Discrimination of Trichosanthis Fructus from Different Geographical Origins Using Near Infrared Spectroscopy Coupled with Chemometric Techniques
Abstract
1. Introduction
2. Results and Discussion
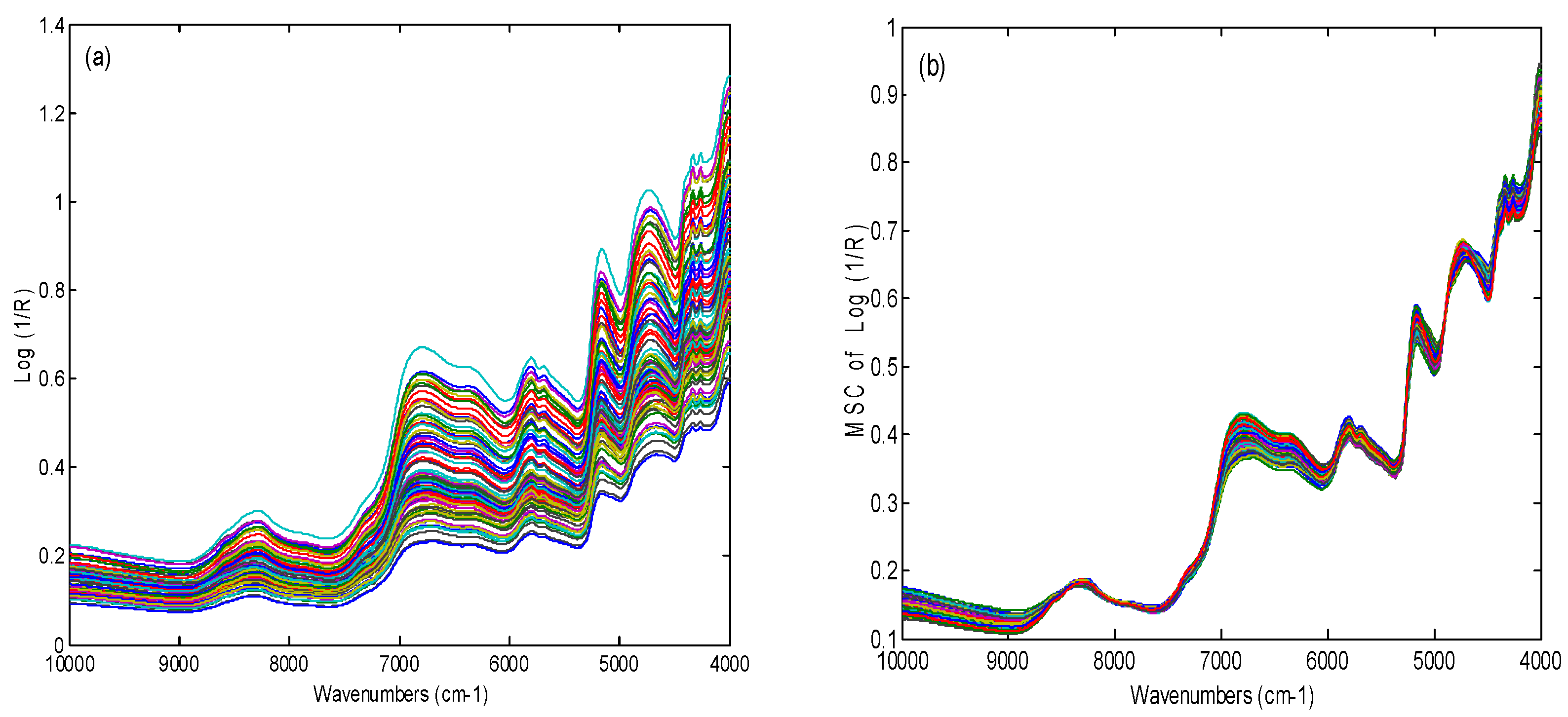
2.1. Spectra Investigation
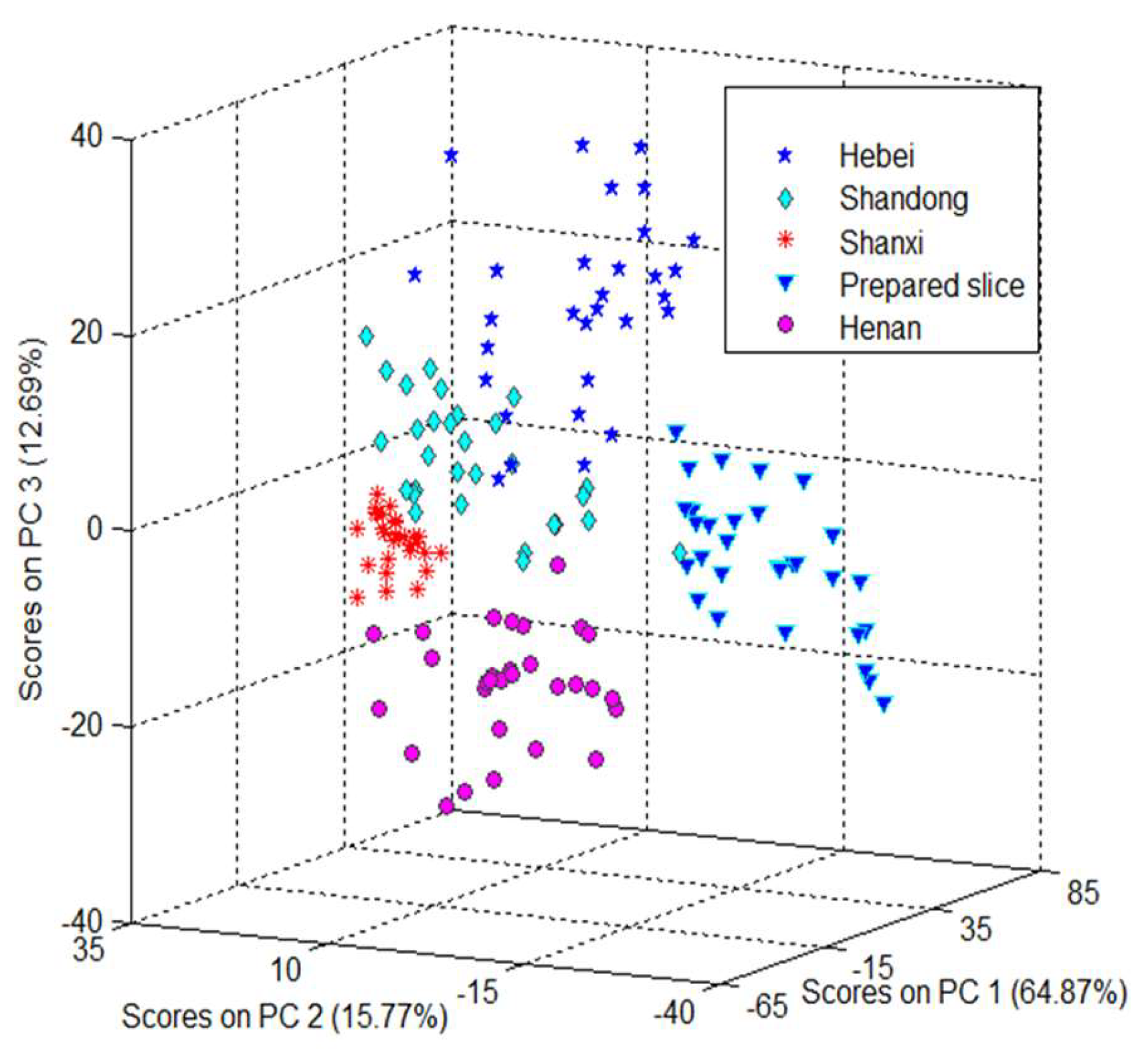
2.2. Principal Component Analysis
2.3. Optimation of Models
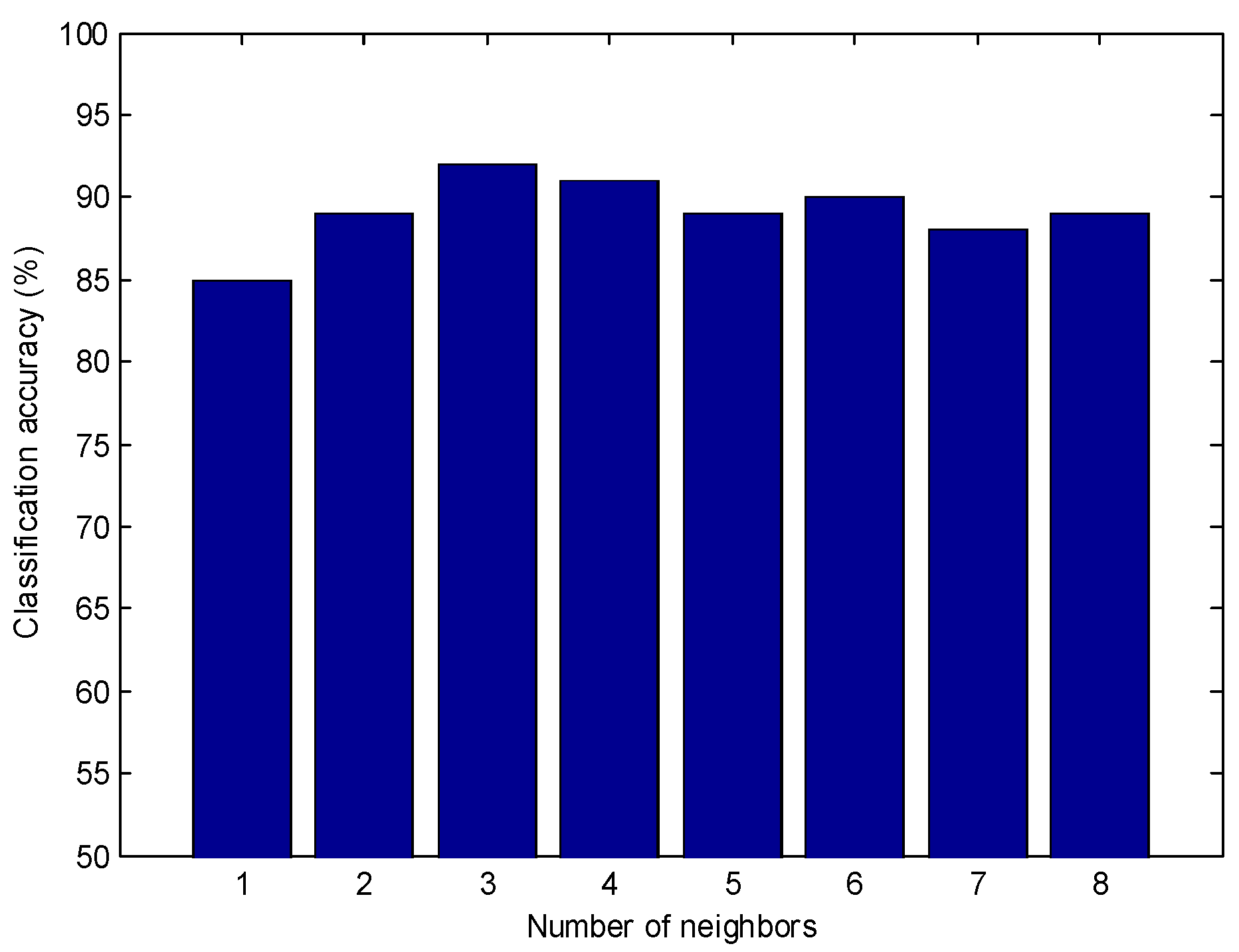
2.3.1. The Establishment of the KNN Model
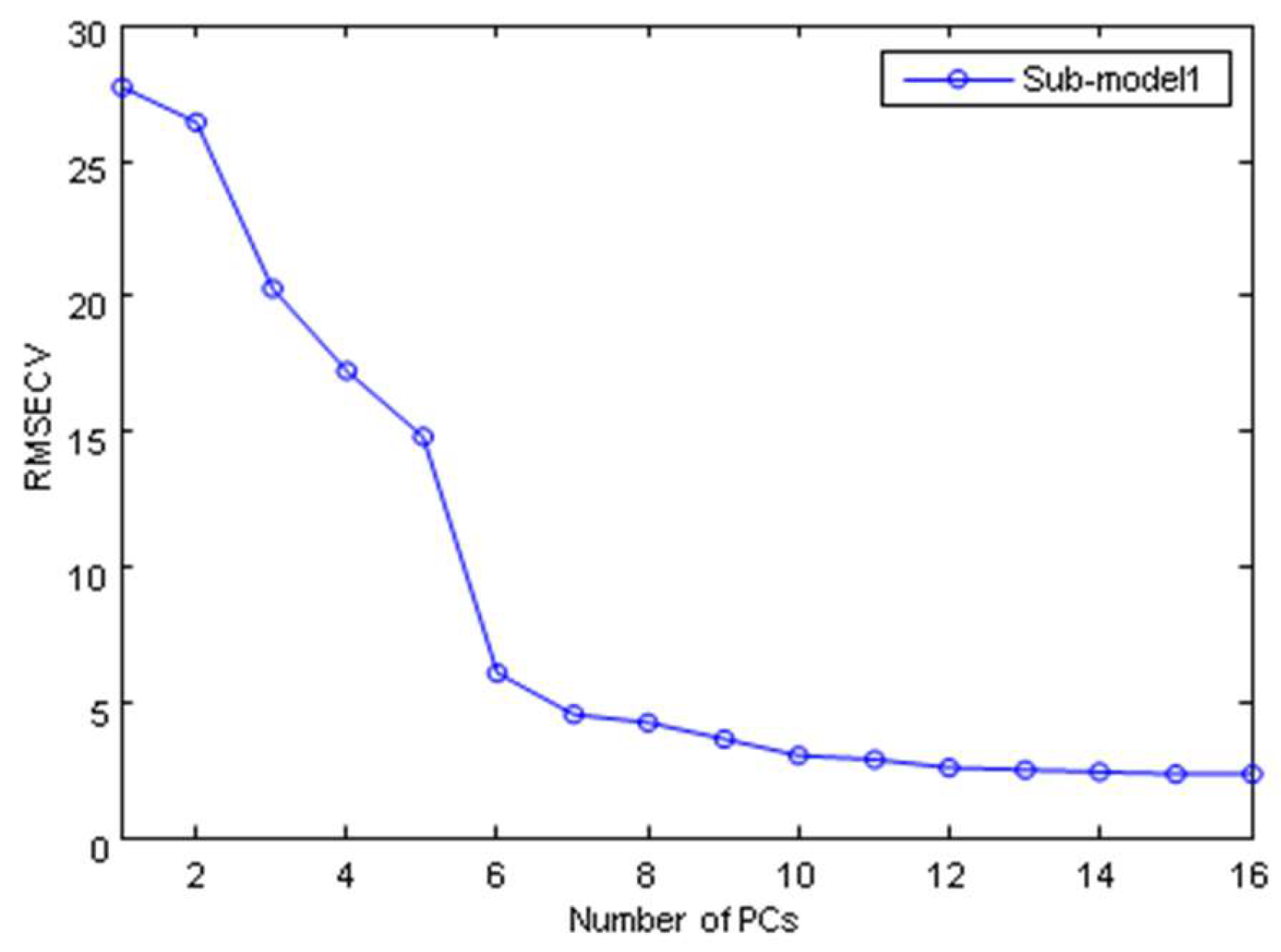
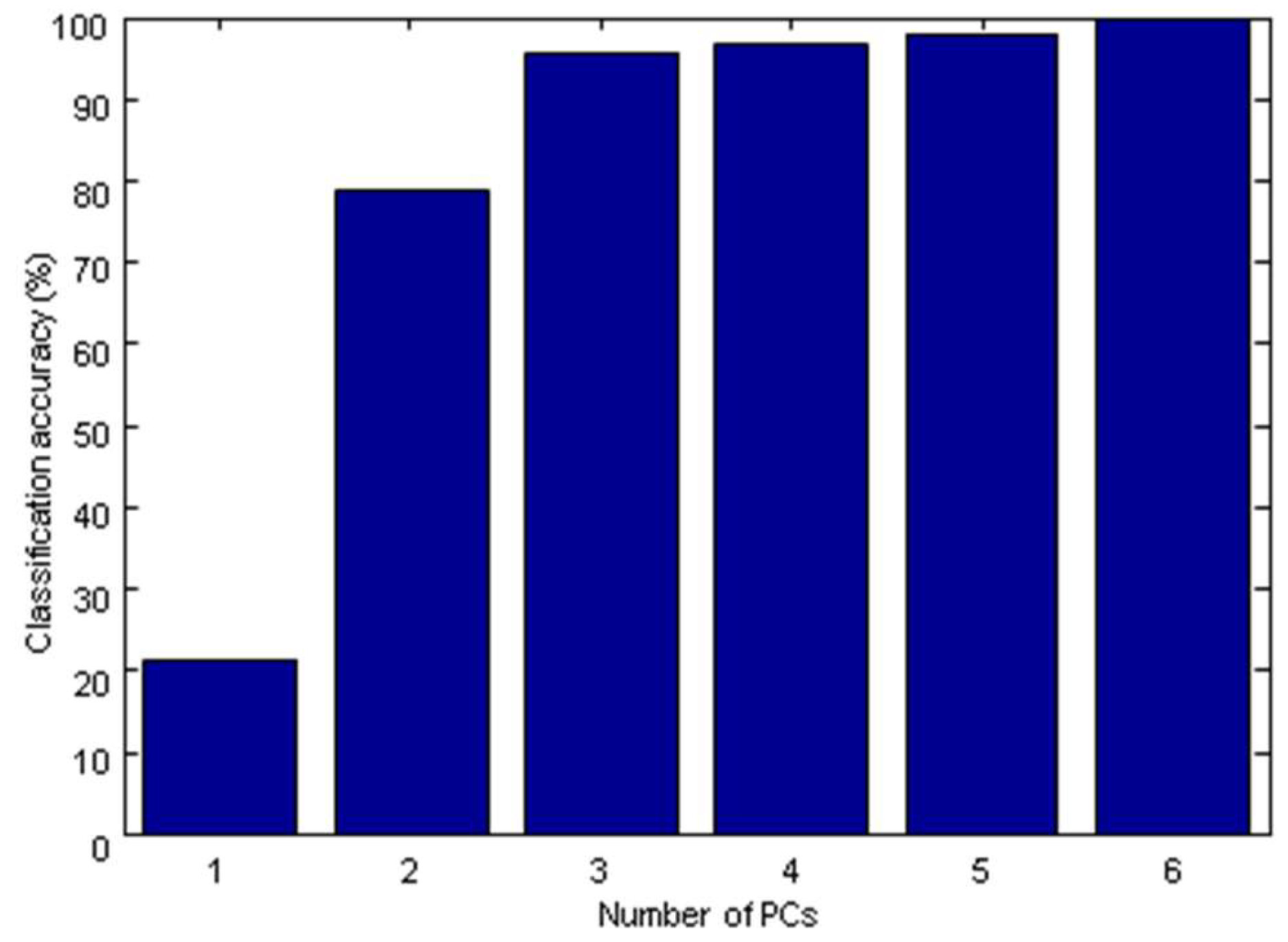
2.3.2. The Establishment of SIMCA Model
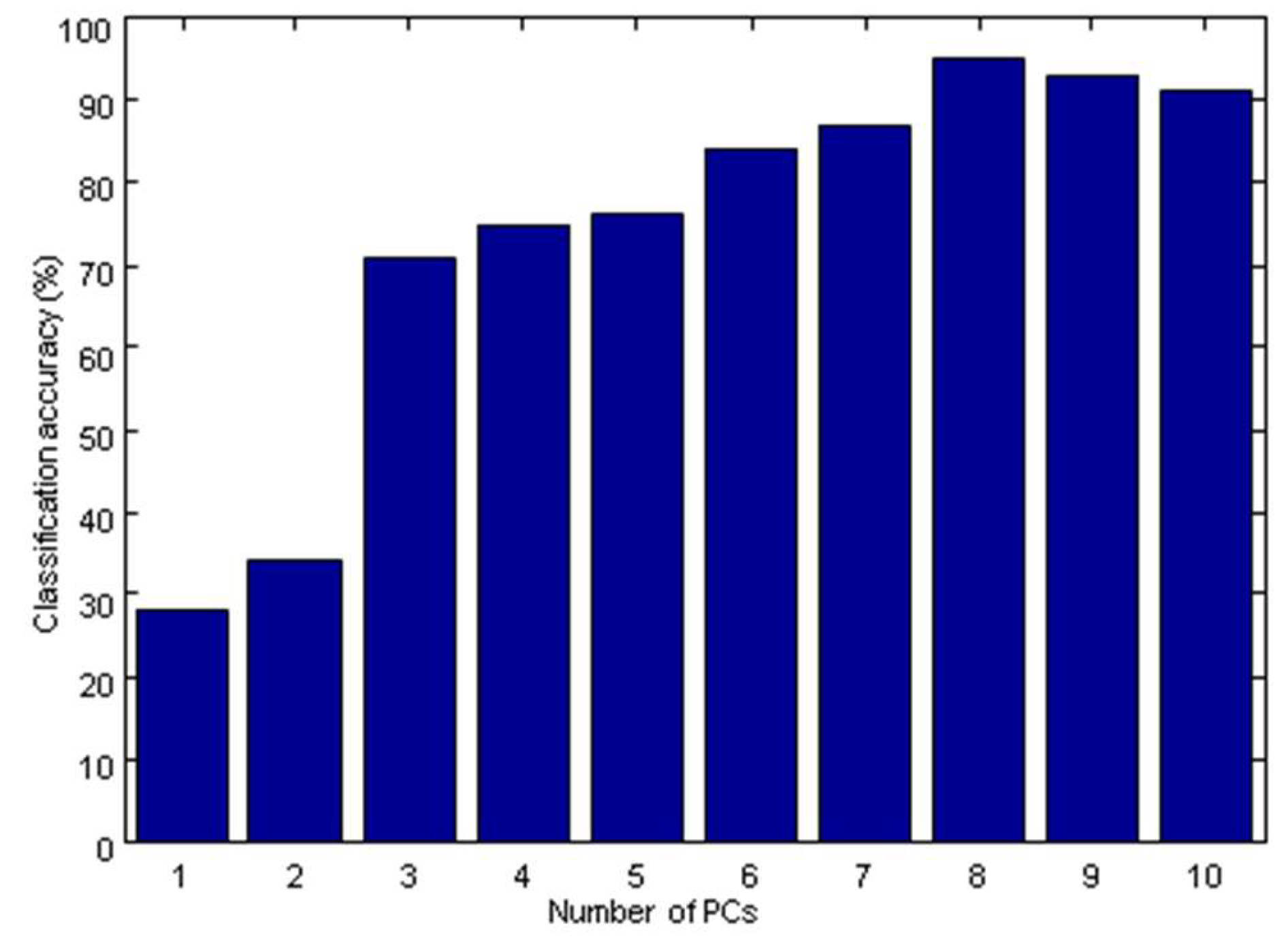
2.3.3. The Establishment of the PLS-DA Model
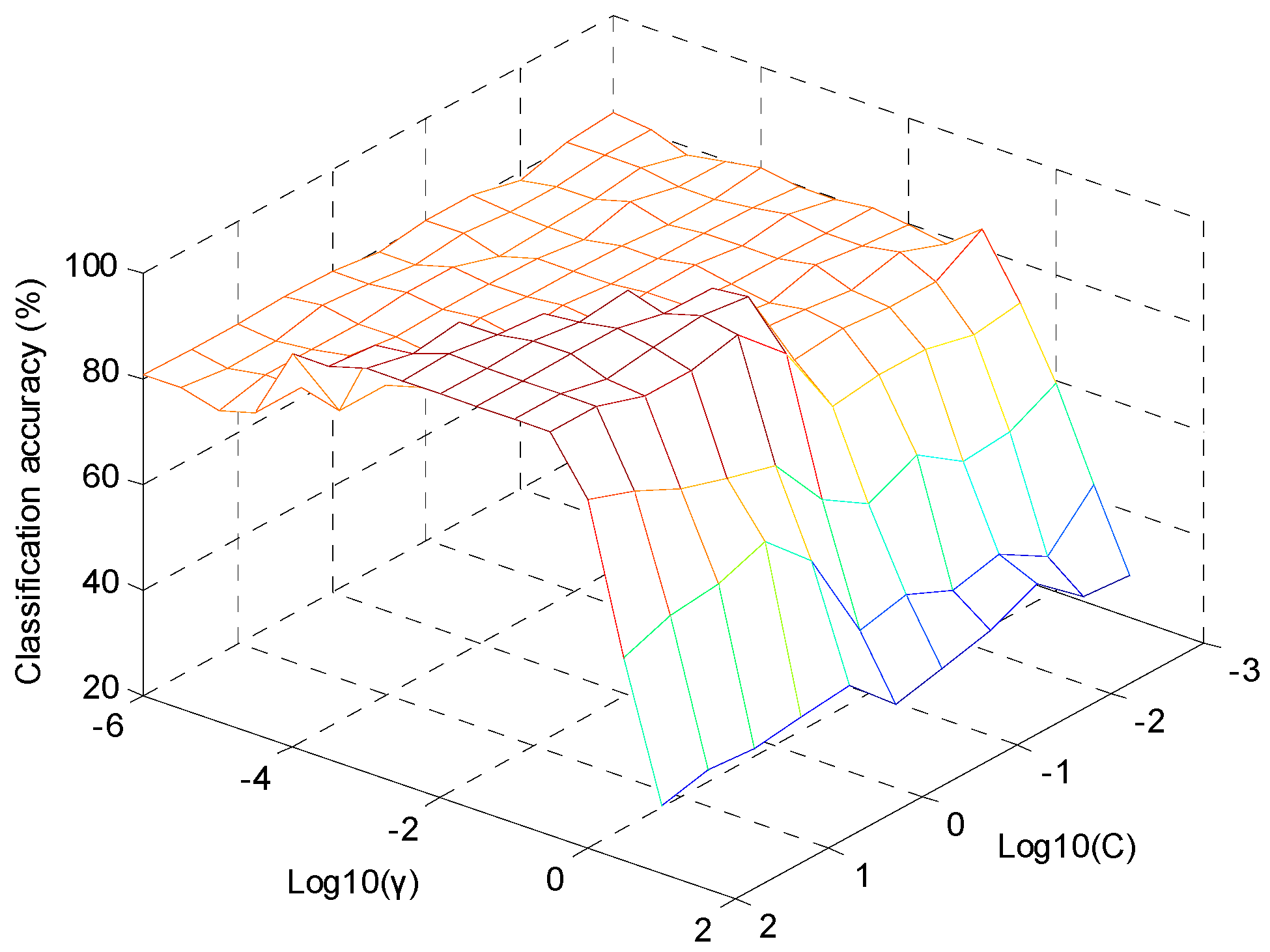
2.3.4. The Establishment of SVM-DA Model
2.4. Comparison of Four Models
3. Materials and Methods
3.1. Sample Preparation
3.2. Spectral Measurement
3.3. Data Analysis
3.4. Chemometrics Study
3.4.1. KNN
3.4.2. SIMCA
3.4.3. PLS-DA
3.4.4. SVM-DA
3.4.5. Model Efficiency Estimation
4. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- The Pharmacopoeia Commission of PRC. Pharmacopoeia of the People’s Republic of China (Part 1); China Medical Science and Technology Press: Beijing, China, 2015; pp. 112–113. [Google Scholar]
- Lu, P.; Shi, W.; Wang, Z.; Song, J. Clinical application and mechanism of Trichosanthis Pericarpium. J. Tradit. Chin. Med. 2013, 54, 1428–1431. [Google Scholar]
- Chu, D.H.; Zhang, Z.Q. Trichosanthis Pericarpium aqueous extract protects H9c2 cardiomyocytes from hypoxia/roxygenation injury by regulating PI3K/Akt/NO pathway. Molecules 2018, 23, 2409. [Google Scholar] [CrossRef] [PubMed]
