New Modified Deoxythymine with Dibranched Tetraethylene Glycol Stabilizes G-Quadruplex Structures
Abstract
1. Introduction
2. Results and Discussion
2.1. Sequence Design to Understand the Effect of Modified Deoxythymines on G-quadruplex Stability
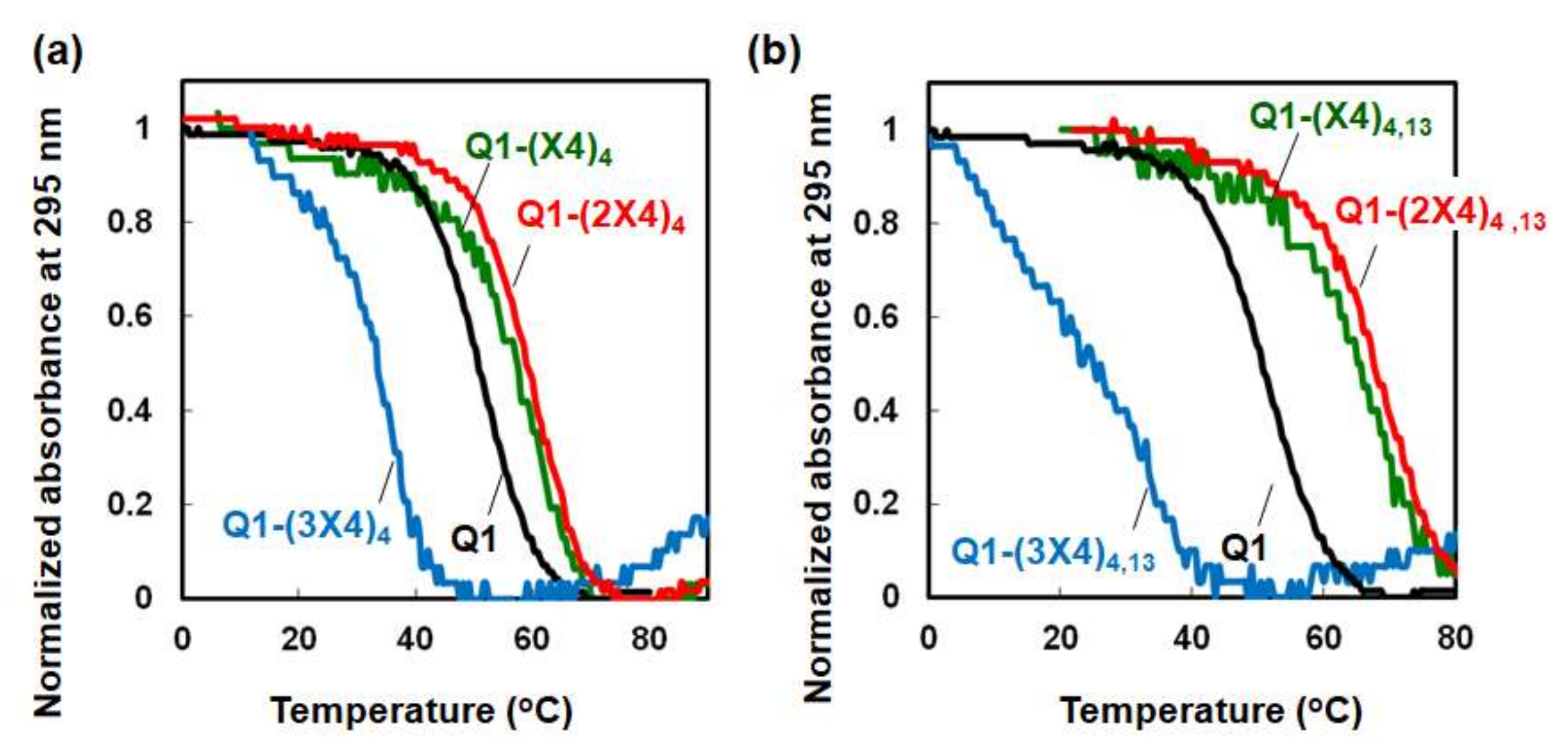
2.2. Thermodynamic Analysis of the Effects of TEG-modified Deoxythymines on G-quadruplex Stability
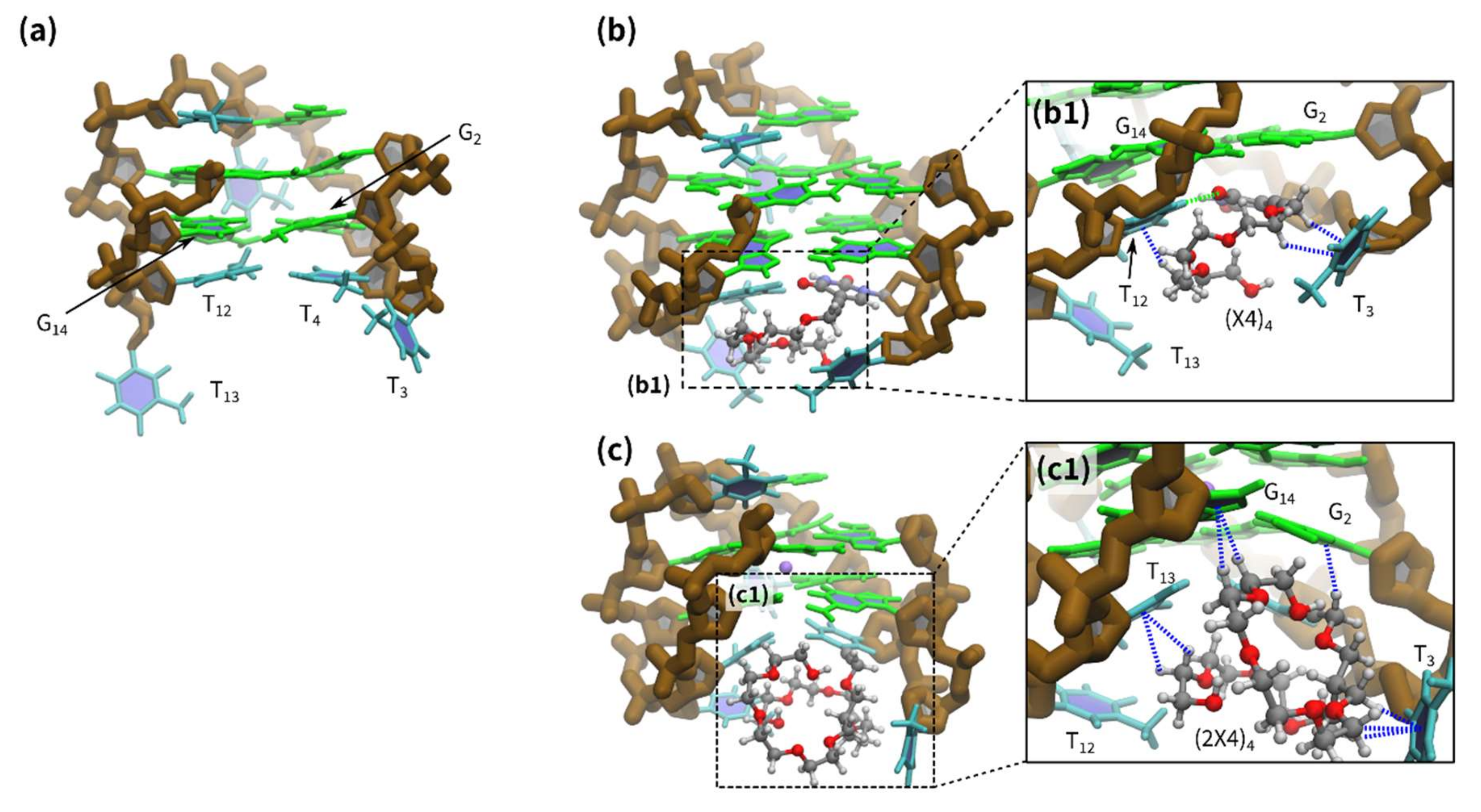
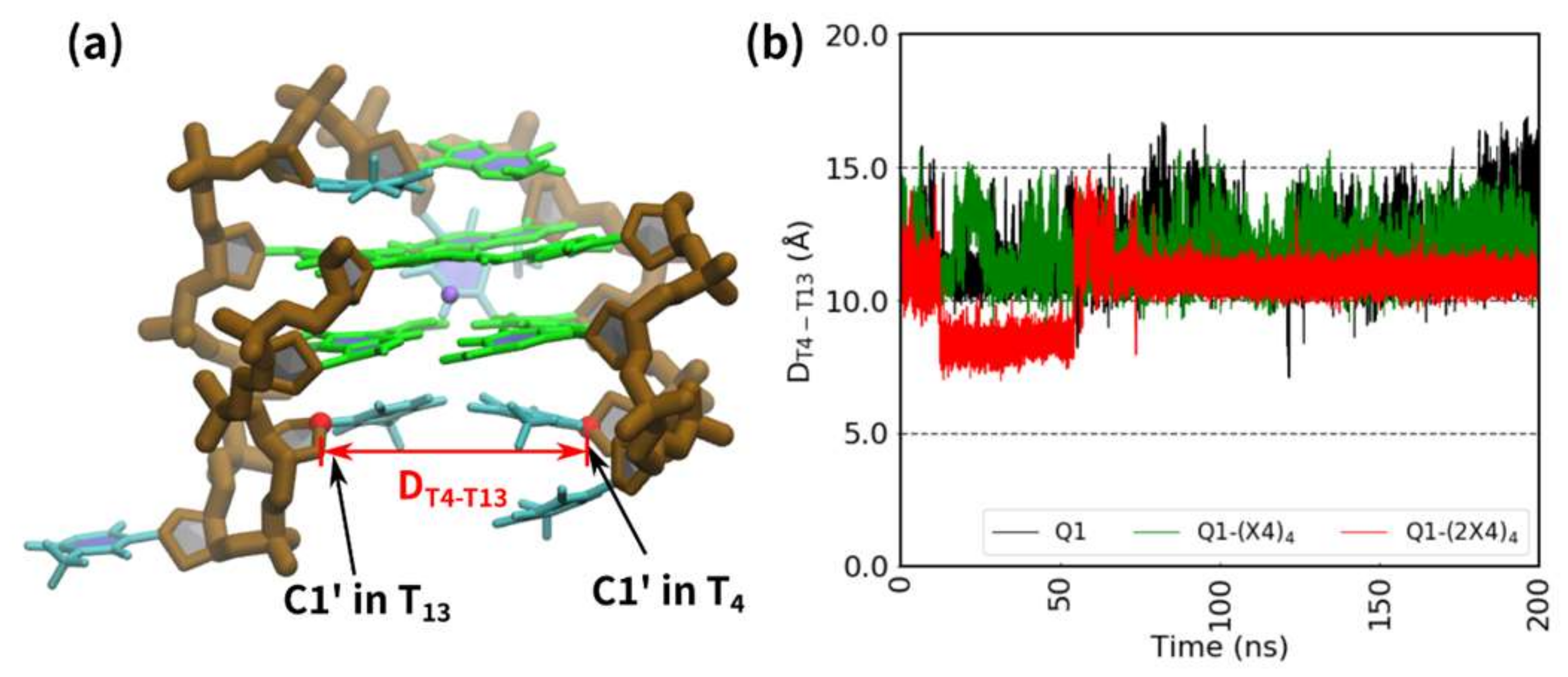
2.3. Molecular Dynamic (MD) Calculations for G-quadruplexes with Modified Deoxythymines
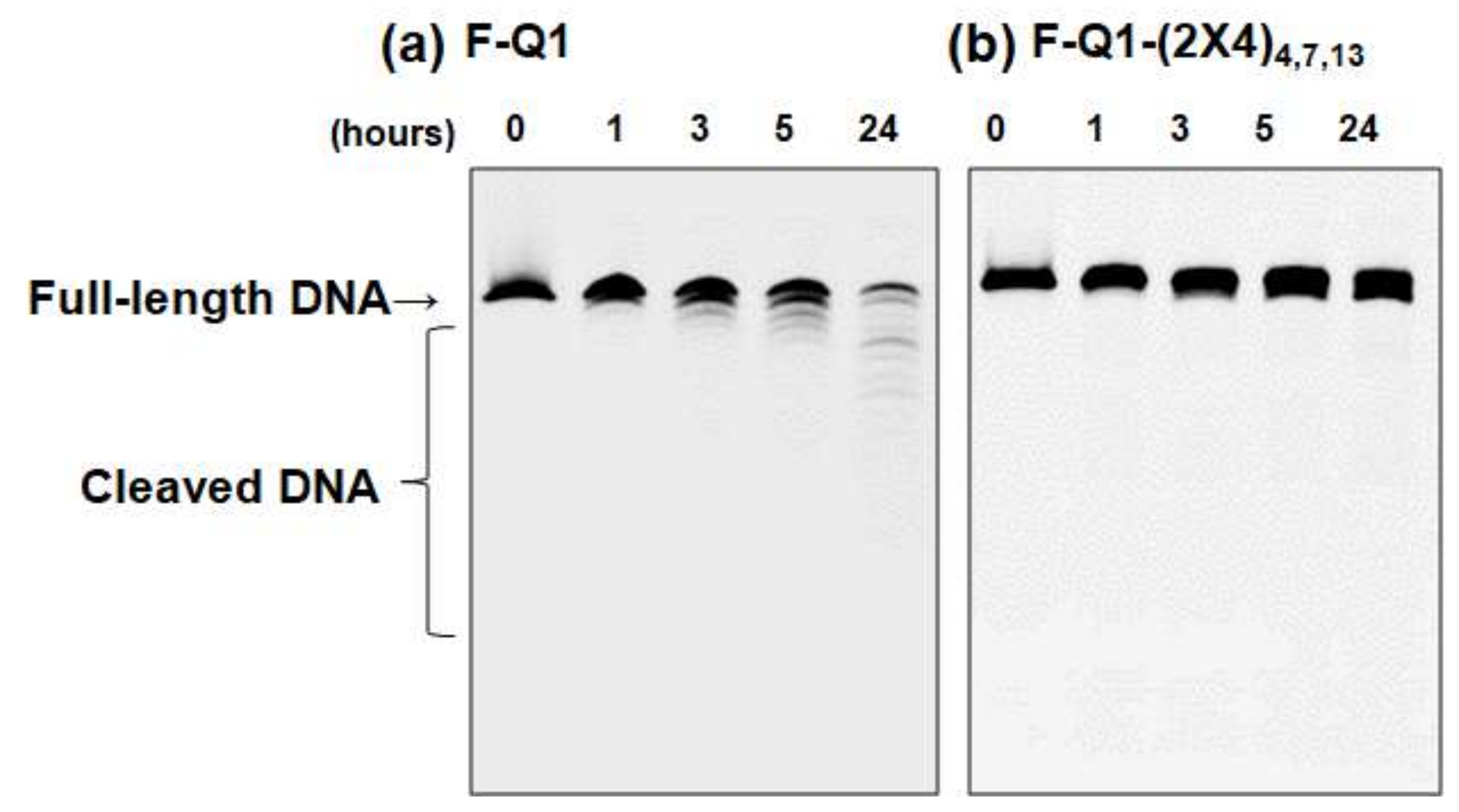
2.4. Modified Bases Improve Nuclease Resistance
3. Materials and Methods
3.1. Chemicals and Materials
3.2. Thermodynamic Analysis
3.3. Molecular Dynamic (MD) Simulations
3.4. Assays for Nuclease Resistance
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Balasubramanian, S.; Hurley, L.H.; Neidle, S. Targeting G-quadruplexes in gene promoters: A novel anticancer strategy? Nat. Rev. Drug Discov. 2011, 10, 261–275. [Google Scholar] [CrossRef] [PubMed]
- Jain, A.; Vale, R.D. RNA phase transitions in repeat expansion disorders. Nature 2017, 546, 243–247. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Yang, J.; Wild, A.T.; Wu, W.H.; Shah, R.; Danussi, C.; Riggins, G.J.; Kannan, K.; Sulman, E.P.; Chan, T.A.; et al. G-quadruplex DNA drives genomic instability and represents a targetable molecular abnormality in ATRX-deficient malignant glioma. Nat. Commun. 2019, 10, 943. [Google Scholar] [CrossRef] [PubMed]
