Phytochemical Diversity in Rhizomes of Three Reynoutria Species and their Antioxidant Activity Correlations Elucidated by LC-ESI-MS/MS Analysis
Abstract
:1. Introduction
2. Results and Discussion
2.1. Mass Spectra Analysis, Annotation and Identification of Major Constituents in Extracts and Fractions
2.1.1. Stilbenoids
2.1.2. Carbohydrates
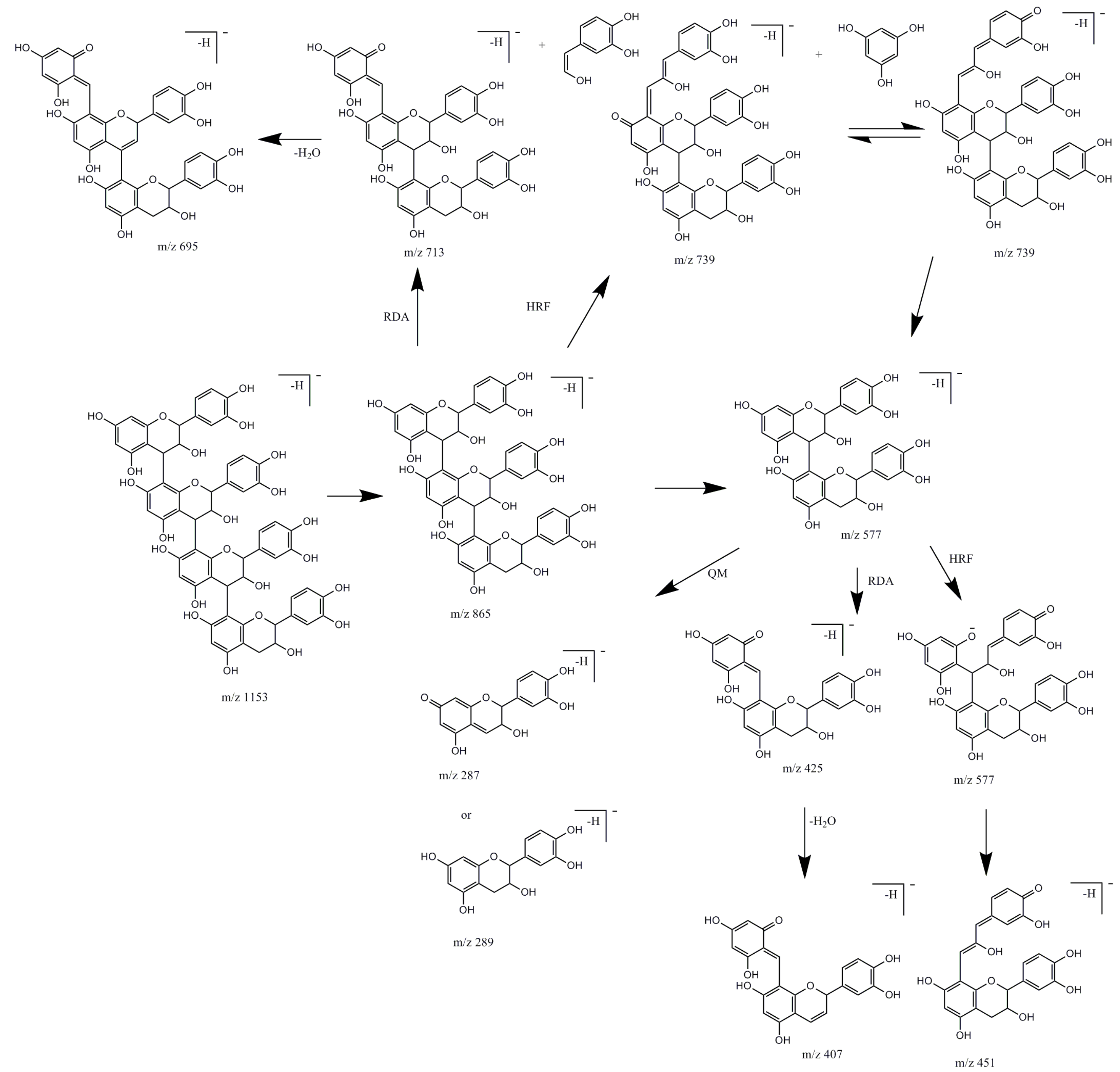
2.1.3. Flavan-3-ols and Procyanidins
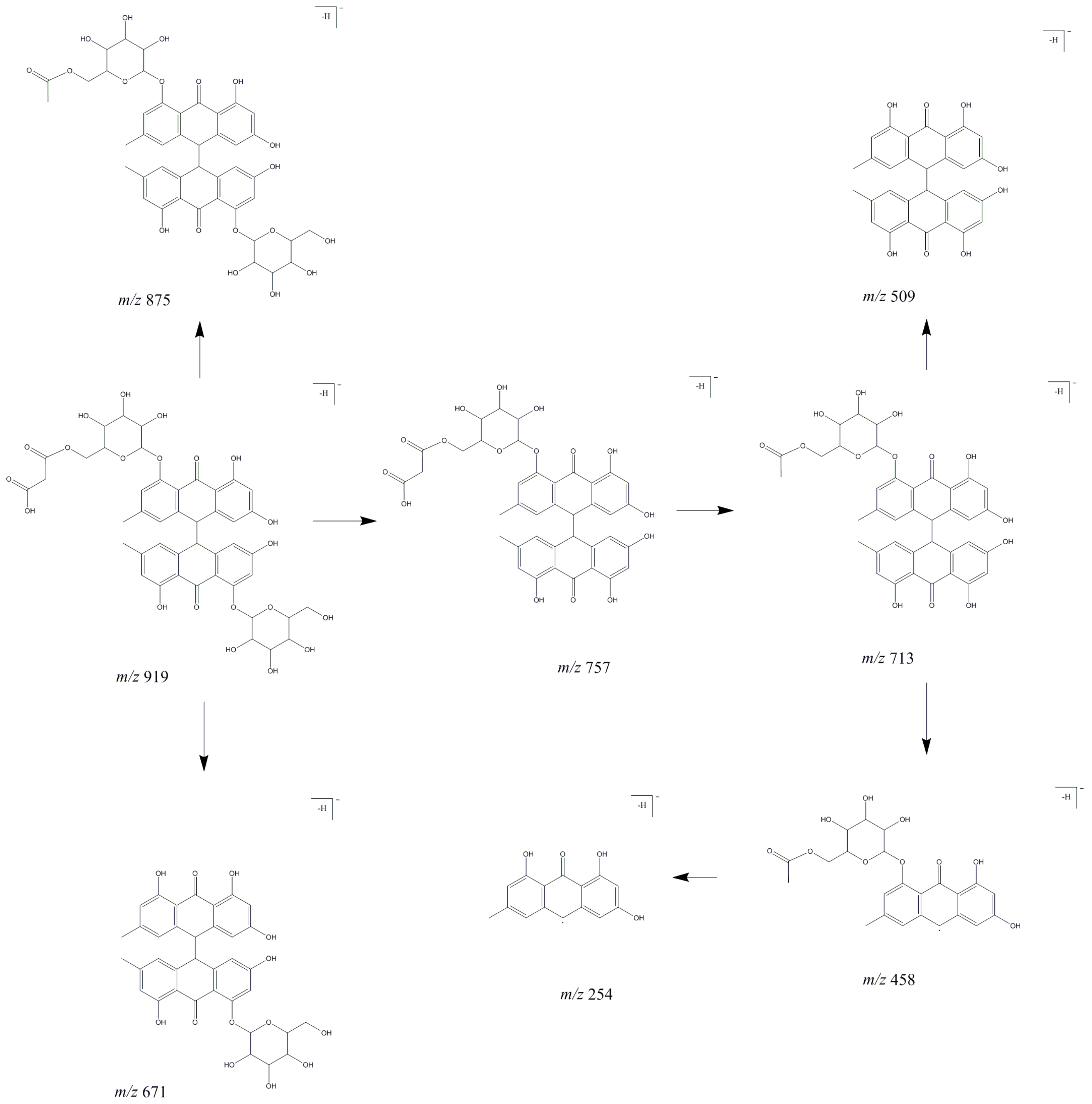
2.1.4. Anthraquinones
2.1.5. Phenylpropanoid Disaccharide Esters
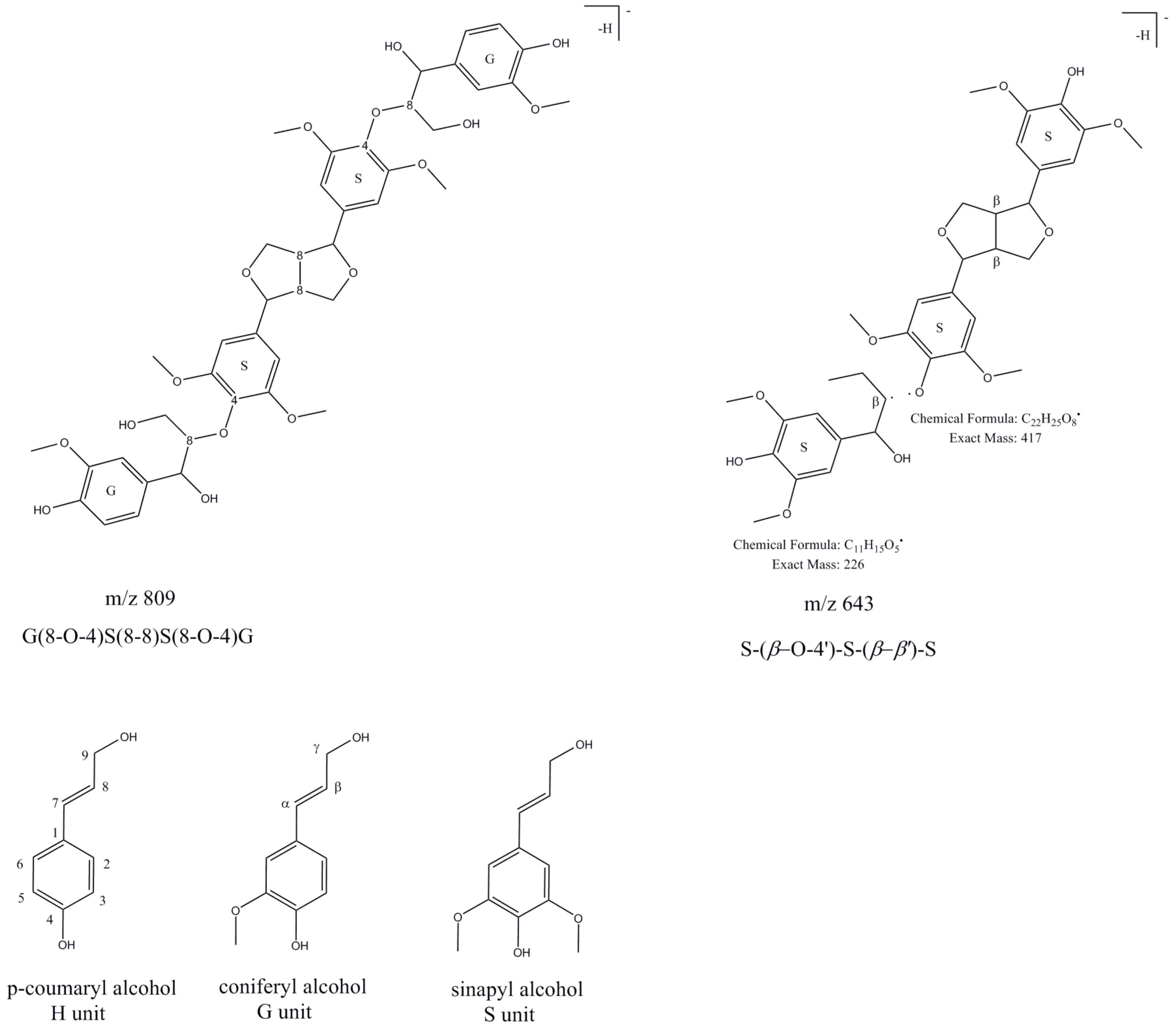
2.1.6. Lignin Oligomers
2.1.7. Other Hydroxycinnamic Acid Derivatives
2.1.8. Naphthalene Derivatives
2.1.9. Other Compounds
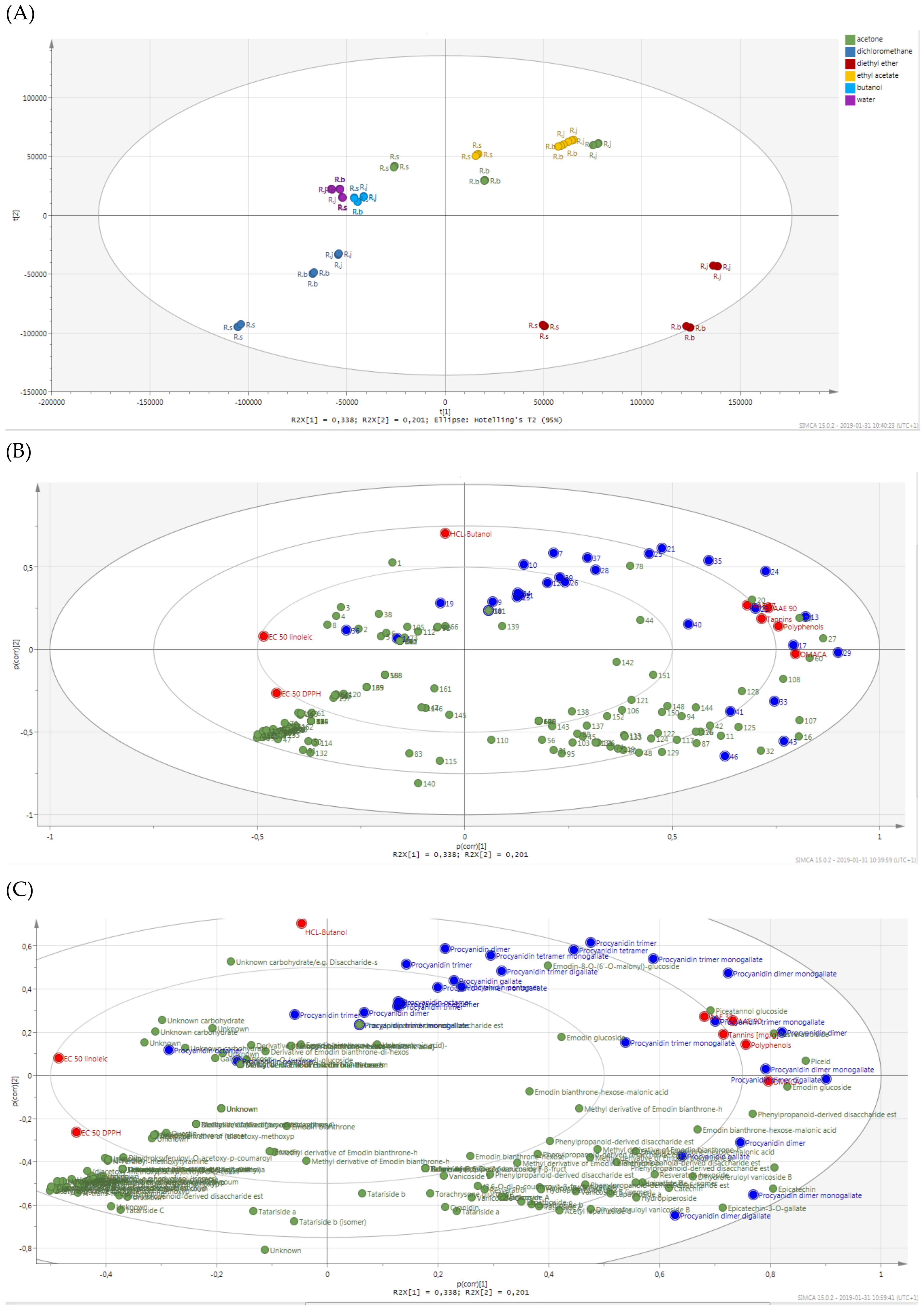
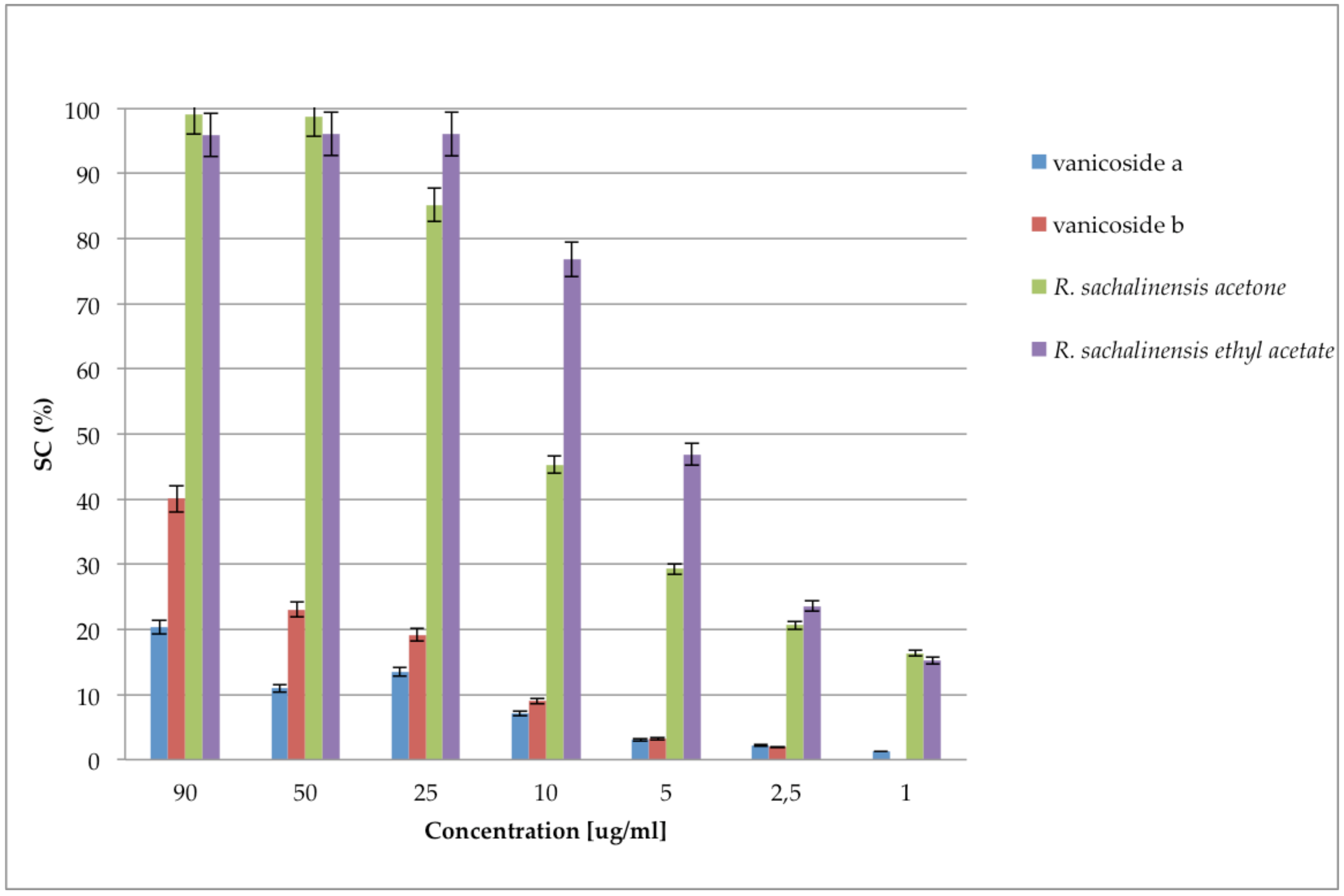
2.2. Antioxidant Activities and Polyphenols Content
3. Materials and Methods
3.1. Plant Material
3.2. Reagents
3.3. DPPH Scavenging Assay
