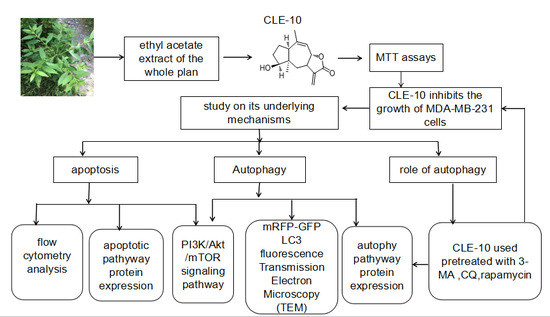
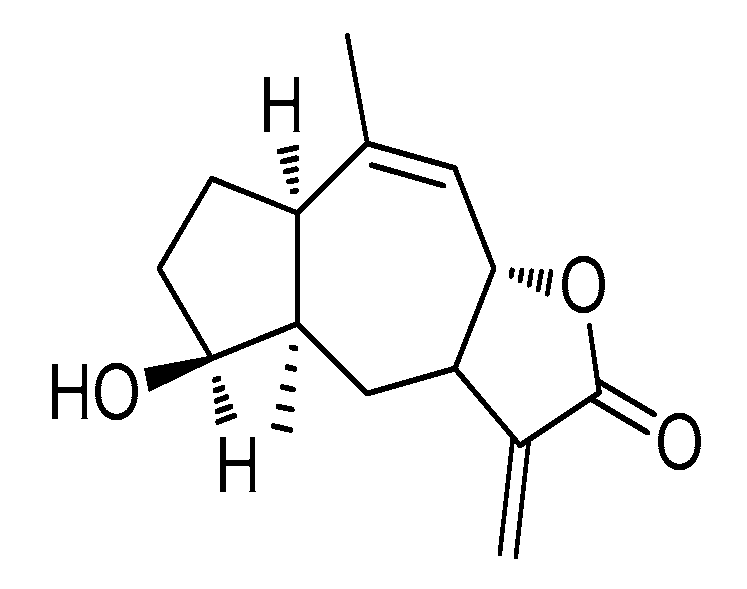
CLE-10 from Carpesium abrotanoides L. Suppresses the Growth of Human Breast Cancer Cells (MDA-MB-231) In Vitro by Inducing Apoptosis and Pro-Death Autophagy Via the PI3K/Akt/mTOR Signaling Pathway
Abstract
1. Introduction
2. Results
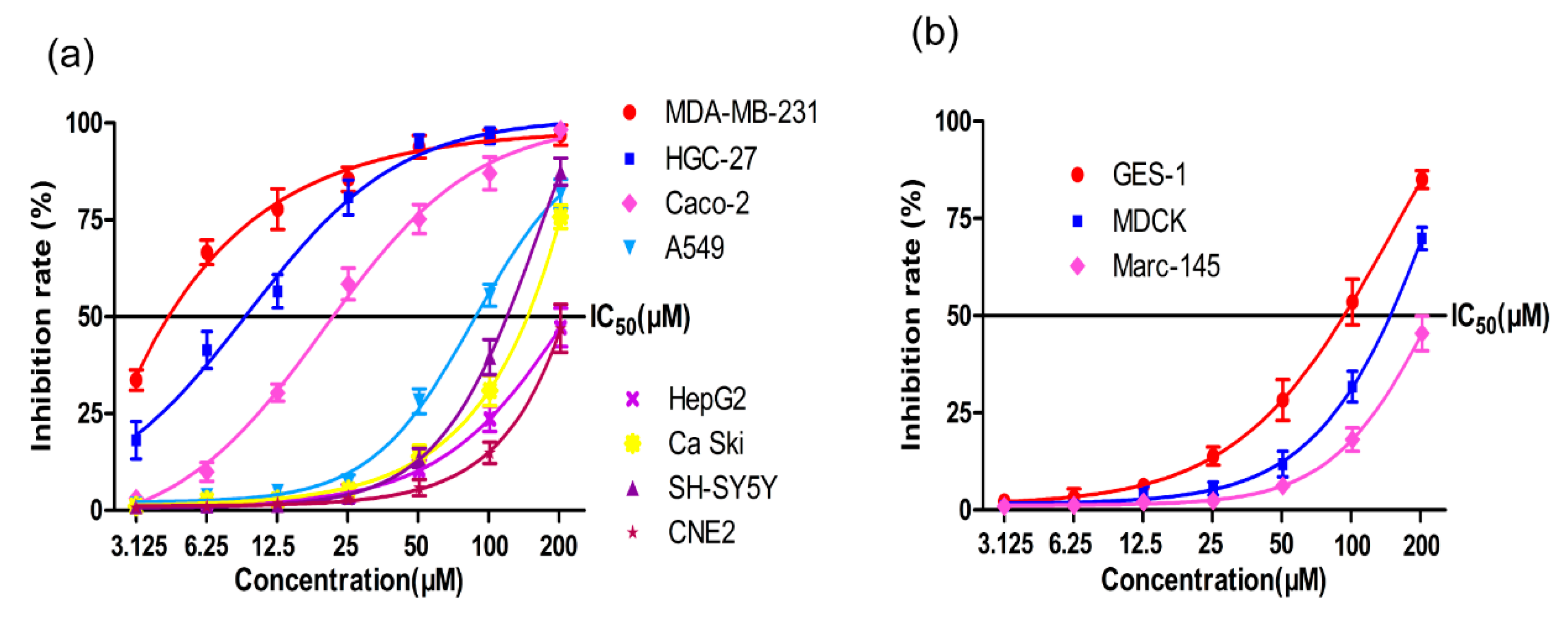
2.1. MTT Assay
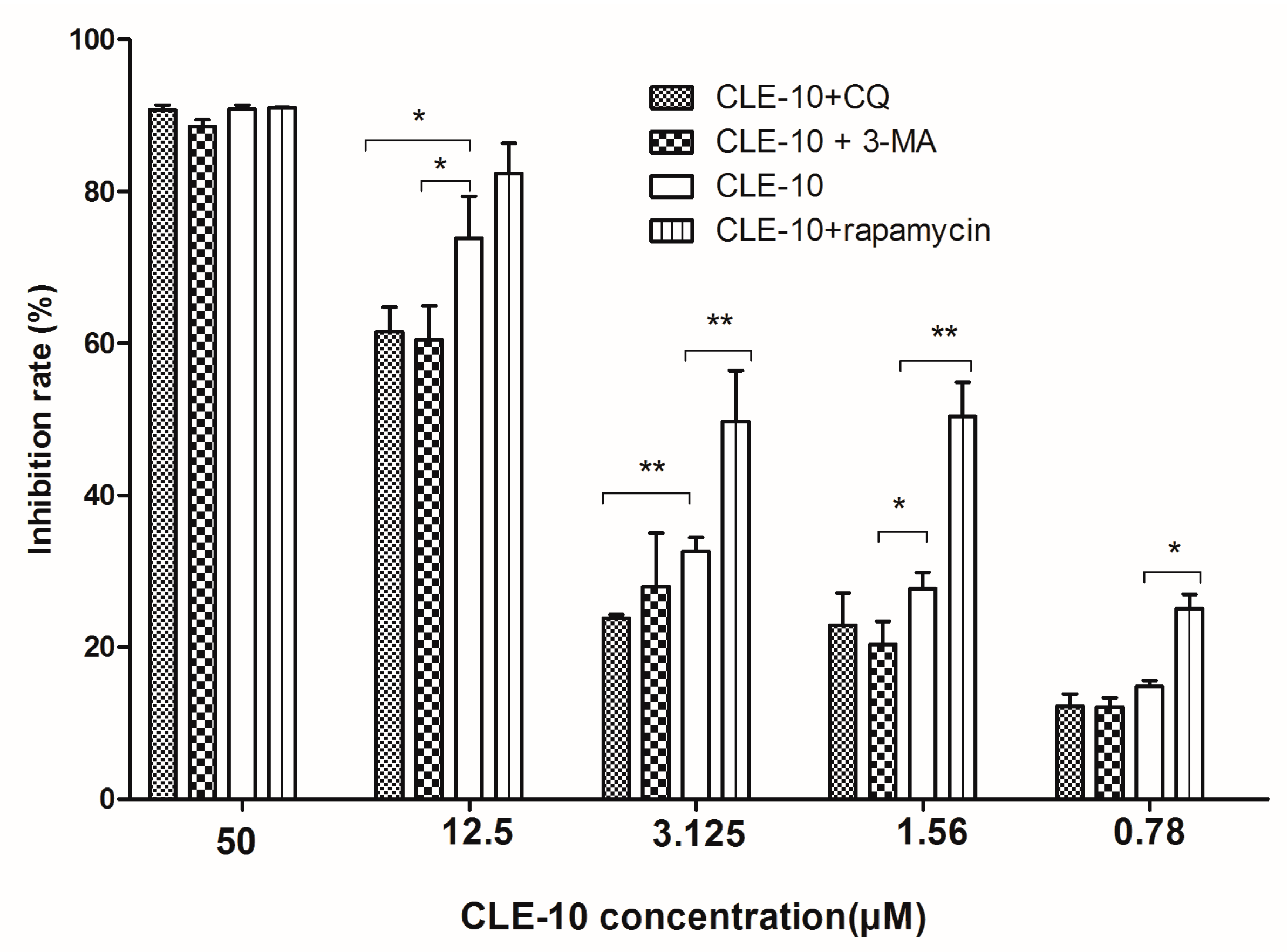
2.2. Inhibition of Autophagy Relieved CLE-10-Induced Cell Death
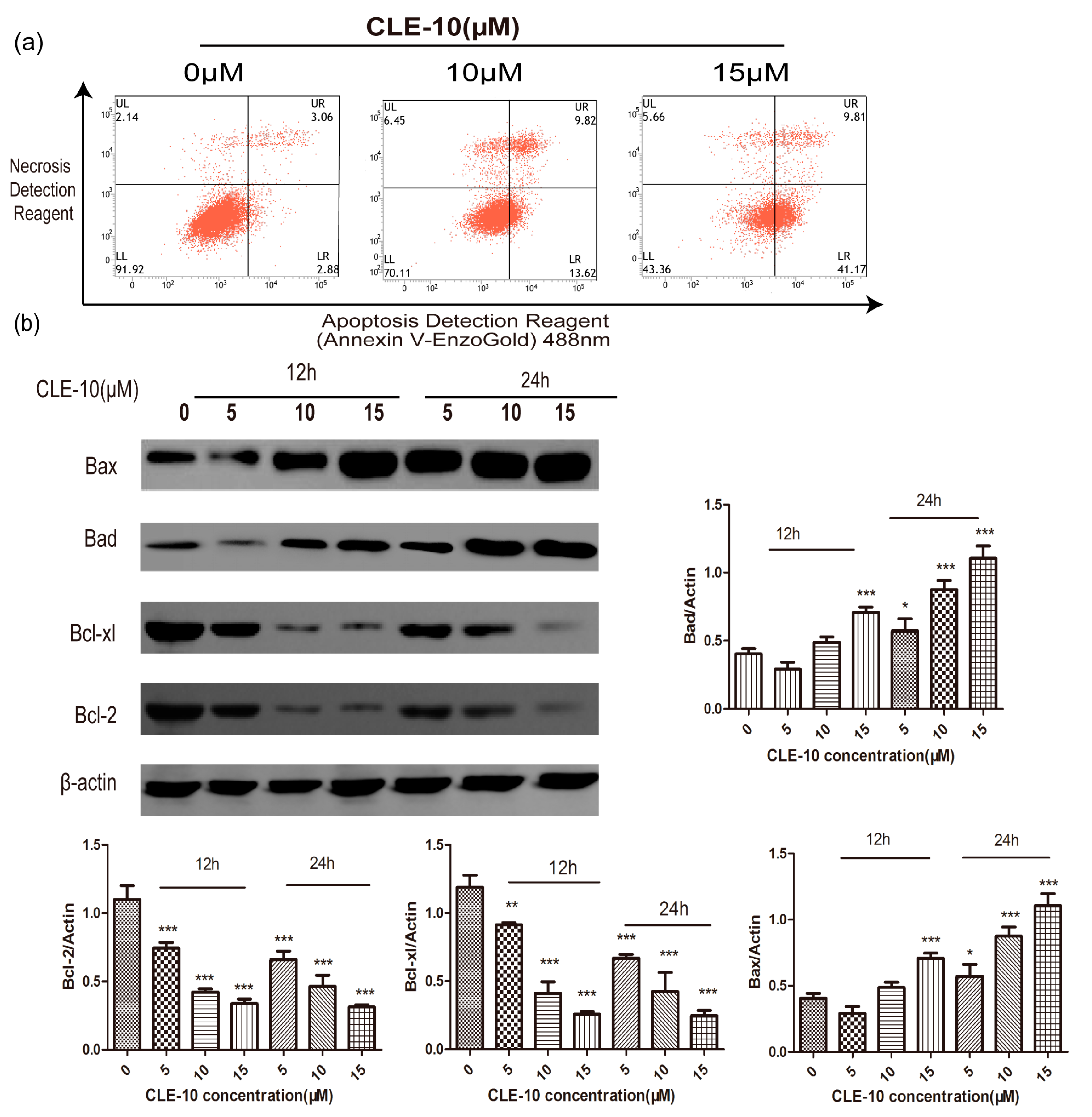
2.3. CLE-10 Induced MDA-MB-231 Cell Apoptosis
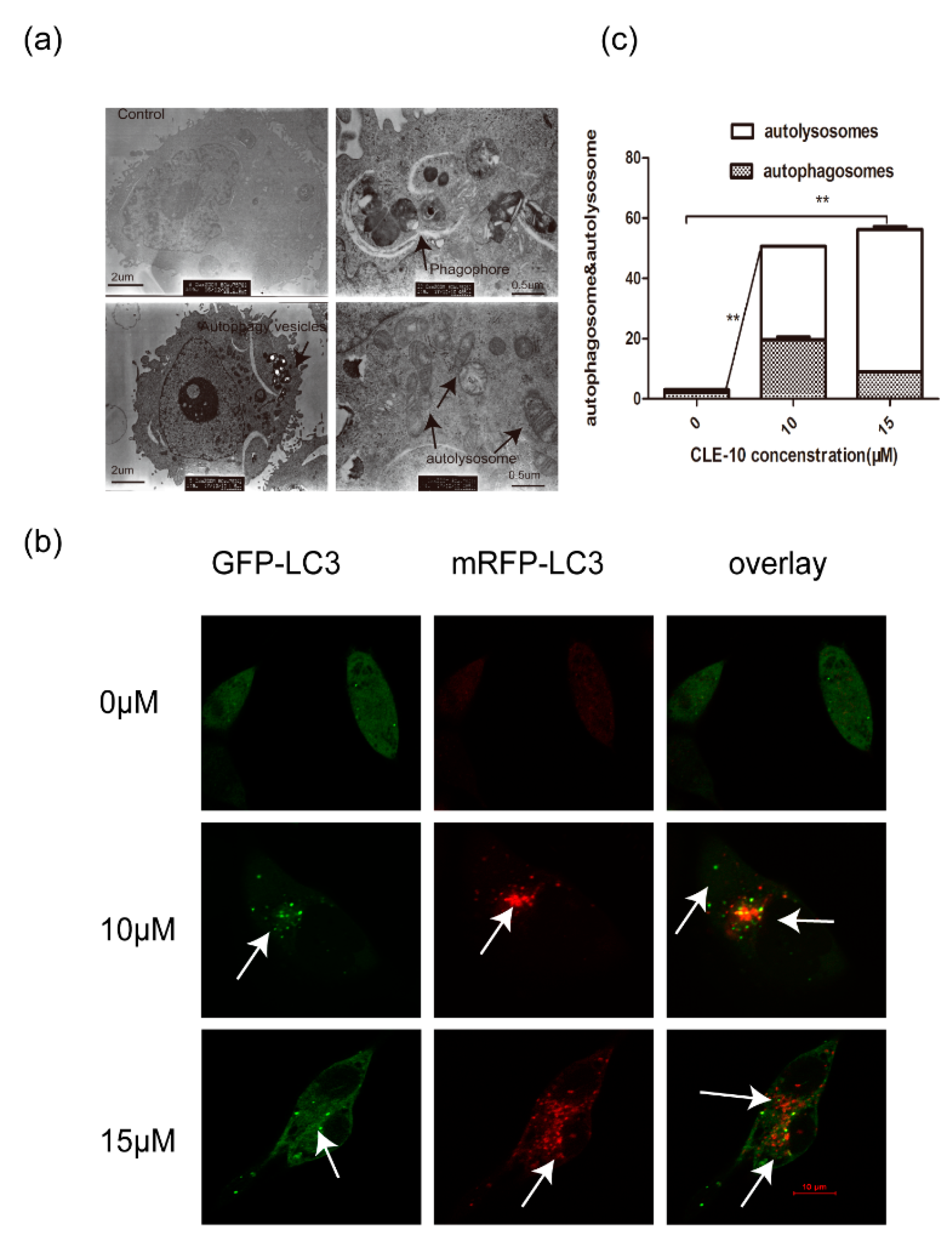
2.4. CLE-10 Induced MDA-MB-231 Cell Autophagy
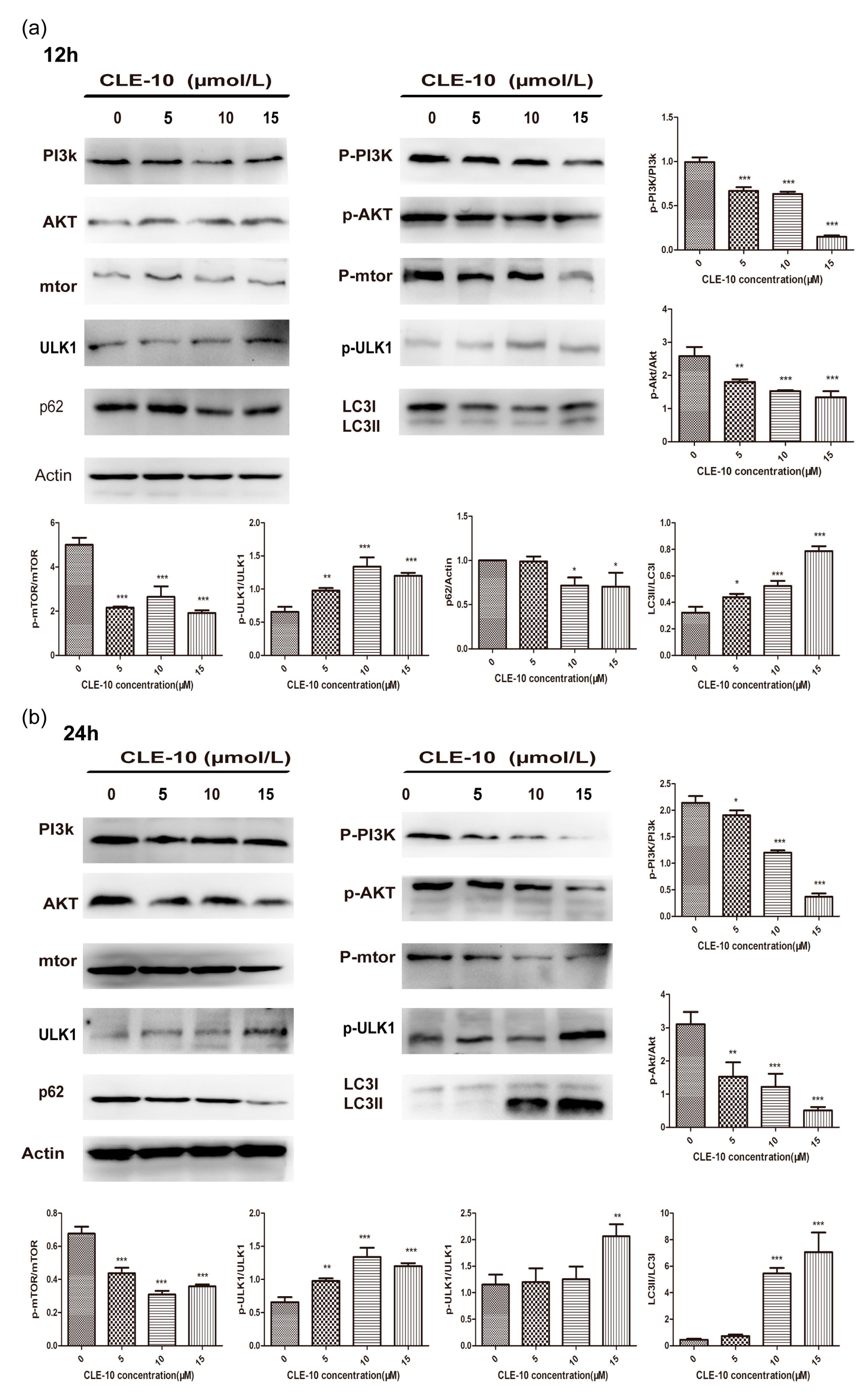
2.5. CLE-10 Inhibited the PI3K/Akt/mTOR Signaling Pathway in MDA-MB-231 Cells
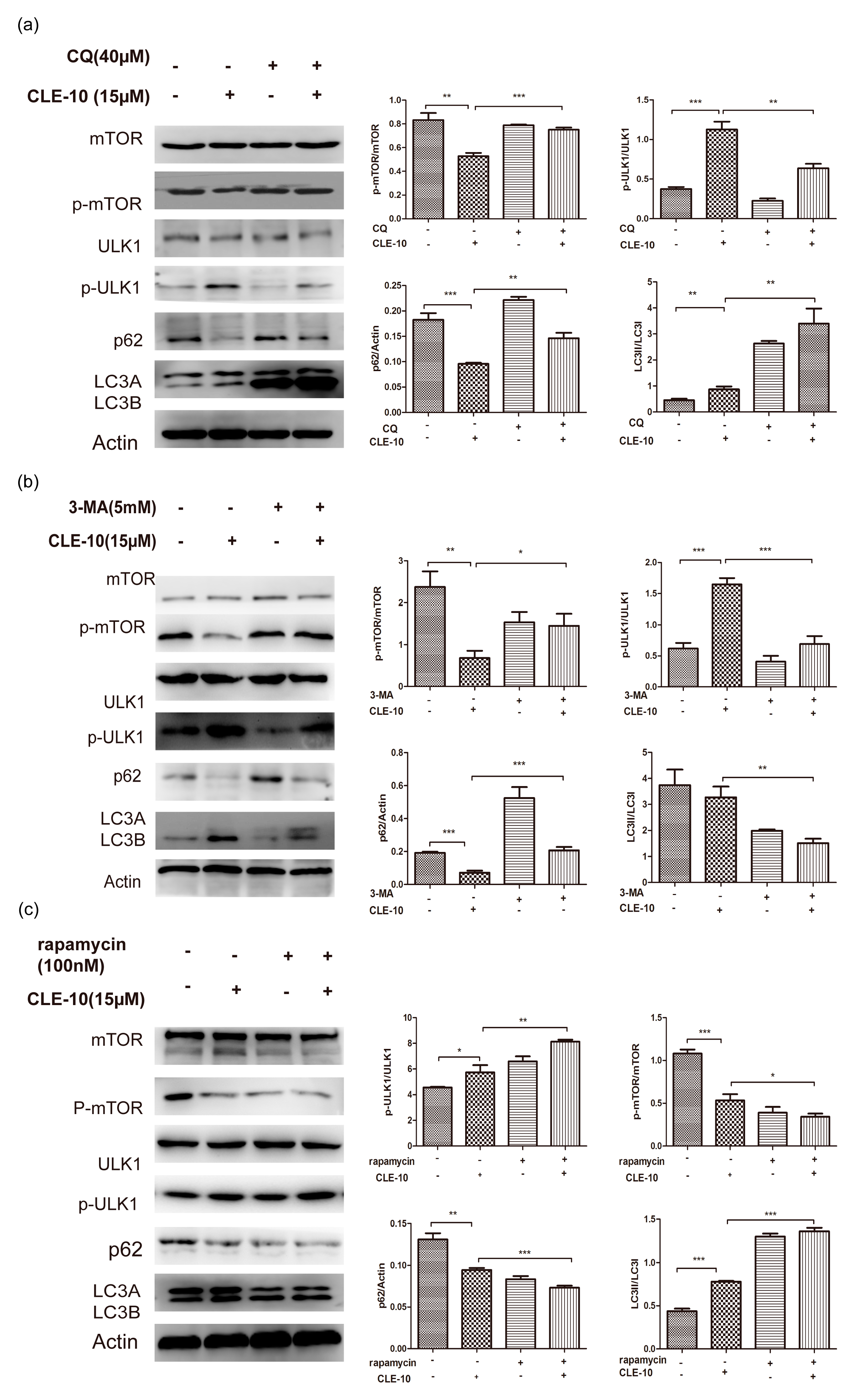
2.6. The Effect of CLE-10 Combined with 3-MA, CQ, or Rapamycin on the Protein Expression of mTOR, ULK1, p62, and LC3
3. Discussion
4. Material and Methods
4.1. Materials
4.2. Cell Lines and Cultures
4.3. MTT Assays
4.4. CLE-10 Pretreated with 3-MA, CQ, Rapamycin Using MTT Assays
4.5. Flow Cytometry (FCM)
4.6. Western Blot Analysis
4.7. Transmission Electron Microscopy (TEM)
