Biophysical Characterization of Epigallocatechin-3-Gallate Effect on the Cardiac Sodium Channel Nav1.5
Abstract
1. Introduction
2. Results
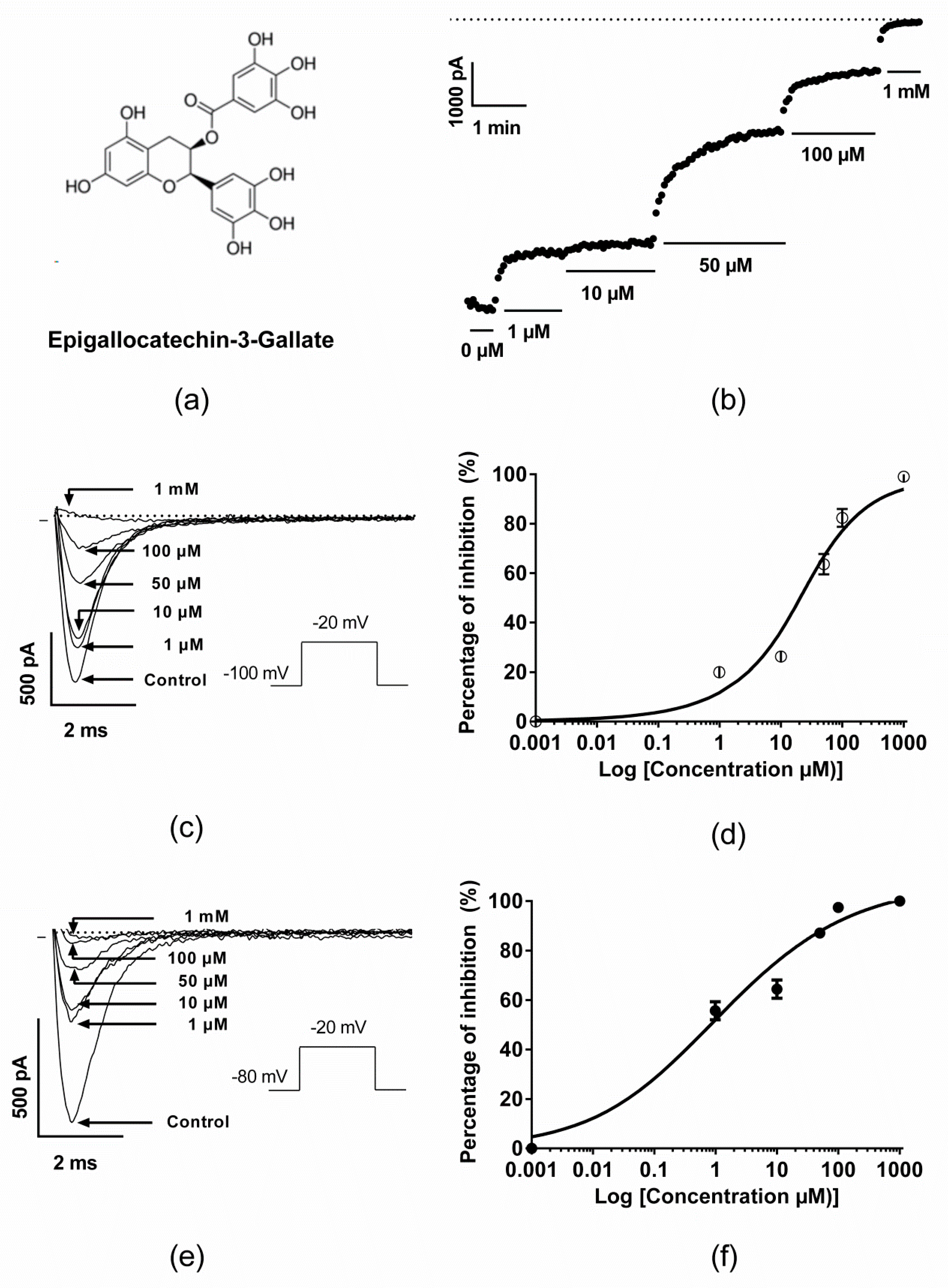
2.1. Inhibitory Effect of EGCG on Human Nav1.5 Channels
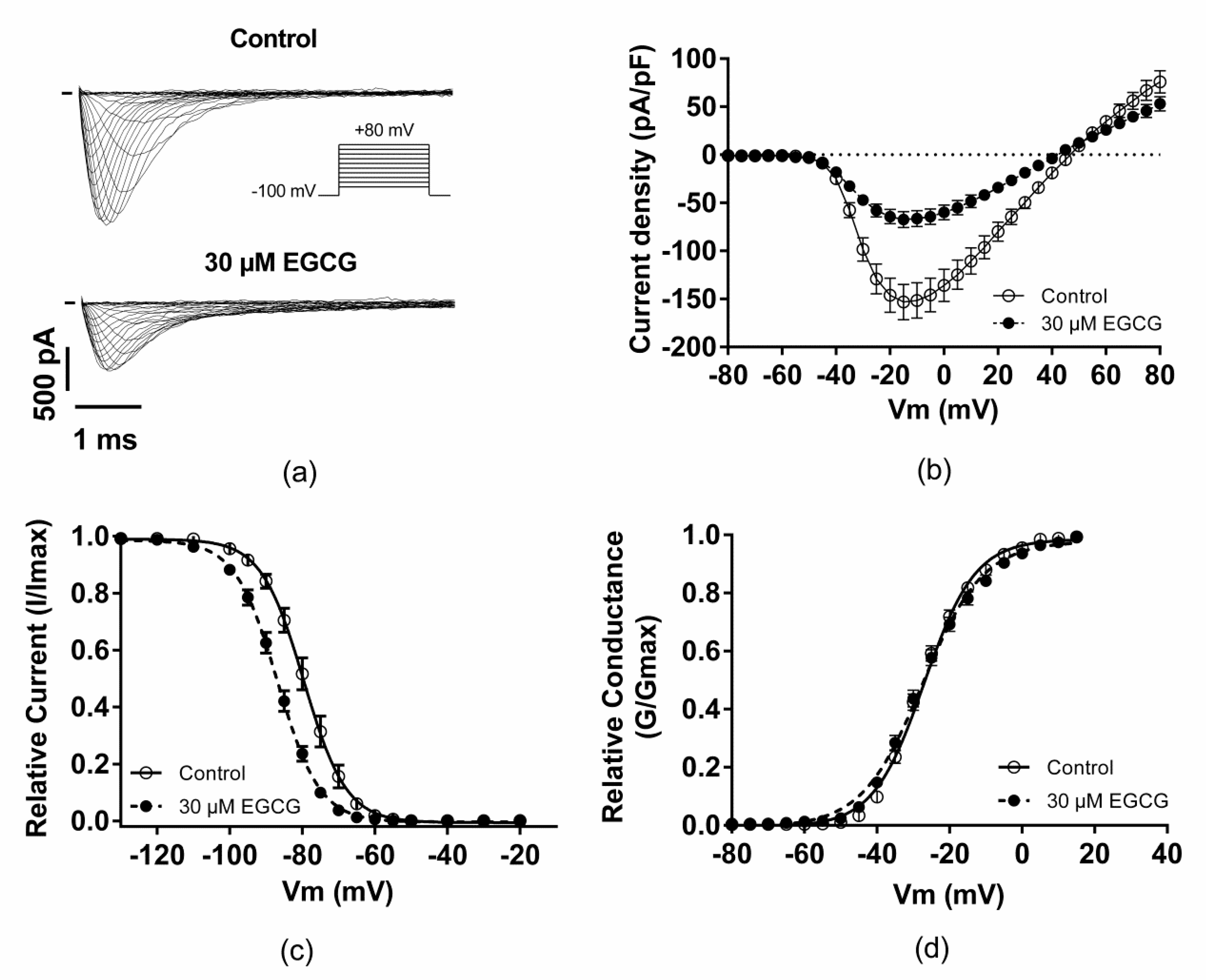
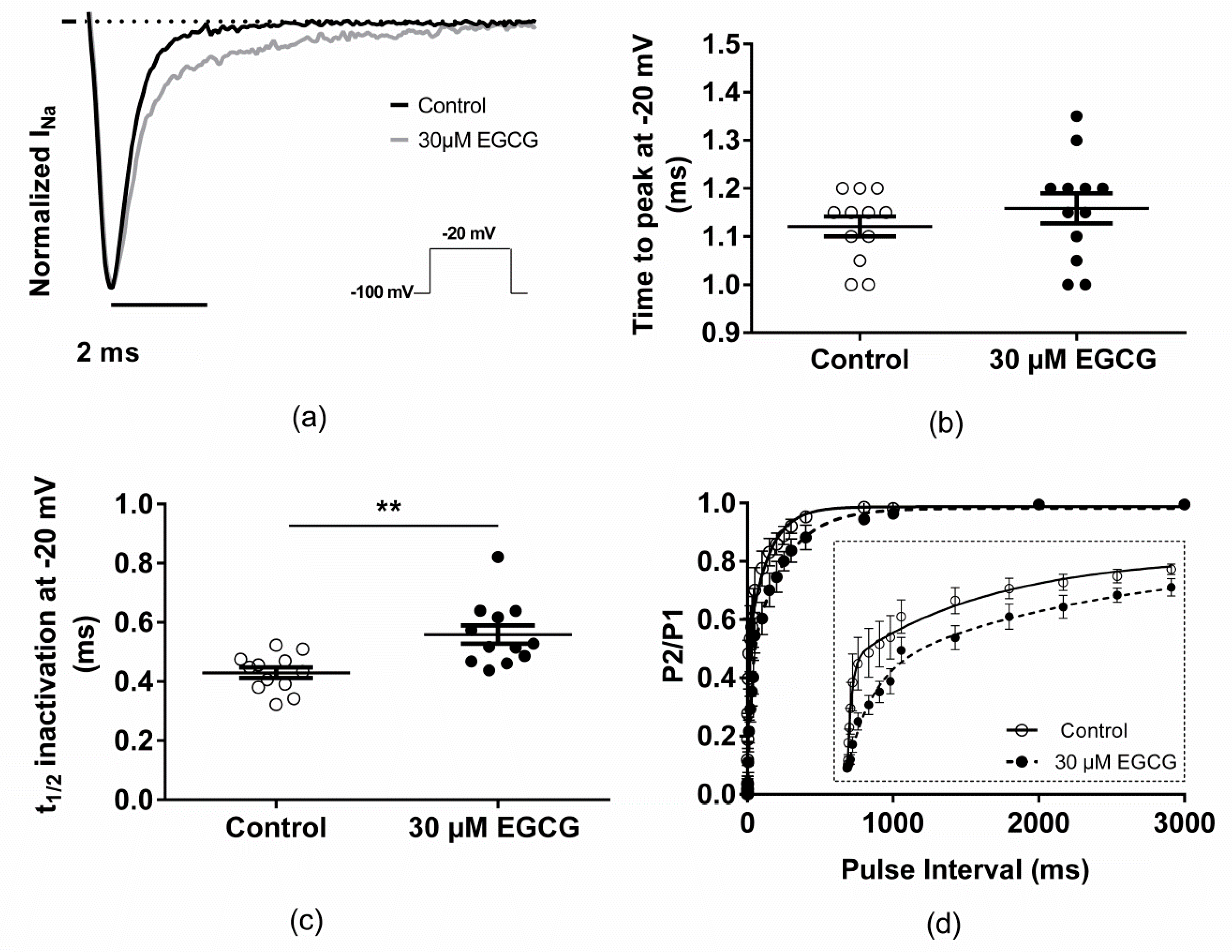
2.2. Effect of EGCG on the Voltage-Dependence Properties of Nav1.5 Channels
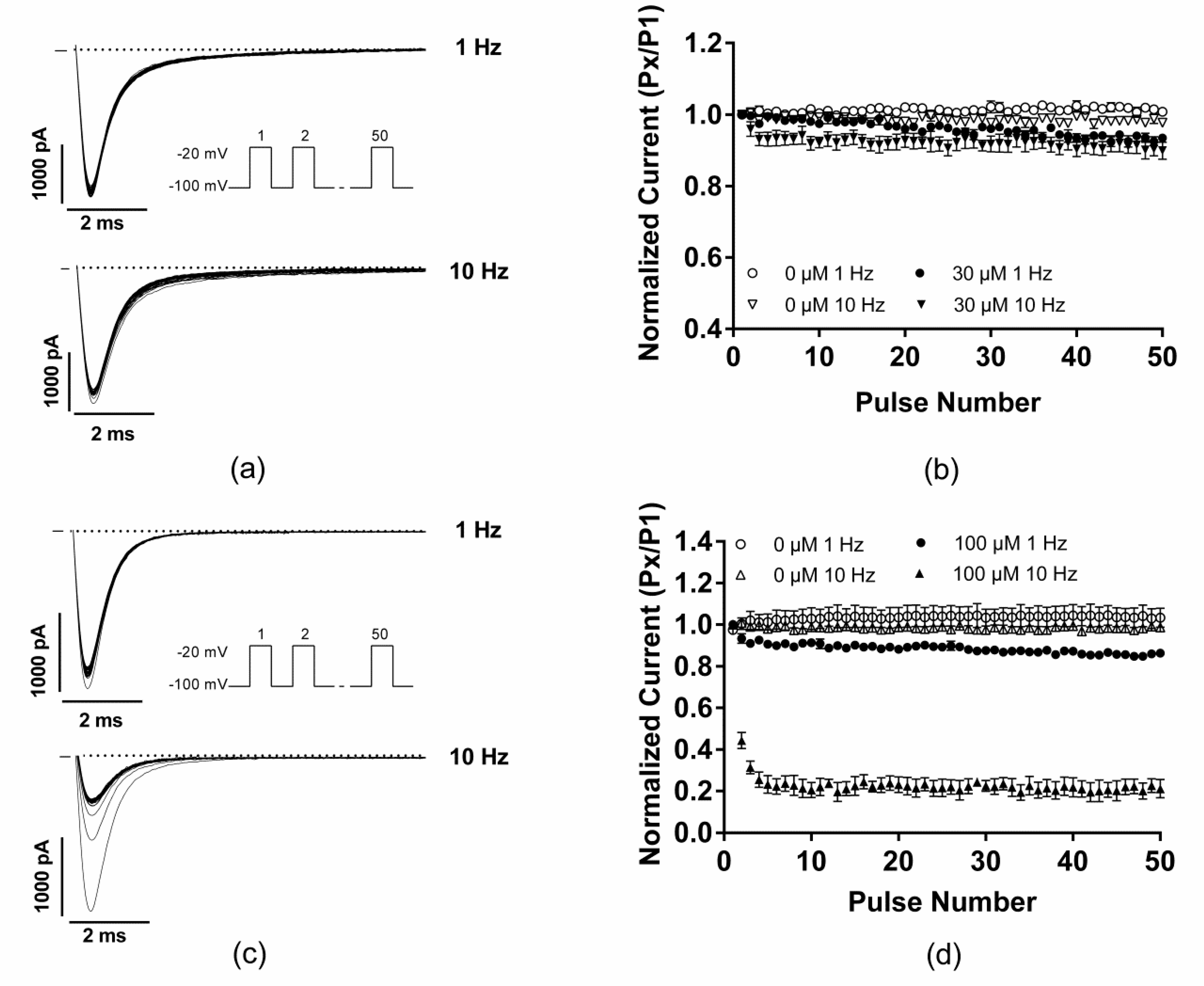
2.3. EGCG did not Produce any Use-Dependent Blockade of Nav1.5 Channels
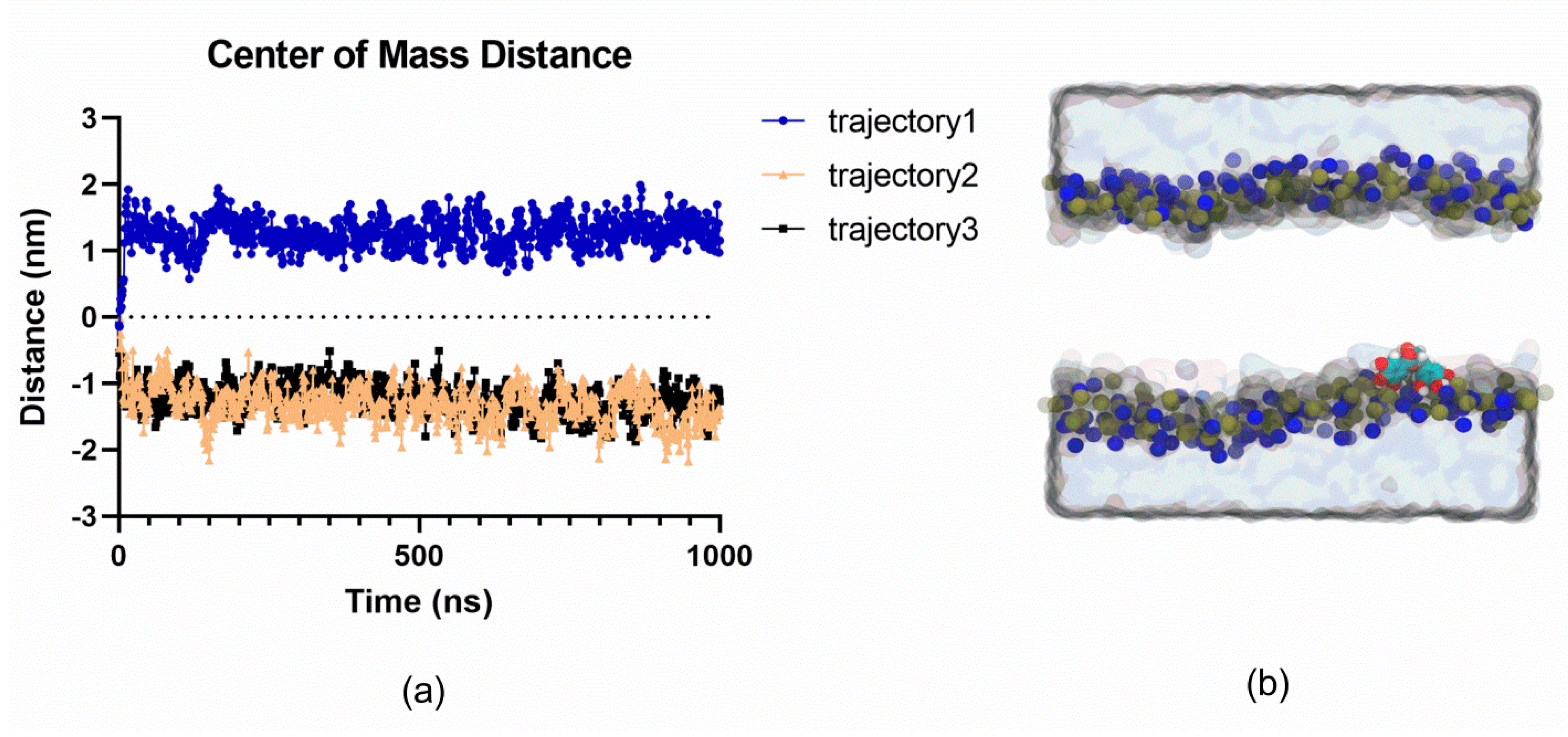
2.4. MD Simulations Suggest that EGCG does not Spontaneously Permeate Cell Membranes
3. Discussion
4. Materials and Methods
4.1. Cell Culture and Transfection
4.2. Cellular Electrophysiology
4.3. Chemicals
4.4. Molecular Dynamics Simulations
4.5. Data Analysis and Statistical Methods
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Chevalier, M.; Vermij, S.H.; Wyler, K.; Gillet, L.; Keller, I.; Abriel, H. Transcriptomic analyses of murine ventricular cardiomyocytes. Sci. Data 2018, 5, 180170. [Google Scholar] [CrossRef] [PubMed]
- Naylor, C.E.; Bagneris, C.; DeCaen, P.G.; Sula, A.; Scaglione, A.; Clapham, D.E.; Wallace, B.A. Molecular basis of ion permeability in a voltage-gated sodium channel. Embo J. 2016, 35, 820–830. [Google Scholar] [CrossRef] [PubMed]
- Payandeh, J.; Scheuer, T.; Zheng, N.; Catterall, W.A. The crystal structure of a voltage-gated sodium channel. Nature 2011, 475, 353–358. [Google Scholar] [CrossRef] [PubMed]
- Savio-Galimberti, E.; Argenziano, M.; Antzelevitch, C. Cardiac Arrhythmias Related to Sodium Channel Dysfunction. Handb. Exp. Pharmacol. 2018, 246, 331–354. [Google Scholar] [CrossRef]
- Wilde, A.A.M.; Amin, A.S. Clinical Spectrum of SCN5A Mutations: Long QT Syndrome, Brugada Syndrome, and Cardiomyopathy. Jacc. Clin. Electrophysiol. 2018, 4, 569–579. [Google Scholar] [CrossRef]
- Laurent, G.; Saal, S.; Amarouch, M.Y.; Beziau, D.M.; Marsman, R.F.; Faivre, L.; Barc, J.; Dina, C.; Bertaux, G.; Barthez, O.; et al. Multifocal ectopic Purkinje-related premature contractions: A new SCN5A-related cardiac channelopathy. J. Am. Coll. Cardiol. 2012, 60, 144–156. [Google Scholar] [CrossRef]
