New Dammarane-Type Saponins from Gynostemma pentaphyllum
Abstract
:1. Introduction
2. Results and Discussion
3. Materials and Methods
3.1. General Experimental Procedures
3.2. Plant Material
3.3. Extraction and Isolation
3.4. Spectroscopic Data (1H- and 13C-NMR spectra of 1-6 were also provided by the supplementary materials)
3.5. Acid Hydrolysis of Dammarane-Type Glycosides
3.6. Antiproliferation Assay
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Razmovski-Naumovski, V.; Huang, T.H.-W.; Tran, V.H.; Li, G.Q.; Duke, C.C.; Roufogalsi, B.D. Chemistry and Pharmacology of Gynostemma pentaphyllum. Phytochem. Rev. 2005, 4, 197–219. [Google Scholar] [CrossRef]
- Shi, G.; Wang, X.; Zhang, H.; Zhang, X.; Zhao, Y. New dammarane-type triterpene saponins from Gynostemma pentaphyllum and their anti-hepatic fibrosis activities in vitro. J. Funct. Foods 2018, 45, 10–14. [Google Scholar] [CrossRef]
- Nguyen, P.H.; Gauhar, R.; Hwang, S.L.; Dao, T.T.; Park, D.C.; Kim, J.E.; Song, H.; Huh, T.L.; Oh, W.K. New dammarane-type glucosides as potential activators of AMP-activated protein kinase (AMPK) from Gynostemma pentaphyllum. Bioorgan. Med. Chem. 2011, 19, 6254–6260. [Google Scholar] [CrossRef] [PubMed]
- Hung, T.M.; Hoang, D.M.; Kim, J.C.; Jang, H.-S.; Ahn, J.S.; Min, B.-S. Protein tyrosine phosphatase 1B inhibitory by dammaranes from Vietnamese Giao-Co-Lam tea. J. Ethnopharmacol. 2009, 124, 240–245. [Google Scholar] [CrossRef]
- Quan, Y.; Qian, M.Z. Effect and mechanism of gypenoside on the inflammatory molecular expression in high-fat induced atherosclerosis rats. Chin. J. Integr. Tradit.l Western Med. 2010, 30, 403–406. [Google Scholar]
- Cai, H.; Liang, Q.; Ge, G. Gypenoside attenuates β-amyloid-induced inflammation in N9 microglial cells via SOCS1 signaling. Neural Plast. 2016, 2016, 1–10. [Google Scholar] [CrossRef]
- Yang, F.; Shi, H.; Zhang, X.; Yang, H.; Zhou, Q.; Yu, L.L. Two new saponins from tetraploid jiaogulan (Gynostemma pentaphyllum), and their anti-inflammatory and α-glucosidase inhibitory activities. Food Chem. 2013, 141, 3606–3613. [Google Scholar] [CrossRef] [PubMed]
- Li, B.; Zhang, X.; Wang, M.; Jiao, L. Characterization and antioxidant activities of acidic polysaccharides from Gynostemma pentaphyllum (Thunb.) Markino. Carbohyd. Polym. 2015, 127, 209–214. [Google Scholar] [CrossRef] [PubMed]
- Jang, H.; Lee, J.W.; Lee, C.; Jin, Q.; Lee, M.K.; Lee, C.K.; Lee, M.K.; Hwang, B.Y. Flavonol glycosides from the aerial parts of Gynostemma pentaphyllum and their antioxidant activity. Arch. Pharm. Res. 2016, 39, 1232. [Google Scholar] [CrossRef]
- Cour, B.L.; Mølgaard, P.; Yi, Z. Traditional Chinese medicine in treatment of hyperlipidaemia. J Ethnopharmacol. 1995, 46, 125–129. [Google Scholar] [CrossRef]