- Tateishi-Karimata, H.; Isono, N.; Sugimoto, N. New insights into transcription fidelity: Thermal stability of non-canonical structures in template DNA regulates transcriptional arrest, pause, and slippage. PLoS ONE 2014, 9, e90580. [Google Scholar] [CrossRef] [PubMed]
- Endoh, T.; Kawasaki, Y.; Sugimoto, N. Suppression of gene expression by G-quadruplexes in open reading frames depends on G-quadruplex stability. Angew. Chem. Int. Ed. Engl. 2013, 52, 5522–5526. [Google Scholar] [CrossRef] [PubMed]
- Kawauchi, K.; Sugimoto, W.; Yasui, T.; Murata, K.; Itoh, K.; Takagi, K.; Tsuruoka, T.; Akamatsu, K.; Tateishi-Karimata, H.; Sugimoto, N.; et al. An anionic phthalocyanine decreases NRAS expression by breaking down its RNA G-quadruplex. Nat. Commun. 2018, 9, 2271. [Google Scholar] [CrossRef]
- Yaku, H.; Murashima, T.; Tateishi-Karimata, H.; Nakano, S.; Miyoshi, D.; Sugimoto, N. Study on effects of molecular crowding on G-quadruplex-ligand binding and ligand-mediated telomerase inhibition. Methods 2013, 64, 19–27. [Google Scholar] [CrossRef]
