Ginkgo biloba Extract Protects against Methotrexate-Induced Hepatotoxicity: A Computational and Pharmacological Approach
Abstract
1. Introduction
2. Material and Methods
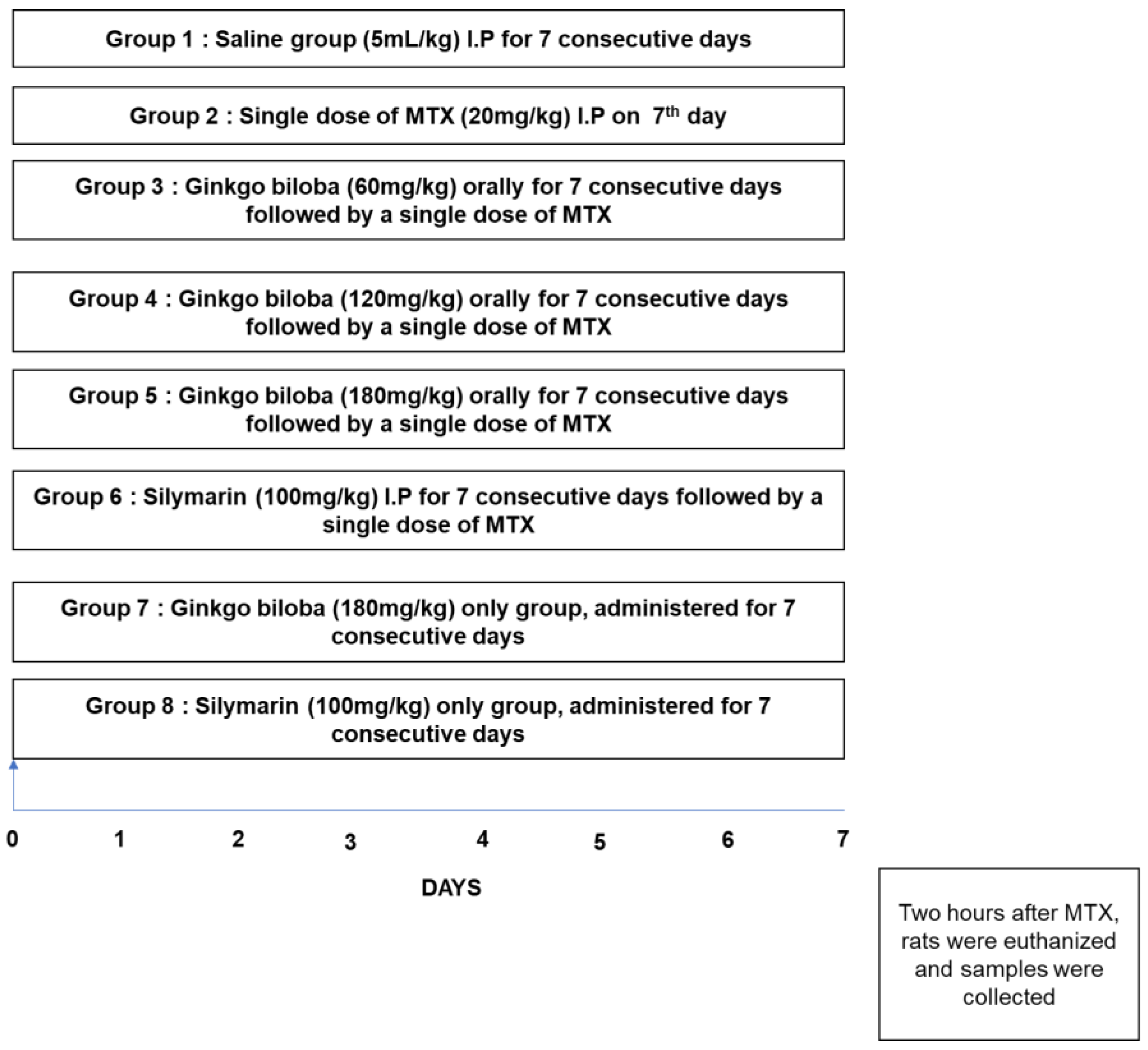
2.1. Animals and Experimental Design
2.2. Hematoxylin and Eosin (H&E) Staining
2.3. Serum Biomarkers Analysis
2.4. Oxidative Enzymes Analysis
2.5. Immunohistochemical Staining and Microscopic Analysis
2.6. Bioinformatics Resources
2.7. Statistical Analysis
3. Results
3.1. Effect of Gb on Liver Biochemical Markers
3.2. Effect of Gb on Fatty Acid Levels
3.3. Effect of Gb on Hepatic Oxidative Stress
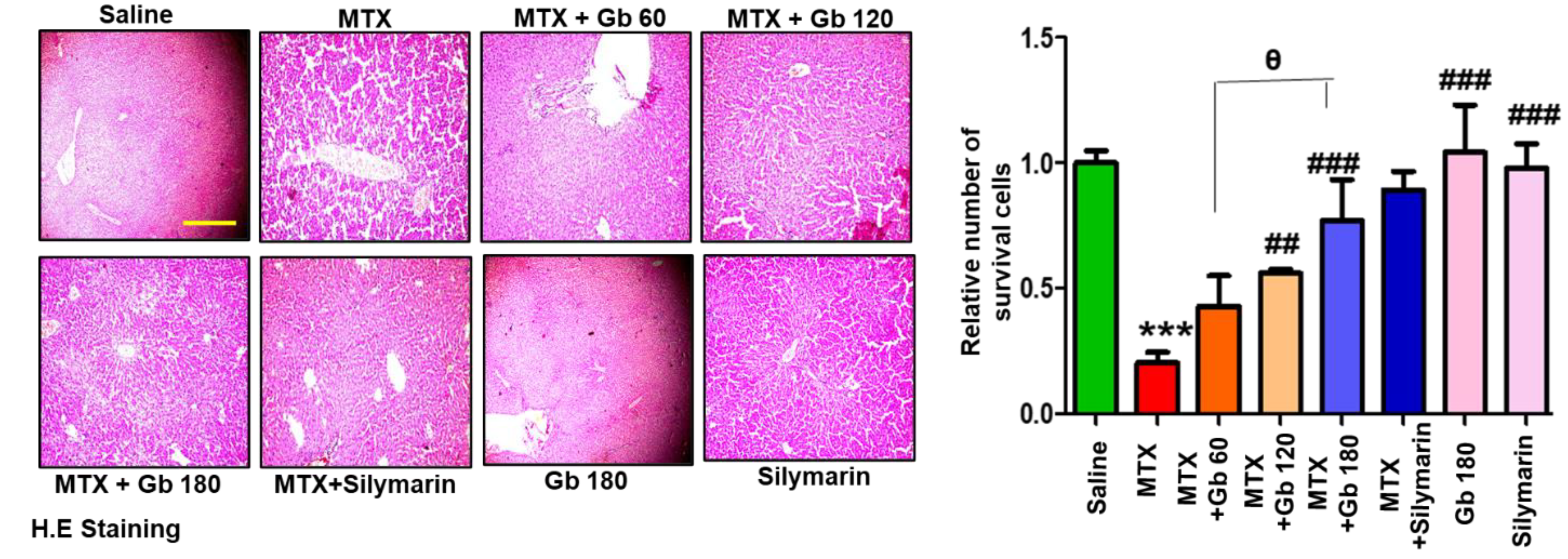
3.4. Effect of Gb Liver Morphology
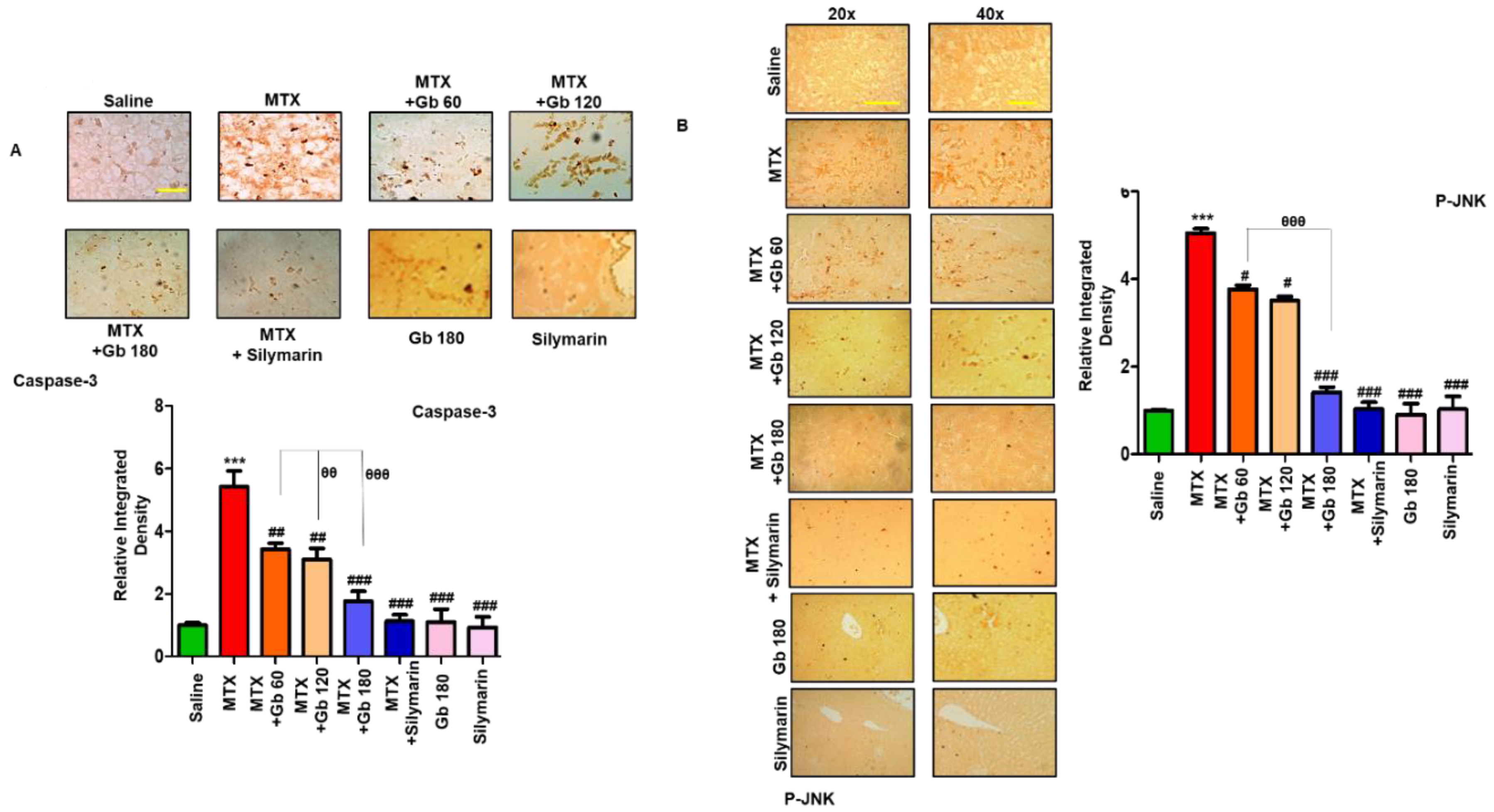
3.5. Gb Attenuated MTX-Induced Liver Apoptosis
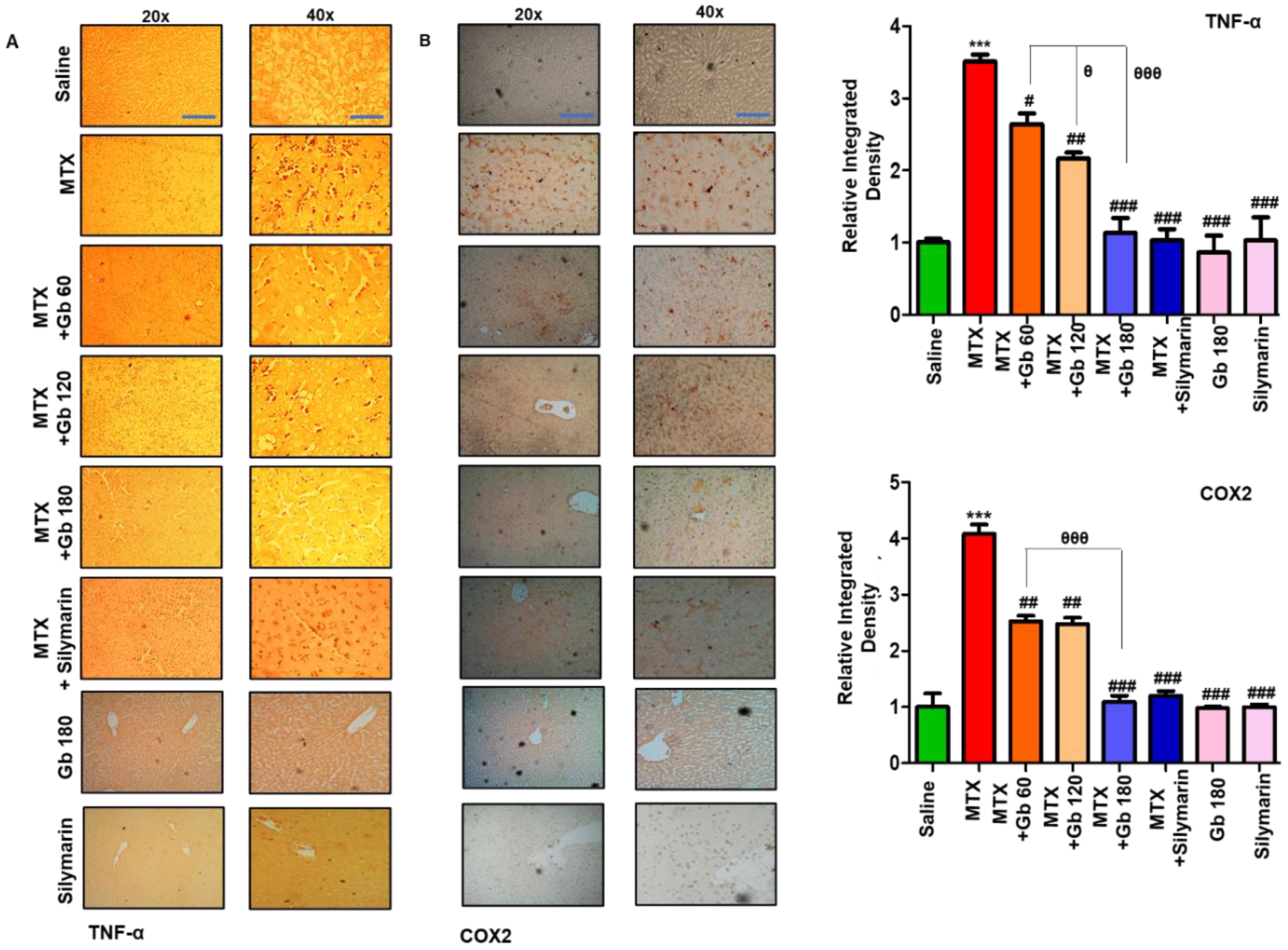
3.6. Gb Attenuated MTX-Induced Inflammatory Mediators in Liver
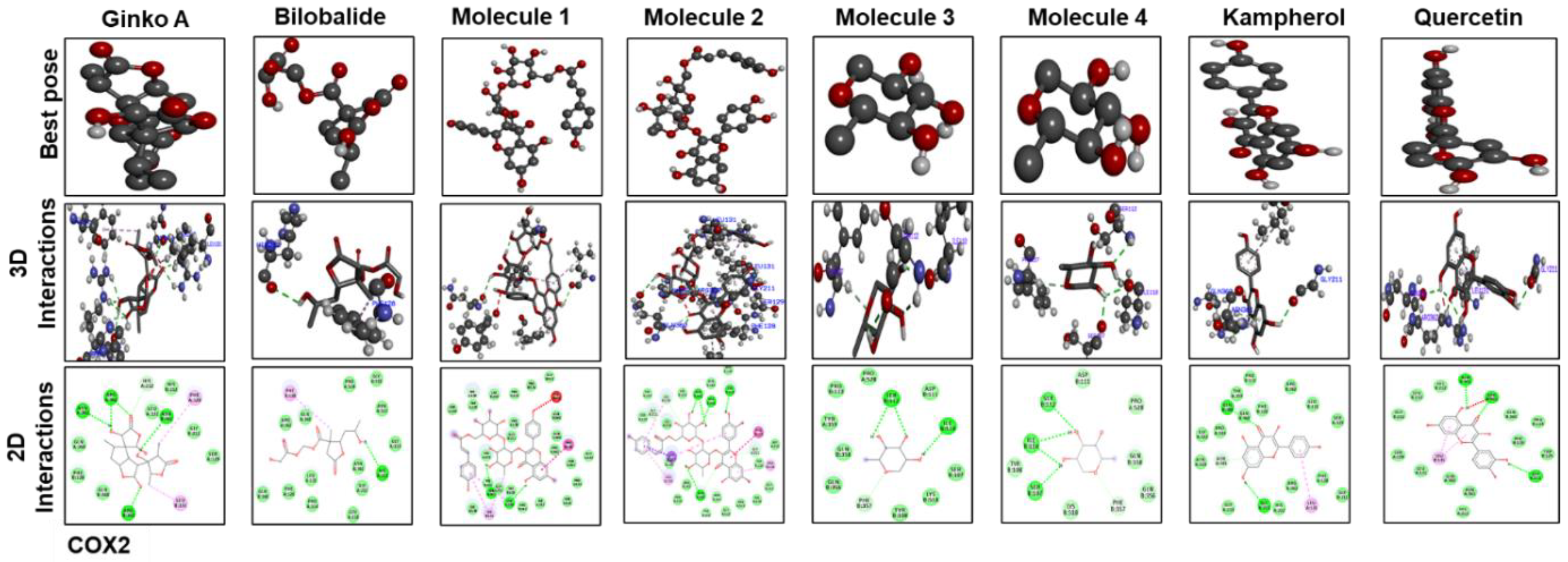
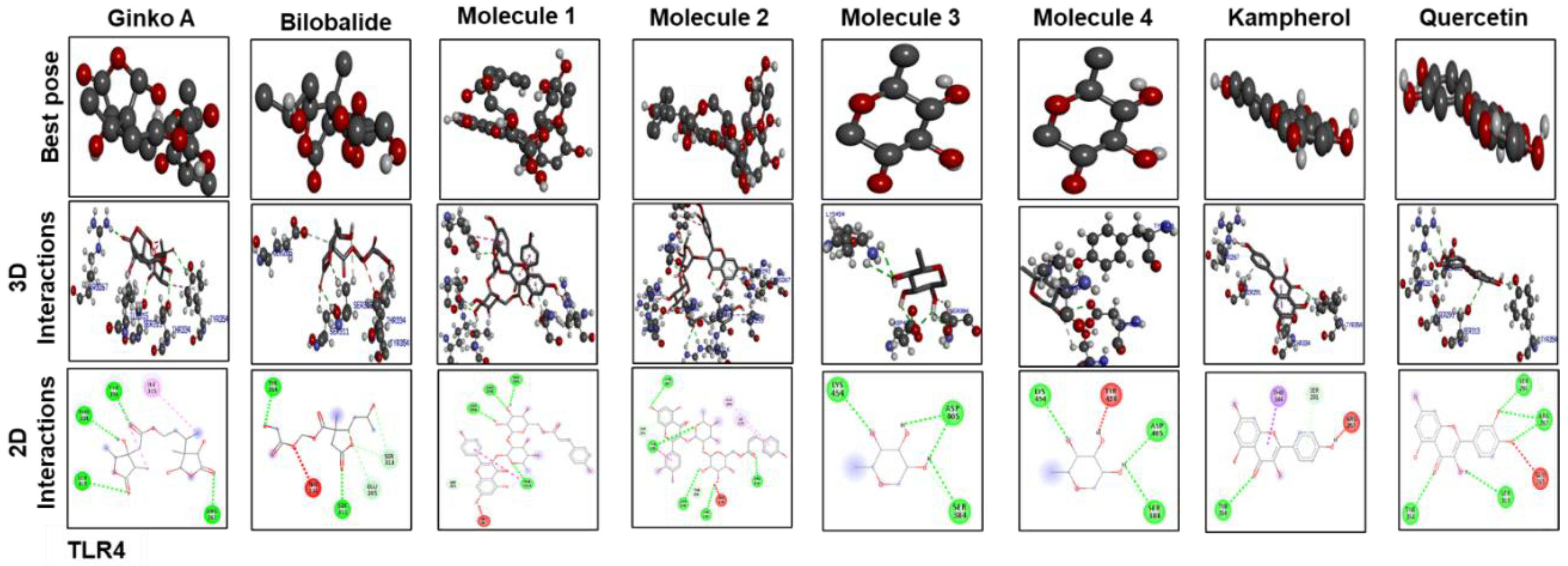
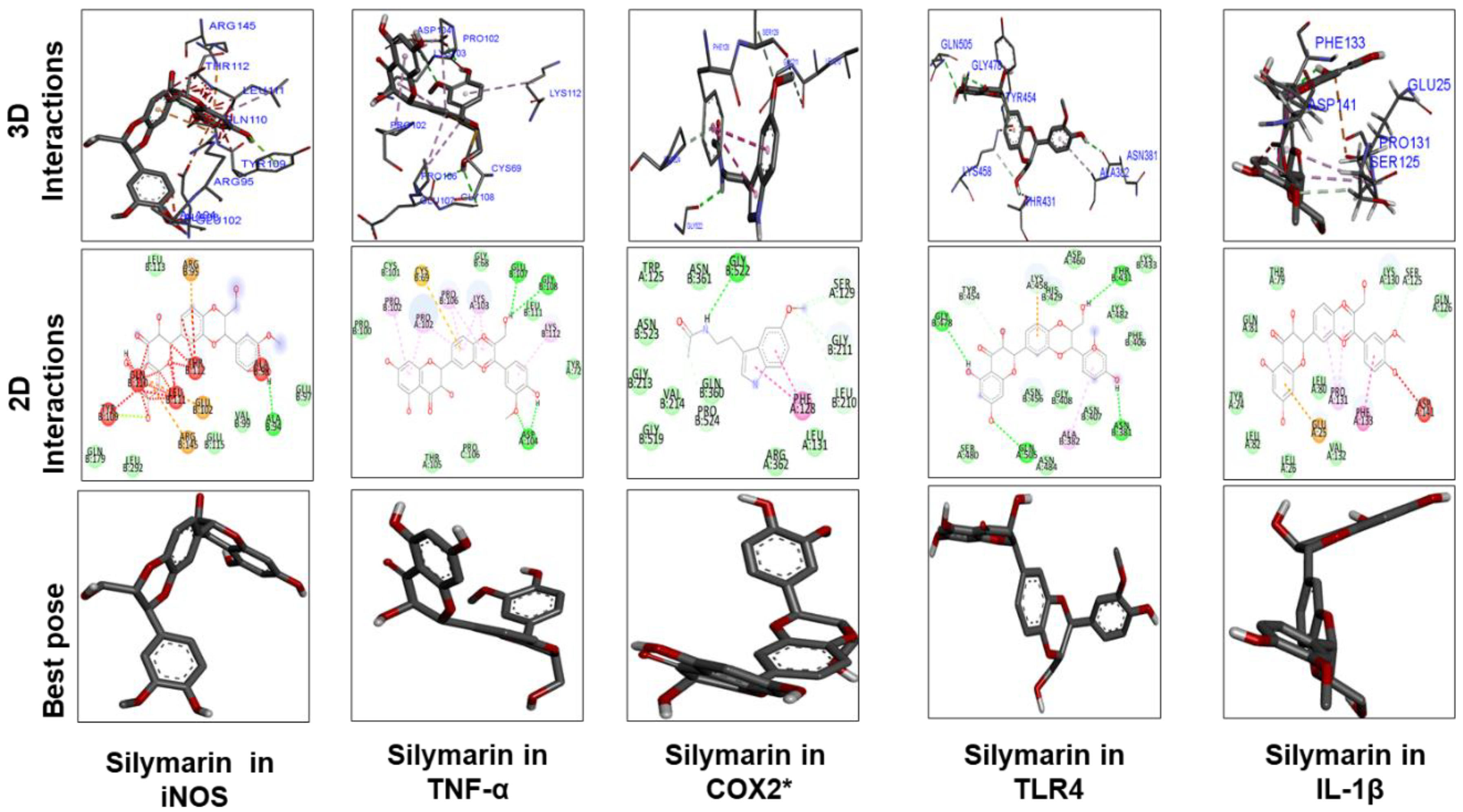
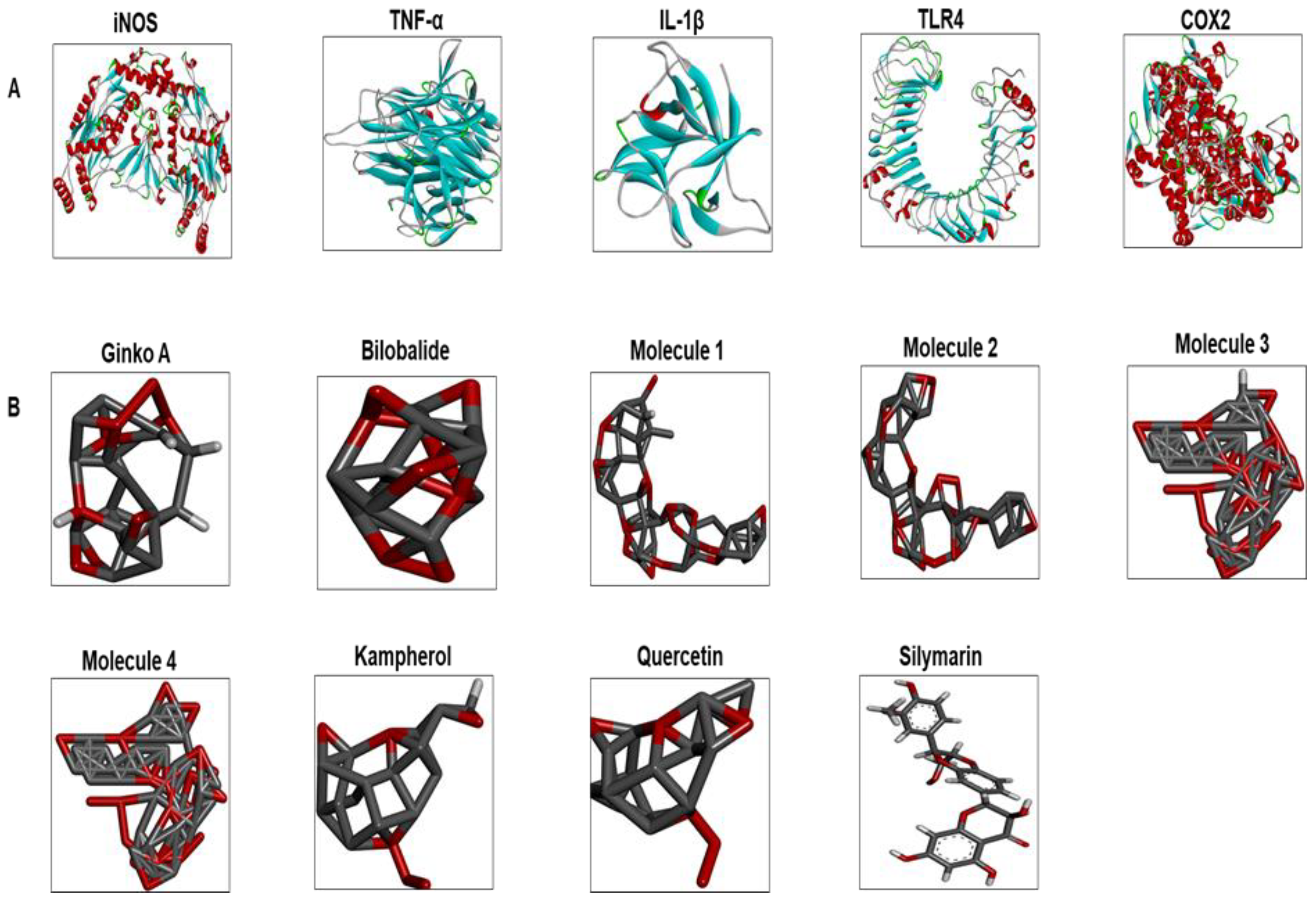
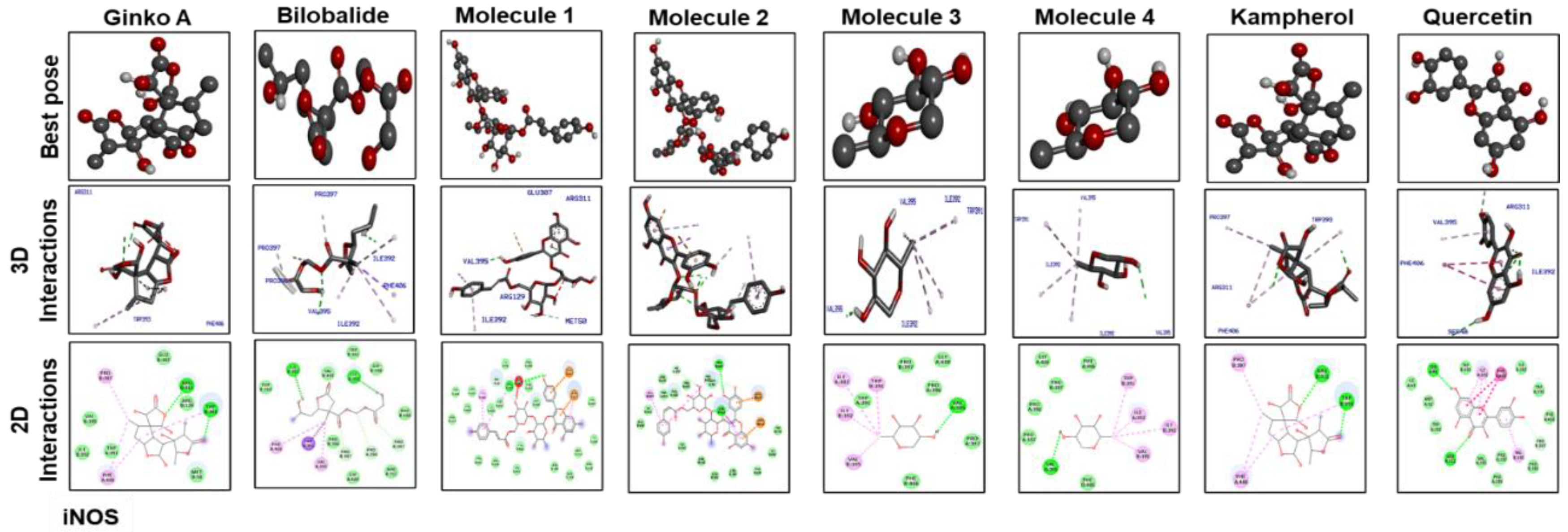
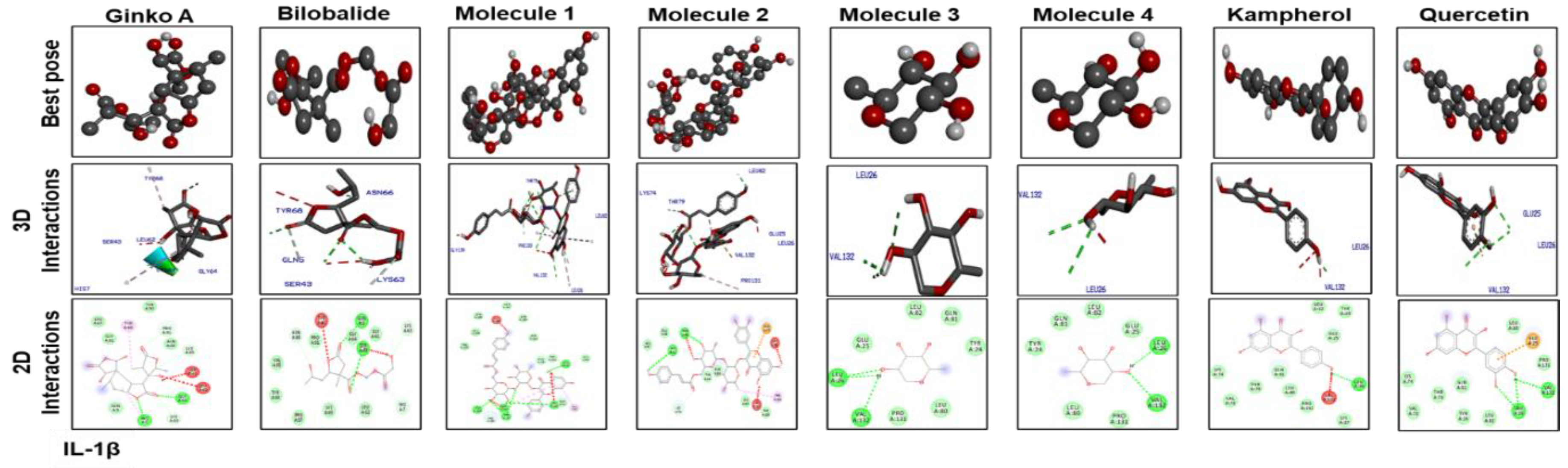
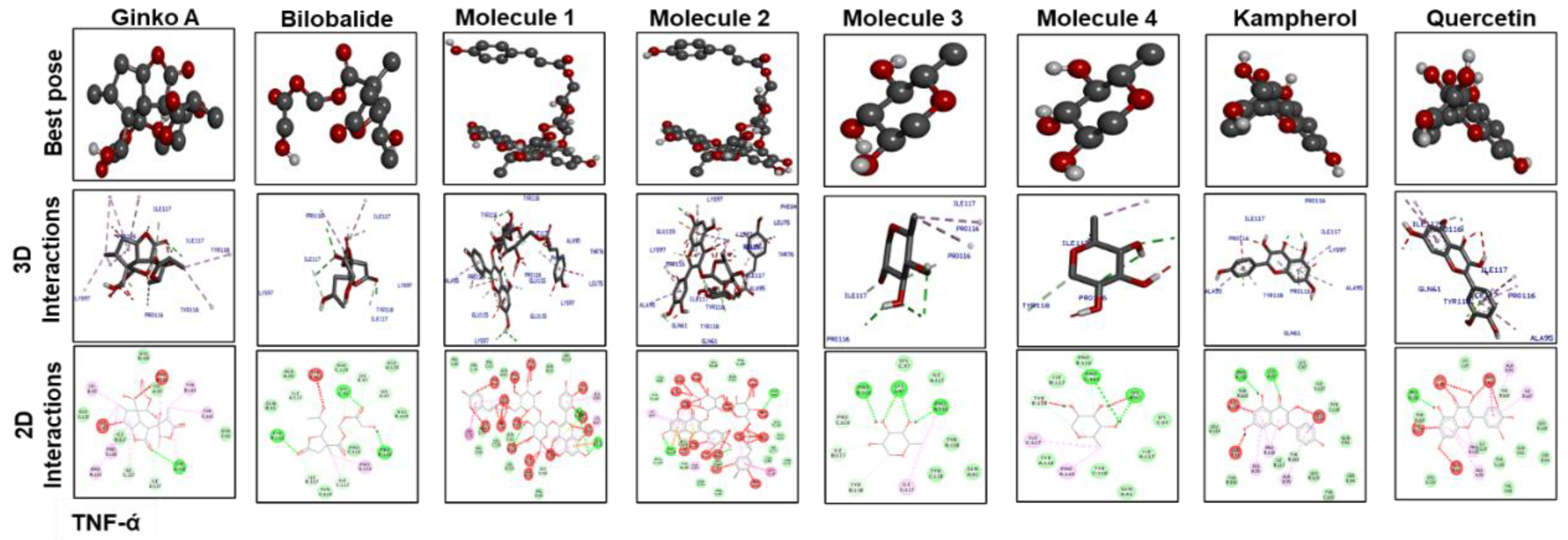
3.7. Docking Studies
| iNOS | IL1-β | TNFα | |||||||
|---|---|---|---|---|---|---|---|---|---|
| Ligand | Binding Energy | # of H Bonds | Residue | Binding Energy | # of H Bonds | Residue | Binding Energy | # of H Bonds | Residue |
| Ginkgolide | –10 | 2 | TRP393, ARG311 | –8.9 | 2 | GLY64, HIS7 | –7.8 | 1 | TYR118 |
| Bilobalide | –9.8 | 2 | ILE392,ILE392 | –9.1 | 2 | GLN5, SER43 | –7.9 | 3 | LYS97, PRO116, TYR118 |
| Molecule 1 | –9.1 | 1 | VAL395 | –6.3 | 6 | LEU26, VAL132, THR79, LEU(80)2, LEU134 | –7.4 | 3 | TYR118, GLU(115)2 |
| Molecule 2 | –8.7 | 3 | TRP393, ARG311(2) | –6.4 | 2 | THR79, LEU82 | –7 | 3 | GLU115, GLN(61)2 |
| Molecule 3 | –8.8 | 1 | VAL395 | –6.2 | 3 | LEU(20)2, VAL132 | –6.3 | 4 | PRO126, LYS(97)2, PRO116 |
| Molecule 4 | –7 | 1 | VAL395 | –6.6 | 2 | LEU20, VAL132 | –5.2 | 2 | PRO116, LYS97 |
