Calcium-Dependent Protein Kinase Genes in Glycyrrhiza Uralensis Appear to be Involved in Promoting the Biosynthesis of Glycyrrhizic Acid and Flavonoids under Salt Stress
Abstract
:1. Introduction
2. Results
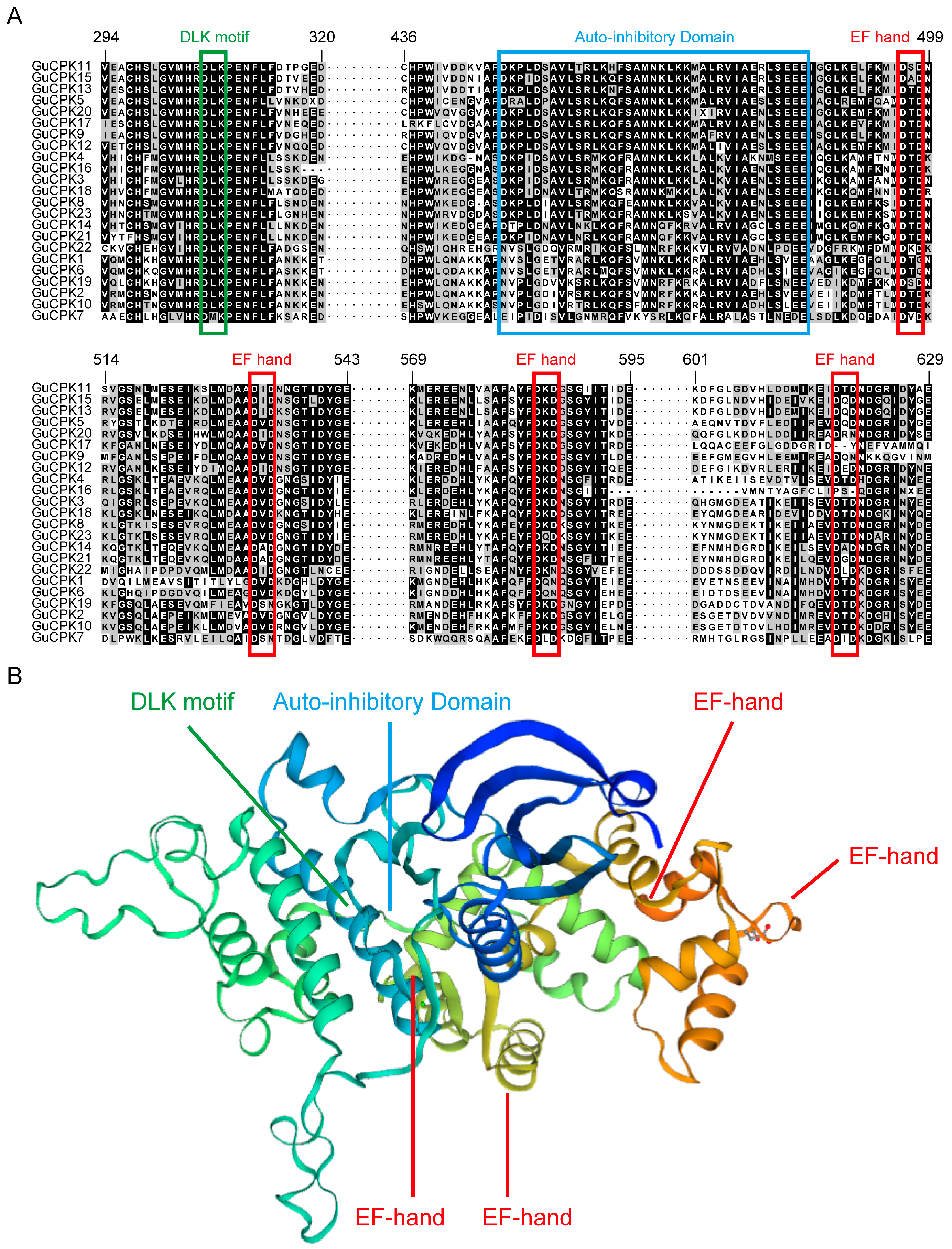
2.1. Identification of CPK Family Genes in G. uralensis
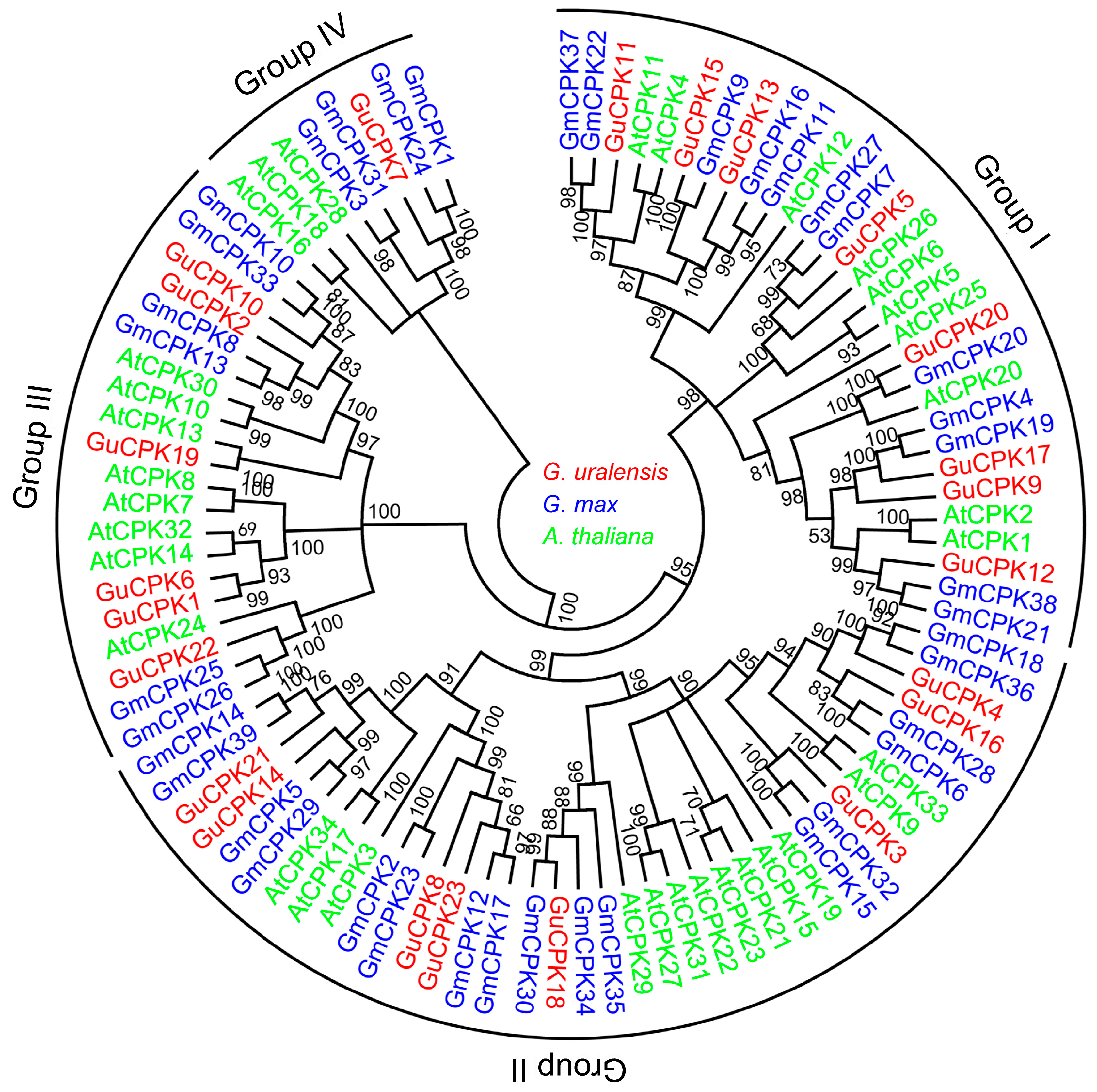
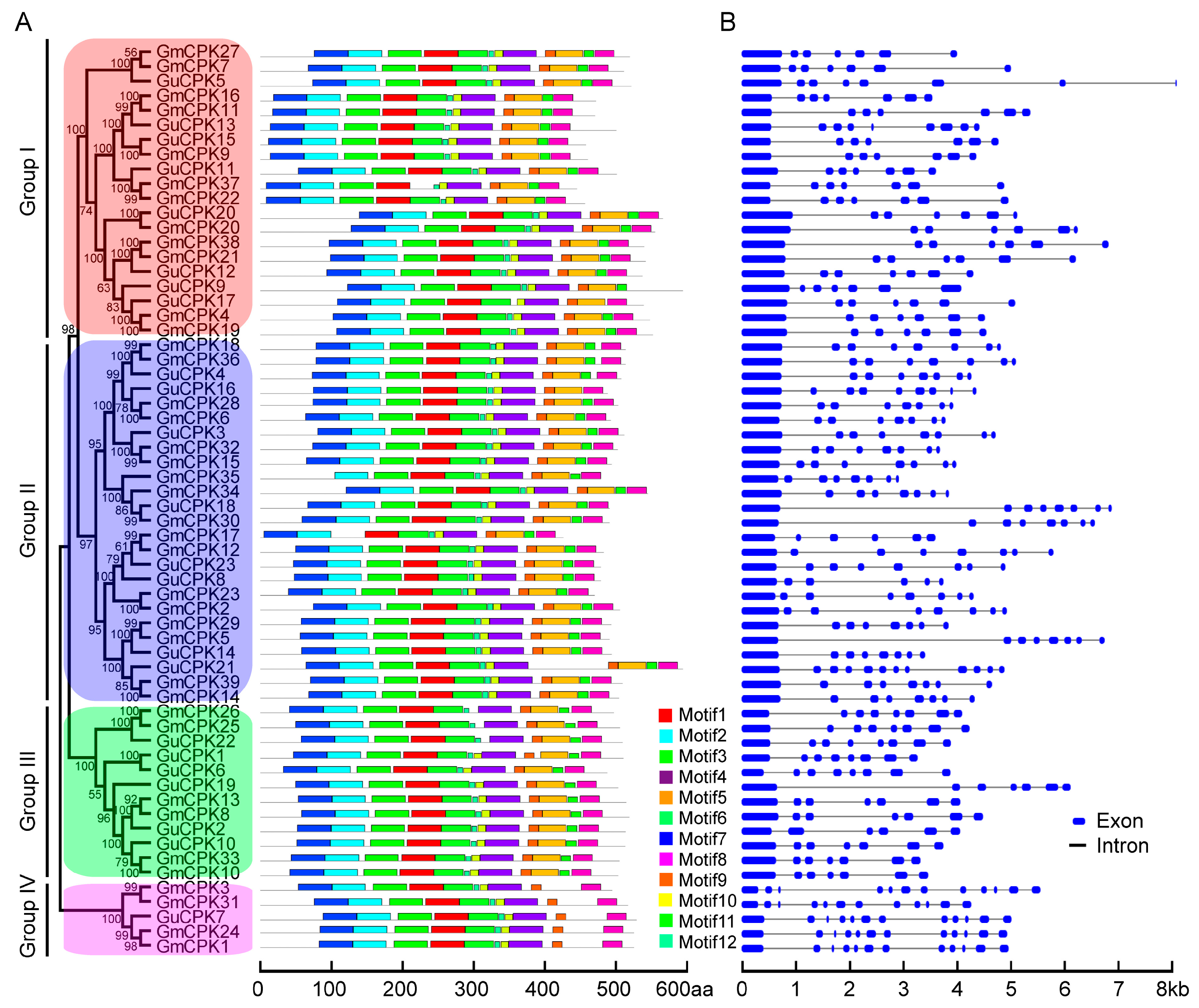
2.2. Phylogenetic Relationship, Gene Structure, and Motif Distribution Analyses of GuCPK Genes
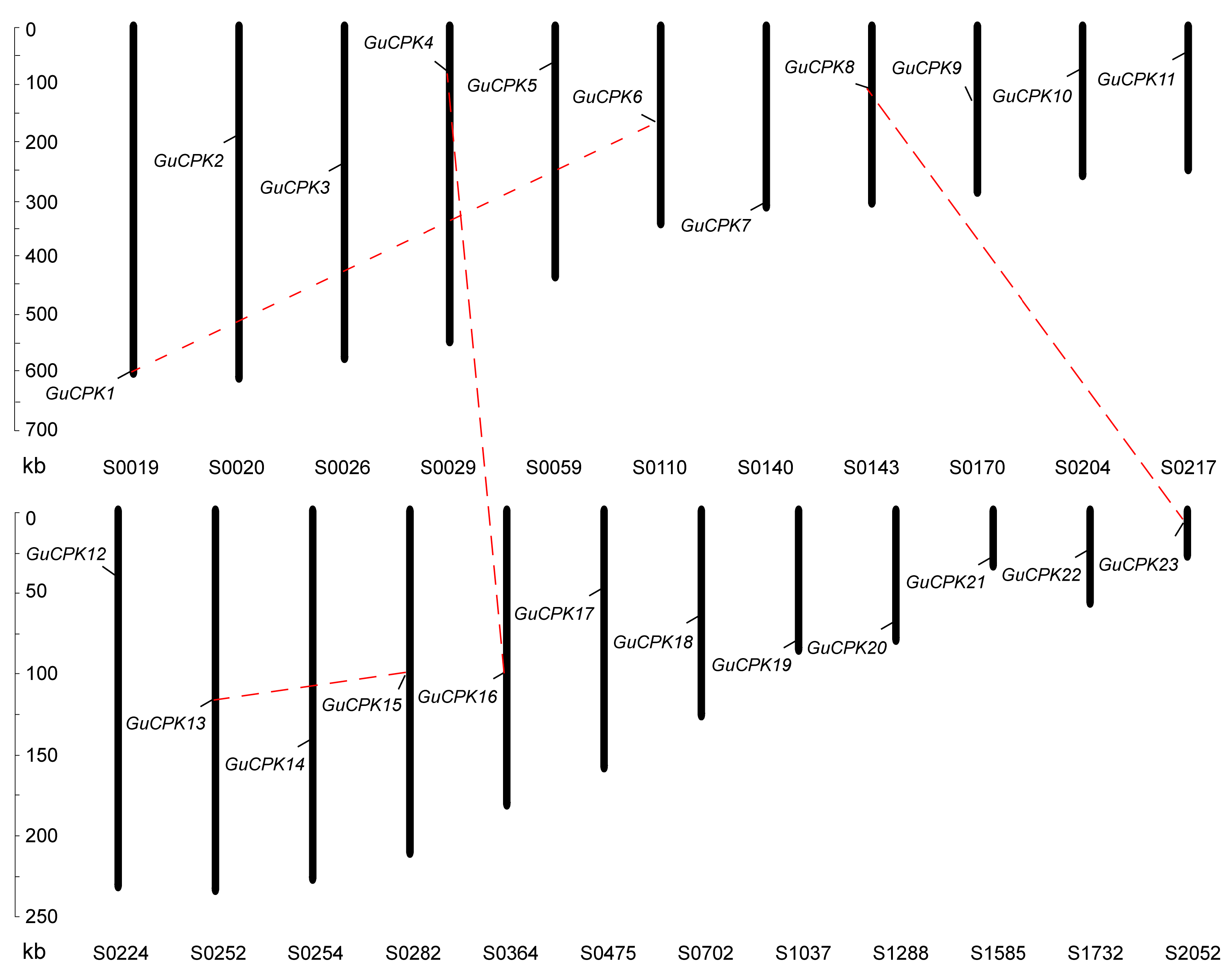
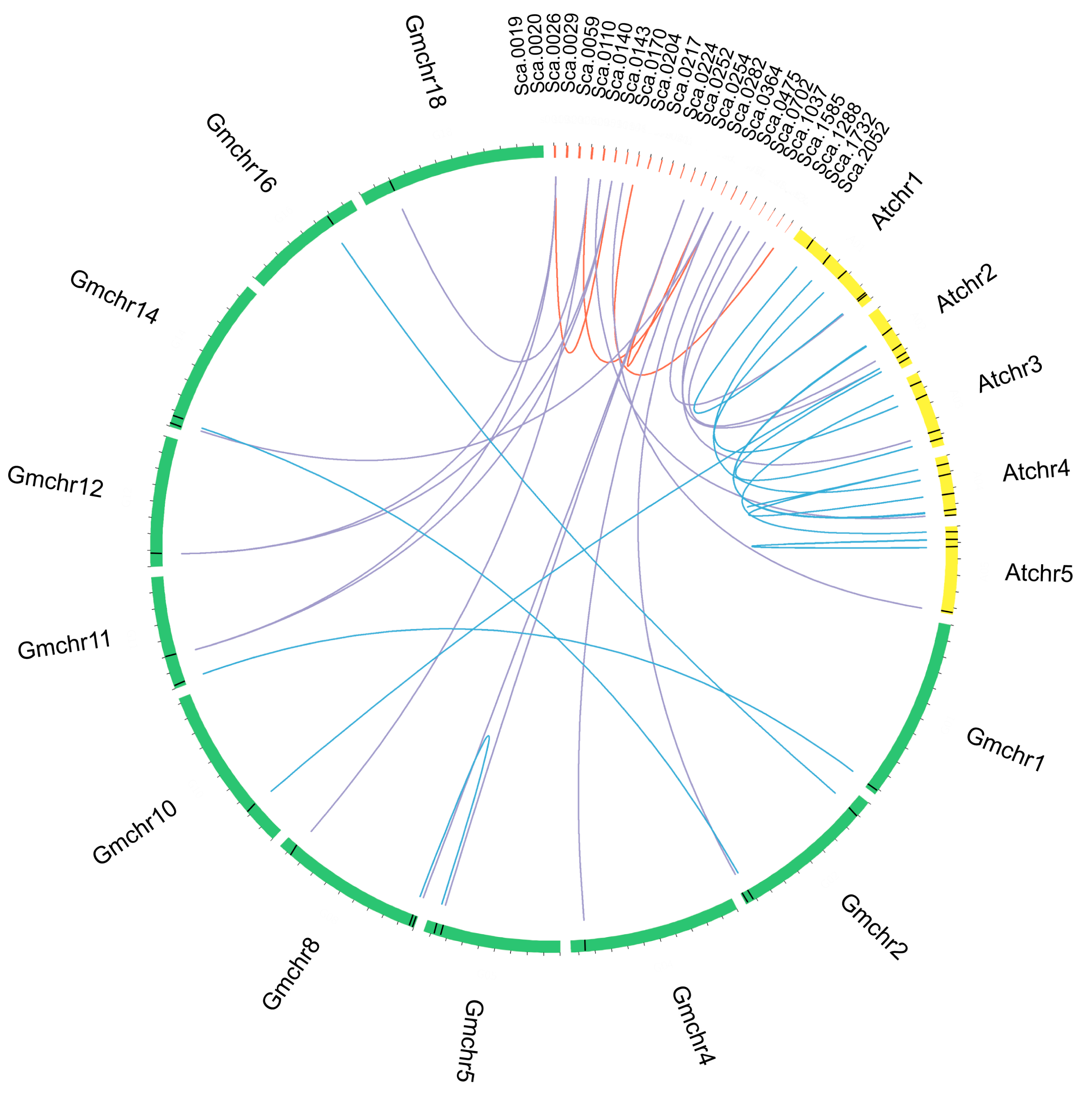
2.3. Duplication Events and Syntenic Analyses of GuCPK Family Gene Members
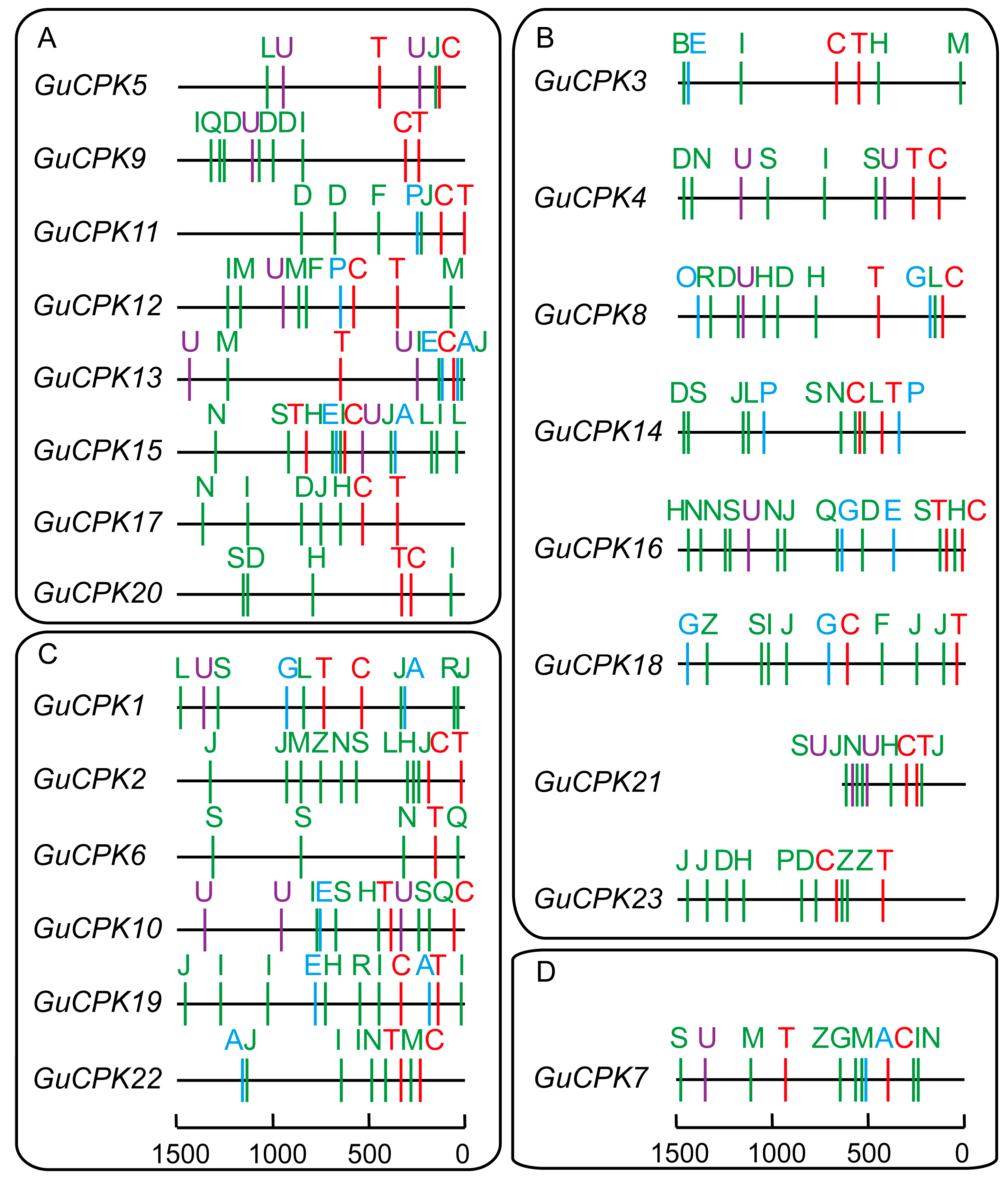
2.4. Analysis of Cis-Acting Elements in the Promoter Region of GuCPK Genes
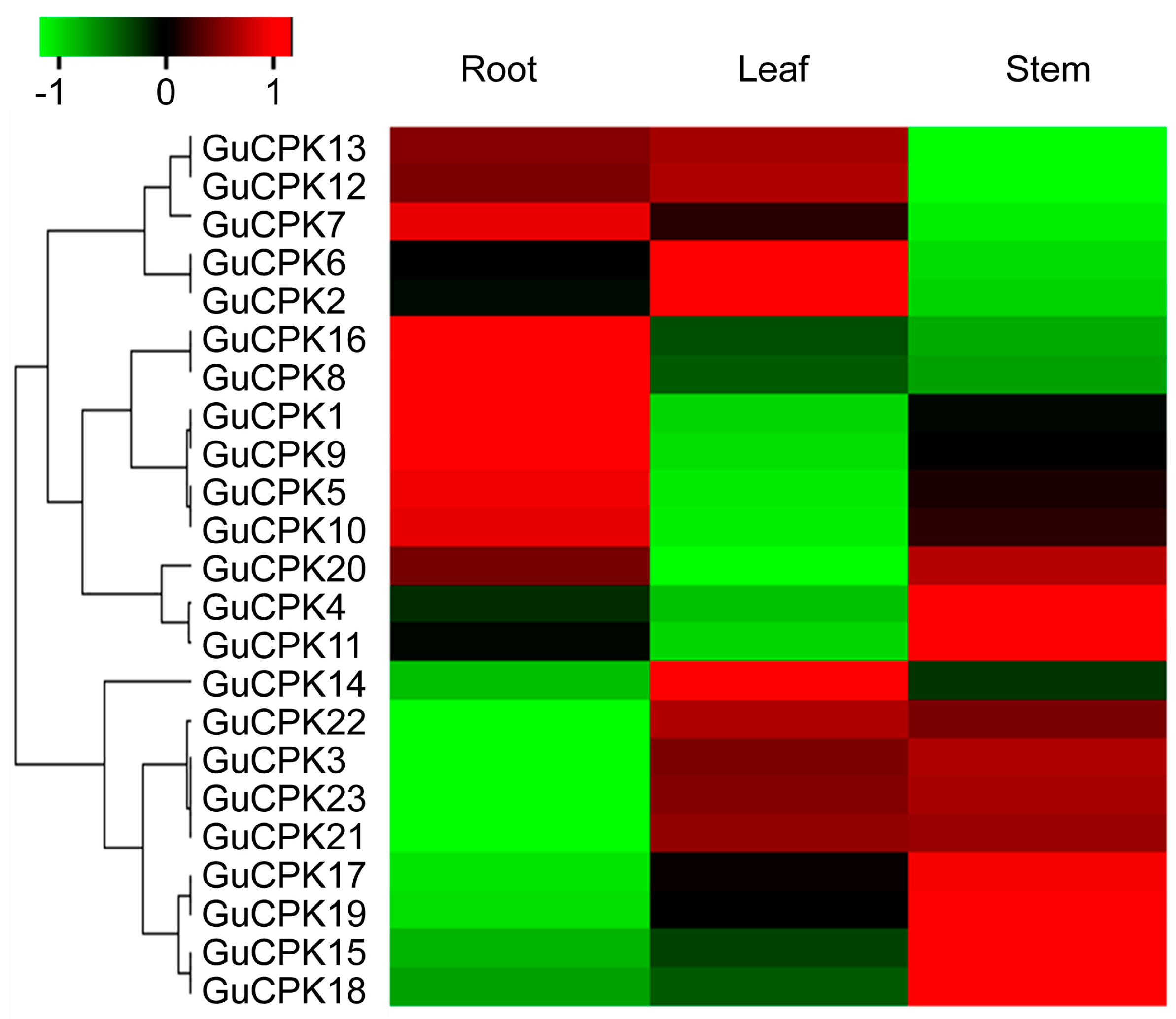
2.5. Tissue-Specific Expression Pattern Analysis of GuCPKs
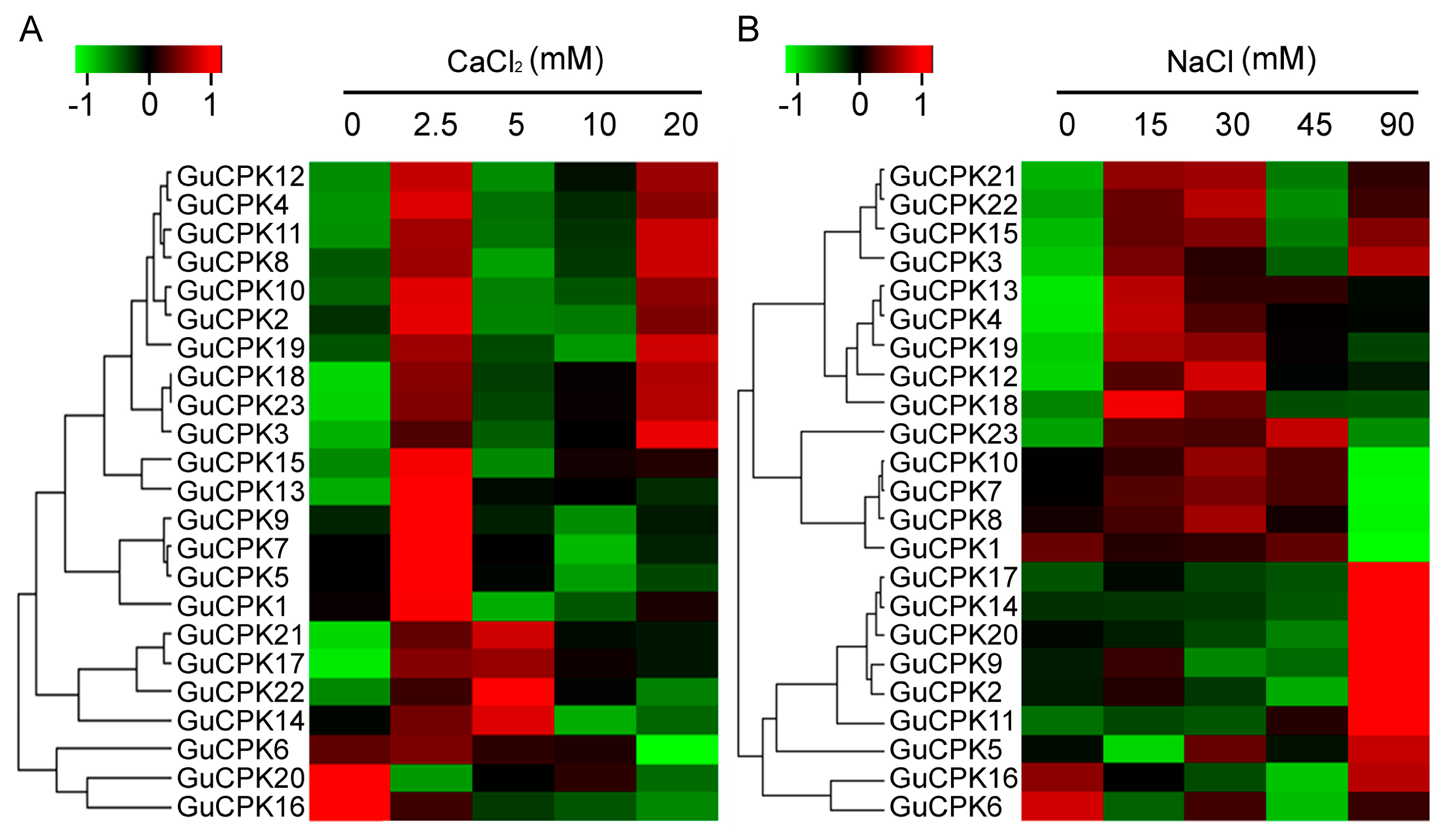
2.6. Expression Profile of GuCPKs in Response to CaCl2 and NaCl Treatments
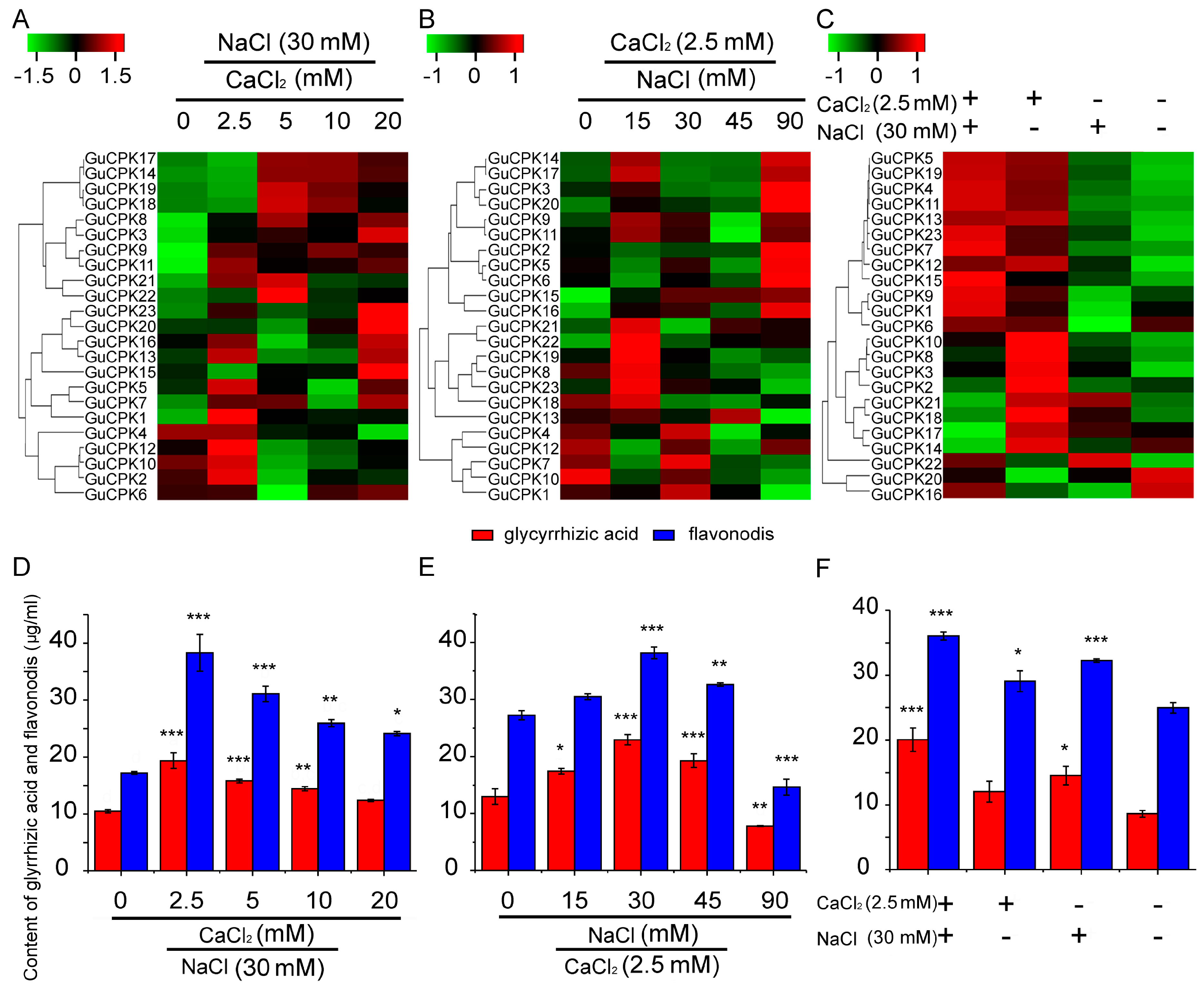
2.7. Correlation Analysis of GuCPKs Expression and Biosynthesis of Glycyrrhizic Acid and Flavonoids of G. Uralensis under CaCl2 and NaCl Treatments
3. Discussion
4. Materials and Methods
4.1. Genome-Wide Identification of GuCPK Family Genes
4.2. Analyses of Phylogenetic Relationship and Gene Structural Organization
4.3. Cis-Element Analysis in GuCPK Promoter Regions
4.4. Plant Materials and Salt Treatment
4.5. Determination of Glycyrrhizic Acid and Flavonoids
4.6. Analysis the Expression Pattern of GuCPKs
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| aa | Amino acid |
| ABA | Abscisic acid |
| ABRE | ABA-responsive element |
| ARE | Anaerobic induction |
| CPK | Calcium-dependent protein kinases |
| DLK | Dual leucine-zipper kinase |
| EF | Effective factor |
| ERF | Ethylene-responsive element |
| GA | Gibberellin |
| GARE | Gibberellin response element |
| GSDS | Gene structure display server |
| HMM | Hidden Markov model |
| HPLC | High-performance liquid chromatography |
| LTR | Low-temperature responsiveness |
| MAG | Monoammonium glycyrrhizinate |
| MBSI | MYB binding site involving in flavonoid biosynthetic gene regulation |
| MEME | Multiple expectation-maximization (Em) for motif elicitation |
| MRE | Mineralocorticoid response element |
| MYB | v-myb avian myeloblastosis viral oncogene homolog |
| MW | Molecular weight |
| ORF | Open reading frame |
