New Cysteine Protease Inhibitors: Electrophilic (Het)arenes and Unexpected Prodrug Identification for the Trypanosoma Protease Rhodesain
Abstract
1. Introduction
2. Results
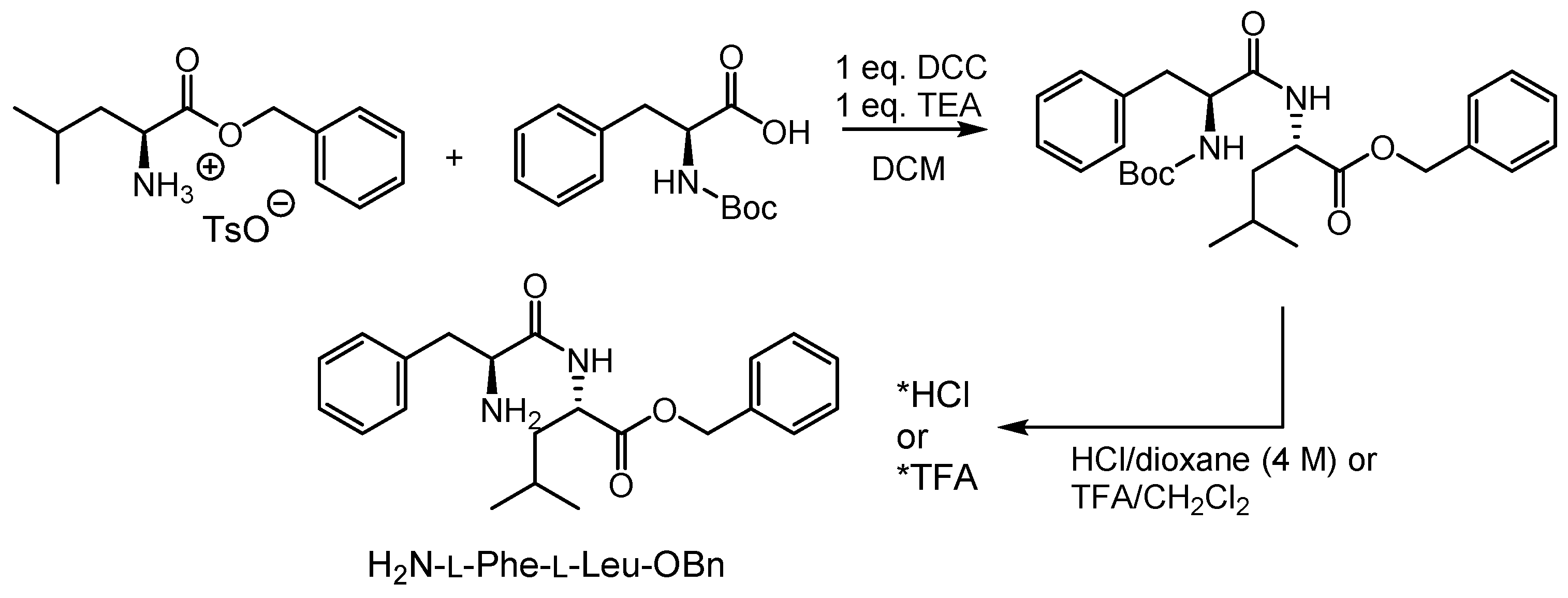
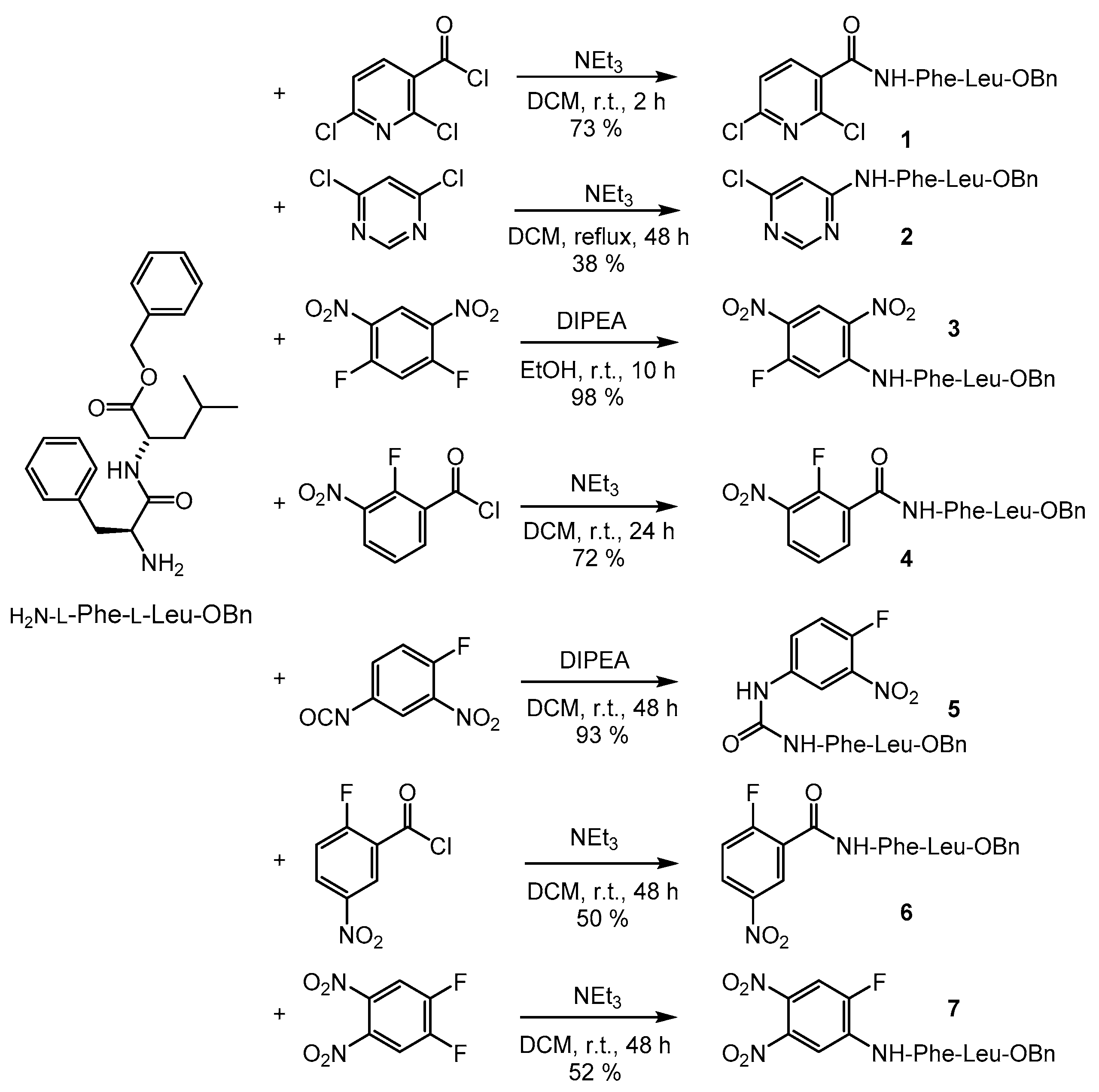
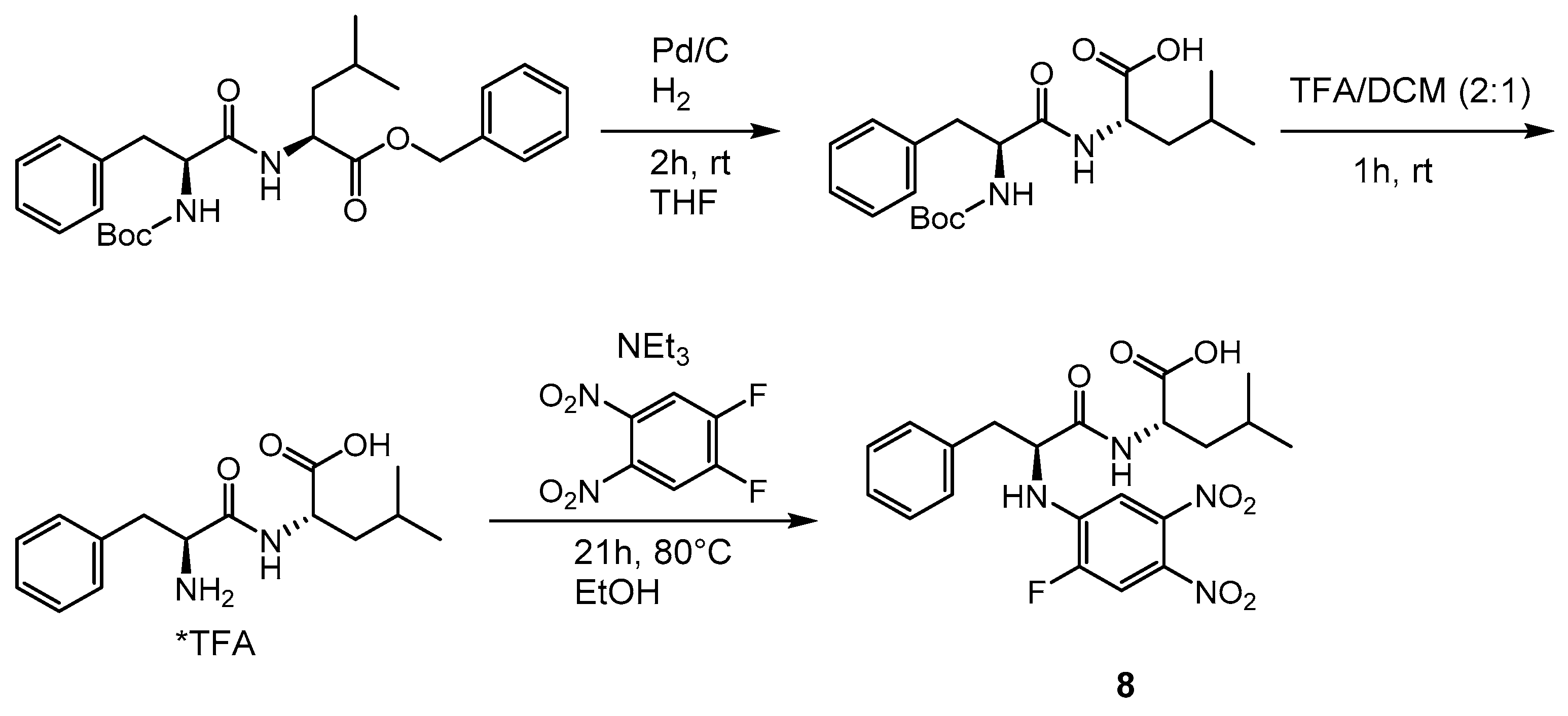
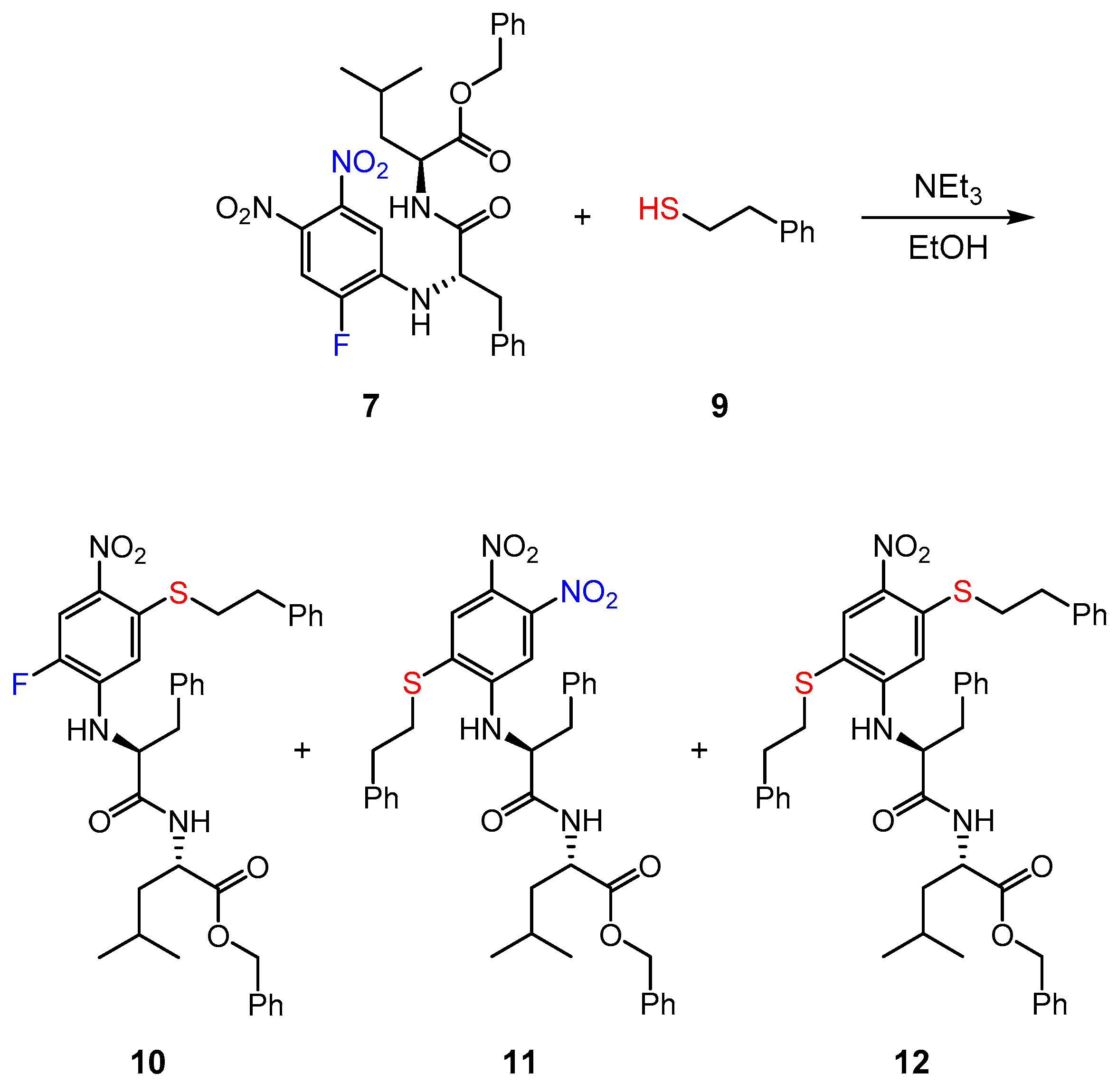
2.1. Syntheses
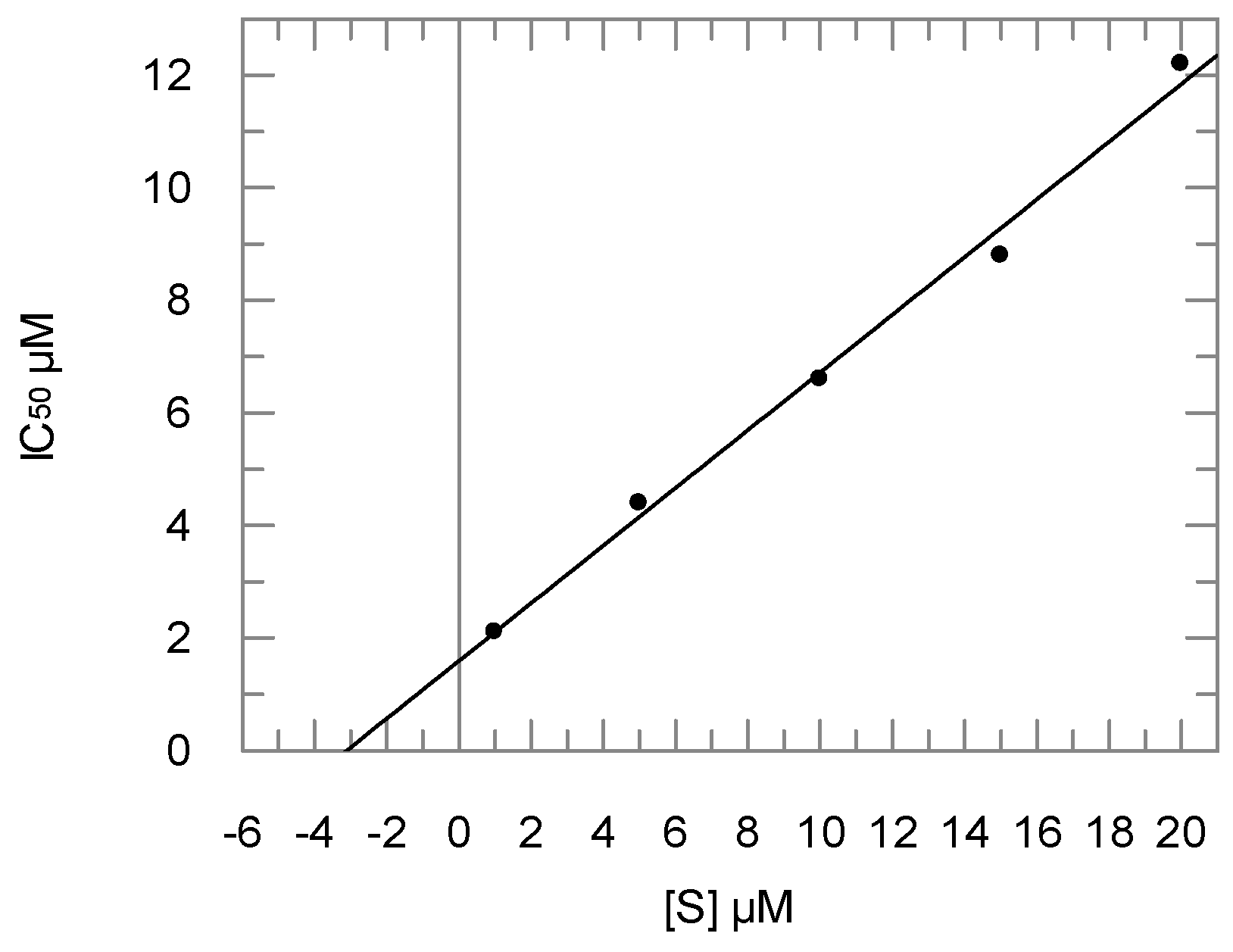
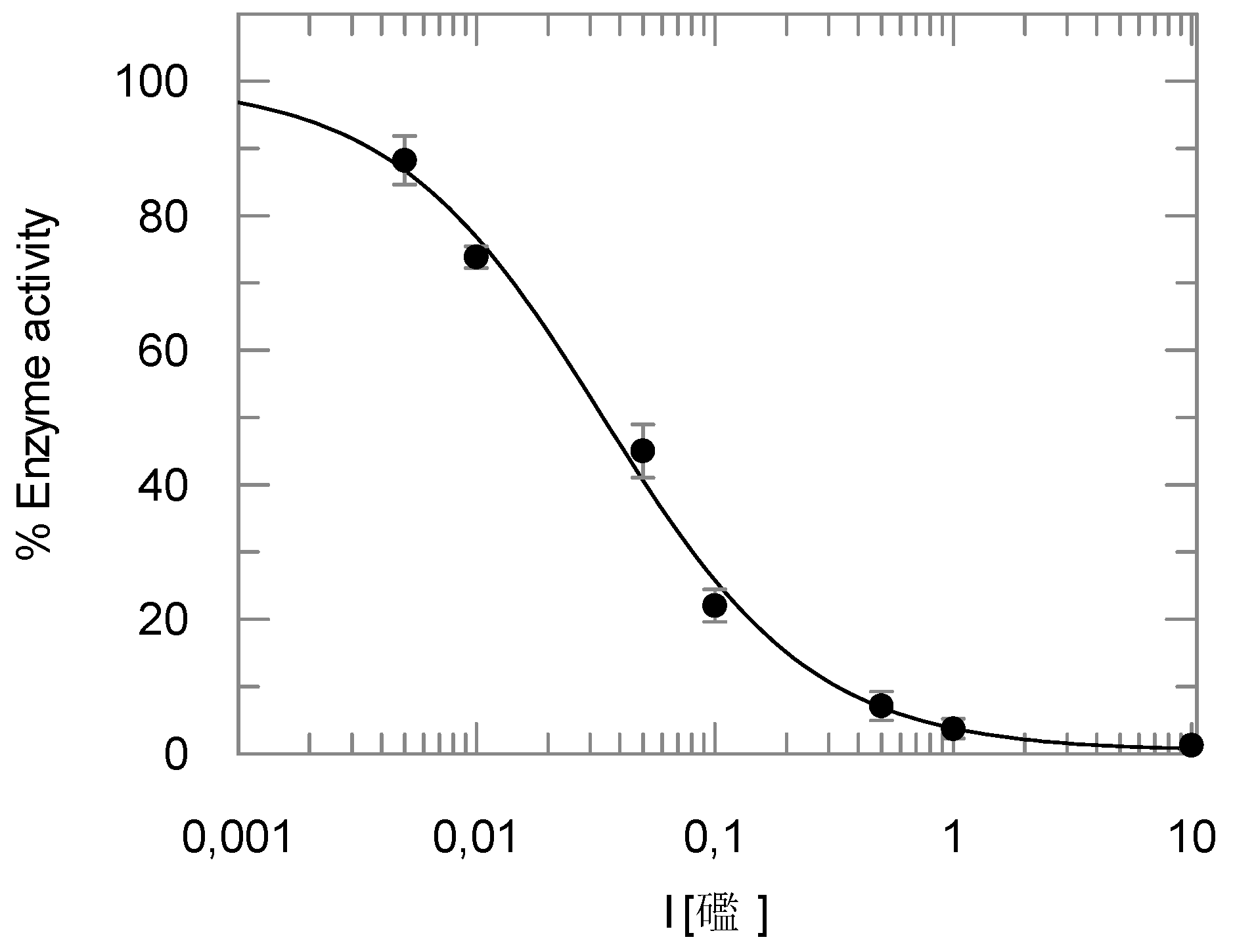
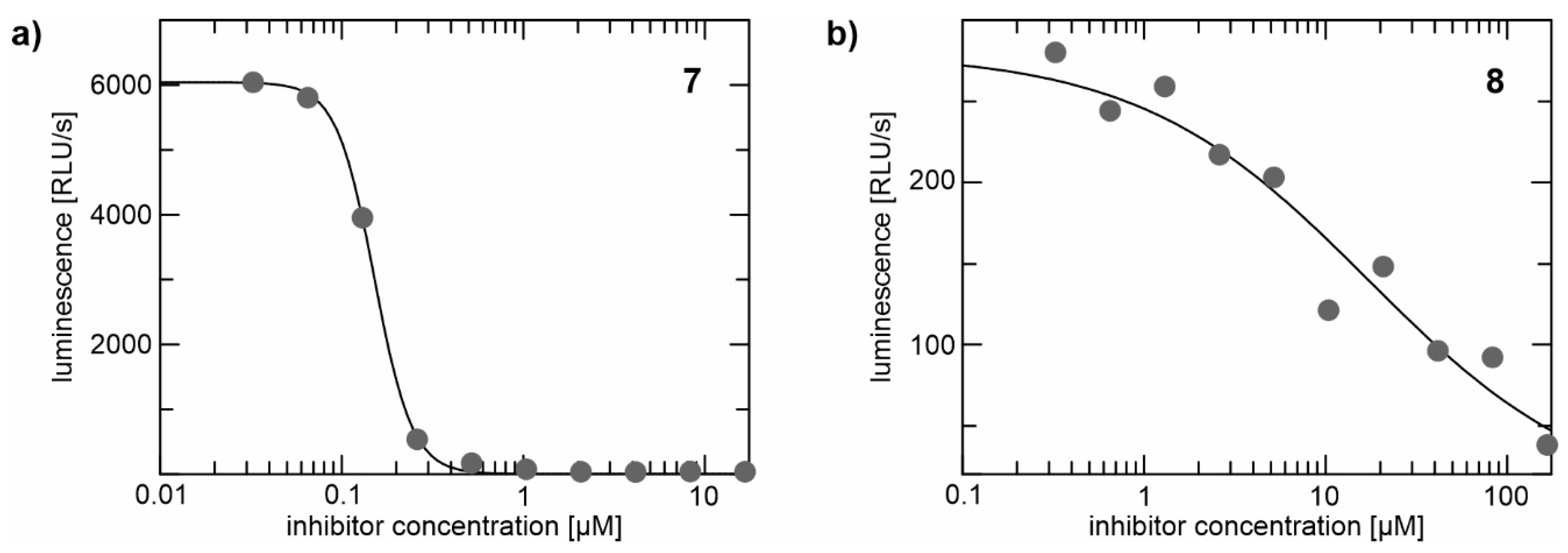
2.2. Enzyme Assays
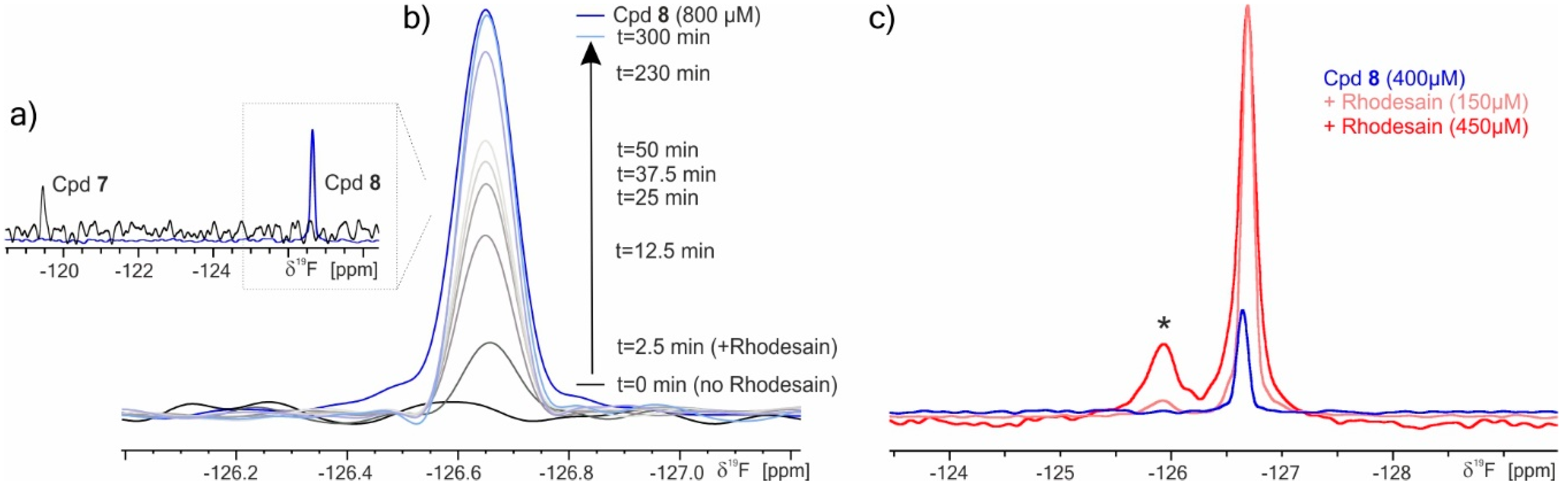
2.3. 19F NMR Spectroscopy
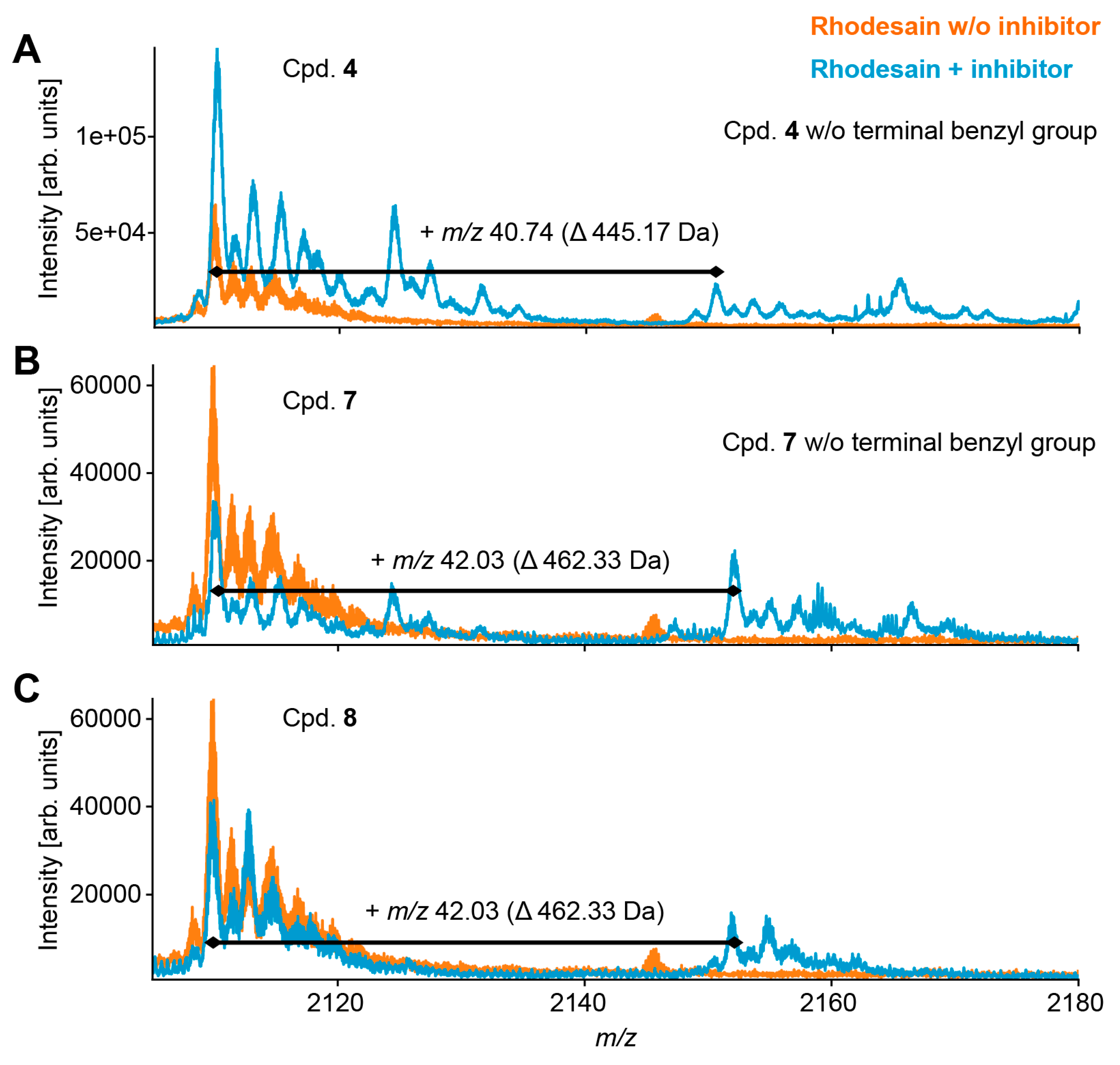
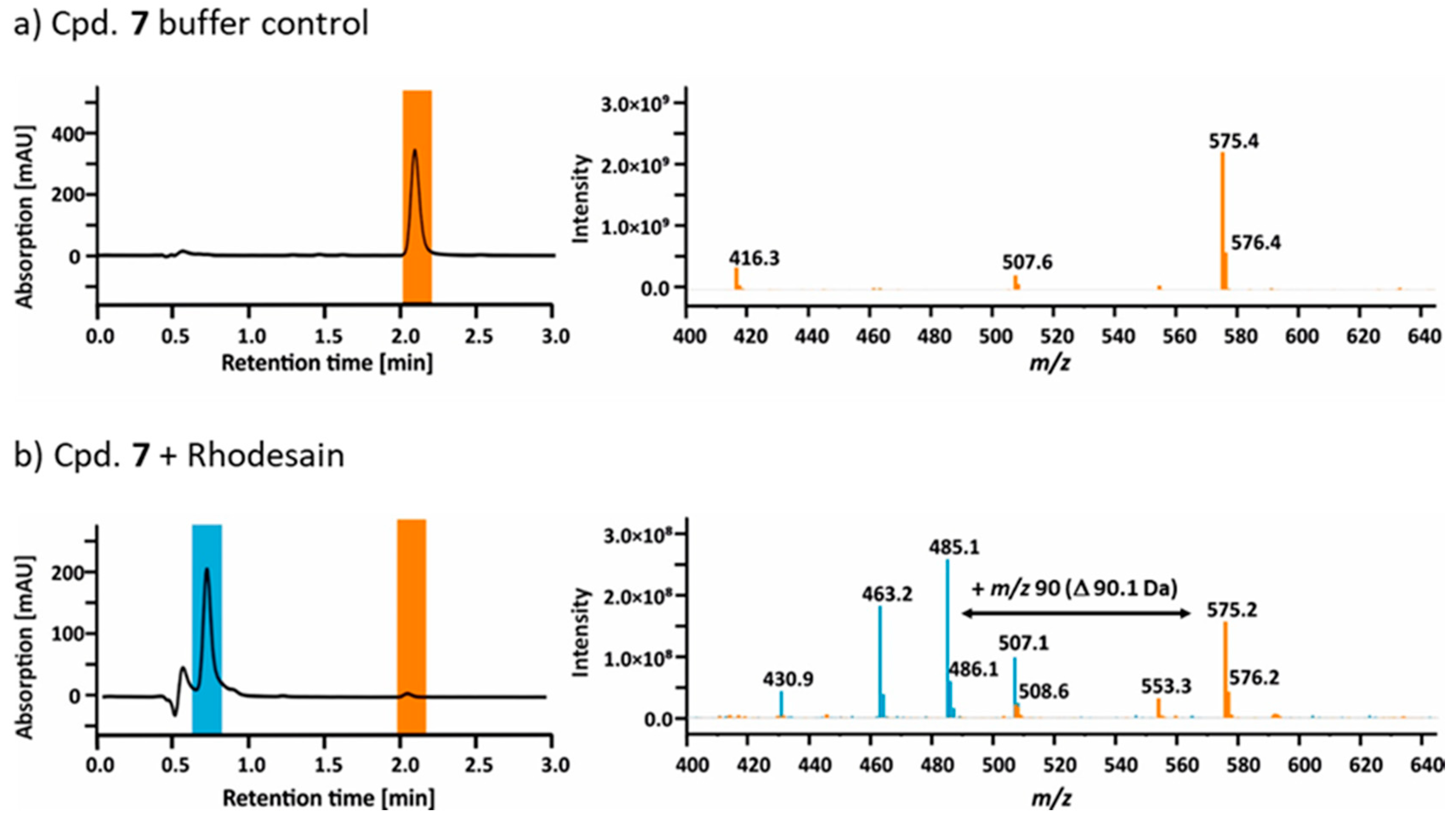
2.4. Mass Spectrometry
2.5. Hydrolysis of Ester 7 by Rhodesain
2.6. Reaction of Inhibitor 7 with a Low Molecular Weight Thiol
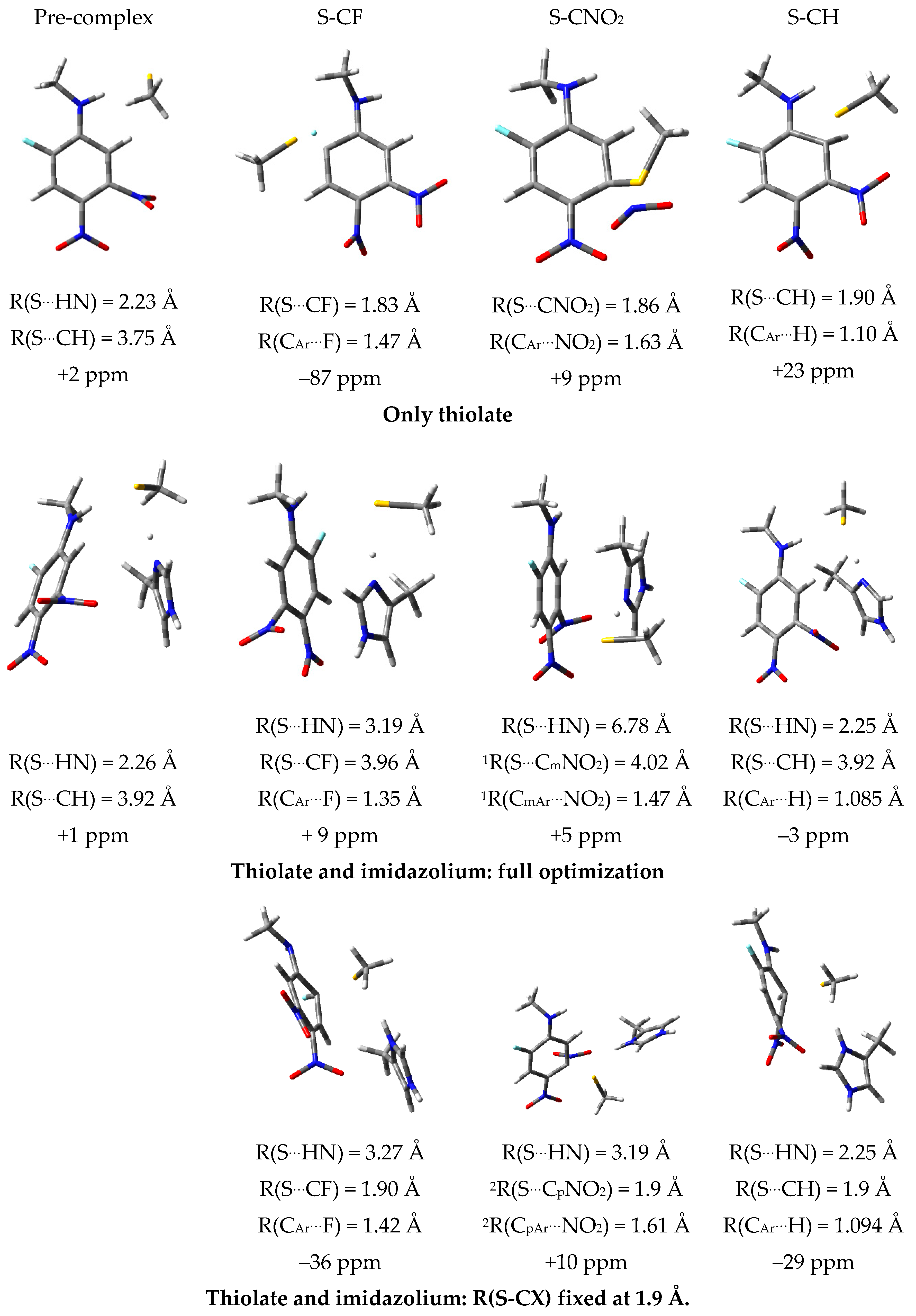
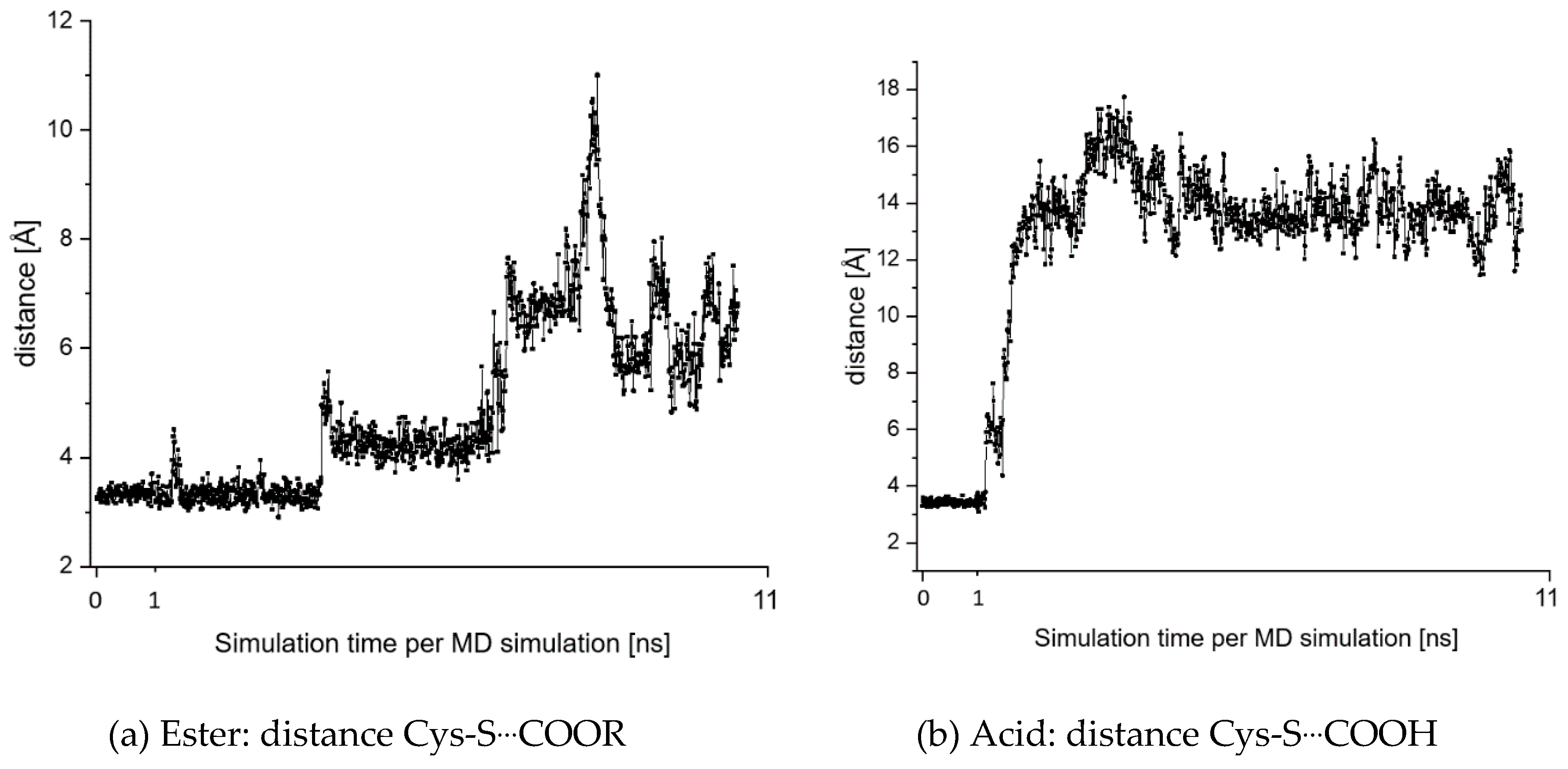
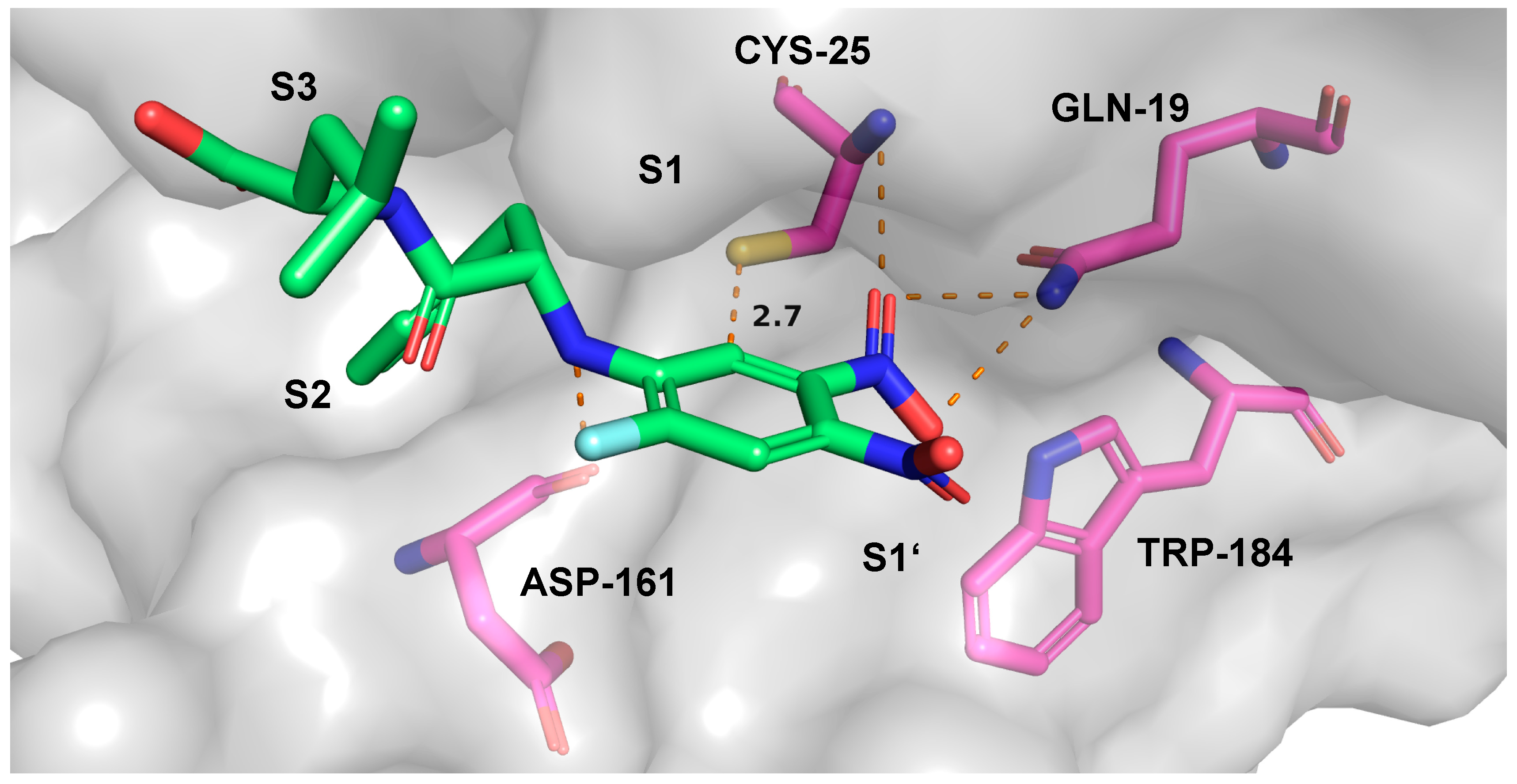
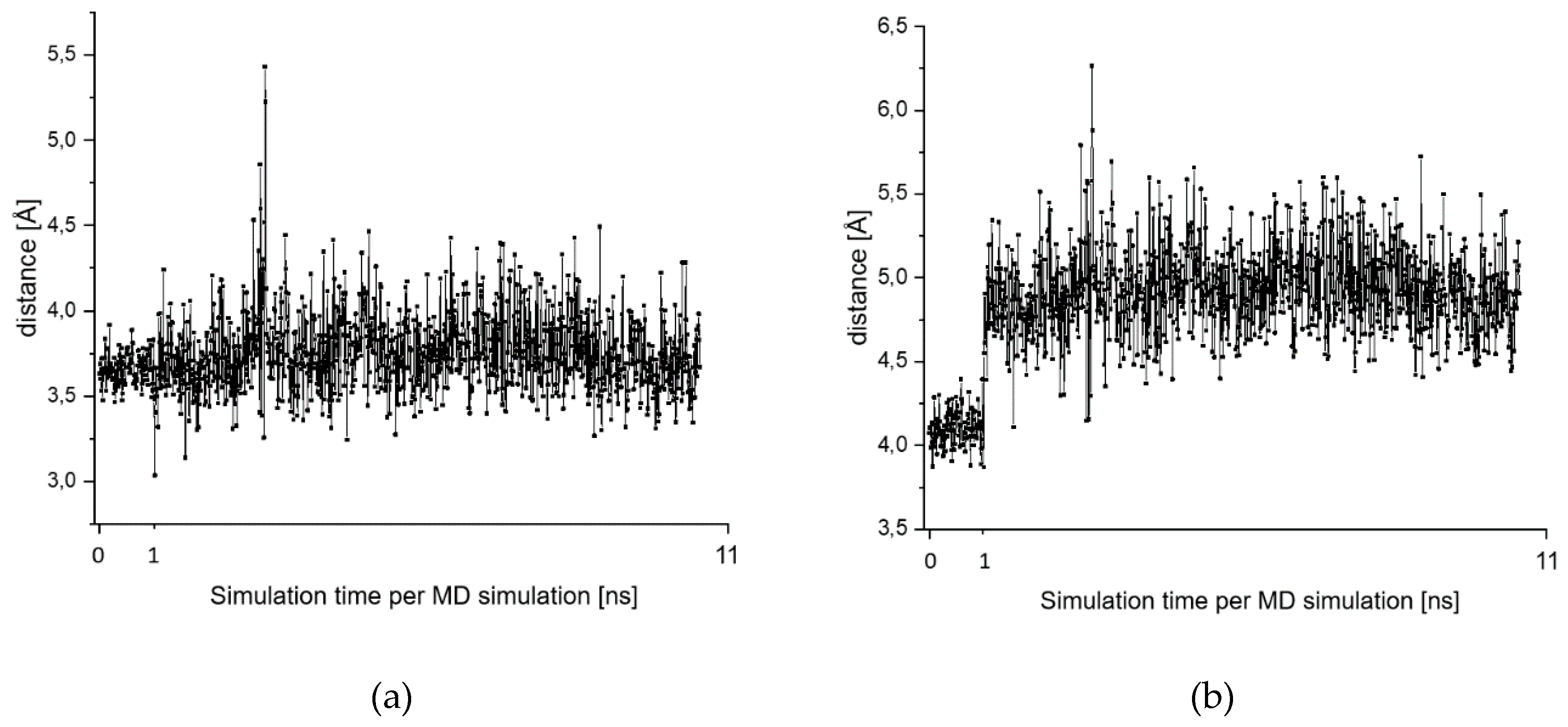
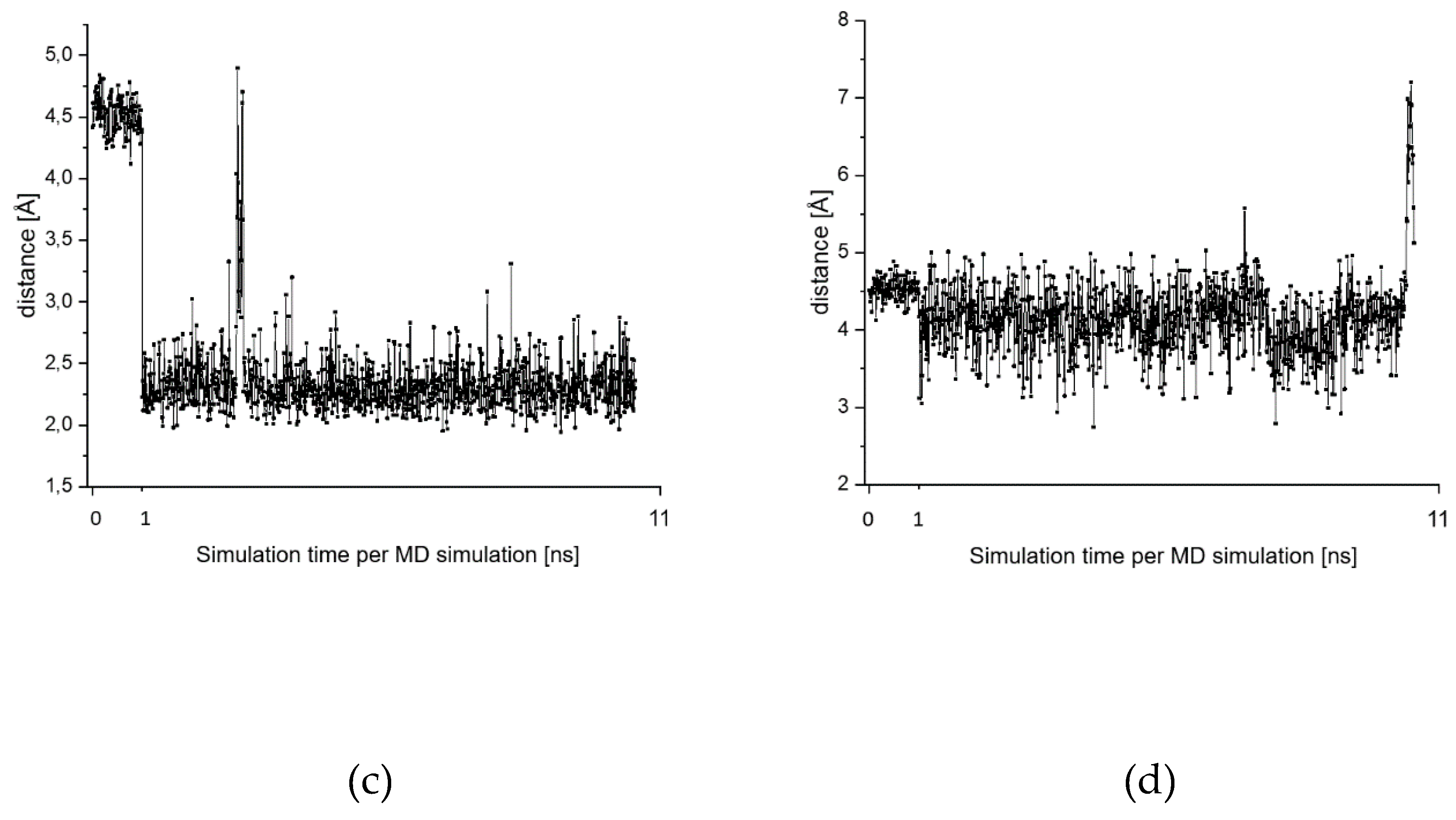
2.7. Theoretical Investigations of the Enzyme-Inhibitor Complex
2.8. T. b. brucei Cell Survival Assay
3. Discussion
4. Materials and Methods
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Bauer, R.A. Covalent inhibitors in drug discovery: From accidental discoveries to avoided liabilities and designed therapies. Drug Discov. Today 2015, 20, 1061–1073. [Google Scholar] [CrossRef] [PubMed]
- Singh, J.; Petter, R.C.; Kluge, A.F. Targeted covalent drugs of the kinase family. Curr. Opin. Chem. Biol. 2010, 14, 475–480. [Google Scholar] [CrossRef] [PubMed]
- Singh, J.; Petter, R.C.; Baillie, T.A.; Whitty, A. The resurgence of covalent drugs. Nat. Rev. Drug Discovery 2011, 10, 307–317. [Google Scholar] [CrossRef] [PubMed]
- Johnson, D.S.; Weerapana, E.; Cravatt, B.F. Strategies for discovering and derisking covalent, irreversible enzyme inhibitors. Future Med. Chem. 2010, 2, 949–964. [Google Scholar] [CrossRef] [PubMed]
