Tri-Cyclic Nucleobase Analogs and Their Ribosides as Substrates of Purine-Nucleoside Phosphorylases. II Guanine and Isoguanine Derivatives
Abstract
1. Introduction
2. Results and Discussion
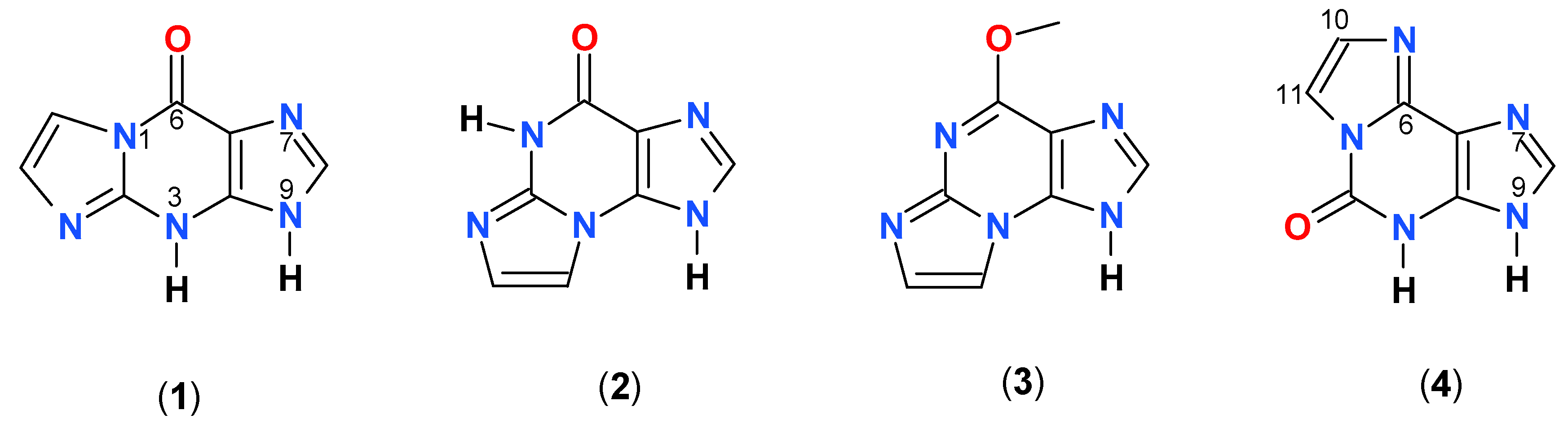
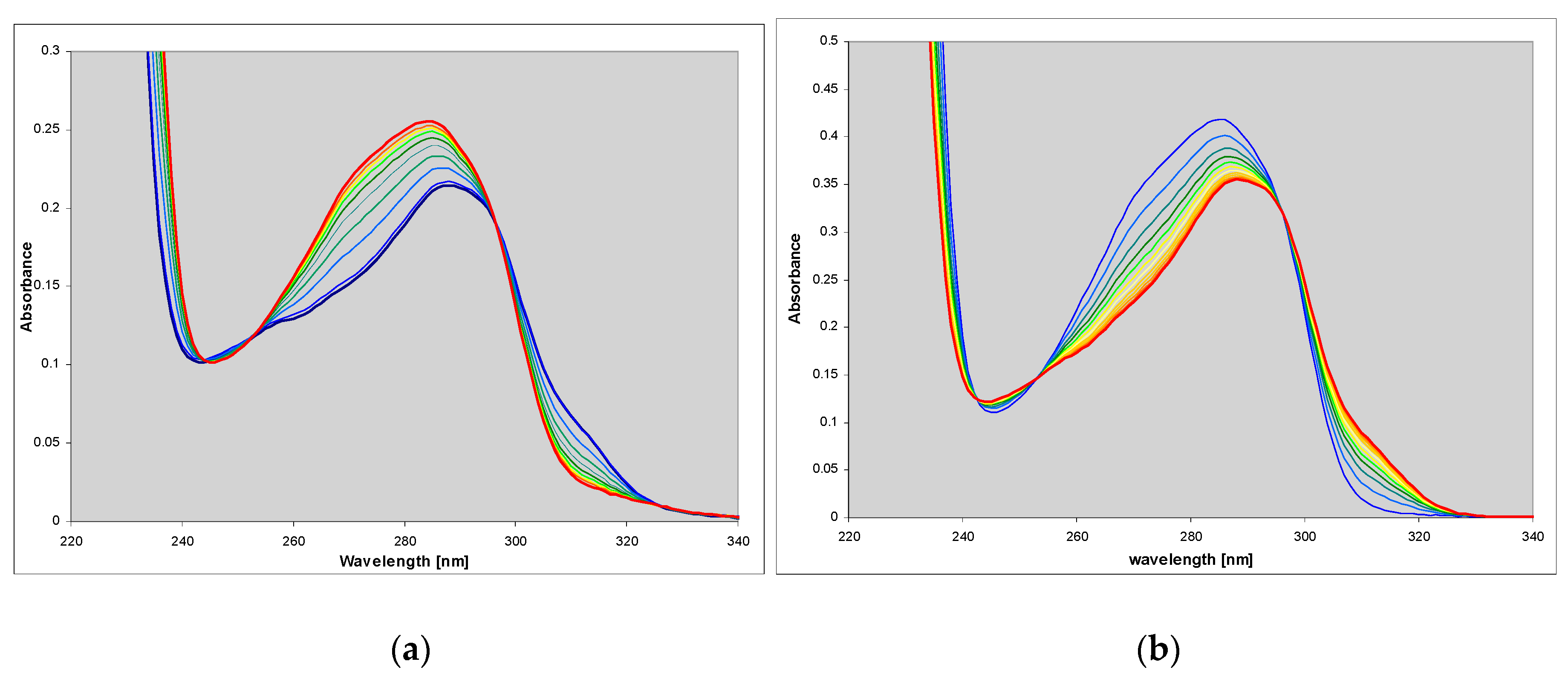
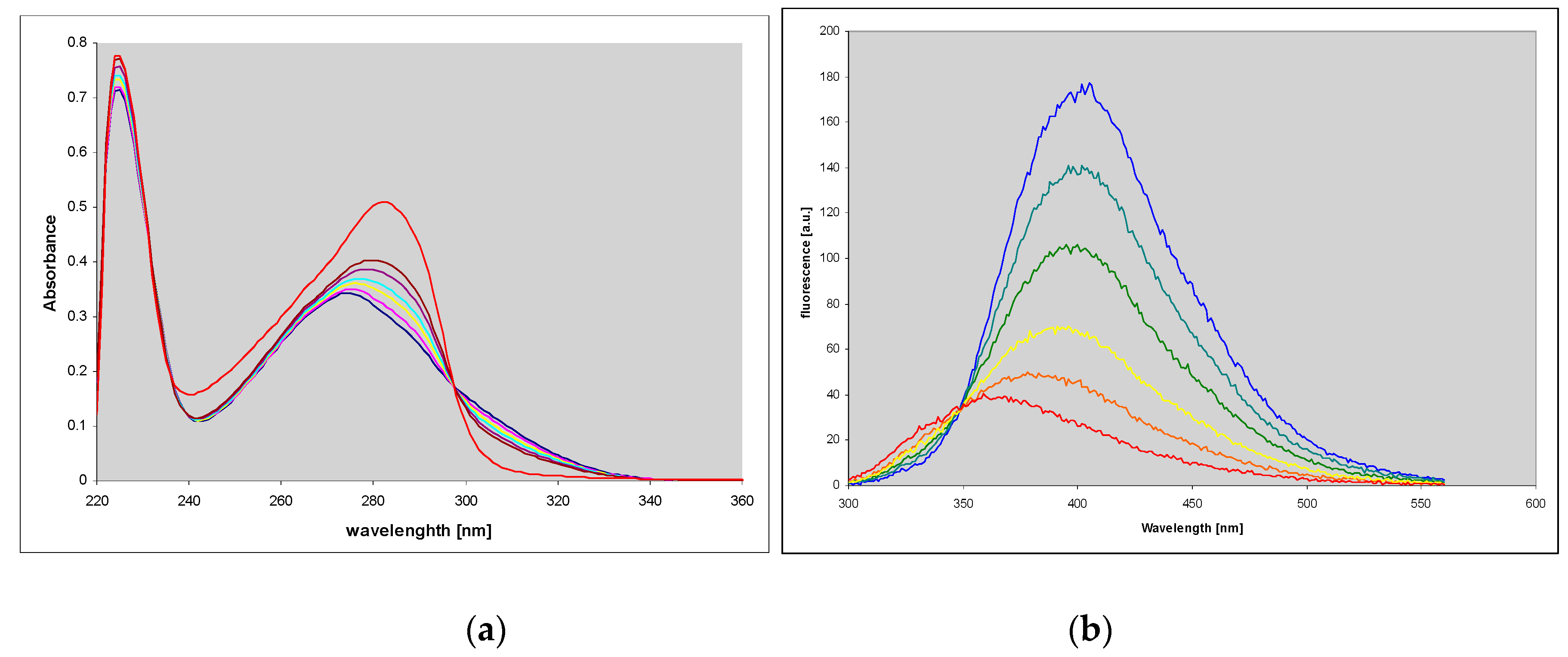
2.1. Properties of 1,N2-ethenoguanine and 1,N2-ethenoguanosine
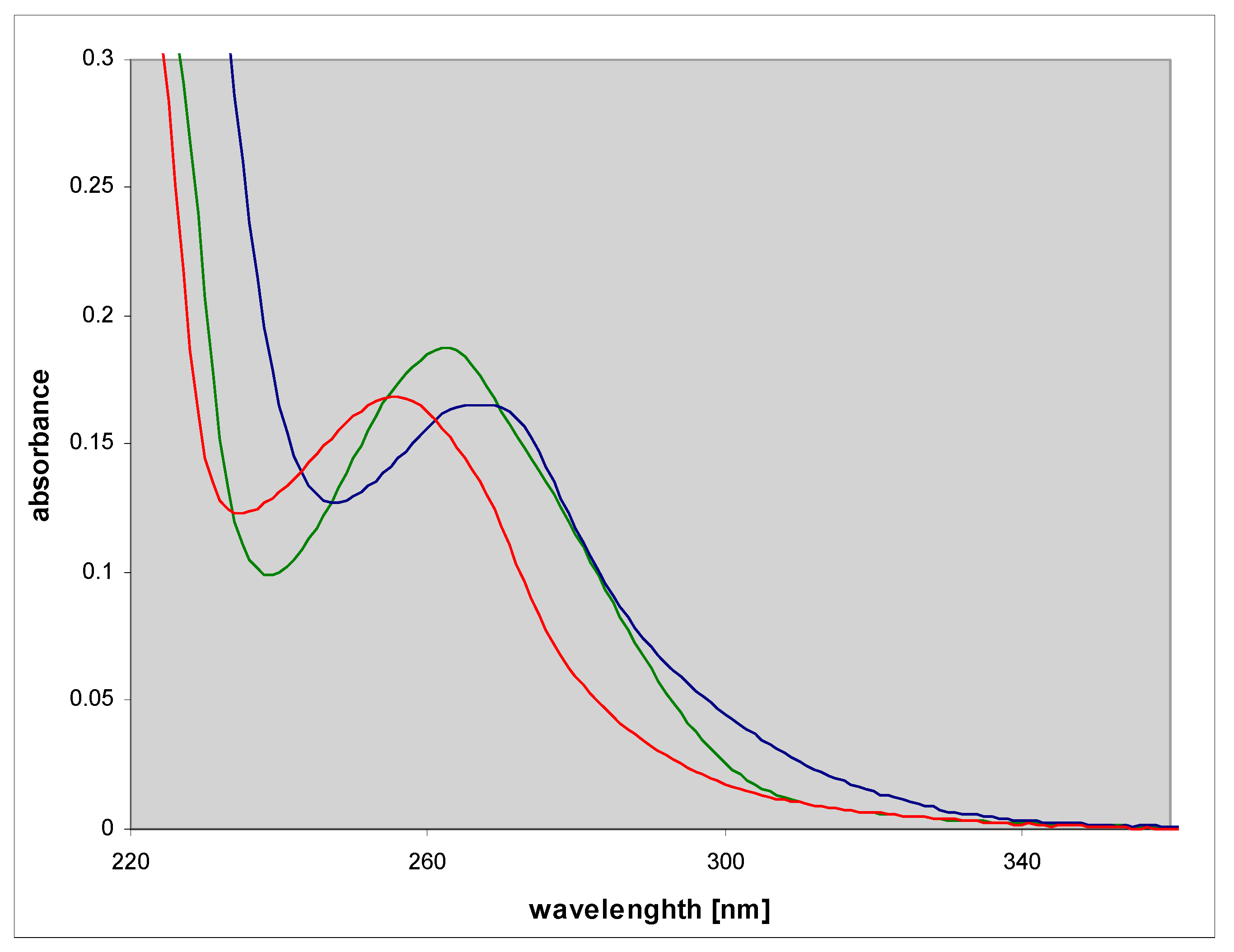
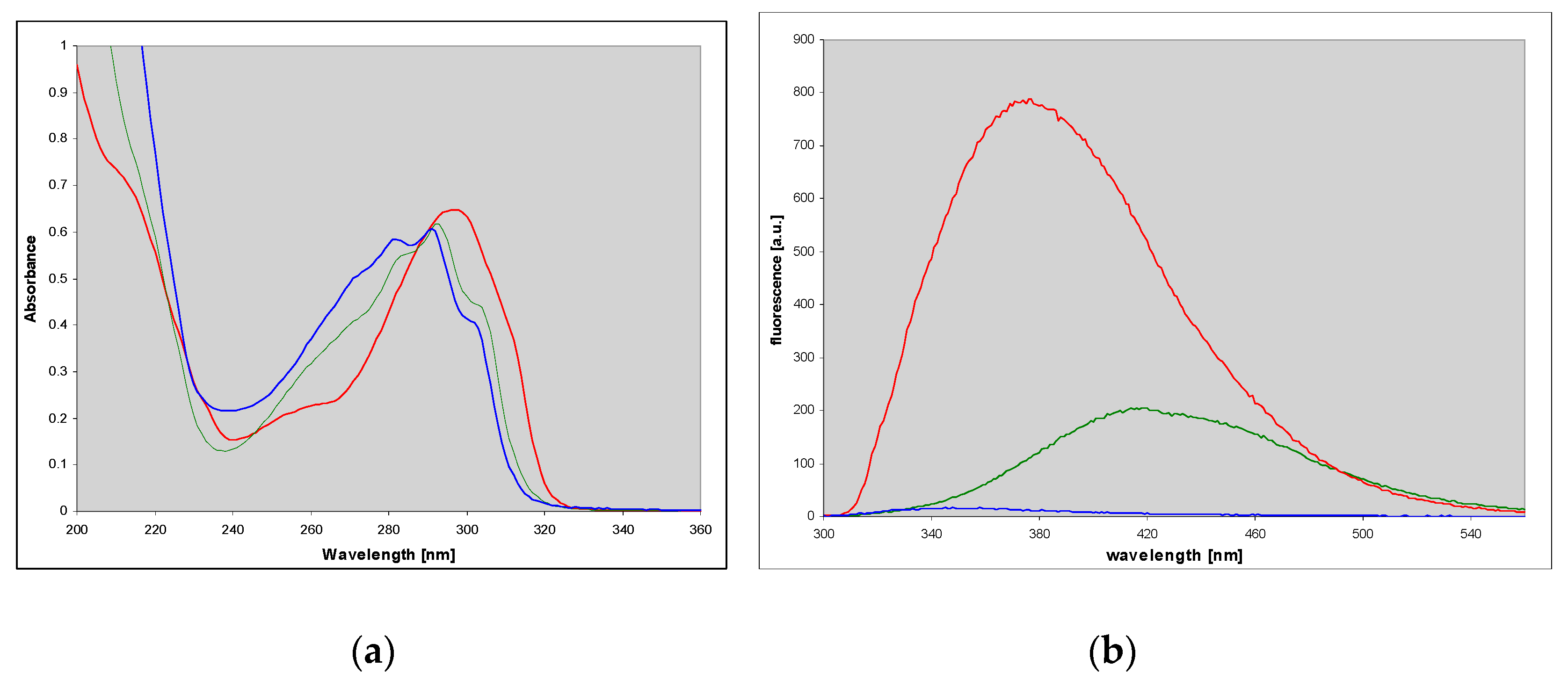
2.2. Spectral Properties of N2,3-ethenoguanine
2.3. N2,3-etheno-O6-methylguanine (3)
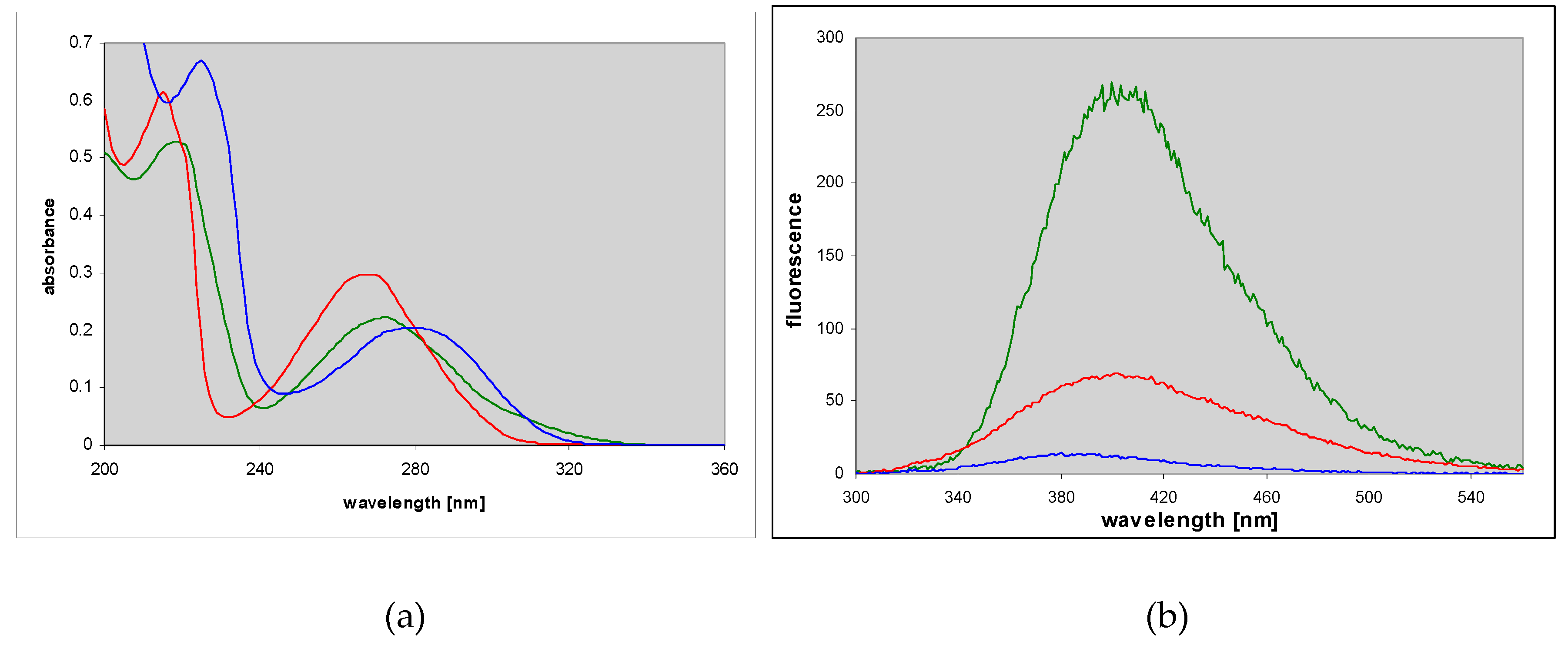
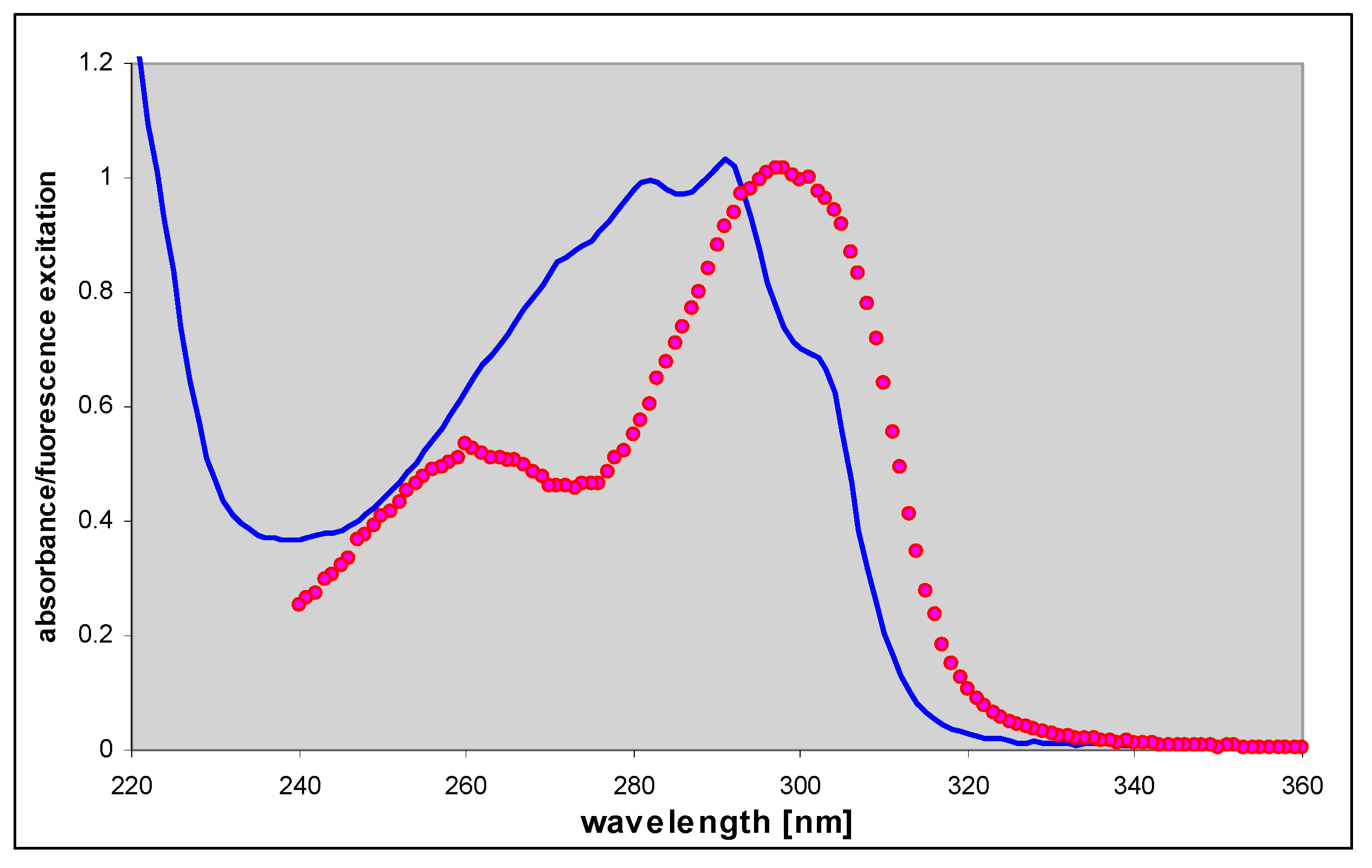
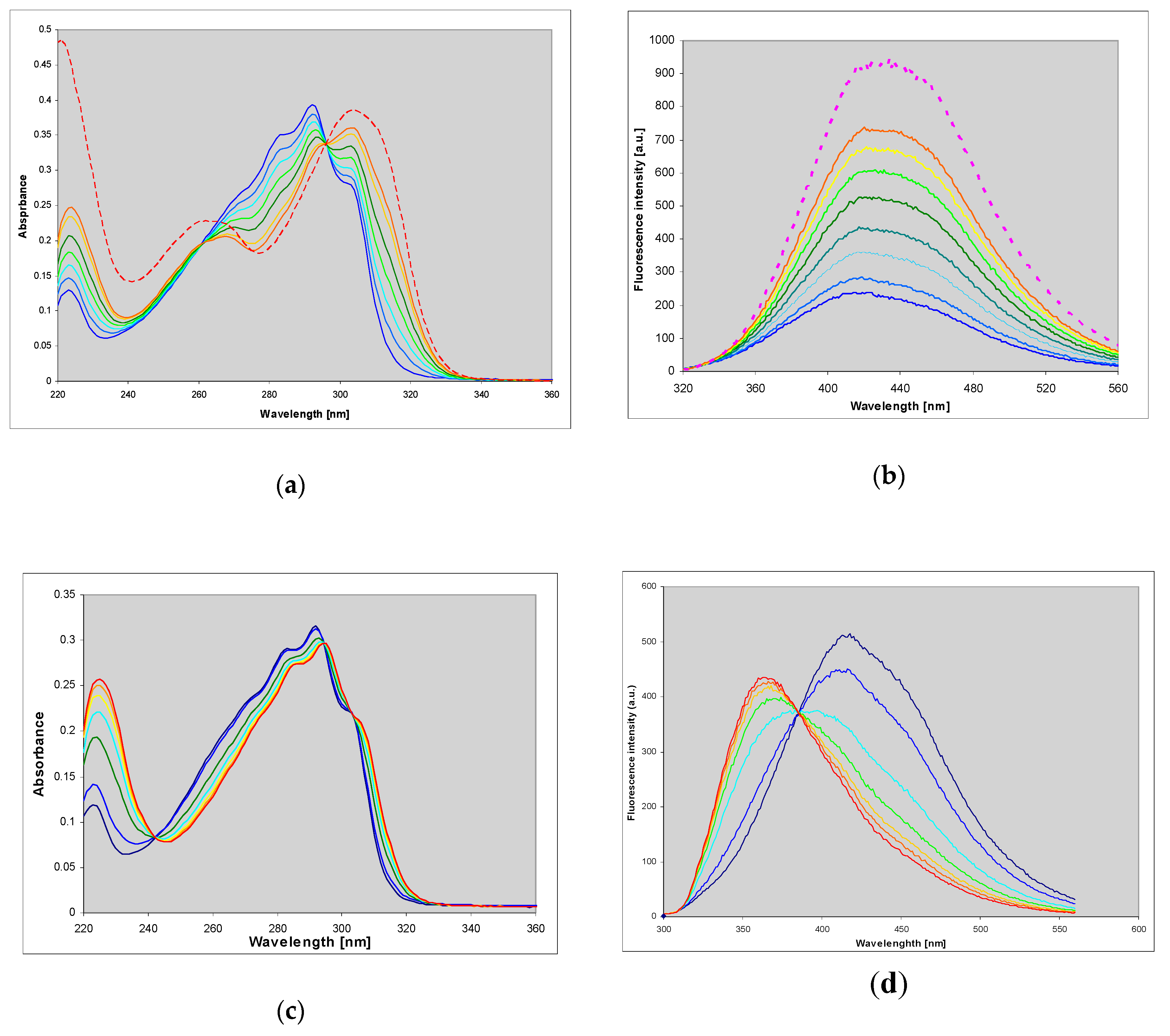
2.4. 1,N6-etheno-isoguanine (4)
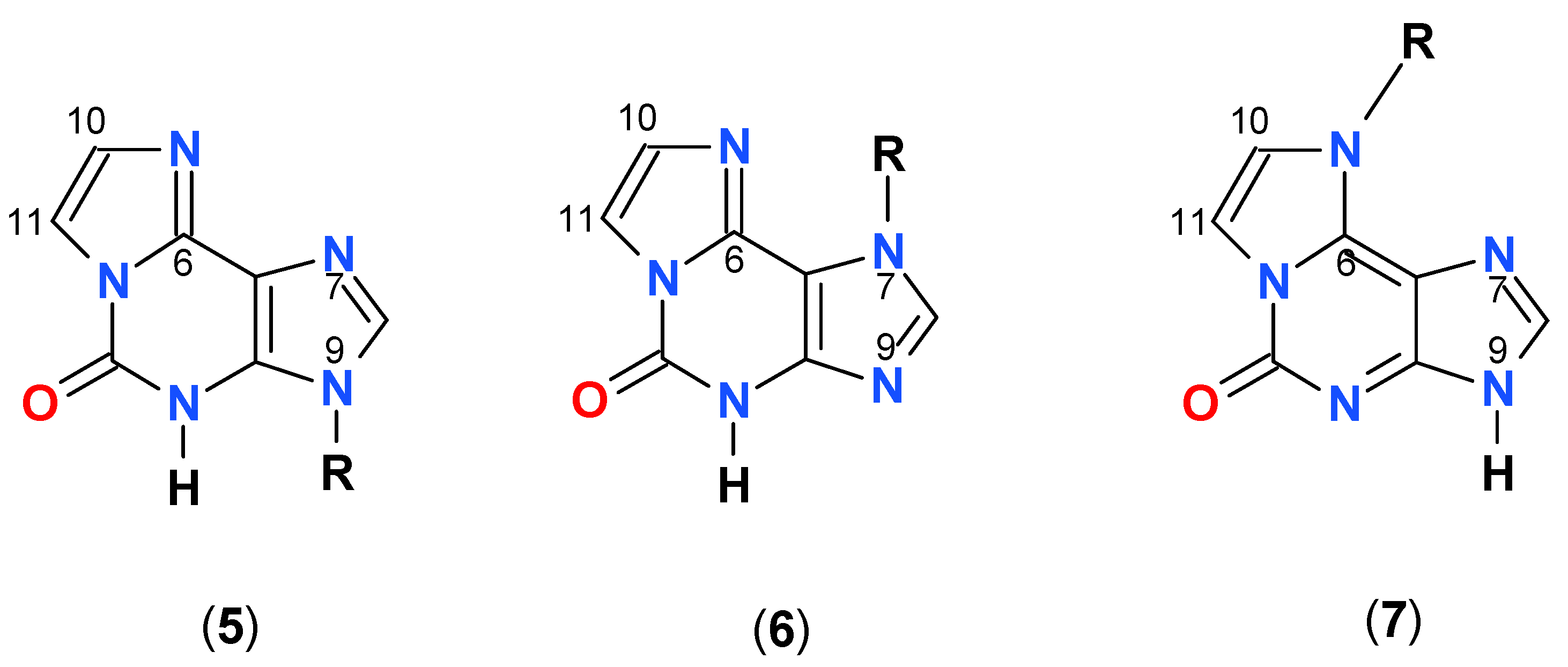
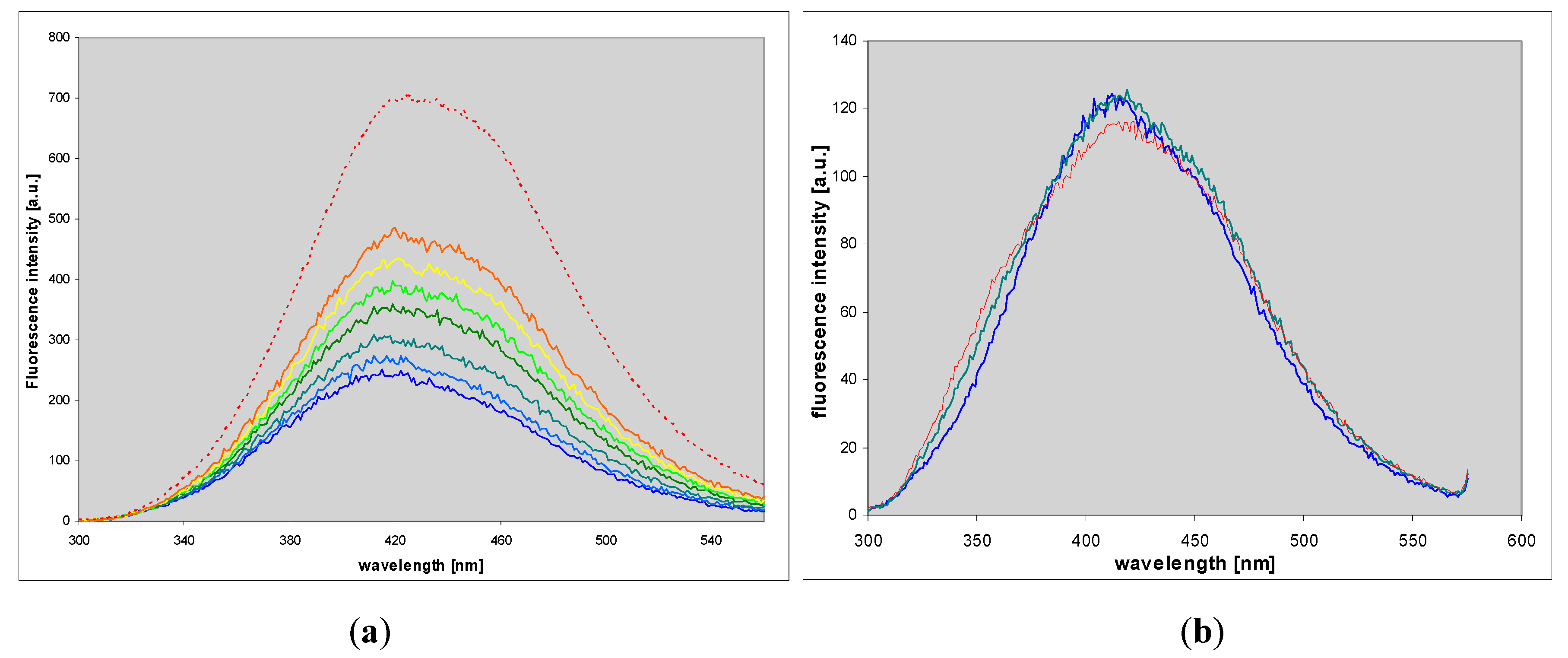
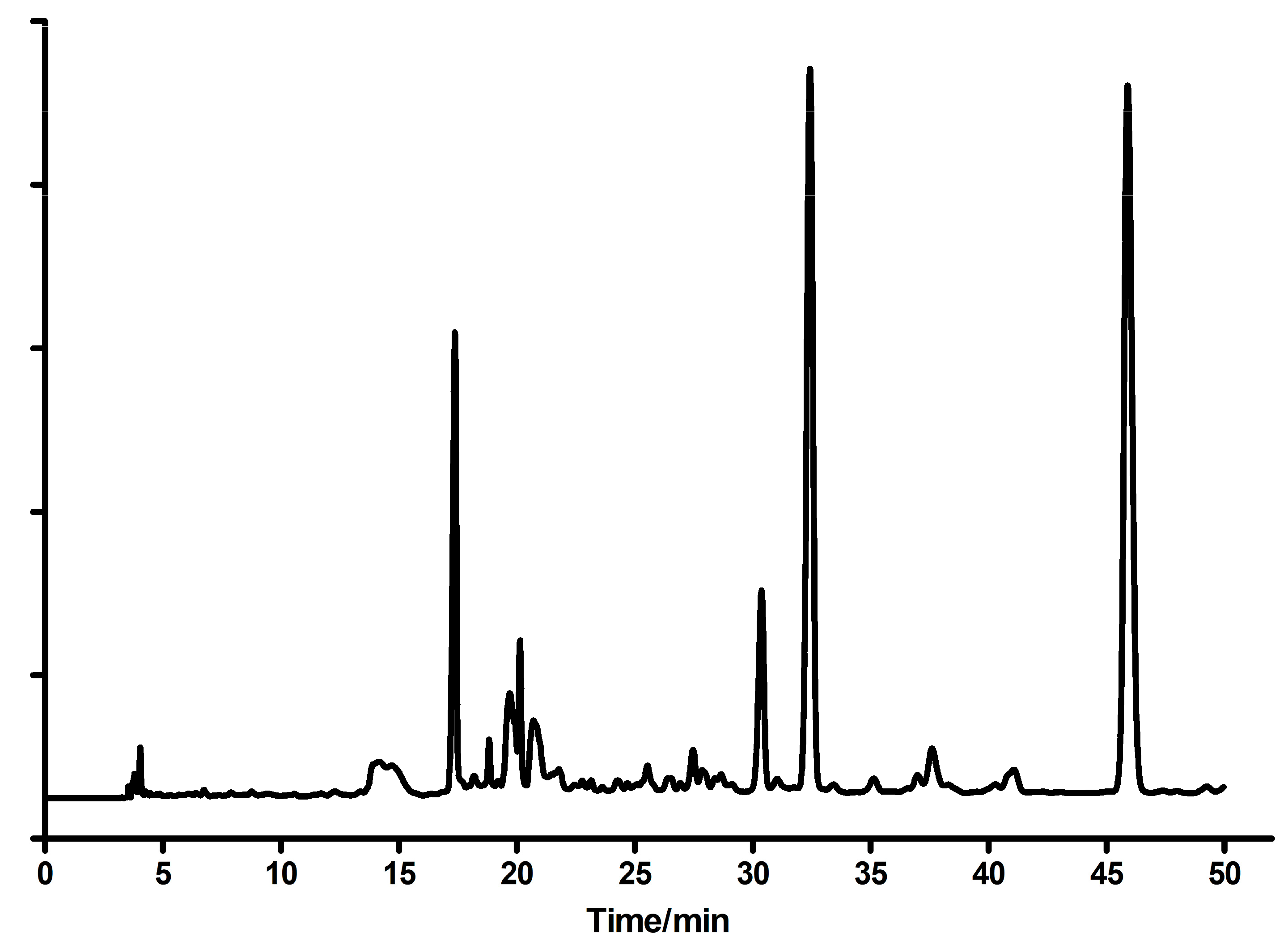
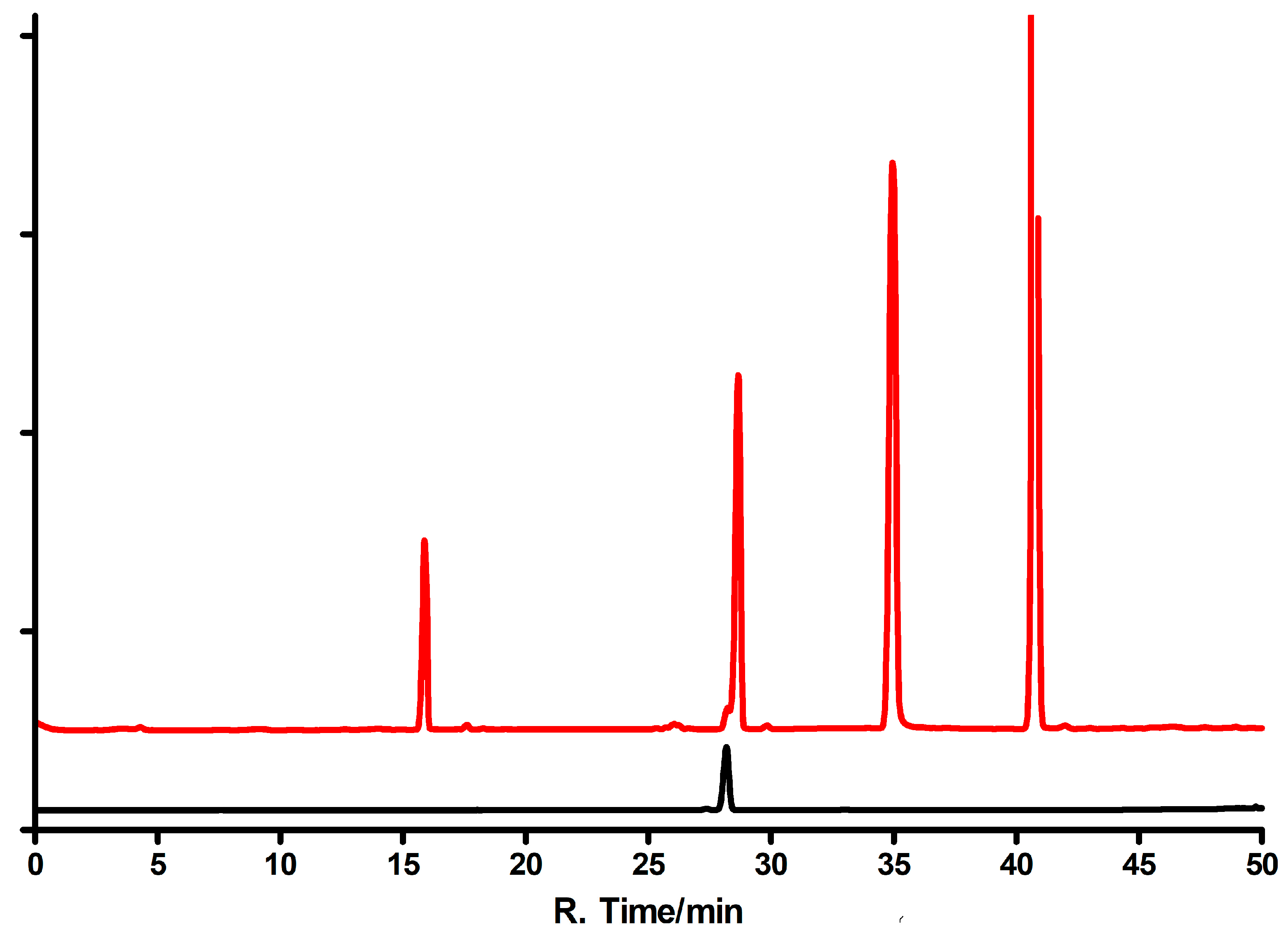
Ribosides of 1,N6-etheno-isoguanine
3. Experimental Section
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
References
- Leonard, N.J. Adenylates: Bound and unbound. Biopolymers 1985, 24, 9–28. [Google Scholar] [CrossRef] [PubMed]
- Leonard, N.J. Etheno-substituted nucleotides and coenzymes: Fluorescence and biological activity. CRC Crit. Revs. Biochem. 1984, 15, 125–199. [Google Scholar] [CrossRef]
- Sattsangi, P.D.; Leonard, N.J.; Frihart, C.R. 1,N2-ethenoguanine and N2,3-ethenoguanine: Synthesis and comparison of the electronic spectral properties of these linear and angular triheterocycles related to the Y bases. J. Org. Chem. 1977, 42, 3292–3296. [Google Scholar] [CrossRef] [PubMed]
- Sattsangi, P.D.; Barrio, J.R.; Leonard, N.J. 1,N6-etheno bridged adenines and adenosines. Alkyl substitution, fluorescence properties, and synthetic applications. J. Am. Chem. Soc. 1980, 102, 770–774. [Google Scholar] [CrossRef]
- Boryski, J. 1,N2-ethenoguanosine: Three methods of synthesis. Nucleosides Nucleotides Nucleic Acids 1990, 9, 803–813. [Google Scholar] [CrossRef]