| pI | Isoelectric point |
| Ser | Serine |
| Thr | Threonine |
References
- Xiao, X.H.; Yang, M.; Sui, J.L.; Qi, J.Y.; Fang, Y.J.; Hu, S.N.; Tang, C.R. The calcium-dependent protein kinase (CDPK) and CDPK-related kinase gene families inHevea brasiliensis-comparison with five other plant species in structure, evolution, and expression. FEBS Open Bio 2017, 7, 4–24. [Google Scholar] [CrossRef] [PubMed]
- Xiong, L.; Schumaker, K.S.; Zhu, J.K. Cell Signaling during Cold, Drought, and Salt Stress. Plant Cell 2002, 14, 165–183. [Google Scholar] [CrossRef]
- Zhu, L.P.; Jin, X.; Xie, Q.L.; Yao, Q.; Wang, X.C.; Li, H.B. Calcium-Dependent Protein Kinase Family Genes Involved in Ethylene-Induced Natural Rubber Production in Different Hevea brasiliensis Cultivars. Int. J. Mol. Sci. 2018, 19, 947. [Google Scholar] [CrossRef]
- Mohanta, T.K.; Occhipinti, A.; Atsbaha Zebelo, S.; Foti, M.; Fliegmann, J.; Bossi, S.; Maffei, M.E.; Bertea, C.M. Ginkgo biloba Responds to Herbivory by Activating Early Signaling and Direct Defenses. PLoS ONE 2012, 7, e32822. [Google Scholar] [CrossRef]
- Monaghan, J.; Matschi, S.; Romeis, T.; Zipfel, C. The calcium-dependent protein kinase CPK28 negatively regulates the BIK1-mediated PAMP-induced calcium burst. Plant Signal. Behav. 2015, 10, 605–615. [Google Scholar] [CrossRef]
- Sanders, D.; Brownlee, C.; Harper, J.F. Communicating with calcium. Plant Cell 1999, 11, 691–706. [Google Scholar] [CrossRef]
- Kudla, J.; Oliver, B.; Kenji, H. Calcium signals: The lead currency of plant information processing. Plant Cell 2010, 22, 541–563. [Google Scholar] [CrossRef]
- Zhang, H.; Liu, W.Z.; Zhang, Y.; Deng, M.; Niu, F.; Yang, B.; Wang, X.; Wang, B.; Liang, W.; Deyholos, M.K.; et al. Identification, expression and interaction analyses of calcium-dependent protein kinase (CPK) genes in canola (Brassica napus L.). BMC Genom. 2014, 15, 211–229. [Google Scholar] [CrossRef] [PubMed]
- Pandey, G.K. Emergence of a novel calcium signaling pathway in plants: CBL-CIPK signaling network. Physiol. Mol. Biol. Plants 2008, 14, 51–70. [Google Scholar] [CrossRef]
- McCormack, E.; Tsai, Y.C.; Braam, J. Handling calcium signaling: Arabidopsis CaMs and CMLs. Trends Plant Sci. 2005, 10, 383–389. [Google Scholar] [CrossRef] [PubMed]
- Hu, Z.; Lv, X.; Xia, X.; Zhou, J.; Shi, K.; Yu, J.; Zhou, Y. Genome-Wide Identification and Expression Analysis of Calcium-dependent Protein Kinase in Tomato. Front. Plant Sci. 2016, 7, e80818. [Google Scholar] [CrossRef]