- Potashman, M.H.; Duggan, M.E. Covalent modifiers: An orthogonal approach to drug design. J. Med. Chem. 2009, 52, 1231–1246. [Google Scholar] [CrossRef] [PubMed]
- Mah, R.; Thomas, J.R.; Shafer, C.M. Drug discovery considerations in the development of covalent inhibitors. Bioorg. Med. Chem. Lett. 2014, 24, 33–39. [Google Scholar] [CrossRef]
- Baillie, T.A. Targeted covalent inhibitors for drug design. Angew. Chem. Int. Ed. 2016, 55, 13408–13421. [Google Scholar] [CrossRef]
- De Cesco, S.; Kurian, J.; Dufresne, C.; Mittermaier, A.K.; Moitessier, N. Covalent inhibitors design and discovery. Eur. J. Med. Chem. 2017, 10, 96–114. [Google Scholar] [CrossRef]
- Johansson, M.H. Reversible Michael additions: Covalent inhibitors and prodrugs. Mini Rev. Med. Chem. 2012, 12, 1330. [Google Scholar]
- Palmer, J.T.; Rasnick, D.; Klaus, J.L.; Broemme, D. Vinyl sulfones as mechanism-based cysteine protease inhibitors. J. Med. Chem. 1995, 38, 3193–3196. [Google Scholar] [CrossRef]
- Ettari, R.; Nizi, E.; Di Francesco, M.E.; Dude, M.-A.; Pradel, G.; Vicik, R.; Schirmeister, T.; Micale, N.; Grasso, S.; Zappala, M. Development of peptidomimetics with a vinyl sulfone warhead as irreversible falcipain-2 inhibitors. J. Med. Chem. 2008, 51, 988–996. [Google Scholar] [CrossRef] [PubMed]
- Ettari, R.; Bonaccorso, C.; Micale, N.; Heindl, C.; Schirmeister, T.; Calabro, M.L.; Grasso, S.; Zappala, M. Development of novel peptidomimetics containing a vinyl sulfone moiety as proteasome inhibitors. ChemMedChem 2011, 6, 1228–1237. [Google Scholar] [CrossRef] [PubMed]