- Hořejši, K.; Pohl, R.; Holy, A. Tricyclic Purine Analogs Derived from 2-Amino-6-chloropurine and 2,6-Diaminopurine and Their Methylated Quaternary Salts. Coll. Czech. Chem. Commun. 2006, 71, 77–90. [Google Scholar] [CrossRef]
- Virta, P.; Holmstrom, T.; Roslund, M.U.; Mattjus, P.; Kronberg, L.; Sjoholm, R. New nucleoside analogs from 2-amino-9-(β-d-ribofuranosyl)-purine. Org. Biomol. Chem. 2004, 2, 821–827. [Google Scholar] [CrossRef] [PubMed]
- Virta, P.; Koch, A.; Roslund, M.U.; Mattjus, P.; Kleinpeter, E.; Kronberg, L.; Sjoholm, R.; Klika, K.D. Synthesis, characterisation and theoretical calculations of 2,6-diaminopurine etheno derivatives. Org. Biomol. Chem. 2005, 3, 2924–2929. [Google Scholar] [CrossRef] [PubMed]
- Golankiewicz, B.; Ostrowski, T. Tricyclic nucleoside analogues as antiherpes agents. Antiviral Res. 2006, 71, 134–140. [Google Scholar] [CrossRef]
- Jahnz-Wechmann, Z.; Framski, G.R.; Januszczyk, P.A.; Boryski, J. Bioactive fused heterocycles: Nucleoside analogs with an additional ring. Eur. J. Med. Chem. 2015, 97, 388–396. [Google Scholar] [CrossRef]
- Jahnz-Wechmann, Z.; Framski, G.R.; Januszczyk, P.A.; Boryski, J. Base-modified nucleosides: Etheno derivatives. Front. Chem. 2016, 4, 19–29. [Google Scholar] [CrossRef] [PubMed]
- Singer, B.; Kuśmierek, J.T. Chemical mutagenesis. Annual Rev. Biochem. 1982, 51, 655–693. [Google Scholar] [CrossRef]
- Bzowska, A.; Kulikowska, E.; Shugar, D. Purine nucleoside phosphorylases: Properties, functions, and clinical aspects. Pharmacol. Ther. 2000, 88, 349–425. [Google Scholar] [CrossRef]
- Wierzchowski, J.; Stachelska-Wierzchowska, A.; Wielgus-Kutrowska, B.; Bzowska, A. 1,N6-ethenoadenine and other fluorescent nucleobase analogues as substrates for purine-nucleoside phosphorylases: Spectroscopic and kinetic studies. Curr. Pharm. Des. 2017, 23, 6972–6990. [Google Scholar]