- Rodriguez, R.; Pantos, G.D.; Goncalves, D.P.; Sanders, J.K.; Balasubramanian, S. Ligand-driven G-quadruplex conformational switching by using an unusual mode of interaction. Angew. Chem. Int. Ed. Engl. 2007, 46, 5405–5407. [Google Scholar] [CrossRef]
- Ma, Y.; Ou, T.M.; Tan, J.H.; Hou, J.Q.; Huang, S.L.; Gu, L.Q.; Huang, Z.S. Synthesis and evaluation of 9-O-substituted berberine derivatives containing aza-aromatic terminal group as highly selective telomeric G-quadruplex stabilizing ligands. Bioorg. Med. Chem. Lett. 2009, 19, 3414–3417. [Google Scholar] [CrossRef]
- Sasaki, Y.; Miyoshi, D.; Sugimoto, N. Regulation of DNA nucleases by molecular crowding. Nucl. Acids Res 2007, 35, 4086–4093. [Google Scholar] [CrossRef]
- Yang, W. Nucleases: Diversity of structure, function and mechanism. Q. Rev. Biophys. 2011, 44, 1–93. [Google Scholar] [CrossRef] [PubMed]
- Rameez, S.; Palmer, A.F. Simple method for preparing poly(ethylene glycol)-surface-conjugated liposome-encapsulated hemoglobins: Physicochemical properties, long-term storage stability, and their reactions with O2, CO, and NO. Langmuir 2011, 27, 8829–8840. [Google Scholar] [CrossRef] [PubMed]
- Hong, K.; Zheng, W.; Baker, A.; Papahadjopoulos, D. Stabilization of cationic liposome-plasmid DNA complexes by polyamines and poly(ethylene glycol)-phospholipid conjugates for efficient in vivo gene delivery. FEBS Lett. 1997, 400, 233–237. [Google Scholar] [CrossRef]