3.4. Phosphomolybdenum Reduction Assay
3.5. Inhibition of Linoleic Acid Peroxidation
3.6. Total Polyphenols and Tannins Content
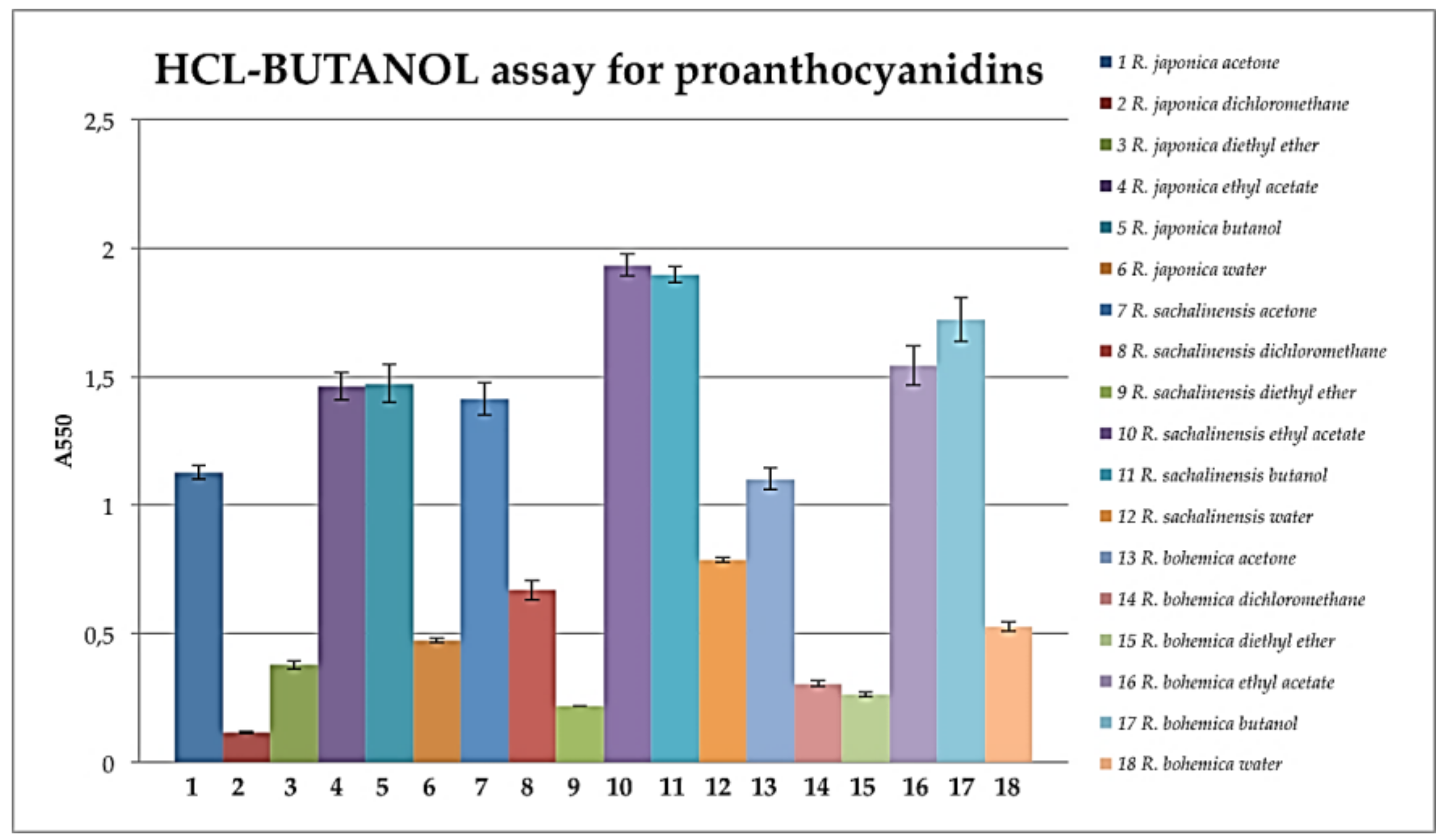
3.7. HCl-Butanol Assay
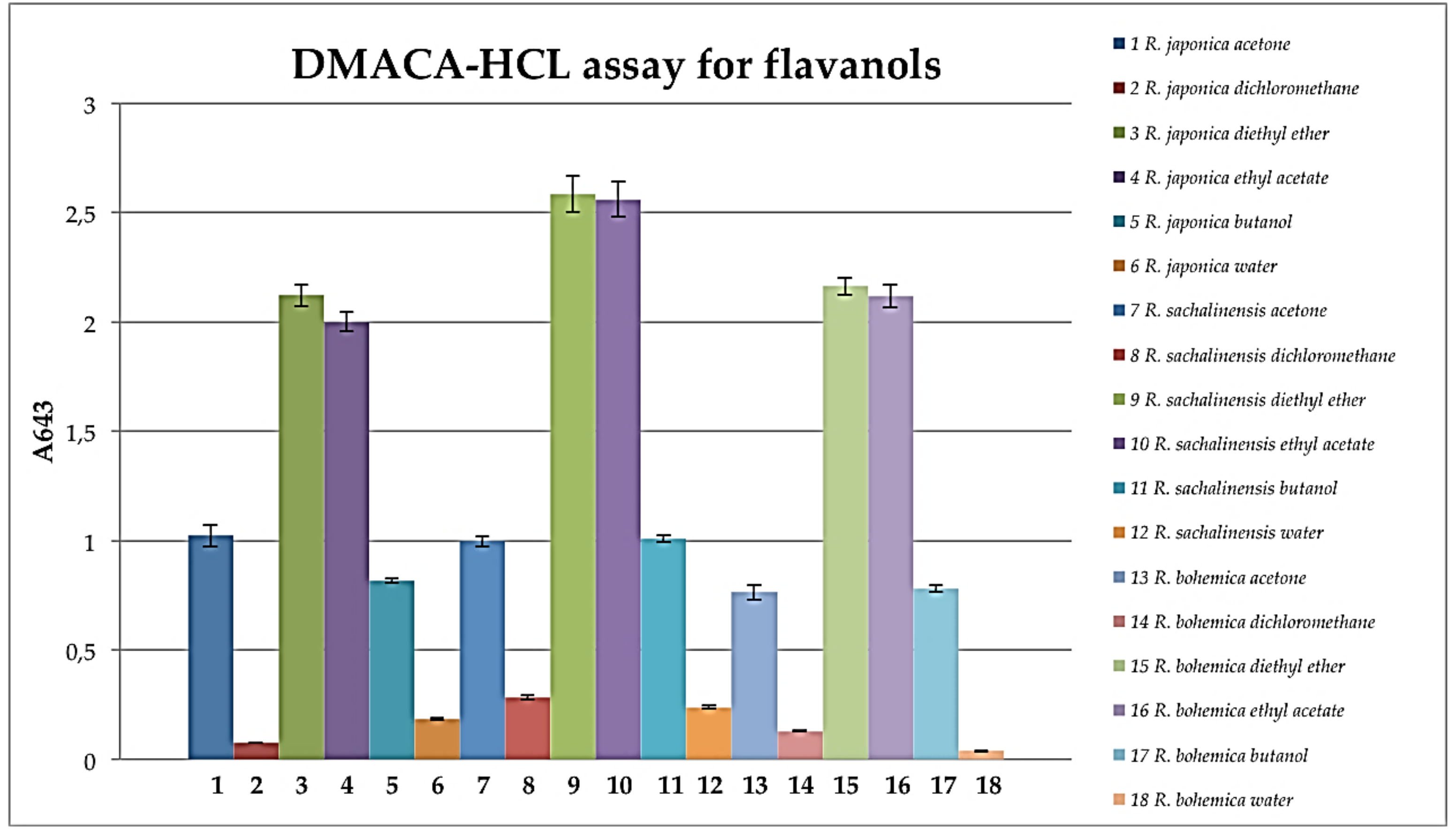
3.8. DMACA-HCl Assay for Flavanols
3.9. HPLC-MS Apparatus
3.10. HPLC-DAD–MS3 Conditions
3.11. Statistical Analysis
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Peng, W.; Qin, R.; Li, X.; Zhou, H. Botany, phytochemistry, pharmacology, and potential application of Polygonum cuspidatum Sieb.et Zucc.: A review. J. Ethnopharmacol. 2013, 148, 729–745. [Google Scholar] [CrossRef] [PubMed]
- Truong, V.L.; Jun, M.; Jeong, W.S. Role of resveratrol in regulation of cellular defense systems against oxidative stress. BioFactors 2018, 44, 36–49. [Google Scholar] [CrossRef] [PubMed]
- European Pharmacopoeia, 9th ed.; Council of Europe: Strasbourg, France, 2017; pp. 1481–1483.