- Swan, H.; Amarouch, M.Y.; Leinonen, J.; Marjamaa, A.; Kucera, J.P.; Laitinen-Forsblom, P.J.; Lahtinen, A.M.; Palotie, A.; Kontula, K.; Toivonen, L.; et al. Gain-of-function mutation of the SCN5A gene causes exercise-induced polymorphic ventricular arrhythmias. Circ. Cardiovasc. Genet. 2014, 7, 771–781. [Google Scholar] [CrossRef]
- Yu, S.; Li, G.; Huang, C.L.; Lei, M.; Wu, L. Late sodium current associated cardiac electrophysiological and mechanical dysfunction. Pflug. Arch. Eur. J. Physiol. 2018, 470, 461–469. [Google Scholar] [CrossRef]
- Mann, S.A.; Castro, M.L.; Ohanian, M.; Guo, G.; Zodgekar, P.; Sheu, A.; Stockhammer, K.; Thompson, T.; Playford, D.; Subbiah, R.; et al. R222Q SCN5A mutation is associated with reversible ventricular ectopy and dilated cardiomyopathy. J. Am. Coll. Cardiol. 2012, 60, 1566–1573. [Google Scholar] [CrossRef]
- Amarouch, M.Y.; Swan, H.; Leinonen, J.; Marjamaa, A.; Lahtinen, A.M.; Kontula, K.; Toivonen, L.; Widen, E.; Abriel, H. Antiarrhythmic Action of Flecainide in Polymorphic Ventricular Arrhythmias Caused by a Gain-of-Function Mutation in the Nav 1.5 Sodium Channel. Ann. Noninvasive Electrocardiol. 2016, 21, 343–351. [Google Scholar] [CrossRef]
- Amarouch, M.Y.; Abriel, H. Cellular hyper-excitability caused by mutations that alter the activation process of voltage-gated sodium channels. Front. Physiol. 2015, 6, 45. [Google Scholar] [CrossRef] [PubMed]
- Saber, S.; Amarouch, M.Y.; Fazelifar, A.F.; Haghjoo, M.; Emkanjoo, Z.; Alizadeh, A.; Houshmand, M.; Gavrilenko, A.V.; Abriel, H.; Zaklyazminskaya, E.V. Complex genetic background in a large family with Brugada syndrome. Physiol. Rep. 2015, 3. [Google Scholar] [CrossRef] [PubMed]
- Kyle, J.W.; Makielski, J.C. Diseases caused by mutations in Nav1.5 interacting proteins. Card. Electrophysiol. Clin. 2014, 6, 797–809. [Google Scholar] [CrossRef] [PubMed]
- Edokobi, N.; Isom, L.L. Voltage-Gated Sodium Channel beta1/beta1B Subunits Regulate Cardiac Physiology and Pathophysiology. Front. Physiol. 2018, 9, 351. [Google Scholar] [CrossRef] [PubMed]
- Salvarani, N.; Crasto, S.; Miragoli, M.; Bertero, A.; Paulis, M.; Kunderfranco, P.; Serio, S.; Forni, A.; Lucarelli, C.; Dal Ferro, M.; et al. The K219T-Lamin mutation induces conduction defects through epigenetic inhibition of SCN5A in human cardiac laminopathy. Nat. Commun. 2019, 10, 2267. [Google Scholar] [CrossRef] [PubMed]