- Mikhailopulo, A.I.; Miroshnikov, A.I. Biologically important nucleosides: Modern trends in biotechnology and application. Mendeleev Commun. 2011, 21, 57–68. [Google Scholar] [CrossRef]
- Kamel, S.; Yehia, H.; Neubauer, P.; Wagner, A. Chapter 1. In Enzymatic and Chemical Synthesis of Nucleic Acid Derivatives; Fernandez-Lucas, J., Camarasa-Rius, M.J., Eds.; Wiley-VCH: Weinheim, Germany, 2019. [Google Scholar]
- Yehia, H.; Kamel, S.; Paulick, K.; Neubauer, P.; Wagner, A. Substrate Spectra of Nucleoside Phosphorylases and their Potential in the Production of Pharmaceutically Active Compounds. Curr. Pharm. Des. 2017, 23, 6913–6935. [Google Scholar] [CrossRef] [PubMed]
- Stachelska-Wierzchowska, A.; Wierzchowski, J.; Bzowska, A.; Wielgus-Kutrowska, B. Tricyclic nitrogen base, 1,N6-Ethenoadenine, and its Ribosides, as Substrates for Purine-Nucleoside Phosphorylases: Spectroscopic and Kinetic Studies. Nucleosides Nucleotides Nucleic Acids 2018, 37, 89–101. [Google Scholar] [CrossRef] [PubMed]
- Schramm, V.L. Transition states, analogues, and drug development. ACS Chem. Biol. 2013, 8, 71–81. [Google Scholar] [CrossRef]
- Kusmierek, J.T.; Singer, B. 1,N2-ethenodeoxyguanosine—Properties and formation in chloroacetaldehyde-treated polynucleotides and DNA. Chem. Res. Toxicol. 1992, 25, 634–638. [Google Scholar] [CrossRef]
- Patra, A.; Su, Y.; Zhang, Q.Q.; Johnson, K.M.; Guengerich, F.P.; Egli, M. Structural and Kinetic Analysis of Miscoding Opposite the DNA Adduct 1,N6-Ethenodeoxyadenosine by Human Translesion DNA Polymerase. J. Biol. Chem. 2016, 291, 14134–14145. [Google Scholar] [CrossRef]
- Choi, J.Y.; Zang, H.; Angel, K.C.; Kozekov, I.D.; Goodenough, A.K.; Rizzo, C.J.; Guengerich, F.P. Translesion synthesis across 1,N2-ethenoguanine by human DNA polymerases. Chem. Res. Toxicol. 2006, 19, 879–886. [Google Scholar] [CrossRef] [PubMed]
- Kusmierek, J.T.; Jensen, D.; Spengler, S.J.; Stolarski, R.; Singer, B. Synthesis and properties of N2,3-ethenoguanosine and N2,3-ethenoguanosine 5′-diphosphate. J. Org. Chem. 1987, 52, 2374–2378. [Google Scholar] [CrossRef]