- Cheng, J.; Huang, L.; Kong, C.C.; He, F. Gualou Xiebai Baijiu decoction alleviates myocardial ischemia reperfusion injury via Akt/GSK-3β signaling pathway. Acad. J. Shanghai Univ. Tradit. Chin. Med. 2018, 32, 82–85, 97. [Google Scholar]
- Liu, J.N.; Xie, X.L.; Yang, T.X.; Liu, M.; Jia, D.S.; Wen, C.X. Study on resources and cultivation progress of Trichosanthis Fructus. Food Res. Dev. 2014, 35, 125–127. [Google Scholar]
- Wu, C.; Chao, Z.M. Process control system about Trichosanthis Fructus quality. Chin. J. Exp. Tradit. Med. Form. 2016, 22, 230–234. [Google Scholar]
- Xin, J.; Zhang, R.C.; Guo, Q.M.; Zhang, Y.Q. Effect of germplasms difference on the content of total saponins and adenosine in Trichosanthis Fructus. Lishizhen Med. Mater. Med. Res. 2015, 26, 2236–2237. [Google Scholar]
- Hao, B.; Pan, L.L.; Yuan, S.X.; Yang, S.; Li, T.T.; Xu, H.F.; Li, X.R. HPLC determination of arginine in Trichosanthes Pericarpium with OPA by pre-column derivatization. Acta Chin. med. Pharm. 2014, 42, 17–19. [Google Scholar]
- Hao, B.; Pan, L.L.; Yuan, S.X.; Yang, S.; Li, T.T.; Xu, H.F.; Li, X.R. Fingerprints of amino acid of Trichosanthis Pericarpium from different varieties and habitats by HPLC. Acta Chin. med. Pharm. 2015, 43, 14–18. [Google Scholar]
- Sun, Z.Y.; Zhou, F.Q.; Guo, Q.M. Analysis of seed protein electrophoresis of farm cultivars of Fructus Trichosanthis from Shandong province. Lishizhen Med. Mater. Med. Res. 2005, 16, 1224–1225. [Google Scholar]
