; Shimatani, H.; Chu, M.L.; Nguyen, V.H.; Okamura, H.; Hamada, F.N. Calcitonin receptors are ancient modulators for rhythms of preferential temperature in insects and body temperature in mammals. Genes Dev. 2018, 32, 140–155. [Google Scholar] [CrossRef] [PubMed]
- Lin, Y.-J.; Seroude, L.; Benzer, S. Extended Life-Span and Stress Resistance in the Drosophila Mutant methuselah. Science 1998, 282, 943–946. [Google Scholar] [CrossRef] [PubMed]
- Cvejic, S.; Zhu, Z.; Felice, S.J.; Berman, Y.; Huang, X.-Y. The endogenous ligand Stunted of the GPCR Methuselah extends lifespan in Drosophila. Nat. Cell Biol. 2004, 6, 540–546. [Google Scholar] [CrossRef]
- Harmar, A.J. Family-B G-protein-coupled receptors. Genome Biol. 2001, 2, 1–3013. [Google Scholar] [CrossRef]
- Mezler, M.; Müller, T.; Raming, K. Cloning and functional expression of GABABreceptors from Drosophila. Eur. J. Neurosci. 2001, 13, 477–486. [Google Scholar] [CrossRef]
- Adler, P.N.; Vinson, C.; Park, W.J.; Conover, S.; Klein, L. Molecular structure of frizzled, a Drosophila tissue polarity gene. Genetics 1990, 126, 401–416. [Google Scholar] [CrossRef]
- I Strutt, D. Asymmetric Localization of Frizzled and the Establishment of Cell Polarity in the Drosophila Wing. Mol. Cell 2001, 7, 367–375. [Google Scholar] [CrossRef]
- Povelones, M.; Nusse, R. The role of the cysteine-rich domain of Frizzled in Wingless-Armadillo signaling. EMBO J. 2005, 24, 3493–3503. [Google Scholar] [CrossRef]
- Heuvel, M.V.D.; Ingham, P.W. smoothened encodes a receptor-like serpentine protein required for hedgehog signalling. Nat. Cell Biol. 1996, 382, 547–551. [Google Scholar] [CrossRef]
- Fan, J.; Liu, Y.; Jia, J. Hh-induced Smoothened conformational switch is mediated by differential phosphorylation at its C-terminal tail in a dose- and position-dependent manner. Dev. Biol. 2012, 366, 172–184. [Google Scholar] [CrossRef]
- Martin, B.L.; Kimelman, D. Wnt Signaling and the Evolution of Embryonic Posterior Development. Curr. Biol. 2009, 19, R215–R219. [Google Scholar] [CrossRef]
- Villarreal, C.M.; Darakananda, K.; Wang, V.R.; Jayaprakash, P.M.; Suzuki, Y. Hedgehog signaling regulates imaginal cell differentiation in a basally branching holometabolous insect. Dev. Biol. 2015, 404, 125–135. [Google Scholar] [CrossRef]
- Abrieux, A.; Duportets, L.; Debernard, S.; Gadenne, C.; Anton, S. The GPCR membrane receptor, DopEcR, mediates the actions of both dopamine and ecdysone to control sex pheromone perception in an insect. Front. Behav. Neurosci. 2014, 8, 312. [Google Scholar] [CrossRef] [PubMed]
- Lark, A.; Kitamoto, T.; Martin, J.-R. Modulation of neuronal activity in the Drosophila mushroom body by DopEcR, a unique dual receptor for ecdysone and dopamine. Biochim. Biophys. Acta (BBA)-Bioenerg. 2017, 1864, 1578–1588. [Google Scholar] [CrossRef] [PubMed]
- Lungchukiet, P.; Zhang, J.; Tobe, S.S.; Bendena, W.G. Quantification of allatostatin receptor mRNA levels in the cockroach, Diploptera punctata, using real-time PCR. J. Insect Physiol. 2008, 54, 981–987. [Google Scholar] [CrossRef] [PubMed]
- Ida, T.; Takahashi, T.; Tominaga, H.; Sato, T.; Kume, K.; Ozaki, M.; Hiraguchi, T.; Maeda, T.; Shiotani, H.; Terajima, S.; et al. Identification of the novel bioactive peptides dRYamide-1 and dRYamide-2, ligands for a neuropeptide Y-like receptor in Drosophila. Biochem. Biophys. Res. Commun. 2011, 410, 872–877. [Google Scholar] [CrossRef]
