Parasitic Protozoa: Unusual Roles for G-Quadruplexes in Early-Diverging Eukaryotes
Abstract
1. Introduction
2. To Begin at the Beginning: A Free-Living Protozoan
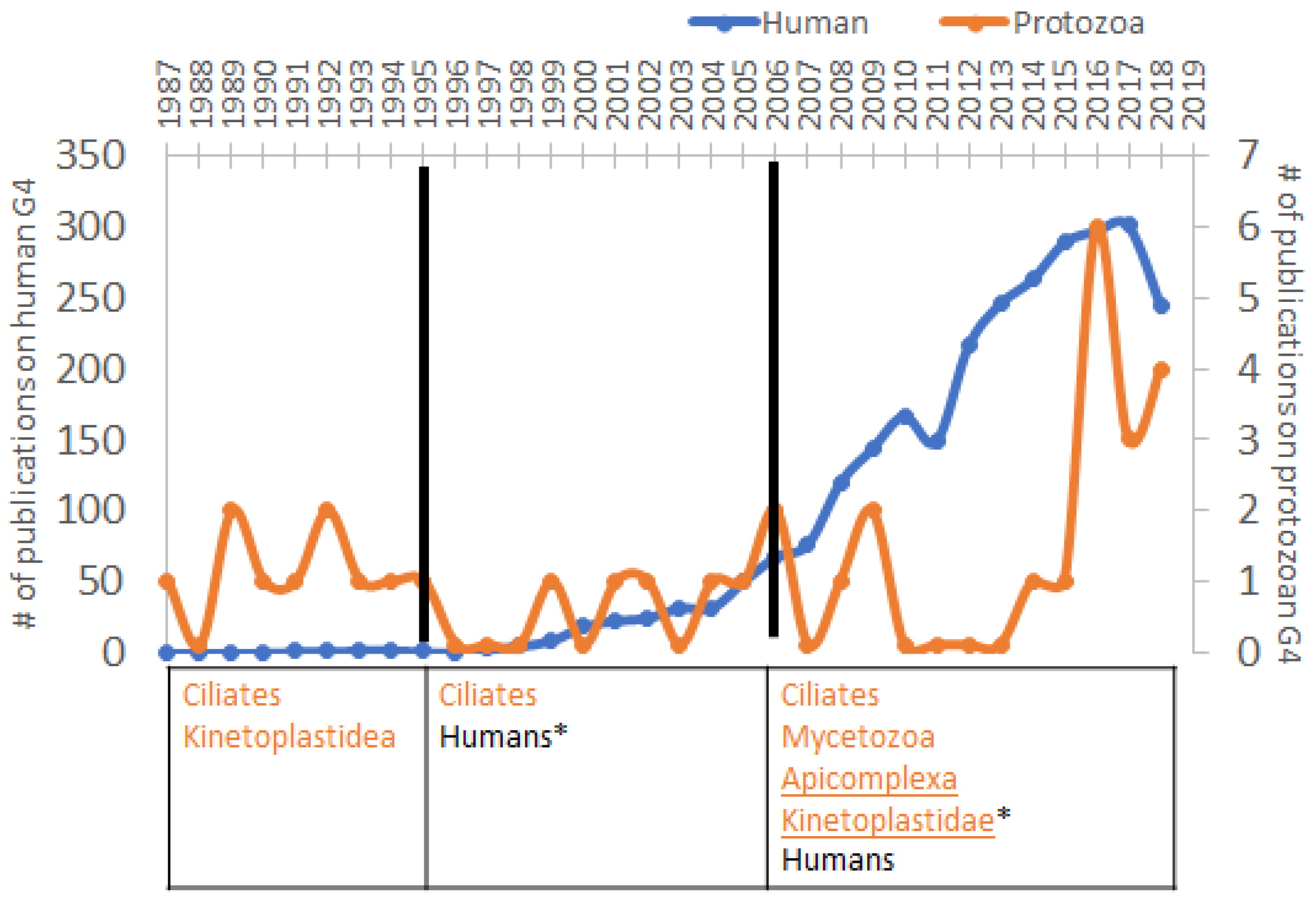
3. Increased Interest in G4s in Humans Correlates with the Diminution of Attention to Protozoa
4. Biological Significance of G4s in Environmental and Pathogenic Protozoans
4.1. G4 Structures and Functions in Protozoa
4.2. G4 Motifs in an AT-Rich Genome: The Example of Plasmodium Falciparum
4.3. G4 Motifs in Transcription Control and RNA Editing in Trypanosomatids
4.4. Targeting the G4 Motifs In Pathogenic Protozoan: A New Strategy in the Design of Anti-Parasitic Drugs
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Watson, J.D.; Crick, F.H.C. Molecular structure of nucleic acids: A structure for deoxyribose nucleic acid. Nature 1953, 171, 737–738. [Google Scholar] [CrossRef] [PubMed]
- Biffi, G.; Tannahill, D.; McCafferty, J.; Balasubramanian, S. Quantitative visualization of DNA G-quadruplex structures in human cells. Nat. Chem. 2013, 5, 182–186. [Google Scholar] [CrossRef] [PubMed]
- Zeraati, M.; Langley, D.B.; Schofield, P.; Moye, A.L.; Rouet, R.; Hughes, W.E.; Bryan, T.M.; Dinger, M.E.; Christ, D. I-motif DNA structures are formed in the nuclei of human cells. Nat. Chem. 2018, 10, 631–637. [Google Scholar] [CrossRef] [PubMed]
- Lopes, J.; Piazza, A.; Le Bermejo, R.; Kriegsman, B.; Colosio, A.; Teulade-Fichou, M.P.; Foiani, M.; Nicolas, A. G-quadruplex-induced instability during leading-strand replication. EMBO J. 2011, 30, 4033–4046. [Google Scholar] [CrossRef] [PubMed]
- Smargiasso, N.; Rosu, F.; Hsia, W.; Colson, P.; Baker, E.S.; Bowers, M.T.; De Pauw, E.; Gabelica, V. G-quadruplex DNA assemblies: Loop length, cation identity, and multimer formation. J. Am. Chem. Soc. 2008, 130, 10208–10216. [Google Scholar] [CrossRef] [PubMed]
- Oka, Y.; Thomas, C.A. The cohering telomeres of Oxytricha. Nucleic Acids Res. 1987, 15, 8877–8898. [Google Scholar] [CrossRef] [PubMed]
- Bedrat, A.; Lacroix, L.; Mergny, J.-L. Re-evaluation of G-quadruplex propensity with G4Hunter. Nucleic Acids Res. 2016, 44, 1746–1759. [Google Scholar] [CrossRef]
- Murat, P.; Balasubramanian, S. Existence and consequences of G-quadruplex structures in DNA. Curr. Opin. Genet. Dev. 2014, 25, 22–29. [Google Scholar] [CrossRef] [PubMed]
- Harris, L.M.; Merrick, C.J. G-quadruplexes in pathogens: A common route to virulence control? PLoS Pathog. 2015, 11, 1–15. [Google Scholar] [CrossRef]