- Cerrone, M.; Lin, X.; Zhang, M.; Agullo-Pascual, E.; Pfenniger, A.; Chkourko Gusky, H.; Novelli, V.; Kim, C.; Tirasawadichai, T.; Judge, D.P.; et al. Missense mutations in plakophilin-2 cause sodium current deficit and associate with a Brugada syndrome phenotype. Circulation 2014, 129, 1092–1103. [Google Scholar] [CrossRef] [PubMed]
- Neubauer, J.; Rougier, J.S.; Abriel, H.; Haas, C. Functional implications of a rare variant in the sodium channel beta1B subunit (SCN1B) in a 5-month-old male sudden infant death syndrome case. Heartrhythm Case Rep. 2018, 4, 187–190. [Google Scholar] [CrossRef]
- Van Hoeijen, D.A.; Blom, M.T.; Tan, H.L. Cardiac sodium channels and inherited electrophysiological disorders: An update on the pharmacotherapy. Expert Opin. Pharmacother. 2014, 15, 1875–1887. [Google Scholar] [CrossRef]
- Mankad, P.; Kalahasty, G. Antiarrhythmic Drugs: Risks and Benefits. Med Clin. North. Am. 2019, 103, 821–834. [Google Scholar] [CrossRef]
- Boukhabza, M.; El Hilaly, J.; Attiya, N.; El-Haidani, A.; Filali-Zegzouti, Y.; Mazouzi, D.; Amarouch, M.Y. In Silico Evaluation of the Potential Antiarrhythmic Effect of Epigallocatechin-3-Gallate on Cardiac Channelopathies. Comput. Math. Methods Med. 2016, 2016, 7861653. [Google Scholar] [CrossRef]
- Chang, J.H.; Chang, S.L.; Hong, P.D.; Chen, P.N.; Hsu, C.H.; Lu, Y.Y.; Chen, Y.C. Epigallocatechin-3-gallate modulates arrhythmogenic activity and calcium homeostasis of left atrium. Int. J. Cardiol. 2017, 236, 174–180. [Google Scholar] [CrossRef] [PubMed]
- Eng, Q.Y.; Thanikachalam, P.V.; Ramamurthy, S. Molecular understanding of Epigallocatechin gallate (EGCG) in cardiovascular and metabolic diseases. J. Ethnopharmacol. 2018, 210, 296–310. [Google Scholar] [CrossRef] [PubMed]
- Kuriyama, S.; Shimazu, T.; Ohmori, K.; Kikuchi, N.; Nakaya, N.; Nishino, Y.; Tsubono, Y.; Tsuji, I. Green tea consumption and mortality due to cardiovascular disease, cancer, and all causes in Japan: The Ohsaki study. Jama 2006, 296, 1255–1265. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, J.; Ogawa, M.; Maejima, Y.; Isobe, K.; Tanaka, H.; Sagesaka, Y.M.; Isobe, M. Tea catechins attenuate chronic ventricular remodeling after myocardial ischemia in rats. J. Mol. Cell. Cardiol. 2007, 42, 432–440. [Google Scholar] [CrossRef]
