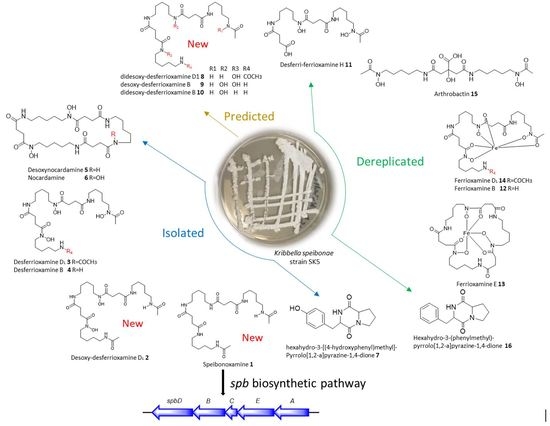
Novel South African Rare Actinomycete Kribbella speibonae Strain SK5: A Prolific Producer of Hydroxamate Siderophores Including New Dehydroxylated Congeners
Abstract
1. Introduction
2. Results and Discussion
2.1. Structure Elucidation
2.2. Molecular Network Analysis
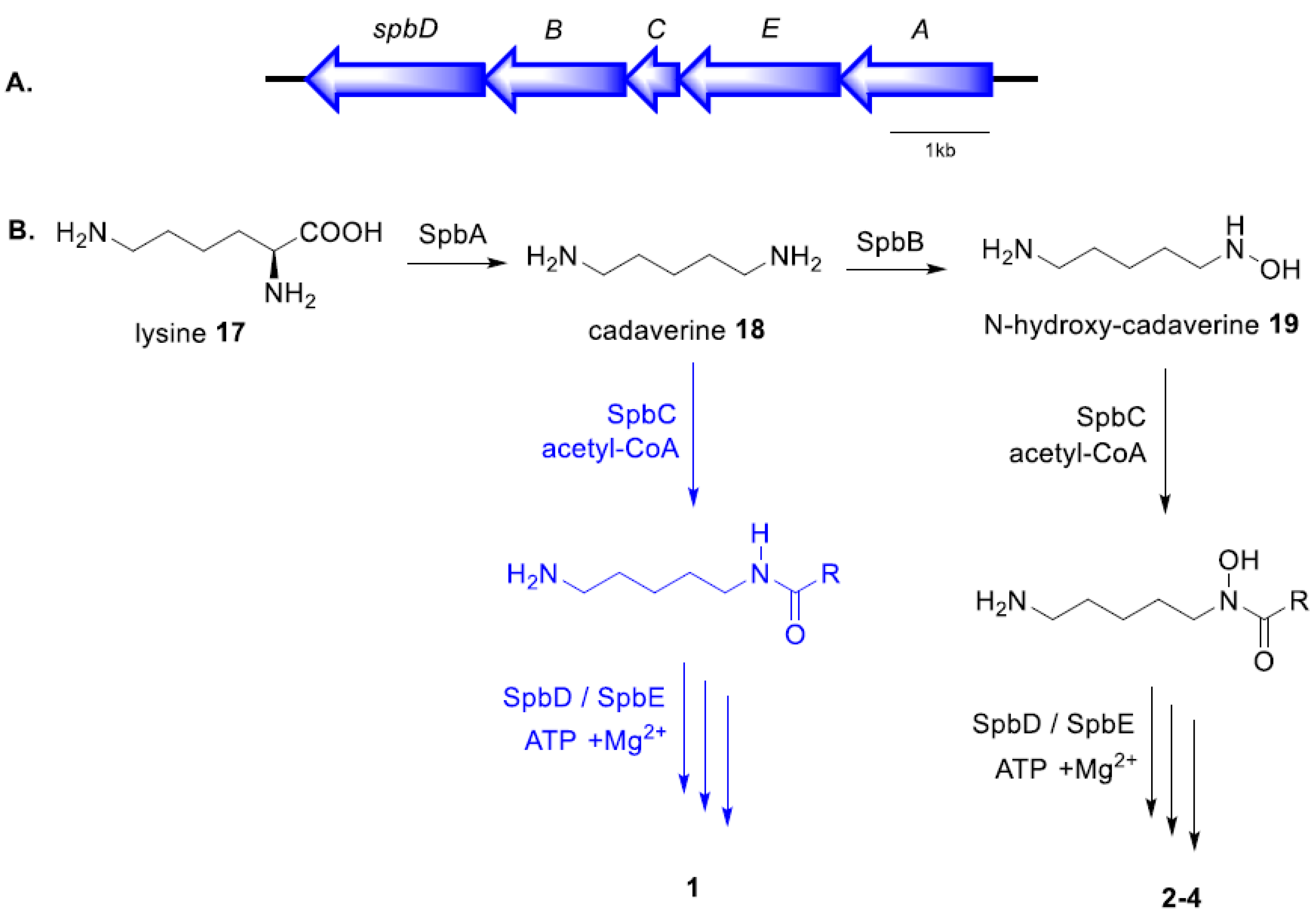
2.3. Proposed Biosynthetic Pathway
3. Materials and Methods
3.1. General Experimental Procedures
3.2. Isolation and Characterization of the Strain
3.3. Fermentation
3.4. HPLC-DAD/HRESIMS Analyses
3.5. Fractionation, Isolation, and Purification of Compounds
3.6. Molecular Networking
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Butler, A.; Theisen, R.M. Iron(III)–siderophore coordination chemistry: Reactivity of marine siderophores. Coord. Chem Rev. 2011, 254, 288–296. [Google Scholar] [CrossRef]
- Weinberg, E.D. Suppression of bacterial biofilm formation by iron limitation. Med. Hypotheses 2004, 63, 863–865. [Google Scholar] [CrossRef] [PubMed]
- Hider, R.C.; Kong, X. Chemistry and biology of siderophores. Nat. Prod. Rep. 2010, 27, 637. [Google Scholar] [CrossRef] [PubMed]
- Ahmed, E.; Holmström, S.J.M. Siderophores in environmental research: Roles and applications. Microb. Biotechnol. 2014, 7, 196–208. [Google Scholar] [CrossRef]
- Rajkumar, M.; Ae, N.; Narasimha, M.; Prasad, V.; Freitas, H. Potential of siderophore-producing bacteria for improving heavy metal phytoextraction. Trends Biotechnol. 2010, 28, 142–149. [Google Scholar] [CrossRef]
- Wu, J.Y.; Srinivas, P.; Pogue, J.M. Cefiderocol: A Novel Agent for the Management of Multidrug-Resistant Gram-Negative Organisms. Infect Dis. Ther. 2020, 9, 17–40. [Google Scholar] [CrossRef]
- Miller, M.J.; Walz, A.J.; Zhu, H.; Wu, C.; Moraski, G.; Ute, M.; Tristani, E.M.; Crumbliss, A.L.; Ferdig, M.T.; Checkley, L.; et al. Design, Synthesis, and Study of a Mycobactin-Artemisinin Conjugate That Has Selective and Potent Activity against Tuberculosis and Malaria. J. Am. Chem. Soc. 2011, 133, 2076–2079. [Google Scholar] [CrossRef] [PubMed]
- Braun, V.; Pramanik, A.; Gwinner, T.; Köberle, M.; Bohn, E. Sideromycins: Tools and antibiotics. BioMetals 2009, 22, 3–13. [Google Scholar] [CrossRef]
- Maglangit, F.; Alrashdi, S.; Renault, J.; Trembleau, L.; Victoria, C.; Tong, M.H.; Wang, S.; Kyeremeh, K.; Deng, H. Characterization of the promiscuous N-acyl CoA transferase, LgoC, in legonoxamine biosynthesis. Org. Biomol. Chem. 2020, 18, 2219–2222. [Google Scholar] [CrossRef]
- Carroll, C.S.; Moore, M.M. Ironing out siderophore biosynthesis: A review of non-ribosomal peptide synthetase (NRPS)-independent siderophore synthetases. Crit. Rev. Biochem. Mol. Biol. 2018, 53, 356–381. [Google Scholar] [CrossRef]