| Kaempferol | –7.4 | 2 | ARG311, TRP393 | –6.5 | 1 | LEU26 | –7.2 | 2 | LYS97, PRO116 |
| Quercetin | –7.8 | 2 | ARG311, SER48 | –6.8 | 3 | VAL132, LEU(26)2 | –8.9 | 1 | PRO116 |
| Silymarin | –9.6 | 1 | ALA94 | –8 | 1 | ASP141 | –9.1 | 4 | GLY108, GLU 107, ASP(164)2 |
| COX2 | TLR4 | |||||
|---|---|---|---|---|---|---|
| Ligand | Binding Energy | # of H Bonds | Residue | Binding Energy | # of H Bonds | Residue |
| Ginkgolide | –6.9 | 4 | ASN(361)2, ARG(3622) | –7.9 | 4 | ARG267, SER313, THR334, TYR354 |
| Bilobalide | –7.3 | 1 | HIS212 | –8.1 | 2 | TYR354, SER311 |
| Molecule 1 | –6.7 | 2 | LEU210, HIS212 | –7.3 | 4 | TYR354, ASP350, ARG359, THR336 |
| Molecule 2 | –8.9 | 5 | ARG362, ASN(361)2, SER129, GLN360 | –8.4 | 5 | ARG267, TYR354, ASP356, SER358, ARG316 |
| Molecule 3 | –6.3 | 3 | SER(112)2, ILE110 | –7.2 | 4 | LYS454, ASP(405)2, SER 384 |
| Molecule 4 | –7.8 | 4 | SER 112, ILE(110)2, SER 107 | –6.6 | 3 | ASP465, SER 384, LYS454 |
| Kaempferol | –6.8 | 1 | GLN 360 | –6.1 | 1 | TYR354 |
| Quercetin | –6.4 | 3 | GLY 211, ARG362, ASN 361 | –7.8 | 4 | TYR354, ARG(267)2, SER291 |
| Silymarin | –5.8 | 1 | GLY522 | –8 | 4 | THR381, GLN505, GLY478, ASN 381 |
4. Discussion
Author Contributions
Funding
Conflicts of Interest
References
- Cichoż-Lach, H.; Michalak, A. Oxidative stress as a crucial factor in liver diseases. World J. Gastroenterol. 2014, 20, 8082–8091. [Google Scholar] [CrossRef] [PubMed]
- Dikici, I.; Mehmetoglu, I.; Dikici, N.; Bitirgen, M.; Kurban, S. Investigation of oxidative stress and some antioxidants in patients with acute and chronic viral hepatitis B and the effect of interferon-α treatment. Clin. Biochem. 2005, 38, 1141–1144. [Google Scholar] [CrossRef] [PubMed]
- Abbès, S.; Ben Salah-Abbès, J.; Jebali, R.; Ben Younes, R.; Oueslati, R. Interaction of aflatoxin B1and fumonisin B1in mice causes immunotoxicity and oxidative stress: Possible protective role using lactic acid bacteria. J. Immunotoxicol. 2015, 13, 46–54. [Google Scholar] [CrossRef] [PubMed]