- Ettari, R.; Cosconati, S.; Amendola, G.; Chouchene, K.; Wagner, A.; Hellmich, U.A.; Ulrich, K.; Krauth-Siegel, R.L.; Wich, P.R.; Schmid, I.; et al. Development of novel peptide-based Michael acceptors targeting rhodesain and falcipain-2 for the treatment of Neglected Tropical Diseases (NTDs). J. Med. Chem. 2017, 60, 6911–6923. [Google Scholar]
- Breuning, A.; Degel, B.; Schulz, F.; Büchold, C.; Stempka, M.; Machon, U.; Gelhaus, C.; Leippe, M.; Leyh, M.; Kisker, C.; et al. Michael acceptor based antiplasmodial and antitrypanosomal cysteine protease inhibitors with unusual amino acids. J. Med. Chem. 2010, 53, 1951–1963. [Google Scholar] [CrossRef] [PubMed]
- Machon, U.; Büchold, C.; Stempka, M.; Schirmeister, T.; Gelhaus, C.; Leippe, M.; Gut, J.; Rosenthal, P.J.; Kisker, C.; Leyh, M.; et al. On-bead screening of a combinatorial fumaric acid derived peptide library yields antiplasmodial cysteine protease inhibitors with unusual peptide sequences. J. Med. Chem. 2009, 52, 5662–5672. [Google Scholar] [CrossRef] [PubMed]
- Ettari, R.; Micale, N.; Schirmeister, T.; Gelhaus, C.; Leippe, M.; Nizi, E.; Di Francesco, M.E.; Grasso, S.; Zappala, M. Novel peptidomimetics containing a vinyl ester moiety as highly potent and selective falcipain-2 inhibitors. J. Med. Chem. 2009, 52, 2157–2160. [Google Scholar] [CrossRef] [PubMed]
- Schirmeister, T.; Kesselring, J.; Jung, S.; Schneider, T.H.; Weickert, A.; Becker, J.; Lee, W.; Bamberger, D.; Wich, P.R.; Distler, U.; et al. Quantum chemical-based protocol for the rational design of covalent inhibitors. J. Am. Chem. Soc. 2016, 138, 8332–8335. [Google Scholar] [CrossRef]
- Ehmke, V.; Quinsaat, J.E.; Rivera-Fuentes, P.; Heindl, C.; Schirmeister, T.; Diederich, F. Tuning and predicting biological affinity: Aryl nitriles as cysteine protease inhibitors. Org. Biomol. Chem. 2012, 10, 5764–5768. [Google Scholar] [CrossRef]