- Guo, Q.M.; Zhou, F.Q.; Yang, J.L.; Gao, H. Study on micro-morphological characters of fruit coats in cultivated Fructus Trichosanthis in Shandong province. Chin. J. Chin. Mater. Med. 2005, 30, 1580–1582. [Google Scholar]
- Sun, Z.Y.; Zhou, F.Q. RAPD analysis on farm cultivars of fruits of Trichosanthes kirilowii from Shandong province. Chin. Tradit. Herb. Drugs 2006, 37, 426–429. [Google Scholar]
- Cao, L.; Huang, Y.N.; Xie, J.; Peng, S.W.; Xu, R.; Zhu, X.Q. Genetic diversity of Trichosanthis Fructus for seeds cultivating. Chin. Tradit. Herb. Drugs 2017, 48, 4316–4322. [Google Scholar]
- Teye, E.; Huang, X.Y.; Dai, H.; Chen, Q.S. Rapid differentiation of Ghana cocoa beans by FT-NIR spectroscopy coupled with multivariate classification. Spectrochim. Acta A Mol. Biomol. Spectrosc. 2013, 114, 183–189. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.Y.; Zhang, C.; Zhu, S.S.; Gao, P.; Feng, L.; He, Y. Non-destructive and rapid variety discrimination and visualization of single grape seed using near-infrared hyperspectral imaging technique and multivariate analysis. Molecules 2018, 22, 1352. [Google Scholar] [CrossRef] [PubMed]
- Reboucas, M.V.; Dos Santos, J.B.; Domingos, D.; Massa, A.R.C.G. Near-infrared spectroscopic prediction of chemical composition of a series of petrochemical process streams for aromatics production. Vib. Spectrosc. 2010, 52, 97–102. [Google Scholar] [CrossRef]
- Jamrógiewicz, M. Application of the near-infrared spectroscopy in the pharmaceutical technology. J. Pharm. Biomed. Anal. 2012, 66, 1–10. [Google Scholar] [CrossRef]