- Mohanta, T.K.; Mohanta, N.; Mohanta, Y.K.; Bae, H. Genome-Wide Identification of Calcium Dependent Protein Kinase Gene Family in Plant Lineage Shows Presence of Novel D-x-D and D-E-L Motifs in EF-Hand Domain. Front. Plant Sci. 2015, 6, 1146–1161. [Google Scholar] [CrossRef]
- Maria, K.; Grazyna, M. Structure and functions of plant calcium-dependent protein kinases. Acta Biochim. Pol. 2007, 54, 219–233. [Google Scholar]
- Harmon, A.C.; Gribskov, M.; Harper, J.F. CDPKs—A kinase for every Ca2+ signal? Trends Plant Sci. 2000, 5, 154–159. [Google Scholar] [CrossRef]
- Hu, W.; Hou, X.; Xia, Z.; Yan, Y.; Wei, Y.; Wang, L.; Zou, M.; Lu, C.; Wang, W.; Peng, M. Genome-wide survey and expression analysis of the calcium-dependent protein kinase gene family in cassava. Mol. Genet. 2016, 291, 241–253. [Google Scholar] [CrossRef]
- Zhu, S.Y.; Yu, X.C.; Wang, X.J.; Zhao, R.; Li, Y.; Fan, R.C.; Shang, Y.; Du, S.Y.; Wang, X.F.; Wu, F.Q.; et al. Two calcium-dependent protein kinases, CPK4 and CPK11, regulate abscisic acid signal transduction in Arabidopsis. Plant Cell 2007, 19, 3019–3036. [Google Scholar] [CrossRef]
- Ma, S.Y.; Wu, W.H. AtCPK23 functions in Arabidopsis responses to drought and salt stresses. Plant Mol. Biol. 2007, 65, 511–518. [Google Scholar] [CrossRef]
- Franz, S.; Ehlert, B.; Liese, A.; Kurth, J.; Cazalé, A.C.; Romeis, T. Calcium-Dependent Protein Kinase CPK21 Functions in Abiotic Stress Response in Arabidopsis thaliana. Mol. Plant 2011, 4, 83–96. [Google Scholar] [CrossRef]
- Zhang, K.; Han, Y.T.; Zhao, F.L.; Hu, Y.; Gao, Y.R.; Ma, Y.F.; Zheng, Y.; Wang, Y.J.; Wen, Y.Q. Genome-wide Identification and Expression Analysis of the CDPK Gene Family in Grape, Vitis spp. BMC Plant Biol. 2015, 15, 164–183. [Google Scholar] [CrossRef]
- Saijo, Y.; Hata, S.; Kyozuka, J.; Shimamoto, K.; Izui, K. Over-expression of a single Ca2+-dependent protein kinase confers both cold and salt/drought tolerance on rice plants. Plant J. 2010, 23, 319–327. [Google Scholar] [CrossRef]
- Asano, T.; Hakata, M.; Nakamura, H.; Aoki, N.; Komatsu, S.; Ichikawa, H.; Hirochika, H.; Ohsugi, R. Functional characterisation of OsCPK21, a calcium-dependent protein kinase that confers salt tolerance in rice. Plant Mol. Biol. 2011, 75, 179–191. [Google Scholar] [CrossRef]