- Gao, D.; Zhao, M.; Qi, X.; Liu, Y.; Li, N.; Liu, Z.; Bian, Y. Hypoglycemic effect of Gynostemma pentaphyllum saponins by enhancing the Nrf2 signaling pathway in STZ-inducing diabetic rats. Arch. Pharm. Res. 2016, 39, 221–230. [Google Scholar] [CrossRef]
- Huyen, V.T.T.; Phan, D.V.; Thang, P.; Hoa, N.K.; Östenson, C.G. Antidiabetic effect of Gynostemma pentaphyllum tea in randomly assigned type 2 diabetic patients. Horm. Metab. Res. 2010, 42, 353–357. [Google Scholar] [CrossRef]
- Li, Y.; Lin, W.; Huang, J.; Xie, Y.; Ma, W. Anti-cancer effects of Gynostemma pentaphyllum (Thunb.) Makino (Jiaogulan). Chinese Med. 2016, 11, 43. [Google Scholar] [CrossRef]
- Yuan, G.; Wei, J.; Zhou, J.; Guo, X.; Yang, M. Apoptosis of human hepatoma cells induced by Gynostemma pentaphyllum Makino. Chin.-Ger. J. Clin. Oncol. 2006, 5, 173–177. [Google Scholar] [CrossRef]
- Tsai, Y.; Lin, C.; Chen, B. Preparative chromatography of flavonoids and saponins in Gynostemma pentaphyllum and their antiproliferation effect on hepatoma cell. Phytomedicine 2010, 18, 2–10. [Google Scholar] [CrossRef]
- Liu, J.; Zhang, L.; Ren, Y.; Gao, Y.; Kang, L.; Qiao, Q. Anticancer and immunoregulatory activity of Gynostemma pentaphyllum polysaccharides in H22 tumor-bearing mice. Int. J. Biol. Macromol. 2014, 69, 1–4. [Google Scholar] [CrossRef]
- Cheng, T.-C.; Lu, J.-F.; Wang, J.-S.; Lin, L.-J.; Kuo, H.-I.; Chen, B.-H. Antiproliferation effect and apoptosis mechanism of prostate cancer cell PC-3 by flavonoids and saponins prepared from Gynostemma pentaphyllum. J. Agr. Food Chem. 2011, 59, 11319–11329. [Google Scholar] [CrossRef]
- Li, Y.; Huang, J.; Lin, W.; Yuan, Z.; Feng, S.; Xie, Y.; Ma, W. In vitro anticancer activity of a nonpolar fraction from Gynostemma pentaphyllum (Thunb.) Makino. Evid-Based Compl. Alt. 2016, 2016, 1–11. [Google Scholar] [CrossRef]
- Wu, P.K.; Tai, W.C.; Choi, R.C.; Tsim, K.W.; Zhou, H.; Liu, X.; Jiang, Z.-H.; Hsiao, W.W. Chemical and DNA authentication of taste variants of Gynostemma pentaphyllum herbal tea. Food Chem. 2011, 128, 70–80. [Google Scholar] [CrossRef]
- Yan, H.; Wang, X.; Wang, Y.; Wang, P.; Xiao, Y. Antiproliferation and anti-migration induced by gypenosides in human colon cancer SW620 and esophageal cancer Eca-109 cells. Hum. Exp. Toxicol. 2013, 33, 522–533. [Google Scholar] [CrossRef]
- Chew, Y.L.; Wong, H.C. Gypenosides, the cancer buster from Gynostemma pentaphyllum (Thunb.) Makino and the apoptotic pathways: A review. Orient. Pharm. Exp. Med. 2016, 16, 153–154. [Google Scholar] [CrossRef]
- Wu, Q.; Jang, M.; Piao, X.-L. Determination by UPLC-MS of four dammarane-type saponins from heat-processed Gynostemma pentaphyllum. Biosci. Biotech. Biochem. 2014, 78, 311–316. [Google Scholar] [CrossRef]
- Piao, X.-L.; Xing, S.-F.; Lou, C.-X.; Chen, D.-J. Novel dammarane saponins from Gynostemma pentaphyllum and their cytotoxic activities against HepG2 cells. Bioorg. Med. Chem. Lett. 2014, 24, 4831–4833. [Google Scholar] [CrossRef]