- Swart, E.C.; Bracht, J.R.; Magrini, V.; Minx, P.; Chen, X.; Zhou, Y.; Khurana, J.S.; Goldman, A.D.; Nowacki, M.; Schotanus, K.; et al. The Oxytricha trifallax macronuclear genome: A complex eukaryotic genome with 16,000 tiny chromosomes. PLoS Biol. 2013, 11, e1001473. [Google Scholar] [CrossRef]
- Eisen, J.A.; Coyne, R.S.; Wu, M.; Wu, D.; Thiagarajan, M.; Wortman, J.R.; Badger, J.H.; Ren, Q.; Amedeo, P.; Jones, K.M.; et al. Macronuclear genome sequence of the ciliate Tetrahymena thermophila, a model eukaryote. PLoS Biol. 2006, 4, 1620–1642. [Google Scholar] [CrossRef]
- Price, C.M.; Cech, T.R. Telomeric DNA-protein interactions of Oxytricha macronuclear DNA. Genes Dev. 1987, 1, 783–793. [Google Scholar] [CrossRef]
- Williamson, J.R.; Raghuraman, M.K.; Cech, T.R. Monovalent cation-induced structure of telomeric DNA: The G-quartet model. Cell 1989, 59, 871–880. [Google Scholar] [CrossRef]
- Sundquist, W.I.; Klug, A. Telomeric DNA dimerizes by formation of guanine tetrads between hairpin loops. Nature 1989, 342, 825–829. [Google Scholar] [CrossRef]
- Raghuraman, M.K.; Cech, T.R. Effect of monovalent cation-induced telomeric DNA structure on the binding of Oxytricha telomeric protein. Nucleic Acids Res. 1990, 18, 4543–4552. [Google Scholar] [CrossRef]
- Hardin, C.C.; Henderson, E.; Watson, T.; Prosser, J.K. Monovalent cation induced structural transitions in telomeric DNAs: G-DNA folding intermediates. Biochemistry 1991, 30, 4460–4472. [Google Scholar] [CrossRef]
- Kang, C.; Zhang, X.; Ratliff, R.; Moyzis, R.; Rich, A. Crystal structure of four-stranded Oxytricha telomeric DNA. Nature 1992, 356, 126–131. [Google Scholar] [CrossRef]
- Schultze, P.; Smith, F.W.; Feigon, J. Refined solution structure of the dimeric quadruplex formed from the Oxytricha telomeric oligonucleotide d(GGGGTTTTGGGG). Structure 1994, 2, 221–233. [Google Scholar] [CrossRef]
- Smith, F.W.; Feigon, J. Quadruplex structure of Oxytricha telomeric DNA oligonucleotides. Nature 1992, 356, 164–168. [Google Scholar] [CrossRef]
- Abu-Ghazalah, R.M.; Macgregor, R.B. Structural polymorphism of the four-repeat Oxytricha nova telomeric DNA sequences. Biophys. Chem. 2009, 141, 180–185. [Google Scholar] [CrossRef]
- Lu, M.; Guo, Q.; Kallenbach, N.R. Thermodynamics of G-tetraplex formation by telomeric DNAs. Biochemistry 1993, 32, 598–601. [Google Scholar] [CrossRef]
- Lukes, J.; Hashimi, H.; Zíková, A. Unexplained complexity of the mitochondrial genome and transcriptome in Kinetoplastid flagellates. Curr. Genet. 2005, 48, 277–299. [Google Scholar] [CrossRef]