- Ehmke, V.; Winkler, E.; Banner, D.W.; Haap, W.; Schweizer, W.B.; Rottmann, M.; Kaiser, M.; Freymond, C.; Brun, R.; Schirmeister, T.; et al. Optimization of triazine nitriles as rhodesain inhibitors: Structure-activity relationships, bioisosteric imidazopyridine nitriles, and X-ray crystal structure analysis with human cathepsin L. ChemMedChem 2013, 8, 967–975. [Google Scholar] [CrossRef]
- Schirmeister, T.; Schmitz, J.; Jung, S.; Schmenger, T.; Krauth-Siegel, R.L.; Gütschow, M. Evaluation of dipeptide nitriles as inhibitors of rhodesain, a major cysteine protease of Trypanosoma brucei. Bioorg. Med. Chem. Lett. 2016, 27, 45–50. [Google Scholar] [CrossRef]
- Giroud, M.; Ivkovic, J.; Martignoni, M.; Fleuti, M.; Trapp, N.; Haap, W.; Kuglstatter, A.; Benz, J.; Kuhn, B.; Schirmeister, T.; et al. Inhibition of the cysteine protease human cathepsin L by triazine nitriles: Amide⋅⋅⋅heteroarene π-stacking interactions and chalcogen bonding in the S3 pocket. ChemMedChem 2017, 12, 257–270. [Google Scholar] [CrossRef] [PubMed]
- Walker, B.; Lynas, J.F. Strategies for the inhibition of serine proteases. CMLS Cell. Mol. Life Sci. 2001, 58, 596–624. [Google Scholar] [CrossRef] [PubMed]
- Siklos, M.; BenAissa, M.; Thatcher, G.R.J. Cysteine proteases as therapeutic targets: Does selectivity matter? A systematic review of calpain and cathepsin inhibitors. Acta Pharm. Sin. B 2015, 5, 506–519. [Google Scholar] [CrossRef] [PubMed]
- Ettari, R.; Zappala, M.; Evers, A.; Gelhaus, C.; Leippe, M.; Schirmeister, T.; Micale, N.; Grasso, S. Synthesis of novel peptidomimetics as inhibitors of protozoan cysteine proteases falcipain-2 and rhodesain. Eur. J. Med. Chem. 2010, 45, 3228–3233. [Google Scholar] [CrossRef]
- Otto, H.-H.; Schirmeister, T. Cysteine proteases and their inhibitors. Chem. Rev. 1997, 97, 133–171. [Google Scholar] [CrossRef]