4.8. Tandem mRFP-GFP-LC3 Transfection
4.9. Statistical Analysis
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Globocan, 2012, Breast Cancer Fact Sheet 2008. Available online: http://globocan.iarc.fr/factsheets/ (accessed on 19 March 2019).
- Jafari, S.H.; Saadatpour, Z.; Salmaninejad, A.; Momeni, F.; Mokhtari, M.; Nahand, J.S.; Rahmati, M.; Mirzaei, H.; Kianmehr, M. Breast cancer diagnosis: Imaging techniques and biochemical markers. J. Cell. Physiol. 2018, 233, 5200–5213. [Google Scholar] [CrossRef]
- Wang, M.; Wang, Y.; Zhong, J. Side population cells and drug resistance in breast cancer. Mol. Med. Rep. 2015, 11, 4297–4302. [Google Scholar] [CrossRef] [PubMed]
- Kiliçarslan, A.; Kahraman, A.; Akkiz, H.; Yildiz Menziletoğlu, S.; Fingas, C.D.; Gerken, G.; Canbay, A. Apoptosis in selected liver diseases. Turk. J. Gastroenterol. 2009, 20, 171–179. [Google Scholar] [CrossRef]
- Tanida, I. Autophagosome formation and molecular mechanism of autophagy. Antioxid. Redox Signal. 2011, 14, 2201–2214. [Google Scholar] [CrossRef] [PubMed]
- Mizushima, N.; Levine, B.; Cuervo, A.M.; Klionsky, D.J. Autophagy fights disease through cellular self-digestion. Nature 2011, 451, 1069–1075. [Google Scholar] [CrossRef]