- Litzinger, D.C.; Buiting, A.M.; van Rooijen, N.; Huang, L. Effect of liposome size on the circulation time and intraorgan distribution of amphipathic poly(ethylene glycol)-containing liposomes. Biochim. Biophys. Acta 1994, 1190, 99–107. [Google Scholar] [CrossRef]
- Katayose, S.; Kataoka, K. Remarkable increase in nuclease resistance of plasmid DNA through supramolecular assembly with poly(ethylene glycol)-poly(L-lysine) block copolymer. J. Pharm. Sci. 1998, 87, 160–163. [Google Scholar] [CrossRef] [PubMed]
- Tateishi-Karimata, H.; Muraoka, T.; Kinbara, K.; Sugimoto, N. G-quadruplexes with tetraethylene glycol-modified deoxythymidines are resistant to nucleases and inhibit HIV-1 reverse transcriptase. ChemBioChem 2016, 17, 1399–1402. [Google Scholar] [CrossRef]
- Tateishi-Karimata, H.; Ohyama, T.; Muraoka, T.; Podbevsek, P.; Wawro, A.M.; Tanaka, S.; Nakano, S.I.; Kinbara, K.; Plavec, J.; Sugimoto, N. Newly characterized interaction stabilizes DNA structure: Oligoethylene glycols stabilize G-quadruplexes CH-pi interactions. Nucl. Acids Res 2017, 45, 7021–7030. [Google Scholar] [CrossRef]
- Puglisi, J.D.; Tinoco, I., Jr. Absorbance melting curves of RNA. Methods Enzym. 1989, 180, 304–325. [Google Scholar]