- Suzuki, J.; Ogawa, M.; Futamatsu, H.; Kosuge, H.; Sagesaka, Y.M.; Isobe, M. Tea catechins improve left ventricular dysfunction, suppress myocardial inflammation and fibrosis, and alter cytokine expression in rat autoimmune myocarditis. Eur. J. Heart Fail. 2007, 9, 152–159. [Google Scholar] [CrossRef]
- Ma, Y.; Hu, Y.; Wu, J.; Wen, J.; Li, S.; Zhang, L.; Zhang, J.; Li, Y.; Li, J. Epigallocatechin-3-gallate inhibits angiotensin II-induced cardiomyocyte hypertrophy via regulating Hippo signaling pathway in H9c2 rat cardiomyocytes. Acta Biochim. Et Biophys. Sin. 2019, 51, 422–430. [Google Scholar] [CrossRef]
- Al Hroob, A.M.; Abukhalil, M.H.; Hussein, O.E.; Mahmoud, A.M. Pathophysiological mechanisms of diabetic cardiomyopathy and the therapeutic potential of epigallocatechin-3-gallate. Biomed. Pharmacother. 2019, 109, 2155–2172. [Google Scholar] [CrossRef]
- Zhang, Q.; Hu, L.; Chen, L.; Li, H.; Wu, J.; Liu, W.; Zhang, M.; Yan, G. (−)-Epigallocatechin-3-gallate, the major green tea catechin, regulates the desensitization of beta1 adrenoceptor via GRK2 in experimental heart failure. Inflammopharmacology 2018, 26, 1081–1091. [Google Scholar] [CrossRef]
- Wei, H.; Meng, Z. Epigallocatechin-3-gallate protects Na+ channels in rat ventricular myocytes against sulfite. Cardiovasc. Toxicol. 2010, 10, 166–173. [Google Scholar] [CrossRef]
- Kang, J.; Cheng, H.; Ji, J.; Incardona, J.; Rampe, D. In vitro electrocardiographic and cardiac ion channel effects of (-)-epigallocatechin-3-gallate, the main catechin of green tea. J. Pharmacol. Exp. Ther. 2010, 334, 619–626. [Google Scholar] [CrossRef]
- Wang, R.; Zhou, W.; Jiang, X. Reaction kinetics of degradation and epimerization of epigallocatechin gallate (EGCG) in aqueous system over a wide temperature range. J. Agric. Food Chem. 2008, 56, 2694–2701. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Jia, Y.Y.; Guo, J.L.; Liu, P.Q.; Jiang, J.M. Effects of (-)-gallocatechin-3-gallate on tetrodotoxin-resistant voltage-gated sodium channels in rat dorsal root ganglion neurons. Int. J. Mol. Sci. 2013, 14, 9779–9789. [Google Scholar] [CrossRef] [PubMed]