- Denton, D.; Nicolson, S.; Kumar, S. Cell death by autophagy: Facts and apparent artefacts. Cell Death Differ. 2012, 19, 87–95. [Google Scholar] [CrossRef]
- Huang, R.; Liu, W. Identifying an essential role of nuclear LC3 for autophagy. Autophagy 2015, 11, 852–853. [Google Scholar] [CrossRef] [PubMed]
- Komatsu, M. Potential role of p62 in tumor development. Autophagy 2011, 7, 1088–1090. [Google Scholar] [CrossRef]
- Islam, M.A.; Sooro, M.A.; Zhang, P. Autophagic regulation of p62 is critical for cancer therapy. Int. J. Mol. Sci. 2018, 19, 1405. [Google Scholar] [CrossRef]
- Bahrami, A.; Khazaei, M.; Shahidsales, S.; Hassanian, S.M.; Hasanzadeh, M.; Maftouh, M.; Ferns, G.A.; Avan, A. The therapeutic potential of PI3k/Akt/mTOR inhibitors in breast cancer: Rational and progress. J. Cell Biochem. 2017, 119, 213–222. [Google Scholar] [CrossRef]
- Hosokawa, N.; Hara, T.; Kaizuka, T.; Kishi, C.; Takamura, A.; Miura, Y.; Lemura, S.I.; Natsume, T.; Takehana, K.; Yamada, N.; et al. Nutrient-dependent mTORC1 association with the ULK1–Atg13–FIP200 complex required for autophagy. Mol. Biol. Cell 2009, 20, 1981–1991. [Google Scholar] [CrossRef]