- Farinati, F.; Cardin, R.; De Maria, N.; Della Libera, G.; Marafin, C.; Lecis, E.; Burra, P.; Floreani, A.; Cecchetto, A.; Naccarato, R. Iron storage, lipid peroxidation and glutathione turnover in chronic anti-HCV positive hepatitis. J. Hepatol. 1995, 22, 449–456. [Google Scholar] [CrossRef]
- Hytiroglou, P.; Theise, N.D.; Schwartz, M.; Mor, E.; Miller, C.; Thung, S.N. Macroregenerative nodules in a series of adult cirrhotic liver explants: Issues of classification and nomenclature. Hepatology 1995, 21, 703–708. [Google Scholar] [PubMed]
- Khan, N.; Abbas, A.; Whang, N.; Balart, L.A.; Bazzano, L.A.; Kelly, T.N. Incidence of Liver Toxicity in Inflammatory Bowel Disease Patients Treated with Methotrexate: A Meta-Analysis of Clinical Trials. Inflamm. Bowel Dis. 2012, 18, 359–367. [Google Scholar] [CrossRef] [PubMed]
- Dalaklioglu, S.; Genc, G.; Aksoy, N.; Akcit, F.; Gumuslu, S. Resveratrol ameliorates methotrexate-induced hepatotoxicity in rats via inhibition of lipid peroxidation. Hum. Exp. Toxicol. 2013, 32, 662–671. [Google Scholar] [CrossRef] [PubMed]
- Singh, D.; Cho, W.C.; Upadhyay, G. Drug-Induced Liver Toxicity and Prevention by Herbal Antioxidants: An Overview. Front. Physiol. 2016, 6, 1397. [Google Scholar] [CrossRef]
- Darwish, S.F.; El-Bakly, W.M.; Arafa, H.M.; El-Dermerdash, E. Targeting TNF-α and NF-κB Activation by Bee Venom: Role in Suppressing Adjuvant Induced Arthritis and Methotrexate Hepatotoxicity in Rats. PLoS ONE 2013, 8, e79284. [Google Scholar] [CrossRef]
- Federico, A.; Dallio, M.; Loguercio, C. Silymarin/Silybin and Chronic Liver Disease: A Marriage of Many Years. Molecules 2017, 22, 191. [Google Scholar] [CrossRef]
- Song, Z.; Deaciuc, I.; Song, M.; Lee, D.Y.-W.; Liu, Y.; Ji, X.; McClain, C. Silymarin Protects Against Acute Ethanol-Induced Hepatotoxicity in Mice. Alcohol. Clin. Exp. Res. 2006, 30, 407–413. [Google Scholar] [CrossRef] [PubMed]
- Cicero, A.F.G.; Fogacci, F.; Banach, M. Botanicals and phytochemicals active on cognitive decline: The clinical evidence. Pharmacol. Res. 2017, 130, 204–212. [Google Scholar] [CrossRef] [PubMed]