- Abdelsayed, M.; Ruprai, M.; Ruben, P.C. The efficacy of Ranolazine on E1784K is altered by temperature and calcium. Sci. Rep. 2018, 8, 3643. [Google Scholar] [CrossRef] [PubMed]
- El-Battrawy, I.; Lang, S.; Zhao, Z.; Akin, I.; Yucel, G.; Meister, S.; Patocskai, B.; Behnes, M.; Rudic, B.; Tulumen, E.; et al. Hyperthermia Influences the Effects of Sodium Channel Blocking Drugs in Human-Induced Pluripotent Stem Cell-Derived Cardiomyocytes. PLoS ONE 2016, 11, e0166143. [Google Scholar] [CrossRef] [PubMed]
- Deng, H.M.; Yin, S.T.; Yan, D.; Tang, M.L.; Li, C.C.; Chen, J.T.; Wang, M.; Ruan, D.Y. Effects of EGCG on voltage-gated sodium channels in primary cultures of rat hippocampal CA1 neurons. Toxicology 2008, 252, 1–8. [Google Scholar] [CrossRef]
- Kim, T.H.; Lim, J.M.; Kim, S.S.; Kim, J.; Park, M.; Song, J.H. Effects of (−) epigallocatechin-3-gallate on Na(+) currents in rat dorsal root ganglion neurons. Eur. J. Pharmacol. 2009, 604, 20–26. [Google Scholar] [CrossRef]
- Salazar, P.B.; de Athayde Moncorvo Collado, A.; Canal-Martinez, V.; Minahk, C.J. Differential inhibition of human erythrocyte acetylcholinesterase by polyphenols epigallocatechin-3-gallate and resveratrol. Relevance of the membrane-bound form. BioFactors 2017, 43, 73–81. [Google Scholar] [CrossRef]
- Ingolfsson, H.I.; Koeppe, R.E., 2nd; Andersen, O.S. Effects of green tea catechins on gramicidin channel function and inferred changes in bilayer properties. Febs Lett. 2011, 585, 3101–3105. [Google Scholar] [CrossRef]
- Gamal El-Din, T.M.; Lenaeus, M.J.; Zheng, N.; Catterall, W.A. Fenestrations control resting-state block of a voltage-gated sodium channel. Proc. Natl. Acad. Sci. USA 2018, 115, 13111–13116. [Google Scholar] [CrossRef]
- Henning, S.M.; Niu, Y.; Lee, N.H.; Thames, G.D.; Minutti, R.R.; Wang, H.; Go, V.L.; Heber, D. Bioavailability and antioxidant activity of tea flavanols after consumption of green tea, black tea, or a green tea extract supplement. Am. J. Clin. Nutr. 2004, 80, 1558–1564. [Google Scholar] [CrossRef]
- Chow, H.H.; Cai, Y.; Hakim, I.A.; Crowell, J.A.; Shahi, F.; Brooks, C.A.; Dorr, R.T.; Hara, Y.; Alberts, D.S. Pharmacokinetics and safety of green tea polyphenols after multiple-dose administration of epigallocatechin gallate and polyphenon E in healthy individuals. Clin. Cancer Res. 2003, 9, 3312–3319. [Google Scholar] [PubMed]