- Deng, X.; Yang, H.; He, X.; Liao, Y.; Zheng, C.; Zhou, Q.; Zhu, C.; Zhang, G.; Gao, J.; Zhou, N. Activation of Bombyx neuropeptide G protein-coupled receptor A4 via a Gαi-dependent signaling pathway by direct interaction with neuropeptide F from silkworm, Bombyx mori. Insect Biochem. Mol. Biol. 2014, 45, 77–88. [Google Scholar] [CrossRef]
- Gross, A.D.; Temeyer, K.B.; Day, T.A.; de León, A.A.P.; Kimber, M.J.; Coats, J.R. Pharmacological characterization of a tyramine receptor from the southern cattle tick, Rhipicephalus (Boophilus) microplus. Insect Biochem. Mol. Biol. 2015, 63, 47–53. [Google Scholar] [CrossRef] [PubMed]
- Collin, C.; Hauser, F.; Krogh-Meyer, P.; Hansen, K.K.; De Valdivia, E.G.; Williamson, M.; Grimmelikhuijzen, C.J. Identification of the Drosophila and Tribolium receptors for the recently discovered insect RYamide neuropeptides. Biochem. Biophys. Res. Commun. 2011, 412, 578–583. [Google Scholar] [CrossRef]
- Qi, Y.-X.; Xia, R.-Y.; Wu, Y.-S.; Stanley, D.; Huang, J.; Ye, G.-Y. Larvae of the small white butterfly, Pieris rapae, express a novel serotonin receptor. J. Neurochem. 2014, 131, 767–777. [Google Scholar] [CrossRef]
- Vleugels, R.; Lenaerts, C.; Broeck, J.V.; Verlinden, H. Signalling properties and pharmacology of a 5-HT7-type serotonin receptor fromTribolium castaneum. Insect Mol. Biol. 2013, 23, 230–243. [Google Scholar] [CrossRef] [PubMed]
- Cao, C.; Sun, L.; Du, H.; Moural, T.W.; Bai, H.; Liu, P.; Zhu, F. Physiological functions of a methuselah-like G protein coupled receptor in Lymantria dispar Linnaeus. Pestic. Biochem. Physiol. 2019, 160, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Pandey, A.; Khatoon, R.; Saini, S.; Vimal, D.; Patel, D.K.; Narayan, G.; Chowdhuri, D.K. Efficacy of methuselah gene mutation toward tolerance of dichlorvos exposure in Drosophila melanogaster. Free. Radic. Biol. Med. 2015, 83, 54–65. [Google Scholar] [CrossRef]
- Sun, Y.; Zou, P.; Yu, X.-Y.; Chen, C.; Yu, J.; Shi, L.-N.; Hong, S.-C.; Zhou, D.; Chang, X.-L.; Wang, W.-J.; et al. Functional characterization of an arrestin gene on insecticide resistance of Culex pipiens pallens. Parasites Vectors 2012, 5, 134. [Google Scholar] [CrossRef] [PubMed]
- Baron, S.; Van Der Merwe, N.A.; Madder, M.; Maritz-Olivier, C. SNP Analysis Infers that Recombination Is Involved in the Evolution of Amitraz Resistance in Rhipicephalus microplus. PLoS ONE 2015, 10, e0131341. [Google Scholar] [CrossRef]
- Meyer, J.M.; Ejendal, K.F.K.; Avramova, L.V.; Garland-Kuntz, E.E.; Giraldo-Calderón, G.I.; Brust, T.F.; Watts, V.J.; Hill, C.A. A “Genome-to-Lead” Approach for Insecticide Discovery: Pharmacological Characterization and Screening of Aedes aegypti D1-like Dopamine Receptors. PLoS Negl. Trop. Dis. 2012, 6, e1478. [Google Scholar] [CrossRef]
- Hill, C.A.; Meyer, J.M.; Ejendal, K.F.; Echeverry, D.F.; Lang, E.G.; Avramova, L.V.; Conley, J.M.; Watts, V.J. Re-invigorating the insecticide discovery pipeline for vector control: GPCRs as targets for the identification of next gen insecticides. Pestic. Biochem. Physiol. 2013, 106, 141–148. [Google Scholar] [CrossRef]
- Nuss, A.B.; Ejendal, K.F.K.; Doyle, T.B.; Meyer, J.M.; Lang, E.G.; Watts, V.J.; Hill, C.A. Dopamine Receptor Antagonists as New Mode-of-Action Insecticide Leads for Control of Aedes and Culex Mosquito Vectors. PLoS Negl. Trop. Dis. 2015, 9, e0003515. [Google Scholar] [CrossRef]