- Inoki, K.; Li, Y.; Zhu, T.Q.; Wu, J.; Guan, K.L. TSC2 is phosphorylated and inhibited by Akt and suppresses mTOR signaling. Nat. Cell Biol. 2002, 4, 648–657. [Google Scholar] [CrossRef]
- Shang, L.B.; Wang, X.D. AMPK and mTOR coordinate the regulation of ULK1 and mammalian autophagy initiation. Autophagy 2011, 7, 924–926. [Google Scholar] [CrossRef]
- Zhang, J.P.; Wang, G.W.; Tian, X.H.; Yang, Y.X.; Liu, Q.X.; Chen, L.P.; Li, H.L.; Zhang, W.D. The genus Carpesium: A review of its ethnopharmacology phytochemistry and pharmacology. J. Ethnopharmacol. 2015, 163, 173–191. [Google Scholar] [CrossRef]
- Feng, J.T.; Hao, W.; Ren, S.X.; He, J.; Yong, L.; Xing, Z. Synthesis and anti-fungal activities of carabrol ester derivatives. J. Agric. Food Chem. 2012, 60, 3817–3823. [Google Scholar] [CrossRef]
- Wang, J.F.; He, W.J.; Zhang, X.X.; Zhao, B.Q.; Liu, Y.H.; Zhou, X.J. Dicarabrol, a new dimeric sesquiterpene from Carpesium abrotanoides L. Bioorg. Med. Chem. Lett. 2015, 25, 4082–4084. [Google Scholar] [CrossRef]