- Ullmann, U.; Haller, J.; Decourt, J.P.; Girault, N.; Girault, J.; Richard-Caudron, A.S.; Pineau, B.; Weber, P. A single ascending dose study of epigallocatechin gallate in healthy volunteers. J. Int. Med Res. 2003, 31, 88–101. [Google Scholar] [CrossRef] [PubMed]
- Lee, M.J.; Maliakal, P.; Chen, L.; Meng, X.; Bondoc, F.Y.; Prabhu, S.; Lambert, G.; Mohr, S.; Yang, C.S. Pharmacokinetics of tea catechins after ingestion of green tea and (-)-epigallocatechin-3-gallate by humans: Formation of different metabolites and individual variability. Cancer Epidemiol. Biomark. Prev. 2002, 11, 1025–1032. [Google Scholar]
- Narumi, K.; Sonoda, J.; Shiotani, K.; Shigeru, M.; Shibata, M.; Kawachi, A.; Tomishige, E.; Sato, K.; Motoya, T. Simultaneous detection of green tea catechins and gallic acid in human serum after ingestion of green tea tablets using ion-pair high-performance liquid chromatography with electrochemical detection. J. Chromatography. Banalytical Technol. Biomed. Life Sci. 2014, 945–946, 147–153. [Google Scholar] [CrossRef] [PubMed]
- Lambert, J.D.; Lee, M.J.; Lu, H.; Meng, X.; Hong, J.J.; Seril, D.N.; Sturgill, M.G.; Yang, C.S. Epigallocatechin-3-gallate is absorbed but extensively glucuronidated following oral administration to mice. J. Nutr. 2003, 133, 4172–4177. [Google Scholar] [CrossRef] [PubMed]
- Yang, C.S.; Wang, X.; Lu, G.; Picinich, S.C. Cancer prevention by tea: Animal studies, molecular mechanisms and human relevance. Nat. Reviews. Cancer 2009, 9, 429–439. [Google Scholar] [CrossRef]
- Lambert, J.D.; Lee, M.J.; Diamond, L.; Ju, J.; Hong, J.; Bose, M.; Newmark, H.L.; Yang, C.S. Dose-dependent levels of epigallocatechin-3-gallate in human colon cancer cells and mouse plasma and tissues. Drug Metab. Dispos. Biol. Fate Chem. 2006, 34, 8–11. [Google Scholar] [CrossRef]
- Nagle, D.G.; Ferreira, D.; Zhou, Y.D. Epigallocatechin-3-gallate (EGCG): Chemical and biomedical perspectives. Phytochemistry 2006, 67, 1849–1855. [Google Scholar] [CrossRef]
- Krupkova, O.; Ferguson, S.J.; Wuertz-Kozak, K. Stability of (-)-epigallocatechin gallate and its activity in liquid formulations and delivery systems. J. Nutr. Biochem. 2016, 37, 1–12. [Google Scholar] [CrossRef]