- Sung, J.-H.; Shah, F.-A.; Cho, E.-H.; Gim, S.-A.; Jeon, S.-J.; Kim, K.-M.; Kim, Y.-M.; Kim, M.-O.; Koh, P.-O. Ginkgo bilobaextract (EGb 761) prevents the ischemic brain injury-induced decrease in parvalbumin expression. Lab. Anim. Res. 2012, 28, 77–82. [Google Scholar] [CrossRef] [PubMed]
- Van Beek, T.A.; Montoro, P. Chemical analysis and quality control of Ginkgo biloba leaves, extracts, and phytopharmaceuticals. J. Chromatogr. A 2009, 1216, 2002–2032. [Google Scholar] [CrossRef] [PubMed]
- Jiang, F.; Dusting, G.J. Natural phenolic compounds as cardiovascular therapeutics: Potential role of their antiinflammatory effects. Curr. Vasc. Pharmacol. 2003, 1, 135–156. [Google Scholar] [CrossRef] [PubMed]
- Salminen, A.; Lehtonen, M.; Suuronen, T.; Kaarniranta, K.; Huuskonen, J. Terpenoids: Natural inhibitors of NF-kappaB signaling with anti-inflammatory and anticancer potential. Cell Mol. Life Sci. 2008, 65, 2979–2999. [Google Scholar] [CrossRef]
- Parimoo, H.A.; Sharma, R.; Patil, R.D.; Sharma, O.P.; Kumar, P.; Kumar, N. Hepatoprotective effect of Ginkgo biloba leaf extract on lantadenes-induced hepatotoxicity in guinea pigs. Toxicon 2014, 81, 1–12. [Google Scholar] [CrossRef]
- Shah, F.-A.; Park, D.-J.; Koh, P.-O. Identification of Proteins Differentially Expressed by Quercetin Treatment in a Middle Cerebral Artery Occlusion Model: A Proteomics Approach. Neurochem. Res. 2018, 43, 1608–1623. [Google Scholar] [CrossRef]
- Ali, A.; Shah, F.A.; Zeb, A.; Malik, I.; Alvi, A.M.; AlKury, L.T.; Rashid, S.; Hussain, I.; Ullah, N.; Khan, A.U.; et al. NF-κB Inhibitors Attenuate MCAO Induced Neurodegeneration and Oxidative Stress-A Reprofiling Approach. Front. Mol. Neurosci. 2020, 13, 33. [Google Scholar] [CrossRef]
- Al Kury, L.T.; Zeb, A.; Abidin, Z.U.; Irshad, N.; Malik, I.; Alvi, A.M.; Khalil, A.A.K.; Ahmad, S.; Faheem, M.; Khan, A.-U.; et al. Neuroprotective effects of melatonin and celecoxib against ethanol-induced neurodegeneration: A computational and pharmacological approach. Drug Des. Dev. Ther. 2019, 13, 2715–2727. [Google Scholar] [CrossRef]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