- Chang, S.C.; Fedeles, B.I.; Wu, J.; Delaney, J.C.; Li, D.Y.; Zhao, L.L.; Christov, P.P.; Yau, E.; Singh, V.; Jost, M. Next-generation sequencing reveals the biological significance of the N2,3-ethenoguanine lesion in vivo. Nucleic Acids Res. 2015, 43, 5489–5500. [Google Scholar] [CrossRef] [PubMed]
- Zhao, L.L.; Pence, M.G.; Christov, P.P.; Wawrzak, Z.; Choi, J.Y.; Rizzo, C.J.; Egli, M.; Guengerich, F.P. Basis of Miscoding of the DNA Adduct N2,3-Ethenoguanine by Human Y-family DNA Polymerases. J. Biol. Chem. 2012, 287, 35516–35526. [Google Scholar] [CrossRef] [PubMed]
- Yarosh, D.B. The role of O6-methylguanine-DNA methyltransferase in cell survival, mutagenesis and carcinogenesis. Mutat. Res. 1985, 145, 1–16. [Google Scholar] [CrossRef][Green Version]
- Rasouli-Nia, A.; Sibghat-Ullah; Mirzayans, R.; Paterson, M.C.; Day, R.S. On the quantitative relationship between O6-methylguanine residues in genomic DNA and production of sister-chromatid exchanges, mutations and lethal events in a Mer- human tumor cell line. Mutat. Res. 1994, 314, 99–113. [Google Scholar] [CrossRef]
- Sepioł, J.; Kazimierczuk, Z.; Shugar, D. Tautomerism of isoguanosine and solvent-induced keto-enol equilibrium. Z. Naturforsch. 1976, 31c, 361–370. [Google Scholar] [CrossRef]
- Cheng, Q.; Compaan, K.R.; Schaffer, H.F., III. Isoguanine formation from adenine. Chem. Eur. J. 2012, 18, 4877–4886. [Google Scholar] [CrossRef]
- Bukowska, A.M.; Kusmierek, J.T. Miscoding properties of isoguanine (2-oxoadenine) studied in an AMV reverse transcriptase in vitro system. Acta Biochim. Polon. 1996, 43, 247–254. [Google Scholar]
- Kemena, A.; Fernandez, M.; Bauman, J.; Keating, M.; Plunkett, W. A sensitive fluorescence assay for quantitation of fludarabine and metabolites in biological-fluids. Clin. Chim. Acta 1991, 200, 95–106. [Google Scholar] [CrossRef]