- Li, N.; Taylor, L.S.; Ferruzzi, M.G.; Mauer, L.J. Kinetic study of catechin stability: Effects of pH, concentration, and temperature. J. Agric. Food Chem. 2012, 60, 12531–12539. [Google Scholar] [CrossRef]
- Jo, S.; Kim, T.; Iyer, V.G.; Im, W. CHARMM-GUI: A web-based graphical user interface for CHARMM. J. Comput. Chem. 2008, 29, 1859–1865. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.; Cheng, X.; Swails, J.M.; Yeom, M.S.; Eastman, P.K.; Lemkul, J.A.; Wei, S.; Buckner, J.; Jeong, J.C.; Qi, Y.; et al. CHARMM-GUI Input Generator for NAMD, GROMACS, AMBER, OpenMM, and CHARMM/OpenMM Simulations Using the CHARMM36 Additive Force Field. J. Chem. Theory Comput. 2016, 12, 405–413. [Google Scholar] [CrossRef] [PubMed]
- Wu, E.L.; Cheng, X.; Jo, S.; Rui, H.; Song, K.C.; Davila-Contreras, E.M.; Qi, Y.; Lee, J.; Monje-Galvan, V.; Venable, R.M.; et al. CHARMM-GUI Membrane Builder toward realistic biological membrane simulations. J. Comput. Chem. 2014, 35, 1997–2004. [Google Scholar] [CrossRef] [PubMed]
- Klauda, J.B.; Venable, R.M.; Freites, J.A.; O’Connor, J.W.; Tobias, D.J.; Mondragon-Ramirez, C.; Vorobyov, I.; MacKerell, A.D., Jr.; Pastor, R.W. Update of the CHARMM all-atom additive force field for lipids: Validation on six lipid types. J. Phys. Chemistry. B 2010, 114, 7830–7843. [Google Scholar] [CrossRef] [PubMed]
- Jorgensen, W.L.; Chandrasekhar, J.; Madura, J.D.; Impey, R.W.; Klein, M.L. Comparison of simple potential functions for simulating liquid water. J. Chem. Phys. 1983, 79, 926–935. [Google Scholar] [CrossRef]
- Parrinello, M.; Rahman, A. Polymorphic transitions in single crystals: A new molecular dynamics method. J. Appl. Phys. 1981, 52, 7182–7190. [Google Scholar] [CrossRef]
- Nosé, S. A unified formulation of the constant temperature molecular dynamics methods. J. Chem. Phys. 1984, 81, 511–519. [Google Scholar] [CrossRef]
- Darden, T.; York, D.; Pedersen, L. Particle mesh Ewald: An N⋅log(N) method for Ewald sums in large systems. J. Chem. Phys. 1993, 98, 10089–10092. [Google Scholar] [CrossRef]
- Abraham, M.J.; Murtola, T.; Schulz, R.; Páll, S.; Smith, J.C.; Hess, B.; Lindahl, E. GROMACS: High performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 2015, 1–2, 19–25. [Google Scholar] [CrossRef]