- Macaya, R.F.; Schultze, P.; Smith, F.W.; Roe, J.A.; Feigon, J. Thrombin-binding DNA aptamer forms a unimolecular quadruplex structure in solution. Proc. Natl. Acad. Sci. USA 1993, 90, 3745–3749. [Google Scholar] [CrossRef]
- Trajkovski, M.; Sket, P.; Plavec, J. Cation localization and movement within DNA thrombin binding aptamer in solution. Org. Biomol. Chem. 2009, 7, 4677–4684. [Google Scholar] [CrossRef]
- Schultze, P.; Macaya, R.F.; Feigon, J. Three-dimensional solution structure of the thrombin-binding DNA aptamer d(GGTTGGTGTGGTTGG). J. Mol. Biol. 1994, 235, 1532–1547. [Google Scholar] [CrossRef] [PubMed]
- Sugimoto, N.; Kierzek, R.; Turner, D.H. Sequence dependence for the energetics of dangling ends and terminal base pairs in ribonucleic acid. Biochemistry 1987, 26, 4554–4558. [Google Scholar] [CrossRef] [PubMed]
- Ohmichi, T.; Nakano, S.; Miyoshi, D.; Sugimoto, N. Long RNA dangling end has large energetic contribution to duplex stability. J. Am. Chem. Soc. 2002, 124, 10367–10372. [Google Scholar] [CrossRef] [PubMed]
- Freier, S.M.; Sugimoto, N.; Sinclair, A.; Alkema, D.; Neilson, T.; Kierzek, R.; Caruthers, M.H.; Turner, D.H. Stability of XGCGCp, GCGCYp, and XGCGCYp helixes: An empirical estimate of the energetics of hydrogen bonds in nucleic acids. Biochemistry 1986, 25, 3214–3219. [Google Scholar] [CrossRef] [PubMed]