- Romeis, T.; Ludwig, A.A.; Martin, R.; Jones, J.D. Calcium-dependent protein kinases play an essential role in a plant defence response. EMBO J. 2014, 20, 5556–5567. [Google Scholar] [CrossRef]
- Candace, M.; Romanowsky, S.M.; Barron, Y.D.; Shilpi, G.; Azuse, C.L.; Amy, C.; Davis, R.M.; Jasmine, H.; Harmon, A.C.; Harper, J.F. Calcium-dependent protein kinases regulate polarized tip growth in pollen tubes. Plant J. Cell Mol. Biol. 2010, 59, 528–539. [Google Scholar]
- Kong, X.; Lv, W.; Jiang, S.; Zhang, D.; Cai, G.; Pan, J.; Li, D. Genome-wide identification and expression analysis of calcium-dependent protein kinase in maize. BMC Genom. 2013, 14, 433. [Google Scholar] [CrossRef]
- Wei, S.; Wei, H.; Deng, X.; Zhang, Y.; Liu, X.; Zhao, X.; Luo, Q.; Jin, Z.; Yin, L.; Zhou, S. A rice calcium-dependent protein kinase OsCPK9 positively regulates drought stress tolerance and spikelet fertility. BMC Plant Biol. 2014, 14, 133–146. [Google Scholar] [CrossRef]
- Morello, L.; Frattini, M.; Gianì, S.; Christou, P.; Breviario, D. Overexpression of the calcium-dependent protein kinase OsCDPK2 in transgenic rice is repressed by light in leaves and disrupts seed development. Transgenic Res. 2000, 9, 453–462. [Google Scholar] [CrossRef]
- Salme, T.; Viiu, P.; Tomas, P.; Jonas, B.; Ameraswar, V.; Triin, D.; Eviatar, N. Bacterial distribution in the rhizosphere of wild barley under contrasting microclimates. PLoS ONE 2011, 6, e17968. [Google Scholar]
- Barea, J.M.; Pozo, M.J.; Azcón, R.; Azcón-Aguilar, C. Microbial co-operation in the rhizosphere. J. Exp. Bot. 2005, 56, 1761–1778. [Google Scholar] [CrossRef] [Green Version]
- Rizzato, G.; Scalabrin, E.; Radaelli, M.; Capodaglio, G.; Piccolo, O. A new exploration of licorice metabolome. Food Chem. 2017, 221, 959–968. [Google Scholar] [CrossRef]
- Rakesh, S.U.; Patil, P.R.; Mane, S.R. Use of natural antioxidants to scavenge free radicals: A major cause of diseases. Int J. Pharmtech Res. 2010, 2, 1074–1081. [Google Scholar]
- Liu, J.; Wu, L.; Wei, S.; Xiang, X.; Su, C.; Peng, J.; Song, Z.; Tao, W.; Yu, Z. Effects of arbuscular mycorrhizal fungi on the growth, nutrient uptake and glycyrrhizin production of licorice (Glycyrrhiza uralensis Fisch). Plant Growth Regul. 2007, 52, 29–39. [Google Scholar] [CrossRef]
- Shibata, S. A drug over the millennia: Pharmacognosy, chemistry, and pharmacology of licorice. Yakugaku Zasshi 2010, 32, 849–862. [Google Scholar] [CrossRef]