- Maglangit, F.; Tong, M.H.; Jaspars, M.; Kyeremeh, K.; Deng, H. Legonoxamines A-B, two new hydroxamate siderophores from the soil bacterium, Streptomyces sp. MA37. Tetrahedron Lett. 2019, 60, 75–79. [Google Scholar] [CrossRef]
- Barry, S.M.; Challis, G.L. Recent advances in siderophore biosynthesis. Curr. Opin. Chem. Biol. 2009, 13, 205–215. [Google Scholar] [CrossRef] [PubMed]
- Pluháček, T.; Lemr, K.; Ghosh, D.; Milde, D.; Novák, J.; Havlíček, V. Characterization of microbial siderophores by mass spectrometry. Mass Spectrom. Rev. 2016, 35, 35–47. [Google Scholar] [CrossRef] [PubMed]
- Subramani, R.; Sipkema, D. Marine Rare Actinomycetes: A Promising Source of Structurally Diverse and Unique Novel Natural Products. Mar. Drugs 2019, 17, 249. [Google Scholar] [CrossRef] [PubMed]
- Igarashi, M.; Sawa, R.; Yamasaki, M.; Hayashi, C.; Umekita, M.; Hatano, M.; Fujiwara, T.; Mizumoto, K.; Nomoto, A. Kribellosides, novel RNA 5′-triphosphatase inhibitors from the rare actinomycete Kribbella sp. MI481-42F6. J. Antibiot. (Tokyo). 2017, 70, 582–589. [Google Scholar] [CrossRef] [PubMed]
- Parte, A.C. LPSN—List of prokaryotic names with standing in nomenclature (Bacterio.net), 20 years on. Int. J. Syst. Evol. Microbiol. 2018, 68, 1825–1829. [Google Scholar] [CrossRef] [PubMed]
- Curtis, S.M.; Norton, I.; Everest, G.J.; Pelser, J.G.; De Kock, M.C.; Meyers, P.R. Development of a Kribbella-specific isolation medium and description of Kribbella capetownensis sp. nov. and Kribbella speibonae sp. nov., isolated from soil. Antonie van Leeuwenhoek 2020, 113, 617–628. [Google Scholar] [CrossRef]
- Zhang, Q.W.; Lin, L.G.; Ye, W.C. Techniques for extraction and isolation of natural products: A comprehensive review. Chinese Med. (UK) 2018, 13, 1–26. [Google Scholar] [CrossRef]
- Feistner, G.J.; Stahl, D.C.; Ann, H. Proferrioxamine Siderophores of Erwinia Amylovora. A Capillary Liquid Chromatographic/Electrospray Tandem Mass Spectrometric Study. Org. Mass Spectrom. 1993, 28, 163–175. [Google Scholar] [CrossRef]
- Lee, H.S.; Hee, J.S.; Kyoung, H.J.; Tae, S.K.; Oh, K.B.; Shin, J. Cyclic peptides of the nocardamine class from a marine-derived bacterium of the genus Streptomyces. J. Nat. Prod. 2005, 68, 623–625. [Google Scholar] [CrossRef]
- Ueki, M.; Suzuki, R.; Takamatsu, S.; Takagi, H.; Uramoto, M.; Ikeda, H.; Osada, H. Nocardamin Production by Streptomyces avermitilis. Actinomycetologica 2009, 23, 34–39. [Google Scholar] [CrossRef]