- Tateishi-Karimata, H.; Pramanik, S.; Nakano, S.; Miyoshi, D.; Sugimoto, N. Dangling ends perturb the stability of RNA duplexes responsive to surrounding conditions. ChemMedChem 2014, 9, 2150–2155. [Google Scholar] [CrossRef]
- Lucas, R.; Gomez-Pinto, I.; Avino, A.; Reina, J.J.; Eritja, R.; Gonzalez, C.; Morales, J.C. Highly polar carbohydrates stack onto DNA duplexes via CH/pi interactions. J. Am. Chem. Soc. 2011, 133, 1909–1916. [Google Scholar] [CrossRef]
- Fujita, H.; Nakajima, K.; Kasahara, Y.; Ozaki, H.; Kuwahara, M. Polymerase-mediated high-density incorporation of amphiphilic functionalities into DNA: Enhancement of nuclease resistance and stability in human serum. Bioorg. Med. Chem. Lett. 2015, 25, 333–336. [Google Scholar] [CrossRef]
- Richards, E.G.; In Fasman, G.D. Handbook of biochemistry and molecular biology. In Nucleic Acids, 3rd ed; CRC Press: Cleveland, OH, USA, 1975. [Google Scholar]
- Tateishi-Karimta, H.; Sugimoto, N. Control of stability and structure of nucleic acids using cosolutes. Methods 2014, 67, 151–158. [Google Scholar] [CrossRef]
- Marathias, V.M.; Bolton, P.H. Structures of the potassium-saturated, 2:1, and intermediate, 1:1, forms of a quadruplex DNA. Nucl. Acids Res. 2000, 28, 1969–1977. [Google Scholar] [CrossRef]
- Discovery Studio, version 2016; Dassault Systèmes BIOVIA: San Diego, CA, USA, 2016.