- Yang, R.; Li, W.; Yuan, B.; Ren, G.; Liu, Y. The genetic and chemical diversity in three original plants of licorice, Glycyrriza uralensis Fisch. Glycyrrhiza inflata Bat. and Glycyrrhiza glabra L. Pak. J. Pharm. Sci. 2018, 31, 525–535. [Google Scholar]
- Cheng, S.H.; Willmann, M.R.; Chen, H.C.; Sheen, J. Calcium signaling through protein kinases. The Arabidopsis calcium-dependent protein kinase gene family. Plant Physiol. 2002, 129, 469–485. [Google Scholar] [CrossRef]
- Liu, H.; Che, Z.; Zeng, X.; Zhou, X.; Sitoe, H.M.; Wang, H.; Yu, D. Genome-wide analysis of calcium-dependent protein kinases and their expression patterns in response to herbivore and wounding stresses in soybean. Funct. Integr. Genom. 2016, 16, 1–13. [Google Scholar] [CrossRef]
- Ray, S.; Agarwal, P.; Arora, R.; Kapoor, S.; Tyagi, A.K. Expression analysis of calcium-dependent protein kinase gene family during reproductive development and abiotic stress conditions in rice (Oryza sativa L. ssp. indica). Mol. Gene Genom. 2007, 278, 493–505. [Google Scholar] [CrossRef]
- Choi, H.I.; Park, H.J.; Park, J.H.; Kim, S.; Im, M.Y.; Seo, H.H.; Kim, Y.W.; Hwang, I.; Kim, S.Y. Arabidopsis calcium-dependent protein kinase AtCPK32 interacts with ABF4, a transcriptional regulator of abscisic acid-responsive gene expression, and modulates its activity. Plant Physiol. 2005, 139, 1750–1761. [Google Scholar] [CrossRef]
- Wang, C.T.; Shao, J.M. Characterization of the ZmCK1 Gene Encoding a Calcium-Dependent Protein Kinase Responsive to Multiple Abiotic Stresses in Maize. Plant Mol. Biol. Report. 2013, 31, 222–230. [Google Scholar] [CrossRef]
- Xu, X.; Liu, M.; Lu, L.; He, M.; Qu, W.; Xu, Q.; Qi, X.; Chen, X. Genome-wide analysis and expression of the calcium-dependent protein kinase gene family in cucumber. Mol. Gene Genom. 2015, 290, 1403–1414. [Google Scholar] [CrossRef]
- Ma, P.; Liu, J.; Yang, X.; Ma, R. Genome-Wide Identification of the Maize Calcium-Dependent Protein Kinase;Gene Family. Appl. Biochem. Biotechnol. 2013, 169, 2111–2125. [Google Scholar] [CrossRef]
- Chen, F.; Fasoli, M.; Tornielli, G.B.; Dal Santo, S.; Pezzotti, M.; Zhang, L.; Cai, B.; Cheng, Z.M. The evolutionary history and diverse physiological roles of the grapevine calcium-dependent protein kinase gene family. PLoS ONE 2013, 8, e80818. [Google Scholar] [CrossRef]
- Berberich, T.; Kusano, T. Cycloheximide induces a subset of low temperature-inducible genes in maize. Mol. Gen. Gent. 1997, 254, 275–283. [Google Scholar] [CrossRef]