- Sá-Carvalho, D.; Traub-Cseko, Y.M. Sequences with high propensity to form G-quartet structures in Kinetoplast DNA from Phytomonas Serpens. Mol. Biochem. Parasitol. 1995, 72, 103–109. [Google Scholar] [CrossRef]
- Schaffitzel, C.; Berger, I.; Postberg, J.; Hanes, J.; Lipps, H.J.; Pluckthun, A. In vitro generated antibodies specific for telomeric guanine-quadruplex DNA react with Stylonychia Lemnae macronuclei. Proc. Natl. Acad. Sci. USA 2001, 98, 8572–8577. [Google Scholar] [CrossRef]
- Prescott, D.M. Restructuring of DNA sequences in the germline genome of Oxytricha. Curr. Opin. Genet. Dev. 1993, 3, 726–729. [Google Scholar] [CrossRef]
- Paeschke, K.; Simonsson, T.; Postberg, J.; Rhodes, D.; Lipps, H.J. Telomere End-Binding Proteins control the formation of G-quadruplex DNA structures in vivo. Nat. Struct. Mol. Biol. 2005, 12, 847–854. [Google Scholar] [CrossRef]
- Oganesian, L.; Moon, I.K.; Bryan, T.M.; Jarstfer, M.B. Extension of G-quadruplex DNA by ciliate telomerase. EMBO J. 2006, 25, 1148–1159. [Google Scholar] [CrossRef]
- Schultze, P.; Hud, N.V.; Smith, F.W.; Feigon, J. The effect of sodium, potassium and ammonium ions on the conformation of the dimeric quadruplex formed by the Oxytricha nova telomere repeat oligonucleotide d(G(4)T(4)G(4)). Nucleic Acids Res. 1999, 27, 3018–3028. [Google Scholar] [CrossRef]
- Haider, S.; Parkinson, G.N.; Neidle, S. Crystal structure of the potassium form of an Oxytricha nova G-quadruplex. J. Mol. Biol. 2002, 320, 189–200. [Google Scholar] [CrossRef]
- Phan, A.T.; Modi, Y.S.; Patel, D.J. Two-repeat Tetrahymena telomeric d(TGGGGTTGGGGT) sequence interconverts between asymmetric dimeric G-quadruplexes in solution. J. Mol. Biol. 2004, 338, 93–102. [Google Scholar] [CrossRef]
- Gill, M.L.; Strobel, S.A.; Loria, J.P. Crystallization and characterization of the thallium form of the Oxytricha nova G-quadruplex. Nucleic Acids Res. 2006, 34, 4506–4514. [Google Scholar] [CrossRef] [PubMed]
- Horvath, M.P.; Schultz, S.C. DNA G-quartets in a 1.86 Å resolution structure of an Oxytricha nova telomeric protein-DNA complex. J. Mol. Biol. 2001, 310, 367–377. [Google Scholar] [CrossRef] [PubMed]
- Nielsen, J.T.; Arar, K.; Petersen, M. Solution structure of a locked nucleic acid modified quadruplex: Introducing the V4 folding topology. Angew. Chemie Int. Ed. 2009, 48, 3099–3103. [Google Scholar] [CrossRef] [PubMed]