- Fang, Q.; Maglangit, F.; Wu, L.; Ebel, R.; Kyeremeh, K.; Andersen, J.H.; Annang, F.; Pérez-Moreno, G.; Reyes, F.; Deng, H. Signalling and Bioactive Metabolites from Streptomyces sp. RK44. Molecules 2020, 25, 460. [Google Scholar] [CrossRef] [PubMed]
- Wang, M.; Carver, J.J.; Phelan, V.V.; Sanchez, L.M.; Garg, N.; Peng, Y.; Nguyen, D.D.; Watrous, J.; Kapono, C.A.; Luzzatto-Knaan, T.; et al. Sharing and community curation of mass spectrometry data with Global Natural Products Social Molecular Networking. Nat. Biotechnol. 2016, 34, 828–837. [Google Scholar] [CrossRef] [PubMed]
- Pukall, R.; Lapidus, A.; Del Rio, T.G.; Copeland, A.; Tice, H.; Cheng, J.F.; Lucas, S.; Chen, F.; Nolan, M.; Labutti, K.; et al. Complete genome sequence of Kribbella flavida type strain (IFO 14399 T). Stand. Genomic Sci. 2010, 2, 186–193. [Google Scholar] [PubMed]
- Ronan, J.L.; Kadi, N.; Mcmahon, S.A.; Naismith, J.H.; Alkhalaf, L.M.; Challis, G.L.; Challis, G.L. Desferrioxamine biosynthesis: Diverse hydroxamate assembly by substrate- tolerant acyl transferase DesC. R. Soc. Publ. 2018, 373. [Google Scholar] [CrossRef] [PubMed]
- Kadi, N.; Song, L.; Challis, G.L. Bisucaberin biosynthesis: An adenylating domain of the BibC multi-enzyme catalyzes cyclodimerization of N-hydroxy-N-succinylcadaverine. Chem. Commun. 2008, 5119–5121. [Google Scholar] [CrossRef]
- Barona-Gómez, F.; Wong, U.; Giannakopulos, A.E.; Derrick, P.J.; Challis, G.L. Identification of a cluster of genes that directs desferrioxamine biosynthesis in Streptomyces coelicolor M145. J. Am. Chem. Soc. 2004, 126, 16282–16283. [Google Scholar] [CrossRef]
- Rütschlin, S.; Böttcher, T. Dissecting the Mechanism of Oligomerization and Macrocyclization Reactions of NRPS-Independent Siderophore Synthetases. Chem. A Eur. J. 2018, 24, 16044–16051. [Google Scholar] [CrossRef]
- Shirling, E.B.; Gottlieb, D. Methods for Characterization of Streptomyces species. Int. J. Syst. Bacteriol. 1966, 16, 313–340. [Google Scholar] [CrossRef]
- Holman, J.D.; Tabb, D.L.; Mallick, P. Employing ProteoWizard to Convert Raw Mass Spectrometry Data. Curr Protoc Bioinforma. 2014, 46, 1–7. [Google Scholar] [CrossRef]
- Otasek, D.; Morris, J.H.; Bouças, J.; Pico, A.R.; Demchak, B. Cytoscape Automation: Empowering workflow-based network analysis. Genome Biol. 2019, 20, 1–15. [Google Scholar] [CrossRef] [PubMed]
Sample Availability: Samples of the compounds are not available from the authors. |


| Protein | Annotated Function | Streptomyces Homologue% Identity/% Similarity | Amino Acid Length |
|---|---|---|---|
| SpbD | IucA/IucC synthetase | 74%/83% | 565 |
| SpbB | Lysine-6-monooxygenase | 80%/88% | 418 |
| SpbC | Acyltransferase | 67%/76% | 156 |
| SpbE | Siderophore synthetase | 78%/84% | 546 |