- Eom, M.R.; Weon, J.B.; Jung, Y.S.; Ryu, G.H.; Yang, W.S.; Ma, C.J. Neuroprotective compounds from Reynoutria sachalinensis. Arch. Pharm. Res. 2017, 40, 704–712. [Google Scholar] [CrossRef] [PubMed]
- Nawrot-Hadzik, I.; Granica, S.; Domaradzki, K.; Pecio, Ł.; Matkowski, A. Isolation and Determination of Phenolic Glycosides and Anthraquinones from Rhizomes of Various Reynoutria Species. Planta Med. 2018, 84, 1118–1126. [Google Scholar] [CrossRef] [PubMed]
- Matkowski, A.; Jamiołkowska-Kozlowska, W.; Nawrot, I. Chinese medicinal herbs as source of antioxidant compounds—Where tradition meets the future. Curr. Med. Chem. 2013, 20, 984–1004. [Google Scholar] [PubMed]
- Lee, M.H.; Kao, L.; Lin, C.C. Comparison of the antioxidant and transmembrane permeative activities of the different Polygonum cuspidatum extracts in phospholipid-based microemulsions. J. Agric. Food Chem. 2011, 59, 9135–9141. [Google Scholar] [CrossRef] [PubMed]
- Ding, X.P.; Zhang, C.L.; Qi, J.; Sun, L.Q.; Qin, M.J.; Yu, B.Y. The Spectrum-Effect integrated fingerprint of Polygonum cuspidatum based on HPLC-diode array detection-flow injection-chemiluminescence. Chin. J. Nat. Med. 2013, 11, 546–552. [Google Scholar] [CrossRef]
- Pan, Y.; Zhang, X.; Wang, H.; Liang, Y.; Zhu, J.; Li, H.; Zhang, Z.; Wu, Q. Antioxidant potential of ethanolic extract of Polygonum cuspidatum and application in peanut oil. Food Chem. 2007, 105, 1518–1524. [Google Scholar] [CrossRef]
- Wang, T.H.; Zhang, J.; Qiu, X.H.; Bai, J.Q.; Gao, Y.H.; Xu, W. Application of ultra-high-performance liquid chromatography coupled with LTQ-orbitrap mass spectrometry for the qualitative and quantitative analysis of Polygonum multiflorum Thumb. and its processed products. Molecules 2016, 21, 40. [Google Scholar] [CrossRef]
- Fu, J.; Wang, M.; Guo, H.; Tian, Y.; Zhang, Z.; Song, R. Profiling of components of rhizoma et radix polygoni cuspidati by high-performance liquid chromatography with ultraviolet diode-array detector and ion trap/time-of-flight mass spectrometric detection. Pharmacogn. Mag. 2015, 11, 486–501. [Google Scholar]
- Rue, E.A.; Rush, M.D.; van Breemen, R.B. Procyanidins: A comprehensive review encompassing structure elucidation via mass spectrometry. Phytochem. Rev. 2018, 17, 1–16. [Google Scholar] [CrossRef]
- Rockenbach, I.I.; Jungfer, E.; Ritter, C.; Santiago-Schübel, B.; Thiele, B.; Fett, R.; Galensa, R. Characterization of flavan-3-ols in seeds of grape pomace by CE, HPLC-DAD-MS n and LC-ESI-FTICR-MS. Food Res. Int. 2012, 48, 848–855. [Google Scholar] [CrossRef]
- Hellström, J.; Sinkkonen, J.; Karonen, M.; Mattila, P. Isolation and structure elucidation of procyanidin oligomers from Saskatoon berries (Amelanchier alnifolia). J. Agric. Food Chem. 2007, 55, 157–164. [Google Scholar] [CrossRef]
- Glavnik, V.; Vovk, I.; Albreht, A. High performance thin-layer chromatography–mass spectrometry of Japanese knotweed flavan-3-ols and proanthocyanidins on silica gel plates. J. Chromatogr. A 2017, 1482, 97–108. [Google Scholar] [CrossRef]
- Fan, P.; Terrier, L.; Hay, A.; Marston, A.; Hostettmann, K. Antioxidant and enzyme inhibition activities and chemical profiles of Polygonum sachalinensis F. Schmidt ex Maxim (Polygonaceae). Fitoterapia 2010, 81, 124–131. [Google Scholar] [CrossRef]
- Hwang, J.T.; Kim, Y.; Jang, H.J.; Oh, H.M.; Lim, C.H.; Lee, S.W.; Rho, M.C. Study of the UV light conversion of feruloyl amides from Portulaca oleracea and their inhibitory effect on IL-6-induced STAT3 activation. Molecules 2016, 21, 865. [Google Scholar] [CrossRef]
- Said, R.B.; Hamed, A.I.; Mahalel, U.A.; Al-Ayed, A.S.; Kowalczyk, M.; Moldoch, J.; Oleszek, W.; Stochmal, A. Tentative characterization of polyphenolic compounds in the male flowers of Phoenix dactylifera by liquid chromatography coupled with mass spectrometry and DFT. Int. J. Mol. Sci. 2017, 18, 512. [Google Scholar]
- Zheng, C.; Hu, C.; Ma, X.; Peng, C.; Zhang, H.; Qin, L. Cytotoxic phenylpropanoid glycosides from Fagopyrum tataricum (L.) Gaertn. Food Chem. 2012, 132, 433–438. [Google Scholar] [CrossRef]
- Wang, L.; Sang, M.; Liu, E.; Banahene, P.O.; Zhang, Y.; Wang, T.; Han, L.; Gao, X. Rapid profiling and pharmacokinetic studies of major compounds in crude extract from Polygonum multiflorum by UHPLC-Q-TOF-MS and UPLC-MS/MS. J. Pharm. Biomed. Anal. 2017, 140, 45–61. [Google Scholar] [CrossRef]
- Xu, W.; Zhang, J.; Huang, Z.; Qiu, X. Identification of new dianthrone glycosides from Polygonum multiflorum Thunb. using high-performance liquid chromatography coupled with LTQ-Orbitrap mass spectrometry detection: A strategy for the rapid detection of new low abundant metabolites from tradi. Anal. Methods 2012, 4, 1806–1812. [Google Scholar] [CrossRef]
- Evtuguin, D.V.; Amado, F.M.L. Application of electrospray ionization mass spectrometry to the elucidation of the primary structure of lignin. Macromol. Biosci. 2003, 3, 339–343. [Google Scholar] [CrossRef]
- Fan, P.; Hay, A.E.; Marston, A.; Lou, H.; Hostettmann, K. Chemical variability of the invasive neophytes Polygonum cuspidatum Sieb. and Zucc. and Polygonum sachalinensis F. Schmidt ex Maxim. Biochem. Syst. Ecol. 2009, 37, 24–34. [Google Scholar] [CrossRef]
- Morreel, K. Profiling of oligolignols reveals monolignol coupling conditions in lignifying poplar xylem. Plant Physiol. 2004, 136, 3537–3549. [Google Scholar] [CrossRef]
- Banoub, J.; Delmas, G.H.; Joly, N.; Mackenzie, G.; Cachet, N.; Benjelloun-Mlayah, B.; Delmas, M. A critique on the structural analysis of lignins and application of novel tandem mass spectrometric strategies to determine lignin sequencing. J. Mass Spectrom. 2015, 50, 5–48. [Google Scholar] [CrossRef]
- Takasaki, M.; Kuroki, S.; Kozuka, M.; Konoshima, T. New phenylpropanoid esters of sucrose from Polygonum lapathifolium. J. Nat. Prod. 2001, 64, 1305–1308. [Google Scholar] [CrossRef]
- Chen, X.; Wang, R.; Liu, B. An update on oligosaccharides and their esters from traditional chinese medicines: Chemical structures and biological activities. Evid. Based. Complement. Alternat. Med. 2015, 2015, 512675. [Google Scholar] [CrossRef]
- Van Kiem, P.; Nhiem, N.X.; Cuong, N.X.; Hoa, T.Q.; Huong, H.T.; Huong, L.M.; Van Minh, C.; Kim, Y.H. New phenylpropanoid esters of sucrose from Polygonum hydropiper and their antioxidant activity. Arch. Pharm. Res. 2008, 31, 1477–1482. [Google Scholar] [CrossRef]
- Ozarowski, M.; Piasecka, A.; Paszel-Jaworska, A.; Chaves, D.S.D.A.; Romaniuk, A.; Rybczynska, M.; Gryszczynska, A.; Sawikowska, A.; Kachlicki, P.; Mikolajczak, P.L.; et al. Comparison of bioactive compounds content in leaf extracts of Passiflora incarnata, P. caerulea and P. alata and in vitro cytotoxic potential on leukemia cell lines. Braz. J. Pharmacogn. 2018, 28, 179–191. [Google Scholar] [CrossRef]
- Nguyen, T.K.O.; Jamali, A.; Grand, E.; Morreel, K.; Marcelo, P.; Gontier, E.; Dauwe, R. Phenylpropanoid profiling reveals a class of hydroxycinnamoyl glucaric acid conjugates in Isatis tinctoria leaves. Phytochemistry 2017, 144, 127–140. [Google Scholar] [CrossRef]
- Zhao, Y.; Lee, M.-J.; Cheung, C.; Ju, J.-H.; Chen, Y.-K.; Liu, B.; Hu, L.-Q.; Yang, C.S. Analysis of multiple metabolites of tocopherols and tocotrienols in mice and humans. J. Agric. Food Chem. 2010, 58, 4844–4852. [Google Scholar] [CrossRef]