- Ky, P.T.; Huong, P.T.; My, T.K.; Anh, P.T.; Kiem, P.V.; Minh, C.V.; Cuong, N.X.; Thao, N.P.; Nhiem, N.X.; Hyun, J.-H.; et al. Dammarane-type saponins from Gynostemma pentaphyllum. Phytochemistry 2010, 71, 994–1001. [Google Scholar] [CrossRef]
- Shi, L.; Lu, F.; Zhao, H.; Zhao, Y.-Q. Two new triterpene saponins from Gynostemma pentaphyllum. J. Asian. Nat. Prod. Res. 2012, 14, 856–861. [Google Scholar] [CrossRef]
- Cui, J.-F.; Eneroth, P.; Bruhn, J. Gynostemma pentaphyllum: Identification of major sapogenins and differentiation from Panax species. Eur. J. Pharm. Sci. 1999, 8, 187–191. [Google Scholar] [CrossRef]
- Zhang, Y.-G.; Zhang, H.-G.; Zhang, G.-Y.; Fan, J.-S.; Li, X.-H.; Liu, Y.-H.; Li, S.-H.; Lian, X.-M.; Tang, Z. Panax notoginseng saponins attenuate atherosclerosis in rats by regulating the blood lipid profile and an anti-inflammatory action. Clin. Exp. Pharmacol. P. 2008, 35, 1238–1244. [Google Scholar] [CrossRef]
- Kuo, Y.-H.; Huang, H.-C.; Kuo, L.-M.Y.; Hsu, Y.-W.; Lee, K.-H.; Chang, F.-R.; Wu, Y.-C. New Dammarane-Type Saponins from the Galls of Sapindus mukorossi. J. Agr. Food Chem. 2005, 53, 4722–4727. [Google Scholar] [CrossRef]
- Chen, D.-J.; Liu, H.-M.; Xing, S.-F.; Piao, X.-L. Cytotoxic activity of gypenosides and gynogenin against non-small cell lung carcinoma A549 cells. Bioorg. Med. Chem. Lett. 2014, 24, 186–191. [Google Scholar] [CrossRef]
- Huang, H.-C.; Wu, M.-D.; Tsai, W.-J.; Liao, S.-C.; Liaw, C.-C.; Hsu, L.-C.; Wu, Y.-C.; Kuo, Y.-H. Triterpenoid saponins from the fruits and galls of Sapindus mukorossi. Phytochemistry 2008, 69, 1609–1616. [Google Scholar] [CrossRef]
- Hung, T.M.; Thu, C.V.; Cuong, T.D.; Hung, N.P.; Kwack, S.J.; Huh, J.-I.; Min, B.S.; Choi, J.S.; Lee, H.K.; Bae, K. Dammarane-type glycosides from Gynostemma pentaphyllumand their effects on IL-4-induced eotaxin expression in human bronchial epithelial cells. J. Nat. Prod. 2010, 73, 192–196. [Google Scholar] [CrossRef]
- Takemoto, T.; Arihara, S.; Yoshikawa, K. Studies on the constituents of cucurbitaceae plants. XIV. Yakugaku Zasshi 1986, 106, 664–670. [Google Scholar] [CrossRef]
- Yoshikawa, K.; Takemoto, T.; Arihara, S. Studies on the constituents of cucurbitaceae plants. XVI. on the saponin constituents of Gynostemma pentaphyllum MAKINO. (11). Yakugaku Zasshi 1987, 107, 262–267. [Google Scholar] [CrossRef]
- Liu, X.; Ye, W.C.; Hsiao, H.W.W.; Che, C.T.; Zhao, S.X. Studies on chemical constituents of Gynostemma pentaphyllum. J. China Pharm. Unive. 2003, 34, 21–23. [Google Scholar]
- Takemoto, T.; Arihara, S.; Yoshikawa, K.; Kawasaki, J.; Nakajima, T.; Okuhira, M. Studies on the constituents of cucurbitaceae plants. XI. on the saponin constituents of Gynostemma pentaphyllum MAKINO (7). Yakugaku Zasshi 1984, 104, 1043–1049. [Google Scholar] [CrossRef]