- Holmén, A.; Albinsson, B.; Nordén, B. Electronic transition dipole moments of the 1,N6-ethenoadenine chromophore. J. Phys. Chem. 1994, 98, 13460–13469. [Google Scholar] [CrossRef]
- Stachelska-Wierzchowska, A.; Wierzchowski, J.; Wielgus-Kutrowska, B.; Mikleušević, G. Enzymatic synthesis of highly fluorescent 8-azapurine ribosides using purine-nucleoside phosphorylase reverse reaction: Variable ribosylation sites. Molecules 2013, 18, 12587–12598. [Google Scholar] [CrossRef]
- Dračínský, M.; Pohl, R. NMR Studies of Purines. Annu. Rep. NMR Spectr. 2014, 82, 59–113. [Google Scholar]
- Markley, J.L.; Bax, A.; Arata, Y.; Hilbers, C.W.; Kaptein, R.; Sykes, B.D.; Wright, P.E.; Wüthrich, K. Recommendations for the presentation of NMR structures of proteins and nucleic acids–IUPAC-IUBMB-IUPAB Inter-Union Task Group on the standardization of data bases of protein and nucleic acid structures determined by NMR spectroscopy. J. Biomol. NMR 1998, 12, 1–23. [Google Scholar] [CrossRef] [PubMed]
- Sinkeldam, R.W.; Greco, N.J.; Tor, Y. Fluorescent analogs of biomolecular building blocks: Design, properties, and applications. Chem. Rev. 2010, 110, 2579–2619. [Google Scholar] [CrossRef]
- Stoeckler, J.D.; Poirot, A.F.; Smith, R.M.; Robert, E.; Parks, R.E.; Ealick, S.E.; Takabayashi, K.; Erion, M.D. Purine nucleoside phosphorylase. 3. Reversal of purine base specificity by site-directed mutagenesis. Biochemistry 1997, 36, 11749–11756. [Google Scholar] [CrossRef]
- Stachelska-Wierzchowska, A.; Wierzchowski, J.; Bzowska, A.; Wielgus-Kutrowska, B. Site-selective ribosylation of fluorescent nucleobase analogs using purine-nucleoside phosphorylase as a catalyst: Effects of point mutations. Molecules 2016, 21, 44. [Google Scholar] [CrossRef]
- Wierzchowski, J.; Mędza, G.; Szabelski, M.; Stachelska-Wierzchowska, A. Properties of 2,6-diamino-8-azapurine, a highly fluorescent purine analog and its N-alkyl derivatives: Tautomerism and excited-state proton transfer reactions. J. Photochem. Photobiol. 2013, 265, 49–57. [Google Scholar] [CrossRef]