| SpbA | PLP-dependent decarboxylase | 86%/93% | 495 |
| Speibonoxamine 1 | Desoxy-Desferrioxamine D1 2 | |||
|---|---|---|---|---|
| Position | 13C | 1H, Mult. (J, Hz) | 13C | 1H, Mult. (J, Hz) |
| 1, 1′ | 23.1, CH3 | 1.77, s | 23.1, CH3 | 1.78, s |
| 2, 2′ | 169.4, C | - | 169.4, C | - |
| 3, 3′, 3” | 38.9, CH2 | 3.00, t (6.15) | 38.9, CH2 | 3.00, m |
| 4, 4′, 4” | 29.3, CH2 | 1.36, m | 29.3, CH2 | 1.38, dd (7.14, 14.24) |
| 5, 5′, 5” | 24.3, CH2 | 1.22, m | 24.0, CH2 | 1.23, m |
| 6, 6′ | 29.3, CH2 | 1.36, m | 26.5, CH2 | 1.50, m |
| 6” | 29.3, CH2 | 1.36, m | 29.3, CH2 | 1.38, dd (7.14, 14.24) |
| 7, 7′ | 38.9, CH2 | 3.00, t (6.15) | 47.6, CH2 | 3.45, t (6.96, 6.96) |
| 7” | 38.9, CH2 | 3.00, t (6.15) | 38.9, CH2 | 3.00, m |
| 8 | 171.7, C | - | 172.4, C | - |
| 8′ | 171.7, C | - | 171.7, C | - |
| 9 | 31.4, CH2 | 2.27, s | 28.1, CH2 | 2.58, m |
| 9′ | 31.4, CH2 | 2.27, s | 29.8, CH2 | 2.40, t (6.89, 6.89) |
| 10 | 31.4, CH2 | 2.27, s | 31.4, CH2 | 2.27, m |
| 10′ | 31.4, CH2 | 2.27, s | 30.4, CH2 | 2.28, m |
| 11 | 171.7, C | - | 174.4, C | - |
| 11′ | 171.7, C | - | 171.2, C | - |
| NH | 7.77 | 7.77 | ||
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Acquah, K.S.; Beukes, D.R.; Warner, D.F.; Meyers, P.R.; Sunassee, S.N.; Maglangit, F.; Deng, H.; Jaspars, M.; Gammon, D.W. Novel South African Rare Actinomycete Kribbella speibonae Strain SK5: A Prolific Producer of Hydroxamate Siderophores Including New Dehydroxylated Congeners. Molecules 2020, 25, 2979. https://doi.org/10.3390/molecules25132979
Acquah KS, Beukes DR, Warner DF, Meyers PR, Sunassee SN, Maglangit F, Deng H, Jaspars M, Gammon DW. Novel South African Rare Actinomycete Kribbella speibonae Strain SK5: A Prolific Producer of Hydroxamate Siderophores Including New Dehydroxylated Congeners. Molecules. 2020; 25(13):2979. https://doi.org/10.3390/molecules25132979
Chicago/Turabian StyleAcquah, Kojo Sekyi, Denzil R. Beukes, Digby F. Warner, Paul R. Meyers, Suthananda N. Sunassee, Fleurdeliz Maglangit, Hai Deng, Marcel Jaspars, and David W. Gammon. 2020. "Novel South African Rare Actinomycete Kribbella speibonae Strain SK5: A Prolific Producer of Hydroxamate Siderophores Including New Dehydroxylated Congeners" Molecules 25, no. 13: 2979. https://doi.org/10.3390/molecules25132979
APA StyleAcquah, K. S., Beukes, D. R., Warner, D. F., Meyers, P. R., Sunassee, S. N., Maglangit, F., Deng, H., Jaspars, M., & Gammon, D. W. (2020). Novel South African Rare Actinomycete Kribbella speibonae Strain SK5: A Prolific Producer of Hydroxamate Siderophores Including New Dehydroxylated Congeners. Molecules, 25(13), 2979. https://doi.org/10.3390/molecules25132979