- Powers, J.C.; Asgian, J.L.; Ekici, O.D.; James, K.E. Irreversible inhibitors of serine, cysteine, and threonine proteases. Chem. Rev. 2002, 102, 4639–4750. [Google Scholar] [CrossRef]
- Schirmeister, T.; Klockow, A. Cysteine protease inhibitors containing small rings. Mini Rev. Med. Chem. 2003, 3, 585–596. [Google Scholar] [CrossRef]
- Vicik, R.; Helten, H.; Schirmeister, T.; Engels, B. Rational design of aziridine containing cysteine protease inhibitors with improved potency - Studies on inhibition mechanism. ChemMedChem 2006, 1, 1021–1028. [Google Scholar] [CrossRef]
- Park, T.; Yang, C.; Yu, Z. Specific inhibitors of lysozyme and peptidases inhibit the growth of the rumen protozoan Entodinium caudatum without decreasing feed digestion or fermentation in vitro. J. Appl. Microbiol. 2019, 127, 670–682. [Google Scholar] [CrossRef]
- Hartmann, K.; Mielczarek, P.; Silberring, J. Synthesis of the novel covalent cysteine proteases inhibitor with iodacetic functional group. Molecules 2020, 25, 813. [Google Scholar] [CrossRef]
- Shannon, D.A.; Banerjee, R.; Webster, E.R.; Bak, D.W.; Wang, C.; Weerapana, E. Investigating the proteome reactivity and selectivity of aryl halides. J. Am. Chem. Soc. 2014, 136, 3330–3333. [Google Scholar] [CrossRef] [PubMed]
- Leesnitzer, L.M.; Parks, D.J.; Bledsoe, R.K.; Cobb, J.E.; Collins, J.L.; Consler, T.G.; Davis, R.G.; Hull-Ryde, E.A.; Lenhard, J.M.; Patel, L.; et al. Functional consequences of cysteine modification in the ligand binding sites of peroxisome proliferator activated receptors by GW9662. Biochemistry 2002, 41, 6640–6650. [Google Scholar] [CrossRef] [PubMed]
- Gehringer, M.; Laufer, S.A. Emerging and re-emerging warheads for Targeted Covalent Inhibitors: Applications in medicinal chemistry and chemical biology. J. Med. Chem. 2019, 62, 5673–5724. [Google Scholar] [CrossRef] [PubMed]