- Harris, R.K.; Becker, E.D.; De Menezes, S.M.C.; Granger, P.; Hoffman, R.E.; Zilm, K.W. Further conventions for NMR shielding and chemical shifts (IUPAC Recommendations 2008). Pure Appl. Chem. 2008, 80, 59–84. [Google Scholar] [CrossRef]
- Goddard, T.D.; Kneller, D.G. SPARKY 3; University of California: San Francisco, CA, USA, 2000. [Google Scholar]
- Mikleuševic, G.; Štefanic, Z.; Narczyk, M.; Wielgus-Kutrowska, B.; Bzowska, A.; Luic, M. Validation of the catalytic mechanism of Escherichia coli purine nucleoside phosphorylase by structural and kinetic studies. Biochimie 2011, 93, 1610–1622. [Google Scholar] [CrossRef]
- Breer, K.; Girstun, A.; Wielgus-Kutrowska, B.; Staron, K.; Bzowska, A. Overexpression, purification and characterization of functional calf purine nucleoside phosphorylase (PNP). Prot. Expr. Purif. 2008, 61, 122–130. [Google Scholar] [CrossRef] [PubMed]
- Calabretta, A.; Leumann, C.J. Base pairing and miscoding properties of 1,N6-ethenoadenine- and 3,N4-ethenocytosine-containing RNA oligonucleotides. Biochemistry 2013, 52, 1990–1997. [Google Scholar] [CrossRef] [PubMed]
- Maciejewska, A.M.; Sokolowska, B.; Nowicki, A.; Kusmierek, J.T. The role of AlkB protein in repair of 1,N6-ethenoadenine in Escherichia coli cells. Mutagenesis 2011, 26, 401–406. [Google Scholar] [CrossRef]
- Mikhailopulo, I.A. Biotechnology of nucleic acid components—State of the art and perspectives. Curr. Org. Chem. 2007, 11, 317–335. [Google Scholar] [CrossRef]
- Ye, W.; Paul, D.; Gao, L.; Seckute, J.; Sangaiah, R.; Jayaraj, K.; Zhang, Z.; Kaminski, P.A.; Ealick, S.E.; Gold, A.; Ball, L.M. Ethenoguanines Undergo Glycosylation by Nucleoside 29-Deoxyribosyltransferases at Non-Natural Sites. PLoS ONE 2014, 9, e115082. [Google Scholar] [CrossRef] [PubMed]


| Compound | pKa ±0.1 | Form* (pH) | UV Absorption | Fluorescence | |||