- Samiei Fard, R.; Matinfar, H.R. Capability of vis-NIR spectroscopy and Landsat 8 spectral data to predict soil heavy metals in polluted agricultural land (Iran). Arab. J. Geosci. 2016, 9, 745. [Google Scholar] [CrossRef]
- Vergouw, C.G.; Botros, L.L.; Roos, P.; Lens, J.W.; Schats, R.; Hompes, P.G.A.; Burns, D.H.; Lambalk, C.B. Metabolomic profiling by near-infrared spectroscopy as a tool to assess embryo viability: a novel, non-invasive method for embryo selection. Hum. Reprod. 2008, 23, 1499–1504. [Google Scholar] [CrossRef] [PubMed]
- Williams, P.; Norris, K. Near-infrared Technology in the Agricultural and Food Industries, 2nd ed.; American Association of Cereal Chemist: St. Paul, MN, USA, 2002. [Google Scholar]
- Lin, H.; Zhao, J.W.; Chen, Q.S.; Zhou, F.; Sun, L. Discrimination of Radix Pseudostellariae according to geographical origins using NIR spectroscopy and support vector data description. Spectrochim. Acta A Mol. Biomol. Spectrosc. 2011, 79, 1381–1385. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Wu, Y.J.; Li, W.; Liu, X.S.; Zheng, J.Y.; Zhang, W.T.; Chen, Y. Determination of geographical origin and icariin content of Herba Epimedii using near infrared spectroscopy and chemometrics. Spectrochim Acta Part A Mol. Biomol. Spectrosc. 2018, 191, 233–240. [Google Scholar] [CrossRef]
- Zuo, Y.M.; Deng, X.H.; Wu, Q. Discrimination of Gastrodia elata from different geographical origin for quality evaluation using newly-build near infrared spectrum coupled with multivariate analysis. Molecules 2018, 23, 1088. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.Z.; Wang, X.H.; Zhu, Y.; Wang, C.Q. Determination of content of quercetin and total flavonoids in Trichosanthis Pericarpium from different areas. Chin. J. Exp. Tradit. Med. Form. 2014, 20, 86–89. [Google Scholar]