- Spokoyny, A.M.; Zou, Y.; Ling, J.J.; Yu, H.; Lin, Y.-S.; Pentelute, B.L.J. A perfluoroaryl-cysteine SNAr chemistry approach to unprotected peptide stapling. J. Am. Chem. Soc. 2013, 135, 5946–5949. [Google Scholar] [CrossRef]
- Dolle, R.A.; Hoyer, D.; Rinker, J.M.; Ross, T.M.; Schmidt, S.J.; Helaszek, C.T.; Ator, M.A. 3-Chloro-4-carboxamido-6-arylpyridazines as non-peptide class of interleukin-1β converting enzyme inhibitor. Bioorg. Med. Chem. Lett. 1997, 7, 1003–1006. [Google Scholar] [CrossRef]
- Shie, J.-J.; Fang, J.-M.; Kuo, C.-J.; Kuo, T.-H.; Liang, P.-H.; Huang, H.-J.; Yang, W.-B.; Lin, C.-H.; Chen, J.-L.; Wu, Y.-T.; et al. Discovery of potent anilide inhibitors against the Severe Acute Respiratory Syndrome 3CL protease. J. Med. Chem. 2005, 48, 4469–4473. [Google Scholar] [CrossRef]
- Li, Z.; Brecher, M.; Deng, Y.-Q.; Zhang, J.; Sakamuru, S.; Liu, B.; Huang, R.; Koetzner, C.A.; Allen, C.A.; Jones, S.A.; et al. Existing drugs as broad-spectrum and potent inhibitors for Zika virus by targeting NS2B-NS3 interaction. Cell Res. 2017, 27, 1046–1064. [Google Scholar] [CrossRef]
- Olson, O.C. Cysteine cathepsin proteases: Regulators of cancer progression and therapeutic response. Nat. Rev. Cancer 2015, 15, 712–729. [Google Scholar] [CrossRef]
- Mohamed, M.M. Cysteine cathepsins: Multifunctional enzymes in cancer. Nat. Rev. Cancer 2006, 6, 764–775. [Google Scholar] [CrossRef]
- Ferreira, L.G.; Andricopulo, A.D. Targeting cysteine proteases in trypanosomatid disease drug discovery. Pharmacol. Ther. 2017, 180, 49–61. [Google Scholar] [CrossRef]
- Ettari, R.; Previti, S.; Tamborini, L.; Cullia, G.; Grasso, S.; Zappala, M. The inhibition of cysteine proteases rhodesain and TbCatB: A valuable approach to treat Human African Trypanosomiasis. Mini Rev. Med. Chem. 2016, 16, 1374–1391. [Google Scholar] [CrossRef] [PubMed]