- Jorgensen, W.L.; Chandrasekhar, J.; Madura, J.D.; Impey, R.W.; Klein, M.L. Comparison of simple potential functions for simulating liquid water. J. Chem. Phys. 1983, 79, 926–935. [Google Scholar] [CrossRef]
- Abraham, M.J.; Murtola, T.; Schulz, R.; Páll, S.; Smith, J.C.; Hess, B.; Lindahl, E. GROMACS: High performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 2015, 1–2, 19–25. [Google Scholar] [CrossRef]
- Maier, J.A.; Martinez, C.; Kasavajhala, K.; Wickstrom, L.; Hauser, K.E.; Simmerling, C. ff14SB: Improving the Accuracy of Protein Side Chain and Backbone Parameters from ff99SB. J. Chem. Theory Comput. 2015, 11, 3696–3713. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Wang, W.; Kollman, P.A.; Case, D.A. Automatic atom type and bond type perception in molecular mechanical calculations. J. Mol. Graph. Model. 2006, 25, 247–260. [Google Scholar] [CrossRef] [PubMed]
- Case, D.A.; Berryman, J.T.; Betz, R.M.; Cerutti, D.S.; Cheatham, T.E., III; Darden, T.A.; Duke, R.E.; Giese, T.J.; Gohlke, H.; Goetz, A.W.; et al. The Amber biomolecular simulation programs. J. Comput. Chem. 2005, 26, 1668–1688. [Google Scholar] [CrossRef]
- Besler, B.H.; Merz, K.M.; Kollman, P.A. Atomic charges derived from semiempirical methods. J. Comput. Chem. 1990, 11, 431–439. [Google Scholar] [CrossRef]
- Frisch, M.J.; Trucks, G.W.; Schlegel, H.B.; Scuseria, G.E.; Robb, M.A.; Cheeseman, J.R.; Scalmani, G.; Barone, V.; Mennucci, B.; Petersson, G.A.; et al. Gaussian09 Revision D.01; Gaussian Inc.: Wallingford, CT, USA, 2013. [Google Scholar]
- Wang, J.; Cieplak, P.; Kollman, P.A. How well does a restrained electrostatic potential (RESP) model perform in calculating conformational energies of organic and biological molecules? J. Comput. Chem. 2000, 21, 1049–1074. [Google Scholar] [CrossRef]
- Sousa da Silva, A.W.; Vranken, W.F. ACPYPE-AnteChamber PYthon Parser interfacE. BMC Res. Notes 2012, 5, 367. [Google Scholar] [CrossRef]
- Hess, B.; Bekker, H.; Berendsen, H.J.C.; Fraaije, J.G.E.M. LINCS: A linear constraint solver for molecular simulations. J. Comput. Chem. 1997, 18, 1463–1472. [Google Scholar] [CrossRef]
- Cheatham, T.E.I.; Miller, J.L.; Fox, T.; Darden, T.A.; Kollman, P.A. Molecular Dynamics Simulations on Solvated Biomolecular Systems: The Particle Mesh Ewald Method Leads to Stable Trajectories of DNA, RNA, and Proteins. J. Am. Chem. Soc. 1995, 117, 4193–4194. [Google Scholar] [CrossRef]