- Zou, C.C.; Zong, Q.N.; Yan, H.Y.; Xie, S.X.; Zhang, M.J.; Zhang, Y.M.; Li, N. HPLC fingerprint and quantitative analysis of 6 components of Fructus Trichosanthis and its steamed products. Chin. Pharm. J. 2017, 52, 597–601. [Google Scholar]
- Sun, W.; Chao, Z.M.; Wang, C.; Wu, X.Y.; Tan, Z.G. Determination of 5-hydroxmethylfurfural in commercial Trichosanthis Fructus by HPLC. Chin. J. Exp. Tradit. Med. Form. 2012, 18, 73–76. [Google Scholar]
- Yan, H.Y.; Zong, Q.N.; Zou, C.C.; Zhang, N. A comparative study on reducing sugar and total sugar in Trichosanthis Fructus and its steamed products. J. Dali Univ. 2018, 3, 71–74. [Google Scholar]
- Chao, Z.M.; He, B.; Zhang, Y.; Akihisa, T. Studies on the chemical constituents of unsaponifiable lipids from the seeds of Trichosanthes kirilowii. Chin. Pharm. J. 2000, 35, 733–736. [Google Scholar]
- Li, X.; Tang, L.Y.; Xu, J.; Yu, X.K.; Yang, H.J.; Li, D.F.; Zhang, Y.; Fan, J.W.; Su, R.Q.; Wu, H.W.; et al. Analysis and identification of chemical components in Trichosanthis Fructus by UPLC-LTQ-Orbitrap-MS. Chin. J. Exp. Tradit. Med. Form. 2019, 25, 201–210. [Google Scholar]
- Ye, X.; Ng, C.C.; Ng, T.B.; Chan, G.H.; Guan, S.; Sha, O. Ribosome-inactivating proteins from root tubers and seeds of Trichosanthes kirilowii and other Trichosanthes species. Protein Pept. Lett. 2016, 23, 699–706. [Google Scholar] [CrossRef] [PubMed]