- Lin, M.-C.; Wang, K.-C.; Lee, S.-S. Transformation of ginsenosides Rg1 and Rb1, and crude sanchi saponins by human intestinal microflora. J. Chin. Chem. Soc. 2001, 48, 113–120. [Google Scholar] [CrossRef]
- Xiang, W.-J.; Guo, C.-Y.; Ma, L.; Hu, L.-H. Dammarane-type glycosides and long chain sesquiterpene glycosides from Gynostemma yixingense. Fitoterapia 2010, 81, 248–252. [Google Scholar] [CrossRef]
- Tomás-Lorente, F.; Garcia-Grau, M.M.; Nieto, J.L.; Tomás-Barberán, F.A. Flavonoids from Cistus ladanifer bee pollen. Phytochemistry 1992, 31, 2027–2029. [Google Scholar] [CrossRef]
- Li, S.-S.; Wu, J.; Chen, L.-G.; Du, H.; Xu, Y.-J.; Wang, L.-J.; Zhang, H.-J.; Zheng, X.-C.; Wang, L.-S. Biogenesis of C-glycosyl flavones and profiling of flavonoid glycosides in lotus (Nelumbo nucifera). PLoS ONE 2014, 9. [Google Scholar] [CrossRef]
- Leite, J.P.V.; Rastrelli, L.; Romussi, G.; Oliveira, A.B.; Vilegas, J.H.Y.; Vilegas, W.; Pizza, C. Isolation and HPLC quantitative analysis of flavonoid glycosides from Brazilian beverages (Maytenus ilicifolia and M.aquifolium). J. Agr. Food Chem. 2001, 49, 3796–3801. [Google Scholar] [CrossRef]
- Sekine, T.; Arai, Y.; Ikegami, F.; Fujii, Y.; Shindo, S.; Yanagisawa, T.; Ishida, Y.; Okonogi, S.; Murakoshi, I. Isolation of camelliaside C from “Tea Seed Cake” and inhibitory effects of its derivatives on arachidonate 5-lipoxygenase. Chem. Pharm. Bull. 1993, 41, 1185–1187. [Google Scholar] [CrossRef]
- Kwon, D.-J.; Bae, Y.-S. Flavonols from the stem bark of Acer komarovii. Chem. Nat. Compd. 2013, 49, 131–132. [Google Scholar] [CrossRef]
- Zhang, H.; Li, X.; Wu, K.; Wang, M.; Liu, P.; Wang, X.; Deng, R. Antioxidant activities and chemical constituents of flavonoids from the flower of Paeonia ostii. Molecules 2016, 22, 5. [Google Scholar] [CrossRef]
- Zhang, J.; Yin, Z.Q.; Liang, J.Y. Flavonoids from Trachelospermum jasminoides. Chem. Nat. Compd. 2013, 49, 507–508. [Google Scholar] [CrossRef]
- Wang, Y.-S.; Liao, Z.; Zhu, H.-K.; Feng, X.-F.; Jiang, K.-M.; Huang, R.; Zhu, N.; Yang, J.-H. Megastigmane O-glucopyranosides from Litsea glutinosa. Chem. Nat. Compd. 2012, 48, 346–349. [Google Scholar] [CrossRef]
- Khan, S.H.; Mosihuzzaman, M.; Nahar, N.; Rashid, M.A.; Rokeya, B.; Ali, L.; Khan, A.K.A. Three megastigmane glycosides from the leaves of Pterospermum semisagittatum. Pharm. Biol. 2003, 41, 512–515. [Google Scholar] [CrossRef]
- Zhang, L.-J.; Chiou, C.-T.; Cheng, J.-J.; Huang, H.-C.; Kuo, L.-M.Y.; Liao, C.-C.; Bastow, K.F.; Lee, K.-H.; Kuo, Y.-H. Cytotoxic polyisoprenyl benzophenonoids from Garcinia subelliptica. J. Nat. Prod. 2010, 73, 557–562. [Google Scholar] [CrossRef]
Sample Availability: Samples of the compounds 1–6 are available from the authors. |
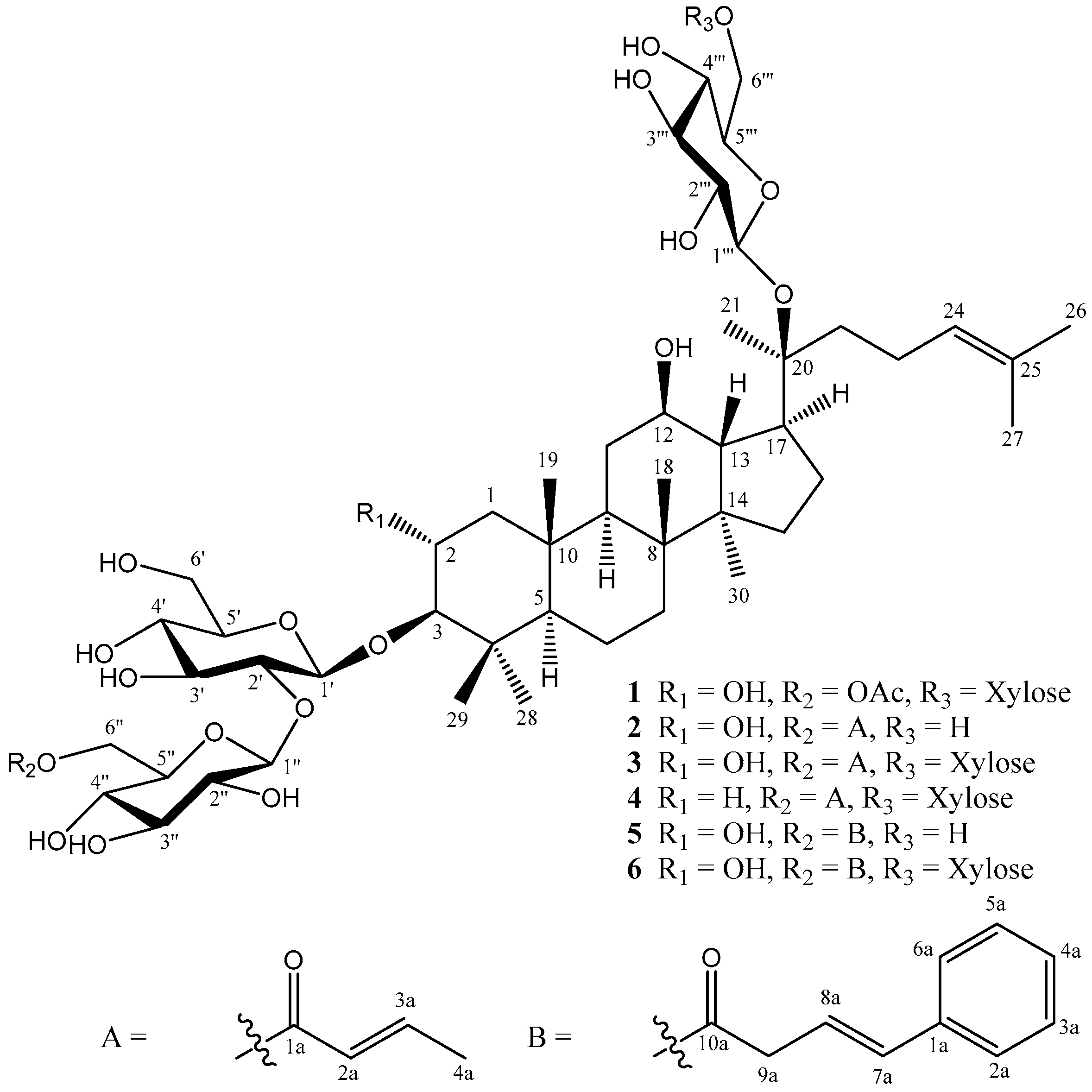
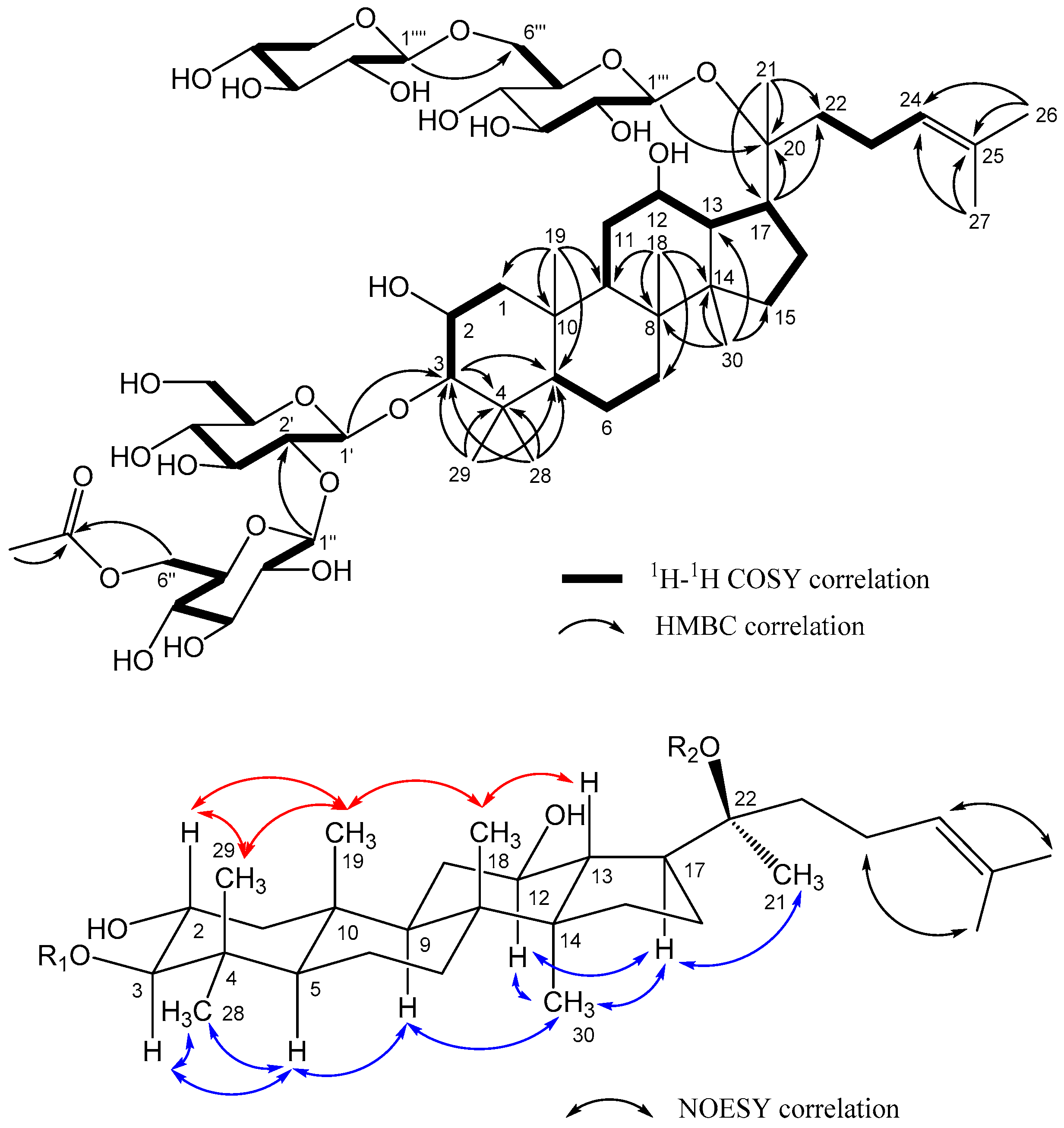
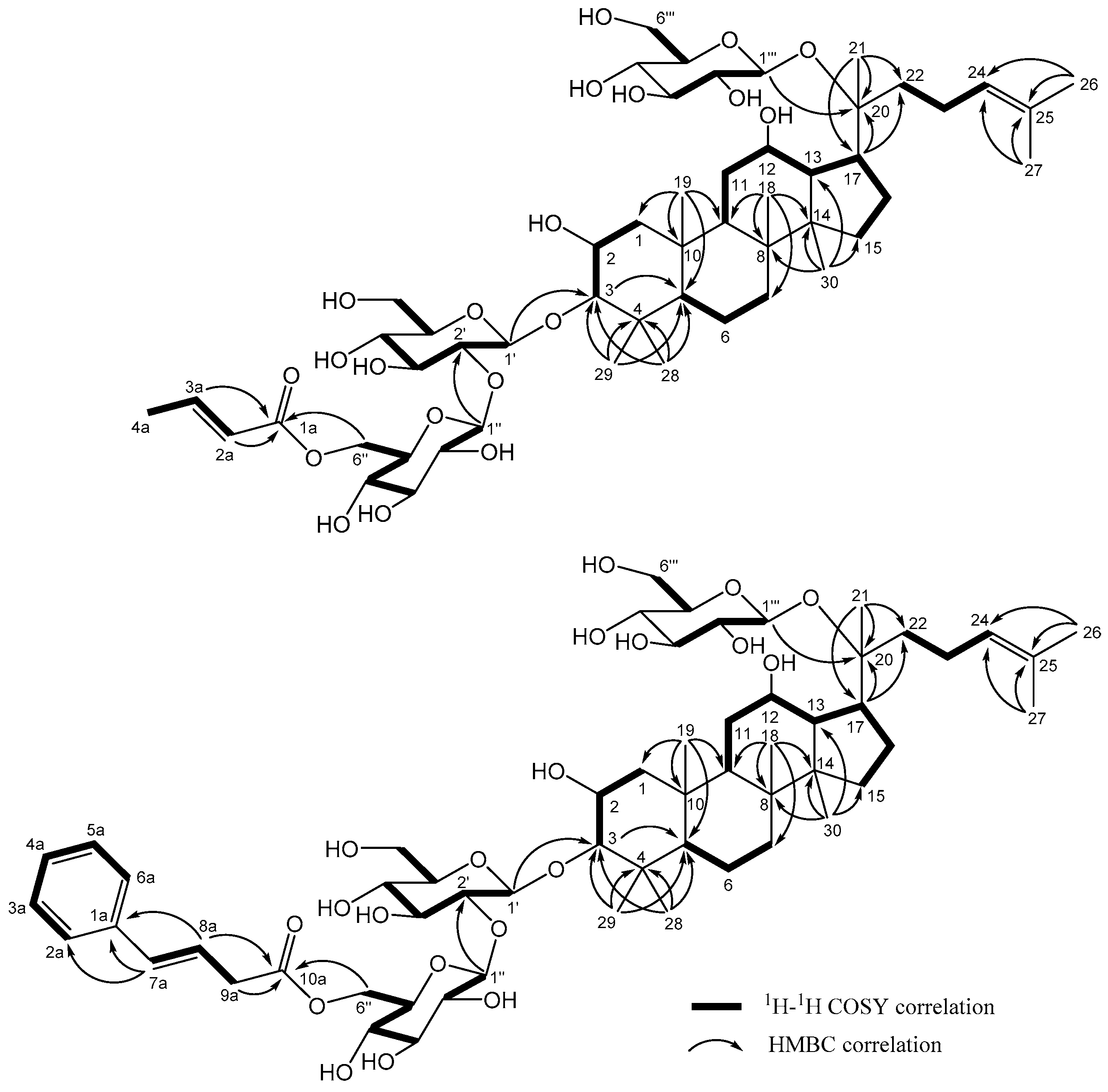