- Hapairai, L.K.; Mysore, K.; Sun, L.; Li, P.; Wang, C.-W.; Scheel, N.D.; Lesnik, A.; Scheel, M.P.; Igiede, J.; Wei, N.; et al. Characterization of an adulticidal and larvicidal interfering RNA pesticide that targets a conserved sequence in Mosquito G protein-coupled dopamine 1 receptor genes. Insect Biochem. Mol. Biol. 2020, 120, 103359. [Google Scholar] [CrossRef] [PubMed]
- Tobin, A.B.; Butcher, A.J.; Kong, K.C. Location, location, location…site-specific GPCR phosphorylation offers a mechanism for cell-type-specific signalling. Trends Pharmacol. Sci. 2008, 29, 413–420. [Google Scholar] [CrossRef] [PubMed]
- Qi, Y.-X.; Xu, G.; Gu, G.-X.; Mao, F.; Ye, G.-Y.; Liu, W.; Huang, J. A new Drosophila octopamine receptor responds to serotonin. Insect Biochem. Mol. Biol. 2017, 90, 61–70. [Google Scholar] [CrossRef] [PubMed]
- Van Hiel, M.B.; Vandersmissen, H.P.; Proost, P.; Broeck, J.V. Cloning, constitutive activity and expression profiling of two receptors related to relaxin receptors in Drosophila melanogaster. Peptides 2015, 68, 83–90. [Google Scholar] [CrossRef]
- Yamanaka, N.; Yamamoto, S.; Žitňan, D.; Watanabe, K.; Kawada, T.; Satake, H.; Kaneko, Y.; Hiruma, K.; Tanaka, Y.; Shinoda, T.; et al. Neuropeptide Receptor Transcriptome Reveals Unidentified Neuroendocrine Pathways. PLoS ONE 2008, 3, e3048. [Google Scholar] [CrossRef] [PubMed]
- Reim, T.; Balfanz, S.; Baumann, A.; Blenau, W.; Thamm, M.; Scheiner, R. AmTAR2: Functional characterization of a honeybee tyramine receptor stimulating adenylyl cyclase activity. Insect Biochem. Mol. Biol. 2017, 80, 91–100. [Google Scholar] [CrossRef]
- Paluzzi, J.-P.; O’Donnell, M.J. Identification, spatial expression analysis and functional characterization of a pyrokinin-1 receptor in the Chagas’ disease vector, Rhodnius prolixus. Mol. Cell. Endocrinol. 2012, 363, 36–45. [Google Scholar] [CrossRef]
- Hamoudi, Z.; Lange, A.B.; Orchard, I. Identification and Characterization of the Corazonin Receptor and Possible Physiological Roles of the Corazonin-Signaling Pathway in Rhodnius prolixus. Front. Neurosci. 2016, 10, 357. [Google Scholar] [CrossRef]
- Paluzzi, J.-P.V.; Ebhatt, G.; Wang, C.-H.J.; Ezandawala, M.; Lange, A.B.; Eorchard, I. Identification, functional characterization, and pharmacological profile of a serotonin type-2b receptor in the medically important insect, Rhodnius prolixus. Front. Neurosci. 2015, 9, 175. [Google Scholar] [CrossRef]
- Sha, K.; Conner, W.C.; Choi, D.Y.; Park, J.H. Characterization, expression, and evolutionary aspects of Corazonin neuropeptide and its receptor from the House Fly, Musca domestica (Diptera: Muscidae). Gene 2012, 497, 191–199. [Google Scholar] [CrossRef]
- Verlinden, H.; Vleugels, R.; Verdonck, R.; Urlacher, E.; Broeck, J.V.; Mercer, A. Pharmacological and signalling properties of a D2-like dopamine receptor (Dop3) in Tribolium castaneum. Insect Biochem. Mol. Biol. 2015, 56, 9–20. [Google Scholar] [CrossRef] [PubMed]
- Stafflinger, E.; Hansen, K.K.; Hauser, F.; Schneider, M.; Cazzamali, G.; Williamson, M.; Grimmelikhuijzen, C.J.P. Cloning and identification of an oxytocin/vasopressin-like receptor and its ligand from insects. Proc. Natl. Acad. Sci. USA 2008, 105, 3262–3267. [Google Scholar] [CrossRef]