- Ritika, D.; Pandey, G.K. Expressional analysis and role of calcium regulated kinases in abiotic stress signaling. Curr. Genomics 2010, 11, 2–13. [Google Scholar]
- Yi, M.; Yichen, Z.; Walker, R.K.; Berkowitz, G.A. Molecular steps in the immune signaling pathway evoked by plant elicitor peptides: Ca2+-dependent protein kinases, nitric oxide, and reactive oxygen species are downstream from the early Ca2+ signal. Plant Physiol. 2013, 163, 1459–1471. [Google Scholar]
- Geiger, D.; Scherzer, S.; Mumm, P.; Stange, A.; Marten, I.; Bauer, H.; Ache, P.; Matschi, S.; Liese, A.; Al-Rasheid, K.A.; et al. Activity of guard cell anion channel SLAC1 is controlled by drought-stress signaling kinase-phosphatase pair. Proc. Natl. Acad. Sci. USA 2009, 106, 21425–21430. [Google Scholar] [CrossRef] [Green Version]
- Ludwig, A.A.; Hiromasa, S.; Georg, F.; Gerald, F.; Otto, M.; Claus, W.; Thomas, B.; Jones, J.D.; Tina, R. Ethylene-mediated cross-talk between calcium-dependent protein kinase and MAPK signaling controls stress responses in plants. Proc. Natl. Acad. Sci. USA 2005, 102, 10736–10741. [Google Scholar] [CrossRef] [Green Version]
- Zou, J.J.; Wei, F.J.; Wang, C.; Wu, J.J.; Ratnasekera, D.; Liu, W.X.; Wu, W.H. Arabidopsis Calcium-Dependent Protein Kinase CPK10 Functions in Abscisic Acid- and Ca2+-Mediated Stomatal Regulation in Response to Drought Stress. Plant Physiol. 2010, 154, 1232–1243. [Google Scholar] [CrossRef]
- Ho, S.L.; Huang, L.F.; Lu, C.A.; He, S.L.; Wang, C.C.; Yu, S.P.; Chen, J.; Yu, S.M. Sugar starvation- and GA-inducible calcium-dependent protein kinase 1 feedback regulates GA biosynthesis and activates a 14-3-3 protein to confer drought tolerance in rice seedlings. Plant Mol. Biol. 2013, 81, 347–361. [Google Scholar] [CrossRef]
- Kamiyoshihara, Y.; Iwata, M.; Fukaya, T.; Tatsuki, M.; Mori, H. Turnover of LeACS2, a wound-inducible 1-aminocyclopropane-1-carboxylic acid synthase in tomato, is regulated by phosphorylation/dephosphorylation. Plant J. 2010, 64, 140–150. [Google Scholar] [CrossRef]
- Egamberdieva, D.; Li, L.; Kristina, L.; Leena, A. A synergistic interaction between salt-tolerant pseudomonas, and mesorhizobium, strains improves growth and symbiotic performance of liquorice (glycyrrhiza uralensis, fish.) under salt stress. Appl. Microbiol. Biotechnol. 2016, 100, 2829–2841. [Google Scholar] [CrossRef]