| No. | 1 | 2 | 3 | 4 | 5 | 6 | ||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| δH (mult, J in Hz) | δC | δH (mult, J in Hz) | δC | δH (mult, J in Hz) | δC | δH (mult, J in Hz) | δC | δH (mult, J in Hz) | δC | δH (mult, J in Hz) | δC | |
| 1 | 2.09 (dd, 5.0, 13.0) | 47.8 | 2.07 (m) | 47.8 | 2.07 (dd, 4.8, 12.6) | 47.8 | 1.70 (m) | 40.2 | 2.03 (dd, 4.2, 12.6) | 47.7 | 2.03 (dd, 4.2, 12.6) | 47.7 |
| 0.87 (m) | 0.91 (t, 8.0) | 0.88 (m) | 0.98 (m) | 0.87 (m) | 0.87 (m) | |||||||
| 2 | 3.72 (m) | 68.2 | 3.70 (m) | 68.2 | 3.69 (m) | 68.2 | 1.68 (m); 1.95 (m) | 27.3 | 3.70 (m) | 68.2 | 3.70 (m) | 68.2 |
| 3 | 2.95 (d, 9.0) | 96.5 | 2.93 (d, 9.5) | 96.5 | 2.93(d, 9.6) | 96.5 | 3.11 (dd, 3.0, 9.0) | 91.0 | 2.92 (d, 9.6) | 96.6 | 2.92 (d, 9.6) | 96.6 |
| 4 | - | 41.8 | - | 41.8 | 41.8 | 40.5 | - | 41.8 | - | 41.8 | ||
| 5 | 0.84 (d, 11.5) | 57.2 | 0.84 (d, 10.5) | 57.2 | 0.82 (m) | 57.2 | 0.74 (d, 11.4) | 57.6 | 0.79 (d, 11.4) | 57.2 | 0.79 (d, 11.4) | 57.2 |
| 6 | 1.49 (m); 1.56 (m) | 19.3 | 1.46 (m); 1.54 (m) | 19.4 | 1.46 (m); 1.53 (m) | 19.4 | 1.43 (m); 1.53 (m) | 19.3 | 1.37 (m); 1.50 (m) | 19.4 | 1.37 (m); 1.50 (m) | 19.4 |
| 7 | 1.57 (m); 1.29, (m) | 35.7 | 1.55 (m); 1.30 (m) | 35.7 | 1.57 (m); 1.28 (m) | 35.7 | 1.55 (m); 1.26 (m) | 35.9 | 1.47 (m); 1.19 (m) | 35.6 | 1.47 (m); 1.19 (m) | 35.6 |
| 8 | - | 41.0 | - | 41.0 | - | 41.0 | - | 41.0 | 40.9 | - | 40.9 | |
| 9 | 1.48 (m) | 51.0 | 1.50 (m) | 51.0 | 1.47 (m) | 51.0 | 1.42 (dd, 3.0, 13.2) | 51.1 | 1.45 (m) | 50.9 | 1.45 (m) | 51.0 |
| 10 | - | 38.8 | - | 38.8 | - | 38.8 | 37.9 | - | 38.8 | - | 38.7 | |
| 11 | 1.82 (m);1.28 (m) | 31.0 | 1.86 (m); 1.28 (m) | 31.1 | 1.82 (m); 1.28 (m) | 31.0 | 1.78 (m); 1.22 (m) | 30.8 | 1.81 (m); 1.22 (m) | 31.1 | 1.81 (m); 1.22, (m) | 30.9 |
| 12 | 3.74 (m) | 71.5 | 3.67 (m) | 71.8 | 3.73 (m) | 71.5 | 3.71 (m) | 71.7 | 3.66 (m) | 71.8 | 3.66 (m) | 71.5 |
| 13 | 1.73 (d, 11.5) | 49.7 | 1.74 (d, 10.5) | 49.8 | 1.73 (d, 10.8) | 49.8 | 1.72 (t, 10.8) | 49.7 | 1.70 (t, 10.8) | 49.7 | 1.70 (t, 10.8) | 49.7 |
| 14 | - | 52.4 | 52.5 | - | 52.4 | - | 52.4 | - | 52.5 | - | 52.4 | |
| 15 | 1.57 (m); 1.03 (m) | 31.5 | 1.58 (m); 1.06 (m) | 31.6 | 1.58 (m); 1.04 (m) | 31.5 | 1.57 (m); 1.03 (m) | 31.5 | 1.55 (m); 1.03 (m) | 31.6 | 1.55 (m); 1.03 (m) | 31.5 |
| 16 | 1.89 (m); 1.33 (m) | 27.3 | 1.92 (m); 1.38 (m) | 27.2 | 1.89 (m); 1.33 (m) | 27.3 | 1.95 (m), 1.32 (m) | 27.3 | 1.91 (m); 1.38 (m) | 27.2 | 1.91 (m); 1.38 (m) | 27.2 |
| 17 | 2.28 (m) | 52.9 | 2.27 (m) | 53.1 | 2.28 (m) | 52.9 | 2.28 (m) | 52.9 | 2.28 (m) | 53.1 | 2.28 (m) | 52.9 |
| 18 | 1.00 (s) | 16.3 | 1.00 (s) | 16.2 | 0.99 (s) | 16.3 | 0.99 (s) | 16.3 | 0.89 (s) | 16.2 | 0.89 (s) | 16.2 |
| 19 | 0.97 (s) | 17.8 | 0.94 (s) | 17.8 | 0.93 (s) | 17.8 | 0.88 (s) | 16.8 | 0.84 (s) | 17.9 | 0.84 (s) | 17.9 |