- Luna, A.S.; Silva, A.P.; Pinho, J.S.A.; Ferré, J.; Boqué, R. Rapid characterization of transgenic and non-transgenic soybean oils by chemometric methods using NIR spectroscopy. Spectrochim Acta Part A Mol. Biomol. Spectrosc. 2013, 100, 115–119. [Google Scholar] [CrossRef] [PubMed]
- Chen, Q.S.; Zhao, J.W.; Zhang, H.D.; Liu, M.H. Application of near infrared reflectance spectroscopy to the identification of tea using SIMCA pattern recognition method. Food Sci. 2006, 27, 186–189. [Google Scholar]
- Lou, Z.Q.; Qin, B. Species Systematization and Quality Evaluation of Commonly Used Chinese Traditional Drugs (Volume 3); Associated Press of Beijing Medical University and Peking Union Medical College: Beijing, China, 1996; pp. 579–680. [Google Scholar]
- Fix, E.; Hodges, J.L. Discriminatory analysis. Nonparametric discrimination: consistency properties. Int. Stat. Rev. 1989, 57, 238–247. [Google Scholar] [CrossRef]
- Coomans, D.; Massart, D.L. Alternative k-nearest neighbour rules in supervised pattern recognition: Part 2. Probabilistic classification on the basis of the kNN method modified for direct density estimation. Anal. Chim. Acta. 1982, 138, 153–165. [Google Scholar] [CrossRef]
- Wold, S. Pattern recognition by means of disjoint principal components models. Pattern Recogn. 1976, 8, 127–139. [Google Scholar] [CrossRef]
- Sjöström, M.; Kowalski, B.R. A comparison of five pattern recognition methods based on the classification results from six real data bases. Anal. Chim. Acta. 1979, 112, 11–30. [Google Scholar] [CrossRef]