- Wiederstein, M.; Sippl, M.J. ProSA-web: Interactive web service for the recognition of errors in three-dimensional structures of proteins. Nucleic Acids Res. 2007, 35, W407–W410. [Google Scholar] [CrossRef] [PubMed]
- Ghadir, M.R.; Riahin, A.A.; Havaspour, A.; Nooranipour, M.; Habibinejad, A.A. The relationship between lipid profile and severity of liver damage in cirrhotic patients. Zahedan J. Res. Med. Sci. 2010, 10, 285–288. [Google Scholar]
- Wada, T.; Penninger, J.M. Mitogen-activated protein kinases in apoptosis regulation. Oncogene 2004, 23, 2838–2849. [Google Scholar] [CrossRef] [PubMed]
- Jin, Z.; El-Deiry, W. Overview of cell death signaling pathways. Cancer Boil. Ther. 2005, 4, 147–171. [Google Scholar] [CrossRef]
- Hollville, E.; Romero, S.E.; Deshmukh, M. Apoptotic cell death regulation in neurons. FEBS J. 2019, 286, 3276–3298. [Google Scholar] [CrossRef]
- Li, X.; Jiang, S.; Tapping, R.I. Toll-like receptor signaling in cell proliferation and survival. Cytokine 2009, 49, 1–9. [Google Scholar] [CrossRef]
- Sass, G.; Koerber, K.; Bang, R.; Guehring, H.; Tiegs, G. Inducible nitric oxide synthase is critical for immune-mediated liver injury in mice. J. Clin. Investig. 2001, 107, 439–447. [Google Scholar] [CrossRef]
- Morgan, S.L.; Oster, R.; Lee, J.Y.; Alarcón, G.S.; Baggott, J.E. The effect of folic acid and folinic acid supplements on purine metabolism in methotrexate-treated rheumatoid arthritis. Arthritis Rheum. 2004, 50, 3104–3111. [Google Scholar] [CrossRef]
- Hashkes, P.J.; Becker, M.L.; Cabral, D.A.; Laxer, R.M.; Paller, A.S.; Rabinovich, C.E.; Turner, D.; Zulian, F. Methotrexate: New Uses for an Old Drug. J. Pediatr. 2014, 164, 231–236. [Google Scholar] [CrossRef]
- Lee, M.-H.; Hong, I.; Kim, M.; Lee, B.-H.; Kim, J.-H.; Kang, K.-S.; Kim, H.-L.; Yoon, B.-I.; Chung, H.; Kong, G.; et al. Gene expression profiles of murine fatty liver induced by the administration of methotrexate. Toxicology 2008, 249, 75–84. [Google Scholar] [CrossRef] [PubMed]