| 20 | - | 84.9 | - | 84.9 | - | 84.9 | - | 85.0 | - | 84.9 | - | 84.9 |
| 21 | 1.36 (s) | 22.4 | 1.34 (s) | 22.8 | 1.35 (s) | 22.4 | 1.35 (s) | 22.4 | 1.33 (s) | 22.9 | 1.33 (s) | 22.4 |
| 22 | 1.79 (m); 1.53(m) | 36.8 | 1.80 (m); 1.60 (m) | 36.7 | 1.79 (m); 1.52 (m) | 36.7 | 1.80 (m); 1.52 (m) | 36.7 | 1.80 (m); 1.61 (m) | 36.7 | 1.80 (m); 1.61 (m) | 36.8 |
| 23 | 2.02 (m); 2.15 (m) | 23.8 | 2.00 (m) | 24.2 | 2.05 (m); 2.16(m) | 23.8 | 2.05 (m); 2.14 (m) | 23.8 | 2.06 (m) | 24.3 | 2.06 (m) | 24.1 |
| 24 | 5.13 (bt, 7.0) | 126.1 | 5.10 (brt, 7.0) | 125.9 | 5.11 (m) | 126.1 | 5.12 (m) | 126.1 | 5.11 (bt, 7.2) | 125.9 | 5.11 (bt, 7.2) | 126.1 |
| 25 | - | 132.2 | - | 132.3 | 132.2 | - | 132.2 | - | 132.3 | - | 132.2 | |
| 26 | 1.68 (s) | 25.9 | 1.68 (s) | 25.9 | 1.68 (s) | 25.9 | 1.68 (s) | 25.9 | 1.69 (s) | 25.9 | 1.69 (s) | 25.9 |
| 27 | 1.62 (s) | 18.0 | 1.62 (s) | 17.9 | 1.62 (s) | 18.0 | 1.62 (s) | 18.0 | 1.62 (s) | 18.0 | 1.62 (s) | 18.0 |
| 28 | 0.86 (s) | 28.6 | 0.82 (s) | 28.5 | 0.82 (s) | 28.5 | 0.77 (s) | 28.4 | 0.83 (s) | 28.6 | 0.83 (s) | 28.6 |
| 29 | 1.10 (s) | 17.7 | 1.08 (s) | 17.8 | 1.07 (s) | 17.79 | 1.02 (s) | 16.7 | 1.07 (s) | 17.9 | 1.07 (s) | 17.9 |
| 30 | 0.92 (s) | 17.4 | 0.92 (s) | 17.2 | 0.91 (s) | 17.3 | 0.91 (s) | 17.4 | 0.89 (s) | 17.1 | 0.89 (s) | 17.3 |
| No. | 1 | 2 | 3 | 4 | 5 | 6 | ||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| δH | δC | δH | δC | δH | δC | δH | δC | δH | δC | δH | δC | |
| 1′ | 4.42 (d, 7.5) | 104.7 | 4.42 (d, 7.5) | 104.6 | 4.41 (d, 7.8) | 104.6 | 4.40 (d, 7.8) | 105.2 | 4.42 (d, 7.8) | 104.6 | 4.41 (d, 7.8) | 104.6 |
| 2′ | 3.56 (m) | 82.4 | 3.55 (m) | 82.7 | 3.56 (m) | 82.7 | 3.45 (m) | 83.4 | 3.54 (dd, 7.8, 9.0) | 83.2 | 3.55 (dd, 7.8, 9.0) | 83.2 |
| 3′ | 3.58 (m) | 78.7 | 3.58 (m) | 78.7 | 3.58 (m) | 78.7 | 3.54 (t, 9.0) | 78.6 | 3.59 (m) | 78.8 | 3.58 (m) | 78.8 |
| 4′ | 3.37 (m) | 70.9 | 3.36 (m) | 70.9 | 3.36 (m) | 70.9 | 3.32 (m) | 71.2 | 3.36 (m) | 70.9 | 3.36 (m) | 70.9 |
| 5′ | 3.34 (m) | 78.0 | 3.19 (m) | 77.9 | 3.34 (m) | 78.0 | 3.23 (m) | 77.5 | 3.20 (m) | 77.9 | 3.35 (m) | 78.0 |
| 6′ | 3.85 (dd, 5.0, 12.0) | 62.3 | 3.85 (dd, 5.0, 12.0) | 62.3 | 3.86 (dd, 1.8, 12.0) | 62.3 | 3.83 (dd, 1.8, 12.0) | 62.7 | 3.86 (dd, 1.8, 12.0) | 62.3 | 3.86 (dd, 1.8, 12.0) | 62.3 |
| 3.66 (dd, 5.0, 12.0) | 3.66 (dd, 5.0, 12.0) | 3.66 (dd, 5.4, 12.0) | 3.65 (dd, 5.4, 12.0) | 3.66 (m) | 3.65 (dd, 5.4, 12.0) | |||||||
| 1″ | 4.71 (d, 7.5) | 105.0 | 4.71 (d, 7.5) | 105.2 | 4.70 (d, 7.8) | 105.2 | 4.62 (d, 7.8) | 105.5 | 4.71 (d, 7.8) | 105.4 | 4.71 (d, 7.8) | 105.4 |
| 2″ | 3.23 (m) | 76.1 | 3.24 (dd, 7.5, 9.0) | 76.1 | 3.24 (t, 8.4) | 76.1 | 3.23 (t, 8.4) | 76.4 | 3.24 (t, 8.4) | 76.1 | 3.23 (t, 8.4) | 76.1 |
| 3″ | 3.35 (m) | 77.9 | 3.35 (m) | 77.9 | 3.35 (m) | 77.9 | 3.35 (m) | 77.7 | 3.35 (m) | 78.0 | 3.34 (m) | 78.1 |