- Mechelke, M.; Herlet, J.; Benz, J.P.; Schwarz, W.H.; Zverlov, V.V.; Liebl, W.; Kornberger, P. HPAEC-PAD for oligosaccharide analysis—Novel insights into analyte sensitivity and response stability. Anal. Bioanal. Chem. 2017, 409, 7169–7181. [Google Scholar] [CrossRef] [PubMed]
- Ye, M.; Han, J.; Chen, H.; Zheng, J.; Guo, D. Analysis of phenolic compounds in rhubarbs using liquid chromatography coupled with electrospray ionization mass spectrometry. J. Am. Soc. Mass Spectrom. 2007, 18, 82–91. [Google Scholar] [CrossRef]
- Wang, G.Y.; Chen, F.F.; Shi, Y.P. Ultra-performance LC-photodiode array-eλ-ESI-MS/MS screening method for the detection of radical-scavenging natural antioxidants from Radix et Rhizoma Rhei. J. Sep. Sci. 2011, 34, 268–277. [Google Scholar] [CrossRef] [PubMed]
- Doherty, W.O.S.; Mousavioun, P.; Fellows, C.M. Value-adding to cellulosic ethanol: Lignin polymers. Ind. Crops Prod. 2011, 33, 259–276. [Google Scholar] [CrossRef]
- Matsuda, S.; Kadota, S.; Tai, T.; Kikuchi, T. Isolation and structures of hedyotisol-A, -B, and -C novel dilignans from Hedyotis lawsoniae. Chem. Pharm. Bull. 1984, 32, 5066–5069. [Google Scholar] [CrossRef]
- Jaiswal, R.; Sovdat, T.; Vivan, F.; Kuhnert, N. Profiling and characterization by LC-MSn of the chlorogenic acids and hydroxycinnamoylshikimate esters in Mateé (Ilex paraguariensis). J. Agric. Food Chem. 2010, 58, 5471–5484. [Google Scholar] [CrossRef] [PubMed]
- Karaköse, H.; Jaiswal, R.; Kuhnert, N. Characterization and quantification of hydroxycinnamate derivatives in Stevia rebaudiana leaves by LC-MSn. J. Agric. Food Chem. 2011, 59, 10143–10150. [Google Scholar] [CrossRef] [PubMed]
- Jaiswal, R.; Matei, M.F.; Golon, A.; Witt, M.; Kuhnert, N. Understanding the fate of chlorogenic acids in coffee roasting using mass spectrometry based targeted and non-targeted analytical strategies. Food Funct. 2012, 3, 976–984. [Google Scholar] [CrossRef]
- Dalzell, S.A.; Kerven, G.L. A rapid method for the measurement of Leucaena spp. proanthocyanidins by the proanthocyanidin (butanol/HCl) assay. J. Sci. Food Agric. 1998, 78, 405–416. [Google Scholar] [CrossRef]
- Scalbert, A. Quantitative methods for the estimation of tannins in plant tissues. In Plant Polyphenols; Hemingway, R.W., Laks, P.E., Eds.; Plenum Press: New York, NY, USA, 1992; Volume 59, pp. 259–280. [Google Scholar]
- Mikulski, D.; Molski, M. Quantitative structure–antioxidant activity relationship of trans-resveratrol oligomers, trans-4,4′-dihydroxystilbene dimer, trans-resveratrol-3-O-glucuronide, glucosides: Trans-piceid, cis-piceid, trans-astringin and trans-resveratrol-4′-O-β-D-glucopyranoside. Eur. J. Med. Chem. 2010, 45, 2366–2380. [Google Scholar] [PubMed]
- Wenzel, E.; Soldo, T.; Erbersdobler, H.; Somoza, V. Bioactivity and metabolism of trans-resveratrol orally administered to Wistar rats. Mol. Nutr. Food Res. 2005, 49, 482–494. [Google Scholar] [CrossRef] [PubMed]
- Lachowicz, S.; Oszmiański, J.; Wojdyło, A.; Cebulak, T.; Hirnle, L.; Siewiński, M. UPLC-PDA-Q/TOF-MS identification of bioactive compounds and on-line UPLC-ABTS assay in Fallopia japonica Houtt. and Fallopia sachalinensis Schmidt leaves and rhizomes grown in Poland. Eur. Food Res. Technol. 2019, 245, 691–706. [Google Scholar] [CrossRef]
- Cos, P.; Bruyne, T.D.; Hermans, N.; Apers, S.; Berghe, D.V.; Vlietinck, A.J. Proanthocyanidins in health care: Current and new trends. Curr. Med. Chem. 2004, 11, 1345–1359. [Google Scholar] [CrossRef]
- Koleckar, V.; Kubikova, K.; Rehakova, Z.; Kuca, K.; Jun, D.; Jahodar, L.; Opletal, L. Condensed and hydrolysable tannins as antioxidants influencing the health. Mini Rev. Med. Chem. 2008, 8, 436–447. [Google Scholar] [CrossRef] [PubMed]
- Castellain, R.C.L.; Gesser, M.; Tonini, F.; Schulte, R.V.; Demessiano, K.Z.; Wolff, F.R.; Delle-Monache, F.; Netz, D.J.A.; Cechinel-Filho, V.; de Freitas, R.A.; et al. Chemical composition, antioxidant and antinociceptive properties of Litchi chinensis leaves. J. Pharm. Pharmacol. 2014, 66, 1796–1807. [Google Scholar] [CrossRef]
- Soobrattee, M.A.; Neergheen, V.S.; Luximon-Ramma, A.; Aruoma, O.I.; Bahorun, T. Phenolics as potential antioxidant therapeutic agents: Mechanism and actions. Mutat. Res. 2005, 579, 200–213. [Google Scholar] [CrossRef]
- Prieto, P.; Pineda, M.; Aguilar, M. Spectrophotometric quantitation of antioxidant capacity through the formation of a phosphomolybdenum complex: Specific application to the determination of vitamin E. Anal. Biochem. 1999, 269, 337–341. [Google Scholar] [CrossRef]
- Luo, L.; Cui, Y.; Cheng, J.; Fang, B.; Wei, Z.; Sun, B. An approach for degradation of grape seed and skin proanthocyanidin polymers into oligomers by sulphurous acid. Food Chem. 2018, 256, 203–211. [Google Scholar] [CrossRef]
- Dallas, C.; Hipolito-Reis, P.; Ricardo-da-Silva, J.M.; Laureano, O. Influence of acetaldehyde, pH, and temperature on transformation of procyanidins in model wine solutions. Am. J. Enol. Vitic. 2003, 54, 119–124. [Google Scholar]
- Fan, P.; Marston, A.; Hay, A.E.; Hostettmann, K. Rapid separation of three glucosylated resveratrol analogues from the invasive plant Polygonum cuspidatum by high-speed countercurrent chromatography. J. Sep. Sci. 2009, 32, 2979–2984. [Google Scholar] [CrossRef]
- Zhang, D.; Li, X.; Hao, D.; Li, G.; Xu, B.; Ma, G.; Su, Z. Systematic purification of polydatin, resveratrol and anthraglycoside B from Polygonum cuspidatum Sieb. et Zucc. Sep. Purif. Technol. 2009, 66, 329–339. [Google Scholar] [CrossRef]
- Fan, P.; Hostettmann, K.; Lou, H. Allelochemicals of the invasive neophyte Polygonum cuspidatum Sieb. & Zucc. (Polygonaceae). Chemoecology 2010, 20, 223–227. [Google Scholar]
- Yang, F.; Zhang, T.; Ito, Y. Large-scale separation of resveratrol, anthraglycoside A and anthraglycoside B from Polygonum cuspidatum Sieb. et Zucc by high-speed counter-current chromatography. J. Chromatogr. A 2001, 919, 443–448. [Google Scholar] [CrossRef]
- Chu, X.; Sun, A.; Liu, R. Preparative isolation and purification of five compounds from the Chinese medicinal herb Polygonum cuspidatum Sieb. et Zucc. by high-speed counter-current chromatography. J. Chromatogr. A 2005, 1097, 33–39. [Google Scholar] [CrossRef]
- Pharmacopoeia of China. Part 1; China Medical Science and Technology Press: Beijing, China, 2005; pp. 180–181.
- Brand-Williams, W.; Cuvelier, M.E.; Berset, C. Use of a free radical method to evaluate antioxidant activity. LWT Food Sci. Technol. 1995, 28, 25–30. [Google Scholar] [CrossRef]
- Woźniak, D.; Dryś, A.; Matkowski, A. Antiradical and antioxidant activity of flavones from Scutellariae baicalensis radix. Nat. Prod. Res. 2015, 29, 1567–1570. [Google Scholar] [CrossRef]
- Nawrot-Hadzik, I.; Granica, S.; Abel, R.; Czapor-Irzabek, H.; Matkowski, A. Analysis of antioxidant polyphenols in loquat leaves using HPLC-based activity profiling. Nat. Prod. Commun. 2016, 12, 163–166. [Google Scholar]
- Li, Y.G.; Tanner, G.; Larkin, P. The DMACA Protocol and the threshold proanthocyanidin content for bloat safety in forage legumes. J. Sci. Food Agric. 1996, 70, 89. [Google Scholar] [CrossRef]
Sample Availability: Samples of the compounds are available from the first author (I.N.-H.) upon request. |
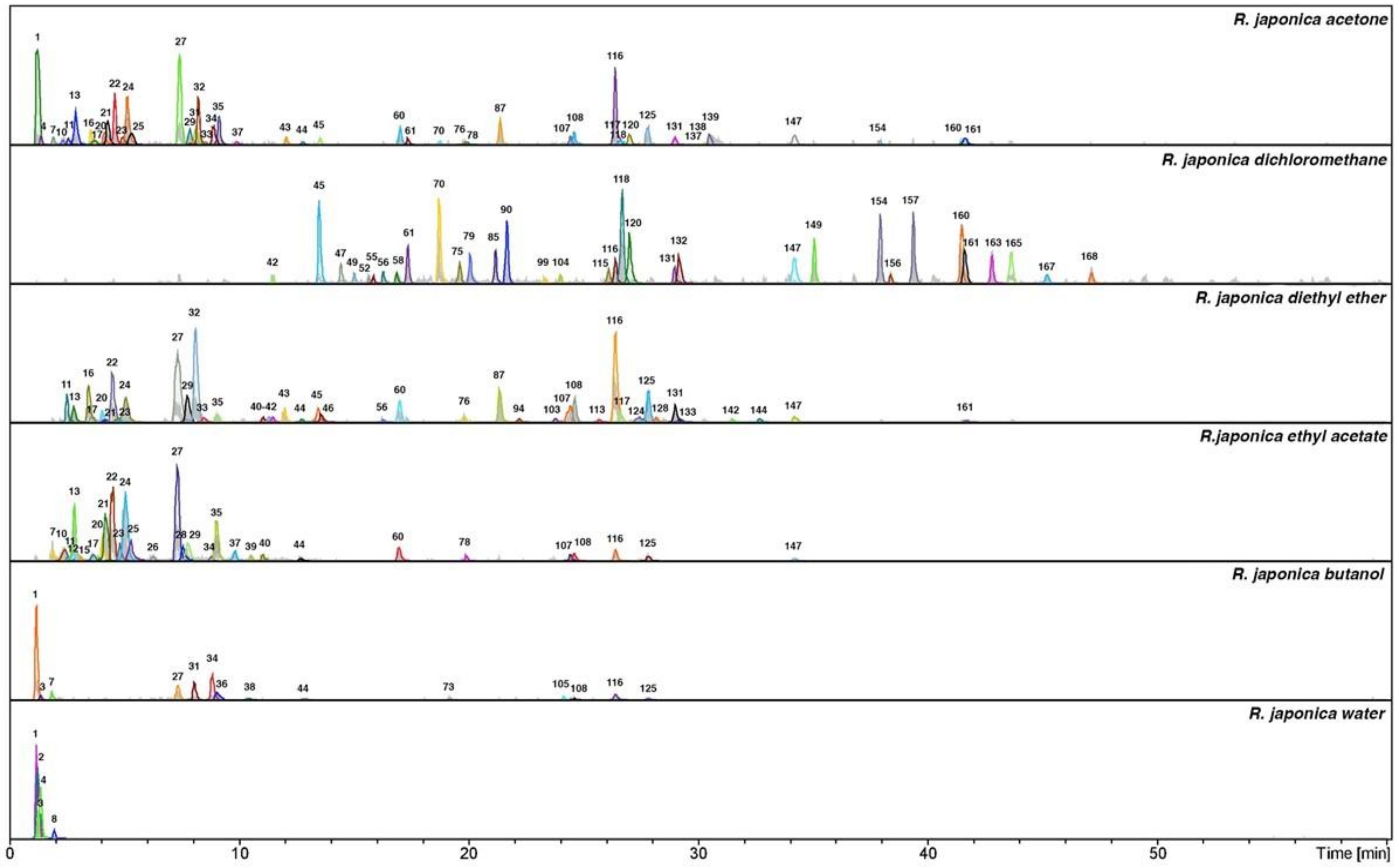
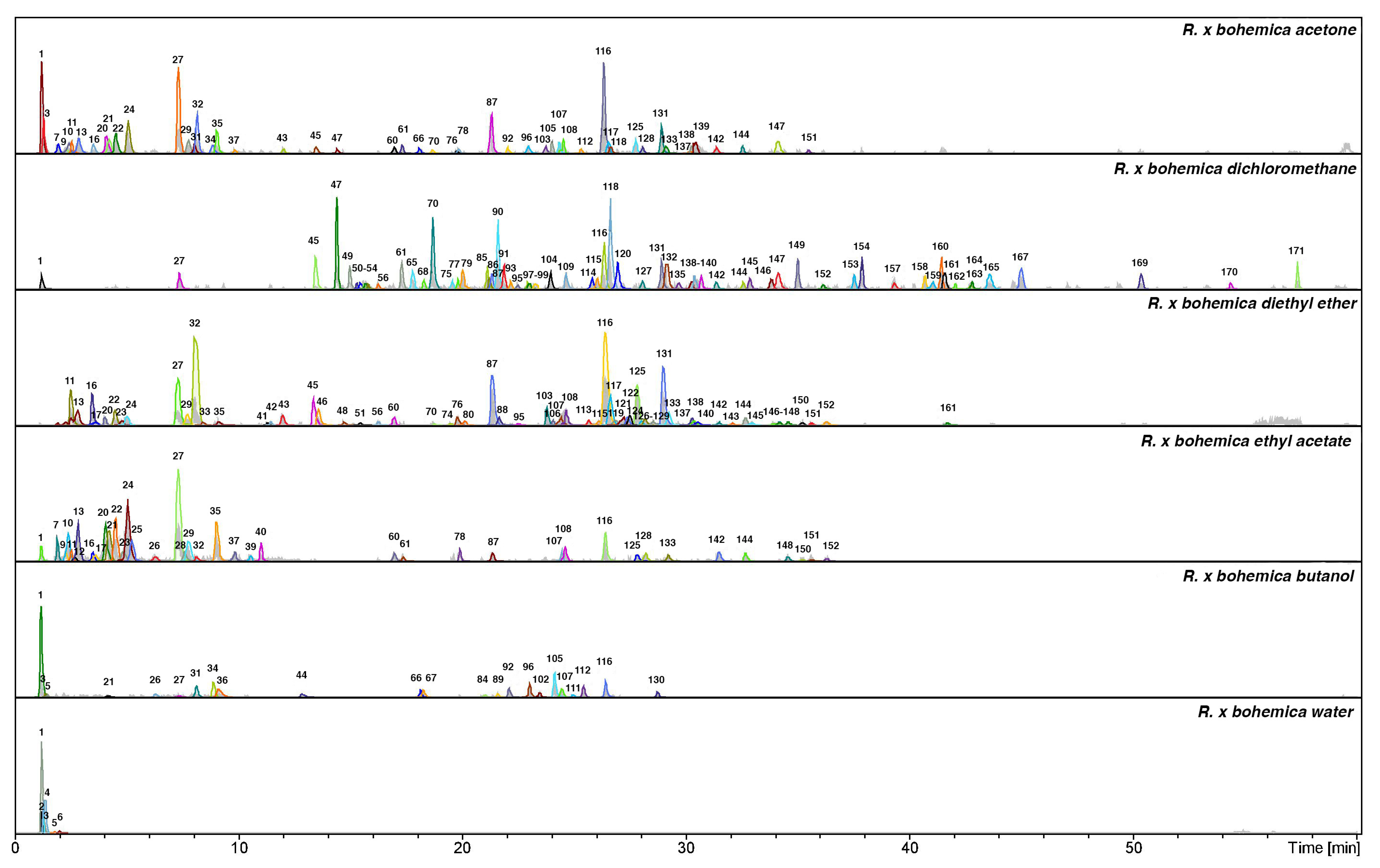
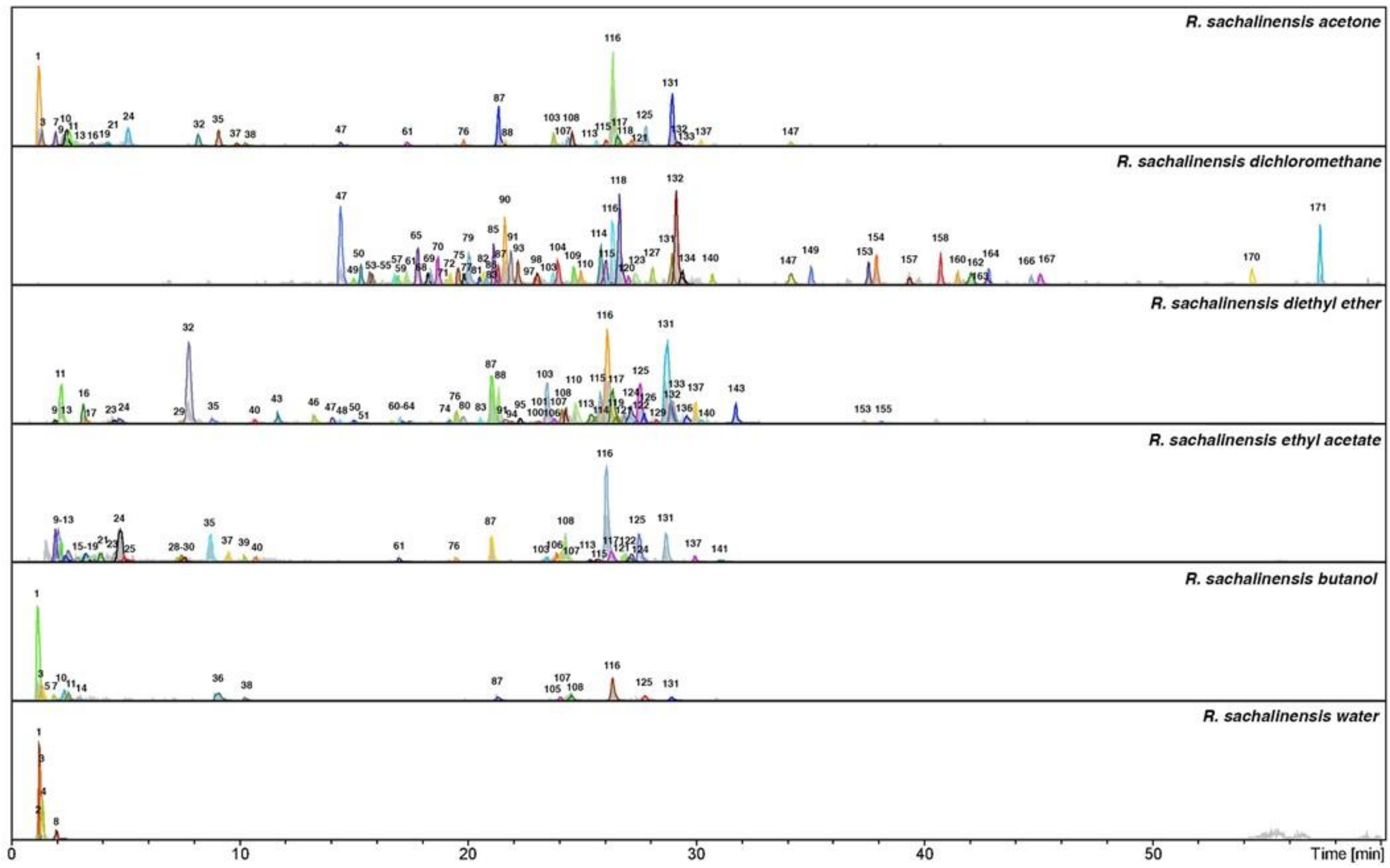

| Nr. | Identification | Rt | λ max (nm) | m/z [M − H]− | MS2 ions | MS3 ions | NL Da | References |
|---|---|---|---|---|---|---|---|---|