- Fischedick, J.T.; Pesic, M.; Podolski-Renic, A.; Bankovic, J.; de Vos, R.C.H.; Peric, M.; Todorovic, S.; Tanic, N. Cytotoxic activity of sesquiterpene lactones from Inula britannica on human cancer cell lines. Phytochem. Lett. 2013, 6, 246–252. [Google Scholar] [CrossRef]
- Li, X.W.; Weng, L.; Gao, X.; Zhao, Y.; Pang, F.; Liu, J.H.; Zhang, H.F.; Hu, J.F. Antiproliferative and apoptotic sesquiterpene lactones from carpesium faberi. Bioorg. Med. Chem. Lett. 2011, 21, 366–372. [Google Scholar] [CrossRef]
- Wang, L.; Qin, W.; Tian, L.; Zhang, X.X.; Lin, F.; Cheng, F.; Chen, J.F.; Liu, C.X.; Guo, Z.Y.; Peter, P.; et al. Caroguaianolide A-E, five new cytotoxic sesquiterpene lactones from Carpesium abrotanoides L. Fitoterapia 2018, 127, 349–355. [Google Scholar] [CrossRef]
- Park, J.H.; Kim, K.P.; Ko, J.J.; Park, K.S. PI3K/Akt/mTOR activation by suppression of ELK3 mediates chemosensitivity of MDA-MB-231 cells to doxorubicin by inhibiting autophagy. Biochem. Biophys. Res. Commun. 2016, 477, 277–282. [Google Scholar] [CrossRef]
- Youle, R.J.; Strasser, A. The Bcl-2 protein family: Opposing activities that mediate cell death. Nat. Rev. Mol. Cell Biol. 2008, 9, 47–59. [Google Scholar] [CrossRef]
- Shen, H.M.; Codogno, P. Autophagic cell death: Loch Ness monster or endangered species? Autophagy 2011, 7, 457–465. [Google Scholar] [CrossRef]