- Wold, S.; Sjöström, M.; Eriksson, L. PLS in chemistry. In The Encyclopedia of Computational Chemistry; Schleyer, P.V.R., Allinger, N.L., Clark, T., Gasteiger, J., Kollman, P.A., Schaefer, H.F., Schreiner, P.R., Eds.; Wiley: Chichester, UK, 1998; pp. 2006–2020. [Google Scholar]
- Dong, G.; Guo, J.; Wang, C.; Chen, Z.L.; Zheng, L.; Zhu, D.Z. The classification of wheat varieties based on near infrared hyperspectral imaging and information fusion. Spectrosc. Spect. Anal. 2015, 35, 3369–3374. [Google Scholar]
- Chen, Q.S.; Zhao, J.W.; Lin, H. Study on discrimination of roast green tea (Camellia sinensis L.) according to geographical origin by FT-NIR spectroscopy and supervised pattern recognition. Spectrochim. Acta Part A Mol. Biomol. Spectrosc. 2009, 72, 845–850. [Google Scholar] [CrossRef]
Sample Availability: Samples of the compounds 5-hydroxymethylfurfural, vanillic acid, and quercetin are available from the authors. |
| Sub-Models | Labels | PCs | Calibration Set | Prediction Set | ||
|---|---|---|---|---|---|---|
| Nright/N0 | CA% | Nright/N0 | CA% | |||
| 1 | Hebei | 6 | 19/20 | 95 | 9/10 | 90 |
| 2 | Shanxi | 4 | 19/20 | 95 | 10/10 | 100 |
| 3 | Shandong | 4 | 17/20 | 85 | 9/10 | 90 |
| 4 | Henan | 5 | 18/20 | 90 | 10/10 | 100 |
| 5 | Prepared slice | 5 | 20/20 | 100 | 10/10 | 100 |
| Classification Models | Optimal Parameters | Classification Accuracy | |
|---|---|---|---|
| Calibration Set (%) | Prediction Set (%) | ||
| KNN | K = 3 | 92 | 94 |
| SIMCA | PCs = 6, 4, 4, 5, 5 | 93 | 96 |
| PLS-DA | PCs = 8 | 95 | 98 |
| SVM-DA | C = 100, γ = 0.00316, PCs = 6 | 100 | 100 |
| Sample No. | Sample Type | Geographic Origins | Geographic Location * | Harvesting Time |