| 1 | Unknown carbohydrate/e.g., Disaccharide-sucrose | 1.2 | ND | 341.15 | 178.82b | 160.81b, 142.78 | 162 | [10] |
| 2 | Unknown carbohydrate | 1.21 | ND | 683.18 | 341.04b | |||
| 3 | Unknown carbohydrate | 1.3 | ND | 781.12 | 439.02b | 420.95, 341.09 [M − 2H]2−, 277.01b, 178.80 | 162 | |
| 4 | Unknown carbohydrate | 1.4 | ND | 781.12 | 439.04b | 421.04, 340.98 [M − 2H]2− b, 276.87, 178.83 | 162 | |
| 5 | Galloyl-glucose | 1.5 | 210, 276 | 331.13 | 270.72, 168.58b | [11] | ||
| 6 | Unknown | 1.8 | 235, 275, 325 | 477.1 | 459.05b, 357.04, 234.83, 150.80 | |||
| 7 | Procyanidin dimer, Type B | 1.9 | 225, 280 | 577.11 | 559.04, 450.99, 424.96b, 407.15, 288.93, 286.97 | 406.90b, 381.02, 272.85 | 152 | [11,12,13] |
| 8 | Unknown | 2.0 | 235, 275, 325 | 439.00b, 425.05 | 344.98, 240.80b | |||
| 9 | Procyanidin dimer, Type B | 2.3 | 225, 280 | 577.13 | 559.04, 450.97, 424.95b, 407.09, 288.93, 286.97 | 406.91b, 381.12, 339.07, 272.90 | 152 | [11,12,13] |
| 10 | Procyanidin trimer, Type B | 2.4 | 225, 280 | 865.19 | 739.14, 695.12b, 577.07, 406.98, 286.87 | [12,13] | ||
| 11 | Catechin * | 2.6 | 225, 280 | 288.99 | 270.90, 244.91b, 204.85, 178.83 | |||
| 12 | Procyanidin trimer monogallate | 2.7 | 225, 280 | 1017.2 | 891.18, 865.18, 847.12, 729.12b, 577.11, 407.07, 287.81 | [13] | ||
| 13 | Procyanidin dimer, Type B | 2.8 | 225, 280 | 577.08 | 559.05, 451.00, 424.96b, 407.00, 288.90, 286.97 | 406.90b, 381.11, 272.87 | 152 | [11,12,13] |
| 14 | Procyanidin pentamer | 3.1 | 225, 280 | 720.55 [M − 2H]2− | 1315.33, 1151.29b, 1027.23, 863.22, 635.05, 577.05, 288.85 | [12] | ||
| 15 | Procyanidin trimer, Type B | 3.2 | 225, 280 | 865.21 | 739.13, 695.14b, 577.08, 407.00, 286.90 | [12,13] | ||
| 16 | Epicatechin * | 3.5 | 225, 280 | 288.82 | 270.76, 244.75b, 230.68, 204.70, 178.65 | |||
| 17 | Procyanidin dimer monogallate | 3.6 | 225, 280 | 729.17 | 577.06b, 425.06, 407.07, 286.92 | 559.05, 450.98, 424.98, 407.00b, 288.90 | 152 | [11,13] |
| 18 | Procyanidin trimer monogallate | 3.7 | 225, 280 | 1017.2 | 865.16b, 847.15, 729.11, 577.06, 406.97 | 847.14, 695.12b, 577.05, 394.95, 286.81 | 152 | [13] |
| 19 | Procyanidin trimer, Type B | 4.0 | 225, 280 | 865.2 | 739.15, 695.14b, 577.07, 406.99, 286.89 | [12,13] | ||
| 20 | Piceatannol glucoside * | 4.1 | 220, 305, 318 | 405.06 | 242.73b | 224.70b, 214.68, 200.69, 184.64, 174.73 | 162 | |
| 21 | Procyanidin trimer, Type B | 4.2 | 225, 280 | 865.19 | 739.15, 695.12b, 577.08, 406.98, 286.87 | [12,13] | ||
| 22 | Resveratroloside * | 4.6 | 219, 304, 315 | 389.07, 435.13 [M + HCOO]− | 389.07b, 226.91 | |||
| 23 | Procyanidin trimer monogallate | 4.8 | 225, 280 | 1017.19 | 865.14, 847.15, 729.16b, 603.09, 559.08, 407.06, 288.89 | 847.07b, 695.02, 575.94, 451.02, 286.80 | 152 | [13] |
| 24 | Procyanidin dimer monogallate | 5.1 | 225, 280 | 729.12 | 577.05, 559.05, 451.00, 441.01, 407.02b, 288.90 | 559.01b, 450.99, 406.95, 288.86 | 152 | [11,13] |
| 25 | Procyanidin tetramer, Type B | 5.3 | 225, 280 | 1153.26 | 1001.20, 983.20, 865.16b, 739.12, 575.09, 449.02 | 983.18b, 804.93, 533.18, 382.95 | 152 | [12,13] |
| 26 | Procyanidin pentamer | 6.3 | 225, 280 | 720.55 [M − 2H]2− | 1315.33, 1151.29b, 1027.23, 863.22, 635.05, 577.05, 288.85 | [12] | ||
| 27 | Piceid * | 7.4 | 218, 308, 318 | 389.12, 435.07 [M + HCOO]− | 226.71b | |||
| 28 | Procyanidin trimer digallate | 7.6 | 225, 280 | 1169.24 | 1151.24, 999.21, 881.22b, 729.18, 603.11, 406.98 | [13] | ||
| 29 | Procyanidin dimer digallate, Type B | 7.7 | 225, 280 | 881.16 | 729.12b, 559.08, 407.01, 288.86 | 603.07, 577.08, 559.07, 451.01b, 407.10, 288.98 | 152 | [13] |
| 30 | Procyanidin trimer monogallate | 7.9 | 225, 280 | 1017.22 | 865.17, 847.15, 729.13b, 603.09, 575.09, 406.98, 286.88 | 847.07b, 739.12, 714.02, 577.04, 448.84, 288.69 | 152 | [13] |
| 31 | Procyanidin heptamer | 8.1 | 225, 280 | 1008.63 [M − 2H]−, 1152.74, 1021.26, 999.18, 631.20, 567.10, 499.09b | 484.00b, 452.98, 419.04, 345.92, 314.85, 288.79 | [12,14] | ||
| 32 | Epicatechin-3-O-gallate * | 8.2 | 220, 280 | 440.95 | 330.82, 302.80, 288.82b, 270.81, 244.82 | |||
| 33 | Procyanidin dimer, Type B | 8.4 | 225, 280 | 577.09 | 559.08, 450.96, 424.94b, 407.06, 288.92 | 406.91b, 381.04, 339.00, 272.85 | 152 | [11,12,13] |
| 34 | Procyanidin octamer | 8.8 | 225, 280 | 1152.70 [M − 2H]−, 901.21, 879.11, 507.02, 439.04b | 423.93b, 392.86, 358.98, 315.84 | [15] | ||
| 35 | Procyanidin trimer monogallate | 9.0 | 225, 280 | 1017.21 | 865.12, 847.15, 729.15b, 603.08, 406.99, 288.90 | 847.15b, 684.05, 518.85, 451.83, 395.07, 301.69 | 152 | [13] |
| 36 | Procyanidin octamer | 9.1 | 225, 280 | 1152.69 [M − 2H]2−, 901.16, 864.10, 845.10, 439.02, 382.90b | 302.80b, 284.83, 176.67 | [15] | ||
| 37 | Procyanidin tetramer monogallate | 9.8 | 225, 280 | 652.11 [M − 2H]2− b, 1305.32 | 1179.22, 1017.23, 863.18, 729.11, 576.00, 567.07b, 440.99, 288.88, 286.86 | [15] | ||
| 38 | Unknown | 10.3 | 225, 280 | 1532.47, 382.90b | 302.77b, 284.88, 178.67 | |||
| 39 | Procyanidin gallate | 10.5 | 225, 280 | 796.27 [M − 3H]3 b,948.11 [M − 2H]2 | 1467.40, 1305.34, 1179.31, 1017.21, 863.17, 729.15b, 440.96, 288.86 | [15] | ||
| 40 | Procyanidin trimer monogallate | 11.0 | 225, 280 | 1017.2 | 891.16, 847.16, 729.14b, 603.07, 559.05, 407.03, 288.87 | [13] | ||
| 41 | Procyanidin gallate | 11.3 | 225, 280 | 660.32, 505.17b | 1151.19, 999.17, 881.14, 584.04b, 440.96, 302.86 | [12,13,15] | ||
| 42 | Resveratrol-hexoside | 11.4 | 219, 304, 315 | 389.06 | 226.71 | |||
| 43 | Procyanidin dimer monogallate | 12.0 | 225, 280 | 729.13 | 711.11, 603.05, 577.04, 559.05, 407.01b, 288.91 | 559.03, 450.97, 406.93b, 288.85 | 152 | [11,13] |
| 44 | Emodin glucoside * | 12.8 | 220, 247, 269, 281, 423 | 431.3 | 268.75b | 239.63, 226.68, 224.72b | 162 | |
| 45 | Resveratrol * | 13.5 | 218, 306, 318 | 226.78 | 184.60, 158.67b, 142.68 | |||
| 46 | Procyanidin dimer digallate, Type B | 13.6 | 225, 280 | 881.13 | 729.11b, 559.12, 407.05, 288.90 | 603.07, 577.08, 559.04, 451.01b, 407.04, 288.98 | 152 | [13] |
| 47 | N-trans-feruloyltyramine * | 14.4 | 220, 281, 323 | 312.08 | 296.97b, 177.83, 134.87 | |||
| 48 | Acetyl lapathoside d | 14.7 | 220, 290, 315 | 675.24 | 633.17, 615.12, 529.07b, 511.12, 487.12, 453.11, 306.97 | 487.04b, 469.19, 306.96 | 146 | [16] |
| 49 | N-Feruloyl-methoxytyramine | 15.0 | 220, 281, 323 | 342.14 | 327.04b, 308.97, 297.01, 177.84, 134.87 | [17] | ||
| 50 | Phenylpropanoid-derived disaccharide ester | 15.3 | 220, 284, 315 | 655.21 | 613.18b, 595.18, 571.16, 553.10, 425.12, 306.99 | |||
| 51 | Cyanidin | 15.4 | 210, 286, 332 | 286.9 | 268.79, 150.59b, 134.71, 124.75, 106.72 | [18] | ||
| 52 | Unknown | 15.6 | 225, 287, 315 | 585.28 | 537.17b, 371.13, 359.13 | |||
| 53 | Unknown | 15.7 | 225, 287, 315 | 583.27 | 535.23b, 369.10, 357.25, 194.91 | |||
| 54 | Unknown | 15.8 | 220, 280 | 685.23 | 643.20b, 625.20, 601.19, 337.12, 192.90 | |||
| 55 | Unknown | 15.9 | 220, 287, 315 | 585.29 | 537.20b, 359.12, 345.14 | |||
| 56 | Torachrysone glucoside * | 16.2 | 225, 267, 325 | 407.16 | 244.87b | 229.97 | 162 | |
| 57 | Unknown | 16.8 | 225, 280 | 371.12 | 327.08b, 297.08 | |||
| 58 | Unknown | 16.85 | 225, 280 | 597.26 | 553.13, 549.23, 383.11b, 371.12, 194.86 | |||
| 59 | Unknown | 16.9 | 225, 280 | 583.31 | 553.20, 369.11b, 357.15, 194.80 | |||
| 60 | Emodin glucoside * | 17.0 | 221, 247, 269, 281, 423 | 431.04 | 310.84, 292.76, 268.75b | 264.71, 240.73, 224.70b | 162 | |