- Verlinden, H.; Vleugels, R.; Marchal, E.; Badisco, L.; Tobback, J.; Pflüger, H.-J.; Blenau, W.; Broeck, J.V. The cloning, phylogenetic relationship and distribution pattern of two new putative GPCR-type octopamine receptors in the desert locust (Schistocerca gregaria). J. Insect Physiol. 2010, 56, 868–875. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Wang, W.; Ma, Q.; Shen, Z.; Zhang, M.; Zhou, N.; Zhang, C. Elevenin signaling modulates body color through the tyrosine-mediated cuticle melanism pathway. FASEB J. 2019, 33, 9731–9741. [Google Scholar] [CrossRef]
- Wu, S.-F.; Xu, G.; Qi, Y.-X.; Xia, R.-Y.; Huang, J.; Ye, G.-Y. Two splicing variants of a novel family of octopamine receptors with different signaling properties. J. Neurochem. 2014, 129, 37–47. [Google Scholar] [CrossRef] [PubMed]
- Xu, G.; Gu, G.-X.; Teng, Z.-W.; Wu, S.-F.; Huang, J.; Song, Q.-S.; Ye, G.-Y.; Fang, Q. Identification and expression profiles of neuropeptides and their G protein-coupled receptors in the rice stem borer Chilo suppressalis. Sci. Rep. 2016, 6, 28976. [Google Scholar] [CrossRef] [PubMed]
- Zhang, F.; Wang, J.; Thakur, K.; Hu, F.; Zhang, J.-G.; Jiang, X.-F.; An, S.-H.; Jiang, H.; Jiang, L.; Wei, Z.-J. Isolation functional characterization of allatotropin receptor from the cotton bollworm, Helicoverpa armigera. Peptides 2019, 122, 169874. [Google Scholar] [CrossRef] [PubMed]
- Pietrantonio, P.V.; Jagge, C.; Taneja-Bageshwar, S.; Nachman, R.J.; Barhoumi, R. The Mosquito Aedes aegypti (L.) leucokinin receptor is a multiligand receptor for the three Aedes kinins. Insect Mol. Biol. 2005, 14, 55–67. [Google Scholar] [CrossRef]
- Esquivel, C.J.; Cassone, B.J.; Piermarini, P.M. Ade novotranscriptome of the Malpighian tubules in non-blood-fed and blood-fed Asian tiger Mosquitoes Aedes albopictus: Insights into diuresis, detoxification, and blood meal processing. PeerJ 2016, 4, e1784. [Google Scholar] [CrossRef]
- Munoz, S.; Guerrero, F.D.; Kellogg, A.; Heekin, A.M.; Leung, M.-Y. Bioinformatic prediction of G protein-coupled receptor encoding sequences from the transcriptome of the foreleg, including the Haller’s organ, of the cattle tick, Rhipicephalus australis. PLoS ONE 2017, 12, e0172326. [Google Scholar] [CrossRef] [PubMed]
- Verlinden, H.; Lismont, E.; Bil, M.; Urlacher, E.; Mercer, A.; Broeck, J.V.; Huybrechts, R. Characterisation of a functional allatotropin receptor in the bumblebee, Bombus terrestris (Hymenoptera, Apidae). Gen. Comp. Endocrinol. 2013, 193, 193–200. [Google Scholar] [CrossRef] [PubMed]
- Kawakami, N.; Miyoshi, K.; Horio, S.; Fukui, H. β2-Adrenergic Receptor-Mediated Histamine H1 Receptor Down-Regulation: Another Possible Advantage of β2 Agonists in Asthmatic Therapy. J. Pharmacol. Sci. 2004, 94, 449–458. [Google Scholar] [CrossRef] [PubMed]
- Drake, M.T.; Shenoy, S.K.; Lefkowitz, R.J. Trafficking of G Protein–Coupled Receptors. Circ. Res. 2006, 99, 570–582. [Google Scholar] [CrossRef]
- Chen, J.-F.; Sonsalla, P.K.; Pedata, F.; Melani, A.; Domenici, M.R.; Popoli, P.; Geiger, J.; Lopes, L.V.; de Mendonça, A. Adenosine A2A receptors and brain injury: Broad spectrum of neuroprotection, multifaceted actions and “fine tuning” modulation. Prog. Neurobiol. 2007, 83, 310–331. [Google Scholar] [CrossRef] [PubMed]
- Duan, W.; Gui, L.; Zhou, Z.; Liu, Y.; Tian, H.; Chen, J.-F.; Zheng, J. Adenosine A2A receptor deficiency exacerbates white matter lesions and cognitive deficits induced by chronic cerebral hypoperfusion in mice. J. Neurol. Sci. 2009, 285, 39–45. [Google Scholar] [CrossRef] [PubMed]
- Liu, N.; Wang, Y.; Li, T.; Feng, X. G-protein coupled receptors (GPCRs): Signaling pathways, characterization and functions in insect physiology and toxicology. Int. J. Mol. Sci. 2021, 22, 5260. [Google Scholar] [CrossRef]