- Zoete, V.; Cuendet, M.A.; Grosdidier, A.; Michielin, O. SwissParam: A fast force field generation tool for small organic molecules. J. Comput. Chem. 2011, 32, 2359–2368. [Google Scholar] [CrossRef]
Sample Availability: Samples of Epigallocatechin-3-Gallate are available from the authors. |
| Control | 30 µM EGCG | |
|---|---|---|
| Peak current densities at −20 mV (pA/pF) | −146.2 ± 17.8; n = 12 | −64.3 ± 7.3 ***; n = 12 |
| V1/2 activation (mV) | −27.4 ± 0.8; n = 12 | −27.4 ± 1; n = 12 |
| Activation slope (mV) | 6.7 ± 0.2; n = 12 | 8 ± 2 ***; n = 12 |
| Time to peak at −20 mV (ms) | 1.12 ± 0.02; n = 12 | 1.16 ± 0.03; n = 12 |
| V1/2 inactivation (mV) | −79.4 ± 1.3; n = 13 | −87 ± 0.9 ***; n = 13 |
| Inactivation slope (mV) | 5 ± 0.2; n = 13 | 5.5 ± 0.2 *; n = 13 |
| t1/2 inactivation at −20 mV (ms) | 0.43 ± 0.02; n = 12 | 0.56 ± 0.03 **; n = 12 |
| Recovery from fast inactivation (ms) | ||
| τfast,1 | 5.1 ± 0.9; n = 5 | 24.3 ± 7.1 *; n = 5 |
| τfast,2 | 110 ± 28.6; n = 5 | 233.7 ± 37.6 *; n = 5 |
| UDB-1 Hz | UDB-10 Hz | |
|---|---|---|
| EGCG (30 µM) | 0.93 ± 0.01 **, n = 5 | 0.90 ± 0.02 *, n = 5 |
| Mexiletine (100 µM) | 0.86 ± 0.01 ***, n = 4 | 0.21 ± 0.04 ***, n = 4 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Amarouch, M.-Y.; Kurt, H.; Delemotte, L.; Abriel, H. Biophysical Characterization of Epigallocatechin-3-Gallate Effect on the Cardiac Sodium Channel Nav1.5. Molecules 2020, 25, 902. https://doi.org/10.3390/molecules25040902
Amarouch M-Y, Kurt H, Delemotte L, Abriel H. Biophysical Characterization of Epigallocatechin-3-Gallate Effect on the Cardiac Sodium Channel Nav1.5. Molecules. 2020; 25(4):902. https://doi.org/10.3390/molecules25040902
Chicago/Turabian StyleAmarouch, Mohamed-Yassine, Han Kurt, Lucie Delemotte, and Hugues Abriel. 2020. "Biophysical Characterization of Epigallocatechin-3-Gallate Effect on the Cardiac Sodium Channel Nav1.5" Molecules 25, no. 4: 902. https://doi.org/10.3390/molecules25040902
APA StyleAmarouch, M.-Y., Kurt, H., Delemotte, L., & Abriel, H. (2020). Biophysical Characterization of Epigallocatechin-3-Gallate Effect on the Cardiac Sodium Channel Nav1.5. Molecules, 25(4), 902. https://doi.org/10.3390/molecules25040902