| 61 | Dihydroksyferuloyl-O-acetoxy-p-coumaroyl-O-caffeoylquinic acid | 17.4 | 214, 282, 325 | 735.27 | 693.19, 559.12b, 541.18 | 517.11b, 499.10, 337.04, 264.90, 192.83 | 176 | [18] |
| 62 | Tatariside e | 17.5 | 220, 290, 315 | 717.39 | 675.19, 571.11b, 529.20, 453.12, 288.94 | 529.06b, 511.05, 469.03, 306.85 | 146 | [19] |
| 63 | Tatariside e | 17.7 | 220, 290, 315 | 717.4 | 675.19, 571.13b, 529.24, 453.10, 288.93 | 529.05b, 511.05, 469.00, 306.85 | 146 | [19] |
| 64 | Unknown | 17.8 | 225, 280 | 314.95 | 299.78b, 270.98, 246.72, 204.68, 178.78 | |||
| 65 | (diacetoxy-methoxyphenyl)acroyl-O-p-coumaroyl-O-caffeoylquinic acid | 17.9 | 214, 282, 325 | 777.25 | 735.22b, 717.25, 693.00, 601.16, 559.13, 337.09 | 559.13b, 541.11, 499.05 | 176 | [18] |
| 66 | Emodin bianthrone-hexose-(malonic acid)-hexose | 18.1 | 220, 278 | 919.21 | 875.23, 757.10, 713.20b, 671.25, 509.08, 458.00 | 713.15b, 509.04, 502.00, 457.99, 253.79 | 162 | [20,21] |
| 67 | Derivative of Emodin bianthrone-hexose-malonic acid | 18.2 | 220, 278 | 1005.23 | 961.13, 917.29, 757.10, 713.23b, 458.10 | [21] | ||
| 68 | Unknown | 18.3 | 225, 280, 325 | 811.36 | 793.32b, 763.38, 745.34, 669.23, 567.21, 389.09, 342.99, 311.93 | |||
| 69 | Unknown | 18.4 | 225, 280, 325 | 597.27 | 549.18b, 401.11, 357.12, 342.12, 194.87 | |||
| 70 | (diacetoxy-methoxyphenyl)acroyl-O-p-coumaroyl-O-caffeoylquinic acid | 18.7 | 214, 282, 325 | 777.26 | 735.24b, 717.25, 693.00, 601.16, 559.20, 337.04 | 559.13b, 541.17, 499.13 | 176 | [18] |
| 71 | Trimer lignin β-O-4-linked S unit with syringaresinol [S-(β-O-4′)-S-(β-β′)-S] | 19.0 | 220, 280 | 643.29 | 613.22, 417.13b, 387.15, 224.93, 194.87 | [22] | ||
| 72 | Tetramer lignin, S-(8-O-4′)-S-(8-O-4′)-S-(8-8′)-S | 19.1 | 220, 280 | 869.39 | 851.34b, 821.34, 697.27, 643.22, 595.21, 417.15, 387.15 | [22] | ||
| 73 | Emodin-O-(sulfonyl)-glucoside | 19.2 | 214, 280, | 511 | 430.99, 268.73b, 240.74, 224.96 | [11,20] | ||
| 74 | Lapathoside c | 19.5 | 220, 290, 315 | 809.28 | 663.13b, 485.07, 322.98 | 517.04, 485.10b, 322.88, 280.89 | 146 | [16,23] |
| 75 | (diacetoxy-methoxyphenyl)acroyl-O-p-coumaroyl-O-caffeoylquinic acid | 19.6 | 214, 282, 325 | 777.3 | 735.24b, 717.13, 693.13, 601.17, 559.00, 337.10 | 559.13b, 541.13, 499.00 | 176 | [18] |
| 76 | Lapathoside c isomer | 19.7 | 220, 290, 315 | 809.28 | 663.13b, 485.07, 322.98 | 517.04, 485.10b, 322.88, 280.89 | 146 | [16,23] |
| 77 | Unknown | 19.8 | 220, 280, 315 | 327.26 | 309.12, 291.10, 228.95b, 210.95, 170.91 | |||
| 78 | Emodin-8-O-(6’-O-malonyl)-glucoside * | 19.81 | 220, 282, 423 | 517.05 | 472.99b, 431.10 | |||
| 79 | Oligolignol-hedyotisol | 20.1 | 220, 280 | 809.36 | 791.33, 773.34, 761.25, 743.33b, 565.21, 417.11 | [24] | ||
| 80 | Tatariside e | 20.2 | 220, 290, 315 | 717.22 | 675.17, 571.09b, 529.10, 511.17, 487.09 | 529.05b, 511.04, 487.03 | 146 | [19] |
| 81 | Derivative of lignin-S(8–8)S | 20.5 | 220, 280 | 641.32 | 623.22, 611.20b, 417.13, 387.08, 347.09, 222.87 | [25] | ||
| 82 | Unknown | 20.7 | 220, 280, 315 | 1035.48 | 1017.45b, 999.38, 969.41, 821.41, 791.35, 595.14 | |||
| 83 | Tatariside a | 20.8 | 220, 290, 315 | 759.22 | 717.21b, 613.13, 571.13, 453.04, 288.94 | 571.09b, 553.10, 529.07, 511.06, 306.71 | 146 | [19] |
| 84 | Methyl derivative of Emodin bianthrone-hexose-(malonic acid)-hexose | 21.0 | 220, 278 | 933.21 | 889.37b, 727.21, 458.06 | [21] | ||
| 85 | Oligolignol-e.g., hedyotisol(isomer) | 21.1 | 220, 280 | 809.37 | 791.34b, 773.25, 761.31, 743.34, 565.21, 417.15 | [24] | ||
| 86 | Acetyl derivative of (diacetoxy-methoxyphenyl) acroyl-O-p-coumaroyl-O-caffeoylquinic acid | 21.2 | 214, 282, 325 | 819.29 | 777.29b, 759.25, 643.19, 601.14, 513.13 | 601.10b, 583.16, 559.07, 337.02 | 176 | [18] |
| 87 | Hydropiperoside * | 21.3 | 220, 290, 315 | 779.26 | 633.16b, 615.19, 487.13, 469.16, 453.09 | 487.12b, 469.16, 453.11, 307.10, 289.03 | 146 | |
| 88 | (3,6-O-di-p-coumaroyl)-β-fructofuranosyl-(2→1)-(2′-O-acetyl-6′-O-feruloyl)-β-glucopyranoside * | 21.5 | 220, 290, 315 | 851.25 | 809.23, 705.20b, 675.20, 527.07 | 663.22b, 645.38, 559.16, 527.16, 485.12 | 146 | |
| 89 | Derivative of Emodin bianthrone-di-hexose | 21.6 | 220, 278 | 1019.22 | 975.25, 931.42b, 889.25, 727.18, 458.06 | [21] | ||
| 90 | Acetyl derivative of (diacetoxy-methoxyphenyl) acroyl-O-p-coumaroyl-O-caffeoylquinic acid | 21.7 | 214, 282, 325 | 819.28 | 777.25b, 759.38, 643.18, 601.14, 513.13 | 601.18b, 583.18, 559.15, 541.11, 337.02 | 176 | [18] |
| 91 | Unknown | 21.9 | 220, 280, 315 | 329.27 | 311.18, 293.12, 228.95b, 210.96, 170.91 | |||
| 92 | Emodin bianthrone-hexose-(malonic acid)-hexose | 22.0 | 220, 278 | 919.2 | 875.24, 757.09, 713.20b, 671.13, 509.06, 458.00 | 713.18b, 508.96, 501.88, 458.03 | 162 | [20,21] |
| 93 | Oligolignol-e.g.,hedyotisol (isomer) | 22.1 | 220, 280 | 809.32 | 791.30, 773.25, 761.28, 743.29, 611.20b, 565.18, 417.19 | [24] | ||
| 94 | Phenylpropanoid-derived disaccharide ester | 22.2 | 220, 290, 315 | 987.31 | 969.39b, 957.50, 851.27, 823.32, 633.18, 453.09 | |||
| 95 | Tatariside a (isomer) | 22.5 | 220, 290, 315 | 759.4 | 717.22,675.16, 613.14b, 571.21, 529.18 | 571.09b, 553.05, 529.06, 511.06 | 146 | [21] |
| 96 | Emodin bianthrone-hexose-(malonic acid)-hexose | 23.0 | 220, 278 | 919.21 | 875.23, 757.10, 713.22b, 671.25, 509.09, 458.13 | 713.16b, 509.00, 501.75, 458.20 | 162 | [20,21] |
| 97 | Unknown | 23.01 | 220, 280, 315 | 837.37 | 819.31, 695.25, 640.23b, 579.18, 347.02 | |||
| 98 | Acetyl derivative of (diacetoxy-methoxyphenyl) acroyl-O-p-coumaroyl-O-caffeoylquinic acid | 23.1 | 220, 288, 325 | 819.26 | 777.28b, 759.38, 643.17, 601.25, 513.13, 361.01 | 601.13b, 583.13, 559.11, 336.97 | 176 | [18] |
| 99 | Acetyl derivative of (diacetoxy-methoxyphenyl) acroyl-O-p-coumaroyl-O-caffeoylquinic acid | 23.3 | 220, 288, 325 | 819.28 | 777.27b, 759.25, 643.17, 601.25, 513.13, 361.04 | 601.15b, 583.10, 559.11, 336.97 | 176 | [18] |
| 100 | Isomer of (3,6-O-di-p-coumaroyl)-β-fructofuranosyl-(2→1)-(2’-O-acetyl-6’-O-feruloyl)-β-glucopyranoside or tatariside d | 23.4 | 220, 290, 315 | 851.39 | 809.24, 705.19b, 663.27, 527.12 | 663.20b, 645.25, 559.13, 527.11, 485.10 | 146 | [19] |
| 101 | Hydropiperoside isomer | 23.45 | 220, 290, 315 | 779.36 | 633.11b, 615.25, 487.06, 469.13, 453.38, 288.86 | 487.06b, 469.18, 453.08, 306.90, 288.88 | 146 | |
| 102 | Methyl derivative of Emodin bianthrone-hexose-(malonic acid)-hexose | 23.5 | 220, 278 | 933.21 | 889.47b, 727.24, 458.09 | [21] | ||
| 103 | Vanicoside C * | 23.8 | 220, 290, 315 | 821.23 | 761.18, 675.16b, 633.19, 529.10, 487.09, 288.87 | 633.15, 529.10b, 453.18, 288.98 | 146 | |
| 104 | Acetyl derivative of (diacetoxy-methoxyphenyl)acroyl-O-p-coumaroyl-O-caffeoylquinic acid | 24.0 | 220, 290, 315 | 819.31 | 777.29b, 759.25, 643.17, 601.25, 583.20, 361.04 | 601.15b, 583.10, 559.11, 337.13 | 176 | [18] |
| 105 | Derivative of Emodin bianthrone-hexose-malonic acid | 24.1 | 220, 278 | 1005.22 | 961.13, 917.29, 757.12, 713.23b, 458.07 | [21] | ||
| 106 | Phenylpropanoid-derived disaccharide esters | 24.2 | 220, 290, 315 | 1181.4 | 1133.38, 1009.38, 955.50b, 809.41, 663.14 | |||
| 107 | Phenylpropanoid-derived disaccharide esters | 24.5 | 220, 290, 315 | 1151.38 | 1133.42, 1103.35, 1009.32, 955.40b, 809.29 | [23] | ||