- Çetin, A.; Kaynar, L.; Koçyigit, I.; Hacioglu, S.K.; Saraymen, R.; Ozturk, A.; Sari, I.; Sağdıç, O.; Sağdiç, O. Role of Grape Seed Extract on Methotrexate Induced Oxidative Stress in Rat Liver. Am. J. Chin. Med. 2008, 36, 861–872. [Google Scholar] [CrossRef] [PubMed]
- Cetinkaya, A.; Bulbuloglu, E.; Kurutaş, E.B.; Kantarceken, B. N-acetylcysteine ameliorates methotrexate-induced oxidative liver damage in rats. Med. Sci. Monit. 2006, 12, 274–278. [Google Scholar]
- Kucinski, I.; Dinan, M.; Kolahgar, G.; Piddini, E. Chronic activation of JNK JAK/STAT and oxidative stress signalling causes the loser cell status. Nat. Commun. 2017, 8, 136. [Google Scholar] [CrossRef] [PubMed]
- Sinha, K.; Das, J.; Pal, P.B.; Sil, P.C. Oxidative stress: The mitochondria-dependent and mitochondria-independent pathways of apoptosis. Arch. Toxicol. 2013, 87, 1157–1180. [Google Scholar] [CrossRef] [PubMed]
- Björkblom, B.; Vainio, J.C.; Hongisto, V.; Herdegen, T.; Courtney, M.; Coffey, E.T. All JNKs Can Kill, but Nuclear Localization Is Critical for Neuronal Death. J. Boil. Chem. 2008, 283, 19704–19713. [Google Scholar] [CrossRef] [PubMed]
- Spurlock, C.F.; Gass, H.M.; Bryant, C.; Wells, B.C.; Olsen, N.J.; Aune, T.M. Methotrexate-mediated inhibition of nuclear factor κB activation by distinct pathways in T cells and fibroblast-like synoviocytes. Rheumatology 2014, 54, 178–187. [Google Scholar] [CrossRef]
- Ho, L.-J.; Hung, L.-F.; Liu, F.-C.; Hou, T.-Y.; Lin, L.-C.; Huang, C.-Y.; Lai, J.-H. Ginkgo biloba Extract Individually Inhibits JNK Activation and Induces c-Jun Degradation in Human Chondrocytes: Potential Therapeutics for Osteoarthritis. PLoS ONE 2013, 8, e82033. [Google Scholar] [CrossRef]
- Gargouri, B.; Carstensen, J.; Bhatia, H.S.; Hüell, M.; Dietz, G.P.; Fiebich, B.L. Anti-neuroinflammatory effects of Ginkgo biloba extract EGb761 in LPS-activated primary microglial cells. Phytomedicine 2018, 44, 45–55. [Google Scholar] [CrossRef]