|---|---|---|---|---|---|---|---|
| λmax {nm} | εmax {M−1cm−1} | λmax {nm} | ϕ | τ {ns} | |||
| 1,N2-ethenoguanine | ~2 ~9.5 | n (6) c (1) ma (11) | 287 291 303 | 9700 ~8600 7200 | - - ~400 | - - <0.01 | - - nd ** |
| N9-β-d-ribofuranosyl- | 8.8 | n (6.5) c (1) ma (11) | 284 nd 307 | 11900 nd 8400 | - - ~400 | - - <0.01 | - - nd |
| N2,3-ethenoguanine | ~2.5; 8.8 | n (6.5) ma (11) c (1) | 261 266 255 | 12850 11300 11500 | 400 - - | 0.02 - - | 1.38 - - |
| N2,3-etheno- O6methylguanine | ~3.5; ~9 | n (6.5) c (2) ma (12) | 272 268 280 | 9900 13300 9100 | 405 395 ~380 | 0.11 0.035 <0.01 | nd nd nd |
| 1,N6-etheno- isoguanine | 3.5; 8.0 | n (6.5) c (2) ma (11) | 292 296 290 | 7000 7300 6900 | 415 385 350 | 0.17 0.44 ~0.01 | 5.15; 3.4 nd nd |
| N9-β-d-ribofuranosyl- | 2.4; 7.1 | n (5.5) c (1.3) ma (11) | 295 294 281 | 7200 ~7000 ~7400 | 415 380 - | 0.34 0.275 <0.01 | 6.1 4.4; 0.81 nd |
| N7-β-d-ribofuranosyl- | 2.8; ~8.5 | n (6) c (2) ma (9) | 294 301 297 | 7100 ~7200 ~6500 | 360 355 365 | 0.036 ~0.4 ~0.2 | 0.8; 5.2 3.5 nd |
| N6-β-d-ribofuranosyl- | ~3; ~8.5 | n (6.5) c (1.7) ma (11) | 303 298 312 | 6900 nd nd | 425 405 450 | 0.66 0.82 ~0.03 | nd nd nd |
| Substrate/Enzyme | Km (µM) | Vmax (relative) * | Approximate Product Ratio: N9: N7-riboside: N6-riboside |
|---|---|---|---|
| 1,N2-ethenoguanine /calf PNP-wt** | nd | <0.5 | predominantly N9 |
| 1,N2-ethenoguanine /calf PNP-mutantN243D | nd | - | - |
| 1,N2-ethenoguanine /E.coli PNP-wt | 48 | 18 | predominantly N9 |
| 1,N2-ethenoguanine /E.coli PNP-mutantD204N | nd | >15 | predominantly N9 |
| N2,3-ethenoguanine /calf PNP-wt | - | <0.1 | - |
| N2,3-ethenoguanine /E.coli PNP-wt | - | <0.1 | - |
| N2,3-etheno-O6-methylguanine /calf PNP-wt | - | <0.1 | - |
| N2,3-etheno-O6-methylguanine /E.coli PNP-wt | ~6 | ~0.3 | Nd |
| 1,N6-ethenoisoguanine /calf PNP-wt | ~160 | 1.6 | 1:1:10 |
| 1,N6-ethenoisoguanine /calf PNP- mutantN243D | nd | ~1 | predominantly N6 |