| 1–30 | Crude drug | Jinan, Shandong | 36°19′ N, 116°19′ E, 127–131 m | Oct 12, 2017 |
| 31–60 | Crude drug | Anyang, Henan | 36°03′ N, 114°23′ E, 68–70 m | Oct 17, 2017 |
| 61–90 | Crude drug | Anguo, Hebei | 38°21′ N, 115°16′ E, 32–33 m | Oct 1, 2017 |
| 91–120 | Crude drug | Houma, Shanxi | 35°19′ N, 111°04′ E, 461–466 m | Oct 4, 2017 |
| 121–150 | Prepared slices | Uncertain | Uncertain | Jan-Mar, 2018 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Xu, L.; Sun, W.; Wu, C.; Ma, Y.; Chao, Z. Discrimination of Trichosanthis Fructus from Different Geographical Origins Using Near Infrared Spectroscopy Coupled with Chemometric Techniques. Molecules 2019, 24, 1550. https://doi.org/10.3390/molecules24081550
Xu L, Sun W, Wu C, Ma Y, Chao Z. Discrimination of Trichosanthis Fructus from Different Geographical Origins Using Near Infrared Spectroscopy Coupled with Chemometric Techniques. Molecules. 2019; 24(8):1550. https://doi.org/10.3390/molecules24081550
Chicago/Turabian StyleXu, Liang, Wen Sun, Cui Wu, Yucui Ma, and Zhimao Chao. 2019. "Discrimination of Trichosanthis Fructus from Different Geographical Origins Using Near Infrared Spectroscopy Coupled with Chemometric Techniques" Molecules 24, no. 8: 1550. https://doi.org/10.3390/molecules24081550
APA StyleXu, L., Sun, W., Wu, C., Ma, Y., & Chao, Z. (2019). Discrimination of Trichosanthis Fructus from Different Geographical Origins Using Near Infrared Spectroscopy Coupled with Chemometric Techniques. Molecules, 24(8), 1550. https://doi.org/10.3390/molecules24081550