- Demkovičová, E.; Bauer, Ľ.; Krafčíková, P.; Tlučková, K.; Tóthova, P.; Halaganová, A.; Valušová, E.; Víglaský, V. Telomeric G-quadruplexes: From human to Tetrahymena repeats. J. Nucleic Acids 2017, 2017. [Google Scholar] [CrossRef] [PubMed]
- Godiska, R.; Yao, M.C. A Programmed Site-specific DNA rearrangement in Tetrahymena thermophila requires flanking polypurine tracts. Cell 1990, 61, 1237–1246. [Google Scholar] [CrossRef]
- Fass, J.N.; Joshi, N.A.; Couvillion, M.T.; Bowen, J.; Gorovsky, M.A.; Hamilton, E.P.; Orias, E.; Hong, K.; Coyne, R.S.; Eisen, J.A.; et al. Genome-scale analysis of programmed DNA elimination sites in Tetrahymena thermophila. G3 (Bethesda). 2011, 1, 515–522. [Google Scholar] [CrossRef]
- Carle, C.M.; Zaher, H.S.; Chalker, D.L. A Parallel G-quadruplex-binding protein regulates the boundaries of DNA elimination events of Tetrahymena thermophila. PLoS Genet. 2016, 12, 1–22. [Google Scholar] [CrossRef]
- Huppert, J.L.; Balasubramanian, S. G-quadruplexes in promoters throughout the human genome. Nucleic Acids Res. 2007, 35, 406–413. [Google Scholar] [CrossRef]
- Scherf, A.; Hernandez-Rivas, R.; Buffet, P.; Bottius, E.; Benatar, C.; Pouvelle, B.; Gysin, J.; Lanzer, M. Antigenic variation in malaria: In situ switching, relaxed and mutually exclusive transcription of var genes during intra-erythrocytic development in Plasmodium falciparum. EMBO J. 1998, 17, 5418–5426. [Google Scholar] [CrossRef]
- Smargiasso, N.; Gabelica, V.; Damblon, C.; Rosu, F.; De Pauw, E.; Teulade-Fichou, M.P.; Rowe, J.A.; Claessens, A. Putative DNA G-quadruplex formation within the promoters of Plasmodium falciparum var genes. BMC Genom. 2009, 10, 1–12. [Google Scholar] [CrossRef]
- Claessens, A.; Harris, L.M.; Stanojcic, S.; Chappell, L.; Stanton, A.; Kuk, N.; Veneziano-Broccia, P.; Sterkers, Y.; Rayner, J.C.; Merrick, C.J. RecQ helicases in the malaria parasite Plasmodium falciparum affect genome stability, gene expression patterns and DNA replication dynamics. PLoS Genet. 2018, 14, e1007490. [Google Scholar] [CrossRef]
- Li, Z.; Yin, S.; Sun, M.; Cheng, X.; Wei, J.; Gilbert, N.; Miao, J.; Cui, L.; Huang, Z.; Dai, X.; et al. DNA helicase RecQ1 regulates mutually exclusive expression of virulence genes in Plasmodium falciparum via heterochromatin alteration. Proc. Natl. Acad. Sci. USA 2019, 116, 3177–3182. [Google Scholar] [CrossRef]
- Claessens, A.; Hamilton, W.L.; Kekre, M.; Otto, T.D.; Faizullabhoy, A.; Rayner, J.C.; Kwiatkowski, D. Generation of antigenic diversity in Plasmodium falciparum by structured rearrangement of var genes during mitosis. PLoS Genet. 2014, 10, e1004812. [Google Scholar] [CrossRef] [PubMed]