- Munz, C. Enhancing immunity through autophagy. Annu. Rev. Immunol. 2009, 27, 423–449. [Google Scholar] [CrossRef]
- Levine, B.; Kroemer, G. Autophagy in the pathogenesis of disease. Cell 2008, 132, 27–42. [Google Scholar] [CrossRef]
- Chen, Y.; Azad, M.B.; Gibson, S.B. Methods for detecting autophagy and determining autophagy-induced cell death. Can. J. Physiol. Pharmacol. 2010, 88, 285–295. [Google Scholar] [CrossRef]
- Fu, H.; Wang, C.; Yang, D.; Zhang, X.; Wei, Z.; Xu, J.; Hu, Z.; Zhang, Y.; Wang, W.; Yan, R.; et al. Curcumin regulates proliferation, autophagy and apoptosis in gastric cancer cells by affecting PI3k and p53 signaling. J. Cell Physiol. 2018, 233, 4634–4642. [Google Scholar] [CrossRef]
- Kumar, D.; Das, B.; Sen, R.; Kundu, P.; Manna, A.; Sarkar, A.; Chowdhury, C.; Chatterjee, M.; Das, P. Andrographolide analogue induces apoptosis and autophagy mediated cell death in U937 cells by inhibition of PI3k/Akt/mTOR pathway. PLoS ONE 2015, 10, e0139657. [Google Scholar] [CrossRef]
- Kenney, D.L.; Benarroch, E.E. The autophagy-lysosomal pathway: General concepts and clinical implications. Neurology 2015, 85, 634–645. [Google Scholar] [CrossRef]