- Kotakadi, V.S.; Jin, Y.; Hofseth, A.B.; Ying, L.; Cui, X.; Volate, S.; Chumanevich, A.; Wood, P.A.; Price, R.L.; McNeal, A.; et al. Ginkgo biloba extract EGb 761 has anti-inflammatory properties and ameliorates colitis in mice by driving effector T cell apoptosis. Carcinogenesis 2008, 29, 1799–1806. [Google Scholar] [CrossRef]
- Chen, D.; Oezguen, N.; Urvil, P.; Ferguson, C.; Dann, S.; Savidge, T.C. Regulation of protein-ligand binding affinity by hydrogen bond pairing. Sci. Adv. 2016, 2, e1501240. [Google Scholar] [CrossRef] [PubMed]
Sample Availability: Samples of the compounds are not available from the authors. |
| Treatment | ALT | AST | TOTAL BILIRUBIN | ALKALINE PHOSPHATASE |
|---|---|---|---|---|
| Saline (5 mL/kg) | 22.0 ± 10 | 26.20 ± 0.58 | 0.78 ± 0.05 | 149 ± 18.09 |
| MTX (20 mg/kg) | 164.80 ± 4.25 *** | 272.2 ± 13.22 *** | 0.92 ± 0.06 * | 158.20 ± 6.23 |
| MTX + Gb (60 mg/kg) | 37.20 ± 1.65 ### | 132.40 ± 1.7 ### | 0.70 ± 0.03 | 127.40 ± 6.61 |
| MTX + Gb (120 mg/kg) | 27.80 ± 1.65 ### | 119.80 ± 1.56 ### | 0.82 ± 0.06 | 136.40 ± 9.41 |
| MTX + Gb (180 mg/kg) | 35.20 ± 6.26 ### | 151.80 ± 12.66 ### | 0.80 ± 0.03 | 166.20 ± 7.21 |
| MTX + Silymarin (100 mg/kg) | 67.80 ± 6.74 | 37.00 ± 9.77 | 0.74 ± 0.05 | 142.80 ± 11.93 |
| Treatment | Triglycerides | Cholesterol | LDL Cholesterol | HDL Cholesterol |
|---|---|---|---|---|
| Saline (5 mL/kg) | 86.40 ± 2.69 | 95.40 ± 6.70 | 31.20 ± 0.66 | 18.00 ± 1.14 |
| MTX (20 mg/kg) | 67.60 ± 1.28* | 70.00 ± 5.75* | 43.00 ± 4.46* | 14.40 ± 1.28* |
| MTX + Gb (60 mg/kg) | 117.40 ± 3.23# | 56.40 ± 2.92 | 19.20 ± 2.72## | 14.20 ± 1.06 |
| MTX + Gb (120 mg/kg) | 102.80 ± 8.49# | 55.80 ± 2.28 | 24.40 ± 2.24## | 11.20 ± 0.58 |
| MTX + Gb (180 mg/kg) | 70.20 ± 5.77 | 63.80 ± 3.51 | 37.00 ± 3.42 | 13.60 ± 0.70 |
| MTX + Silymarin (100 mg/kg) | 88.20 ± 5.67 | 74.80 ± 2.08 | 37.80 ± 3.30 | 15.00 ± 0.31 |
| Treatment | GST µmol CDNB Conjugate/min/mg of Protein | NO μmol/mg | GSH µmol/mg of Protein |
|---|---|---|---|
| Saline (5 mL/kg) | 55 ± 0.7 | 17 ± 2.5 | 85 ± 1.6 |
| MTX (20 mg/kg) | 8.6 ± 0.5*** | 100 ± 2.11*** | 30.3 ± 1.5*** |