| 4″ | 3.29 (m) | 71.4 | 3.32 (m) | 71.4 | 3.32 (m) | 71.4 | 3.32 (m) | 71.3 | 3.33 (m) | 71.4 | 3.33 (m) | 71.3 |
| 5″ | 3.43 (m) | 75.3 | 3.45 (m) | 75.3 | 3.45 (m) | 75.3 | 3.44 (m) | 75.5 | 3.46 (dd, 1.8, 5.4) | 75.2 | 3.46 (m) | 75.3 |
| 6″ | 4.15 (dd, 5.0, 12.0) | 65.0 | 4.18 (dd, 5.0, 12.0) | 64.7 | 4.18 (dd, 4.8, 12.0) | 64.7 | 4.18 (dd, 4.8, 12.0) | 64.7 | 4.20 (dd, 5.4, 12.0) | 65.2 | 4.20 (dd, 4.8, 12.0) | 65.1 |
| 4.31 (dd, 5.0, 12.0) | 4.37 (dd, 5.0, 12.0) | 4.36 (dd, 1.8, 12.0) | 4.37 (dd, 1.8, 12.0) | 4.38 (dd, 1.8, 12.0) | 4.38 (dd, 1.8, 12.0) | |||||||
| 1a | 172.8 | 168.1 | 168.1 | 168.2 | 138.4 | 138.4 | ||||||
| 2a | 2.04 (s) | 20.9 | 5.88 (dq, 1.5, 15.5) | 123.6 | 5.88 (dq, 1.5, 15.6) | 123.5 | 5.89 (dq, 1.8, 15.6) | 123.5 | 7.38 (d, 7.2) | 127.4 | 7.38 (d, 7.2) | 127.3 |
| 3a | 7.00 (dq, 7.0, 15.5) | 146.6 | 7.00 (dq, 7.2, 15.5) | 146.6 | 7.00 (dq, 7.2, 15.6) | 146.6 | 7.29 (t, 7.2) | 129.6 | 7.29 (t, 7.2) | 129.7 | ||
| 4a | 1.89 (dd, 1.5, 7.0) | 18.1 | 1.89 (dd, 1.8, 7.2) | 18.1 | 1.89 (dd, 1.8, 7.2) | 18.1 | 7.20 (t, 7.2) | 128.6 | 7.21 (t, 7.2) | 128.6 | ||
| 5a | 7.29 (t, 7.2) | 129.6 | 7.29 (t, 7.2) | 129.7 | ||||||||
| 6a | 7.38 (d, 7.2) | 127.4 | 7.38 (d, 7.2) | 127.3 | ||||||||
| 7a | 6.52 (d, 16.2) | 134.6 | 6.52 (d, 16.2) | 134.6 | ||||||||
| 8a | 6.33 (dd, 7.2, 16.2) | 122.7 | 6.33 (dd, 7.2, 16.2) | 122.7 | ||||||||
| 9a | 3.30 (m) | 38.7 | 3.30 (m) | 38.7 | ||||||||
| 10a | 173.4 | 173.4 | ||||||||||
| 1’’’ | 4.56 (d, 8.0) | 98.1 | 4.60 (d, 8.0) | 98.3 | 4.56 (d, 7.8) | 98.1 | 4.56 (d, 8.4) | 98.1 | 4.59 (d, 7.8) | 98.3 | 4.56 (d, 7.8) | 98.1 |
| 2’’’ | 3.12 (m) | 75.3 | 3.08 (m) | 75.4 | 3.11 (m) | 75.3 | 3.11 (m) | 75.3 | 3.07 (m) | 75.4 | 3.12 (m) | 75.3 |
| 3’’’ | 3.33 (m) | 78.6 | 3.34 (m) | 78.3 | 3.32 (m) | 78.6 | 3.32 (m) | 78.6 | 3.35 (m) | 78.2 | 3.33 (m) | 78.6 |
| 4’’’ | 3.31 (m) | 71.4 | 3.32 (m) | 71.2 | 3.32 (m) | 71.4 | 3.32 (m) | 71.4 | 3.33 (m) | 71.2 | 3.32 (m) | 71.4 |
| 5’’’ | 3.32 (m) | 76.7 | 3.18 (m) | 78.0 | 3.39 (m) | 76.7 | 3.39 (m) | 76.7 | 3.20 (m) | 77.9 | 3.38 (m) | 76.7 |
| 6’’’ | 3.73 (dd, 5.5, 11.5) | 70.1 | 3.64 (dd, 5.5, 11.5) | 62.5 | 3.72 (dd, 5.4, 11.4) | 70.1 | 3.73 (m) | 70.1 | 3.64 (m) | 62.5 | 3.73 (dd, 5.4, 11.4) | 70.1 |
| 4.00 (dd, 2.0, 11.5) | 3.77 (dd, 2.0, 11.5) | 4.00 (dd, 2.4, 11.4) | 4.00 (dd, 1.8, 11.4) | 3.77 (dd, 2.4, 12.0) | 4.00 (dd, 1.8, 11.4) | |||||||
| 1’’’’ | 4.30 (d, 7.5) | 105.5 | 4.29 (d, 7.2) | 105.6 | 4.29 (d, 7.8) | 105.6 | 4.29 (d, 7.8) | 105.6 | ||||
| 2’’’’ | 3.20 (m) | 74.8 | 3.20 (m) | 74.8 | 3.19 (d, 7.2) | 74.8 | 3.20 (d, 7.2) | 74.8 | ||||
| 3’’’’ | 3.30 (m) | 77.5 | 3.30 (m) | 77.5 | 3.29 (m) | 77.5 | 3.30 (m) | 77.5 | ||||
| 4’’’’ | 3.47 (m) | 71.2 | 3.46 (m) | 71.2 | 3.47 (m) | 71.2 | 3.46 (m) | 71.2 | ||||
| 5’’’’ | 3.18 (m) | 66.8 | 3.18 (m) | 66.8 | 3.18 (m) | 66.8 | 3.18 (m) | 66.8 | ||||