- Lee, Y.K.; Lee, J.A. Role of the mammalian Atg8/LC3 family in autophagy: Differential and compensatory roles in the spatiotemporal regulation of autophagy. Bmb. Rep. 2016, 49, 424–430. [Google Scholar] [CrossRef]
- Tanida, I.; Minematsu-Ikeguchi, N.; Ueno, T.; Kominami, E. Lysosomal turnover, but not a cellular level of endogenous LC3 is a marker for autophagy. Autophagy 2005, 1, 84–91. [Google Scholar] [CrossRef]
- Wei, J.L.; Lin, Y.; Wei, F.H.; Lin, J.G.; Zi, G.X.; Hong, L.W.; Chen, Y.; Liu, H.F. P62 links the autophagy pathway and the ubiqutin–proteasome system upon ubiquitinated protein degradation. Cell Mol. Biol. Lett. 2016, 21, 29. [Google Scholar] [CrossRef]
- Su, H.; Wang, X. Autophagy and p62 in cardiac protein quality control. Autophagy 2011, 7, 1382–1383. [Google Scholar] [CrossRef]
Sample Availability: Samples of the compounds are available from the authors. |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tian, L.; Cheng, F.; Wang, L.; Qin, W.; Zou, K.; Chen, J. CLE-10 from Carpesium abrotanoides L. Suppresses the Growth of Human Breast Cancer Cells (MDA-MB-231) In Vitro by Inducing Apoptosis and Pro-Death Autophagy Via the PI3K/Akt/mTOR Signaling Pathway. Molecules 2019, 24, 1091. https://doi.org/10.3390/molecules24061091