| 1,N6-ethenoisoguanine /E.coli PNP-wt | 98 | ~24 | 1:4:0.1 |
| 1,N6-ethenoisoguanine /E.coli PNP-mutantD204N | nd | >5 | 5:5:0 |
| Atom | Sample/Chemical Shift (PPM) | ||
|---|---|---|---|
| N6-riboside | N7-riboside | N9-riboside | |
| H1’ | 6.511 | 5.987 | 5.712 |
| H8 | 7.822 | 8.262 | 7.753 |
| H10 * | 7.775 | 7.301/7.79 | 7.212/7.520 |
| H11 ** | 7.827 | ||
| C1’ | 90.293 | 89.833 | 89.114 |
| C5 | NA | 104.600 | NA |
| C6 | 138.682 | 137.918 | 141.788 |
| C8 | 137.799 | 140.026 | 135.525 |
| C10 | 113.386 | 130.645/113.699 | NA/111.558 |
| C11 | 116.951 | ||
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Stachelska-Wierzchowska, A.; Wierzchowski, J.; Górka, M.; Bzowska, A.; Wielgus-Kutrowska, B. Tri-Cyclic Nucleobase Analogs and Their Ribosides as Substrates of Purine-Nucleoside Phosphorylases. II Guanine and Isoguanine Derivatives. Molecules 2019, 24, 1493. https://doi.org/10.3390/molecules24081493
Stachelska-Wierzchowska A, Wierzchowski J, Górka M, Bzowska A, Wielgus-Kutrowska B. Tri-Cyclic Nucleobase Analogs and Their Ribosides as Substrates of Purine-Nucleoside Phosphorylases. II Guanine and Isoguanine Derivatives. Molecules. 2019; 24(8):1493. https://doi.org/10.3390/molecules24081493
Chicago/Turabian StyleStachelska-Wierzchowska, Alicja, Jacek Wierzchowski, Michał Górka, Agnieszka Bzowska, and Beata Wielgus-Kutrowska. 2019. "Tri-Cyclic Nucleobase Analogs and Their Ribosides as Substrates of Purine-Nucleoside Phosphorylases. II Guanine and Isoguanine Derivatives" Molecules 24, no. 8: 1493. https://doi.org/10.3390/molecules24081493
APA StyleStachelska-Wierzchowska, A., Wierzchowski, J., Górka, M., Bzowska, A., & Wielgus-Kutrowska, B. (2019). Tri-Cyclic Nucleobase Analogs and Their Ribosides as Substrates of Purine-Nucleoside Phosphorylases. II Guanine and Isoguanine Derivatives. Molecules, 24(8), 1493. https://doi.org/10.3390/molecules24081493