| 108 | Phenylpropanoid-derived disaccharide esters | 24.6 | 220, 290, 315 | 1151.4 | 1133.38, 1103.38, 1009.33, 955.39b, 809.29 | [23] | ||
| 109 | Unknown | 24.7 | 220, 280, 315 | 623.28 | 591.21, 551.26, 486.13, 460.17b, 352.16, 297.07 | |||
| 110 | Tatariside b * | 25.0 | 220, 290, 315 | 893.27 | 851.24, 747.22b, 705.27, 687.33, 569.19 | 705.24b, 687.25, 663.22, 569.16, 527.18, 322.96 | 146 | |
| 111 | Methyl derivative of Emodin bianthrone-hexose-(malonic acid)-hexose | 25.1 | 220, 278 | 933.2 | 889.42b, 727.19, 685.20, 416.06 | [21] | ||
| 112 | Derivative of Emodin bianthrone-di-hexose | 25.4 | 220, 278 | 1019.24 | 975.25, 931.43b, 889.25, 727.20, 458.07 | [21] | ||
| 113 | Vanicoside B (isomer) | 25.6 | 220, 290, 315 | 955.37 | 809.26b, 663.19 | 663.26b, 485.20, 453.09 | 146 | |
| 114 | Unknown | 25.8 | 220, 280, 315 | 801.29 | 759.25b, 741.50, 655.19, 613.25, 571.13, 331.05 | 613.18b, 595.13, 571.15, 553.12, 330.95 | 146 | |
| 115 | Tatariside b (isomer) | 26.0 | 220, 290, 315 | 893.28 | 851.27, 747.21b, 705.29, 687.31, 569.18 | 705.26b, 687.37, 663.34, 569.23, 527.31, 322.98 | 146 | |
| 116 | Vanicoside B * | 26.4 | 220, 290, 315 | 955.29 | 809.22b, 663.20, 453.05 | 663.21b, 485.20, 323.05 | 146 | |
| 117 | Lapathoside a | 26.6 | 220, 290, 315 | 985.3 | 839.24b, 809.24, 663.22, 483.12 | 663.20b, 485.08, 322.85 | 176 | [16,23] |
| 118 | Diacetyl derivative of (diacetoxy-methoxyphenyl)acroyl-O-p-coumaroyl-O-caffeoylquinic acid | 26.7 | 220, 288, 325 | 861.3 | 819.29b, 801.25, 777.25, 759.25, 685.20, 643.17, 601.20, 583.18, 559.25, 513.17, 361.01 | 643.19b, 625.18, 601.15, 583.15 | 176 | [18] |
| 119 | Lapathoside b | 26.8 | 220, 290, 315 | 1015.31 | 869.23, 839.23b, 693.19, 663.22, 483.15 | 693.23, 663.20b, 645.28, 499.09, 322.89 | 176 | [26] |
| 120 | Questin * | 27.0 | 222, 286, 430 | 282.94 | 267.89, 239.85b | |||
| 121 | Phenylpropanoid-derived disaccharide esters | 27.1 | 220, 290, 315 | 1193.48 | 1175.45, 1145.50, 1051.38, 997.44b, 851.31, 821.30 | |||
| 122 | Phenylpropanoid-derived disaccharide esters | 27.2 | 220, 290, 315 | 1163.41 | 1145.45b, 1133.51, 999.37, 955.30, 851.15, 809.28 | |||
| 123 | Diacetyl derivative of (diacetoxy-methoxyphenyl) acroyl-O-p-coumaroyl-O-caffeoylquinic acid | 27.3 | 220, 288, 325 | 861.32 | 819.29b, 801.25, 777.25, 759.25, 685.20, 643.17, 601.20, 583.18, 559.25, 513.17, 361.01 | 643.17b, 625.18, 601.15, 583.15 | 176 | [18] |
| 124 | Vanicoside B (isomer) | 27.4 | 220, 290, 315 | 955.28 | 809.20b, 663.19, 453.04 | 663.23b, 485.20, 323.06 | 146 | |
| 125 | Dihydroferuloyl vanicoside B | 27.8 | 220, 290, 315 | 1133.4 | 1115.49b, 1103.65, 997.32, 969.37 | [16,23] | ||
| 126 | Unknown | 28.0 | 220, 290, 315 | 1071.38 | 1053.46b, 1041.64, 935.32, 907.40, 866.38, 717.11 | |||
| 127 | Diacetyl derivative of (diacetoxy-methoxyphenyl)acroyl-O-p-coumaroyl-O-caffeoylquinic acid | 28.1 | 220, 288, 325 | 861.32 | 819.29b, 801.25, 777.25, 759.25, 685.20, 643.17, 601.20, 583.18, 559.25, 513.17, 361.01 | 643.17b, 625.18, 601.15, 583.15 | 176 | [18] |
| 128 | Emodin bianthrone-hexose-malonic acid | 28.2 | 220, 278, 350 | 757.14 | 713.25b, 509.10, 458.12 | [21] | ||
| 129 | Dihydroferuloyl vanicoside B | 28.5 | 220, 290, 315 | 1133.38 | 1115.49b, 1103.50, 997.33, 969.38 | [16,23] | ||
| 130 | Derivative of Emodin bianthrone-di-hexose | 28.7 | 220, 278 | 1019.24 | 975.38, 931.43b, 889.25, 727.20, 458.07 | [21] | ||
| 131 | Vanicoside A * | 29.0 | 220, 290, 315 | 997.31 | 955.29, 851.24b, 821.28, 705.21, 453.05 | 809.24, 705.29b, 663.48, 527.22 | 146 | |
| 132 | Tatariside C | 29.1 | 220, 290, 315 | 935.27 | 893.27, 789.22b, 747.32, 705.29, 611.17, 569.18 | 747.26b, 705.23, 611.26, 569.22 | 146 | [19,27] |
| 133 | Hydropiperoside b | 29.2 | 220, 290, 315 | 1027.3 | 985.38, 967.30, 881.25b, 851.23, 705.20, 453.09 | 809.19, 705.20b, 663.20, 527.08, 453.06, 322.96 | 176 | [28] |
| 134 | Derivative of (diacetoxy-methoxyphenyl)acroyl-O-p-coumaroyl-O-caffeoylquinic acid | 29.4 | 220, 285, 325 | 965.36 | 923.31, 819.26, 789.29b, 747.22, 643.21 | 777.31b, 643.08, 611.15, 569.05, 361.06 | 146 | [18] |
| 135 | Derivative of (diacetoxy-methoxyphenyl) acroyl-O-p-coumaroyl-O-caffeoylquinic acid | 29.7 | 220, 285, 325 | 995.37 | 953.33, 819.23b, 777.25, 759.13, 611.24 | 777.23b, 735.18, 643.29, 611.16, 569.18 | 176 | [18] |
| 136 | Isomer vanicoside A/vanicoside F | 29.9 | 220, 290, 315 | 997.32 | 955.29, 851.24b, 821.28, 705.21, 453.06 | 809.22, 705.27b, 663.31, 527.20, 323.01 | 146 | |
| 137 | Phenylpropanoid-derived disaccharide esters | 30.3 | 220, 290, 315 | 1175.43 | 1157.52b, 1145.61, 1039.33, 1011.37 | |||
| 138 | Emodin bianthrone-hexose | 30.35 | 220, 278, 350 | 671.17 | 653.18, 509.09, 416.08b, 253.95 | 491.01, 253.88b | 162 | [21] |
| 139 | Unknown | 30.4 | 220, 265, 325 | 324.99b, 244.93 | 244.88 | |||
| 140 | Unknown | 30.7 | 220, 265, 325 | 1113.43 | 1095.45b, 1083.45, 977.29, 949.33 | |||
| 141 | Phenylpropanoid-derived disaccharide esters | 31.4 | 220, 290, 315 | 954.33 [M − 3H]3 | 881.20 [M − 2H]2, 809.20, 779.22b | 633.09b, 486.99 | 176 | [23] |
| 142 | Emodin bianthrone-hexose-malonic acid | 31.5 | 220, 278, 350 | 757.16 | 713.25b, 671.25, 509.10, 502.00, 458.12 | [21] | ||
| 143 | Vanicoside E | 32.1 | 220, 290, 315 | 1039.31 | 997.24, 893.25b, 747.30, 453.05 | 851.27, 747.28b, 705.40, 569.24, 304.91 | 146 | [27,28] |
| 144 | Emodin bianthrone-hexose-malonic acid | 32.7 | 220, 278, 350 | 757.16 | 713.21b, 671.19, 509.11, 502.00, 458.12 | [21] | ||
| 145 | Methyl derivative of Emodin bianthrone-hexose | 33.0 | 220, 278, 350 | 685.18 | 416.07b, 253.92 | [21] | ||
| 146 | Methyl derivative of Emodin bianthrone-hexose | 34.0 | 220, 278, 350 | 685.17 | 416.07b, 253.92 | [21] | ||
| 147 | Emodin * | 34.2 | 220, 248, 265, 288, 430 | 268.89 | 240.81, 224.93b, 181.68 | |||
| 148 | Methyl derivative of Emodin bianthrone-hexose-malonic acid | 34.6 | 220, 278, 350 | 771.14 | 727.22b, 502.05, 458.07 | [21] | ||
| 149 | Unknown | 35.0 | 220, 278, 350 | 721.41 | 675.39b, 397.10 | |||
| 150 | Methyl derivative of Emodin bianthrone-hexose-malonic acid | 35.2 | 220, 278, 350 | 771.15 | 727.24b, 502.05, 458.07 | [21] | ||
| 151 | Methyl derivative of Emodin bianthrone-hexose-malonic acid | 35.6 | 220, 278, 350 | 771.14 | 727.23b, 502.04, 458.08 | [21] | ||
| 152 | Methyl derivative of Emodin bianthrone-hexose-malonic acid | 36.3 | 220, 278, 350 | 771.15 | 727.23b, 502.04, 458.08 | [21] | ||
| 153 | Unknown | 37.5 | 225, 280, 325 | 647.37b, 1203.74 | 601.34b, 341.1 | |||
| 154 | Unknown | 37.9 | 225, 280, 325 | 723.42 | 677.40, 397.09 | |||
| 155 | Unknown | 38.3 | 220, 278, 350 | 369.18 | 351.12, 311.02, 292.99b, 210.79, 170.76 | |||
| 156 | Unknown | 38.4 | 225, 280, 325 | 559.35 | 513.28b, 277.15, 252.98 | |||
| 157 | Unknown | 39.4 | 225, 280, 325 | 559.36 | 513.29b, 277.16, 253.01 | |||
| 158 | Unknown | 40.7 | 225, 275 | 649.39 | 603.37 | |||
| 159 | Isovitexin/vitexin diglucoside | 41.0 | 269, 333 | 755.39 | 593.25, 575.29b, 477.06, 431.21 | 533.25, 503.21, 431.19b, 413.28 | 162 | [29,30] |
| 160 | Unknown | 41.5 | 220, 278, 360 | 725.45 | 679.43b, 397.09 | |||
| 161 | Emodin bianthrone | 41.6 | 220, 278, 360 | 509.14 | 491.08, 253.88b | [21] | ||
| 162 | Unknown | 42.1 | 225, 280, 325 | 295.19 | 277.08b, 194.94, 170.90 | |||
| 163 | Unknown | 42.7 | 225, 280, 325 | 561.59 | 515.32b, 279.20, 253.00 | |||
| 164 | Unknown | 42.8 | 225, 280, 325 | 625.39 | 579.36 | |||