- Zhang, X.; Zhang, W.; Lang, D.; Cui, J.; Li, Y. Silicon improves salt tolerance of glycyrrhiza uralensis fisch. by ameliorating osmotic and oxidative stresses and improving phytohormonal balance. Environ. Sci. Pollut. Res. Int. 2018, 25, 1–17. [Google Scholar] [CrossRef]
- Taarit, M.B.; Mouna; Msaada, K.; Hosni, K.; Marzouk, B. Physiological changes and essential oil composition of clary sage (Salvia sclarea L.) rosette leaves as affected by salinity. Acta Physiol. Plant. 2011, 33, 153–162. [Google Scholar] [CrossRef]
- Shao, Y.H.; Gao, J.L.; Wu, X.W.; Qian, L.; Wang, J.G.; Ping, D.; Lai, X.P. Effect of salt treatment on growth, isoenzymes and metabolites of Andrographis paniculata (Burm. f.) Nees. Acta Physiol. Plant. 2015, 37, 35–43. [Google Scholar] [CrossRef]
- Pan, Y.; Wu, L.J.; Yu, Z.L. Effect of salt and drought stress on antioxidant enzymes activities and SOD isoenzymes of liquorice (Glycyrrhiza uralensis Fisch). Plant Growth Regul. 2006, 49, 157–165. [Google Scholar] [CrossRef]
- Zhao, L.Y.; Deng, X.P.; Lun, S. The Response Mechanism of Active Oxygen Species Removing System to Drought Stress. Acta Bot. Boreali-Occident. Sin. 2005, 25, 413–418. [Google Scholar]
- Gandhi, N.M.; Maurya, D.K.; Salvi, V.; Kapoor, S.; Mukherjee, T.; Nair, C.K. Radioprotection of DNA by glycyrrhizic acid through scavenging free radicals. J. Radiat. Res. 2004, 45, 461–468. [Google Scholar] [CrossRef]
- Li, W.D.; Hou, J.L.; Wang, W.Q.; Liu, C.L.; Xing, D. Effect of water deficit on biomass production and accumulation of secondary metabolites in roots of Glycyrrhiza uralensis. Russ. J. Plant Physiol. 2011, 58, 538–542. [Google Scholar] [CrossRef]
- Ren, J.; Wen, L.; Gao, X.; Jin, C.; Xue, Y.; Yao, X. CSS-Palm 2.0: An updated software for palmitoylation sites prediction. Protein Eng. Des. Sel. 2008, 21, 639–644. [Google Scholar] [CrossRef]
- Jin, X.; Zhu, L.; Yao, Q.; Meng, X.; Ding, G.; Wang, D.; Xie, Q.; Tong, Z.; Tao, C.; Yu, L.; et al. Expression Profiling of Mitogen-Activated Protein Kinase Genes Reveals Their Evolutionary and Functional Diversity in Different Rubber Tree (Hevea brasiliensis) Cultivars. Genes 2017, 8, 261. [Google Scholar] [CrossRef]
- Xin, S.; Tao, C.; Li, H. Cloning and Functional Analysis of the Promoter of an Ascorbate Oxidase Gene from Gossypium hirsutum. PLoS ONE 2016, 11, e0161695. [Google Scholar] [CrossRef]