- Stanton, A.; Harris, L.M.; Graham, G.; Merrick, C.J. Recombination events among virulence genes in malaria parasites are associated with G-quadruplex-forming DNA motifs. BMC Genom. 2016, 17, 1–16. [Google Scholar] [CrossRef]
- Bhartiya, D.; Chawla, V.; Ghosh, S.; Shankar, R.; Kumar, N. Genome-wide regulatory dynamics of G-quadruplexes in human malaria parasite Plasmodium falciparum. Genomics 2016, 108, 224–231. [Google Scholar] [CrossRef]
- Guédin, A.; Lin, L.Y.; Armane, S.; Lacroix, L.; Mergny, J.-L.; Thore, S.; Yatsunyk, L.A. Quadruplexes in ‘Dicty’: Crystal structure of a four-quartet G-quadruplex formed by G-rich motif found in the Dictyostelium discoideum genome. Nucleic Acids Res. 2018, 46, 5297–5307. [Google Scholar] [CrossRef]
- Kaufer, A.; Ellis, J.; Stark, D.; Barratt, J. The evolution of Trypanosomatid taxonomy. Parasites and Vectors 2017, 10, 1–17. [Google Scholar] [CrossRef]
- Genest, P.-A.; Baugh, L.; Taipale, A.; Zhao, W.; Jan, S.; van Luenen, H.G.A.M.; Korlach, J.; Clark, T.; Luong, K.; Boitano, M.; et al. Defining the sequence requirements for the positioning of base J in DNA using SMRT sequencing. Nucleic Acids Res. 2015, 43, 2102–2115. [Google Scholar] [CrossRef]
- Biffi, G.; Di Antonio, M.; Tannahill, D.; Balasubramanian, S. Visualization and selective chemical targeting of RNA G-quadruplex structures in the cytoplasm of human cells. Nat. Chem. 2014, 6, 75–80. [Google Scholar] [CrossRef]
- Aphasizhev, R.; Aphasizheva, I. Mitochondrial RNA editing in Trypanosomes: Small RNAs in control. Biochimie 2014, 100, 125–131. [Google Scholar] [CrossRef]
- Leeder, W.M.; Voigt, C.; Brecht, M.; Göringer, H.U. The RNA chaperone activity of the Trypanosoma brucei editosome raises the dynamic of bound pre-mRNAs. Sci. Rep. 2016, 6, 1–11. [Google Scholar] [CrossRef]
- Leeder, W.-M.; Hummel, N.F.C.; Göringer, H.U. Multiple G-quartet structures in pre-edited mRNAs suggest evolutionary driving force for RNA editing in Trypanosomes. Sci. Rep. 2016, 6, 29810. [Google Scholar] [CrossRef]
- Barthwal, R.; Tariq, Z. Molecular recognition of parallel G-quadruplex [d-(TTGGGGT)]4 containing Tetrahymena telomeric DNA sequence by anticancer drug daunomycin: NMR-based structure and thermal stability. Molecules 2018, 23, 2266. [Google Scholar] [CrossRef]
- Platella, C.; Riccardi, C.; Montesarchio, D.; Roviello, G.N.; Musumeci, D. G-quadruplex-based aptamers against protein targets in therapy and diagnostics. Biochim. Biophys. Acta Gen. Subj. 2017, 1861, 1429–1447. [Google Scholar] [CrossRef]