| 165 | Emodin bianthrone isomer | 43.6 | 220, 278, 360 | 509.14 | 491.06, 253.88b | [21] | ||
| 166 | Unknown | 44.7 | 225, 280, 325 | 651.41 | 605.4 | |||
| 167 | Unknown | 45.2 | 220, 278, 350 | 757.4 | 595.30, 577.30, 477.05b, 433.22, 279.16 | 535.27, 505.24, 475.23, 433.22b, 279.13 | 162 | |
| 168 | Unknown | 47.2 | 225, 280, 325 | 563.39 | 517.34b, 281.21, 253.00 | |||
| 169 | Methyl derivative of emodin bianthrone | 50.4 | 220, 278, 360 | 523.18 | 253.89 | [21] | ||
| 170 | Alpha-carboxyethylhydroxychroman | 54.4 | 292 | 277.19 | 259.13, 233.06b | [31] | ||
| 171 | Unknown | 57.4 | 220, 278, 350 | 279.2 | 261.11b, 233.17 |
| Fraction | Radical Scavenging Activity DPPH (EC50 µg/mL) | Reducing Power AAE (%) 37 °C | Reducing Power AAE (%) 90 °C | LA-Peroxidation (IC50 µg/mL) | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| R.j | R.s | R.b | R.j | R.s | R.b | R.j | R.s | R.b | R.j | R.s | R.b | |
| Acetone | 9.6 ± 0.5 | 8.7 ± 0.4 | 12.6 ± 0.7 | 6.5 ± 0.3 | 6.0 ± 0.3 | 6.4 ± 0.2 | 28.5 ± 1.1 | 27.9 ± 1.0 | 21.4 ± 1.6 | 80.3 ± 2.8 | 71.6 ± 2.6 | 68.9 ± 1.6 |
| Dichloromethane | 202.1 ± 5.6 | 56.5 ± 3.9 | 63.3 ± 2.9 | 2.6 ± 0.2 | 1.8 ± 0.1 | 1.6 ± 0.06 | 11.2 ± 0.1 | 12.2 ± 0.6 | 10.8 ± 0.4 | 401.8 ± 12.7 | 112.2 ± 2.5 | 153.6 ± 6.0 |
| Diethyl ether | 9.3 ± 0.4 | 10.2 ± 0.8 | 8.8 ± 0.3 | 10.2 ± 0.5 | 8.3 ± 0.4 | 10.9 ± 0.4 | 35.0 ± 1.6 | 32.6 ± 1.2 | 35.4 ± 1.1 | 63.8 ± 2.6 | 67.3 ± 1.4 | 52.1 ± 2.6 |
| Ethyl acetate | 6.5 ± 0.4 | 4.7 ± 0.3 | 6.2 ± 0.1 | 13.9 ± 0.3 | 16.2 ± 0.2 | 16.6 ± 0.2 | 38.8 ± 1.3 | 44.7 ± 1.3 | 36.5 ± 1.7 | 45.7 ± 1.9 | 32.3 ± 1.7 | 40.6 ± 1.4 |
| Butanol | 9.1 ± 0.3 | 6.9 ± 0.2 | 8.1 ± 0.3 | 6.6 ± 0.2 | 8.2 ± 0.2 | 8.1 ± 0.2 | 29.0 ± 1.1 | 29.4 ± 0.8 | 25.7 ± 1.2 | 93.2 ± 3.5 | 66.2 ± 2.6 | 113.4 ± 4.2 |
| Water | 58.0 ± 2.5 | 35.0 ± 0.5 | 57.3 ± 2.3 | 0.6 ± 0.02 | 1.5 ± 0.05 | 0.1 ± 0.01 | 13.6 ± 0.4 | 16.9 ± 0.4 | 12.8 ± 0.2 | 650.7 ± 10.6 | 635.6 ± 17.8 | 690.1 ± 9.0 |
| Fraction | TPC Total Polyphenols [GAE] mg/g Fraction | Tannins Content [GAE] mg/g Fraction | ||||
|---|---|---|---|---|---|---|
| R.j | R.s | R.b | R.j | R.s | R.b | |
| Acetone | 324.1 ± 9.8 | 317.7 ± 14.1 | 487.7 ± 11.9 | 233.3 ± 6.4 | 264.0 ± 7.0 | 360.0 ± 6.5 |
| Dichloromethane | 96.4 ± 5.6 | 22.7 ± 0.9 | 81.1 ± 2.7 | 61.0 ± 2.9 | 13.0 ± 0.4 | 60.3 ± 2.7 |
| Diethyl ether | 469.1 ± 3.0 | 355.1 ± 17.1 | 615.4 ± 6.7 | 338.6 ± 17.2 | 241.6 ± 11.3 | 509.3 ± 19.8 |
| Ethyl acetate | 583.4 ± 6.5 | 640.7 ± 11.0 | 642.9 ± 8.9 | 484.3 ± 19.1 | 528.3 ± 16.9 | 510.5 ± 15.8 |
| Butanol | 307.1 ± 6.9 | 352.7 ± 7.0 | 286.1 ±6.0 | 258.0 ± 9.6 | 315.0 ± 7.4 | 243.0 ± 10.4 |
| Water | 28.7 ± 1.5 | 65.4 ± 4.5 | 29.7 ± 2.2 | 23.6 ± 1.1 | 46.6 ± 2.0 | 29.3 ± 0.6 |
| Variable | LA-Peroxidation EC50 | DPPH EC50 | Reducing Power AAE 37 °C | Reducing Power AAE 90 °C | Total Polyphenols | Tannins | DMACA | HCl-Butanol |
|---|---|---|---|---|---|---|---|---|
| LA-Peroxidation EC50 | 1000 | 0.751 | −0.904 | −0.874 | −0.823 | −0.804 | −0.938 | −0.300 |
| DPPH EC50 | 0.751 | 1000 | −0.843 | −0.869 | −0.663 | −0.742 | −0.757 | −0.736 |
| Reducing power AAE 37 °C | −0.904 | −0.843 | 1000 | 0.899 | 0.781 | 0.819 | 0.877 | 0.400 |
| Reducing power AAE 90 °C | −0.874 | −0.869 | 0.899 | 1000 | 0.795 | 0.810 | 0.917 | 0.411 |
| Total polyphenols | −0.823 | −0.663 | 0.781 | 0.795 | 1000 | 0.939 | 0.779 | 0.259 |
| Tannins | −0.804 | −0.742 | 0.819 | 0.810 | 0.939 | 1000 | 0.738 | 0.378 |
| DMACA | −0.938 | −0.757 | 0.877 | 0.917 | 0.779 | 0.738 | 1000 | 0.272 |
| HCL-Butanol | −0.300 | −0.736 | 0.400 | 0.411 | 0.259 | 0.378 | 0.272 | 1000 |
| Nr. | Identification | EC50linoleic | EC50 DPPH | AAE 37 | AAE 90 |
|---|---|---|---|---|---|
| 9 | Procyanidin dimer | 0.563 | 0.552 | 0.458 | 0.484 |
| 10 | Procyanidin trimer | 0.63 | 0.68 | 0.62 | 0.572 |
| 11 | Catechin | 0.611 | 0.305 | 0.373 | 0.502 |
| 12 | Procyanidin trimer monogallate | 0.635 | 0.646 | 0.665 | 0.601 |
| 13 | Procyanidin dimer | 0.554 | 0.536 | 0.664 | 0.645 |
| 15 | Procyanidin trimer | 0.555 | 0.571 | 0.536 | 0.527 |
| 17 | Procyanidin dimer monogallate | 0.763 | 0.655 | 0.762 | 0.795 |
| 18 | Procyanidin trimer monogallate | 0.494 | 0.504 | 0.446 | 0.445 |
| 20 | Piceatannol glucoside | 0.432 | 0.389 | 0.588 | 0.446 |
| 21 | Procyanidin trimer | 0.48 | 0.512 | 0.6 | 0.501 |
| 22 | Resveratrolside | 0.342 | 0.353 | 0.499 | 0.491 |
| 23 | Procyanidin trimer monogallate | 0.781 | 0.697 | 0.806 | 0.783 |
| 24 | Procyanidin dimer monogallate | 0.687 | 0.684 | 0.758 | 0.734 |
| 25 | Procyanidin tetramer | 0.481 | 0.526 | 0.608 | 0.518 |
| 26 | Procyanidin pentamer | 0.35 | 0.438 | 0.584 | 0.387 |
| 27 | Piceid | 0.34 | 0.319 | 0.48 | 0.466 |
| 28 | Procyanidin trimer digallate | 0.592 | 0.598 | 0.717 | 0.585 |
| 29 | Procyanidin dimer digallate | 0.477 | 0.414 | 0.592 | 0.583 |
| 30 | Procyanidin trimer monogallate | 0.494 | 0.504 | 0.446 | 0.445 |
| 35 | Procyanidin trimer monogallate | 0.746 | 0.719 | 0.764 | 0.721 |
| 37 | Procyanidin tetramer monogallate | 0.682 | 0.701 | 0.729 | 0.643 |
| 39 | Procyanidin gallate | 0.666 | 0.669 | 0.724 | 0.618 |
| 40 | Procyanidin trimer monogallate | 0.716 | 0.561 | 0.753 | 0.636 |
| 78 | Emodin-8-O-(6’-O-malonyl)-glucoside | 0.37 | 0.349 | 0.496 | 0.316 |
| 87 | Hydropiperoside | 0.541 | 0.212 | 0.264 | 0.395 |
| 106 | Phenylpropanoid-derived disaccharide esters | 0.659 | 0.391 | 0.424 | 0.509 |
| 107 | Phenylpropanoid-derived disaccharide esters | 0.511 | 0.366 | 0.458 | 0.561 |
| 108 | Phenylpropanoid-derived disaccharide esters | 0.704 | 0.477 | 0.631 | 0.719 |
| 113 | Vanicoside B (isomer) | 0.501 | 0.166 | 0.198 | 0.338 |
| 116 | Vanicoside B | 0.618 | 0.315 | 0.349 | 0.473 |
| 117 | Lapathoside a | 0.537 | 0.209 | 0.263 | 0.394 |
| 121 | Phenylpropanoid-derived disaccharide esters | 0.579 | 0.41 | 0.407 | 0.511 |
| 122 | Phenylpropanoid-derived disaccharide esters | 0.556 | 0.289 | 0.358 | 0.447 |
| 124 | Vanicoside B (isomer) | 0.564 | 0.217 | 0.284 | 0.403 |
| 125 | Dihydroferuloylvanicoside B | 0.624 | 0.341 | 0.39 | 0.54 |
| 141 | Phenylpropanoid-derived disaccharide esters | 0.494 | 0.504 | 0.446 | 0.445 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Nawrot-Hadzik, I.; Ślusarczyk, S.; Granica, S.; Hadzik, J.; Matkowski, A. Phytochemical Diversity in Rhizomes of Three Reynoutria Species and their Antioxidant Activity Correlations Elucidated by LC-ESI-MS/MS Analysis. Molecules 2019, 24, 1136. https://doi.org/10.3390/molecules24061136
Nawrot-Hadzik I, Ślusarczyk S, Granica S, Hadzik J, Matkowski A. Phytochemical Diversity in Rhizomes of Three Reynoutria Species and their Antioxidant Activity Correlations Elucidated by LC-ESI-MS/MS Analysis. Molecules. 2019; 24(6):1136. https://doi.org/10.3390/molecules24061136
Chicago/Turabian StyleNawrot-Hadzik, Izabela, Sylwester Ślusarczyk, Sebastian Granica, Jakub Hadzik, and Adam Matkowski. 2019. "Phytochemical Diversity in Rhizomes of Three Reynoutria Species and their Antioxidant Activity Correlations Elucidated by LC-ESI-MS/MS Analysis" Molecules 24, no. 6: 1136. https://doi.org/10.3390/molecules24061136
APA StyleNawrot-Hadzik, I., Ślusarczyk, S., Granica, S., Hadzik, J., & Matkowski, A. (2019). Phytochemical Diversity in Rhizomes of Three Reynoutria Species and their Antioxidant Activity Correlations Elucidated by LC-ESI-MS/MS Analysis. Molecules, 24(6), 1136. https://doi.org/10.3390/molecules24061136