- Darden, T.; York, D.; Pedersen, L. Particle mesh Ewald: An N⋅log(N) method for Ewald sums in large systems. J. Chem. Phys. 1993, 98, 10089–10092. [Google Scholar] [CrossRef]
Sample Availability: Not available. |


| Name | Sequences (5′-3′) a |
|---|---|
| Q1 | GGTTGGTGTGGTTGG |
| Q1-(X4)4 | GGT(X4)GGTGTGGTTGG |
| Q1-(X4)4,13 | GGT(X4)GGTGTGGT(X4)GG |
| Q1-(2X4)4 | GGT(2X4)GGTGTGGTTGG |
| Q1-(2X4)7 | GGTTGG(2X4)GTGGTTGG |
| Q1-(2X4)4,13 | GGT(2X4)GGTGTGGT(2X4)GG |
| Q1-(3X4)4 | GGT(3X4)GGTGTGGTTGG |
| Q1-(3X4)4,13 | GGT(3X4)GGTGTGGT(3X4)GG |
| F-Q1 b | F-GGTTGGTGTGGTTGG |
| F-Q1-(2X4)4,7,13 b | F-GGT(2X4)GG(2X4)GTGGT(2X4)GG |
| F-Q1-(3X4)4,7,13 b | F-GGT(3X4)GG(3X4)GTGGT(3X4)GG |
| Sequence | ∆ H° (kcal mol−1) | T∆S° (kcal mol−1) | ∆G°25 (kcal mol−1) | Tmb (°C) |
|---|---|---|---|---|
| Q1-(2X4)4 | −45.3 ± 3.3 | −39.9 ± 2.9 | −5.4 ± 0.4 | 61.6 |
| Q1-(2X4)7 | −38.2 ± 8.7 | −34.5 ± 1.6 | −3.7 ± 0.7 | 53.2 |
| Q1-(2X4)4,13 | −50.5 ± 1.5 | −43.4 ± 5.4 | −7.1 ± 0.1 | 70.1 |
| Q1-(3X4)4 | −72.9 ± 3.3 | −69.7 ± 2.9 | −3.2 ± 0.4 | 36.4 |
| Q1-(3X4)4,13 | n.d. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tateishi-Karimata, H.; Ohyama, T.; Muraoka, T.; Tanaka, S.; Kinbara, K.; Sugimoto, N. New Modified Deoxythymine with Dibranched Tetraethylene Glycol Stabilizes G-Quadruplex Structures. Molecules 2020, 25, 705. https://doi.org/10.3390/molecules25030705
Tateishi-Karimata H, Ohyama T, Muraoka T, Tanaka S, Kinbara K, Sugimoto N. New Modified Deoxythymine with Dibranched Tetraethylene Glycol Stabilizes G-Quadruplex Structures. Molecules. 2020; 25(3):705. https://doi.org/10.3390/molecules25030705
Chicago/Turabian StyleTateishi-Karimata, Hisae, Tatsuya Ohyama, Takahiro Muraoka, Shigenori Tanaka, Kazushi Kinbara, and Naoki Sugimoto. 2020. "New Modified Deoxythymine with Dibranched Tetraethylene Glycol Stabilizes G-Quadruplex Structures" Molecules 25, no. 3: 705. https://doi.org/10.3390/molecules25030705
APA StyleTateishi-Karimata, H., Ohyama, T., Muraoka, T., Tanaka, S., Kinbara, K., & Sugimoto, N. (2020). New Modified Deoxythymine with Dibranched Tetraethylene Glycol Stabilizes G-Quadruplex Structures. Molecules, 25(3), 705. https://doi.org/10.3390/molecules25030705