| Insect Species | Total Number of Genes | Class A Gene Number | Class B Gene Number | Class C Gene Number | Class F Gene Number | Source of Genome Information | Reference |
|---|---|---|---|---|---|---|---|
| D. melanogaster | ~200 | >70 | ~20 | ~5 | ~5 | https://flybase.org/ (accessed on 7 May 2021) | [84,85] |
| An. gambiae | ~276 | 81 | 21 | 8 | ~8 | https://www.ncbi.nlm.nih.gov/genome/46?genome_assembly_id=22679 (accessed on 7 May 2021) | [86] |
| Ae. aegypti | 135 | 89 | 24 | 8 | 11 | https://www.ncbi.nlm.nih.gov/genome/44?genome_assembly_id=322291 (accessed on 7 May 2021) | [80] |
| B. mori | ~90 | ~70 | ~7 | ~8 | ~4 | https://www.ncbi.nlm.nih.gov/genome/76?genome_assembly_id=1491718 (accessed on 7 May 2021) | [87] |
| A. mellifera | ~50 | ~31 | ~4 | Not clear | Not clear | https://www.ncbi.nlm.nih.gov/genome/48?genome_assembly_id=403979 (accessed on 7 May 2021) | [17] |
| L. cuprina | 197 | 73 | 18 | 9 | Not clear | https://www.ncbi.nlm.nih.gov/genome/12732?genome_assembly_id=358015 (accessed on 7 May 2021) | [88] |
| Cx. quinquefasciatus | 115 | 52 | 4 | Not clear | Not clear | https://www.ncbi.nlm.nih.gov/genome/393?genome_assembly_id=1502880 (accessed on 7 May 2021) | [42] |
| M. domestica | 94 | 55 | 27 | 4 | Not clear | https://www.ncbi.nlm.nih.gov/genome/14461?genome_assembly_id=44793 (accessed on 7 May 2021) | [72] |
| Receptor Group | Receptor Name | Classes | Species | Function | Reference |
|---|---|---|---|---|---|
| 5-HT receptors | Trica5-HT7 R | Class A | Tribolium castaneum | Insect’s neural processes | [134] |
| Adipokinetic hormone receptor | Akh receptor | Class A | Bactrocera dorsalis | Lipid mobilization | [62] |
| Adipokinetic hormone receptor | Akh receptor | Class A | D. melanogaster | Lipid mobilization | [51] |
| Adipokinetic hormone receptor | Akh receptor | Class A | Nilaparvata lugens | Lipid mobilization | [53] |
| Allatostatin receptor | AstAR1 | Class A | D. melanogaster | Metamorphosis | [98] |
| Allatostatin receptor | DAR-1/DAR-2 | Class A | D. melanogaster | Feeding modulation | [99] |
| Allatostatin receptor | Dippu-AstR | Class A | Diploptera punctata | Juvenile hormone synthesis | [128] |
| Arginine vasopressin-like receptor | AVPL receptor | Class A | T. castaneum | Diuretic signaling pathway | [92] |
| Calcitonin receptors | GPCRCAL1 | Class A | Ae. aegypti | primary urine secretion | [66] |
| CCHa2 receptor | CCHa2-R | Class A | D. melanogaster | Insulin production | [105] |
| Diapause hormone receptor | DH-R | Class A | Ae. aegypti | Development | [39] |
| Diapause hormone receptor | Bommo-DHR | Class A | B. mori | Development | [45] |
| Diapause hormone receptor | HzDHr | Class A | Helicoverpa zea | Development | [41] |
| Dopamine receptor | Dop1R2, DmDopEcR | Class A | D. melanogaster | Morphogenesis | [34,97] |
| Dopamine receptor | DopEcR | Class A | D. melanogaster | Mushroom and locomotor activity | [127] |
| Dopamine receptor | DopEcR | Class A | D. melanogaster | Ethanol-induced sedation | [68] |
| Dopamine receptor | AipsDopEcR | Class A | Agrotis ipsilon | Sexual activity regulation | [60,126] |
| Dopamine receptor | DopEcR | Class A | Helicoverpa armigera | Morphogenesis | [35] |
| Dopamine receptor | D2R | Class A | T. castaneum | Morphogenesis | [29] |
| Leucokinin receptor | LKr | Class A | D. melanogaster | Feeding modulation | [100,103] |