| MTX + Gb (60 mg/kg) | 19.94 ± 0.13# | 92.18 ± 2.15 | 68 ± 0.5# |
| MTX + Gb (120 mg/kg) | 25.98 ± 0.3# | 78.28 ± 5.7## | 142 ± 6.8### |
| MTX + Gb (180 mg/kg) | 31.24 ± 0.6# | 58.30 ± 0.90### | 161.5 ± 1### |
| MTX + Silymarin (100 mg/kg) | 77.8 ± 0.5 | 20.567 ± 0.8 | 90.23 ± 0.3 |
| Histopathology | Saline | MTX | MTX+ Gb 60 | MTX+ Gb 120 | MTX+ Gb 180 | MTX+ Silymarin | Gb 180 | Silymarin |
|---|---|---|---|---|---|---|---|---|
| Hepatoportal and Sinusoidal Cogestion | − | ++ | + | +/− | − | − | − | − |
| Apoptosis | − | +++ | ++ | + | + | − | − | − |
| Necrotic damage | − | ++ | + | − | − | − | − | − |
| Inflammatory Infiltrate | − | ++ | + | − | − | +/− | − | − |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Al Kury, L.T.; Dayyan, F.; Ali Shah, F.; Malik, Z.; Khalil, A.A.K.; Alattar, A.; Alshaman, R.; Ali, A.; Khan, Z. Ginkgo biloba Extract Protects against Methotrexate-Induced Hepatotoxicity: A Computational and Pharmacological Approach. Molecules 2020, 25, 2540. https://doi.org/10.3390/molecules25112540
Al Kury LT, Dayyan F, Ali Shah F, Malik Z, Khalil AAK, Alattar A, Alshaman R, Ali A, Khan Z. Ginkgo biloba Extract Protects against Methotrexate-Induced Hepatotoxicity: A Computational and Pharmacological Approach. Molecules. 2020; 25(11):2540. https://doi.org/10.3390/molecules25112540
Chicago/Turabian StyleAl Kury, Lina Tariq, Fazli Dayyan, Fawad Ali Shah, Zulkifal Malik, Atif Ali Khan Khalil, Abdullah Alattar, Reem Alshaman, Amjad Ali, and Zahid Khan. 2020. "Ginkgo biloba Extract Protects against Methotrexate-Induced Hepatotoxicity: A Computational and Pharmacological Approach" Molecules 25, no. 11: 2540. https://doi.org/10.3390/molecules25112540
APA StyleAl Kury, L. T., Dayyan, F., Ali Shah, F., Malik, Z., Khalil, A. A. K., Alattar, A., Alshaman, R., Ali, A., & Khan, Z. (2020). Ginkgo biloba Extract Protects against Methotrexate-Induced Hepatotoxicity: A Computational and Pharmacological Approach. Molecules, 25(11), 2540. https://doi.org/10.3390/molecules25112540