Tian L, Cheng F, Wang L, Qin W, Zou K, Chen J. CLE-10 from Carpesium abrotanoides L. Suppresses the Growth of Human Breast Cancer Cells (MDA-MB-231) In Vitro by Inducing Apoptosis and Pro-Death Autophagy Via the PI3K/Akt/mTOR Signaling Pathway. Molecules. 2019; 24(6):1091. https://doi.org/10.3390/molecules24061091
Chicago/Turabian StyleTian, Li, Fan Cheng, Lei Wang, Wen Qin, Kun Zou, and Jianfeng Chen. 2019. "CLE-10 from Carpesium abrotanoides L. Suppresses the Growth of Human Breast Cancer Cells (MDA-MB-231) In Vitro by Inducing Apoptosis and Pro-Death Autophagy Via the PI3K/Akt/mTOR Signaling Pathway" Molecules 24, no. 6: 1091. https://doi.org/10.3390/molecules24061091
APA StyleTian, L., Cheng, F., Wang, L., Qin, W., Zou, K., & Chen, J. (2019). CLE-10 from Carpesium abrotanoides L. Suppresses the Growth of Human Breast Cancer Cells (MDA-MB-231) In Vitro by Inducing Apoptosis and Pro-Death Autophagy Via the PI3K/Akt/mTOR Signaling Pathway. Molecules, 24(6), 1091. https://doi.org/10.3390/molecules24061091