| Myosuppressin receptors | CG8985/CG13803 | Class A | D. melanogaster | visceral muscles inhibition | [108] |
| Neuropeptide receptors | GPCR-B2 | Class A | B. mori | Ecdysone synthesis | [33] |
| Neuropeptide receptors | Schgr-sNPFR | Class A | Schistocerca gregaria | Feeding behavior | [50] |
| Neuropeptide Drosulfakinin receptor | CCKLR-17D1 | Class A | D. melanogaster | Fighting behavior | [104] |
| Orphan receptor | DLGR2 | Class A | D. melanogaster | Bursicon bioactivity | [25] |
| Orphan receptor | BNGR-A4 receptor | Class A | B. mori | Food intake and growth | [130] |
| Rhodopsin receptors | Rh2 | Class A | T. castaneum | Reproduction | [22] |
| Sex peptide receptor | SPR | Class A | D. melanogaster | Reproductive behavior | [101] |
| SIFamide receptor | SIFaR | Class A | D. melanogaster | Reproductive behavior | [102] |
| Tyramine receptor | TAR1 | Class A | Rhipicephalus (Boophilus) microplus | Development of antiparasitic | [131] |
| Corticotropin releasing factor receptor | CG12370 | Class B | D. melanogaster | Water balance | [113] |
| Diuretic hormone receptors | DH31R | Class B | D. melanogaster | temperature regulation and homeostasis | [114] |
| Methuselah receptor | mth | Class B | D. melanogaster | Oxidative stress resistance | [136] |
| Methuselah receptor | Ldmthl1 | Class B | Lymantria dispar | Insect longevity | [135] |
| Metabotropic GABA receptors | D-GABABR1, R2 and R3 | Class C | D. melanogaster | Central nervous system | [118] |
| Receptor Name | Gene | Class | Species | Insecticide | References |
|---|---|---|---|---|---|
| Calcitonin receptor | CPIJ014419 | Class A | Cx. quinquefasciatus | Permethrin | [42] |
| Pteropsin | CPIJ014334 | Class A | Cx. quinquefasciatus | Permethrin | [42] |
| Conserved hypothetical protein | CPIJ019111 | Not clear yet | Cx. quinquefasciatus | Permethrin | [42] |
| Leucokinin receptor | LOC101891982 | Class A | M. domestica | Imidacloprid | [72] |
| Opsin receptor | LOC101900880, LOC101900148 | Class A | M. domestica | Imidacloprid | [72] |
| Methuselah-like receptor | LOC101889292, LOC101899380, LOC105262457, LOC101894839 | Class B | M. domestica | Imidacloprid | [72] |
| Dopamine receptor | LOC101896361 | Class A | M. domestica | Imidacloprid | [72] |
| Crustacean cardioactive peptide receptor | LOC101898141 | Class A | M. domestica | Imidacloprid | [72] |
| Methuselah-like GPCR | Ldmthl1 | Class B | L. dispar | Deltamethrin | [135] |
| Arrestin | HQ833831 | Cx. pipiens | Deltamethrin | [137] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, N.; Li, T.; Wang, Y.; Liu, S. G-Protein Coupled Receptors (GPCRs) in Insects—A Potential Target for New Insecticide Development. Molecules 2021, 26, 2993. https://doi.org/10.3390/molecules26102993
Liu N, Li T, Wang Y, Liu S. G-Protein Coupled Receptors (GPCRs) in Insects—A Potential Target for New Insecticide Development. Molecules. 2021; 26(10):2993. https://doi.org/10.3390/molecules26102993
Chicago/Turabian StyleLiu, Nannan, Ting Li, Yifan Wang, and Shikai Liu. 2021. "G-Protein Coupled Receptors (GPCRs) in Insects—A Potential Target for New Insecticide Development" Molecules 26, no. 10: 2993. https://doi.org/10.3390/molecules26102993
APA StyleLiu, N., Li, T., Wang, Y., & Liu, S. (2021). G-Protein Coupled Receptors (GPCRs) in Insects—A Potential Target for New Insecticide Development. Molecules, 26(10), 2993. https://doi.org/10.3390/molecules26102993