- Lescot, M.; Déhais, P.; Thijs, G.; Marchal, K.; Moreau, Y.; Van de Peer, Y.; Rouzé, P.; Rombauts, S. PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res. 2002, 30, 325–327. [Google Scholar] [CrossRef] [PubMed] [Green Version]
Sample Availability: Samples of the Glycyrrhiza uralensis materials are available from the authors. |

| Paralogous Genes | Kaa | Ksb | Ka/Ks | Selective Pressure |
|---|---|---|---|---|
| GuCPK1–GuCPK6 | 0.0584 | 0.6518 | 0.0895 | Purifying selection |
| GuCPK4–GuCPK16 | 0.0798 | 0.4052 | 0.1969 | Purifying selection |
| GuCPK8–GuCPK23 | 0.0760 | 0.5474 | 0.1388 | Purifying selection |
| GuCPK13–GuCPK15 | 0.0627 | 0.5245 | 0.1195 | Purifying selection |
| Testing Group | Mt b | M1 c | M2 d | Χ2 | Pe |
|---|---|---|---|---|---|
| GuCPK4/GuCPK16 with GmCPK6 | 398 | 39 | 19 | 6.90 | 0.00864 |
| GuCPK1/GuCPK6 with GmCPK25 | 296 | 9 | 16 | 1.96 | 0.16151 |
| GuCPK1/GuCPK6 with GmCPK26 | 300 | 9 | 16 | 1.96 | 0.16151 |
| GuCPK8/GuCPK23 with GmCPK23 | 422 | 19 | 24 | 0.58 | 0.44577 |
| GuCPK13/GuCPK15 with GmCPK9 | 426 | 32 | 11 | 10.26 | 0.00136 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tong, X.; Cao, A.; Wang, F.; Chen, X.; Xie, S.; Shen, H.; Jin, X.; Li, H. Calcium-Dependent Protein Kinase Genes in Glycyrrhiza Uralensis Appear to be Involved in Promoting the Biosynthesis of Glycyrrhizic Acid and Flavonoids under Salt Stress. Molecules 2019, 24, 1837. https://doi.org/10.3390/molecules24091837
Tong X, Cao A, Wang F, Chen X, Xie S, Shen H, Jin X, Li H. Calcium-Dependent Protein Kinase Genes in Glycyrrhiza Uralensis Appear to be Involved in Promoting the Biosynthesis of Glycyrrhizic Acid and Flavonoids under Salt Stress. Molecules. 2019; 24(9):1837. https://doi.org/10.3390/molecules24091837
Chicago/Turabian StyleTong, Xuechen, Aiping Cao, Fei Wang, Xifeng Chen, Shuangquan Xie, Haitao Shen, Xiang Jin, and Hongbin Li. 2019. "Calcium-Dependent Protein Kinase Genes in Glycyrrhiza Uralensis Appear to be Involved in Promoting the Biosynthesis of Glycyrrhizic Acid and Flavonoids under Salt Stress" Molecules 24, no. 9: 1837. https://doi.org/10.3390/molecules24091837
APA StyleTong, X., Cao, A., Wang, F., Chen, X., Xie, S., Shen, H., Jin, X., & Li, H. (2019). Calcium-Dependent Protein Kinase Genes in Glycyrrhiza Uralensis Appear to be Involved in Promoting the Biosynthesis of Glycyrrhizic Acid and Flavonoids under Salt Stress. Molecules, 24(9), 1837. https://doi.org/10.3390/molecules24091837