- Musumeci, D.; Riccardi, C.; Montesarchio, D. G-Quadruplex forming oligonucleotides as anti-HIV agents. Molecules 2015, 20, 17511–17532. [Google Scholar] [CrossRef]
- De Cian, A.; Grellier, P.; Mouray, E.; Depoix, D.; Bertrand, H.; Monchaud, D.; Teulade-Fichou, M.P.; Mergny, J.L.; Alberti, P. Plasmodium telomeric sequences: Structure, stability and quadruplex targeting by small compounds. Chembiochem 2008, 9, 2730–2739. [Google Scholar] [CrossRef]
- Calvo, E.P.; Wasserman, M. G-quadruplex ligands: Potent inhibitors of telomerase activity and cell proliferation in Plasmodium falciparum. Mol. Biochem. Parasitol. 2016, 207, 33–38. [Google Scholar] [CrossRef]
- Anas, M.; Sharma, R.; Dhamodharan, V.; Pradeepkumar, P.I.; Manhas, A.; Srivastava, K.; Ahmed, S.; Kumar, N. Investigating pharmacological targeting of G-quadruplexes in the human malaria parasite. Biochemistry 2017, 56, 6691–6699. [Google Scholar] [CrossRef] [PubMed]
- Belmonte-Reche, E.; Martínez-García, M.; Guédin, A.; Zuffo, M.; Arévalo-Ruiz, M.; Doria, F.; Campos-Salinas, J.; Maynadier, M.; López-Rubio, J.J.; Freccero, M.; et al. G-quadruplex identification in the genome of protozoan parasites points to naphthalene diimide ligands as new antiparasitic agents. J. Med. Chem. 2018, 61, 1231–1240. [Google Scholar] [CrossRef]
- Harris, L.M.; Monsell, K.R.; Noulin, F.; Famodimu, M.T.; Smargiasso, N.; Damblon, C.; Horrocks, P.; Merrick, C.J. G-Quadruplex DNA motifs in the malaria parasite Plasmodium falciparum and their potential as novel antimalarial drug targets. Antimicrob. Agents Chemother. 2018, 62, 1–14. [Google Scholar] [CrossRef]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Dumetz, F.; Merrick, C.J. Parasitic Protozoa: Unusual Roles for G-Quadruplexes in Early-Diverging Eukaryotes. Molecules 2019, 24, 1339. https://doi.org/10.3390/molecules24071339
Dumetz F, Merrick CJ. Parasitic Protozoa: Unusual Roles for G-Quadruplexes in Early-Diverging Eukaryotes. Molecules. 2019; 24(7):1339. https://doi.org/10.3390/molecules24071339
Chicago/Turabian StyleDumetz, Franck, and Catherine J. Merrick. 2019. "Parasitic Protozoa: Unusual Roles for G-Quadruplexes in Early-Diverging Eukaryotes" Molecules 24, no. 7: 1339. https://doi.org/10.3390/molecules24071339
APA StyleDumetz, F., & Merrick, C. J. (2019). Parasitic Protozoa: Unusual Roles for G-Quadruplexes in Early-Diverging Eukaryotes. Molecules, 24(7), 1339. https://doi.org/10.3390/molecules24071339