| 4.00 (dd, 2.0, 11.5) | 3.84 (dd, 5.4, 11.4) | 3.84 (dd, 5.4, 11.4) | 3.84 (dd, 5.4, 11.4) | |||||||||
| Cmpd. | A549 Cell Line | HepG2 Cell Line | Number of Sugars | ||
|---|---|---|---|---|---|
| Inhibition (%) a | EC50 (μM) | Inhibition (%) | EC50 (μM) | ||
| 1 | 16.4 ± 3.97 | (-) b | 38.9 ± 3.82 | (-) | 4 |
| 2 | 84.1 ± 7.97 | 59.4 ± 2.51 | 83.0 ± 3.91 | 60.4 ± 0.63 | 3 |
| 3 | 23.3 ± 6.20 | (-) | 44.0 ± 2.28 | (-) | 4 |
| 4 | 17.0 ± 2.00 | (-) | 46.4 ± 2.60 | (-) | 4 |
| 5 | 42.7 ± 1.41 | (-) | 71.1 ± 0.60 | 29.3 ± 0.26 | 3 |
| 6 | 29.7 ± 5.34 | (-) | 55.5 ± 6.45 | 54.2 ± 2.07 | 4 |
| 7 | 37.1 ± 0.78 | (-) | 53.8 ± 3.43 | 89.1 ± 3.75 | 3 |
| 8 | 6.5 ± 5.16 | (-) | 30.7 ± 5.32 | (-) | 3 |
| 9 | 14.5 ± 6.04 | (-) | 31.1 ± 7.76 | (-) | 4 |
| 10 | 20.9 ± 2.62 | (-) | 44.1 ± 1.96 | (-) | 3 |
| 11 | 94.4 ± 0.28 | 70.1 ± 2.34 | 93.1 ± 0.99 | 76.2 ± 2.10 | 2 |
| 12 | 27.8 ± 11.35 | (-) | 60.4 ± 6.34 | 100.7 ± 1.36 | 2 |
| 13 | 65.1 ± 7.29 | 87.3 ± 3.39 | 73.3 ± 1.81 | 68.4 ± 0.57 | 2 |
| 14 | 17.9 ± 2.72 | (-) | 24.5 ± 2.79 | (-) | 4 |
| 15 | 43.2 ± 3.67 | 73.8 ± 2.86 | 56.5 ± 1.55 | 75.4 ± 1.30 | 3 |
| 16 | 25.2 ± 1.35 | (-) | 37.3 ± 0.53 | (-) | 4 |
| 17 | 13.1 ± 2.88 | (-) | 11.5 ± 1.17 | (-) | - |
| 18 | 16.1 ± 5.32 | (-) | 9.4 ± 3.02 | (-) | - |
| 19 | 19.3 ± 1.40 | (-) | 32.5 ± 3.83 | (-) | - |
| 20 | 13.0 ± 3.40 | (-) | 18.0 ± 2.99 | (-) | - |
| 21 | 15.5 ± 2.73 | (-) | 29.2 ± 2.45 | (-) | - |
| 22 | 22.9 ± 12.78 | (-) | 26.6 ± 6.30 | (-) | - |
| 23 | 21.7 ± 9.46 | (-) | 31.2 ± 3.45 | (-) | - |
| 24 | 10.9 ± 6.06 | (-) | 25.2 ± 3.65 | (-) | - |
| 25 | 5.6 ± 3.12 | (-) | 6.5 ± 3.17 | (-) | - |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chen, P.-Y.; Chang, C.-C.; Huang, H.-C.; Zhang, L.-J.; Liaw, C.-C.; Lin, Y.-C.; Nguyen, N.-L.; Vo, T.-H.; Cheng, Y.-Y.; Morris-Natschke, S.L.; et al. New Dammarane-Type Saponins from Gynostemma pentaphyllum. Molecules 2019, 24, 1375. https://doi.org/10.3390/molecules24071375
Chen P-Y, Chang C-C, Huang H-C, Zhang L-J, Liaw C-C, Lin Y-C, Nguyen N-L, Vo T-H, Cheng Y-Y, Morris-Natschke SL, et al. New Dammarane-Type Saponins from Gynostemma pentaphyllum. Molecules. 2019; 24(7):1375. https://doi.org/10.3390/molecules24071375
Chicago/Turabian StyleChen, Po-Yen, Chih-Chao Chang, Hui-Chi Huang, Li-Jie Zhang, Chia-Ching Liaw, Yu-Chi Lin, Nham-Linh Nguyen, Thanh-Hoa Vo, Yung-Yi Cheng, Susan L. Morris-Natschke, and et al. 2019. "New Dammarane-Type Saponins from Gynostemma pentaphyllum" Molecules 24, no. 7: 1375. https://doi.org/10.3390/molecules24071375
APA StyleChen, P.-Y., Chang, C.-C., Huang, H.-C., Zhang, L.-J., Liaw, C.-C., Lin, Y.-C., Nguyen, N.-L., Vo, T.-H., Cheng, Y.-Y., Morris-Natschke, S. L., Lee, K.-H., & Kuo, Y.-H. (2019). New Dammarane-Type Saponins from Gynostemma pentaphyllum. Molecules, 24(7), 1375. https://doi.org/10.3390/molecules24071375