- Vicik, R.; Busemann, M.; Gelhaus, C.; Stiefl, N.; Scheiber, J.; Schmitz, W.; Schulz, F.; Mladenovic, M.; Engels, B.; Leippe, M.; et al. Aziridide-based inhibitors of cathepsin L: Synthesis, inhibition activity, and docking studies. ChemMedChem 2006, 1, 1126–1141. [Google Scholar] [CrossRef] [PubMed]
- Barthels, F.; Marincola, G.; Marciniak, T.; Konhäuser, M.; Hammerschmidt, S.; Bierlmeier, J.; Distler, U.; Wich, P.R.; Tenzer, S.; Schwarzer, D.; et al. Asymmetric disulfanylbenzamides as irreversible and selective inhibitors of Staphylococcus aureus sortase A. ChemMedChem 2020. accepted. [Google Scholar] [CrossRef] [PubMed]
- Dixon, M. The determination of enzyme inhibitor constants. Biochem. J. 1953, 55, 170–171. [Google Scholar] [CrossRef]
- Dixon, M. The graphical determination of Km and Ki. Biochem. J. 1972, 129, 197–202. [Google Scholar] [CrossRef]
- Cheng, Y.-C.; Prusoff, W.H. Relationship between the inhibition constant (Ki) and the concentration of inhibitor which causes 50 percent inhibition (I50) of an enzymatic reaction. Biochem. Pharmacol. 1973, 22, 3099–3108. [Google Scholar]
- Nitsche, C.; Holloway, S.; Schirmeister, T.; Klein, C.D. Biochemistry and medicinal chemistry of the Dengue virus protease. Chem. Rev. 2014, 114, 11348–11381. [Google Scholar] [CrossRef]
- Millies, B.; Hammerstein, v.F.; Gellert, A.; Hammerschmidt, S.; Barthels, F.; Göppel, U.; Immerheiser, M.; Elgner, F.; Jung, N.; Basic, M.; et al. Proline-based allosteric inhibitors of Zika and Dengue virus NS2B/NS3 proteases. J. Med. Chem. 2019, 62, 11359–11382. [Google Scholar] [CrossRef]
- Barrett, A.J.; Woessner, J.F.; Rawlings, N.D. (Eds.) Handbook of Proteolytic Enzymes; Elsevier: London, UK, 2012; Volume 2, 984p, Available online: https://www.sciencedirect.com/book/9780120796113/handbook-of-proteolytic-enzymes (accessed on 23 March 2020).
- Kerr, I.D.; Lee, J.H.; Farady, C.J.; Marion, R.; Rickert, M.; Sajid, M.; Pandey, K.C.; Caffrey, C.R.; Legac, J.; Hansell, E.; et al. Vinyl sulfones as antiparasitic agents and a structural basis for drug design. J. Biol. Chem. 2009, 284, 25697–25703. [Google Scholar] [CrossRef]
- Stewart, J.J.P. Optimization of parameters for semiempirical methods V: Modification of NDDO approximations and application to 70 elements. J. Mol. Model. 2007, 13, 1173–1213. [Google Scholar] [CrossRef]
- Halgren, T.A. Merck molecular force field. I. Basis, form, scope, parameterization and performance of MMFF94. J. Comput. Chem. 1996, 17, 490–519. [Google Scholar]
- Vosko, S.H.; Wilk, L.; Nusair, M. Accurate spin-dependent electron liquid correlation energies for local spin density calculations: A critical analysis. Can. J. Phys. 1980, 58, 1200–1211. [Google Scholar] [CrossRef]
- Lee, C.; Yang, W.; Parr, R.G. Development of the Colle-Salvetti correlation-energy formula into a functional of the electron density. Phys. Rev. B 1988, 37, 785–789. [Google Scholar] [CrossRef] [PubMed]
- Becke, A.D. Density-functional thermochemistry. III. The role of exact exchange. J. Chem. Phys. 1993, 98, 5648–5652. [Google Scholar] [CrossRef]
- Stephens, P.J.; Devlin, F.J.; Chabalowski, C.F.; Frisch, M.J. Ab initio calculation of vibrational absorption and circular dichroism spectra using density functional force fields. J. Phys. Chem. 1994, 98, 11623–11627. [Google Scholar] [CrossRef]
- Krishnan, R.; Binkley, J.S.; Seeger, R.; Pople, J.A. Self-consistent molecular orbital methods. XX. A basis set for correlated wave functions. J. Chem. Phys. 1980, 72, 650–654. [Google Scholar] [CrossRef]
- Clark, T.; Chandrasekhar, J.; Spitznagel, G.W.; Schleyer, P.v.R. Efficient diffuse function-augmented basis-sets for anion calculations. 3. The 3-21+G basis set for 1st-row elements, Li-F. J. Comput. Chem. 1983, 4, 294–301. [Google Scholar]
- Frisch, M.J.; Pople, J.A.; Binkley, J.S. Self-consistent molecular orbital methods 25. Supplementary functions for Gaussian basis sets. J. Chem. Phys. 1984, 80, 3265–3269. [Google Scholar] [CrossRef]
- Adamo, C.; Barone, V. Exchange functionals with improved long-range behavior and adiabatic connection methods without adjustable parameters: The mPW and mPW1PW models. J. Chem. Phys. 1998, 108, 664–675. [Google Scholar] [CrossRef]
- Ditchfield, R. Self-consistent perturbation theory of diamagnetism. Mol. Physics 1974, 27, 789–807. [Google Scholar] [CrossRef]
- Tomasi, J.; Mennucci, B.; Cancès, E. The IEF version of the PCM solvation method: An overview of a new method addressed to study molecular solutes at the QM ab initio level. J. Mol. Struct.: Theochem 1999, 464, 211–226. [Google Scholar] [CrossRef]
- Grimblat, N.; Zanardi, M.M.; Sarotti, A.M. Beyond DP4: An improved probability for the stereochemical assignment of isomeric compounds using quantum chemical calculations of NMR shifts. J. Org. Chem. 2015, 80, 12526–12534. [Google Scholar] [CrossRef] [PubMed]
- Helten, H.; Schirmeister, T.; Engels, B. Theoretical studies about the influence of different ring substituents on the nucleophilic ring opening of three-membered heterocycles and possible implications for the mechanisms of cysteine protease inhibitors. J. Org. Chem. 2005, 70, 233–237. [Google Scholar] [CrossRef] [PubMed]
- Helten, H.; Schirmeister, T.; Engels, B. Model calculations about the influence of protic environments on the alkylation step of epoxide, aziridine, and thiirane based cysteine protease inhibitors. J. Phys. Chem. A 2004, 108, 7691–7701. [Google Scholar] [CrossRef]
- Buback, V.; Mladenovic, M.; Engels, B.; Schirmeister, T. Rational design of improved aziridine-based inhibitors of cysteine proteases. J. Phys. Chem. B 2009, 113, 5282–5289. [Google Scholar] [CrossRef]
- Chai, J.-D.; Head-Gordon, M. Long-Range Corrected Hybrid Density Functionals with Damped Atom−Atom Dispersion Corrections. Phys. Chem. Chem. Phys. 2008, 10, 6615–6621. [Google Scholar] [CrossRef]
- Grimme, S. Improved second-order Moller-Plesset perturbation theory by separate scaling of parallel- and antiparallel-spin pair correlation energies. J. Phys. Chem. 2003, 118, 9095–9102. [Google Scholar] [CrossRef]
- Gaussian 16, Revision A.03; Gaussian, Inc.: Wallingford, CT, USA, 2016.
- LeadIT/FlexX. Version 2.3.2; BioSolveIT GmbH: St. Augustin, Germany, 2018. [Google Scholar]
- Mladenovic, M.; Junold, K.; Fink, R.F.; Thiel, W.; Schirmeister, T.; Engels, B. Atomistic insights into the inhibition of cysteine proteases: First QM/MM calculations clarifying the regiospecificity and the inhibition potency of epoxide- and aziridine-based inhibitors. J. Phys. Chem. B 2008, 112, 5458–5469. [Google Scholar] [CrossRef]
- Mladenovic, M.; Schirmeister, T.; Thiel, S.; Thiel, W.; Engels, B. The importance of the active site histidine for the activity of epoxide- or aziridine-based inhibitors of cysteine proteases. ChemMedChem 2007, 2, 120–128. [Google Scholar] [CrossRef]
- Mladenovic, M.; Ansorg, K.; Fink, R.F.; Thiel, W.; Schirmeister, T.; Engels, B. Atomistic Insights into the Inhibition of Cysteine Proteases: First QM/MM Calculations Clarifying the Stereoselectivity of Epoxide-Based Inhibitors. J. Phys. Chem. B 2008, 112, 11798–11808. [Google Scholar] [CrossRef]
- Wagner, A.; Le, T.A.; Brennich, M.; Klein, P.; Bader, N.; Diehl, E.; Paszek, D.; Weickhmann, A.K.; Dirdjaja, N.; Krauth-Siegel, R.L.; et al. Inhibitor-induced dimerization of an essential oxidoreductase from African trypanosomes. Angew. Chem. Int. Ed. 2019, 58, 3640–3644. [Google Scholar] [CrossRef] [PubMed]
- Vicik, R.; Hoerr, V.; Glaser, M.; Schultheis, M.; Hansell, E.; McKerrow, J.H.; Holzgrabe, U.; Caffrey, C.R.; Ponte-Sucre, A.; Moll, H.; et al. Aziridine-2,3-dicarboxylate inhibitors targeting the major cysteine protease of Trypanosoma brucei as lead trypanocidal agents. Bioorg. Med. Chem. Lett. 2006, 16, 2753–2757. [Google Scholar] [CrossRef] [PubMed]
Sample Availability: Samples of the compounds are available from the authors. |


| Cpd. | Cath. L | Cath. B | Rhod. | DENV PR | Sortase A |
|---|---|---|---|---|---|
| % inh./Ki | % inh./Ki | % inh./Ki | % inh./Ki | % inh./Ki | |
| 1 | 57/nd | 16/nd | nd/nd | 19/nd | nd/nd |
| 2 | 56/nd | 33/nd | nd/nd | 24/nd | nd/nd |
| 3 | 60/nd | 22/nd | nd/nd | 28/nd | nd/nd |
| 4 | 99/0.66 | 20/nd | 75/0.29 | 10/nd | 11/nd |
| 5 | 8/nd | 10/nd | nd/nd | 11/nd | nd/nd |
| 6 | 50/nd | 21/nd | nd/nd | 12/nd | nd/nd |
| 7 | 98/1.6 | 63/4.4 | 45/nd | 31/nd | 16/nd |
| Structure | ωB97XD | MP2 | SCS-MP2 |
|---|---|---|---|
| Pre-complex | –8.8/–9.0/ | –12.3/–16.0/- | –10.3/–12.8/ |
| S-CH | +0.8/–11.5/–6.8 | –6.6/–16.0/–12.8 | –4.1/–12.9/–9.3 |
| S-CF | –3.3/–13.1/–3.1 | –13.2/–20.3/–12.5 | –11.1/–16.5/–10.1 |
| S-NO2 | –2.7/–9.3/+3.3 | –11.9/–17.0/–5.3 | –9.1/–13.3/–2.9 |
| Starting Structure | R(S-CNO2) | R(S-CH) | ΔE |
|---|---|---|---|
| MD 6 Frame 963 conformer with short Cys-S…H-N distance | 3.7 | 0.0 | |
| 3.3 | 1.4 | ||
| 3.1 | 1.7 | ||
| 2.9 | 2.5 | ||
| MD 6 Frame 1096 conformer with short Cys-S…H-N distance | 3.7 | 0.0 | |
| 3.3 | 0.7 | ||
| 3.1 | 0.8 | ||
| 2.9 | 2.6 | ||
| MD 6 Frame 542 conformer with short Cys-S…H-N distance | 4.9 | 0.0 | |
| 4.5 | 1.2 | ||
| 4.1 | 3.4 | ||
| 3.9 | 0.4 | ||
| 3.5 | 5.3 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Klein, P.; Johe, P.; Wagner, A.; Jung, S.; Kühlborn, J.; Barthels, F.; Tenzer, S.; Distler, U.; Waigel, W.; Engels, B.; et al. New Cysteine Protease Inhibitors: Electrophilic (Het)arenes and Unexpected Prodrug Identification for the Trypanosoma Protease Rhodesain. Molecules 2020, 25, 1451. https://doi.org/10.3390/molecules25061451
Klein P, Johe P, Wagner A, Jung S, Kühlborn J, Barthels F, Tenzer S, Distler U, Waigel W, Engels B, et al. New Cysteine Protease Inhibitors: Electrophilic (Het)arenes and Unexpected Prodrug Identification for the Trypanosoma Protease Rhodesain. Molecules. 2020; 25(6):1451. https://doi.org/10.3390/molecules25061451
Chicago/Turabian StyleKlein, Philipp, Patrick Johe, Annika Wagner, Sascha Jung, Jonas Kühlborn, Fabian Barthels, Stefan Tenzer, Ute Distler, Waldemar Waigel, Bernd Engels, and et al. 2020. "New Cysteine Protease Inhibitors: Electrophilic (Het)arenes and Unexpected Prodrug Identification for the Trypanosoma Protease Rhodesain" Molecules 25, no. 6: 1451. https://doi.org/10.3390/molecules25061451
APA StyleKlein, P., Johe, P., Wagner, A., Jung, S., Kühlborn, J., Barthels, F., Tenzer, S., Distler, U., Waigel, W., Engels, B., Hellmich, U. A., Opatz, T., & Schirmeister, T. (2020). New Cysteine Protease Inhibitors: Electrophilic (Het)arenes and Unexpected Prodrug Identification for the Trypanosoma Protease Rhodesain. Molecules, 25(6), 1451. https://doi.org/10.3390/molecules25061451





